Parallel and Hierarchical Mode Association Clustering with an R Package Modalclust
|
|
|
- Alexander McCormick
- 6 years ago
- Views:
Transcription
1 Parallel and Hierarchical Mode Association Clustering with an R Package Modalclust Surajit Ray and Yansong Cheng Department of Mathematics and Statistics Boston University 111 Cummington Street, Boston, MA 02215, USA Abstract Modalclust is an R package which performs Hierarchical Mode Association Clustering (HMAC) along with its parallel implementation (PHMAC) over several processors. Modal clustering techniques are especially designed to efficiently extract clusters in high dimensions with arbitrary density shapes. Further, clustering is performed over several resolutions and the results are summarized as a hierarchical tree, thus providing a model based multi resolution cluster analysis. Finally we implement a novel parallel implementation of HMAC which performs the clustering job over several processors thereby dramatically increasing the speed of clustering procedure especially for large data sets. This package also provides a number of functions for visualizing clusters in high dimensions, which can also be used with other clustering softwares. Keywords: Hierarchical mode association clustering, Modal clustering, Modal EM, Nonparametric density, Parallel computing, R, Clustering of Complex Data, Robust analysis. 1. Introduction Cluster analysis is a ubiquitous technique in statistical analysis that has been widely used in multiple disciplines for many years. Historically cluster analysis techniques have been approached from either a fully parametric view, e.g. mixture model based clustering, or a distribution free approach, e.g. linkage based hierarchical clustering. While the parametric paradigm provides the inferential framework and accounts for the sampling variability, it often lacks the flexibility to accommodate complex clusters and are often not scalable to high dimensional data. On the other hand, the distribution free approaches are usually fast and capable of uncovering complex clusters by making use of different distance measures. However, the inferential framework is distinctly Preprint submitted to Computational Statistics & Data Analysis June 2, 2011
2 missing in the distribution free clustering techniques. Accordingly most clustering packages in R (R Development Core Team, 2010) also fall under the two above mentioned groups of clustering techniques. This paper describes a software program for cluster analysis that can knead the strengths of these two seemingly different approaches and develop a framework of parallel implementation for clustering techniques. For most model based approaches to clustering the following limitations are well recognized in the literature: (i) the number of clusters has to be specified (ii) the mixing densities have to be specified, and as estimating the parameters of the mixture models is often computationally very expensive, we are often forced to limit our choices to simple distributions such as Gaussian; (iii) computational speed is inadequate especially in high dimensions and this coupled with the complexity of the proposed model often limits the use of model-based techniques either theoretically or computationally; (v) it is not straightforward to extend model-based clustering to uncover heterogeneity at multiple resolutions, similar to the one offered by to the model free linkage based hierarchical clustering. Influential work towards resolving the first three issues has been carried out in Fraley and Raftery (1998), Fraley (1998), Fraley and Raftery (1999, 2002b,a) Ray and Lindsay (2005) and Ray and Lindsay (2008). Many previous approaches have focused on model selection either for choosing the number of components or for determining the covariance structure of the density under consideration or both. They work efficiently if the underlying distribution is chosen correctly, but none of these model based approaches is designed to handle a completely arbitrary underlying distribution (see Figure 5 for one such example). That is, we think that limitations due to issues (iii) and (iv), above often necessitates the use of model-free techniques. The hierarchical mode association clustering HMAC (Li et al., 2007), which is constructed by first determining modes of the high-dimensional density and then associating sample points to those modes, is the first multivariate model based clustering approach resolving many of the drawbacks of standard model-based clustering. Specifically, it can accommodate flexible subpopulation structures at multiple resolutions while retaining the desired natural inferential framework of parametric mixtures. Modalclust is the package implemented in R (R Development Core Team, 2010) for carrying out HMAC along with its parallel implementation (PHMAC) over several processors. Though mode-counting or mode hunting has been extensively used as a clustering technique (see 2
3 Silverman, 1981, 1983, Hartigan and Hartigan, 1985, Cheng and Hall, 1998, 1999, Hartigan, 1988, and references therein), most implementation are limited to univariate data. Generalization to higher dimensions was limited both due to the computational complexity of finding modes in higher dimension and the lack of any natural framework to study the inferential properties of modes in higher dimensions. The HMAC provides a computationally fast iterative algorithm for calculating the modes and thereby providing a clustering approach which is scalable to high dimensions. This article provides the description of the R-package that implements HMAC and additionally provides an wide array visualization tools for representing clusters in high dimensions. Further, we propose a novel parallel implementation of the approach which dramatically reduces the computational time especially for large data sets, both in data dimensions and the number of observations. This paper is organized as follows: Section 2 briefly introduces the algorithm of Modal Expectation Maximization (MEM) and builds the notion of mode association clustering technique. Section 3 describes a parallel computing framework of HMAC along with computing time comparisons. Section 4 illustrates the implementation of clustering functions in the R-package Modalclust along with examples of the plotting functions especially designed for objects of class hmac. Section 5 provides the conclusion and discussion. 2. Modal EM and HMAC The main challenge of using mode-based clustering in high dimensions is the cost of computing modes, which are mathematically evaluated as local maximas of the density function with support on R D, D being the data dimension. Traditional techniques of finding local maxima, such as hill climbing works well for univariate data. But multivariate hill climbing is computationally expensive thereby limiting its use in high dimensions. Li et al. (2007) proposed an algorithm that solves a local maximum of a kernel density by ascending iterations starting from the data points. Since the algorithm is very similar to Expectation Maximization (EM) algorithm, it is named as Modal Expectation Maximization (MEM). Given a multivariate kernel K, let the density of the data be given by f(x Σ) = n i=1 1 n K(x x i Σ), where Σ is the matrix of smoothing parameters. Now, given any initial value x (0), the MEM solves a local maximum of the mixture density by alternating the following two steps until it meets some user defined stopping criterion. Step 1. Let p i = K(x x i Σ) nf(x (r) ), i = 1,..., n. 3
4 Step 2. Update x (r+1) = argmax x n p i log f i (x). i=1 Details of convergence of the MEM approach can be found in Li et al. (2007). The above iterative steps provide a computationally simpler approach for hill climbing from any starting point x R D by exploiting the properties of density functions. Further, in the special case of Gaussians kernels, i.e., K(x x i Σ) = φ(x x i, Σ), where φ( ) is the pdf of a Gaussian distribution, the update of x (r+1) is simply x (r+1) = n p i x i. allowing us to avoid the numerical optimization of Step 2. i=1 Now we present the HMAC algorithm. First we scale the data and use a kernel density estimator, with a normal kernel to estimate the density of the data. The variance of the kernel, Σ is a diagonal matrix with all entries σ 2 denoted by D(σ 2 ), thus σ 2 is the single smoothing parameter for all the dimensions. The choice of the smoothing parameter is an area of research in itself. In the present version of the program we incorporate the strategy of using pseudo degrees of freedom, first proposed in Lindsay et al. (2008) and then developed in Lindsay et al. (2011). Their strategy provides us with a range of smoothing parameters and exploring them from finest to coarsest resolution provides the user with the desired hierarchical clustering. First we describe the steps of Mode Association Clustering (MAC) for a single bandwidth σ 2. Algorithm for MAC for a single bandwidth Step 1. Given a set of data S = {x 1,x 2,...,x n }, x i R d form kernel density f(x S,σ 2 ) = n i=1 1 n φ( x x i,d(σ 2 ) ). (1) Step 2. Use f(x S,σ 2 ) as the density function. Use each x i, i = 1,2,...,n, as the initial value in the MEM algorithm and find a mode of f(x S,σ 2 ). Let the mode identified by starting from x i be M σ (x i ). Step 3. Extract distinctive values from the set {M σ (x i ),i = 1,2,...,n} to form a set G. Label the elements in G from 1 to G. In practice, due to finite precision, two modes are regarded equal if their distance is below a threshold. Step 4. If M σ (x i ) equals the k th element in G, x i is put in the k th cluster. 4
5 We note that when the bandwidth σ increases, the kernel density estimator f(x S, σ 2 ) in (1) becomes smoother, and thus more points tend to climb to the same mode. This suggests a natural approach for hierarchical organization (or nesting ) of our MAC clusters. Thus, given a range of bandwidths σ 1 < σ 2 < < σ L, The clustering can be performed in the following bottom-up manner. First we perform MAC at the smallest bandwidth σ 1. At any bandwidth σ l, the clustering representatives in G l 1 obtained from the preceding bandwidth are fed into MAC using the density f(x S, σ 2 l ). The modes identified at this level form a new set of cluster representatives G l. This procedure is repeated across all σ l s. This preserves the hierarchy of clusters and thus the name Hierarchical Mode Association Clustering (HMAC). To summarize we present the HMAC procedure in the following box. Algorithm of HMAC for a sequence of bandwidths Step 1. Start with the data G 0 = {x 1,...,x n } and set level l = 0 and initialize the mode association of the i th data point as P 0 (x i ) = i. Step 2. l l + 1. Step 3. Form kernel density as in (1) using σ 2 l. Step 4. Cluster the points in G l 1 by using density f(x S,σl 2 ). Let the set of distinct modes obtained be G l. Step 5. If P l 1 (x i ) = k and the k th element in G l 1 is clustered to the k th mode in G l, then P l (x i ) = k. In another word, the cluster of x i at level l is determined by its cluster representative in G l 1. Step 6. Stop if l = L, otherwise go to Step Parallel HMAC The HMAC approach, though scalable to higher number of dimensions becomes computationally expensive when the number of objects (n) becomes large. The first level of HMAC requires one to start the MEM from each observation and find its modal association with respect to the overall density. This step alone becomes computationally expensive when n becomes large. Note that the hierarchical steps for level l = 2 onwards only needs to start the MEM from the modes of the previous level G l 1 and hence the computational cost does not increase at the rate of n. Fortunately the mode association clustering provides a natural framework of proposing a divide and conquer algorithm of clustering. One can simply partition the data into m partitions, perform modal clustering on each of those partitions and then pool the modes obtained from each of these 5
6 partitions and allow the modes to form the L 1 top layers of modes. If the user has access to multiple computing cores on the same machine or several processors of a shared memory computing cluster, the divide and conquer algorithm can be seamlessly parallelized. This section provides the details of the parallel version of HMAC and its application together with some comparisons of performance of the parallel approach. First we describe the main framework of parallel HMAC and then we provide details on regarding how to choose the partitions. Algorithm for parallel HMAC (PHMAC) in multiple dimensions Step 1. Let G 0 = {x 1,..., x n }. Partition the data (n objects) into m partitions G j o, j = 1, 2,...,m. Step 2. Perform HMAC on each of these partitions at the lowest resolution, i.e, using σ 1 and get the modes G j 1, j = 1, 2,...,m. Step 3. Pool the modes from each partitioned to form G 1 = Step 4. Perform HMAC starting from Step 2 and l = 1 and obtain the final hierarchical clustering. m j=1 G j 1 A demonstration of different steps of parallel clustering with four random partitions is given in Figure 1. The original data set is partitioned into four random partitions and initial modal clustering is performed within the partitions. In the next step the modes of each of these partitions are merged to form the overall modal clusters in Figure 1(c) Discussion on parallel clustering In general model based clustering is hard to parallelize. But modes have a natural hierarchy and it is computationally easy to merge modes from different partitions. But one needs to decide on how to best perform the partition and how many partitions to use. In this paper we will just provide some guidelines regarding the two choices without exploring their optimality in details. In the absence of any other knowledge one should randomly partition the data. Other choices include partitioning data based on certain coordinates which form a natural clustering and then taking products of a few of those coordinates to build the overall partition. This strategy might increase the computational speed by restricting the modes within a relatively homogeneous set of observations. Another choice might be to sample the data and build partitions based on the modes 6
7 Partition 1 Partition 2 V Partition 3 Partition 4 V1 (a) (b) Partition 1 Partition 2 Merge modes from partitions Partition 3 Partition 4 (c) (d) Figure 1: Steps in parallel HMAC procedure for a simulated data (a) shows the simulated data with four clusters along with the contour plot, where the color indicates the final clustering using phmac; (b) shows the four random partitions of the unlabeled data along with the modes (red asterisk) at each partition; (c) shows the mode obtained from the four partitions; (d) shows the final modes (green triangles) starting from the modes of the partitioned data. 7
8 of the sampled data. The number of partitions one should use to minimize the computational time is a complex function of the number of available processors, the number of observations to be clustered and the choice of smoothing parameter. If using more partitions, one might speed up the first step but would run the risk of having to merge more modes at the next level, where the hill climbing is done from the collection of modes from each partitions with respect to the overall density. On the other hand for large n if one chooses too few partitions or no partitions, one would require huge computational cost at the lowest level. Finally the choice of smoothing parameter will also determine how many modes one needs to start from at the merged level. Next we compare the computing speed for parallel versus serial clustering using 1, 2, 4, 8 and 12 multicore processors. Tests were performed on a 64 bits 4 Quad Core AMD 8384 (2.7 Ghz each core), with 16 GB RAM running Linux Centos 5 and R version From Table 1, it is clear to observe that parallel computing significantly increases the computing speed. Because the nonparametric density estimate in (1) is a sum of kernels centered at every data point, the amount of computation to identify the mode associated with a single point grows linearly with n, the size of the data set. The computational complexity of clustering all the data by HMAC is thus quadratic in n. Suppose we have p processors. The computing complexity for the initial level for serial HMAC is n 2 and for parallel HMAC is thus ( n p )2. But as discussed before, we can see that the computational speed is not a monotonically decreasing function of the number of processors. Theoretically, it is true that more processors can reduce the computing complexity at the initial step. However, in practice, if the data set is not sufficiently large, using more processors may not save time as it may produce a large number of modes in the pooled stage. When the n=10,000 or n=50,000 including more processors provides dramatic decrease in computing time, where as for n=2000 there is a clear dip in time elapsed when using 4 or 8 processors instead of the maximum 12 processors. For n=50,000 the decrease in computing time from using one processor to using 12 processors is more than 40 folds ( see Figure 2) but even if the user is able to use just two processors the computing time is reduced to one third of what a single processor would take. Even for n=20,000 the advantage of using 12 processors is almost 30 fold, whereas for n=2,000 the advantage is only 8 fold. In fact the lowest time is actually clocked by 8 processors for n=2,000 but using all 12 processors does not increase the time very much. These comparisons show the potential for parallelizing the modal 8
9 clustering algorithm and its inherent use of clustering high throughput data. Table 1: Comparison of computing time (elapsed time in seconds) using different number of processors Number of processors Data dimensions n=2,000, d= n=2,000, d= n=2,000, d= n=10,000, d= n=10,000, d= n=50,000, d= Fold increase in time N=2,000 N=10,000 N=50, Number of processors Figure 2: Comparison of fold increase in time of clustering two dimensional data of different sample sizes with respect to using 12 processors 9
10 4. Example of using R-package Modalclust In this section we will demonstrate the functions and plotting tools available in the Modalclust package Modal Clustering First we provide an example of performing modal clustering to extract the subpopulations in the logcta20 dataset. The description of the dataset is given in the package and the scatter plot along with its smooth density is provided in Figure 3. First we use the following commands to download and install the package: > install.packages( Modalclust ) > library(modalclust) Using the following commands, we can get the standard (serial) HMAC and parallel HMAC using two processors for logcta20 data. > logcta20.hmac <-phmac(logcta20,npart=1,parallel=f) > logcta20p2.hmac <-phmac(logcta20,npart=2,parallel=t) Colors represent different clusters V V V1 Figure 3: Scatter plot of logcta20 data V1 Level 4 with number of clusters 3 Figure 4: HMAC implied clusters of logcta20 data 10
11 Both implementations result in the final clustering given in Figure 4, which clearly identifies the three distinct subpopluation which has been biologically validated. Other model based clustering methods in R, such as Mclust, kmeans could not capture the subpopulation structure as the individual density are non-normal in shape. Distance based clustering method e.g. hclust with a range of linkage functions performed even poorly. By default the function selects an interesting range of smoothing parameters with ten σ 2 values and the final clustering only shows the results from the levels which produced merging from previous its level. For example for the logcta20 the smoothing parameters chosen automatically are > logcta20.hmac$sigma [1] which are chosen using the pseudo degrees of freedom criterion mentioned in Section 2. Though we started with 10 different smoothing levels the final clustering shows only 6 different levels along with a decreasing number of hierarchical cluster > logcta20.hmac$level [1] > logcta20.hmac$n.cluster [1] The user can also provide smoothing levels using the option sigmaselect in phmac. There is also the option of starting the phmac from user defined modes instead of the original data points. This option comes handy if the user wishes to merge clusters obtained from other model based clustering e.g. Mclust, kmeans Some Examples of Plotting There are several plotting functions in Modalclust which can be used to visualize the output from the function phmac. The plotting functions are defined on object class hmac, which is the default class of a phmac output. These plot functions will be illustrated through a data set named disc2d, composed of 400 observations representing the shape of two half discs. The scatter plot of disc2d along with its contours is given in Figure 5. 11
12 V1 V Contour plot Figure 5: The scatter plot of disc2d data along with its probability contours HMAC Dendrogram clustering level Figure 6: Hierarchical tree (Dendrogram) of the disc2d data showing the clustering at four levels of smoothing 12
13 First we will discuss the standard plot function for an object of class hmac. This unique and extremely informative plot shows the hierarchical tree obtained from modal clustering. It is available for any dimensions and can be obtained by > data(disc2d.hmac) > plot(disc2d.hmac) The dendrogram obtained for the disc2d data is given in Figure 6. The y-axis gives the different levels and the tree displays the merging at different levels. There are several options available for drawing the tree including starting the tree from a specific level, drawing the tree only up to a desired number of clusters and comparison of the clustering results with user defined clusters. Output from this tree is used for other plotting functions available in this package. Hard clustering produced by HMAC Colors represent different clusters Soft clustering produced by HMAC Colors represent different clusters boundary points V V V V1 Figure 7: Hard clustering for disc2d data at level 3 Figure 8: Soft clustering for disc2d data at level 2 The rest of the plotting functions are mainly designed for visualizing clustering results for two dimensional data, although one can provide multivariate extensions of the functions by considering all possible pairwise dimensions. One can obtain the hard clustering of the data for each level using the command > hard.hmac(disc2d.hmac) 13
14 Alternatively the user can specify the smoothing level or the number of desired clusters and obtain the corresponding hard clustering of the data. For example the plot in Figure 7 can be obtained by either of the two commands > hard.hmac(disc2d.hmac,n.cluster=2) > hard.hmac(disc2d.hmac,level=3) Another function allows the user to visualize the soft clustering of the data based on the posterior probability of each observation belonging to the clusters at a specified level. For example the plot in Figure 8 can be obtained by using > soft.hmac(disc2d.hmac,n.cluster=3) The plot enables us to visualize the probabilistic clustering of the three cluster model. User can specify a probability threshold for assigning observations which clearly belong to a cluster or lies in the boundary of more than one clusters. Points having posterior probability below the user specified boundlevel (default value 0.4) are assigned as boundary points and colored in gray. In Figure 8 we have five boundary points among the 400 original observations that are clustered. Additionally at any specified level or cluster size the plot=false option in hard.hmac returns the cluster membership. Similarly plot=false option in soft.hmac returns a list that contains the posterior probability of each observation and boundary points. > disc2d.2clust <- hard.hmac(disc2d.hmac,n.cluster=2,plot=f) > disc2d.2clust.soft <- soft.hmac(disc2d.hmac,n.cluster=2,plot=f) Finally we demonstrate another very useful function for choosing a cluster dynamically from a two dimensional plot. The function choose.cluster allows the user to click on any part of a two dimensional plot and dynamically select the cluster that point will belong to. One can start the display by invoking the command > choose.cluster(disc2d.hmac,n.cluster=2) which will open up a graphical window with the scatter plot as displayed in the left panel of Figure 9. After the user clicks a point anywhere near the upper disc the points in the cluster consisting of upper disc will light up as in the right panel of Figure 9. 14
15 V V Choose a point by V1 clicking on the graph Stop the program by click anywhere outside the graph Choose a point by V1 clicking on the graph Stop the program by click anywhere outside the graph Figure 9: Graphical display for choosing the cluster using the function choose.cluster for the logcta20 data at level 3 with 2 clusters. The left panel displays the plot before the click and the right panel highlights the points after the user pointer click at the arrow head ( ). If the user clicks any existing data point, all other points belonging to the same cluster will light up. One can stop the program by clicking anywhere outside the plot area. This is an extremely useful function and can be used in merging clusters based on expert opinion or to design semisupervised clustering. 5. Discussion Modalclust performs a hierarchical model based clustering allowing for arbitrary density shapes.. Parallel computing can dramatically increase the computing speed by splitting the data and running the HMAC simultaneously on multicore processors. Plotting functions give the user a comprehensive visualizing and understanding of the clustering result. One future work from this stage would be increase computing speed, especially for large data set. From Section 3, it is clear to see, parallel computing increases the computing speed a lot. That relies on the computing equipment. If one user has no multicore or a few multicore processors available, it will take a lot of the computing resources when clustering large data sets. One potential way to solve the computing speed problem is using k-means or other faster clustering techniques 15
16 initially, and using the HMAC from the centers of each cluster of initial clustering results. For example, if we have a data set with 20,000 observations, we can use k-means clustering and choose a certain number of centers, like 200 centers and run k-means clustering first. And then we start from the centers of 200 clusters and clustering by HMAC. Theoretically it is a sub-optimal way compared with running HMAC for all points. In practice, it is very useful to reduce the computing costs and still obtain the right clustering. In addition, we are currently working on an implementation of modal clustering for online or streaming data, where the goal would be to update an existing cluster with the new data without storing all the original data points and allowing for creation of new clusters and merging of existing clusters. Sources, binaries and documentation of Modalclust are available for download from the Comprehensive R Archive Network under the GNU Public License. References Cheng, M.-Y., Hall, P., Calibrating the excess mass and dip tests of modality. J. R. Stat. Soc. Ser. B Stat. Methodol. 60 (3), Cheng, M.-Y., Hall, P., Mode testing in difficult cases. Ann. Statist. 27 (4), Fraley, C., Algorithms for model-based Gaussian hierarchical clustering. SIAM Journal on Scientific Computing 20, Fraley, C., Raftery, A. E., How many clusters? Which clustering method? - Answers via model-based cluster analysis. Computer Journal 41, Fraley, C., Raftery, A. E., MCLUST: Software for model-based cluster analysis. Journal of Classification 16, Fraley, C., Raftery, A. E., October 2002a. MCLUST: Software for model-based clustering, density estimation, and discriminant analysis. Technical Report 415, University of Washington, Department of Statistics. Fraley, C., Raftery, A. E., 2002b. Model-based clustering, discriminant analysis and density estimation. Journal of the American Statistical Association 97, Hartigan, J. A., The span test for unimodality. In: Bock, H. H. (Ed.), Classification and Related Methods of Data Analysis. Elsevier/North-Holland [Elsevier Science Publishing Co., New York; North- Holland Publishing Co., Amsterdam], pp
17 Hartigan, J. A., Hartigan, P. M., The dip test of unimodality. Ann. Statist. 13 (1), Li, J., Ray, S., Lindsay, B., A nonparametric statistical approach to clustering via mode identification. Journal of Machine Learning Research 8, Lindsay, B. G., Markatou, M., Ray, S., Spectral degrees of freedom and highdimensional smoothing. Tech. rep., Under Preparation. Lindsay, B. G., Markatou, M., Ray, S., Yang, K., Chen, S.-C., Quadratic distances on probabilities: a unified foundation. Ann. Statist. 36 (2), R Development Core Team, R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria, ISBN URL Ray, S., Lindsay, B., The topography of multivariate normal mixtures. Ann. Statist. 33 (5), Ray, S., Lindsay, B. G., Model selection in high dimensions: a quadratic-risk-based approach. J. Roy. Statist. Soc. Ser. B 70 (1), Silverman, B. W., Using kernel density estimates to investigate multimodality. J. Roy. Statist. Soc. Ser. B 43 (1), Silverman, B. W., Some properties of a test for multimodality based on kernel density estimates. In: Probability, statistics and analysis. Vol. 79 of London Math. Soc. Lecture Note Ser. Cambridge Univ. Press, Cambridge, pp
Clustering and Classification of Functional Data
 Clustering and Classification of Functional Data Université de Montréal, Département de mathématiques et de statistique, Montréal, 2008 Surajit Ray Boston University (Dept of Mathematics and Statistics)
Clustering and Classification of Functional Data Université de Montréal, Département de mathématiques et de statistique, Montréal, 2008 Surajit Ray Boston University (Dept of Mathematics and Statistics)
Clustering Lecture 5: Mixture Model
 Clustering Lecture 5: Mixture Model Jing Gao SUNY Buffalo 1 Outline Basics Motivation, definition, evaluation Methods Partitional Hierarchical Density-based Mixture model Spectral methods Advanced topics
Clustering Lecture 5: Mixture Model Jing Gao SUNY Buffalo 1 Outline Basics Motivation, definition, evaluation Methods Partitional Hierarchical Density-based Mixture model Spectral methods Advanced topics
MSA220 - Statistical Learning for Big Data
 MSA220 - Statistical Learning for Big Data Lecture 13 Rebecka Jörnsten Mathematical Sciences University of Gothenburg and Chalmers University of Technology Clustering Explorative analysis - finding groups
MSA220 - Statistical Learning for Big Data Lecture 13 Rebecka Jörnsten Mathematical Sciences University of Gothenburg and Chalmers University of Technology Clustering Explorative analysis - finding groups
MultiDimensional Signal Processing Master Degree in Ingegneria delle Telecomunicazioni A.A
 MultiDimensional Signal Processing Master Degree in Ingegneria delle Telecomunicazioni A.A. 205-206 Pietro Guccione, PhD DEI - DIPARTIMENTO DI INGEGNERIA ELETTRICA E DELL INFORMAZIONE POLITECNICO DI BARI
MultiDimensional Signal Processing Master Degree in Ingegneria delle Telecomunicazioni A.A. 205-206 Pietro Guccione, PhD DEI - DIPARTIMENTO DI INGEGNERIA ELETTRICA E DELL INFORMAZIONE POLITECNICO DI BARI
A Nonparametric Statistical Approach to Clustering via Mode Identification
 A Nonparametric Statistical Approach to Clustering via Mode Identification Jia Li Surajit Ray Bruce G. Lindsay Abstract In this paper, we develop a mode-based clustering approach applying new optimization
A Nonparametric Statistical Approach to Clustering via Mode Identification Jia Li Surajit Ray Bruce G. Lindsay Abstract In this paper, we develop a mode-based clustering approach applying new optimization
CS 2750 Machine Learning. Lecture 19. Clustering. CS 2750 Machine Learning. Clustering. Groups together similar instances in the data sample
 Lecture 9 Clustering Milos Hauskrecht milos@cs.pitt.edu 539 Sennott Square Clustering Groups together similar instances in the data sample Basic clustering problem: distribute data into k different groups
Lecture 9 Clustering Milos Hauskrecht milos@cs.pitt.edu 539 Sennott Square Clustering Groups together similar instances in the data sample Basic clustering problem: distribute data into k different groups
Hard clustering. Each object is assigned to one and only one cluster. Hierarchical clustering is usually hard. Soft (fuzzy) clustering
 An unsupervised machine learning problem Grouping a set of objects in such a way that objects in the same group (a cluster) are more similar (in some sense or another) to each other than to those in other
An unsupervised machine learning problem Grouping a set of objects in such a way that objects in the same group (a cluster) are more similar (in some sense or another) to each other than to those in other
Expectation Maximization (EM) and Gaussian Mixture Models
 Expectation Maximization (EM) and Gaussian Mixture Models Reference: The Elements of Statistical Learning, by T. Hastie, R. Tibshirani, J. Friedman, Springer 1 2 3 4 5 6 7 8 Unsupervised Learning Motivation
Expectation Maximization (EM) and Gaussian Mixture Models Reference: The Elements of Statistical Learning, by T. Hastie, R. Tibshirani, J. Friedman, Springer 1 2 3 4 5 6 7 8 Unsupervised Learning Motivation
Understanding Clustering Supervising the unsupervised
 Understanding Clustering Supervising the unsupervised Janu Verma IBM T.J. Watson Research Center, New York http://jverma.github.io/ jverma@us.ibm.com @januverma Clustering Grouping together similar data
Understanding Clustering Supervising the unsupervised Janu Verma IBM T.J. Watson Research Center, New York http://jverma.github.io/ jverma@us.ibm.com @januverma Clustering Grouping together similar data
Cluster Analysis. Jia Li Department of Statistics Penn State University. Summer School in Statistics for Astronomers IV June 9-14, 2008
 Cluster Analysis Jia Li Department of Statistics Penn State University Summer School in Statistics for Astronomers IV June 9-1, 8 1 Clustering A basic tool in data mining/pattern recognition: Divide a
Cluster Analysis Jia Li Department of Statistics Penn State University Summer School in Statistics for Astronomers IV June 9-1, 8 1 Clustering A basic tool in data mining/pattern recognition: Divide a
CS 1675 Introduction to Machine Learning Lecture 18. Clustering. Clustering. Groups together similar instances in the data sample
 CS 1675 Introduction to Machine Learning Lecture 18 Clustering Milos Hauskrecht milos@cs.pitt.edu 539 Sennott Square Clustering Groups together similar instances in the data sample Basic clustering problem:
CS 1675 Introduction to Machine Learning Lecture 18 Clustering Milos Hauskrecht milos@cs.pitt.edu 539 Sennott Square Clustering Groups together similar instances in the data sample Basic clustering problem:
Clustering CS 550: Machine Learning
 Clustering CS 550: Machine Learning This slide set mainly uses the slides given in the following links: http://www-users.cs.umn.edu/~kumar/dmbook/ch8.pdf http://www-users.cs.umn.edu/~kumar/dmbook/dmslides/chap8_basic_cluster_analysis.pdf
Clustering CS 550: Machine Learning This slide set mainly uses the slides given in the following links: http://www-users.cs.umn.edu/~kumar/dmbook/ch8.pdf http://www-users.cs.umn.edu/~kumar/dmbook/dmslides/chap8_basic_cluster_analysis.pdf
CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS
 CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS CHAPTER 4 CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS 4.1 Introduction Optical character recognition is one of
CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS CHAPTER 4 CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS 4.1 Introduction Optical character recognition is one of
Dynamic Thresholding for Image Analysis
 Dynamic Thresholding for Image Analysis Statistical Consulting Report for Edward Chan Clean Energy Research Center University of British Columbia by Libo Lu Department of Statistics University of British
Dynamic Thresholding for Image Analysis Statistical Consulting Report for Edward Chan Clean Energy Research Center University of British Columbia by Libo Lu Department of Statistics University of British
CS 229 Midterm Review
 CS 229 Midterm Review Course Staff Fall 2018 11/2/2018 Outline Today: SVMs Kernels Tree Ensembles EM Algorithm / Mixture Models [ Focus on building intuition, less so on solving specific problems. Ask
CS 229 Midterm Review Course Staff Fall 2018 11/2/2018 Outline Today: SVMs Kernels Tree Ensembles EM Algorithm / Mixture Models [ Focus on building intuition, less so on solving specific problems. Ask
SYDE Winter 2011 Introduction to Pattern Recognition. Clustering
 SYDE 372 - Winter 2011 Introduction to Pattern Recognition Clustering Alexander Wong Department of Systems Design Engineering University of Waterloo Outline 1 2 3 4 5 All the approaches we have learned
SYDE 372 - Winter 2011 Introduction to Pattern Recognition Clustering Alexander Wong Department of Systems Design Engineering University of Waterloo Outline 1 2 3 4 5 All the approaches we have learned
Clustering and Visualisation of Data
 Clustering and Visualisation of Data Hiroshi Shimodaira January-March 28 Cluster analysis aims to partition a data set into meaningful or useful groups, based on distances between data points. In some
Clustering and Visualisation of Data Hiroshi Shimodaira January-March 28 Cluster analysis aims to partition a data set into meaningful or useful groups, based on distances between data points. In some
Classification. Vladimir Curic. Centre for Image Analysis Swedish University of Agricultural Sciences Uppsala University
 Classification Vladimir Curic Centre for Image Analysis Swedish University of Agricultural Sciences Uppsala University Outline An overview on classification Basics of classification How to choose appropriate
Classification Vladimir Curic Centre for Image Analysis Swedish University of Agricultural Sciences Uppsala University Outline An overview on classification Basics of classification How to choose appropriate
Cluster Analysis. Debashis Ghosh Department of Statistics Penn State University (based on slides from Jia Li, Dept. of Statistics)
 Cluster Analysis Debashis Ghosh Department of Statistics Penn State University (based on slides from Jia Li, Dept. of Statistics) Summer School in Statistics for Astronomers June 1-6, 9 Clustering: Intuition
Cluster Analysis Debashis Ghosh Department of Statistics Penn State University (based on slides from Jia Li, Dept. of Statistics) Summer School in Statistics for Astronomers June 1-6, 9 Clustering: Intuition
Chapter 6 Continued: Partitioning Methods
 Chapter 6 Continued: Partitioning Methods Partitioning methods fix the number of clusters k and seek the best possible partition for that k. The goal is to choose the partition which gives the optimal
Chapter 6 Continued: Partitioning Methods Partitioning methods fix the number of clusters k and seek the best possible partition for that k. The goal is to choose the partition which gives the optimal
Region-based Segmentation
 Region-based Segmentation Image Segmentation Group similar components (such as, pixels in an image, image frames in a video) to obtain a compact representation. Applications: Finding tumors, veins, etc.
Region-based Segmentation Image Segmentation Group similar components (such as, pixels in an image, image frames in a video) to obtain a compact representation. Applications: Finding tumors, veins, etc.
Machine Learning and Data Mining. Clustering (1): Basics. Kalev Kask
 Machine Learning and Data Mining Clustering (1): Basics Kalev Kask Unsupervised learning Supervised learning Predict target value ( y ) given features ( x ) Unsupervised learning Understand patterns of
Machine Learning and Data Mining Clustering (1): Basics Kalev Kask Unsupervised learning Supervised learning Predict target value ( y ) given features ( x ) Unsupervised learning Understand patterns of
Motivation. Technical Background
 Handling Outliers through Agglomerative Clustering with Full Model Maximum Likelihood Estimation, with Application to Flow Cytometry Mark Gordon, Justin Li, Kevin Matzen, Bryce Wiedenbeck Motivation Clustering
Handling Outliers through Agglomerative Clustering with Full Model Maximum Likelihood Estimation, with Application to Flow Cytometry Mark Gordon, Justin Li, Kevin Matzen, Bryce Wiedenbeck Motivation Clustering
Approaches to Clustering
 Clustering A basic tool in data mining/pattern recognition: Divide a set of data into groups. Samples in one cluster are close and clusters are far apart. 6 5 4 3 1 1 3 1 1 3 4 5 6 Motivations: Discover
Clustering A basic tool in data mining/pattern recognition: Divide a set of data into groups. Samples in one cluster are close and clusters are far apart. 6 5 4 3 1 1 3 1 1 3 4 5 6 Motivations: Discover
Clustering and Dissimilarity Measures. Clustering. Dissimilarity Measures. Cluster Analysis. Perceptually-Inspired Measures
 Clustering and Dissimilarity Measures Clustering APR Course, Delft, The Netherlands Marco Loog May 19, 2008 1 What salient structures exist in the data? How many clusters? May 19, 2008 2 Cluster Analysis
Clustering and Dissimilarity Measures Clustering APR Course, Delft, The Netherlands Marco Loog May 19, 2008 1 What salient structures exist in the data? How many clusters? May 19, 2008 2 Cluster Analysis
Data Clustering Hierarchical Clustering, Density based clustering Grid based clustering
 Data Clustering Hierarchical Clustering, Density based clustering Grid based clustering Team 2 Prof. Anita Wasilewska CSE 634 Data Mining All Sources Used for the Presentation Olson CF. Parallel algorithms
Data Clustering Hierarchical Clustering, Density based clustering Grid based clustering Team 2 Prof. Anita Wasilewska CSE 634 Data Mining All Sources Used for the Presentation Olson CF. Parallel algorithms
Unsupervised Learning : Clustering
 Unsupervised Learning : Clustering Things to be Addressed Traditional Learning Models. Cluster Analysis K-means Clustering Algorithm Drawbacks of traditional clustering algorithms. Clustering as a complex
Unsupervised Learning : Clustering Things to be Addressed Traditional Learning Models. Cluster Analysis K-means Clustering Algorithm Drawbacks of traditional clustering algorithms. Clustering as a complex
DATA MINING LECTURE 7. Hierarchical Clustering, DBSCAN The EM Algorithm
 DATA MINING LECTURE 7 Hierarchical Clustering, DBSCAN The EM Algorithm CLUSTERING What is a Clustering? In general a grouping of objects such that the objects in a group (cluster) are similar (or related)
DATA MINING LECTURE 7 Hierarchical Clustering, DBSCAN The EM Algorithm CLUSTERING What is a Clustering? In general a grouping of objects such that the objects in a group (cluster) are similar (or related)
Segmentation of Images
 Segmentation of Images SEGMENTATION If an image has been preprocessed appropriately to remove noise and artifacts, segmentation is often the key step in interpreting the image. Image segmentation is a
Segmentation of Images SEGMENTATION If an image has been preprocessed appropriately to remove noise and artifacts, segmentation is often the key step in interpreting the image. Image segmentation is a
Note Set 4: Finite Mixture Models and the EM Algorithm
 Note Set 4: Finite Mixture Models and the EM Algorithm Padhraic Smyth, Department of Computer Science University of California, Irvine Finite Mixture Models A finite mixture model with K components, for
Note Set 4: Finite Mixture Models and the EM Algorithm Padhraic Smyth, Department of Computer Science University of California, Irvine Finite Mixture Models A finite mixture model with K components, for
Clustering: Classic Methods and Modern Views
 Clustering: Classic Methods and Modern Views Marina Meilă University of Washington mmp@stat.washington.edu June 22, 2015 Lorentz Center Workshop on Clusters, Games and Axioms Outline Paradigms for clustering
Clustering: Classic Methods and Modern Views Marina Meilă University of Washington mmp@stat.washington.edu June 22, 2015 Lorentz Center Workshop on Clusters, Games and Axioms Outline Paradigms for clustering
Network Traffic Measurements and Analysis
 DEIB - Politecnico di Milano Fall, 2017 Introduction Often, we have only a set of features x = x 1, x 2,, x n, but no associated response y. Therefore we are not interested in prediction nor classification,
DEIB - Politecnico di Milano Fall, 2017 Introduction Often, we have only a set of features x = x 1, x 2,, x n, but no associated response y. Therefore we are not interested in prediction nor classification,
Clustering: K-means and Kernel K-means
 Clustering: K-means and Kernel K-means Piyush Rai Machine Learning (CS771A) Aug 31, 2016 Machine Learning (CS771A) Clustering: K-means and Kernel K-means 1 Clustering Usually an unsupervised learning problem
Clustering: K-means and Kernel K-means Piyush Rai Machine Learning (CS771A) Aug 31, 2016 Machine Learning (CS771A) Clustering: K-means and Kernel K-means 1 Clustering Usually an unsupervised learning problem
Data Mining Chapter 9: Descriptive Modeling Fall 2011 Ming Li Department of Computer Science and Technology Nanjing University
 Data Mining Chapter 9: Descriptive Modeling Fall 2011 Ming Li Department of Computer Science and Technology Nanjing University Descriptive model A descriptive model presents the main features of the data
Data Mining Chapter 9: Descriptive Modeling Fall 2011 Ming Li Department of Computer Science and Technology Nanjing University Descriptive model A descriptive model presents the main features of the data
Mixture Models and the EM Algorithm
 Mixture Models and the EM Algorithm Padhraic Smyth, Department of Computer Science University of California, Irvine c 2017 1 Finite Mixture Models Say we have a data set D = {x 1,..., x N } where x i is
Mixture Models and the EM Algorithm Padhraic Smyth, Department of Computer Science University of California, Irvine c 2017 1 Finite Mixture Models Say we have a data set D = {x 1,..., x N } where x i is
Clustering. Supervised vs. Unsupervised Learning
 Clustering Supervised vs. Unsupervised Learning So far we have assumed that the training samples used to design the classifier were labeled by their class membership (supervised learning) We assume now
Clustering Supervised vs. Unsupervised Learning So far we have assumed that the training samples used to design the classifier were labeled by their class membership (supervised learning) We assume now
Smoothing Dissimilarities for Cluster Analysis: Binary Data and Functional Data
 Smoothing Dissimilarities for Cluster Analysis: Binary Data and unctional Data David B. University of South Carolina Department of Statistics Joint work with Zhimin Chen University of South Carolina Current
Smoothing Dissimilarities for Cluster Analysis: Binary Data and unctional Data David B. University of South Carolina Department of Statistics Joint work with Zhimin Chen University of South Carolina Current
INF 4300 Classification III Anne Solberg The agenda today:
 INF 4300 Classification III Anne Solberg 28.10.15 The agenda today: More on estimating classifier accuracy Curse of dimensionality and simple feature selection knn-classification K-means clustering 28.10.15
INF 4300 Classification III Anne Solberg 28.10.15 The agenda today: More on estimating classifier accuracy Curse of dimensionality and simple feature selection knn-classification K-means clustering 28.10.15
Overview Citation. ML Introduction. Overview Schedule. ML Intro Dataset. Introduction to Semi-Supervised Learning Review 10/4/2010
 INFORMATICS SEMINAR SEPT. 27 & OCT. 4, 2010 Introduction to Semi-Supervised Learning Review 2 Overview Citation X. Zhu and A.B. Goldberg, Introduction to Semi- Supervised Learning, Morgan & Claypool Publishers,
INFORMATICS SEMINAR SEPT. 27 & OCT. 4, 2010 Introduction to Semi-Supervised Learning Review 2 Overview Citation X. Zhu and A.B. Goldberg, Introduction to Semi- Supervised Learning, Morgan & Claypool Publishers,
Nonparametric Regression
 Nonparametric Regression John Fox Department of Sociology McMaster University 1280 Main Street West Hamilton, Ontario Canada L8S 4M4 jfox@mcmaster.ca February 2004 Abstract Nonparametric regression analysis
Nonparametric Regression John Fox Department of Sociology McMaster University 1280 Main Street West Hamilton, Ontario Canada L8S 4M4 jfox@mcmaster.ca February 2004 Abstract Nonparametric regression analysis
Cluster Analysis. Ying Shen, SSE, Tongji University
 Cluster Analysis Ying Shen, SSE, Tongji University Cluster analysis Cluster analysis groups data objects based only on the attributes in the data. The main objective is that The objects within a group
Cluster Analysis Ying Shen, SSE, Tongji University Cluster analysis Cluster analysis groups data objects based only on the attributes in the data. The main objective is that The objects within a group
Unsupervised Learning and Clustering
 Unsupervised Learning and Clustering Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Spring 2009 CS 551, Spring 2009 c 2009, Selim Aksoy (Bilkent University)
Unsupervised Learning and Clustering Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Spring 2009 CS 551, Spring 2009 c 2009, Selim Aksoy (Bilkent University)
Clustering. CS294 Practical Machine Learning Junming Yin 10/09/06
 Clustering CS294 Practical Machine Learning Junming Yin 10/09/06 Outline Introduction Unsupervised learning What is clustering? Application Dissimilarity (similarity) of objects Clustering algorithm K-means,
Clustering CS294 Practical Machine Learning Junming Yin 10/09/06 Outline Introduction Unsupervised learning What is clustering? Application Dissimilarity (similarity) of objects Clustering algorithm K-means,
Fall 09, Homework 5
 5-38 Fall 09, Homework 5 Due: Wednesday, November 8th, beginning of the class You can work in a group of up to two people. This group does not need to be the same group as for the other homeworks. You
5-38 Fall 09, Homework 5 Due: Wednesday, November 8th, beginning of the class You can work in a group of up to two people. This group does not need to be the same group as for the other homeworks. You
Random projection for non-gaussian mixture models
 Random projection for non-gaussian mixture models Győző Gidófalvi Department of Computer Science and Engineering University of California, San Diego La Jolla, CA 92037 gyozo@cs.ucsd.edu Abstract Recently,
Random projection for non-gaussian mixture models Győző Gidófalvi Department of Computer Science and Engineering University of California, San Diego La Jolla, CA 92037 gyozo@cs.ucsd.edu Abstract Recently,
Introduction to Pattern Recognition Part II. Selim Aksoy Bilkent University Department of Computer Engineering
 Introduction to Pattern Recognition Part II Selim Aksoy Bilkent University Department of Computer Engineering saksoy@cs.bilkent.edu.tr RETINA Pattern Recognition Tutorial, Summer 2005 Overview Statistical
Introduction to Pattern Recognition Part II Selim Aksoy Bilkent University Department of Computer Engineering saksoy@cs.bilkent.edu.tr RETINA Pattern Recognition Tutorial, Summer 2005 Overview Statistical
COMP5318 Knowledge Management & Data Mining Assignment 1
 COMP538 Knowledge Management & Data Mining Assignment Enoch Lau SID 20045765 7 May 2007 Abstract 5.5 Scalability............... 5 Clustering is a fundamental task in data mining that aims to place similar
COMP538 Knowledge Management & Data Mining Assignment Enoch Lau SID 20045765 7 May 2007 Abstract 5.5 Scalability............... 5 Clustering is a fundamental task in data mining that aims to place similar
Cluster Analysis. Angela Montanari and Laura Anderlucci
 Cluster Analysis Angela Montanari and Laura Anderlucci 1 Introduction Clustering a set of n objects into k groups is usually moved by the aim of identifying internally homogenous groups according to a
Cluster Analysis Angela Montanari and Laura Anderlucci 1 Introduction Clustering a set of n objects into k groups is usually moved by the aim of identifying internally homogenous groups according to a
CSE 5243 INTRO. TO DATA MINING
 CSE 5243 INTRO. TO DATA MINING Cluster Analysis: Basic Concepts and Methods Huan Sun, CSE@The Ohio State University 09/25/2017 Slides adapted from UIUC CS412, Fall 2017, by Prof. Jiawei Han 2 Chapter 10.
CSE 5243 INTRO. TO DATA MINING Cluster Analysis: Basic Concepts and Methods Huan Sun, CSE@The Ohio State University 09/25/2017 Slides adapted from UIUC CS412, Fall 2017, by Prof. Jiawei Han 2 Chapter 10.
CSE 5243 INTRO. TO DATA MINING
 CSE 5243 INTRO. TO DATA MINING Cluster Analysis: Basic Concepts and Methods Huan Sun, CSE@The Ohio State University Slides adapted from UIUC CS412, Fall 2017, by Prof. Jiawei Han 2 Chapter 10. Cluster
CSE 5243 INTRO. TO DATA MINING Cluster Analysis: Basic Concepts and Methods Huan Sun, CSE@The Ohio State University Slides adapted from UIUC CS412, Fall 2017, by Prof. Jiawei Han 2 Chapter 10. Cluster
Density estimation. In density estimation problems, we are given a random from an unknown density. Our objective is to estimate
 Density estimation In density estimation problems, we are given a random sample from an unknown density Our objective is to estimate? Applications Classification If we estimate the density for each class,
Density estimation In density estimation problems, we are given a random sample from an unknown density Our objective is to estimate? Applications Classification If we estimate the density for each class,
Machine Learning Lecture 3
 Machine Learning Lecture 3 Probability Density Estimation II 19.10.2017 Bastian Leibe RWTH Aachen http://www.vision.rwth-aachen.de leibe@vision.rwth-aachen.de Announcements Exam dates We re in the process
Machine Learning Lecture 3 Probability Density Estimation II 19.10.2017 Bastian Leibe RWTH Aachen http://www.vision.rwth-aachen.de leibe@vision.rwth-aachen.de Announcements Exam dates We re in the process
COMP 551 Applied Machine Learning Lecture 13: Unsupervised learning
 COMP 551 Applied Machine Learning Lecture 13: Unsupervised learning Associate Instructor: Herke van Hoof (herke.vanhoof@mail.mcgill.ca) Slides mostly by: (jpineau@cs.mcgill.ca) Class web page: www.cs.mcgill.ca/~jpineau/comp551
COMP 551 Applied Machine Learning Lecture 13: Unsupervised learning Associate Instructor: Herke van Hoof (herke.vanhoof@mail.mcgill.ca) Slides mostly by: (jpineau@cs.mcgill.ca) Class web page: www.cs.mcgill.ca/~jpineau/comp551
Points Lines Connected points X-Y Scatter. X-Y Matrix Star Plot Histogram Box Plot. Bar Group Bar Stacked H-Bar Grouped H-Bar Stacked
 Plotting Menu: QCExpert Plotting Module graphs offers various tools for visualization of uni- and multivariate data. Settings and options in different types of graphs allow for modifications and customizations
Plotting Menu: QCExpert Plotting Module graphs offers various tools for visualization of uni- and multivariate data. Settings and options in different types of graphs allow for modifications and customizations
Pattern Recognition. Kjell Elenius. Speech, Music and Hearing KTH. March 29, 2007 Speech recognition
 Pattern Recognition Kjell Elenius Speech, Music and Hearing KTH March 29, 2007 Speech recognition 2007 1 Ch 4. Pattern Recognition 1(3) Bayes Decision Theory Minimum-Error-Rate Decision Rules Discriminant
Pattern Recognition Kjell Elenius Speech, Music and Hearing KTH March 29, 2007 Speech recognition 2007 1 Ch 4. Pattern Recognition 1(3) Bayes Decision Theory Minimum-Error-Rate Decision Rules Discriminant
Cluster Analysis. Summer School on Geocomputation. 27 June July 2011 Vysoké Pole
 Cluster Analysis Summer School on Geocomputation 27 June 2011 2 July 2011 Vysoké Pole Lecture delivered by: doc. Mgr. Radoslav Harman, PhD. Faculty of Mathematics, Physics and Informatics Comenius University,
Cluster Analysis Summer School on Geocomputation 27 June 2011 2 July 2011 Vysoké Pole Lecture delivered by: doc. Mgr. Radoslav Harman, PhD. Faculty of Mathematics, Physics and Informatics Comenius University,
Data Mining 4. Cluster Analysis
 Data Mining 4. Cluster Analysis 4.5 Spring 2010 Instructor: Dr. Masoud Yaghini Introduction DBSCAN Algorithm OPTICS Algorithm DENCLUE Algorithm References Outline Introduction Introduction Density-based
Data Mining 4. Cluster Analysis 4.5 Spring 2010 Instructor: Dr. Masoud Yaghini Introduction DBSCAN Algorithm OPTICS Algorithm DENCLUE Algorithm References Outline Introduction Introduction Density-based
( ) =cov X Y = W PRINCIPAL COMPONENT ANALYSIS. Eigenvectors of the covariance matrix are the principal components
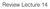 Review Lecture 14 ! PRINCIPAL COMPONENT ANALYSIS Eigenvectors of the covariance matrix are the principal components 1. =cov X Top K principal components are the eigenvectors with K largest eigenvalues
Review Lecture 14 ! PRINCIPAL COMPONENT ANALYSIS Eigenvectors of the covariance matrix are the principal components 1. =cov X Top K principal components are the eigenvectors with K largest eigenvalues
Parametrizing the easiness of machine learning problems. Sanjoy Dasgupta, UC San Diego
 Parametrizing the easiness of machine learning problems Sanjoy Dasgupta, UC San Diego Outline Linear separators Mixture models Nonparametric clustering Nonparametric classification and regression Nearest
Parametrizing the easiness of machine learning problems Sanjoy Dasgupta, UC San Diego Outline Linear separators Mixture models Nonparametric clustering Nonparametric classification and regression Nearest
K-means and Hierarchical Clustering
 K-means and Hierarchical Clustering Xiaohui Xie University of California, Irvine K-means and Hierarchical Clustering p.1/18 Clustering Given n data points X = {x 1, x 2,, x n }. Clustering is the partitioning
K-means and Hierarchical Clustering Xiaohui Xie University of California, Irvine K-means and Hierarchical Clustering p.1/18 Clustering Given n data points X = {x 1, x 2,, x n }. Clustering is the partitioning
ADVANCED MACHINE LEARNING MACHINE LEARNING. Kernel for Clustering kernel K-Means
 1 MACHINE LEARNING Kernel for Clustering ernel K-Means Outline of Today s Lecture 1. Review principle and steps of K-Means algorithm. Derive ernel version of K-means 3. Exercise: Discuss the geometrical
1 MACHINE LEARNING Kernel for Clustering ernel K-Means Outline of Today s Lecture 1. Review principle and steps of K-Means algorithm. Derive ernel version of K-means 3. Exercise: Discuss the geometrical
Supervised vs. Unsupervised Learning
 Clustering Supervised vs. Unsupervised Learning So far we have assumed that the training samples used to design the classifier were labeled by their class membership (supervised learning) We assume now
Clustering Supervised vs. Unsupervised Learning So far we have assumed that the training samples used to design the classifier were labeled by their class membership (supervised learning) We assume now
Clustering. SC4/SM4 Data Mining and Machine Learning, Hilary Term 2017 Dino Sejdinovic
 Clustering SC4/SM4 Data Mining and Machine Learning, Hilary Term 2017 Dino Sejdinovic Clustering is one of the fundamental and ubiquitous tasks in exploratory data analysis a first intuition about the
Clustering SC4/SM4 Data Mining and Machine Learning, Hilary Term 2017 Dino Sejdinovic Clustering is one of the fundamental and ubiquitous tasks in exploratory data analysis a first intuition about the
Homework. Gaussian, Bishop 2.3 Non-parametric, Bishop 2.5 Linear regression Pod-cast lecture on-line. Next lectures:
 Homework Gaussian, Bishop 2.3 Non-parametric, Bishop 2.5 Linear regression 3.0-3.2 Pod-cast lecture on-line Next lectures: I posted a rough plan. It is flexible though so please come with suggestions Bayes
Homework Gaussian, Bishop 2.3 Non-parametric, Bishop 2.5 Linear regression 3.0-3.2 Pod-cast lecture on-line Next lectures: I posted a rough plan. It is flexible though so please come with suggestions Bayes
International Journal Of Engineering And Computer Science ISSN: Volume 5 Issue 11 Nov. 2016, Page No.
 www.ijecs.in International Journal Of Engineering And Computer Science ISSN: 2319-7242 Volume 5 Issue 11 Nov. 2016, Page No. 19054-19062 Review on K-Mode Clustering Antara Prakash, Simran Kalera, Archisha
www.ijecs.in International Journal Of Engineering And Computer Science ISSN: 2319-7242 Volume 5 Issue 11 Nov. 2016, Page No. 19054-19062 Review on K-Mode Clustering Antara Prakash, Simran Kalera, Archisha
The exam is closed book, closed notes except your one-page cheat sheet.
 CS 189 Fall 2015 Introduction to Machine Learning Final Please do not turn over the page before you are instructed to do so. You have 2 hours and 50 minutes. Please write your initials on the top-right
CS 189 Fall 2015 Introduction to Machine Learning Final Please do not turn over the page before you are instructed to do so. You have 2 hours and 50 minutes. Please write your initials on the top-right
Pattern Clustering with Similarity Measures
 Pattern Clustering with Similarity Measures Akula Ratna Babu 1, Miriyala Markandeyulu 2, Bussa V R R Nagarjuna 3 1 Pursuing M.Tech(CSE), Vignan s Lara Institute of Technology and Science, Vadlamudi, Guntur,
Pattern Clustering with Similarity Measures Akula Ratna Babu 1, Miriyala Markandeyulu 2, Bussa V R R Nagarjuna 3 1 Pursuing M.Tech(CSE), Vignan s Lara Institute of Technology and Science, Vadlamudi, Guntur,
An Introduction to Cluster Analysis. Zhaoxia Yu Department of Statistics Vice Chair of Undergraduate Affairs
 An Introduction to Cluster Analysis Zhaoxia Yu Department of Statistics Vice Chair of Undergraduate Affairs zhaoxia@ics.uci.edu 1 What can you say about the figure? signal C 0.0 0.5 1.0 1500 subjects Two
An Introduction to Cluster Analysis Zhaoxia Yu Department of Statistics Vice Chair of Undergraduate Affairs zhaoxia@ics.uci.edu 1 What can you say about the figure? signal C 0.0 0.5 1.0 1500 subjects Two
Clustering algorithms
 Clustering algorithms Machine Learning Hamid Beigy Sharif University of Technology Fall 1393 Hamid Beigy (Sharif University of Technology) Clustering algorithms Fall 1393 1 / 22 Table of contents 1 Supervised
Clustering algorithms Machine Learning Hamid Beigy Sharif University of Technology Fall 1393 Hamid Beigy (Sharif University of Technology) Clustering algorithms Fall 1393 1 / 22 Table of contents 1 Supervised
The Curse of Dimensionality
 The Curse of Dimensionality ACAS 2002 p1/66 Curse of Dimensionality The basic idea of the curse of dimensionality is that high dimensional data is difficult to work with for several reasons: Adding more
The Curse of Dimensionality ACAS 2002 p1/66 Curse of Dimensionality The basic idea of the curse of dimensionality is that high dimensional data is difficult to work with for several reasons: Adding more
Unsupervised Learning
 Unsupervised Learning Unsupervised learning Until now, we have assumed our training samples are labeled by their category membership. Methods that use labeled samples are said to be supervised. However,
Unsupervised Learning Unsupervised learning Until now, we have assumed our training samples are labeled by their category membership. Methods that use labeled samples are said to be supervised. However,
An Introduction to PDF Estimation and Clustering
 Sigmedia, Electronic Engineering Dept., Trinity College, Dublin. 1 An Introduction to PDF Estimation and Clustering David Corrigan corrigad@tcd.ie Electrical and Electronic Engineering Dept., University
Sigmedia, Electronic Engineering Dept., Trinity College, Dublin. 1 An Introduction to PDF Estimation and Clustering David Corrigan corrigad@tcd.ie Electrical and Electronic Engineering Dept., University
Instance-based Learning CE-717: Machine Learning Sharif University of Technology. M. Soleymani Fall 2015
 Instance-based Learning CE-717: Machine Learning Sharif University of Technology M. Soleymani Fall 2015 Outline Non-parametric approach Unsupervised: Non-parametric density estimation Parzen Windows K-Nearest
Instance-based Learning CE-717: Machine Learning Sharif University of Technology M. Soleymani Fall 2015 Outline Non-parametric approach Unsupervised: Non-parametric density estimation Parzen Windows K-Nearest
Classification: Linear Discriminant Functions
 Classification: Linear Discriminant Functions CE-725: Statistical Pattern Recognition Sharif University of Technology Spring 2013 Soleymani Outline Discriminant functions Linear Discriminant functions
Classification: Linear Discriminant Functions CE-725: Statistical Pattern Recognition Sharif University of Technology Spring 2013 Soleymani Outline Discriminant functions Linear Discriminant functions
Unsupervised Learning
 Outline Unsupervised Learning Basic concepts K-means algorithm Representation of clusters Hierarchical clustering Distance functions Which clustering algorithm to use? NN Supervised learning vs. unsupervised
Outline Unsupervised Learning Basic concepts K-means algorithm Representation of clusters Hierarchical clustering Distance functions Which clustering algorithm to use? NN Supervised learning vs. unsupervised
Data Mining Cluster Analysis: Basic Concepts and Algorithms. Slides From Lecture Notes for Chapter 8. Introduction to Data Mining
 Data Mining Cluster Analysis: Basic Concepts and Algorithms Slides From Lecture Notes for Chapter 8 Introduction to Data Mining by Tan, Steinbach, Kumar Tan,Steinbach, Kumar Introduction to Data Mining
Data Mining Cluster Analysis: Basic Concepts and Algorithms Slides From Lecture Notes for Chapter 8 Introduction to Data Mining by Tan, Steinbach, Kumar Tan,Steinbach, Kumar Introduction to Data Mining
CHAPTER 6 MODIFIED FUZZY TECHNIQUES BASED IMAGE SEGMENTATION
 CHAPTER 6 MODIFIED FUZZY TECHNIQUES BASED IMAGE SEGMENTATION 6.1 INTRODUCTION Fuzzy logic based computational techniques are becoming increasingly important in the medical image analysis arena. The significant
CHAPTER 6 MODIFIED FUZZY TECHNIQUES BASED IMAGE SEGMENTATION 6.1 INTRODUCTION Fuzzy logic based computational techniques are becoming increasingly important in the medical image analysis arena. The significant
Cluster Analysis. Mu-Chun Su. Department of Computer Science and Information Engineering National Central University 2003/3/11 1
 Cluster Analysis Mu-Chun Su Department of Computer Science and Information Engineering National Central University 2003/3/11 1 Introduction Cluster analysis is the formal study of algorithms and methods
Cluster Analysis Mu-Chun Su Department of Computer Science and Information Engineering National Central University 2003/3/11 1 Introduction Cluster analysis is the formal study of algorithms and methods
DATA DEPTH AND ITS APPLICATIONS IN CLASSIFICATION
 DATA DEPTH AND ITS APPLICATIONS IN CLASSIFICATION Ondrej Vencalek Department of Mathematical Analysis and Applications of Mathematics Palacky University Olomouc, CZECH REPUBLIC e-mail: ondrej.vencalek@upol.cz
DATA DEPTH AND ITS APPLICATIONS IN CLASSIFICATION Ondrej Vencalek Department of Mathematical Analysis and Applications of Mathematics Palacky University Olomouc, CZECH REPUBLIC e-mail: ondrej.vencalek@upol.cz
Divide and Conquer Kernel Ridge Regression
 Divide and Conquer Kernel Ridge Regression Yuchen Zhang John Duchi Martin Wainwright University of California, Berkeley COLT 2013 Yuchen Zhang (UC Berkeley) Divide and Conquer KRR COLT 2013 1 / 15 Problem
Divide and Conquer Kernel Ridge Regression Yuchen Zhang John Duchi Martin Wainwright University of California, Berkeley COLT 2013 Yuchen Zhang (UC Berkeley) Divide and Conquer KRR COLT 2013 1 / 15 Problem
GPUML: Graphical processors for speeding up kernel machines
 GPUML: Graphical processors for speeding up kernel machines http://www.umiacs.umd.edu/~balajiv/gpuml.htm Balaji Vasan Srinivasan, Qi Hu, Ramani Duraiswami Department of Computer Science, University of
GPUML: Graphical processors for speeding up kernel machines http://www.umiacs.umd.edu/~balajiv/gpuml.htm Balaji Vasan Srinivasan, Qi Hu, Ramani Duraiswami Department of Computer Science, University of
Machine Learning for OR & FE
 Machine Learning for OR & FE Unsupervised Learning: Clustering Martin Haugh Department of Industrial Engineering and Operations Research Columbia University Email: martin.b.haugh@gmail.com (Some material
Machine Learning for OR & FE Unsupervised Learning: Clustering Martin Haugh Department of Industrial Engineering and Operations Research Columbia University Email: martin.b.haugh@gmail.com (Some material
K-Means Clustering 3/3/17
 K-Means Clustering 3/3/17 Unsupervised Learning We have a collection of unlabeled data points. We want to find underlying structure in the data. Examples: Identify groups of similar data points. Clustering
K-Means Clustering 3/3/17 Unsupervised Learning We have a collection of unlabeled data points. We want to find underlying structure in the data. Examples: Identify groups of similar data points. Clustering
Parallel Algorithms on Clusters of Multicores: Comparing Message Passing vs Hybrid Programming
 Parallel Algorithms on Clusters of Multicores: Comparing Message Passing vs Hybrid Programming Fabiana Leibovich, Laura De Giusti, and Marcelo Naiouf Instituto de Investigación en Informática LIDI (III-LIDI),
Parallel Algorithms on Clusters of Multicores: Comparing Message Passing vs Hybrid Programming Fabiana Leibovich, Laura De Giusti, and Marcelo Naiouf Instituto de Investigación en Informática LIDI (III-LIDI),
Lecture 11: E-M and MeanShift. CAP 5415 Fall 2007
 Lecture 11: E-M and MeanShift CAP 5415 Fall 2007 Review on Segmentation by Clustering Each Pixel Data Vector Example (From Comanciu and Meer) Review of k-means Let's find three clusters in this data These
Lecture 11: E-M and MeanShift CAP 5415 Fall 2007 Review on Segmentation by Clustering Each Pixel Data Vector Example (From Comanciu and Meer) Review of k-means Let's find three clusters in this data These
Lecture 10: Semantic Segmentation and Clustering
 Lecture 10: Semantic Segmentation and Clustering Vineet Kosaraju, Davy Ragland, Adrien Truong, Effie Nehoran, Maneekwan Toyungyernsub Department of Computer Science Stanford University Stanford, CA 94305
Lecture 10: Semantic Segmentation and Clustering Vineet Kosaraju, Davy Ragland, Adrien Truong, Effie Nehoran, Maneekwan Toyungyernsub Department of Computer Science Stanford University Stanford, CA 94305
Hierarchical Multi level Approach to graph clustering
 Hierarchical Multi level Approach to graph clustering by: Neda Shahidi neda@cs.utexas.edu Cesar mantilla, cesar.mantilla@mail.utexas.edu Advisor: Dr. Inderjit Dhillon Introduction Data sets can be presented
Hierarchical Multi level Approach to graph clustering by: Neda Shahidi neda@cs.utexas.edu Cesar mantilla, cesar.mantilla@mail.utexas.edu Advisor: Dr. Inderjit Dhillon Introduction Data sets can be presented
Robust PDF Table Locator
 Robust PDF Table Locator December 17, 2016 1 Introduction Data scientists rely on an abundance of tabular data stored in easy-to-machine-read formats like.csv files. Unfortunately, most government records
Robust PDF Table Locator December 17, 2016 1 Introduction Data scientists rely on an abundance of tabular data stored in easy-to-machine-read formats like.csv files. Unfortunately, most government records
Machine Learning Lecture 3
 Many slides adapted from B. Schiele Machine Learning Lecture 3 Probability Density Estimation II 26.04.2016 Bastian Leibe RWTH Aachen http://www.vision.rwth-aachen.de leibe@vision.rwth-aachen.de Course
Many slides adapted from B. Schiele Machine Learning Lecture 3 Probability Density Estimation II 26.04.2016 Bastian Leibe RWTH Aachen http://www.vision.rwth-aachen.de leibe@vision.rwth-aachen.de Course
Using Statistical Techniques to Improve the QC Process of Swell Noise Filtering
 Using Statistical Techniques to Improve the QC Process of Swell Noise Filtering A. Spanos* (Petroleum Geo-Services) & M. Bekara (PGS - Petroleum Geo- Services) SUMMARY The current approach for the quality
Using Statistical Techniques to Improve the QC Process of Swell Noise Filtering A. Spanos* (Petroleum Geo-Services) & M. Bekara (PGS - Petroleum Geo- Services) SUMMARY The current approach for the quality
The K-modes and Laplacian K-modes algorithms for clustering
 The K-modes and Laplacian K-modes algorithms for clustering Miguel Á. Carreira-Perpiñán Electrical Engineering and Computer Science University of California, Merced http://faculty.ucmerced.edu/mcarreira-perpinan
The K-modes and Laplacian K-modes algorithms for clustering Miguel Á. Carreira-Perpiñán Electrical Engineering and Computer Science University of California, Merced http://faculty.ucmerced.edu/mcarreira-perpinan
Machine Learning Lecture 3
 Course Outline Machine Learning Lecture 3 Fundamentals (2 weeks) Bayes Decision Theory Probability Density Estimation Probability Density Estimation II 26.04.206 Discriminative Approaches (5 weeks) Linear
Course Outline Machine Learning Lecture 3 Fundamentals (2 weeks) Bayes Decision Theory Probability Density Estimation Probability Density Estimation II 26.04.206 Discriminative Approaches (5 weeks) Linear
An Efficient Model Selection for Gaussian Mixture Model in a Bayesian Framework
 IEEE SIGNAL PROCESSING LETTERS, VOL. XX, NO. XX, XXX 23 An Efficient Model Selection for Gaussian Mixture Model in a Bayesian Framework Ji Won Yoon arxiv:37.99v [cs.lg] 3 Jul 23 Abstract In order to cluster
IEEE SIGNAL PROCESSING LETTERS, VOL. XX, NO. XX, XXX 23 An Efficient Model Selection for Gaussian Mixture Model in a Bayesian Framework Ji Won Yoon arxiv:37.99v [cs.lg] 3 Jul 23 Abstract In order to cluster
BBS654 Data Mining. Pinar Duygulu. Slides are adapted from Nazli Ikizler
 BBS654 Data Mining Pinar Duygulu Slides are adapted from Nazli Ikizler 1 Classification Classification systems: Supervised learning Make a rational prediction given evidence There are several methods for
BBS654 Data Mining Pinar Duygulu Slides are adapted from Nazli Ikizler 1 Classification Classification systems: Supervised learning Make a rational prediction given evidence There are several methods for
Unsupervised Learning and Clustering
 Unsupervised Learning and Clustering Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Spring 2008 CS 551, Spring 2008 c 2008, Selim Aksoy (Bilkent University)
Unsupervised Learning and Clustering Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Spring 2008 CS 551, Spring 2008 c 2008, Selim Aksoy (Bilkent University)
Including the Size of Regions in Image Segmentation by Region Based Graph
 International Journal of Emerging Engineering Research and Technology Volume 3, Issue 4, April 2015, PP 81-85 ISSN 2349-4395 (Print) & ISSN 2349-4409 (Online) Including the Size of Regions in Image Segmentation
International Journal of Emerging Engineering Research and Technology Volume 3, Issue 4, April 2015, PP 81-85 ISSN 2349-4395 (Print) & ISSN 2349-4409 (Online) Including the Size of Regions in Image Segmentation
Developing a Data Driven System for Computational Neuroscience
 Developing a Data Driven System for Computational Neuroscience Ross Snider and Yongming Zhu Montana State University, Bozeman MT 59717, USA Abstract. A data driven system implies the need to integrate
Developing a Data Driven System for Computational Neuroscience Ross Snider and Yongming Zhu Montana State University, Bozeman MT 59717, USA Abstract. A data driven system implies the need to integrate
Machine Learning. Unsupervised Learning. Manfred Huber
 Machine Learning Unsupervised Learning Manfred Huber 2015 1 Unsupervised Learning In supervised learning the training data provides desired target output for learning In unsupervised learning the training
Machine Learning Unsupervised Learning Manfred Huber 2015 1 Unsupervised Learning In supervised learning the training data provides desired target output for learning In unsupervised learning the training
Search Engines. Information Retrieval in Practice
 Search Engines Information Retrieval in Practice All slides Addison Wesley, 2008 Classification and Clustering Classification and clustering are classical pattern recognition / machine learning problems
Search Engines Information Retrieval in Practice All slides Addison Wesley, 2008 Classification and Clustering Classification and clustering are classical pattern recognition / machine learning problems
Summer School in Statistics for Astronomers & Physicists June 15-17, Cluster Analysis
 Summer School in Statistics for Astronomers & Physicists June 15-17, 2005 Session on Computational Algorithms for Astrostatistics Cluster Analysis Max Buot Department of Statistics Carnegie-Mellon University
Summer School in Statistics for Astronomers & Physicists June 15-17, 2005 Session on Computational Algorithms for Astrostatistics Cluster Analysis Max Buot Department of Statistics Carnegie-Mellon University
