marked Package Vignette
|
|
|
- Amber Baldwin
- 6 years ago
- Views:
Transcription
1 marked Package Vignette Jeff L. Laake, Devin S. Johnson, and Paul B. Conn Jan 16, 2013 Summary We describe an R package called marked for analysis of mark-recapture data. Currently, the package is capable of fitting Cormack-Jolly-Seber (CJS) models with maximum likelihood estimation (MLE) and Bayesian Markov Chain Monte Carlo (MCMC) methods. In addition, Jolly-Seber (JS) models can be fitted with MLE. Additional models will be added as the package is updated. We provide some example analyses with the well-known dipper data and compare run timing and results with MARK. For analysis with several thousand or more capture histories and several time-varying covariates, the run times with marked are substantially less. By providing this open source software, we hope to expand the analyst s toolbox and enble the knowledgeable user to understand fully the models and software. Introduction Currently the most comprehensive software for analysis of capture-recapture data is MARK (White and Burnham 1999). MARK is a FORTRAN program for fitting capture-recapture models that are manually constructed with a graphical user interface. RMark (Laake and Rexstad 2008) is an R package that constructs models for MARK with user-specified formulas to replace the manual model creation. With RMark and MARK most currently available capture-recapture models can be fitted and manipulated in R. Additional R packages for analysis of capture-recapture data have been made available including Rcapture (Baillargeon and Rivest 2007), mra (McDonald et al. 2005), secr (Borchers and Efford 2008), BTSPAS (Schwarz et al. 2009), SPACECAP (Royle et al. 2009), and BaSTA (Colchero et al. 2012). Rcapture fits closed and open models in a log-linear framework. The mra package 1
2 fits Cormack-Jolly-Seber (CJS) and the Huggins closed model with a regression approach to model specification. The secr and SPACECAP packages provide spatially explicit modeling of closed capture-recapture data and BTSPAS fits time-stratified Petersen models in a Bayesian framework. BaSTA estimates survival with covariates from capture-recapture/recovery data in a Bayesian framework when many individuals are of unknown age. Each package is designed for a unique niche or structure. We believe these alternative packages in R are useful because they expand the analyst s toolbox and the code is open source which enables the knowledgeable user to understand fully what the software is doing. Also, this independent innovation provides testing for existing software and potential gains in developer knowledge to improve existing software. We developed this R package we named marked for analysis with marked animals in contrast to the R package unmarked (Fiske and Chandler 2011) that focuses on analysis with unmarked animals. The original impetus for the package was to implement the CJS model using the hierarchical likelihood construction described by Pledger et al. (2003) and to improve on execution times with RMark/MARK (White and Burnham 1999;Laake and Rexstad 2008) for analysis of our own large data sets with many time-varying individual (animal-specific) covariates. Subsequently, we implemented the Jolly-Seber model with the Schwarz and Arnason (1996) POPAN structure where the hierarchical likelihood construction idea extended to the entry of animals into the population. We also added a Bayesian Markov Chain Monte Carlo (MCMC) implementation of the CJS model based on the approach used by Albert and Chib (1993) for analyzing binary data with a probit regression model. Background We assume you know and understand mark-recapture terminology and models. For background material on capture-recapture and MARK refer to org/software/mark/docs/book/. In the appendix, we provide a comparison of the MARK and marked data and model structures and details on likelihood construction for the CJS and JS models. Our focus in the vignette will be to provide examples on the use of marked. Dipper data example Anyone who has used RMark will find it easy to use marked because it has nearly identical syntax and structure with a few minor differences. The structure of the 2
3 input dataframe for marked is also identical to RMark. Any dataframe with a character field named ch containing the capture history will work with marked. If the capture history represents more than one animal then the dataframe should contain the additional numeric field named freq, which is the number of animals represented by that capture history. However, it is not necessary to accumulate capture histories in the input data because the process.data step will accumulate duplicated records by default. There can be any number of additional fields, some of which can be used as covariates in the analysis. Some of the functions in marked and RMark have the same name (e.g., process.data, make.design.data), so only one of the packages should be loaded in R to avoid aliasing and resulting errors. We will start by fitting the simplest CJS model with constant φ and p. The dipper data contain 294 records with the capture history (ch) and a factor variable sex with values Female or Male. Models are fitted with the function crm (capturerecapture model). After a call to library to attach the package, and data to retrieve the dipper data from the package, the model is fitted with crm and assigned to the object named model: > library(marked) > library(ggplot2) > data(dipper) > model=crm(dipper) Elapsed time in minutes: For this example, we are using the default CJS model and a default formula of 1 (constant) for each parameter. The function crm calls three functions in turn: 1) process.data to process the data, 2) make.design.data to create the list of parameterspecific dataframes, and 3) cjs, to fit the CJS model with the defined formulas. The code reports progress and some results as it executes. We ll suppress these messages in later examples but show them here to explain some of the messages. When it processes the data, it collapses the 294 rows into 55 which are the unique number of rows in the data including ch and sex. After processing, it creates the design data and the design matrices for each parameter. Prior to the optimization, it also collapses the histories further from 55 to 32 which it can do here because sex is not used in either formula. The final steps are to compute initial values for the parameters and find the MLEs with the selected optimization method(s). As it progresses, the value of -2log-likelihood is reported every 100 evaluations of the objective function. It is not shown above because there were fewer than 100 function evaluations for this example. A brief listing of the model results is obtained by typing model which invokes the function print.crm because class(model)= crm. 3
4 > model crm Model Summary Npar : 2-2lnL: AIC : Beta Estimate Phi.(Intercept) p.(intercept) The output includes the number of parameters (npar), -2log-likelihood, Akaike s Information Criterion (AIC ), and the estimates for φ and p. Estimates of precision are not shown because the default is hessian=false and it must be set to TRUE to get estimates of precision. This default was chosen because the marked package does not count parameters from the hessian, so there is no need to compute it for each model as with MARK. The hessian may never be needed if the model is clearly inferior. Also, this allows the model to be fitted again from the final or different estimates to check for convergence without the penalty of computing the hessian at the final values each time. Separate functions (cjs.hessian and js.hessian) are provided to compute and store in the model the variance-covariance matrix from the hessian at the final estimates as shown below: > model=cjs.hessian(model) > model crm Model Summary Npar : 2-2lnL: AIC : Beta Estimate se lcl ucl Phi.(Intercept) p.(intercept)
5 Once the hessian has been computed, printing the model will display the standard errors and 95% normal confidence intervals for the parameter estimates on the link scale (e.g., logit for φ and p). You can set hessian=true in the call to crm if you want to compute it when the model is fitted. You ll never fit only one model to data, so the most efficient approach is to call process.data and make.design.data separately and pass the results to crm so they can be used for each fitted model as shown below: > dipper.proc=process.data(dipper) > dipper.ddl=make.design.data(dipper.proc) > Phi.sex=list(formula=~sex) > model=crm(dipper.proc,dipper.ddl,model.parameters=list(phi=phi.sex), + accumulate=false) Collapsing capture history records is controlled by the accumulate arguments in process.data and crm. In the above example, accumulate was TRUE for the process.data step but it was turned off for the model fit because it would have not resulted in any accumulation with the sex term in the model. Typically the default values are optimal but if you are fitting many models and most are complex models, then it may save time to accumulate in the process.data step but not for the model fitting step. If you fit more than a few models, use crm.wrapper rather than crm. It fits a set of models and returns a list with a model selection table that summarizes the fit of all the models. By default, crm.wrapper stores the model results externally and in the list it only stores the names of the files containing the models. If you set external=false, then it will store the model results in the list as shown in the example below. > dipper.proc=process.data(dipper) > dipper.ddl=make.design.data(dipper.proc) > fit.models=function() + { + Phi.sex=list(formula=~sex) + Phi.time=list(formula=~time) + p.sex=list(formula=~sex) + p.dot=list(formula=~1) + cml=create.model.list(c("phi","p")) + results=crm.wrapper(cml,data=dipper.proc, ddl=dipper.ddl, + external=false,accumulate=false) 5
6 + return(results) + } > dipper.models=fit.models() The model selection table is displayed with: > dipper.models model npar AIC DeltaAIC weight neg2lnl 1 Phi(~sex)p(~1) Phi(~time)p(~1) Phi(~sex)p(~sex) Phi(~time)p(~sex) convergence A non-zero value for convergence means the model did not converge. If the models are not stored externally, an individual model can be extracted from the list with either the model number which is listed in the model table or with the model name which is the model formula specifications pasted together as shown below: > dipper.models[[1]] crm Model Summary Npar : 3-2lnL: AIC : Beta Estimate Phi.(Intercept) Phi.sexMale p.(intercept) > dipper.models[["phi.sex.p.dot"]] 6
7 crm Model Summary Npar : 3-2lnL: AIC : Beta Estimate Phi.(Intercept) Phi.sexMale p.(intercept) If the models are stored externally, they can be retrieved with the function load.model as shown below: > model=load.model(dipper.models[[1]]) > model crm Model Summary Npar : 3-2lnL: AIC : Beta Estimate Phi.(Intercept) Phi.sexMale p.(intercept) To make the analysis more interesting, we will add some covariates for φ and p. For φ, we will add a static covariate weight which is a random value between 1 and 10. For φ, we also add a time-varying covariate Flood which is the same for all dippers but varies by time with a 0 value for times 1,4,5,6 and a value of 1 for times 2 and 3. For p, we will add a time-varying individual covariate td (trap dependence) which is the 0/1 value of the capture from the previous occasion. Static covariates are entered in the dataframe in a single column and time-varying covariates have a column and name for each occasion with the appropriate time appended at the end of each name. In this case, Flood will be Flood1,...,Flood6 and for td it will be 7
8 td2,...,td7 because time for φ is based on the time at the beginning of the interval and for p it is the time for the capture occasion. Below the names of the covariates in the dataframe are shown after they are created: > data(dipper) > # Add a dummy weight field which are random values from 1 to 10 > set.seed(123) > dipper$weight=round(runif(nrow(dipper),0,9),0)+1 > # Add Flood covariate > Flood=matrix(rep(c(0,1,1,0,0,0),each=nrow(dipper)),ncol=6) > colnames(flood)=paste("flood",1:6,sep="") > dipper=cbind(dipper,flood) > # Add td covariate, but exclude first release as a capture > # splitch and process.ch are functions in the marked package > td=splitch(dipper$ch) > td=td[,1:6] > releaseocc=process.ch(dipper$ch)$first > releaseocc=cbind(1:length(releaseocc),releaseocc) > releaseocc=releaseocc[releaseocc[,2]<nchar(dipper$ch[1]),] > td[releaseocc]=0 > colnames(td)=paste("td",2:7,sep="") > dipper=cbind(dipper,td) > # show names > names(dipper) [1] "ch" "sex" "weight" "Flood1" "Flood2" "Flood3" "Flood4" [8] "Flood5" "Flood6" "td2" "td3" "td4" "td5" "td6" [15] "td7" Next we process the data with the default CJS model and make the design data with parameters that identify which covariates to use for each parameter. By default, each covariate in the dataframe is added to the design data of each parameter but the argument static can be used to limit the variables appended and the time-varying argument is needed to identify which covariates vary by time and should be collapsed into a single column. If we did not include the static argument for φ, then weight and each of the td columns would also be included and for p, weight and each of the Flood columns would be included. If td was added to the time-varying argument for φ and Flood was added for p, then we would have also needed to add td1 and Flood7 due to the different time labels for φ and p. We have no intention to use those 8
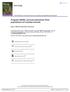
9 variables so it is best to specify what is needed. After specifying the design lists for φ and p, they are used in the call to make.design.data and the resulting dataframe names are shown below: > # Process data > dipper.proc=process.data(dipper) > # Create design data with static and time varying covariates > design.phi=list(static=c("weight"),time.varying=c("flood")) > design.p=list(static=c("sex"),time.varying=c("td"), + age.bins=c(0,1,20)) > design.parameters=list(phi=design.phi,p=design.p) > ddl=make.design.data(dipper.proc,parameters=design.parameters) > names(ddl$phi) > names(ddl$p) Next we define the models for φ and p that we want to fit and call crm. > Phi.sfw=list(formula=~Flood+weight) > p.ast=list(formula=~age+sex+td) > model=crm(dipper.proc,ddl,hessian=true, + model.parameters=list(phi=phi.sfw,p=p.ast)) Below we create a range of data values to compute predicted φ values and then plot the results for Flood and non-flood years for a range of weights. Not surprising that the slope for weight is nearly 0 because the weight values were generated randomly. > newdipper=expand.grid(sex=c("male","female"),weight=1:10,flood1=0, + Flood2=1,Flood3=1,Flood4=0,Flood5=0,Flood6=0,td2=0,td3=c(0,1), + td4=c(0,1),td5=c(0,1),td6=c(0,1),td7=c(0,1)) > reals=predict(model,newdata=newdipper,se=true) > reals$phi$flood=factor(reals$phi$flood,labels=c("non-flood","flood")) > ggplot(reals$phi,aes(weight,estimate,ymin=lcl,ymax=ucl))+ + geom_errorbar(width=0.2)+geom_point()+geom_line()+ + xlab("\nweight")+ylab("survival\n")+facet_grid(flood~.) 9
10 Survival Non flood Flood Weight Fitting an MCMC CJS model (probitcjs) to the dipper data is quite similar to the MLE models with the added arguments burnin and iter to control the number of burnin iterations and the number of iterations to perform after the burnin process. The following fits Phi( Flood)p( sex+time) with low values set for burnin and iter to reduce the vignette build time. > data(dipper) > # Add Flood covariate > Flood=matrix(rep(c(0,1,1,0,0,0),each=nrow(dipper)),ncol=6) > colnames(flood)=paste("flood",1:6,sep="") > dipper=cbind(dipper,flood) > design.parameters=list(phi=list(time.varying="flood")) > model.parameters=list(phi=list(formula=~flood), + p=list(formula=~time+sex)) > MCMCfit=crm(dipper,model="probitCJS", 10
11 + model.parameters=model.parameters, + design.parameters=design.parameters, + burnin=1000,iter=5000) Approximate time till completion: 0.6 minutes 10 % completed 20 % completed 30 % completed 40 % completed 50 % completed 60 % completed 70 % completed 80 % completed 90 % completed 100 % completed Elapsed time in minutes: The results shown are the mode,mean,standard deviation, and 95% highest posterior density interval for the beta parameters and unique φ and p real values. > # beta estimates > MCMCfit crm Model Summary Npar : 9 Beta mode mean sd CI.lower CI.upper Phi.(Intercept) Phi.Flood p.(intercept) p.time p.time p.time p.time p.time p.sexmale
12 > # real estimates > MCMCfit$results$reals $Phi Flood occ mode mean sd CI.lower CI.upper $p time sex occ mode mean sd CI.lower CI.upper 1 7 Female Male Female Male Female Male Female Male Female Male Female Male References Albert, J. and Chib, S. (1993). Bayesian-analysis of binary and polychotomous reponse data. Journal of the American Statistical Association, 88(422): Baillargeon, S. and Rivest, L. P. (2007). Rcapture: Loglinear models for capturerecapture in R. Journal of Statistical Software, 19(5):1 31. Borchers, D. L. and Efford, M. G. (2008). Spatially Explicit Maximum Likelihood Methods for Capture-Recapture Studies. Biometrics, 64(2): Colchero, F., Jones, O. R., and Rebke, M. (2012). BaSTA: an R package for Bayesian estimation of age-specific survival from incomplete mark-recapture/recovery data with covariates. Methods in Ecology and Evolution, 3: Fiske, I. J. and Chandler, R. B. (2011). unmarked : An R Package for fitting hierarchical models of wildlife occurrence and abundance. 43(10):
13 Laake, J. and Rexstad, E. (2008). RMark an alternative approach to building linear models in MARK. In Cooch, E. and White, G. C., editors, Program MARK: A Gentle Introduction. Manly, B. F. J. and Parr, M. J. (1968). A new method of estimating population size, survivorship, and birth rate from recapture data. Transactions of the society for British Entomology, 18: McDonald, T. L., Amstrup, S. C., Regehr, E. V., and Manly, B. F. J. (2005). Examples. In Amstrup, S. C., McDonald, T. L., and Manly, B. F. J., editors, Handbook of capture-recapture analysis, pages Princeton University Press, Princeton, New Jersey USA. Pledger, S., Pollock, K. H., and Norris, J. L. (2003). Open capture-recapture models with heterogeneity: I. Cormack-Jolly-Seber model. Biometrics, 59(4): Royle, J. A., Karanth, K. U., Gopalaswamy, A. M., and Kumar, N. S. (2009). Bayesian inference in camera trapping studies for a class of spatial capturerecapture models. Ecology, 90(11): Schwarz, C. J. and Arnason, A. N. (1996). A general methodology for the analysis of capture-recapture experiments in open populations. Biometrics, 52(3): Schwarz, C. J., Pickard, D., Marine, K., and Bonner., S. J. (2009). Juvenile salmonid outmigrant monitoring evaluation, Phase II, september Unpublished Report prepared for the Trinity River Restoration Program, Weaverville, CA. White, G. C. and Burnham, K. P. (1999). Program MARK: survival estimation from populations of marked animals. Bird Study, 46: Appendix Model construction The manner in which models are constructed in the marked package is different than in MARK. We start by describing how MARK specifies and constructs models and how the marked package differs. The likelihood for CJS/JS is a product multinomial and the multinomial cell probabilities are functions of the parameters. We will focus on CJS which has two types of parameters: p (capture/recapture probability) and 13
14 φ (apparent survival). Each cell is associated with a release cohort and recapture time (occasion). The cohorts can be further split into groups based on categorical variables (e.g., sex). In MARK, a parameter index matrix (PIM) is used to specify the parameters. Each cell can have a unique index for each type of parameter type (e.g., p and φ). As a very simple example, we will consider a CJS model for a single group with k=4 occasions. For this structure the PIMS are triangular with a row for each cohort released at each occasion (time) and the column representing the occasions (times) following the release. Using an unique index for each potential parameter, the PIMS would be: φ p Time Time Cohort Cohort The index 5 represents survival of the second release cohort between occasions 3 and 4 and index 8 represents recapture probability of the first release cohort on the third occasion. A separate set of PIMS is created for each group defined in the data. For this example, if there were g groups there would be g*12 possible unique indices. Some reduced models with constraints on parameters can be constructed by modifying the PIMS. For example, a model with constant survival and capture probability (Phi(.)p(.)) could be specified by setting all of the PIM values to 1 for the φ PIM and 2 for the p PIM. Not all reduced models can be specified by modifying the PIMS and more generally reduced models are constructed with a design matrix which is a set of linear constraints. Each parameter index specifies a row in the design matrix which can be used to apply constraints on the parameters and to relate covariates to the parameters. The PIM approach minimizes the work of manually creating a design matrix by reducing the set of potential parameters and thus the number of rows in the design matrix. For our example, there would be 12 rows in the design matrix and each column in the design matrix (X) represents one of the parameters in the vectorβ. If a logit link is used, the real parameters θ (e.g., φ and p in CJS) are: 1 θ = (1) 1 + exp( Xβ) To specify the Phi(.)p(.) model, the design matrix (X) would have 12 rows and 2 columns. The first column in X would have a 1 in the first six rows (indices 1-6) and 14
15 a 0 in the last six rows (indices 7-12). The second column would have 0 in the first six rows and a 1 in the last 6 rows. The resulting model would have two parameters with the first representing φ and the second p. To automate the creation of design matrices with formula, the RMark package creates design data which are data about the parameters like cohort, time, group, and any related variables. However, having a single X becomes problematic with individual covariates (variables that differ for animals in the same group/cohort). In MARK, these individual covariates are entered into the design matrix as a covariate name. When the real parameters are computed, the name of the covariate is replaced with the value for an animal. If there are only a few individual covariates and small number of animals, the time required for replacement and computation is minor, but when the individual covariates differ in time, there is a different covariate name for each time and most rows in the design matrix must be recomputed which results in longer execution times for large models. Even if computation time is not limiting, we agree with McDonald et al. (2005) that the PIMs used in MARK can be quite confusing to the novice user because they are an additional layer of abstraction from the data. PIMS are useful for manual model creation, but they become an unnecessary nuisance when models and design matrices are created with formula. Explicit PIMS can be avoided by having an underlying real parameter for each animal for each occasion regardless of the release cohort. It can be viewed as a rectangular matrix with a column for each animal and a row for each occasion. That was the solution of McDonald et al. (2005) who in their mra package use a rectangular covariate matrices as predictor for the real parameters as in standard regression. We use a similar approach that we believe is simpler for the user. For a specified model structure (e.g., CJS), the marked package creates from the raw data, a list of dataframes. Each dataframe contains the covariate data for a type of parameter in the model (e.g., φ and p). The dataframes contain a record for each animal-occasion and the columns are the covariates. Each record contains the covariate values for that animal-occasion. If the covariate is constant across time then the value is the same value in each record for an animal and the value may be different for each occasion if the covariate is time-varying. Time-varying covariates must be named in the raw data in a certain manner and specified in the processing step. Each column in the dataframe is a vector that is equivalent to one of the covariate matrices in mra; however, rather than specifying the model as a set of covariate matrices, the standard R model formula (e.g., time+sex) can be used with the dataframe for each parameter which is even closer to a typical regression analysis. A separate dataframe is created for each parameter to allow different data (e.g., 15
16 time values for φ and p) and number of occasions (e.g., φ and p in CJS) for each parameter. For φ and p in the CJS model, there are n(k 1) rows for n animals and k 1 occasions for each parameter but time is labeled by 1 to k 1 for φ and 2 to k for p. For the JS model, there are n(k 1) records for survival probability and entry probability, but for capture probability there are nk records because the initial capture event is modeled. The parameter-specific animal-occasion dataframes are automatically created from the user s data which contain a single record per animal containing the capture history and any covariates. Some covariates like cohort, age, and time are generated automatically for each record. Other static and time-varying covariates specified in the data are added. Static variables (e.g., sex) are repeated for each occasion for each animal. Time-varying covariates are specified in the data using a naming convention of vt where v is the covariate name and t is a numeric value for the time (occasion) (e.g., td19 contains the value of covariate td at time 19). The time-varying covariates are collapsed in the animal-occasion dataframes to a single column with the name identified by v and each record contains the appropriate value for the occasion. All of this is transparent to the user who only needs to specify a dataframe, the type of model (e.g., CJS), the variables that should be treated as time-varying, and the formula for each parameter. With the R function model.matrix, the formula is applied to the dataframe to create the design matrix for each parameter (e.g, X φ,x p ). They are each equivalent to a portion of the design matrix in MARK for an animal which are then stacked on top of one another to make a matrix for all animals. For maximum likelihood estimation (MLE), equation 1 (or similar inverse link function) is applied and the resulting vector is converted to a matrix with n rows and a column for each required occasion (k 1 or k). For Bayesian MCMC inference in marked, onlyxβ are needed for updating. Model fitting Currently there are only three types of models implemented in the marked package: 1) CJS, 2) JS, and 3) probitcjs. The first two are based on MLE and the third is an MCMC implementation as described by (Johnson et al in prep). The likelihoods for CJS and JS were developed hierarchically as described by Pledger et al. (2003) for CJS. For simplicity we only consider a single animal to avoid the additional subscript. Let ω be a capture history vector having value ω j = 1 when the animal was initially captured and released or recaptured at occasion j and ω j = 0 if the animal was not captured on occasion j, for occasions j =1,...k. The probability of 16
17 observing a particular capture history P r(ω) for CJS can be divided into two pieces (P r(ω) = P r(ω 1 )P r(ω 2 ): 1) ω 1 is the portion of the capture history from the initial release (i.e., first 1) to the last occasion it was sighted (i.e., last 1), and 2) ω 2 is from the last 1 to the last occasion. P r(ω 1 ) is easily computed because the time period when the animal was available for capture is known but P r(ω 2 ) is more difficult to compute because it is unknown whether the animal survived until the last occasion, or if it died, when it died. Typically, P r(ω 2 ) has been computed recursively (Nichols 2005). A hierarchical construction is more direct and understandable. As described in Pledger et al. (2003), we let f be the occasion an animal was released and let d be the occasion after which the animal is no longer available to be recaptured due to death or termination of the study. Then P r(ω f) = d P r(ω f, d)p r(d f) where the conditional probability P r(ω f, d) is: P r(ω f, d) = and the departure probability is P r(d f) = d j=f+1 ( d 1 p ω j j (1 p j) (1 ω j) ) φ j (1 φ d ) j=f Viewed in this way, it is possible to construct the likelihood values for all of the observations with a set of matrices as we describe in the Appendix. The hierarchical approach is easily extended to JS. Now the capture history has an additional component with P r(ω) = P r(ω 0 )P r(ω 1 )P r(ω 2 ) where ω 0 is the portion of the capture history vector from the first occasion (j =1) to the occasion the animal was first seen (j=f ). The P r(ω 0 ) is similar to P r(ω 2 ) but now it is e, the occasion at which the animal was first available for capture (i.e., entered in the prior interval) that is unknown. The CJS likelihood provides P r(ω f) so we only need to compute P r(ω 0 ) = f 1 e=0 P r(ω 0 e)π e where π e is the probability of an animal enters the population in the interval between occasion e and e+1 ( k 1 e=1 π e = 1 π 0 ; pent parameters in MARK and specified as β by Schwarz and Arnason (1996)) and ( f 1 ) P r(ω 0 e) = (1 p j ) j=e+1 The JS likelihood also contains a component for animals that entered but were never seen. The details for the JS likelihood construction are provided in the Appendix. 17 p f
18 The MLEs are obtained numerically by finding the minimum of the negative log-likelihood using optimization methods provided through the optimx R package (Nash and ). The default optimization method is BFGS but you can specify alternate methods and several methods used independently or in sequence as described by Nash and (). Initial values for parameters can either be provided as a constant (e.g., 0) or as a vector from the results of a previously fitted similar model. If initial values are not specified then they are computed using general linear models (GLM) that provide approximations. Using the underlying idea in Manly and Parr (1968) we compute initial estimates for capture-probability p for occasions 2 to k 1 using a binomial GLM with the formula for p which is fitted to a sequence of Bernoulli random variables that are a subset of the capture history values y ij i = 1,..., n and j = f i + 1,..., l i 1 where f i and l i are the first and last occasions the i th animal was seen. A similar but more ad-hoc idea is used for φ. We know the animal is alive between f i and l i and assume that the animal dies at occasion l i + 1 k. We use a binomial GLM with the formula for φ fitted to the yij i = 1,..., n and j = f i + 1,..., l i + 1 k where yij=1 for j = f i + 1,..., l i and yij=0 for j = l i + 1 k. For φ and p, a logit link is used for MLE and a probit link for MCMC. For JS, a log link is used for f 0, the number never captured and the estimate of super-population size is the total number of individual animals plus f 0. Also, for JS a multinomial logit link is used for π e. The initial value is set to 0 for any β that is not estimated (e.g., p K in CJS or p 1,π e, N for JS). Likelihood Details Here we provide some details for the likelihoods that are computed in cjs.lnl and cjs.f for the CJS model and in js.lnl for the JS model. Let φ,m p be n k matrices which are functions of the parameters whereφ ij is the survival probability for animal i from occasion j 1 to j (φ i1 = 1 ), m ij is the probability of dying in the interval j to j + 1 (m ik = 1) and p ij is the capture probability for animal i on occasion j (p i1 = 0). In addition we define a series of matrices and vectors computed from the data in the function process.ch. Let C be an n k matrix of the capture history values and f i and l i are the first and last occasions the i th animal was seen. Derived from those values are n k matrices L, F and F + which contain 0 except that L ij = 1 j l i,f ij = 1 j f i and F + ij = 1 j > f i. The likelihood calculation has the following steps where the matrix multiplication is element-wise and 1 and represents an n k matrix where each element is 1: 1. Construct the n k matrix φ = 1 F + + φf + which is the φ matrix modified such that φ ij = 1 for j f i. 18
19 2. Construct the n k matrix φ from φ where φ ij = j i=1 φ ij. 3. Construct the n k matrix p = 1 F + + F + (Cp + (1 C)(1 p)). 4. Construct the n k matrix p from p where p ij = j i=1 p ij. 5. Compute n 1 vector of probabilities of observed capture histories P r(ω i ) i = 1,..., n which are the sums of the rows of LMφ p. 6. Compute the log-likelihood n i=1 ln(p r(ω i)) The R code translates from the mathematics quite literally as: > Phiprime=1-Fplus + Phi*Fplus > Phistar=t(apply(Phiprime,1,cumprod)) > pprime=(1-fplus)+fplus*(c*p+(1-c)*(1-p)) > pstar=t(apply(pprime,1,cumprod)) > pomega=rowsums(l*m*phistar*pstar) > lnl=sum(log(pomega)) While this code is simple it is faster if the apply is replaced with a loop because there are far more rows than occasions or replaced with compiled code which we did with a FORTRAN subroutine (cjs.f called from cjs.lnl). The Jolly-Seber likelihood can be partitioned into 3 components: 1) CJS likelihood for P r(ω 1 )P r(ω 2 ) treating the first 1 as a release, 2) a likelihood component for P r(ω 0 ); entry and first observation, 3) a component for those that entered before each occasion but were never seen. We define an additional n k matrix π which are the entry probabilities into the population (specified as β by Schwarz and Arnason (1996)) with the obvious constraint that k 1 j=0 π ij = 1 and N g g = 1,..., G is the abundance of animals in each of the defined G groups (e.g., male/female) that were in the population at some time (super-population size). N g = n g + fg 0 where n g is the number observed in the group and fg 0 are the estimated number of animals in the group that were never seen. The same hierarchical approach used for CJS can be used for the second component. Construct the capture history probability for a given entry time and then sum over all possible entry times.. The second component is constructed with the following steps: 1. Construct the n k matrix E = (1 p)φ(1 F ) + F, 2. Construct the n k matrix E where E ij = k l=j E il, 19
20 3. Compute n 1 vector of probabilities P r(ω ) which are the sums of the rows of E (1 F + )π multiplied by the vector p ifi i = 1,..., n, 4. Compute the log-likelihood n i=1 ln(p r(ω i)). For the third likelihood component for missed animals, we constructed G k dummy capture histories of all 0 s except for a 1 at the occasion the animals entered. From the CJS portion of the code, we obtained p 0 gj the probability that an animal released in group g on occasion j would never be captured. The final log-likelihood component is: [ G k ] fg 0 ln π g,j 1 (1 p gj )p 0 gj + ln(n g!) ln(fg 0!) g=1 j=1 The JS log-likelihood is the total of the 3 components plus G g=1 k j=1 ln(n gj!) which do not depend on the parameters but is added after optimization in function js, to be consistent with the output from MARK. With the structure we have used, the design matrices can become quite large and available memory could become limiting. For the design matrices, we use sparse matrices with the R package Matrix (citation). In addition, for MLE analysis, we use the following to reduce the required memory: 1. We reduce the data to n n individuals by aggregating records with identical data and using the frequencies (f 1,..., f n ; n = n i=1 f i) in the likelihood. 2. We construct the design matrix X incrementally with a user-specified size of data chunk that is processed at one time. 3. We retain only the rows of X which are unique for the design and including any fixed parameters which can be animal-specific. For Bayesian MCMC inference, we cannot aggregate records but we reduce the required memory by: 1. Eliminating the unused animal-occasion data for occasions prior to and including the release occasion for the animal, and 2. Storing only the unique values of the real parameters which are unique rows of X φ and X p. 20
Data input for secr. Murray Efford May 5, 2010
 Data input for secr Murray Efford May 5, 2010 Data for analysis in secr must be prepared as an object of class capthist which includes both the detector layout and the capture data. The structure of a
Data input for secr Murray Efford May 5, 2010 Data for analysis in secr must be prepared as an object of class capthist which includes both the detector layout and the capture data. The structure of a
Package marked. December 6, 2016
 Version 1.1.13 Date 2016-12-5 Package marked December 6, 2016 Title Mark-Recapture Analysis for Survival and Abundance Estimation Author Jeff Laake , Devin Johnson ,
Version 1.1.13 Date 2016-12-5 Package marked December 6, 2016 Title Mark-Recapture Analysis for Survival and Abundance Estimation Author Jeff Laake , Devin Johnson ,
RMark Workshop Notes 2009
 I. What are MARK and RMark? a. What is MARK and what does it do? MARK is freely available software that was written by Dr. Gary White at Colorado State University (http://welcome.warnercnr.colostate.edu/~gwhite/mark/mark.htm,
I. What are MARK and RMark? a. What is MARK and what does it do? MARK is freely available software that was written by Dr. Gary White at Colorado State University (http://welcome.warnercnr.colostate.edu/~gwhite/mark/mark.htm,
Program MARK: survival estimation from populations of marked animals
 Bird Study ISSN: 0006-657 (Print) 19-6705 (Online) Journal homepage: http://www.tandfonline.com/loi/tbis0 Program MARK: survival estimation from populations of marked animals Gary C. White & Kenneth P.
Bird Study ISSN: 0006-657 (Print) 19-6705 (Online) Journal homepage: http://www.tandfonline.com/loi/tbis0 Program MARK: survival estimation from populations of marked animals Gary C. White & Kenneth P.
Multi-session models in secr 3.1 Murray Efford
 Multi-session models in secr 3.1 Murray Efford 2018-02-23 Contents Input of session-specific data....................................... 1 Detections............................................... 1 Session-specific
Multi-session models in secr 3.1 Murray Efford 2018-02-23 Contents Input of session-specific data....................................... 1 Detections............................................... 1 Session-specific
A. Using the data provided above, calculate the sampling variance and standard error for S for each week s data.
 WILD 502 Lab 1 Estimating Survival when Animal Fates are Known Today s lab will give you hands-on experience with estimating survival rates using logistic regression to estimate the parameters in a variety
WILD 502 Lab 1 Estimating Survival when Animal Fates are Known Today s lab will give you hands-on experience with estimating survival rates using logistic regression to estimate the parameters in a variety
Individual Covariates
 WILD 502 Lab 2 Ŝ from Known-fate Data with Individual Covariates Today s lab presents material that will allow you to handle additional complexity in analysis of survival data. The lab deals with estimation
WILD 502 Lab 2 Ŝ from Known-fate Data with Individual Covariates Today s lab presents material that will allow you to handle additional complexity in analysis of survival data. The lab deals with estimation
Estimation of Item Response Models
 Estimation of Item Response Models Lecture #5 ICPSR Item Response Theory Workshop Lecture #5: 1of 39 The Big Picture of Estimation ESTIMATOR = Maximum Likelihood; Mplus Any questions? answers Lecture #5:
Estimation of Item Response Models Lecture #5 ICPSR Item Response Theory Workshop Lecture #5: 1of 39 The Big Picture of Estimation ESTIMATOR = Maximum Likelihood; Mplus Any questions? answers Lecture #5:
Adjustment for varying effort in secr 3.1
 Adjustment for varying effort in secr 3.1 Murray Efford 2018-02-25 Contents Background 1 Previous methods 2 Linear hazard models 2 Implementation in secr 2 Data entry.................................................
Adjustment for varying effort in secr 3.1 Murray Efford 2018-02-25 Contents Background 1 Previous methods 2 Linear hazard models 2 Implementation in secr 2 Data entry.................................................
Bayesian Time-Stratified-Petersen estimators for abundance. Sampling Protocol. Simon J. Bonner (UBC) Carl James Schwarz (SFU)
 Bayesian Time-Stratified-Petersen estimators for abundance Sampling Protocol Simon J. Bonner (UBC) Carl James Schwarz (SFU) cschwarz@stat.sfu.ca Simple-Petersen or Stratified-Petersen methods are often
Bayesian Time-Stratified-Petersen estimators for abundance Sampling Protocol Simon J. Bonner (UBC) Carl James Schwarz (SFU) cschwarz@stat.sfu.ca Simple-Petersen or Stratified-Petersen methods are often
Data input for secr. Murray Efford Introduction 1. Text formats - general 2. Capture data format 2. Detector layout format 3
 Data input for secr Murray Efford 2017-11-29 Contents Introduction 1 Text formats - general 2 Capture data format 2 Detector layout format 3 Using read.capthist 4 When read.capthist is inadequate... 5
Data input for secr Murray Efford 2017-11-29 Contents Introduction 1 Text formats - general 2 Capture data format 2 Detector layout format 3 Using read.capthist 4 When read.capthist is inadequate... 5
Samuel Coolidge, Dan Simon, Dennis Shasha, Technical Report NYU/CIMS/TR
 Detecting Missing and Spurious Edges in Large, Dense Networks Using Parallel Computing Samuel Coolidge, sam.r.coolidge@gmail.com Dan Simon, des480@nyu.edu Dennis Shasha, shasha@cims.nyu.edu Technical Report
Detecting Missing and Spurious Edges in Large, Dense Networks Using Parallel Computing Samuel Coolidge, sam.r.coolidge@gmail.com Dan Simon, des480@nyu.edu Dennis Shasha, shasha@cims.nyu.edu Technical Report
CHAPTER 1 INTRODUCTION
 Introduction CHAPTER 1 INTRODUCTION Mplus is a statistical modeling program that provides researchers with a flexible tool to analyze their data. Mplus offers researchers a wide choice of models, estimators,
Introduction CHAPTER 1 INTRODUCTION Mplus is a statistical modeling program that provides researchers with a flexible tool to analyze their data. Mplus offers researchers a wide choice of models, estimators,
Generalized least squares (GLS) estimates of the level-2 coefficients,
 Contents 1 Conceptual and Statistical Background for Two-Level Models...7 1.1 The general two-level model... 7 1.1.1 Level-1 model... 8 1.1.2 Level-2 model... 8 1.2 Parameter estimation... 9 1.3 Empirical
Contents 1 Conceptual and Statistical Background for Two-Level Models...7 1.1 The general two-level model... 7 1.1.1 Level-1 model... 8 1.1.2 Level-2 model... 8 1.2 Parameter estimation... 9 1.3 Empirical
Fathom Dynamic Data TM Version 2 Specifications
 Data Sources Fathom Dynamic Data TM Version 2 Specifications Use data from one of the many sample documents that come with Fathom. Enter your own data by typing into a case table. Paste data from other
Data Sources Fathom Dynamic Data TM Version 2 Specifications Use data from one of the many sample documents that come with Fathom. Enter your own data by typing into a case table. Paste data from other
opencr open population capture recapture
 opencr 1.2 - open population capture recapture Murray Efford 2018-05-25 Contents 1 Outline 2 1.1 Model types.............................................. 2 1.2 Data..................................................
opencr 1.2 - open population capture recapture Murray Efford 2018-05-25 Contents 1 Outline 2 1.1 Model types.............................................. 2 1.2 Data..................................................
Assessing the Quality of the Natural Cubic Spline Approximation
 Assessing the Quality of the Natural Cubic Spline Approximation AHMET SEZER ANADOLU UNIVERSITY Department of Statisticss Yunus Emre Kampusu Eskisehir TURKEY ahsst12@yahoo.com Abstract: In large samples,
Assessing the Quality of the Natural Cubic Spline Approximation AHMET SEZER ANADOLU UNIVERSITY Department of Statisticss Yunus Emre Kampusu Eskisehir TURKEY ahsst12@yahoo.com Abstract: In large samples,
Alternative parameterisations of detection parameters in secr 3.0
 Alternative parameterisations of detection parameters in secr 3.0 Murray Efford 2017-04-10 Contents Theory.................................................... 1 Implementation...............................................
Alternative parameterisations of detection parameters in secr 3.0 Murray Efford 2017-04-10 Contents Theory.................................................... 1 Implementation...............................................
GAMs semi-parametric GLMs. Simon Wood Mathematical Sciences, University of Bath, U.K.
 GAMs semi-parametric GLMs Simon Wood Mathematical Sciences, University of Bath, U.K. Generalized linear models, GLM 1. A GLM models a univariate response, y i as g{e(y i )} = X i β where y i Exponential
GAMs semi-parametric GLMs Simon Wood Mathematical Sciences, University of Bath, U.K. Generalized linear models, GLM 1. A GLM models a univariate response, y i as g{e(y i )} = X i β where y i Exponential
Frequently Asked Questions Updated 2006 (TRIM version 3.51) PREPARING DATA & RUNNING TRIM
 Frequently Asked Questions Updated 2006 (TRIM version 3.51) PREPARING DATA & RUNNING TRIM * Which directories are used for input files and output files? See menu-item "Options" and page 22 in the manual.
Frequently Asked Questions Updated 2006 (TRIM version 3.51) PREPARING DATA & RUNNING TRIM * Which directories are used for input files and output files? See menu-item "Options" and page 22 in the manual.
Approximate Bayesian Computation using Auxiliary Models
 Approximate Bayesian Computation using Auxiliary Models Tony Pettitt Co-authors Chris Drovandi, Malcolm Faddy Queensland University of Technology Brisbane MCQMC February 2012 Tony Pettitt () ABC using
Approximate Bayesian Computation using Auxiliary Models Tony Pettitt Co-authors Chris Drovandi, Malcolm Faddy Queensland University of Technology Brisbane MCQMC February 2012 Tony Pettitt () ABC using
Bayesian Time-Stratified-Petersen Estimators for Abundance
 Bayesian Time-Stratified-Petersen Estimators for Abundance Simon Bonner 1 Carl James Schwarz 2 1 Department of Statistics University of Kentucky Lexington, KY 2 Department of Statistics and Actuarial Science
Bayesian Time-Stratified-Petersen Estimators for Abundance Simon Bonner 1 Carl James Schwarz 2 1 Department of Statistics University of Kentucky Lexington, KY 2 Department of Statistics and Actuarial Science
Monte Carlo for Spatial Models
 Monte Carlo for Spatial Models Murali Haran Department of Statistics Penn State University Penn State Computational Science Lectures April 2007 Spatial Models Lots of scientific questions involve analyzing
Monte Carlo for Spatial Models Murali Haran Department of Statistics Penn State University Penn State Computational Science Lectures April 2007 Spatial Models Lots of scientific questions involve analyzing
STATISTICS (STAT) Statistics (STAT) 1
 Statistics (STAT) 1 STATISTICS (STAT) STAT 2013 Elementary Statistics (A) Prerequisites: MATH 1483 or MATH 1513, each with a grade of "C" or better; or an acceptable placement score (see placement.okstate.edu).
Statistics (STAT) 1 STATISTICS (STAT) STAT 2013 Elementary Statistics (A) Prerequisites: MATH 1483 or MATH 1513, each with a grade of "C" or better; or an acceptable placement score (see placement.okstate.edu).
1. Estimation equations for strip transect sampling, using notation consistent with that used to
 Web-based Supplementary Materials for Line Transect Methods for Plant Surveys by S.T. Buckland, D.L. Borchers, A. Johnston, P.A. Henrys and T.A. Marques Web Appendix A. Introduction In this on-line appendix,
Web-based Supplementary Materials for Line Transect Methods for Plant Surveys by S.T. Buckland, D.L. Borchers, A. Johnston, P.A. Henrys and T.A. Marques Web Appendix A. Introduction In this on-line appendix,
Dynamic Thresholding for Image Analysis
 Dynamic Thresholding for Image Analysis Statistical Consulting Report for Edward Chan Clean Energy Research Center University of British Columbia by Libo Lu Department of Statistics University of British
Dynamic Thresholding for Image Analysis Statistical Consulting Report for Edward Chan Clean Energy Research Center University of British Columbia by Libo Lu Department of Statistics University of British
R package mcll for Monte Carlo local likelihood estimation
 R package mcll for Monte Carlo local likelihood estimation Minjeong Jeon University of California, Berkeley Sophia Rabe-Hesketh University of California, Berkeley February 4, 2013 Cari Kaufman University
R package mcll for Monte Carlo local likelihood estimation Minjeong Jeon University of California, Berkeley Sophia Rabe-Hesketh University of California, Berkeley February 4, 2013 Cari Kaufman University
2. A Bernoulli distribution has the following likelihood function for a data set D: N 1 N 1 + N 0
 Machine Learning Fall 2015 Homework 1 Homework must be submitted electronically following the instructions on the course homepage. Make sure to explain you reasoning or show your derivations. Except for
Machine Learning Fall 2015 Homework 1 Homework must be submitted electronically following the instructions on the course homepage. Make sure to explain you reasoning or show your derivations. Except for
Towards the mother of all models: customised construction of the mark recapture likelihood function
 Animal Biodiversity and Conservation 27.1 (2004) 177 Towards the mother of all models: customised construction of the mark recapture likelihood function R. J. Barker & G. C. White Barker, R. J. & White,
Animal Biodiversity and Conservation 27.1 (2004) 177 Towards the mother of all models: customised construction of the mark recapture likelihood function R. J. Barker & G. C. White Barker, R. J. & White,
Show how the LG-Syntax can be generated from a GUI model. Modify the LG-Equations to specify a different LC regression model
 Tutorial #S1: Getting Started with LG-Syntax DemoData = 'conjoint.sav' This tutorial introduces the use of the LG-Syntax module, an add-on to the Advanced version of Latent GOLD. In this tutorial we utilize
Tutorial #S1: Getting Started with LG-Syntax DemoData = 'conjoint.sav' This tutorial introduces the use of the LG-Syntax module, an add-on to the Advanced version of Latent GOLD. In this tutorial we utilize
A GENERAL GIBBS SAMPLING ALGORITHM FOR ANALYZING LINEAR MODELS USING THE SAS SYSTEM
 A GENERAL GIBBS SAMPLING ALGORITHM FOR ANALYZING LINEAR MODELS USING THE SAS SYSTEM Jayawant Mandrekar, Daniel J. Sargent, Paul J. Novotny, Jeff A. Sloan Mayo Clinic, Rochester, MN 55905 ABSTRACT A general
A GENERAL GIBBS SAMPLING ALGORITHM FOR ANALYZING LINEAR MODELS USING THE SAS SYSTEM Jayawant Mandrekar, Daniel J. Sargent, Paul J. Novotny, Jeff A. Sloan Mayo Clinic, Rochester, MN 55905 ABSTRACT A general
Frequencies, Unequal Variance Weights, and Sampling Weights: Similarities and Differences in SAS
 ABSTRACT Paper 1938-2018 Frequencies, Unequal Variance Weights, and Sampling Weights: Similarities and Differences in SAS Robert M. Lucas, Robert M. Lucas Consulting, Fort Collins, CO, USA There is confusion
ABSTRACT Paper 1938-2018 Frequencies, Unequal Variance Weights, and Sampling Weights: Similarities and Differences in SAS Robert M. Lucas, Robert M. Lucas Consulting, Fort Collins, CO, USA There is confusion
Package BaSTA. November 8, 2015
 Type Package Package BaSTA November 8, 2015 Title Age-Specific Survival Analysis from Incomplete Capture-Recapture/Recovery Data Version 1.9.4 Date 2015-11-07 Author Fernando Colchero, Owen Jones, Maren
Type Package Package BaSTA November 8, 2015 Title Age-Specific Survival Analysis from Incomplete Capture-Recapture/Recovery Data Version 1.9.4 Date 2015-11-07 Author Fernando Colchero, Owen Jones, Maren
BeviMed Guide. Daniel Greene
 BeviMed Guide Daniel Greene 1 Introduction BeviMed [1] is a procedure for evaluating the evidence of association between allele configurations across rare variants, typically within a genomic locus, and
BeviMed Guide Daniel Greene 1 Introduction BeviMed [1] is a procedure for evaluating the evidence of association between allele configurations across rare variants, typically within a genomic locus, and
FMA901F: Machine Learning Lecture 3: Linear Models for Regression. Cristian Sminchisescu
 FMA901F: Machine Learning Lecture 3: Linear Models for Regression Cristian Sminchisescu Machine Learning: Frequentist vs. Bayesian In the frequentist setting, we seek a fixed parameter (vector), with value(s)
FMA901F: Machine Learning Lecture 3: Linear Models for Regression Cristian Sminchisescu Machine Learning: Frequentist vs. Bayesian In the frequentist setting, we seek a fixed parameter (vector), with value(s)
Clustering Relational Data using the Infinite Relational Model
 Clustering Relational Data using the Infinite Relational Model Ana Daglis Supervised by: Matthew Ludkin September 4, 2015 Ana Daglis Clustering Data using the Infinite Relational Model September 4, 2015
Clustering Relational Data using the Infinite Relational Model Ana Daglis Supervised by: Matthew Ludkin September 4, 2015 Ana Daglis Clustering Data using the Infinite Relational Model September 4, 2015
Workshop 8: Model selection
 Workshop 8: Model selection Selecting among candidate models requires a criterion for evaluating and comparing models, and a strategy for searching the possibilities. In this workshop we will explore some
Workshop 8: Model selection Selecting among candidate models requires a criterion for evaluating and comparing models, and a strategy for searching the possibilities. In this workshop we will explore some
A noninformative Bayesian approach to small area estimation
 A noninformative Bayesian approach to small area estimation Glen Meeden School of Statistics University of Minnesota Minneapolis, MN 55455 glen@stat.umn.edu September 2001 Revised May 2002 Research supported
A noninformative Bayesian approach to small area estimation Glen Meeden School of Statistics University of Minnesota Minneapolis, MN 55455 glen@stat.umn.edu September 2001 Revised May 2002 Research supported
DENSITY 5.0. Program DENSITY is a Windows application for spatially explicit capture recapture (SECR).
 DENSITY 5.0 Copyright (C) 2007-2014 Murray Efford Program DENSITY is a Windows application for spatially explicit capture recapture (SECR). www.otago.ac.nz/density See also Google group 'secr' This software
DENSITY 5.0 Copyright (C) 2007-2014 Murray Efford Program DENSITY is a Windows application for spatially explicit capture recapture (SECR). www.otago.ac.nz/density See also Google group 'secr' This software
CHAPTER 7 EXAMPLES: MIXTURE MODELING WITH CROSS- SECTIONAL DATA
 Examples: Mixture Modeling With Cross-Sectional Data CHAPTER 7 EXAMPLES: MIXTURE MODELING WITH CROSS- SECTIONAL DATA Mixture modeling refers to modeling with categorical latent variables that represent
Examples: Mixture Modeling With Cross-Sectional Data CHAPTER 7 EXAMPLES: MIXTURE MODELING WITH CROSS- SECTIONAL DATA Mixture modeling refers to modeling with categorical latent variables that represent
PIANOS requirements specifications
 PIANOS requirements specifications Group Linja Helsinki 7th September 2005 Software Engineering Project UNIVERSITY OF HELSINKI Department of Computer Science Course 581260 Software Engineering Project
PIANOS requirements specifications Group Linja Helsinki 7th September 2005 Software Engineering Project UNIVERSITY OF HELSINKI Department of Computer Science Course 581260 Software Engineering Project
MULTISTATE MODELS FOR ECOLOGICAL
 MULTISTATE MODELS FOR ECOLOGICAL DATA RACHEL MCCREA DICE SEMINAR CANTERBURY, OCTOBER 2015 COLLABORATORS Mike Hudson Hannah Worthington Ming Zhou Diana Cole Richard Griffiths Ruth King Eleni Matechou OUTLINE
MULTISTATE MODELS FOR ECOLOGICAL DATA RACHEL MCCREA DICE SEMINAR CANTERBURY, OCTOBER 2015 COLLABORATORS Mike Hudson Hannah Worthington Ming Zhou Diana Cole Richard Griffiths Ruth King Eleni Matechou OUTLINE
Multiple imputation using chained equations: Issues and guidance for practice
 Multiple imputation using chained equations: Issues and guidance for practice Ian R. White, Patrick Royston and Angela M. Wood http://onlinelibrary.wiley.com/doi/10.1002/sim.4067/full By Gabrielle Simoneau
Multiple imputation using chained equations: Issues and guidance for practice Ian R. White, Patrick Royston and Angela M. Wood http://onlinelibrary.wiley.com/doi/10.1002/sim.4067/full By Gabrielle Simoneau
Machine Learning (BSMC-GA 4439) Wenke Liu
 Machine Learning (BSMC-GA 4439) Wenke Liu 01-25-2018 Outline Background Defining proximity Clustering methods Determining number of clusters Other approaches Cluster analysis as unsupervised Learning Unsupervised
Machine Learning (BSMC-GA 4439) Wenke Liu 01-25-2018 Outline Background Defining proximity Clustering methods Determining number of clusters Other approaches Cluster analysis as unsupervised Learning Unsupervised
Inference for Generalized Linear Mixed Models
 Inference for Generalized Linear Mixed Models Christina Knudson, Ph.D. University of St. Thomas October 18, 2018 Reviewing the Linear Model The usual linear model assumptions: responses normally distributed
Inference for Generalized Linear Mixed Models Christina Knudson, Ph.D. University of St. Thomas October 18, 2018 Reviewing the Linear Model The usual linear model assumptions: responses normally distributed
INLA: an introduction
 INLA: an introduction Håvard Rue 1 Norwegian University of Science and Technology Trondheim, Norway May 2009 1 Joint work with S.Martino (Trondheim) and N.Chopin (Paris) Latent Gaussian models Background
INLA: an introduction Håvard Rue 1 Norwegian University of Science and Technology Trondheim, Norway May 2009 1 Joint work with S.Martino (Trondheim) and N.Chopin (Paris) Latent Gaussian models Background
RJaCGH, a package for analysis of
 RJaCGH, a package for analysis of CGH arrays with Reversible Jump MCMC 1. CGH Arrays: Biological problem: Changes in number of DNA copies are associated to cancer activity. Microarray technology: Oscar
RJaCGH, a package for analysis of CGH arrays with Reversible Jump MCMC 1. CGH Arrays: Biological problem: Changes in number of DNA copies are associated to cancer activity. Microarray technology: Oscar
The Rcapture Package
 Type Package The Rcapture Package May 3, 2007 Title Loglinear Models in Capture-Recapture Experiments Version 1.0 Date 2007-05-02 Author Sophie Baillargeon and Louis-Paul Rivest
Type Package The Rcapture Package May 3, 2007 Title Loglinear Models in Capture-Recapture Experiments Version 1.0 Date 2007-05-02 Author Sophie Baillargeon and Louis-Paul Rivest
NCSS Statistical Software
 Chapter 327 Geometric Regression Introduction Geometric regression is a special case of negative binomial regression in which the dispersion parameter is set to one. It is similar to regular multiple regression
Chapter 327 Geometric Regression Introduction Geometric regression is a special case of negative binomial regression in which the dispersion parameter is set to one. It is similar to regular multiple regression
An Efficient Model Selection for Gaussian Mixture Model in a Bayesian Framework
 IEEE SIGNAL PROCESSING LETTERS, VOL. XX, NO. XX, XXX 23 An Efficient Model Selection for Gaussian Mixture Model in a Bayesian Framework Ji Won Yoon arxiv:37.99v [cs.lg] 3 Jul 23 Abstract In order to cluster
IEEE SIGNAL PROCESSING LETTERS, VOL. XX, NO. XX, XXX 23 An Efficient Model Selection for Gaussian Mixture Model in a Bayesian Framework Ji Won Yoon arxiv:37.99v [cs.lg] 3 Jul 23 Abstract In order to cluster
Smoothing Population Size Estimates for Time-Stratified Mark-Recapture. Experiments using Bayesian P-Splines
 Biometrics 000, 000 000 DOI: 000 000 0000 Smoothing Population Size Estimates for Time-Stratified Mark-Recapture Experiments using Bayesian P-Splines Simon J Bonner 1 and Carl J Schwarz 2 1 Department
Biometrics 000, 000 000 DOI: 000 000 0000 Smoothing Population Size Estimates for Time-Stratified Mark-Recapture Experiments using Bayesian P-Splines Simon J Bonner 1 and Carl J Schwarz 2 1 Department
in this course) ˆ Y =time to event, follow-up curtailed: covered under ˆ Missing at random (MAR) a
 Chapter 3 Missing Data 3.1 Types of Missing Data ˆ Missing completely at random (MCAR) ˆ Missing at random (MAR) a ˆ Informative missing (non-ignorable non-response) See 1, 38, 59 for an introduction to
Chapter 3 Missing Data 3.1 Types of Missing Data ˆ Missing completely at random (MCAR) ˆ Missing at random (MAR) a ˆ Informative missing (non-ignorable non-response) See 1, 38, 59 for an introduction to
An Introduction to Model-Fitting with the R package glmm
 An Introduction to Model-Fitting with the R package glmm Christina Knudson November 7, 2017 Contents 1 Introduction 2 2 Formatting the Data 2 3 Fitting the Model 4 4 Reading the Model Summary 6 5 Isolating
An Introduction to Model-Fitting with the R package glmm Christina Knudson November 7, 2017 Contents 1 Introduction 2 2 Formatting the Data 2 3 Fitting the Model 4 4 Reading the Model Summary 6 5 Isolating
Bootstrapping Methods
 Bootstrapping Methods example of a Monte Carlo method these are one Monte Carlo statistical method some Bayesian statistical methods are Monte Carlo we can also simulate models using Monte Carlo methods
Bootstrapping Methods example of a Monte Carlo method these are one Monte Carlo statistical method some Bayesian statistical methods are Monte Carlo we can also simulate models using Monte Carlo methods
STEPHEN WOLFRAM MATHEMATICADO. Fourth Edition WOLFRAM MEDIA CAMBRIDGE UNIVERSITY PRESS
 STEPHEN WOLFRAM MATHEMATICADO OO Fourth Edition WOLFRAM MEDIA CAMBRIDGE UNIVERSITY PRESS Table of Contents XXI a section new for Version 3 a section new for Version 4 a section substantially modified for
STEPHEN WOLFRAM MATHEMATICADO OO Fourth Edition WOLFRAM MEDIA CAMBRIDGE UNIVERSITY PRESS Table of Contents XXI a section new for Version 3 a section new for Version 4 a section substantially modified for
Simulating from the Polya posterior by Glen Meeden, March 06
 1 Introduction Simulating from the Polya posterior by Glen Meeden, glen@stat.umn.edu March 06 The Polya posterior is an objective Bayesian approach to finite population sampling. In its simplest form it
1 Introduction Simulating from the Polya posterior by Glen Meeden, glen@stat.umn.edu March 06 The Polya posterior is an objective Bayesian approach to finite population sampling. In its simplest form it
SampleSize 3.0 USER S MANUAL
 SampleSize 3.0 USER S MANUAL SAMPLE SIZE CALCULATIONS FOR FISH AND WILDLIFE SURVIVAL STUDIES COLUMBIA BASIN RESEARCH SCHOOL OF AQUATIC AND FISHERY SCIENCES UNIVERSITY OF WASHINGTON Program SampleSize
SampleSize 3.0 USER S MANUAL SAMPLE SIZE CALCULATIONS FOR FISH AND WILDLIFE SURVIVAL STUDIES COLUMBIA BASIN RESEARCH SCHOOL OF AQUATIC AND FISHERY SCIENCES UNIVERSITY OF WASHINGTON Program SampleSize
Expectation-Maximization Methods in Population Analysis. Robert J. Bauer, Ph.D. ICON plc.
 Expectation-Maximization Methods in Population Analysis Robert J. Bauer, Ph.D. ICON plc. 1 Objective The objective of this tutorial is to briefly describe the statistical basis of Expectation-Maximization
Expectation-Maximization Methods in Population Analysis Robert J. Bauer, Ph.D. ICON plc. 1 Objective The objective of this tutorial is to briefly describe the statistical basis of Expectation-Maximization
CHAPTER 18. Mark-resight models. Brett McClintock, NOAA National Marine Mammal Laboratory, Seattle, Washington, USA
 CHAPTER 18 Mark-resight models Brett McClintock, NOAA National Marine Mammal Laboratory, Seattle, Washington, USA Mark-resight methods constitute a slightly different type of data than found in traditional
CHAPTER 18 Mark-resight models Brett McClintock, NOAA National Marine Mammal Laboratory, Seattle, Washington, USA Mark-resight methods constitute a slightly different type of data than found in traditional
Regression. Dr. G. Bharadwaja Kumar VIT Chennai
 Regression Dr. G. Bharadwaja Kumar VIT Chennai Introduction Statistical models normally specify how one set of variables, called dependent variables, functionally depend on another set of variables, called
Regression Dr. G. Bharadwaja Kumar VIT Chennai Introduction Statistical models normally specify how one set of variables, called dependent variables, functionally depend on another set of variables, called
ECE521: Week 11, Lecture March 2017: HMM learning/inference. With thanks to Russ Salakhutdinov
 ECE521: Week 11, Lecture 20 27 March 2017: HMM learning/inference With thanks to Russ Salakhutdinov Examples of other perspectives Murphy 17.4 End of Russell & Norvig 15.2 (Artificial Intelligence: A Modern
ECE521: Week 11, Lecture 20 27 March 2017: HMM learning/inference With thanks to Russ Salakhutdinov Examples of other perspectives Murphy 17.4 End of Russell & Norvig 15.2 (Artificial Intelligence: A Modern
Generalized Additive Model
 Generalized Additive Model by Huimin Liu Department of Mathematics and Statistics University of Minnesota Duluth, Duluth, MN 55812 December 2008 Table of Contents Abstract... 2 Chapter 1 Introduction 1.1
Generalized Additive Model by Huimin Liu Department of Mathematics and Statistics University of Minnesota Duluth, Duluth, MN 55812 December 2008 Table of Contents Abstract... 2 Chapter 1 Introduction 1.1
Zero-Inflated Poisson Regression
 Chapter 329 Zero-Inflated Poisson Regression Introduction The zero-inflated Poisson (ZIP) regression is used for count data that exhibit overdispersion and excess zeros. The data distribution combines
Chapter 329 Zero-Inflated Poisson Regression Introduction The zero-inflated Poisson (ZIP) regression is used for count data that exhibit overdispersion and excess zeros. The data distribution combines
Supplementary Notes on Multiple Imputation. Stephen du Toit and Gerhard Mels Scientific Software International
 Supplementary Notes on Multiple Imputation. Stephen du Toit and Gerhard Mels Scientific Software International Part A: Comparison with FIML in the case of normal data. Stephen du Toit Multivariate data
Supplementary Notes on Multiple Imputation. Stephen du Toit and Gerhard Mels Scientific Software International Part A: Comparison with FIML in the case of normal data. Stephen du Toit Multivariate data
Acknowledgments. Acronyms
 Acknowledgments Preface Acronyms xi xiii xv 1 Basic Tools 1 1.1 Goals of inference 1 1.1.1 Population or process? 1 1.1.2 Probability samples 2 1.1.3 Sampling weights 3 1.1.4 Design effects. 5 1.2 An introduction
Acknowledgments Preface Acronyms xi xiii xv 1 Basic Tools 1 1.1 Goals of inference 1 1.1.1 Population or process? 1 1.1.2 Probability samples 2 1.1.3 Sampling weights 3 1.1.4 Design effects. 5 1.2 An introduction
Package RWildbook. April 6, 2018
 Type Package Package RWildbook April 6, 2018 Title Interface for the 'Wildbook' Wildlife Data Management Framework Version 0.9.3 Provides an interface with the 'Wildbook' mark-recapture ecological database
Type Package Package RWildbook April 6, 2018 Title Interface for the 'Wildbook' Wildlife Data Management Framework Version 0.9.3 Provides an interface with the 'Wildbook' mark-recapture ecological database
Project Report: "Bayesian Spam Filter"
 Humboldt-Universität zu Berlin Lehrstuhl für Maschinelles Lernen Sommersemester 2016 Maschinelles Lernen 1 Project Report: "Bayesian E-Mail Spam Filter" The Bayesians Sabine Bertram, Carolina Gumuljo,
Humboldt-Universität zu Berlin Lehrstuhl für Maschinelles Lernen Sommersemester 2016 Maschinelles Lernen 1 Project Report: "Bayesian E-Mail Spam Filter" The Bayesians Sabine Bertram, Carolina Gumuljo,
Stata goes BUGS (via R)
 (via R) Department of Political Science I University of Mannheim 31. March 2006 4th German Stata Users Group press ctrl + l to start presentation Problem 1 Problem You are a Stata user... and have a complicated
(via R) Department of Political Science I University of Mannheim 31. March 2006 4th German Stata Users Group press ctrl + l to start presentation Problem 1 Problem You are a Stata user... and have a complicated
International Graduate School of Genetic and Molecular Epidemiology (GAME) Computing Notes and Introduction to Stata
 International Graduate School of Genetic and Molecular Epidemiology (GAME) Computing Notes and Introduction to Stata Paul Dickman September 2003 1 A brief introduction to Stata Starting the Stata program
International Graduate School of Genetic and Molecular Epidemiology (GAME) Computing Notes and Introduction to Stata Paul Dickman September 2003 1 A brief introduction to Stata Starting the Stata program
Package GWRM. R topics documented: July 31, Type Package
 Type Package Package GWRM July 31, 2017 Title Generalized Waring Regression Model for Count Data Version 2.1.0.3 Date 2017-07-18 Maintainer Antonio Jose Saez-Castillo Depends R (>= 3.0.0)
Type Package Package GWRM July 31, 2017 Title Generalized Waring Regression Model for Count Data Version 2.1.0.3 Date 2017-07-18 Maintainer Antonio Jose Saez-Castillo Depends R (>= 3.0.0)
Discovery of the Source of Contaminant Release
 Discovery of the Source of Contaminant Release Devina Sanjaya 1 Henry Qin Introduction Computer ability to model contaminant release events and predict the source of release in real time is crucial in
Discovery of the Source of Contaminant Release Devina Sanjaya 1 Henry Qin Introduction Computer ability to model contaminant release events and predict the source of release in real time is crucial in
Introducing Categorical Data/Variables (pp )
 Notation: Means pencil-and-paper QUIZ Means coding QUIZ Definition: Feature Engineering (FE) = the process of transforming the data to an optimal representation for a given application. Scaling (see Chs.
Notation: Means pencil-and-paper QUIZ Means coding QUIZ Definition: Feature Engineering (FE) = the process of transforming the data to an optimal representation for a given application. Scaling (see Chs.
A New Method for Estimation of Resource Selection Probability Function
 Tools and Technology Article A New Method for Estimation of Resource Selection Probability Function SUBHASH R. LELE, 1 Department of Mathematical and Statistical Sciences, University of Alberta, Edmonton,
Tools and Technology Article A New Method for Estimation of Resource Selection Probability Function SUBHASH R. LELE, 1 Department of Mathematical and Statistical Sciences, University of Alberta, Edmonton,
Univariate Extreme Value Analysis. 1 Block Maxima. Practice problems using the extremes ( 2.0 5) package. 1. Pearson Type III distribution
 Univariate Extreme Value Analysis Practice problems using the extremes ( 2.0 5) package. 1 Block Maxima 1. Pearson Type III distribution (a) Simulate 100 maxima from samples of size 1000 from the gamma
Univariate Extreme Value Analysis Practice problems using the extremes ( 2.0 5) package. 1 Block Maxima 1. Pearson Type III distribution (a) Simulate 100 maxima from samples of size 1000 from the gamma
Monte Carlo Methods and Statistical Computing: My Personal E
 Monte Carlo Methods and Statistical Computing: My Personal Experience Department of Mathematics & Statistics Indian Institute of Technology Kanpur November 29, 2014 Outline Preface 1 Preface 2 3 4 5 6
Monte Carlo Methods and Statistical Computing: My Personal Experience Department of Mathematics & Statistics Indian Institute of Technology Kanpur November 29, 2014 Outline Preface 1 Preface 2 3 4 5 6
MODEL SELECTION AND MODEL AVERAGING IN THE PRESENCE OF MISSING VALUES
 UNIVERSITY OF GLASGOW MODEL SELECTION AND MODEL AVERAGING IN THE PRESENCE OF MISSING VALUES by KHUNESWARI GOPAL PILLAY A thesis submitted in partial fulfillment for the degree of Doctor of Philosophy in
UNIVERSITY OF GLASGOW MODEL SELECTION AND MODEL AVERAGING IN THE PRESENCE OF MISSING VALUES by KHUNESWARI GOPAL PILLAY A thesis submitted in partial fulfillment for the degree of Doctor of Philosophy in
Strategies for Modeling Two Categorical Variables with Multiple Category Choices
 003 Joint Statistical Meetings - Section on Survey Research Methods Strategies for Modeling Two Categorical Variables with Multiple Category Choices Christopher R. Bilder Department of Statistics, University
003 Joint Statistical Meetings - Section on Survey Research Methods Strategies for Modeling Two Categorical Variables with Multiple Category Choices Christopher R. Bilder Department of Statistics, University
Computer vision: models, learning and inference. Chapter 10 Graphical Models
 Computer vision: models, learning and inference Chapter 10 Graphical Models Independence Two variables x 1 and x 2 are independent if their joint probability distribution factorizes as Pr(x 1, x 2 )=Pr(x
Computer vision: models, learning and inference Chapter 10 Graphical Models Independence Two variables x 1 and x 2 are independent if their joint probability distribution factorizes as Pr(x 1, x 2 )=Pr(x
Package bayescl. April 14, 2017
 Package bayescl April 14, 2017 Version 0.0.1 Date 2017-04-10 Title Bayesian Inference on a GPU using OpenCL Author Rok Cesnovar, Erik Strumbelj Maintainer Rok Cesnovar Description
Package bayescl April 14, 2017 Version 0.0.1 Date 2017-04-10 Title Bayesian Inference on a GPU using OpenCL Author Rok Cesnovar, Erik Strumbelj Maintainer Rok Cesnovar Description
3.6 Sample code: yrbs_data <- read.spss("yrbs07.sav",to.data.frame=true)
 InJanuary2009,CDCproducedareportSoftwareforAnalyisofYRBSdata, describingtheuseofsas,sudaan,stata,spss,andepiinfoforanalyzingdatafrom theyouthriskbehaviorssurvey. ThisreportprovidesthesameinformationforRandthesurveypackage.Thetextof
InJanuary2009,CDCproducedareportSoftwareforAnalyisofYRBSdata, describingtheuseofsas,sudaan,stata,spss,andepiinfoforanalyzingdatafrom theyouthriskbehaviorssurvey. ThisreportprovidesthesameinformationforRandthesurveypackage.Thetextof
Chapter 15 Mixed Models. Chapter Table of Contents. Introduction Split Plot Experiment Clustered Data References...
 Chapter 15 Mixed Models Chapter Table of Contents Introduction...309 Split Plot Experiment...311 Clustered Data...320 References...326 308 Chapter 15. Mixed Models Chapter 15 Mixed Models Introduction
Chapter 15 Mixed Models Chapter Table of Contents Introduction...309 Split Plot Experiment...311 Clustered Data...320 References...326 308 Chapter 15. Mixed Models Chapter 15 Mixed Models Introduction
Mixture Models and the EM Algorithm
 Mixture Models and the EM Algorithm Padhraic Smyth, Department of Computer Science University of California, Irvine c 2017 1 Finite Mixture Models Say we have a data set D = {x 1,..., x N } where x i is
Mixture Models and the EM Algorithm Padhraic Smyth, Department of Computer Science University of California, Irvine c 2017 1 Finite Mixture Models Say we have a data set D = {x 1,..., x N } where x i is
Chapter 7: Dual Modeling in the Presence of Constant Variance
 Chapter 7: Dual Modeling in the Presence of Constant Variance 7.A Introduction An underlying premise of regression analysis is that a given response variable changes systematically and smoothly due to
Chapter 7: Dual Modeling in the Presence of Constant Variance 7.A Introduction An underlying premise of regression analysis is that a given response variable changes systematically and smoothly due to
MCMC Methods for data modeling
 MCMC Methods for data modeling Kenneth Scerri Department of Automatic Control and Systems Engineering Introduction 1. Symposium on Data Modelling 2. Outline: a. Definition and uses of MCMC b. MCMC algorithms
MCMC Methods for data modeling Kenneth Scerri Department of Automatic Control and Systems Engineering Introduction 1. Symposium on Data Modelling 2. Outline: a. Definition and uses of MCMC b. MCMC algorithms
Bayesian model selection and diagnostics
 Bayesian model selection and diagnostics A typical Bayesian analysis compares a handful of models. Example 1: Consider the spline model for the motorcycle data, how many basis functions? Example 2: Consider
Bayesian model selection and diagnostics A typical Bayesian analysis compares a handful of models. Example 1: Consider the spline model for the motorcycle data, how many basis functions? Example 2: Consider
Analysis of Incomplete Multivariate Data
 Analysis of Incomplete Multivariate Data J. L. Schafer Department of Statistics The Pennsylvania State University USA CHAPMAN & HALL/CRC A CR.C Press Company Boca Raton London New York Washington, D.C.
Analysis of Incomplete Multivariate Data J. L. Schafer Department of Statistics The Pennsylvania State University USA CHAPMAN & HALL/CRC A CR.C Press Company Boca Raton London New York Washington, D.C.
The Cross-Entropy Method for Mathematical Programming
 The Cross-Entropy Method for Mathematical Programming Dirk P. Kroese Reuven Y. Rubinstein Department of Mathematics, The University of Queensland, Australia Faculty of Industrial Engineering and Management,
The Cross-Entropy Method for Mathematical Programming Dirk P. Kroese Reuven Y. Rubinstein Department of Mathematics, The University of Queensland, Australia Faculty of Industrial Engineering and Management,
Package mcemglm. November 29, 2015
 Type Package Package mcemglm November 29, 2015 Title Maximum Likelihood Estimation for Generalized Linear Mixed Models Version 1.1 Date 2015-11-28 Author Felipe Acosta Archila Maintainer Maximum likelihood
Type Package Package mcemglm November 29, 2015 Title Maximum Likelihood Estimation for Generalized Linear Mixed Models Version 1.1 Date 2015-11-28 Author Felipe Acosta Archila Maintainer Maximum likelihood
Multiple-imputation analysis using Stata s mi command
 Multiple-imputation analysis using Stata s mi command Yulia Marchenko Senior Statistician StataCorp LP 2009 UK Stata Users Group Meeting Yulia Marchenko (StataCorp) Multiple-imputation analysis using mi
Multiple-imputation analysis using Stata s mi command Yulia Marchenko Senior Statistician StataCorp LP 2009 UK Stata Users Group Meeting Yulia Marchenko (StataCorp) Multiple-imputation analysis using mi
Correctly Compute Complex Samples Statistics
 SPSS Complex Samples 15.0 Specifications Correctly Compute Complex Samples Statistics When you conduct sample surveys, use a statistics package dedicated to producing correct estimates for complex sample
SPSS Complex Samples 15.0 Specifications Correctly Compute Complex Samples Statistics When you conduct sample surveys, use a statistics package dedicated to producing correct estimates for complex sample
Ludwig Fahrmeir Gerhard Tute. Statistical odelling Based on Generalized Linear Model. íecond Edition. . Springer
 Ludwig Fahrmeir Gerhard Tute Statistical odelling Based on Generalized Linear Model íecond Edition. Springer Preface to the Second Edition Preface to the First Edition List of Examples List of Figures
Ludwig Fahrmeir Gerhard Tute Statistical odelling Based on Generalized Linear Model íecond Edition. Springer Preface to the Second Edition Preface to the First Edition List of Examples List of Figures
Edge-exchangeable graphs and sparsity
 Edge-exchangeable graphs and sparsity Tamara Broderick Department of EECS Massachusetts Institute of Technology tbroderick@csail.mit.edu Diana Cai Department of Statistics University of Chicago dcai@uchicago.edu
Edge-exchangeable graphs and sparsity Tamara Broderick Department of EECS Massachusetts Institute of Technology tbroderick@csail.mit.edu Diana Cai Department of Statistics University of Chicago dcai@uchicago.edu
The GLMMGibbs Package
 The GLMMGibbs Package April 22, 2002 Version 0.5-1 Author Jonathan Myles and David Clayton Maintainer Jonathan Myles Depends R (>= 1.0) Date 2001/22/01 Title
The GLMMGibbs Package April 22, 2002 Version 0.5-1 Author Jonathan Myles and David Clayton Maintainer Jonathan Myles Depends R (>= 1.0) Date 2001/22/01 Title
Machine Learning (BSMC-GA 4439) Wenke Liu
 Machine Learning (BSMC-GA 4439) Wenke Liu 01-31-017 Outline Background Defining proximity Clustering methods Determining number of clusters Comparing two solutions Cluster analysis as unsupervised Learning
Machine Learning (BSMC-GA 4439) Wenke Liu 01-31-017 Outline Background Defining proximity Clustering methods Determining number of clusters Comparing two solutions Cluster analysis as unsupervised Learning
Convexization in Markov Chain Monte Carlo
 in Markov Chain Monte Carlo 1 IBM T. J. Watson Yorktown Heights, NY 2 Department of Aerospace Engineering Technion, Israel August 23, 2011 Problem Statement MCMC processes in general are governed by non
in Markov Chain Monte Carlo 1 IBM T. J. Watson Yorktown Heights, NY 2 Department of Aerospace Engineering Technion, Israel August 23, 2011 Problem Statement MCMC processes in general are governed by non
Package survivalmpl. December 11, 2017
 Package survivalmpl December 11, 2017 Title Penalised Maximum Likelihood for Survival Analysis Models Version 0.2 Date 2017-10-13 Author Dominique-Laurent Couturier, Jun Ma, Stephane Heritier, Maurizio
Package survivalmpl December 11, 2017 Title Penalised Maximum Likelihood for Survival Analysis Models Version 0.2 Date 2017-10-13 Author Dominique-Laurent Couturier, Jun Ma, Stephane Heritier, Maurizio
CS281 Section 9: Graph Models and Practical MCMC
 CS281 Section 9: Graph Models and Practical MCMC Scott Linderman November 11, 213 Now that we have a few MCMC inference algorithms in our toolbox, let s try them out on some random graph models. Graphs
CS281 Section 9: Graph Models and Practical MCMC Scott Linderman November 11, 213 Now that we have a few MCMC inference algorithms in our toolbox, let s try them out on some random graph models. Graphs
Markov Network Structure Learning
 Markov Network Structure Learning Sargur Srihari srihari@cedar.buffalo.edu 1 Topics Structure Learning as model selection Structure Learning using Independence Tests Score based Learning: Hypothesis Spaces
Markov Network Structure Learning Sargur Srihari srihari@cedar.buffalo.edu 1 Topics Structure Learning as model selection Structure Learning using Independence Tests Score based Learning: Hypothesis Spaces
Computational Statistics The basics of maximum likelihood estimation, Bayesian estimation, object recognitions
 Computational Statistics The basics of maximum likelihood estimation, Bayesian estimation, object recognitions Thomas Giraud Simon Chabot October 12, 2013 Contents 1 Discriminant analysis 3 1.1 Main idea................................
Computational Statistics The basics of maximum likelihood estimation, Bayesian estimation, object recognitions Thomas Giraud Simon Chabot October 12, 2013 Contents 1 Discriminant analysis 3 1.1 Main idea................................
Modeling Criminal Careers as Departures From a Unimodal Population Age-Crime Curve: The Case of Marijuana Use
 Modeling Criminal Careers as Departures From a Unimodal Population Curve: The Case of Marijuana Use Donatello Telesca, Elena A. Erosheva, Derek A. Kreader, & Ross Matsueda April 15, 2014 extends Telesca
Modeling Criminal Careers as Departures From a Unimodal Population Curve: The Case of Marijuana Use Donatello Telesca, Elena A. Erosheva, Derek A. Kreader, & Ross Matsueda April 15, 2014 extends Telesca
