Breast Cancer Detection and Diagnosis in Dynamic Contrast-Enhanced Magnetic Resonance Imaging
|
|
|
- Kimberly Lane
- 6 years ago
- Views:
Transcription
1 Breast Cancer Detection and Diagnosis in Dynamic Contrast-Enhanced Magnetic Resonance Imaging Xi Liang Submitted in total fulfilment of the requirements of the degree of Doctor of Philosophy August 2013 Department of Computing and Information Systems University of Melbourne Produced on archival quality paper
2
3 Abstract Dynamic contrast-enhanced magnetic resonance imaging (DCE-MRI) of the breast is a medical imaging tool used to detect and diagnose breast disease. A DCE-MR image is a series of three-dimensional (3D) breast MRI scans. It is acquired to form a 4D image (3D spatial + time), before and after the injection of paramagnetic contrast agents. DCE-MRI allows the analysis of the intensity variation of magnetic resonance (MR) signals, before and after the injection of contrast agents over time. The interpretation of 4D DCE-MRI images can be time consuming due to the amount of information involved. Motion artifacts in between the image scans further complicate the diagnosis. A DCE-MR image includes a large amount of data and it is challenging to interpret even for an experienced radiologist. Therefore, a computer-aided diagnosis (CAD) system is desirable in assisting the diagnosis of abnormal findings in the DCE-MR image. We propose a fully automated CAD system that is comprised of five novel components: a new image registration method to recover motions in between MR image acquisitions, a novel lesion detection method to identify all suspicious regions, a new lesion segmentation method to draw lesion contours and a novel lesion feature characterization method. We then classify the automatically detected lesions using our proposed features. The following lists the challenges found in most CAD systems and the contributions in our CAD system of breast DCE-MRI. 1. Image registration. One challenge in the interpretation of DCE- MRI is motion artifacts which cause the pattern of tissue enhancement to be unreliable. Image registration is used to recover rigid and nonrigid motions between the 3D image sequences in a 4D breast
4 DCE-MRI. Most existing b-spline based registration methods require lesion segmentation in breast DCE-MRI to preserve the lesion volume before performing the registration. An automatic regularization coefficients generation method is proposed in the b-spline based registration of the breast DCE-MRI, where the tumor regions are transformed in a rigid fashion. Our method does not perform lesion segmentation but computes a map to reflect the tissue rigidity. In the evaluation of our proposed coefficients, the registration methods using our coefficients for rigidity terms are compared against manually assigned coefficients of the rigidity terms and smoothness terms. The evaluation is performed on 30 synthetic and 40 clinical pairs of preand post-contrast MRI scans. The results show that the tumor volumes can be well-preserved by using a rigidity term (2.25±4.48% of volume changes) compared to a smoothness term (22.47% ± 20.1%). In our dataset, the volume preservation performance by using our automatically generated coefficients is comparable to the manually assigned rigidity coefficients (2.29% ± 13.25%), and show no significant difference in volume changes (p > 0.05). 2. Lesion detection. After the motions have been corrected by our registration method, we locate the region of interest (ROI) using our lesion detection method. The aim is to highlight the suspicious ROIs to reduce the ROI searching time and the possibility of overlooking small regions by radiologists. A low signal-to-noise ratio is a general challenge in lesion detection of MRI. In addition, the value ranges of a feature of normal tissue in a patient can overlap with that of malignant tissue in another patient, e.g. tissue intensity values, enhancement et al.. Most existing lesion detection methods face the problem of high false positive rate due to blood vessels or motion artifacts. In our method, we locate suspicious lesions by applying a threshold on essential features. The features are normalized to reduce the variation between patients. We then exclude blood vessel or motion artifacts from the initial results by applying filters that can differentiate them from other tissues. In the evaluation of the system
5 on 21 patients with 50 lesions, all were successfully detected with 5.04 false positive regions per breast. 3. Lesion segmentation. One of the main challenges of existing lesion segmentation methods in breast DCE-MRI is that they require the size of the ROI that encloses a lesion to be small in order to successfully segment the lesion. We propose a lesion segmentation method based on naive Bayes and Markov random field. Our method also requires a ROI generated by a user, but the method is not sensitive to the size of the ROI. In our method, the ROI selected in a DCE-MR image is modeled as a connected graph with local Markov properties where each voxel of the image is regarded as a node. Three edge potentials of the graph are proposed to encourage the smoothness of the segmented regions. In the validation on 72 lesions, our method performs better than a baseline fuzzy-c-means method and another closely related method in segmenting lesions in breast MRI by showing a higher overlap with the ground truth. 4. Feature analysis and lesion classification. The challenge of feature analysis in breast DCE-MRI is that different types of lesions can share similar features. In our study, we extract various morphological, textural and kinetic features of the lesions and apply three classifiers to label them. In the morphological feature analysis, we propose minimum volume enclosing ellipsoid (MVEE) based features to measure the similarity of between a lesion and its MVEE. In statistical testing on 72 lesion, the MVEE-based features are significant in differentiating malignant from benign lesions. 5. CAD applications. The proposed CAD system is versatile. We show two scenarios in which a radiologist makes use of the system. In the first scenario, a user selects a rectangular region of interest (ROI) as input and the CAD automatically localizes and classifies the lesion in the ROI as benign or malignant. In another scenario, the CAD system acts as a second reader which fully and automatically identifies all malignant regions. At the time of writing, this is the first automated
6 CAD system that is capable carrying out all these processes without any human interaction. In this thesis, we evaluated the proposed image registration, lesion detection, lesion segmentation, feature extraction and lesion classification using a relatively small database which makes conclusions on generalizability difficult. In the future work, the system requires clinical testing on a large dataset in order to advance this breast MRI CAD to reduce the image interpretation time, eliminate unnecessary biopsy and improve the cancer identification sensitivity for radiologists.
7 To my family, who support me in everything that I do.
8 Declaration This is to certify that the thesis comprises only my original work towards the PhD. due acknowledgement has been made in the text to all other material used, the thesis is fewer than 100,000 words in length, exclusive of tables, maps, bibliographies and appendices. Xi Liang
9 Acknowledgements I cannot expressed enough my deeply heartfelt gratitude to my principal adviser, Professor Rao Kotagiri. He is the funniest adviser and one of the smartest people I know. He has the magic to find the solution to problems that seems impossible to solve. Rao was and remains my best role model for a scientist, mentor and teacher. I hope that I can be as enthusiastic and energetic as Rao. I thank him for keeping a sense of humor at those times I lost mine and for keeping the office door forever open. And I thank him for cheering me up through all the tough periods and for supporting me in taking risks in scientific research during my PhD. I would also like to thank Dr. Qing Yang who saw the need for the fully automated computer-aided diagnosis tool for breast MRI. He also generously provided his company s software so I could perform scientific research. I would like to thank Dr. Helen Frazer who was my clinical adviser. She has helped me immensely in my research and graded many medical images in many precious hours as a hospital doctor. I would like to thank my PhD committee members: Prof. James Bailey and AProf. Andrew Turpin for their helpful comments and suggestions. National ICT Australia (NICTA) funded this thesis and I would like to thank them for the generous support in providing a living allowance, and conference and student exchange traveling grants. I would like to thank all the friends who supported me in my long PhD journey. I would run out the pages if I try to list them all, so I will name just a few of them. I would like to thank Dana for being my mentor in the first year of my PhD. I am grateful to Andrey for being a supportive and caring friend who listened to all my problems in scientific research with a big smile, especially in the tough second year of my PhD. I would like
10 to thank Uyen, Juan, Pallab, Weijie and Lei for all the fruitful discussions over medical imaging and I am very grateful to Goce and Vajay for all their supports in the IEEE student community. I would like to thank Mahtab and Miriam for the discussions about female scientists and engineers. I would also like to thank Saurab, Tom, Naomi and Jeffery for the numerous reviews of my publications and thesis. Last but not least, I would like to give deep felt thanks to Jessie, Chris, Suzie and Jolie for all the joy and support that you brought and bring to me. I love you. The last thank-you is to my mother Qinrong and my partner Haiming. There are no words to describe my gratitude and love for you.
11 Publications Xi Liang, Kotagiri Ramamohanara, Helen Frazer. A lesion detection system in breast DCE-MRI. (Submitted, 2014). (Chapter 4) Xi Liang, Kotagiri Ramamohanara, Helen Frazer, and Qing Yang. Lesion segmentation in dynamic contrast enhanced MRI of breast. In tje International Conference on Digital Image Computing Techniques and Applications (DICTA), pages 1-8. IEEE, (Chapter 5) Xi Liang, Kotagiri Ramamohanarao, Helen Frazer, and Qing Yang. A lesion shape and margin characterization method in dynamic contrast enhanced magnetic resonance imaging of breast. In the 9th IEEE International Symposium on Biomedical Imaging (ISBI), pages IEEE, (Chapter 6) Xi Liang, Kotagiri Ramamohanarao, Qing Yang, Alexander Pitman, and Marious Staring. A practical method to compute coefficients for regularization term in nonrigid registration of DCE-MRI. In the 20th Annual Meeting and Exhibition of the International Society for Magnetic Resonance in Medicine, page 3035, (Chapter 3) Xi Liang, Kotagiri Ramamohanarao, Qing Yang, Alexander Pitman, and Marius Staring. Generating regularization coefficients in nonrigid registration of DCE-MRI of breast. Workshop on Breast Image Analysis, pages 1-8, (Chapter 3)
12 Contents Contents List of Figures List of Tables x xiv xviii 1 Introduction Magnetic resonance imaging Excitement and relaxation Spatial encoding Contrast agents Dynamic contrast-enhanced MRI of breast BI-RADS R lexicon Computer-aided diagnosis (CAD) system and our CAD system Image registration Lesion localization Feature analysis and classification Versatility of CAD system Thesis structure DCE-MRI Database Imaging protocol Lesion statistics Breast registration Introduction x
13 CONTENTS 3.2 Methodology Rigid transformation Nonrigid transformation Similarity metric Regularization term Choice of fixed and moving images Experiments and results Real and synthetic images Evaluation method Registration settings Results Summary Lesion detection Introduction Methodology Pre-processing Initial lesion identification False positive reduction Performance analysis Experiment and results Discussion Summary Lesion segmentation Introduction Methodology Feature extraction Feature normalization Markov random field Experiments Materials Segmentation schemes xi
14 CONTENTS Feature selection and normalization Evaluation method Results Summary Feature analysis and lesion classification Introduction Kinetic features Pharmacokinetic model Three-time points (3TP) model Time-intensity curve based kinetic features Extract representative region to generate curve Morphological features Textural features Classification Methodology Kinetic features Textural features Morphological features Classifier Experiments and results Feature significance test Classification schemes ANN settings ANN classification results RF settings RF classification results Summary Computer-aided diagnosis (CAD) system Introduction Scenario ANN settings xii
15 CONTENTS ANN classification results RF settings RF classification results Scenario ANN settings ANN classification results RF settings RF classification results Summary Conclusions Contributions and limitations Image registration Lesion detection Lesion segmentation Lesion feature analysis and classifications Future work BI-RADS descriptor mapping Analysis on non-mass lesions Lesion retrieval Evaluation on generalizability Appendix 141 Index 144 Bibliography 146 xiii
16 List of Figures H atom behaves like a minuscule magnet, called a nuclear spin Alignment of spins and the generated net magnetization A RF pulse flips the net magnetization 90 from the longitudinal axis to the transverse plane The net-magnetization regains along Z-axis and finally aligns with Z-axis Illustration of T 1 and T 2 relaxation time A fat-suppressed T 1 weighted image and a normal T 2 weighted image Slice selection in z-axis, phase encoding in the y-axis and frequency encoding in the x-axis A slice of an MR image with and without contrast agent Pre-contrast, post-contrast and subtraction MR images Initial and delayed enhancement patterns defined in BI-RADS R A malignant lesion and its time-intensity curve A benign lesion and its time-intensity curve A lesion with a spiculated margin and irregular shape and a lesion with a heterogeneous internal enhancement pattern The general work flow of a CAD system The work flow of the proposed CAD system Maximum intensity projection (MIP) of a subtraction image of a postfrom pre-contrast image Lesion volume (cm 3 ) distribution Image registration framework Illustration of a rigid and a nonrigid registrations xiv
17 LIST OF FIGURES 3.3 The workflow in computing a tissue stiffness map as regularization coefficients Demonstrative slices of three real images that are used to generate synthetic images Example slices from an original post-contrast image, a rigidity and a nonrigidly transformed synthetic post-contrast images The choice of weight parameter w in Equation (3.1) Breast extraction Intermediate images in lesion detection Maximum intensity projection (MIP) images show the process of false positive reduction True positives using SVM with different combinations of CC and C False positives using SVM with different combinations of CC and C MIP (maximum intensity projection) image to show the initial lesion identification and false positive reduction result False positive reduction ROC curves that show the performance of lesion detection sensitivity against false positive detections per lesion by varying the output class probabilities in SVM Two cases of which all lesions are correctly identified without any false positives A case with 41 isolated false positives A case with 17 true lesions and 30 false positive detections A case with numerous foci in both breasts A 4D ROI (3D spatial + time dimensions) Kinetic features defined based on time-intensity curve: peak enhancement (PE), time-to-peak (TTP), wash-out rate (WOR) and wash-in rate (WIR) of a voxel Demonstrative slices showing peak intensity (PI), peak enhancement (PE), wash-in rate (WIR), wash-out rate (WOR), enhancement (E) at all post-contrast images xv
18 LIST OF FIGURES 5.4 Node graphs of NB-MRF-F and NB-MRF-NP. NB: naive Bays. MRF: Markov random field. F and NP: the edge potentials are computed using node features and node potentials The segmentation results using NB-MRF-F Prototype enhancement curves found by segmenting lesion voxels and non-lesion voxels within the ROI The normalized histogram of E t 2 in lesion and non-lesion groups using the proposed and fixed range normalization methods. E t 2 : enhancement in a post-contrast image that is taken at approximate second minute after the injection of contrast agent The normalized histogram of E t 5 in the lesion and non-lesion groups using the proposed and fixed range normalization methods The normalized histogram of peak enhancement in the lesion and nonlesion groups using the proposed and fixed range normalization methods The normalized histogram of peak intensity in the lesion and nonlesion groups using the proposed and fixed range normalization methods The normalized histogram of wash-in rate in the lesion and non-lesion groups using the proposed and fixed range normalization methods ROC curves of all the segmentation methods Lesion volume distributions of successfully segmented (overlap 40%) and missed lesions by using FCM, GMM, GMM-MRF and NB Lesion volume distributions, mean and standard deviation of successfully segmented (overlap 40%) and missed lesions (overlap 40%) by using NB-MRF-F, NB-MRF-I and NB-MRF-NP Two-compartment pharmacokinetic model Color-coding scheme for creating the color hue and color intensity of the three-time point (3TP) model TIC of 9 clustered in a malignant lesion computed by FCM TIC of 9 clusters in a benign lesion computed by FCM method Normalized radial length (NRL) of a lesion Illustration of NRL and NSD The possible approaches to make use of the CAD system xvi
19 LIST OF FIGURES 7.2 Flow chart in scenario Flow chart in scenario xvii
20 List of Tables 1.1 Morphological descriptors for mass-like finding Morphological descriptors for non-mass-like finding Lesion composition The formulation of synthetic images from real images Ground-truth for registration using synthetic images Registration results on breasts in synthetic images Registration results on lesion regions in synthetic images Evaluation results for 39 clinical images The rules defined in studies from Ertas et al. [21; 22; 23] Lesion composition Experiment results Lesion composition Feature values computed using the fixed range and proposed normalization methods on the lesion and non-lesion voxels The Jaccard index of features: enhancement at the 2 and 5 time phrase (E t 2, E t 5 ), peak intensity (PI), (peak enhancement) PE and wash-in rate (WIR) on lesion and non-lesion groups using the proposed and fixed range normalization methods Segmentation results of AUC and overlap rate with the ground-truth The statistics on the successfully segmented (Y) and misclassified (N) lesions xviii
21 LIST OF TABLES 6.1 Existing computer-aided diagnosis classification methods and their results Lesion composition in the evaluation of the feature analysis and lesion classification Mann-Whitney U-test on the gross shape features: volume, surface, compactness and volume overlap rate (VOR) Mann-Whitney U-test results of NSD- and NRL-based margin features. NSD: normalized surface distance. NRL: normalized radius length Kinetic feature significance test using the Mann-Whitney U-test Textural feature significance analysis using the Mann-Whitney U-test ANN classification results using kinetic, textural, morphological NSD and NRL features Random forest classification results using kinetic, textural, morphological (NSD- and NRL-based) features Lesion number and the method by which the lesion is segmented (manually or automatically) in the training, tuning and testing dataset in the ANN classification in scenario ANN classification results in the scenario 1 using the kinetic, textural, NRL-based morphological, NSD-based morphological features and the combination of the all Lesion number and the lesion segmentation method (manual or automated) in the training and testing dataset in RF classification in scenario Random forest classification results in scenario 1 using the kinetic, textural, NRL-based morphological, NSD-based morphological features and a combination of all of the above Composition of group P and Q Lesion number and the method in which the lesions are segmented manually or automatically in the training, tuning and testing datasets in the ANN classifications in scenario xix
22 LIST OF TABLES 7.7 ANN classification results in scenario 2 using the kinetic, textural, NRL-based morphological, NSD-based morphological and a combination of all of the above Lesion number and the method in which the lesions are segmented manually or automatically in the training and testing dataset in random forest (RF) classification in scenario Random forest classification results in scenario 2 using the kinetic, textural, NRL-based morphological, NSD-based morphological and a combination of all features False positive (FP) number and FP at 100% recall in labeling a region as a lesion or non-lesion xx
23 Chapter 1 Introduction Breast cancer ranks as the second highest cause of death by cancer in women after lung cancer [41]. The major goals of breast cancer diagnosis are early detection of malignancy, which can reduce the likelihood of spread [99] and differentiation of breast cancer tissue from artifacts caused by other breast diseases. Mammography remains the primary screening method employed in the early detection of breast cancer. Screening mammography has reduced breast cancer mortality [35], however, it is less sensitive in detecting cancer in young women, women with dense breast tissues and those who are a BRCA mutation carrier [51; 91]. Many of the breast cancer cases that are missed by mammography are found to be better detected by dynamic contrast-enhanced magnetic resonance imaging (DCE- MRI) [9; 93]. DCE-MRI is complementary to mammography in the diagnosis of breast cancer and plays an important factor in the decision as to whether to pursue conserving surgery or mastectomy [7]. DCE-MRI has been shown to be the most sensitive screening methodology for detecting breast cancer in women with dense breast tissue [92] and those with a high familial risk [52; 110]. The American Cancer Society has recommended that a woman who is an untested first-degree relative of a BRCA carrier and with a cumulative lifetime risk of breast cancer of 20-25% or greater should attend an annual MRI screening. In this chapter, we will introduce magnetic resonance imaging (MRI), explain how a radiologist makes diagnosis using DCE-MRI. We introduce the challenges in interpreting and assisting image interpretation faced by a radiologist and outline that major components in the computer-aided diagnosis (CAD) system of DCE-MRI. In the end, 1
24 we present a fully automated CAD system, of which each component is novel. 1.1 Magnetic resonance imaging Magnetic resonance imaging (MRI) is both flexible and capable of showing the spatial distribution of tissue characteristics. The human body is primarily fat and water and as such, it contains many hydrogen (H) atoms. An MR image measures the interaction between the hydrogen nucleus (proton) 1 H and radio frequency (RF) magnetic fields. A MR scanner generates RF fields that are absorbed by the 1 H protons, and then the protons re-emit certain RF fields that are detected by a receiver in the MR scanner. Therefore, the distribution of water and fat molecules are responsible for the signal in an MRI. An 1 H proton behaves like a minuscule magnet and naturally generates a weak magnetic signal. This is called a nuclear spin, as shown in Figure 1.1. The sum of all the tiny magnetic fields of each spin is called as the net magnetization. The net magnetization is comprised of two orthogonal components, a longitudinal component (M Z ) and a transverse plane component (M XY ). In the natural state of the spins, their direction is randomly distributed and the sum of all spins in both transverse and longitudinal plane gives a null net magnetization. Figure 1.1: 1 H atom behaves like a minuscule magnet, called a nuclear spin. After the 1 H protons are subjected in an external magnetic field B 0 produced by the MR scanner, they align themselves with the field parallelly or anti-parallelly, at a low or high energy state, as shown in Figure 1.2a. As a result, the 1 H protons generate 2
25 a net magnetization in the longitudinal direction (M Z ) and the transverse components are canceled, as shown in Figure 1.2b. The MR scanner acquires image through manipulating the net magnetization field by applying an electromagnetic field in the radio frequency band and measures the interactive behavior of the protons. N N N S S N N S N S N S B0 B0 B0 B0 B0 B0 B0 B0 B0 S N N S S N S N N S S S (a) Alignment (b) Net magnetization Figure 1.2: (a) Spins align to an external magnetic field parallelly at a low energy state and anti-parallelly at a high state. (b) Spins generate a net magnetization along z (longitudinal) direction. The transverse components are canceled out Excitement and relaxation The net magnetization can be rotated in the direction of the XY (transverse) plane, that is perpendicular to the Z axis, by sending an RF pulse with a certain strength and frequency for a certain period of time as shown in Figure 1.3. The net magnetization flips from the Z axis onto the XY plane as the protons absorbed energy from the RF pulse in the excitement process. At this moment, all spins are in phase which means they are all precessing together around the external magnet field direction. The precession refers to the spin rotation of the transverse component along the longitudinal axis. When the spins are in phase, they generate maximum net magnetization in the transverse plane M XY and the net magnetization reduces when they precess out of phase (i.e. they do not precess together). After the excitement, when the RF pulse stops, the protons release the absorbed energy and generate a RF wave that can be detected by a receiver in the MR scanner. 3
26 RF pulse Z Y X Figure 1.3: A RF pulse flips the net magnetization 90 from the longitudinal axis to the transverse plane. Figure 1.4 illustrates the process of relaxation where the overall M Z recovers and the M XY decays or reduces. T 1 relaxation is defined as a time constant it takes for the longitudinal net magnetization (M z ) to regain 63% (1 1 ) of the original net magnetization by releasing the e absorbed energy to the surrounding tissues, as shown in Figure 1.5a. The density of 1 H protons in different tissues vary which reflect as image contrast in MRI. For example, 1 H protons in fat bind more tightly compared to those in water. Therefore, the 1 H protons in fat release their energy to the surrounding tissues at a faster rate than those in water. At the time of the image acquisition, water molecules with a longer T 1 will only partially recover and hence appear darker in the T 1 weighted MR image. In comparison, fat molecules with a shorter T 1 recover more quickly and present as brighter in the same image. While the 1 H is recovering to its original magnetization along the z-axis, the net magnetization along the transverse plane (M XY ) decays towards zero. The protons change the state of a total in-phase situation to a total out-of-phase situation. T 2 relaxation is defined as the time constant it takes for the proton to de-phase to 37% of the original value, as shown in Figure 1.5b. Similar to T 1, different tissues also have a different T 2. For instance, fat molecules de-phase faster than water molecules. In the T 2 weighted image, molecules with short T 2 have a low signal in the MR image, hence the fat molecules look darker than the water molecules. In addition to the measurement of the T 1 and T 2 tissue characteristics, some mag- 4
27 Z Z RF wave RF wave B B Y X Y X (a) time 1 (b) time 2 Z Z RF wave RF wave B B Y X Y X (c) time 3 (d) time 4 Figure 1.4: The net-magnetization regains along Z-axis and finally aligns with Z-axis from time 1 to time 4. netization techniques can eliminate the effect of fat tissues on the image when the fat signal obscures a tissue of interest. The Figure 1.6a shows a fat-suppressed T 1 image, in which fibroglandular tissue has a higher intensity value than fat tissue. Figure 1.6b shows its corresponding non-fat-suppressed T 2 image where fat tissue is brighter than the fibroglandular tissue which provides complementary information when needed Spatial encoding A voxel in MRI is a single volume element that contains protons. To localize the voxel, spatial information needs to be encoded into the MR signals by selecting the desired slice, encoding along rows and following this along columns. A magnetic field gradient is a variation in the magnetic field with respect to position. Spatial encoding 5
28 Mz recovery Mxy decay 100% 100% 63% 37% T Time (msec) T Time (msec) (a) T 1 (b) T 2 Figure 1.5: Illustration of (a) T 1 and (b) T 2 relaxation time. The horizontal axis is time (miliseconds) and the vertical axis is the percentage of the M Z (a) recovers to its original value and the M XY. (b) decays to zero. (a) Fat-suppressed T 1 (b) Non-fat-suppressed T 2 Figure 1.6: (a) Fat-suppressed T 1 weighted image, in which fibroglandular tissue has a higher intensity than fat tissue. (b) the corresponding non-fat-suppressed T 2 image in which fat tissue is bright than the fibroglandular tissue. is accomplished by superimposing gradient fields. The gradient coils in a MR scanner generate a spatially variate magnetic field so that the 1 H protons at a different location precess at frequencies unique to their location, allowing the MRI scanner to reconstruct an MR image. An RF pulse at a certain frequency that can excite the 1 H protons is called Larmor 6
29 frequency. It is affected by the local value of the magnetic field. In an MRI scanner, the main magnetic field in the Z-axis is uniform and static. Additional magnetic fields can be temporarily superimposed on the main field which makes the Larmor frequencies of the 1 H protons vary. A particular slice of protons can be excited or flipped by applying a RF pulse at the Larmor frequency. Figure 1.7a shows the magnetic gradient in the Z-axis. After the slice is selected, a gradient in the y-axis is applied to differentiate the protons. They are in different rows based on the phase of the protons. In the phase encoding, the transverse magnetization of the different rows are rotated to different positions along the y-axis. After each row in the slice is identified, an RF pulse with different frequency is applied to generate a gradient in the column direction or tje x- axis. This process in called frequency encoding. Figure 1.7b demonstrates the phase encoding and frequency encoding by applying field gradients in the x-axis and y-axis respectively. Z X Z Y (a) Slice selection (b) Phase and frequency encoding Figure 1.7: Illustration of (a) a slice selection in the z-axis by temporarily superimposing additional magnetic fields in the z-axis, (b) phase encoding in the y-axis by applying gradient magnetic fields in the y-axis, and frequency encoding in the x-axis by sending an RF pulse with a different frequency in the x-axis Contrast agents Despite the 3D information generated by an MRI, the contrast in some MR images is still insufficient in visually distinguishing between normal and abnormal structures 7
30 in some body parts, such as the breast. Magnetic resonance contrast agents are compounds that are able to dramatically change the T 1 and T 2 relaxation times. The contrast agent is usually injected intravenously and several MRI sequences are acquired before and after the injection of the contrast agent. This allows the observation of the intake and depletion pattern of the contrast agent. The T 1 and T 2 contrast agents can interact with 1 H spins and reduce the corresponding relaxation time, thus producing different MR signals, usually targeting the relaxation time in the blood. Figure 1.8 show a MR breast image with and without contrast agent. (a) No contrast agent (b) With contrast agent Figure 1.8: (a) A slice of an MR image without contrast agent. (b) A slice of an MR image of the same region with contrast agents. 1.2 Dynamic contrast-enhanced MRI of breast Angiogenesis is the process where new blood vessels are formed from pre-existing vessels. It is an important process that occurs in both normal and abnormal body tissues. The normal tissues control the process of angiogenesis by stimulating or inhibiting the blood vessel formation. However, cancer tissues lose the ability to regulate the angiogenesis process. The angiogenesis can aggressively grow blood vessels to feed the malignant tissues. Therefore, the process of angiogenesis is a vital indication of the growth and development of a malignant lesion. Malignant tissues require a larger amount of blood supply than in normal body tissues which results in increased angio- 8
31 genesis. There is higher and denser growth of micro-blood vessels around the malignant lesion, to support the aggressive growth and spread of the lesion. A large number of new vessels are formed in cancer tissue at a much higher rate than that in normal tissues. The blood walls in most cancer tissues are not properly constructed and thus are highly permeable. This is what makes cancer detection using dynamic contrastenhanced magnetic resonance imaging (DCE-MRI) effective, where a contrast agent is used to differentiate tissues with higher blood concentrations than normal tissues. A DCE-MR image consists of several MRI-scans over time, as the contrast agent is injected into the blood stream and dissipates in the body. Each MRI scan is a threedimensional image. The first scan, known as the pre-contrast image, serves as a baseline. Once the contrast agent has been injected, the subsequent MRI scans of the breast tissue performed at a regular time interval, known as post-contrast images, show the absorption and dissipation of the contrast agent in the breast tissues. Different tissues have different micro-vascular permeability characteristics. A DCE-MRI allows the examination of the micro-vascular permeability and density by observing and quantifying the absorption and depletion contrast agent into breast tissues. As malignant tissues have increased blood supply, higher concentrations of contrast material will be absorbed by these tissues and appears brighter than other tissues in the post-contrast images soon after the injection of the contrast agents. Malignant tissues usually have poor micro-vascular structure, where the contrast agent can leak into the surrounding tissues faster. Figure 1.9 shows a pre-contrast MR sequence, a first post-contrast MR sequence and the subtraction image of the two. The lesion is marked by a square in all of the figures. It can be seen in the subtraction image 1.9c that the lesion is much more enhanced compared to other tissues in the breast. 1.3 BI-RADS R lexicon The Breast Imaging Reporting and Database System (BI-RADS R ) [79] developed by the American College of Radiology provides a reporting standard and lexicon for abnormal enhancement findings in the breast DCE-MRI. Radiologists diagnose breast cancer using DCE-MRI images by finding abnormal enhancements. The BI-RADS R defines abnormal enhancements as enhancements of higher signal intensity compared to the surrounding normal glandular tissue in a contrast-enhanced image [79]. There 9
32 (a) Pre-contrast image (b) Post-contrast image (c) Subtraction image Figure 1.9: Pre-contrast (a), post-contrast (b) and the subtraction (c) images of the image (b) from the image (a) with an enhanced lesion labeled by the square mark. are three types of enhancement findings: focus (< 5mm), mass and non-mass-like enhancements. Radiologists perform morphological analysis on the shape, margin (surface of lesions) and spread of the suspected finding of cancerous tissue, and kinetic analysis on the flow of blood to identify malignant tissues and their types of cancer. The diagnosis report is based on the descriptors (e.g. shape, mass margin) in the BI- RADS R lexicon. Radiologists perform kinetic analysis on the flow of blood in a region of interest to differentiate malignant from benign tissues. The time intensity curve (TIC) is estimated from the rate of absorption and dissipation of the amount of contrast material 10
33 Relative enhancement in a region in the post-contrast images. The estimation of TIC requires the same tissues to be located at the same position at different time steps. Motion artifacts, such as respiratory and cardiac motions, can lead to the inaccurate TIC of a region. TIC is described in BI-RADS R by its initial and delayed phase enhancement patterns, as shown in Figure The initial enhancement patterns are slow, medium or rapid. Delayed enhancement patterns include persistent (intensity increase over time), plateau (no intensity change over time) and washout (intensity decrease over time). Early phase Later phase Persistent ±20% Plateau Washout t t t Time Figure 1.10: Initial and delayed enhancement patterns defined in BI-RADS R. Cancer tissues usually present rapid patterns in the up-take and release of contrast material (washout). Figure 6.3 shows the pre- and post-contrast images of a cancer and its corresponding TIC. Benign lesions usually have a slower initial enhancement and plateau or persistent delayed enhancement patterns. Figure 6.4 shows pre- and post-contrast images of a benign lesion and the derived TIC of the lesion. In addition to the kinetic descriptors, BI-RADS R also defines the morphological descriptors for mass and non-mass lesions. Table 1.1 shows different types of shape, margin and internal enhancement that are more likely to be observed in malignant and benign mass lesions. Malignant lesions usually have an irregular shape and margin due to the aggressive growth of the cancer tissue. A malignant lesion with a spiculated 11
34 (a) Pre and post-contrast image of a malignant lesion 1500 Intensity Cancer normal 0 t0 t1 t2 t3 t4 t5 Time (b) Time-intensity curve (TIC) Figure 1.11: A malignant lesion (a) and its time-intensity curve (b). margin and irregular shape is shown in Figure 1.13a, and another with a heterogeneous internal enhancement pattern is shown in Figure 1.13b. Malignant Benign Shape lobulated, irregular round, oval Margin irregular, spiculated smooth Internal Enhancement Heterogeneous Homogeneous Table 1.1: Morphological descriptors for mass-like finding. Malignant Benign Internal Enhancement Clumped Stippled, regional, multiple regional, diffuse, reticular Bilateral enhancement Asymmetric Symmetric Table 1.2: Morphological descriptors for non-mass-like finding. 12
35 (a) Pre and post-contrast image of a benign lesion Intensity Benign normal 200 t0 t1 t2 t3 t4 t5 Time (b) Time-intensity curve (TIC) Figure 1.12: A benign lesion (a) and its time-intensity curve (b). In BI-RADS R system, internal and bilateral enhancement patterns are used to characterize the morphological features of a non-mass lesion. Table 1.2 shows the descriptors that are observed in malignant and benign findings. For example, if a patient has a lesion that presents clumped internal enhancement in one side of the breast, the lesion is more likely to be malignant than benign. 1.4 Computer-aided diagnosis (CAD) system and our CAD system The DCE-MRI of a breast involves the analysis of large volume of data. Commercial and non-commercial computer-aided diagnosis (CAD) systems are available to assist radiologists in interpreting images. The word diagnosis can refer to a broad range of analysis of the image examination results. CAD systems are designed to support but not replace radiologists in the process of diagnosis. 13
36 (a) spiculated margin (b) heterogeneous enhancement Figure 1.13: (a) A lesion with a spiculated margin and irregular shape. (b) A lesion with a heterogeneous internal enhancement pattern. A non-commercial CAD system can refer to a computer-aided detection (CADe) or a computer-aided diagnosis (CADx) system [33]. A CADe system is used to identify suspicious tissues. A CADx system typically assumes that these suspicious tissues in a breast have been detected either manually or by a CADe system. It then extracts features of the lesions and labels them as benign or malignant using these features. Popular commercial CAD systems include CADstream R (Merge Healthcare Inc., Chicago, IL, US) and DynaCAD R (Invivo, Gainesville, FL, US). These systems provide various functions, including visualization, reformatting, motion correction, colorcoded kinetic mapping, time-intensity curve plotting, lesion segmentation, morphological feature analysis and lesion evaluations etc. Most functions provided in these CAD systems allow radiologists to interact with the system. The flowchart of a typical CAD is illustrated in Figure It consists of four main functions: (1) using image registration to reduce motion artifacts in pre- and post-contrast MR images such that the computation of kinetic features are reliable, (2) selecting a region (most likely manually) of interest that encloses a lesion and segmenting the lesion volume, (3) computing morphological and kinetic features of the lesion, (4) selecting the most important features for classifying the lesion as benign or malignant. 14
37 Input: DCE-MRI Image Registration Reduce motion exist in between MR images Lesion localization Localized and segment a lesion inside a ROI Feature analysis Extract all features of the lesions Lesion classification Classify the lesions using selected features Output: lesion evaluation result Figure 1.14: The general work flow of a CAD system. It is challenging to preserve the volume and texture of lesions in recovering the motions between pre- and post-contrast images. Assuming the motions have been corrected, it is still a difficult task to automatically detect and segment the lesion in a DCE-MRI due to the noise in an MR image and the large variation of tissue features among patients. After the lesions are segmented, they can be labeled using these lesion features. The accumulation of errors in each component can lead to a poor performance in labeling an automatically detected lesion in a DCE-MR image. In this thesis, we propose a fully automated CAD system in which the major components are novel. This system includes functions that are found in CADe and CADx systems. As far as we know, our system is the first that performs tasks in both CADe and CADx systems. Our system takes a 4D DCE-MRI breast image as an input and automatically detects and labels all suspicious tissues in the breast image as malignant or non-malignant. Our CAD system includes five novel components, as shown in Figure In this chapter, we briefly introduce each component of the CAD system. A more 15
38 DCE MRI Chapter 2 Automated Image registration Manual Chapter 3 Automated Lesion detection Manual Chapter 4 Automated Lesion segmentation Manual Chapter 5 Feature analysis Automated Lesion classification Manual Chapter 6 Automated Manual Diagnosis report Figure 1.15: The work flow of the proposed CAD system: image registration to recover the motions, ROI detection to identify suspicious regions, lesion segmentation to draw the contour of a lesion and lesion classification to label a lesion as malignant or benign. Each component (task) can be performed manually or automatically. detailed literature review and discussions of each component will be presented in later chapters Image registration It usually takes several minutes to acquire all DCE-MRI scans of a breast, including the time taken in injecting contrast agents. Various motions can occur in between MRI acquisitions due to the subject breathing or adjusting their body positions. The motion artifacts can cause the kinetic analysis to be unreliable or even impossible to perform. Image registration is commonly applied in reducing motion artifacts between multiple images of the same body tissue, to obtain spatially consistent regions of interest. Unavoidable respiratory and cardiac motions that appear on DCE-MRI images con- 16
39 tribute to inaccurate estimation of the time intensity curves (TIC) in suspicious regions, where the TIC is used in kinetic analysis to identify lesion malignancy. In general, a widely used image registration algorithm has been demonstrated to be able to correct the overall motions in the whole breast images [89]. The challenge is to achieve a good alignment over the local suspicious lesions while not compromising the overall breast image [87; 104]. Most lesions are known to be relatively rigid or stiff in nature while other breast tissues are nonrigid or soft. In the existing registration studies, additional information regarding the tissue stiffness/rigidity through a lesion segmentation is used to ensure a good local alignment over lesions [64; 88; 100]. However, manual lesion segmentation is time consuming. An automated segmentation method either requires the absence of motions in the image or requires a preliminary registration to correct motions if they exist. Therefore, it is impractical to perform lesion segmentation or localization in order to perform an effective image registration. There is a need to develop an algorithm that is able to align both lesion and non-lesion tissues without having to perform lesion segmentation. We propose a weighting method of the regularization term in B-spline based nonrigid registration method. The aim of the weighting strategy is to preserve the lesion information while correcting the overall motions over the whole breast. We have published our method in [60; 61]. A more comprehensive introduction to the methodology and validation on the image registration will be presented in Chapter Lesion localization After the motions have been recovered, the tissue locations in different MR sequences are spatially consistent. Given there are few or no motion artifacts to complicate the comparison of the post-contrast image with the pre-contrast image, the detailed scan of the abnormal enhancement can be easily seen using some visualization tools [81]. Maximum intensity projection (MIP) is a visualization method that projects the voxels in a 3D image with maximum intensity to a 2D image. Radiologists make use of an MIP of the subtraction image of a post- from pre-contrast image to locate regions of interest. Figure 1.16 shows such an example. In cases that require visualizations of higher details, true three-dimensional reconstructions are used. Some CAD systems are capable of identifying and highlighting the tissues with 17
40 suspicious TIC patterns by superimposing color-encoded maps on original images to label their tissue attributes [81], e.g. malignant tissues are labeled in red and normal tissues in blue. However, some malignant and benign tissues share similar TIC patterns, which could lead to false-positive diagnosis by radiologists using CAD systems [6]. Figure 1.16: Maximum intensity projection (MIP) of a subtraction image of a postfrom pre-contrast image An accurate lesion segmentation or delineation plays an important role in the computerized assessment of lesion malignancy by quantitatively analyzing the morphological, textural and kinetic features of the lesions. Manual lesion segmentation is labor intensive and is also subject to inter-observer and intra-observer variations [10]. Automated segmentation enables consistent, efficient and reproducible lesion delineation. Fuzzy c-means (FCM)-based segmentation [5; 15; 58; 80; 98] is a commonly used baseline method that is used to be compared with other methods in segmenting breast lesions in DCE-MRI. Most existing segmentation algorithms require users to place a bounding box as an ROI or a seed point over the lesion. Therefore, there is a need to investigate automated selection of regions of interest that contain lesions, or lesion region detection. There are several studies that investigated in lesion detection in the whole breast [8; 21; 22; 23; 30; 108]. These are able to identify most lesions but the false positive detections are still high. Radiologists are required to manually rule out those regions. 18
41 Therefore, there is a need to investigate a lesion detection method with high sensitivity and low false positive detections in order to achieve a fully automated CAD system. There are very few studies available in investigating fully automated lesion detection systems in breast DCE-MRI. We propose a novel algorithm to identify breast lesions aiming to have a high sensitivity while having a reasonable false positive detection rate (less than 20%). We expect that with false positive ratio of 20% would potentially reduce the workload for the clinicians by 80%. Chapter 4 will present a more detailed literature review of lesion detection methods. In addition to lesion detection from the whole original image, we also propose a novel lesion segmentation method to delineate the lesion margin from a region of interest that is selected either manually or automatically. In our segmentation method, each ROI is presented as a connected graph with a local Markov property. We have published this method in [58] and present it in Chapter Feature analysis and classification Once the lesion has been localized or segmented accurately, various properties are computed to characterize the lesion. Kinetic features are extensively investigated and used as a conventional method in the evaluation of the performance of various classifiers [6; 16; 20; 105; 111]. Although kinetic features provide valuable insight into the diagnosis of lesion, there is a significant overlap in the enhancement patterns between benign and malignant lesions. More recently, morphological features are computed and evaluated in the lesion analysis [1; 2; 63; 73; 77; 78]. Similar to kinetic features, some malignant cancers are reported to show benign-like morphologies [82]. After all lesion features have been extracted, various classifiers such as KNN, decision trees, support vector machines and neural network can be applied to label each lesion as benign or malignant. The prediction results can be presented in various ways, such as the probability of the diagnosis or as a binary recommendation. Lesion feature analysis and classification is usually called a CAD system in the academic research community and there have been a number of active investigations in recent years [2; 32; 53; 56; 63; 68; 73; 77; 78; 85; 86; 113]. We also propose and have previously published a shape and margin feature characterization method [59] that is presented in Chapter 6 of this thesis. 19
42 1.4.4 Versatility of CAD system In our CAD system, a user can choose to employ a component or a combination of several components automatically or manually based on the task requirement. For instance, if a series of DCE-MRI sequences present no obvious motions and the lesion enhancement is much more enhanced than other breast tissue, e.g. fat and glandular tissue, a radiologist may choose to skip registration and ROI selection and only perform lesion segmentation and feature analysis to assist his/her diagnosis. Within the DCE-MRI academic research community, breast registration, lesion selection and segmentation are usually excluded from the scope of CAD, and are regarded as pre-processing methods of these CAD systems. Therefore, the input of the CAD systems in the academic research community usually is the segmented lesions from the DCE-MR images. Ideally a CAD system should be capable of taking the original DCE-MR images as input and generating images that are labeled as benign or malignant. This is a challenge in building a fully automated CAD that can interpret a DCE-MR image as a second reader. Such CAD systems for mammography are approved by the FDA for the detection of breast cancer, including R2 ImageChecker CAD R (Hologic, Inc., Bedford, MA, America) and SecondLook Digital CAD R (icad, Inc., Nashua, NH, America). In this thesis, we validate two approaches using our CAD system in DCE-MR image interpretation. The evaluation will be presented in Chapter Thesis structure In this thesis, Chapter 2 presents all images used in examining and validating each component of our CAD system. Chapter 3 presents our registration method which is an extension of our previous publications [60; 61]. The validation of the registration method in this thesis is evaluated using a larger dataset than that in [60; 61]. A new lesion detection method is proposed in Chapter 4. Chapter 5 presents our published lesion segmentation algorithm [58]. Our published feature extraction and lesion classification systems [59] are presented in Chapter 6. The evaluation of the fully automated CAD is presented in Chapter 7. Chapter 8 concludes the thesis. 20
43 Chapter 2 DCE-MRI Database There were 39 patients or cases enrolled in our study. The patients had 83 lesions in total that were used in evaluating each novel component or method of our CAD system. The images were collected between 2006 and 2011 at St Vincent s Hospital in Melbourne, Australia. 21
44 We were able to collect tje nreast DCE-MR images from 39 patients between 2006 and 2011 at St Vincent s Hospital in Melbourne, Australia. Our study was rated as a Minimal Risk Project and an Ethics Clearance for this project was granted in 2011 by the Engineering Human Ethics Advisory Group at the University of Melbourne, Australia. Thirty-nine patients were used in evaluating each novel component of our CAD system. All the image protocols and disease statistics of these patients are presented in this chapter to avoid duplications in later chapters. 2.1 Imaging protocol All DCE-MR breast images were acquired with a Siemens Avanto 1.5T MR system. There were 20 patients from whom T 1 - and T 2 -weighted images were available while 19 patients have only T 1 -weighted images. All T 1 -weighted images in the breast DCE-MRI database are fat-saturated. The protocols of these images are largely the same, except for those obtained between June 2010 and July 2011 for which a slightly different DCE sequencing protocol was used. During this time, a high-resolution three-minute Sagital volume was inserted in the DCE sequences. Therefore the time of image acquisition was increased from 6.32 minutes to 8.29 minutes. The first protocol was T R = 4.19 ms, T E = 1.22ms, field of view = 340 mm and flip angle = 10. The voxel dimensions were approximately mm 3. The temporal resolution was 1.02 minutes. There were six measurements or image scans in total, one pre-contrast sequence, followed by a 20 seconds pause for contrast injection, then five sequences after the injection. The second protocol had T R = 4.82ms, T E = 1.79ms, field of view = 380mm and flip angle = 10. The voxel dimensions were approximately mm 3. The temporal resolution was 1.01 minutes. The total acquisition time was approximately 8.29 minutes, starting with a pre-contrast sequence, a 25-second pause for contrast injection, two more post contrast sequences, a three-minute long high-resolution Sagital sequence (not part of DCE for contrast uptake analysis) and two further sequences to complete the DCE series. There were six image scans in both protocols. The kinetic analysis of the images using the second protocol was performed on only five of the six images, excluding the 22
45 high-resolution sequence taken in the fourth acquisition. This sequence was mainly used for morphological analysis purposes, as the high resolution allows more accurate shape and margin analysis especially in cases where the lesions were small. For 20 of the 39 patients, T 2 -weighted non-fat-saturated fast inversion recovery (STIR) sequences were available, in addition to the T 1 -weighted dynamic MR sequences. The T 2 -weighted STIR sequence was measured with T R = 9290 ms, T E = 89ms, field of view = 350 mm and flip angle = 150. The voxel dimensions were approximately mm Lesion statistics There were 83 breast lesions in the 39 patients, including 28 malignant and 44 benign lesions, 3 foci and 8 lesions of unknown type. The lesion composition is listed in Table 2.1. All malignant lesions were histologically verified. Only 30 benign lesions were biopsy-proven while the remainders were diagnosed as benign. The 8 lesions of unknown type were not classified due to the lack of ground-truth. Malignant Benign Type Number Breast carcinoma 1 16 DCIS 6 IDC/DCIS 2 IDC 1 ILC 3 Cyst 2 Fibroadenoma 40 Benign papillary lesion 1 Adenoma 1 Foci 3 Unknown 8 Total 83 Table 2.1: Lesion composition. The lesion volume distribution is plotted in Figure 2.1. Most of the lesions are small, with 65 lesions (78.5%) having a volume of less than 1 cm 3. The volume of 23
46 benign lesions ranges between 0.04 cm 3 to 5.34 cm 3 with a mean of 0.55 cm 3. The volume of malignant lesions ranges from 0.06 cm 3 to 9.69 cm 3 with a mean of 2.17 cm 3. number of lesions lesion volume (cm 3 ) Figure 2.1: Lesion volume (cm 3 ) distribution 24
47 Chapter 3 Breast registration The first component of our CAD system is image registration, which aligns two images such that they are spatially consistent. In this chapter, we review the existing registration methods that are used in motion correction of breast DCE-MRI. We then propose a method to automatically compute the regularization coefficients in B-spline based nonrigid registration without a need of lesion segmentation. Both synthetic and real DCE-MR images are used to evaluate our regularization coefficients against others that require segmenting lesions manually in B-spline based registrations. In our dataset, the performance of the preservation of lesion volumes by using our automatically generated coefficients is comparable to the manually assigned rigidity coefficients. This chapter is an extension of our previous publications 12. The methodology presented in this chapter is the same as previously published but it is evaluated using a larger dataset (as stated in Chapter 2). 1 Xi Liang, Kotagiri Ramamohanarao, Qing Yang, Alexander Pitman, and Marious Staring. A practical method to compute coefficients for regularization term in nonrigid registration of DCE-MRI. In 20th Annual Meeting and Exhibition of International Society for Magnetic Resonance in Medicine, page 3035, Xi Liang, Kotagiri Ramamohanarao, Qing Yang, Alexander Pitman, and Marius Staring. Generating regularization coefficients in nonrigid registration of DCE-MRI of breast. Workshop on Breast Image Analysis, pages 1-8,
48 3.1 Introduction In breast imaging, image registration serves as a prerequisite for a quality image interpretation, the fusion of different imaging modalities, and surgical planning. Image registration can be formulated as an optimization problem that aims to minimize the differences between multiple images such that they are spatially or temporally aligned. It is composed of four main components: a transformation model to align two images, an image similarity metric to measure the quality of alignment, a transformation regulator to penalize undesirable deformations (over-fitting) and an optimizer to compute transformation parameters. The image registration framework is shown in Figure 3.1. Registration framework Fixed image Moving image Image similarity Metric Image interpolator Transform Cost function optimizer Transform parameters Registered image Figure 3.1: Image registration framework. A transformation model is used to align the moving image to the fixed image. An image similarity metric in the cost function measures the quality of the alignment of the two images, serving as the driving force of the optimization. The regularizer serves as a soft constraint to penalize undesirable deformations. The optimizer computes the transformation parameters. Image registration can be categorized based on its transformation model that determines the complexity and flexibility of a registration. Rotation, translation, or their combinations, are used to model global motions in rigid transformations. Affine trans- 26
49 formations, such as scaling, shears, and skews, are also used in the modeling process. In breast DCE-MRI, rigid and affine registrations are typically applied on a whole breast volume in order to recover motions caused by adjusting body positions during image acquisitions. However, breast tissue is non-rigid in nature, as it dilates and contracts with breathing and cardiac motions. Rigid or affine transformations are not able to capture these local deformations. Consequently, nonrigid (or elastic) transformations are developed to model such local motions, such as B-spline based [89] and thin-plate spline based [74] models. Rigid and nonrigid registration are illustrated in Figure 3.2, showing the registration by two faces, one fixed (Figure 3.2 (a)) and the other moving (Figure 3.2 (b)). The moving image is rigidly registered to the fixed image globally by computing and applying a transition (Figure 3.2 (c)) and a rotation (Figure 3.2 (d)) to the moving image, resulting in a rigidly registered image (Figure 3.2 (e)). After the initial rigid registration, a nonrigid registration is performed to allow local deformations as shown in Figure 3.2 (f). One of widely used nonrigid registration algorithms in breast DCE-MRI is that of Rueckert et al. [89] which is based on B-spline and free-form deformations (FFD). There are a number of extensions or refinements of this work [87; 100; 104]. There are also other nonrigid registration algorithms, such as curvature-based registration [26] and diffusive flow methods [18; 29]. Image registration can be classified based on the feature space used in computing image dissimilarities. The feature space is the information extracted from an image that is used for matching, such as pixel intensities [87; 89; 104], edges [67] or surfaces. The main focus of registration techniques is the measurement of intensity difference. Mutual information (MI) [71; 83] is a popular similarity metrics in b-spline based nonrigid registration of breast DCE-MRI. The parameters of a transformation model can be approximated or computed by maximizing the MI between two images. There are two main reasons that lead to intensity differences. One is motion artifacts that occur between the image acquisitions. The other is the tissue signal enhancements in post-contrast images caused by the injection of contrast agents. Therefore, minimizing the image intensity differences not only reduces the motion artifacts, but may also change the volume of the enhancing regions [87; 104]. Regularization terms are used to penalize undesirable deformations in registrations. Volume preserving terms are used to preserve the volume of regions of interest [36; 37; 27
50 (a) fixed image (b) moving image (c) transition (d) rotation (e) rigidly registered (f) nonrigidly registered Figure 3.2: Illustration of a rigid and a nonrigid registrations. In the rigid registration, a transition and a rotation is applied on the moving image to align it to the fixed image. In the nonrigid registration, the image is locally deformed as illustrated by the grid. (a) fixed image. (b) moving image. (c) transition of the moving image. (d) rotation after transition. (e) rigidly registered result (solid) overlaid on fixed image (dotted). (f) final nonrigid registration result (solid) overlaid on the fixed image (dotted). 87; 104]. The assumption is that the total time of image acquisitions of a DCE-MR image is short enough that motion artifacts do not lead to volume changes in the breast or lesion. The incompressibility term is applied on the whole breast with the same weight for all breast tissue [87; 104]. Spatial-variant rigidity terms have been proposed to preserve the rigidity of different breast tissue [64; 88; 100]. In the DCE-MR application to a kidney, a local rigidity constraint was applied to preserve the rigidity of the kidney tissue [90]. These terms require a coefficient or weight to determine the tissue stiffness of each voxel. While 28
51 applying the coefficients in the registration of DCE-MRI of breasts, lesions are usually assumed to be rigid and other tissue (e.g. fat) is assumed to be relatively soft. Therefore some kind of lesion segmentation is required in order to build a binary stiffness map. Manual segmentation of the lesions is usually regarded as the most accurate method but can be extremely time consuming. Automated lesion segmentation, however, requires a preliminary successful registration to correct motion artifacts to make use of kinetic features. Therefore, to apply a rigidity term in the registration of the DCE-MRI of breasts, it is desirable to build a robust and reliable coefficient map for the breast tissue. A regularization term with robust term coefficients is proposed in this chapter to correct motions by applying b-spline based registration while preserving lesion rigidity in breast DCE-MRI. In the evaluation of our method, three b-spline based registration schemes are compared to investigate the performance of motion recovery over the whole breast and lesion regions. All three schemes applied the same method of measuring image similarity, varying only in the regularization terms used. One scheme employs a smoothness term in the registration. Another applies a rigidity term with binary coefficients generated by performing a manual lesion segmentation. In the evaluation, the registrations using the proposed rigidity coefficients shows comparable performance to those that are assigned manually in both the whole breast and the lesion regions. 3.2 Methodology Image registration is defined as a problem of finding a spatial transformation T relating two images of dimension d, one of which is fixed f(x) : Ω F R d R and the other moving m(x) : Ω M R d R. In this research, intensity-based image registration is employed. It is formulated as an optimization problem in which a cost function C is minimized with respect to the spatial transformation T. The cost function is composed of two components, a image similarity cost function and a regularization cost function. A similarity metric S defines the quality of the match and serves as the driving force of the optimization. A regularizer R is introduced to avoid overfitting in the transformation parameter approximation. This penalizes the undesirable 29
52 deformations as a soft constraint. The cost function is formalized as C(T ; f, m) = S(T ; f, m) + wr(t ), (3.1) where w balances the similarity metric against the regularizer which is usually empirically determined. The formalization of Equation (3.1) is applied in many studies [87; 89; 100; 104] and is also adapted in this chapter. In the following section, we will discuss each component in the registration algorithm applied in our study in detail, including rigid transformation (Section 3.2.1), nonrigid transformation (Section 3.2.2), the image similarity metrics (Section 3.2.3), registration terms and their coefficients (Section 3.2.4), and the choice of fixed/moving images in a DCE-MRI (Section 3.2.5) Rigid transformation Rigid registration is usually regarded as an initial pre-processing for a local motion reduction. We apply rigid transformations to model global breast motions. The motions are characterized by six degrees of freedom, modeling a combination of three rotations and three translations. Given the domain of the moving image Ω M = {x = (x 1, x 2, x 3 ) 0 x 1 < X 1, 0 x 2 < X 2, 0 x 3 < X 3 }, the rigid transformation model is written as θ 1,1 θ 1,2 θ 1,3 T θ (x 1, x 2, x 3 ) = θ 2,1 θ 2,2 θ 2,3 x 1 x 2 + θ 1,4 θ 2,4, (3.2) θ 3,1 θ 3,2 θ 3,3 x 3 θ 3,4 where the coefficients θ are the parameters to approximate the global motions Nonrigid transformation Local motions are modeled by a B-spline based transformation model [54; 55], called free-form deformation (FFD) [89]. FFD transforms the moving image by manipulating 30
53 the control points µ i,j,k of mesh n x1 n x2 n x3 overlaid on it and is defined as, T µ (x) = B l (u)b m (v)b n (w)µ (i+l),(j+m),(k+n), (3.3) l=0 m=0 n=0 where i = x 1 /n x1 1, k = x 2 /n x2 1, j = x 3 /n x3 1, u = x 1 /n x1 x 1 /n x1, v = x 2 /n x2 x 2 /n x2, w = x 3 /n x3 x 3 /n x3, B l is the l-th B-spline basis functions, B 0 (u) = (1 u) 3 /6, B 1 (u) = (3u 3 + 6u 2 + 4)/6, B 2 (u) = ( 3u 3 + 3u 2 + 3u + 1)/6, B 3 (u) = u 3 /6. The control points µ are the parameters of the nonrigid registration and changing them locally affects the deformations Similarity metric A similarity metric measures the alignment of the transformed moving image to the fixed image by comparing the intensity differences between the two. Mutual information (MI) is commonly used to measure the alignment of images of different modalities [69; 70; 83]. Normalized MI is defined as NMI(A, B) = H(A) + H(B) H(A, B), (3.4) H(A, B) where H(A) and H(B) are the marginal entropy of A and B and H(A, B) is their joint entropy, which is computed from the joint histogram of A and B. The NMI of two unrelated images is close to 0 and perfectly aligned images are maximized to 1. NMI is employed as the similarity metric in this study Regularization term FFD based nonrigid registration recovers the motions between two MRI scans by minimizing the image intensity differences between the two. There are two main reasons 31
54 that lead to the intensity differences: one caused by the motions between two MR image acquisitions, the other is the intensity enhancement caused by contrast agent injections in between the pre- and post-contrast images, especially on lesion and vessel tissue. Therefore, a desirable registration in DCE-MRI should be capable of reducing the intensity differences due to the motion artifacts while preserving the intensity enhancement. Various regularization terms are used in image registration to discourage undesirable deformations in aligning the moving image to the fixed image, including thin plate bending energy [89] to encourage the smooth deformations, an incompressibility term [87] to preserve tissue volumes, and an orthogonality term [88] to preserve tissue stiffness. Staring et al. [100] transform and integrate these three regularization terms into one term which has been shown to achieve better tumor rigidity preservation than individuals when applied on 3D CT thorax images taken at different times. These CT images were acquired at different times and include both rigid and nonrigid motions. In this research, a combination of two terms, affinity and orthonormality, as described in [100], is applied as a soft constraint in the cost function to preserve the tissue stiffness in breast MRI. There are at least two conditions that must hold for a transformation T (x) to be a rigid transformation. This section discusses a 2D matrix (transformation) as an example to illustrate the regularization term. For a transformation T (x) = u(x) + x to be rigid at location x where u(x) is the deformation, it must hold that u(x) = u(x) + x = Rx + t, (3.5) ( ) r 1,1 r 1,2 where R = is a rotation matrix and t = (t 1, t 2 ) is a translation vector. Two conditions on T (x) can be derived, called affinity (AC) and r 2,1 r 2,2 orthonormality (OC).3 A rigid transformation is a special form of affine transformation. For any affine transformation, in this case the rigid transformation, the second derivatives of u to x are zero, written as AC k,i,j (x) = 2 u k (x) x i x j = 0, (3.6) 32
55 for all k, i, j {1, 2}. The affinity term is similar to the smoothness constraint applied by Rueckert et al. [89]. Both terms encourage the local smoothness of the nonrigid transformation. The rotation matrix R is also orthonormal where R T R is an identity matrix. When R being a orthonormal matrix, for all i, j {1, 2} it much hold that OC i,j (x) = 2 r k,i r k,j δ i,j = 0, (3.7) k=1 where δ is the Kronecker delta function of { 0, if i j δ i,j = 1, if i = j, (3.8) From Equation (3.5) it follows that Equation (3.7) can be rewritten as u i (x) x j = r i,j δ i,j. (3.9) OC i,j (x) = 2 k=1 ( u k(x) x i + δ k,i )( u k(x) x j + δ k,j ) δ i,j = 0, (3.10) by replacing the rotation coefficients r k,i in Equation (3.7) with u k(x) x i to Equation (3.9). + δ k,i according The final regularization term is defined as the weighted sum of the AC and OC squared: 1 R(T ) = γ(x + T (x)) (3.11) x Ω M γ(x + T (x)) x Ω { M } w AC AC k,i,j (x) 2 + w OC OC i,j (x) 2, k,i,j where w AC and w OC are constant weights of the terms, and γ( ) is a coefficient mapping function to be applied on each voxel in the moving image which suggests i,j 33
56 the weight of the penalty is applied on the voxel. This formalization of the term is slightly different from that used in the study by Staring [100], in which an additional properness condition is included. The properness condition is introduced to ensure that the determinant of the orthonormal matrix is 1. In this condition the rotation is a proper rotation without inversion (mirroring). However, an inversion rotation is unlikely to occur in the nonrigid transformation given the affinity and orthonormal penalty have been applied. The simplest mapping function of γ( ) is a constant c in which all voxels in the moving image Ω M are equally penalized, shown as γ(x) = c. (3.12) For example, Rohlfing et al. [87] applied a uniformly weighted incompressibility term on all breast tissue. A binary mapping function requires a segmentation of various tissues and a weight to be assigned based on the stiffness of the tissue types. For instance by performing lesion segmentation in a DCE-MRI, the weight on lesion tissue is assigned to a constant and non-lesion tissue is 0, written as { c x lesion tissue γ(x) = (3.13) 0 otherwise. Another kind of function maps the intensity of an image to a certain range where the intensity values can imply the tissue types, such that the solid tissue have higher values while soft tissue have lower values. Ruan et al. [88] transform the intensity of a CT image into a range where most of voxel intensities are either 1 (bone tissue) or 0 (air). In DCE-MRI, voxel intensity does not have a clear one-to-one mapping to tissue types or stiffness. Therefore it is hard to determine the tissue type solely based on the intensity values. Most lesions in a breast are more rigid than healthy tissue in breast DCE-MRI and lesion tissue are usually significantly more enhanced in post-contrast images. Therefore, the tissue enhancement value is one indication of the stiffness of the lesion and healthy tissue. Based on the assumption that lesions get enhanced in postcontrast images, we obtain the regularization coefficients of each voxel by applying a 34
57 sigmoid function on the absolute differences between a post- and pre-contrast image. The resulting coefficient map has a higher value on more enhanced regions and lower value on less enhanced regions. This requires that the difference image is of a high quality with no obvious motion artifacts, since the regions of high intensity should be caused by enhancement not the motion artifacts. Therefore, a preliminary registration is necessary to remove the enhancement due to the motion artifacts in the difference image. Unlike the common registration practice where the moving image is registered to the fixed image, the preliminary registration aligns the fixed image to the moving image to create the regularization coefficients. There are later on in registering the moving image to the fixed image. Figure 3.3 illustrates the computation pipeline of the regularization coefficients. Given a fixed image f(x) and a moving image m(x), a preliminary rigid and then nonrigid (FFD) registration is performed such that f(x) is registered to m(x), obtaining a registered image f (x). An absolute difference of the registered fixed image from the moving image shown in Figure 3.3(b) is obtained by subtracting f (x) from m(x) and then taking the absolute value. The resulting image is then smoothed by a Gaussian filter (σ = 2), as shown in Figure 3.3(c). A sigmoid function is then applied on the smoothed difference image, resulting in a stiffness image (Figure 3.3(d)). The sigmoid function t( ) transforms the smoothed difference image s(x) to a new range (0-1) with a center α and scale β: t(x) = max min s(x) β (1 + e ( α ) ) + min, (3.14) where max and min are the maximum and minimum intensity of the difference image, α and β are determined by performing a k-means clustering method on the smoothed difference image and partitioning it into k groups with various intensity means. The highest intensity mean value is assigned to β and the standard deviation of that cluster is assigned to α. Therefore, the only user-defined parameter in the regularization term is the number of clusters k. The performance of the registration is demonstrated to be insensitive to the value of k in the range of 2 to 5 in the evaluation study in Section In the last step of the pipeline, the resulting image of the sigmoid filter is dilated 35
58 Fixed image fdfdfsfdsfd Moving image Register to using FFD Registered fixed image Image subtraction Smooth filter Sigmoid filter Dilation filter Tissue stiffness map (a) Work flow (b) Difference (c) Smoothed (d) Sigmoid Figure 3.3: The workflow in computing a tissue stiffness map as regularization coefficients. such that the surrounding area of the enhanced regions is also covered. The motivation is that the volume or shape of the lesion might have shrunk in the pre-registration. The dilation will ensure that the original area is covered. 36
59 3.2.5 Choice of fixed and moving images A 4D DCE-MRI includes a pre-contrast image and several post-contrast images after injecting contrast agents into the veins. In this study, the post-contrast image taken at two minutes after the contrast injection is selected as the fixed image in a registration, while the corresponding pre-contrast image and the remaining post-contrast images are aligned to it in a pair-wise fashion. In this study, both affinity and orthonormal terms are applied in the regularization term in registering the pre-contrast images to post-contrast images. However, only the affinity (smoothness) term is applied in the regularization term in registering between post-contrast images where the weight of orthonormal weight w OC in Equation (3.11) is set as zero. The reason for not using the orthonormal term is that the intensity differences among the post-contrast images are not as significant as those in between pre- and post-contrast images. 3.3 Experiments and results The registration framework in [89] is adapted in the evaluation, starting with a linear registration to correct global motions and then a B-spline based nonrigid registration to correct local motions. Various studies have demonstrated the validity this framework [71; 96; 100; 104], however, there is no comprehensive evaluation on the effectiveness of rigidity regularization in gross breast motion correction and lesion shape and volume preservation. Given the motion correction is usually the first component in the pipeline of a CAD system, it is important that the lesion is not distorted during the process. Lesion shape or volume changes can cause later steps in the pipeline, such as segmentation or analytis of the lesions, to be unreliable or even inaccurate. Therefore, we focus on evaluating the effectiveness of motion correction of lesions as well as the whole breast tissue. We apply different methods to compute rigidity terms, especially their coefficients, in registering the pre-contrast images to the post-contrast images. There are three registration schemes tested in registering pre-contrast images to their corresponding post-contrast images. The main differences in the methods are their regularization term or to be more precise, the coefficient γ(x) in the terms. In the first method, there is no rigidity term applied in the nonrigid registration, called the NR 37
60 scheme, serving as a baseline. Not using a regularization term is equivalent to setting the coefficients of the rigidity term to γ(x) = 0. In the second method we apply the rigidity term in which the tissue stiffness map or term coefficients are generated by manually segmenting lesions, called MR scheme. In the last method, we employ the proposed automated method (AR) to compute coefficients for rigidity terms. The cluster numbers k = 2, 3, 4, 5 are tested in determining the parameters of the sigmoid function Real and synthetic images In this study, synthetic and clinical images are used in the evaluation. There are 39 preand post-contrast image pairs included in the evaluation. The imaging protocol, lesion composition and volume distributions for all patients are specified in Chapter 2. In the evaluation of our method, synthetic pre- and post-contrast image pairs with known deformations are the ground-truth of the real DCE-MR image. Three clinical DCE-MR breast images without obvious motions are carefully selected to generate 3 10 synthetic images with simulated deformations. The images include small, medium and large volume sizes of lesions (1.5cm 3, 11.8cm 3, 22.3cm 3 ) or enhancement patterns (homogeneous, heterogeneous). Some demonstrative slices of the images are shown in Figure 3.4. A pre- and second post-contrast image pair (p 0, q 0 ) in each clinical DCE-MR breast image series is used in building a synthetic image pair (p 1, q 1 ). The enhanced lesions s 0 in the image pair are manually segmented from the difference images of the postcontrasts from the corresponding pre-contrast images. The lesion volume, location and intensity are used as ground truth in the validation of motion recovery effectiveness of the registration methods over lesion regions. In the deformation simulation, we randomly generate two rigid transformations (T r1, T r2 ), and two B-spline transformations with a grid point space of 10mm and 20mm. We later update these B-spline transformations to T b1, T b2 such that the lesions are rigidly deformed by enforcing the related control points to be zero. Note the B- spline grid space in the motion simulation is designed to be different from that in the registration to recover the motion. The motivation for this is to avoid the evaluation bias. We subsequently compose all the transformations to form the ground-truth T gt = 38
61 (a) pre (b) post (c) difference (d) pre (e) post (f) difference (g) pre (h) post (i) difference Figure 3.4: Demonstrative slices of three real images that are used to generate synthetic images. The columns from left to right show the pre-, post- and subtracted images. T b1 T r1 T b2 T r2, which are used to construct synthetic pre- and post-contrast images p 1, q 1 and lesion mask s 1. The formulation of the synthetic images and the resulting images in registration are listed in Table 3.1. We show some demonstrative results of rigidity and nonrigidly deformed images on post-contrast images in Figure 3.5 since lesions are not visible in pre-contrast images. However, all the simulated deformations are applied on the pre-contrast images in the evaluation. There are two types of ground-truth available in the synthetic images pairs: the original pre-contrast image and the known simulated deformations as shown in Table 3.2. The ideal registered pre-contrast images and lesions (p 2, s 2 ) are the same as 39
62 pre post lesion real p 0 q 0 s synthetic p 1 = T gt (p 0 ) q 0 s 1 = T gt (s 0 ) registered p 2 = T est (p 1 ) q 0 s 2 = T est (s 1 ) Table 3.1: The formulation of synthetic images from real images. (a) Original (b) Rigid (c) Nonrigid Figure 3.5: Example slices from (a) original post-contrast image, (b) rigidity and (c) nonrigidly transformed synthetic post-contrast images. the original pre-contrast images and lesions (p 0, s 0 ). The ideal estimated deformation u est (x) = T est (x) x is able to recover the simulated deformation u gt (x) = T gt (x) x. type result ground-truth intensity p 2 q 0 s 2 s 0 deformation u exp (x) T gt(x) x Table 3.2: Ground-truth for registration using synthetic images Evaluation method Target Registration Error (TRE) [96] is used to evaluate the degree of alignment between two corresponding voxels in terms of deformations: TRE = 1 (x + u est (x)) T gt (x), (3.15) N x Ω 40
63 where N is the number of voxels, T gt is the simulated transformation, u est is the estimated deformations obtained from various registration schemes. A smaller TRE value suggests that a registration can better recover the simulated motion. We also measure the recovery of the motion in synthetic pre-contrast images by measuring the intensity similarity with the corresponding original pre-contrast images, using root mean squared error (RMS) and normalized correlation (NC): RMS(A, B) = 1 N (A i B i ) N 2 (3.16) i=1 N i=1 NC(A, B) = (A i B i ) N (3.17) i=1 A2 i N i=1 B2 i where A i, B i is the intensity of the i-th voxel of images A and B, and N is the total number of voxels considered. Smaller RMS values and larger NC values suggest higher image similarity and correlation hence better registration performance. To evaluate the lesion preservation, we also compute changes of lesion volume by applying the estimated transformation T est on the lesion mask s 1. In testing of the real pre- and post-contrast image set, there is no ground-truth available, so the normalized mutual information is used to estimate the motion correction performance Registration settings We implemented and computed the rigidity coefficient map using ITK. We applied Elastix registration software [46] to perform all registrations by using the coefficient map we generated. All registration schemes start with an initial rigid registration to correct global breast motions. Three-level resolutions are used in the rigid registration to avoid the local minimum in the optimization. After that, an FFD-based nonrigid registration is applied to reduce local motion artifacts. In the four-level resolution scheme, the grid spacing is decreased from 64, 32, 16 to 8 voxels. The weight parameter w in Equation (3.1) in each scheme is determined by finding the value that can preserve the lesion volume while maintaining a comparable or better result than the unconstrained 41
64 FFD scheme and the computation time is less than 15 minutes. Based on the observation of performance and selection characteristics on synthetic cases, w = 1.5 is applied on both synthetic and real data in the experiments. One example is shown in Figure 3.6. The constant weights w AC = 100 and w OC = 1 in Equation (3.11) are empirically determined to balance affinity and orthonormal terms in the regularizer. Joint histogram is used to estimate the normalized mutual information in computing the similarity metric. A stochastic gradient descent optimizer is used to approximate the transformation parameters. 4.6 TRE (breast) Vol change 0.08 TRE (breast) Vol change w value Figure 3.6: The choice of weight parameter w in Equation (3.1) in AR(k=5) in a synthetic image pair. The value range of w [0.1, 1.5]. The running time of w > 1.5 is more than 15 minutes Results Table 3.3 shows the registration performance over the whole breast by presenting the deformation recovery (TRE) and intensity recovery (RMS and NC) values computed using the ground-truth. When using the known deformations as ground-truth, the TRE value reduces from an average of 3.42 voxels (1.98 mm 3 ) in the NR scheme (no rigidity term) to 1.5 voxels (0.87 mm 3 ) in the MR scheme. The MR scheme applied the rigid- 42
65 ity coefficients that were manually generated in the rigidity term. The AR scheme obtained the smallest TRE of 1.32 voxels (0.76 mm 3 ), in which the proposed coefficients are applied. Therefore, using a rigidity term in a b-spline based nonrigid registration can reduce the deformation error (TRE), especially when using the proposed method to compute the term coefficients. The improvement in registration performance by using rigidity terms is also reflected by a smaller dis-similarity (RMS and NC) between the registered and the original pre-contrast images. There are no significant differences in RMS or NC values across AR(k = 2, 3, 4, 5) and MR schemes, with p > 0.05 using Mann-Whitney U test. Therefore, the AR and MR schemes show similar performance in improving registration performance over the whole breast in the synthetic images. It also proves that our methods in computing term coefficients are not sensitive to the number of clusters that are used to compute the parameters in the sigmoid function. TRE RMS NC NR 3.42± ± ±0.01 AR(2) 1.32± ± ±0.01 AR(3) 1.55± ± ±0.01 AR(4) 1.60± ± ±0.01 AR(5) 1.60± ± ±0.01 MR 1.50± ± ±0.01 Table 3.3: Registration results on breasts in synthetic images. Table 3.4 presents the registration performance over the lesion regions. Changes in lesion volume are also measured in addition to registration errors and image similarities. As with the performance obtained over the whole breast, known deformations and intensity are better recovered by using a rigidity term, with either manually generated or automatically computed (AR) term coefficients. Compared to AR(2,3), the simulated deformations are recovered better in AR(4,5) and MR by having a lower TRE value of 0.1 voxel on average. The better performance of AR(4,5) and MR is also reflected in the intensity recovery of a smaller dis-similarity (RMS) and higher correlation (NC) as well. In addition, they outperform AR(2,3) in terms of volume preservation of lesions. Therefore, AR(4,5) and MR show a better registration performance over the lesion regions in terms of the recovery of the known deformations and image intensity as well as the volume preservation of lesions. 43
66 TRE RMS NC Vol change NR 1.09± ± ± ±11% AR(2) 0.16± ± ±0.04 4±4% AR(3) 0.13± ± ±0.03 3±2% AR(4) 0.10± ± ±0.03 1±1% AR(5) 0.10± ± ±0.02 1±1% MR 0.10± ± ±0.02 1±1% Table 3.4: Registration results on lesion regions in synthetic images. Based on the analysis of registration performance over the breast and lesion regions using the synthetic dataset, AR(4,5) and MR show a similar performance. AR(5) takes more time to run compared to AR(4) but shows a similar performance. Therefore, k = 4 is used to compute the rigidity term coefficients in the evaluation on the real image set. Unlike the synthetic image set in which the ground-truth is available, we only measure the similarity of the registered pre-contrast image with the post-contrast image to evaluate the performance over breasts and volume changes over lesion regions. As shown in Table 3.5, all schemes show the same NMI over breasts. However, the AR(4) and MR methods reduce the volume changes and there is no statistical significant differences between the two in the volume changes using Mann-Whitney U-test on the real image set. Breast regions NMI lesion regions Vol change NR -0.97± ±22.1% AR(4) -0.97± ±4.5% MR -0.97± ±3.3% Table 3.5: Evaluation results for 39 clinical images. Based on the analysis of the evaluation results in both synthetic and real images, the registration scheme using the proposed rigidity coefficients shows comparable performance to those that are assigned manually in both the whole breast and the lesion regions. 44
67 3.4 Summary In B-spline based nonrigid registration of breast DCE-MRI, it is important to preserve the information of breasts and lesions during the registration, as motion correction is required for accurate lesion segmentation, feature analysis and classification. In this study, we adapted a B-spline based nonrigid registration which has been extensively applied on DCE-MR breast images and has been shown to be able to effectively recover motions [71; 96; 104]. However, there is no comprehensive evaluation on the effectiveness of rigidity regularization in terms of gross breast motion correction, lesion shape and volume preservation. In addition, current spatial variant rigidity regularization terms require some kind of tissue segmentation to assign appropriate weights to different types of tissue. However, in lesion segmentation, a pre-requisite registration is required to obtain reliable kinetic features. We have proposed a method to automatically compute regularization coefficients using a sigmoid mapping method on an absolute differences of images obtained by a pre-registration. Moreover, the simulation of a pre- and post-contrast breast image pair was discussed. In the evaluation of our approach on synthetic and clinical images, the spatial variant rigidity constraint was shown to be able to improve registration performance over the whole breast as well as local lesion regions by showing a smaller target registration error and higher similarity between image acquisitions. Given no ground-truth was available for the clinical images, only changes in lesion volume were measured. As a result, the volume preservation of lesions by using our automatically generated coefficients is comparable to the manually assigned rigidity coefficients. Note that our focus was not to create a better lesion segmentation method, but to create a method that shows a comparable result in computing coefficients of a rigidity term. In the proposed method, we assumed that the enhanced tissue in a breast is more rigid than that is less enhanced, e.g. tumor tissue is much more enhanced than fat tissue, however, this is not always the case. Therefore, our method cannot be applied to lesions that are deformed significantly during the image acquisitions. In addition, non-enhancing or less enhanced lesion might have small regularization coefficients which make it difficult to preserve their shape or volume. 45
68 Chapter 4 Lesion detection In the previous chapter we demonstrated that a registration method using our proposed regularization term and its coefficients is able to correct motions between MR acquisitions while preserving lesion information. In this chapter, we propose a fully automated lesion detection algorithm on the whole DCE-MRI to identify regions containing suspicious features that are typically present in breast lesions. In the evaluation on 21 patients, we implemented and compared our method against another method in which it had shown good results in a different study. As a result, our method is able to achieve a better performance than the other method. It could be a future work to evaluate our method using a larger dataset. 46
69 4.1 Introduction In the interpretation of a breast MR of a patient, radiologists usually go through the patient s history, including previous images if provided, such as mammograms, ultrasounds and MRIs. After that, they may go through the DCE-MR sequences to check whether they are valid, whether they contain any severe motion artifacts or any metal artifacts, and whether the enhancement level is high enough to make a diagnosis. The subtraction image of a post-contract from the pre-contrast image is also useful in identifying and assessing abnormal findings. The selected post-contrast volume is usually the one that is acquired about two minutes after an injection of contrast agent. This is because most cancer tissues are known to reach their peak enhancement around this time. Radiologists also frequently refer to a maximum intensity projection (MIP) image to identify suspicious regions and characterize the distribution of enhanced tissues. The maximum intensity projection of a 3D MR volume consists of projecting the voxel with the highest intensity value on the visualization plane throughout the volume parallel onto the plane of the projection to form a 2D image. MIP enhances the 3D nature of the highly enhanced tissues of clinical importance, making them stand out from other tissues with low enhancement. Radiologists might also look at the MIP image to make a diagnosis on a breast MR case where there is little enhancement shown in the MIP image. In a DCE-MR image with extensive background enhancement, radiologists go through the pre-, post-contrast or subtraction contrast image slice-by-slice to closely analyze all suspicious regions. They also refer to various MR images of different protocols when necessary, such as T 2 images. Most radiologists make use of various computer-aided diagnosis (CAD) tools in the MR interpretation. Most commercial CAD systems provide a color-coded kinetic map that can be turned on and off when needed, such as CADstream R (Merge Healthcare Inc., Chicago, Illinois, USA), or DynaCAD R (Invivo, Gainesville, Florida, USA). The map is usually overlaid on a selected 3D MR volume to highlight the regions with certain kinetic features. For example, the regions with high initial enhancement and washout pattern are usually coded in red. The regions with a persistent time-signal pattern are coded in green. Using the color-coded map reduces the time required by radiologists to identify and characterize suspicious tissues. However, accurately identifying lesions and lesion differential analysis and diagnosis are difficult tasks and there 47
70 can be variations in the diagnosis opinions among radiologists. Therefore, an automated lesion detection system is necessary to enable a standard and reliable quantitative analysis. There are also circumstances where this system is even more valuable. For instance, in complex cases where there is extensive parenchymal enhancement, some small lesions with similar enhancement might be readily buried or shadowed. Another example is where there are numerous foci that are scattered or gathered in a DCE-MR image, and it can be time-consuming to identify suspicious lesions among the clusters. In this case, it is also difficult to record the exact location of the lesion among the numerous foci. This can make it hard for radiologists to review the diagnosis report in the follow-up studies. It is desirable to have an accurate lesion detection system to highlight suspicious regions and make a record associated with the diagnosis report. In addition to the clinical requirements and interests in practice, a lesion detection system is also needed to carry out other technical tasks in a CAD pipeline. There are efforts being made in computerized analysis and characterization of kinetic features [16; 20; 105; 111] and more recently morphological features [1; 2; 63; 73; 77; 78] of breast lesions. Feature analysis requires the selection of regions of interest (ROI) breast lesions in this case before quantitative measurements can be made. In addition, lesion segmentation has to be performed to extract more reliable and accurate lesion features. In the current literature, most regions of interest are selected manually [31] or semi-automatically [62]. Very few of them are carried out automatically without any user interaction. Three studies from Ertas et al. [21; 22; 23] achieved high sensitivity (equal or close to 100%) with low false positive rate on non-fat saturated DCE-MR images. In comparison, the false positive rate is very high on all existing studies when applied on fat saturated MR images [30; 108]. In clinical practice, the advantage of using fat-saturated dynamic sequences is that lesions are clearly visible. As such, there is a need to investigate detection of lesions in fat-saturated DCE-MR sequences. Ertas et al. [22] proposed a breast segmentation method using cellular neural networks and a lesion detection method using 3D template matching. The method started with identifying tissues that met the following requirements: 1. Showing 40% and higher maximum enhancement at two minutes after injection of contrast agents or, 48
71 2. Showing 60% and higher maximum enhancement at three minutes after injection of contrast agents or, 3. Showing 150% and higher maximum enhancement at eight minutes after injection of contrast agents. After identifying all tissues that met the requirements, two empirically determined thresholds were carefully chosen in order to get a balance between lesion detection sensitivity and false positive identifications. Firstly, a threshold on a normalized maximum intensity-time ratio map (nmitr) was applied to remove fatty tissues, muscles and parenchymal tissues. The map was then convolved with a 3D kernel to enhance compact structures, and the final lesion detection result was obtained by applying another threshold given as: nmitr 0.33 and convoluted result In the evaluation of 19 patients with 39 lesions, the system identified all of the lesions with 12 false positive detections in total. In their later work [23], Ertas et al.applied a similar system and included more rules to reduce false positives. After applying a threshold = 0.33 on the nmitr map, the same 3D kernel was convoluted on the map. The threshold applied on the convolution result was computed using fuzzy c-partitioning. In the phase of false positive reduction, a ROI meeting the following rules was labeled as a true lesion, volume 0.08cm 3, and 0.08cm 3 < volume < 1.00cm 3, eccentricity > 0.91, or volume 1.00cm 3, eccentricity > 0.97 The dataset included 39 patients with 76 mass lesions. The minimum lesion volume was 0.08cm 3 in the test dataset. In the test of the system they detected 97% of the lesions with an average false positive detection rate at 0.43 per lesion and 0.68 per case. However, the values in the rules defined in this system were carefully tuned to the testing data. Therefore, the performance of the system cannot be guaranteed due to the bias introduced by the testing data. 49
72 In their most recent study by Ertas et al. [21], a similar framework was adapted and some of the threshold and region exclusion rules were changed. In this study, instead of applying a user-defined threshold on an nmitr map, they performed a principle component analysis on the time-intensity curve for each voxel. A threshold equal to 75% of the cumulative Gaussian histogram of principal component values was used as the threshold to be applied on the most significant principal component map. The map was then convoluted with the 3D kernel filter. Another threshold = 0 was applied on the filtered image for the initial lesion localization. A method of false positive reduction was employed, similar to that in the previous study but with different values, 0.08cm 3 volume < 0.97cm 3 and eccentricity 0.95 or, volume 0.97cm 3 and eccentricity The system was tested on 24 women with 54 mass lesions and 23 healthy women with no lesion recorded. For a detection sensitivity of 100%, the false positive identification was 1.02/lesion and 1.17/case. Table 4.1 summarizes all the rules applied in the work by Ertas et al. [21; 22; 23]. Vignati et al. [108] and Giannini et al. [30] used a similar approach in their lesion detection systems for different fat-saturated DCE-MR images. They applied a global threshold on a subtracted mean intensity projection image to detect breast lesions. The regions with high intensity variations in post-contrast volumes and high oscillation in the last and second last sequences were excluded. Vignati et al. [108] tested the algorithm on 13 fat-saturated MR cases. It detected 25 out of 27 lesions with 26 false positive identifications per case. Giannini [30] tested the algorithm on both fat-saturated and non-fat-saturated data sets. The system detected 80% lesions in the fat-saturated group with a median of 11 false positives per case and 90% lesions in the non-fat-saturated group at a cost of a median of 16 false positives per case. The detection systems proposed by Ertas et al. [21; 22; 23] were all tested on nonfat-saturated MR images. They achieved high lesion detection sensitivity with less than 1 false positive per lesion and per case. However, the systems required multiple carefully tuned parameters in their thresholds and rules based on the examined dataset. Other studies that were performed on fat-saturated images showed lower lesion detection sensitivity and more false positive identifications [30; 108]. It is therefore desirable to develop a system that does not require user interaction or manual tuning of 50
73 [22] [23] [21] ME > 40% at 2 min, or ME > 60% at 3 min, or ME > 150% at 8 min, or nmitr 0.33, and convoluted nmitr 0.47 nmitr 0.33, and ME > 20% at 1 min, or ME > 40% at 2 min, or ME > 158% at 8 min, or V 0.08, or 0.08cm 3 < V < 1.00cm 3, Ecc > 0.91, or V 1.00cm 3, Ecc > 0.97 cgh > 75%, and 0.08cm 3 V < 0.97cm 3, Ecc > 0.95, or V > 0.97cm 3, Ecc > 0.86 Table 4.1: The rules defined in studies from Ertas et al. [21; 22; 23]. ME: maximum enhancement. nmitr: normalized maximum intensity time ratio. Ecc: eccentricity. V: volume. cgh: cumulative Gaussian histogram. parameters. In this chapter, a fully automated lesion detection system is proposed. It achieves the highest lesion detection sensitivity and lowest false positive detection rate compared to other studies on fat-saturated DCE-MR breast images [30; 108]. 4.2 Methodology There are three components in the lesion detection system: a pre-processing phase to reduce motion artifacts and extract breast regions, an initial lesion detection by applying automatically computed thresholds, and a false positive reduction phase Pre-processing Lesion identification in DCE-MR images usually makes use of kinetic information to extract suspicious regions. Motion artifacts can introduce inaccurate kinetic features that might cause false positive detections or mis-identification of lesions in a detection 51
74 system. The system first performs motion correction in between MR volume acquisitions of each patient by applying the image registration method proposed in Chapter 3. As another important part of image pre-processing, the whole breast is extracted by removing air and uninteresting tissues from the MR sequences. We apply a breast segmentation method [109] based on a Hessian-based sheetness filter [28]. The algorithm was demonstrated to be able to successfully extract breast regions on non-fat saturated 3D MR volumes. However, our DCE-MR sequences are fat-saturated volumes where the algorithm is not suitable. The solution to this problem is discussed in Section 4.3. Figure 4.1 (a) shows a representative slice from a post-contrast image. Figure 4.1 (b) superimposes a mask of the breast on the post-contrast slice. This mask removes the breast skin that is usually highly enhanced and might contribute to false positives. (a) post-contrast (b) breast extraction Figure 4.1: The left image shows a slice from a second post-contrast MR volume. The right image shows the extracted breast mask overlaid on the left image Initial lesion identification We compute subtraction image intensity, enhancement integral and wash-in rate for each pixel of a DCE-MR image. The three feature maps are used to represent kinetic information about the image. We also explored other kinetic information that was shown to be not as robust or effective in differentiating lesion and non-lesion tissue as these three features, such as wash-out rate, maximum enhancement and minimum enhancement etc. Each voxel of the feature maps show the corresponding kinetic information of that voxel in the DCE-MR image. Wash-in rate (WIR) measures how fast a tissue can absorb the contrast agent in 52
75 (a) integral (b) integral label (c) WIR (d) WIR label (e) Subtraction (f) Subtraction label (g) Ground-truth (h) Overlap Figure 4.2: The images on the left of the top three rows show the same slice of integral, WIR and subtraction feature maps. The images on the right of the the top three rows show the regions select by applying threshold on the corresponding feature maps. The image on the left at the bottom shows the ground-truth of the lesion location. The image on the right at the bottom shows the overlap of the three images above it. 53
76 approximately 2 minutes (t 2 ) after the injection of the contrast agent. The wash-in rate of a voxel x is defined as WIR(x) = I t 2 (x) I t0 (x) (4.1) where t 0 is the acquisition time of the pre-contrast image I t0, and I t2 is the post-contrast image which is taken after approximately 2 minutes. Since WIR is obtained through division, tissues of the low intensity value are more prone to noise or motion artifacts such that WIR may have a large value. Therefore, the WIR map is smoothed by a 3D Gaussian kernel with σ = 3. Figure 4.2 (c) shows a representative slice from a WIR map. The integral of the enhancement rate is the area under the normalized signalintensity curve over time. It shows the accumulated relative enhancement overtime which is less effected by division. Let n be the last time point of a DCE-MR sequence, the integral of the voxel x is defined as integral(x) = n i=1 I ti (x) I t0 (x) I t0 (x) (4.2) Similar to the WIR map, a 3D Gaussian kernel with σ = 3 is employed on the integral map. Figure 4.2 (a) shows the integral map for the slice. In addition to the integral and WIR maps, subtraction image intensity which is defined in Equation (4.3) is also used as a feature. This is because both integral and WIR are computed by dividing some of the post-contrast image from the pre-contrast image where some tissues with low intensity might have a larger value. The subtraction image can be used to overcome this problem and also provides information on the enhancement. Again, the post-contrast image is chosen at around 2 minutes. An example is shown in Figure 4.2 (e). Subtraction(x) = I t2 (x) I t0 (x) (4.3) As the feature values of the same tissue can vary significantly amongst patients, we convert the feature values computed in Equation (4.1) (4.2) (4.3) to percentile values by computing the cumulative histogram within the kinetic map of that patient. This normalizes the feature values of the same tissues among different patients. 54
77 We then apply a threshold on each kinetic map. This threshold is automatically generated from a training set. In the training set, we have a mask of lesion tissue and we can compute the mean feature value for each lesion. This value is then converted to a percentile in that breast, which is the cumulative histogram value. The reason for this is that the enhancement pattern of the contrast agent is subject to large physiological variation, especially depending on differences in vascular permeability. We compute the percentile for all lesions in the corresponding breasts. Some basic statistics can be applied to determine a value to be used as a threshold, such as the mean or median standard deviation. We chose to use the minimum percentile value as a threshold in order to detect all lesions. When applying this percentile on a new case, the percentile will be converted to be the feature value in that kinetic map. That feature value is used as the threshold on the new image to generate a binary image. The same technique is applied on all feature maps (integral, WIR and SUB), resulting in three binary images as shown in Figures 4.2 (d), 4.2 (b) and 4.2 (f). We then take the overlap of the three binary images as shown in Figure 4.2 (g). The small regions are removed to exclude noise or regions that are not of clinical importance. Up to this stage, we have obtained an initial detection result by applying automatically computed threshold on kinetic feature maps. The intuition is to find all the important regions with kinetic features of concern. It is known that blood vessels can have similar kinetic features as lesions. Therefore these need to be removed at the later stage. There are other enhanced regions in addition to lesions, such as parenchymal enhancement that will be picked up at this stage False positive reduction The aim of the false positive reduction process is to remove false positive regions, which are mainly comprised of blood vessels, motion artifacts, parenchymal enhancements and other enhanced tissues. In the initial lesion identification phase, only kinetic information is employed to remove background tissues. We include morphological features in this step to further remove other uninterested regions. Blood vessels account for a large portion of the regions extracted in the initial lesion identification. Blood vessels are long and thin in general, so a filter [28] that can enhance linear structures is applied to the binary mask. Figure 4.3a shows the 55
78 (a) Initial identification (b) Vesselness filter (c) Blobness filter (d) Detection result Figure 4.3: Maximum intensity projection (MIP) images show the process of false positive reduction. (a) Shows the MIP image of the initial identification result. (b) and (c) show the MIP image of results applying a vesselness filter and blobness filter on the initial identification results correspondingly. (d) Shows the final lesion detection results after applying SVM classification using extracted features of each region. initial lesion identification result in a MIP image and Figure 4.3b shows the MIP of the resulting image after applying the vesselness filter. The structure marked by an arrow is a fraction of a vessel that is more enhanced than other structures. Another filter is applied on the binary overlap image to enhance the regions with a more regular shape, like a blob. We named it a blobness filter. It is an N N 3 matrix defined in Equation (4.4) where N is determined by the pixel size in an image. The size is computed in Equation (4.5) to capture the smallest lesion of clinical importance 56
79 which is 5mm in diameter. For example N = 8, the blobness filter can be defined as, M(1) = M(2) = M(3) = N = 5mm pixel size in mm (4.4) (4.5) Ertas et al. [22] applied a similar 3D filter of size on a normalized kinetic map to enhance small lesions. They applied a carefully selected threshold (47%) on the filtered image to identify lesions. We have a very different usage of the filter. In 57
80 our case, the filter is applied on the initial identification mask to extract region features to be used in a classifier later on. We extract 6 features in each region generated in the initial identification phase. We compute the mean values of integral, WIR and subtraction intensity of the extracted regions. The kinetic features are the cumulative histogram (percentile) of these mean values in each patient. We compute the mean values of the regions in the resulting images processed by the vesselness and blobness filter. The volume is also used as a feature. The 6 features in each region are used in a support vector machine (SVM) classifier with a linear kernel which is implemented using MATLAB to reduce false positives. Figure 4.3 (d) demonstrates the final detection result after applying SVM to reduce false positive identifications. We apply leave-one-out validation to compute the false positive reduction number for each patient. In each validation, one patient is used as a test set. The remaining patients are partitioned into two set, all regions obtained in the initial detection from 19 patients are used for building the SVM module and the regions detected in the remaining 1 patient for choosing the best soft margin C i for each lesion i, defined as: C i = l C1 i CC i + (1 l) CC i (4.6) where lesion label l = 1 and non-lesion label l = 0. To choose the optimal soft margin values, we employ grid search for the parameters CC and C1 in the ranges of [5-20] and [ ] respectively, see Figure 4.4 for true positives and Figure 4.5 for false positives. In order to appreciate the processing of the two main phases, initial lesion identification and false positive reduction using SVM, we display the processing results by superimposing the result on the MIP of a subtraction image. Figure 4.6 (a) shows the MIP of a subtraction image. There are multiple foci on the left breast. The lesion location ground-truth is shown in Figure 4.6 (b) and is colored in red. In the initial identification result shown in Figure 4.6c, the lesion is successfully identified. There is a blood vessel, a focus on the left breast and 4 other unknown enhancements extracted on the right breast at this stage. After employing false positive reduction, the blood vessel and some enhanced regions are excluded in the final detection results as shown 58
81 TP CC C Figure 4.4: True positives using SVM with different combinations of CC and C FP CC C Figure 4.5: False positives using SVM with different combinations of CC and C1. in Figure 4.6d. It can be seen that there is still one false positive detection in each breast. 59
82 (a) MIP subtraction image (b) ground-truth (c) threshold results (d) After false positive reduction Figure 4.6: MIP image to show the initial lesion identification and false positive reduction result. (a) Shows a MIP subtraction image of a representative patient. (b) shows the ground-truth of lesion location colored in red. (c) shows the initial lesion identification. (d) shows the final lesion detection result after applying false positive reduction using SVM. The arrow shows the true lesion Performance analysis The performance of the lesion detection system is evaluated using ROC curves by comparing the lesion detection sensitivity against average false positive (FP) detections per lesion. sen lesion = sen case = FP lesion = FP case = number of lesion detected number of lesions number of cases where at least one lesion is detected number of cases number of false positive detections number of lesions number of false positive detections number of cases (4.7) (4.8) (4.9) (4.10) 60
83 The false positives per lesion (FP lesion ) suggest the number of false detections to be included in the result in order to detect a lesion. It sets the price on a true lesion being detected by the number of false detections. The false positives per case (FP case ) suggests the number of false ROIs in each case. This gives an idea how much extra effort manual effort that a radiologist has to put into each breast DCE-MRI, in order to identify the remaining lesions that are not detected by our system. 4.3 Experiment and results There are 39 patients available in the experiment. All of these patients have fatsaturated DCE-MR sequences but only 21 of them have the corresponding non-fatsaturated T2 -weighted images. The imaging protocol is as described in Chapter 2. As the breast extraction algorithm used in this system only works on non-fat-saturated images, we therefore can only choose these 21 cases with the corresponding T2 - weighted non-fat-saturated MR sequence available. For each patient, we registered the T2 -weighted non-fat-saturated MR sequence to the fat-saturated T 1 -weighted precontrast MR sequence by bspline-based nonrigid registration. We then extracted the breast mask from the registered non-fat-saturated MR sequence which is consistent with the breast location in the corresponding T 1 -weighted DCE-MR sequences. In the 21 valid patients, there are in total 50 lesions, 21 malignant lesions and 29 benign lesions. The lesion composition is listed in Table 4.2. There are 39 biopsyproven lesions, 21 malignant lesions, 15 fibroadenoma and 1 adenoma. The remaining lesions are benign lesions by diagnosis. The ground-truth for each of the lesions is then manually segmented and edited by a radiologist. A ROI is identified as a true lesion if the overlap with the ground-truth is greater than 40%, otherwise it is a false positive detection. We compared our system with the state-of-the-art method by Giannini et al. [30] for lesion detection in fat-saturated breast MR images. We implemented the algorithm [30] on their study [108]. We did not consider algorithms [21; 22; 23] that were designed for non-fat-saturated DCE-MRI. Our lesion detection system identified 600 suspicious lesion regions in the initial detection phase, comprised of 50 true lesions and 550 normal regions as false positive. In the false positive reduction phase, we excluded 298 normal regions from the ini- 61
84 type number breast carcinoma 1 16 IDC/DCIS 1 IDC 1 ILC 3 fibroadenoma 25 benign papillary lesion 1 adenoma 1 unknown 2 total 50 Table 4.2: Lesion composition Case 1 Case 2 Case 3 Case 4 Case 5 Case 6 Case 7 Case 8 Case 9 Case 10 Case 11 Case 12 Case 13 Case 14 Case 15 Case 16 Case 17 Case 18 Case 19 Figure 4.7: False positive reduction. Detection region # Reduction # Case 20 Case 21 tial detection results. The ROC curve in Figure 4.8 plots the lesion detection sensitivity against false positives by varying the output class membership probabilities in SVM. In order to achieve 100% sensitivity per lesion, the cost is 5.04 FPs per lesion and 12 FPs per case on average. The number of excluded false positives and detected regions for each case in shown in Figure 4.7.The quantitative measurements are listed in Table 4.3. In comparison, the lesion detection sensitivity obtained from Giannini et al. is not satisfactory based on the low sensitivity. The reason could be that the threshold defined in their system is not robust and the performance varies significantly among different dataset. 62
85 sensitivity per lesion false positive detections per lesion Figure 4.8: ROC curves that show the performance of lesion detection sensitivity against false positive detections per lesion by varying the output class probabilities in SVM. Study sen case sen lesion FP case FP lesion Proposed method 100% 100% Giannini [30] 85.7% 34% Table 4.3: Columns from left to right: study group, sensitivity per case (sen case ), sensitivity per lesion (sen lesion ), average false positive (FP) detections per case (FP case ) and average FPs per lesion (FP lesion ). The first row shows the result from our study, where we achieved the best performance in all measurements, except for the FP case. 4.4 Discussion In the experiments of our lesion detection system, there are two cases with no false positives, as shown in Figure 4.9. The images in the left column show maximum intensity projection of the subtraction image of the second post-contrast image from the pre-contract image. The images on the right show the detection result colored in red and they are superimposed on the corresponding MIP images on the left. 63
86 (a) MIP of subtraction image (b) detection result (c) MIP of subtraction image (d) detection result Figure 4.9: Two cases in which all lesions are correctly identified without any false positives. The images in the left column are the maximum intensity projection (MIP) of the subtraction image of the second post-contrast image from the pre-contrast image. The images on the right are the MIP of labels superimposed on the images on the right. The case in the top row is an extremely difficult case with extensive parenchymal enhancement according to the comments in the image interpretation report. The case in Figure 4.9c is a more complex case compared to the case in Figure 4.9a. It is an extremely difficult case with extensive parenchymal enhancement as stated in the patient diagnosis report. It can be seen from the MIP image in Figure 4.9a that there are extensive dense parenchymal enhancement and multiple small cysts in both breasts that can be challenging to radiologists as well. Our system has shown a high sensitivity and no false positives in this difficult case. It suggests that this system can potentially assist radiologists in challenging cases with extensive parenchymal enhancements. There are 3 patients with over 30 false detections. One of the patients has a combination of cysts and benign appearing diffuse stippled parenchymal enhancements. In the results of this case, our system detected 39 isolated regions which were labeled as false positive regions. The MIP image in Figure 4.10a shows the distribution of stippled parenchymal enhancements. The ground-truth in Figure 4.10b shows the location 64
87 (a) MIP of subtraction image (b) Ground-truth (c) Detection results Figure 4.10: A case with 41 isolated false positives. This case has numerous simple and complex cysts and benign appearing diffuse stippled enhancement. The image on the top is maximum intensity projection (MIP) of a subtraction image of the second postcontrast image from the pre-contrast image. The second row shows the MIP of groundtruth superimposed on the image on the right. Note that the cysts are not labeled in the ground-truth. The last row shows the detection results. 65
88 of a mass lesion. The cysts documented in the diagnosis report are not drawn in the ground-truth. The enhancement level of the lesion is very similar with that in some other tissues. The detection result is shown in Figure 4.10c in which a thick vessel is included in the detection result. The diffused stippled enhanced normal parenchymal tissues are also included. It seems that our detection system performs relatively poorly in cases with stippled parenchymal enhancements, especially in the cases where they are even more enhanced than lesions. This is caused by our detection system assuming that any suspicious regions are from the most enhanced regions. Another case in Figure 4.11a has 17 labeled lesions in total in the ground-truth in 4.11b. In the diagnosis report, this case was cited as an extremely complex MR. This is a good example of where our system is shown to be helpful. It can be very time consuming for a radiologist to identify and list the location of each individual lesion in a report. It is also hard to recover the lesion location based on the textual report of the image if a review is required later on. In addition, our automated lesion detection system can allow the extraction of lesion features. After aligning the image in the follow-up study to the image in the previous study using a registration, the detected lesions can be related to each other to monitor the lesion development, such as their shape, volume and kinetic patterns. The system is especial helpful in cases where there are numerous lesions needing to be monitored. In this case, the system showed 30 false positive detections with the majority being parenchymal enhancement, however some of these false positives could be true lesions. The remark from the radiologist was that the marked lesions were not exhaustive as it took too much time to identify each of the small lesions, there possibly could be more small lesions in the patient. Although the system shows quite a large number of false positives, the statistics might be a good characterization of the parenchymal enhancement pattern. This is useful in the follow-up (longitudinal) studies as it can be used to measure the changes of parenchymal enhancement patterns if there are any. The regions detected in the follow-up study can be removed if they are identified as false positives in the previous study, meaning that false positives can be reduced in the follow-up studies. In the last case (Figure 4.12a), there is an enhanced, spiculated mass with surrounding architectural distortion that measures 2.4cm in diameter and is labeled in Figure 4.12b. There are extensive foci that are scattered diffusely throughout the both 66
89 (a) MIP of subtraction image (b) Ground-truth (c) Detection results Figure 4.11: A case with 17 true lesions and 30 false positive detections, in which there are numerous simple and complex cysts and benign appearing diffuse stippled enhancement. The image on the top is the maximum intensity projection (MIP) of the subtraction image of the second post-contrast image from the pre-contrast image. The second row shows the MIP of ground-truth superimposed on the image on the right. Note that cysts are not labeled in the ground-truth. The last row shows the detection results. 67
90 breasts. In the results as shown in Figure 4.12c there are 35 false positives in this case, in which most of them are foci. In the measurements of the false positives, most of them are slightly larger than 5mm in diameter. The system could not eliminate these from the detections as any region larger than 5mm is of clinical importance. We also noticed that there is one thick blood vessel in each breast that is labeled as a lesion. Based on the observations, our lesion detection system has a 100% sensitivity in detecting benign and malignant lesions. However, it shows poor performance in cases with stippled parenchymal enhancement or foci, which are the major contribution to the false positives. The reason is that the volume of these regions is very small, however their largest diameter is still greater than 5mm, which makes them clinically important. Our system considers each individual region independently in the process of detection. Therefore, our system cannot exclude those clusters of enhanced regions without the knowledge that these clusters are stippled parenchymal enhancements. Future work in this area would incorporate the spatial information in between regions in the lesion detection system, in order to reduce the false positives that are caused by stippled parenchymal enhancement. 4.5 Summary In this chapter, we proposed an automated lesion detection system that simulated the interpretation procedure of a radiologist. This system is able to carry out tasks that a radiologist is not trained for, such as making classifications based on quantitative tissue features. These features are hard for a human being to capture through visual images only. The motivation of the study for clinical practice is to reduce the image interpretation time that is required by radiologists and potentially improve diagnosis performance, especially in complicated cases with extensive enhancements, numerous foci or small lesions. Another motivation is to enable later tasks to be performed in a CAD system. Our lesion detection system began with motion correction and breast tissue extraction. The initial identification of the suspicious regions was obtained by applying automatically computed thresholds on three kinetic feature maps. A SVM classifier was then employed on these regions to reduce false positive detections using both kinetic and morphological features. The evaluation results of 100% detection sensitivity and 68
91 (a) MIP of subtraction image (b) ground-truth (c) detection results Figure 4.12: A case with numerous foci in both breasts. The image on the top is the maximum intensity projection (MIP) of the subtraction image of the second postcontrast image from the pre-contrast image. The second row shows the MIP of groundtruth superimposed on the image on the right. The last row shows the detection results in which two thick blood vessels and most of larger foci are labeled as lesion enhancements 69
92 5.04 FPs/lesion show that our system achieves the best performance in fat-saturated DCE-MR breast images at the time of writing of the thesis. There are four main contributions in this study: 1. We applied a combination of kinetic features that were able to provide a good initial lesion identification results. Most of the kinetic features require some form of normalization by dividing from the intensity value in pre-contrast images. In this study, we used the subtraction intensity, that we observed was able to compensate other division based kinetic features. 2. The thresholds that were automatically computed by were in the form of a cumulative histogram value, which was proved to be able to tolerate large signal or feature variations amongst patients. This could avoid the problem of having to carefully tune parameters based on dataset. 3. We applied a combination of morphological features in the SVM classifier that were demonstrated to be able to reduce false positives. 4. By using our dataset, the experiments showed the the system was able to achieve a better performance than an automated lesion detection systems [30] in fatsaturated DCE-MR breast images. Although the proposed system is able to identify all the suspicious regions in our database. It is still radiologists decision what is the next procedure need to be carried out. When these highlighted regions are examined, the radiologists may discard many of them and only those that are considered suspicious will be further investigated. It could be a future work to evaluate whether this system can reduce the interpretation time and improve the diagnosis accuracy for radiologists. We observed that most of the false positives come from stippled diffused parenchymal enhancements and foci. These regions are clinically important based on BI-RADS standard. There are different practices amongst hospitals and/or radiologists in terms of dealing with these enhancements and foci. In our study, each of these isolated regions was counted as a false positive. In future work, the spatial relationship between these regions can be used to label them as a single cluster of parenchymal enhancement. This is also what we observed in the patient diagnosis report. One limitation of this study is that the 70
93 experiment is carried out using a small dataset that was acquired in the same medical center with similar imaging settings. Therefore, it is difficult to conclude on generalizability of the method. It could be a future work to evaluate this method using a larger database. 71
94 Chapter 5 Lesion segmentation In the previous chapter, our lesion detection method successfully identified all lesions with a reasonable amount of false positives in breasts. In this chapter, a lesion segmentation method is proposed to annotate lesion tissues in a user-defined region of interest. In this method, a region of interest in a DCE-MR image is modeled as a connected graph with local Markov properties of which each voxel is a node. Three edge potentials of the graph are designed to encourage the smoothness and continuity of the segmented regions. This chapter is based on an extension of our previous publication 1. Our segmentation method is the same as that discussed in our paper, however it is evaluated using a larger dataset as stated in Chapter 2. 1 Xi Liang, Kotagiri Ramamohanara, Helen Frazer, and Qing Yang. Lesion segmentation in dynamic contrast enhanced MRI of breast. In International Conference on Digital Image Computing Techniques and Applications (DICTA), pages 1-8. IEEE,
95 5.1 Introduction Interpreting breast DCE-MR images is challenging even for a radiologist, due to the large amount of information contained in 4D (3D + time) images. Various computeraided diagnosis (CAD) systems have been developed to allow faster and more accurate image interpretation and diagnosis [2; 32; 53; 56; 63; 68; 73; 77; 78; 85; 86; 113]. Automated lesion segmentation plays a key role in many other tasks in a CAD system, such as in the evaluation of lesion development, lesion detection, lesion characterization and lesion classification. Segmenting breast lesions in a DCE-MR image is challenging due to the inherent low signal-to-noise ratios, motion artifacts as well as high inter-patient variability. A number efforts are efforts have been made to address this problem. There have been a variety of studies of threshold-based methods on signal intensity of kinetics have been studied [19; 65; 72]. Lucas-Quesada et al. [65] employed a semiautomated method in a slice-by-slice manner. They computed a map for each voxel to represent the similarity of local signal enhancements to a representative time-intensity curve. This map was extracted in a user-selected ROI that encloses a lesion and then binarized by applying a user-defined threshold. Meinel et al. [72] proposed a threshold based segmentation method on a subtraction image. The threshold was extracted from rays that started from a point with the highest intensity radiating to the boundary of a user-defined ROI. In the post-processing stage of this lesion segmentation method, hole closing was applied to include non-enhancing or less enhancing areas (e.g. necrosis). Cui et al. [19] proposed a marker-controlled watershed method to segment lesions. A lesion was isolated on a 2D slice of a breast DCE-MR image that contained the largest area of the lesion. A Gaussian mixture model computed the markers or threshold of the background and the lesion intensities. The watershed method was then applied to the adjacent slices until there were no lesion tissues included in the next slice. The performance of the method was moderate on lesions of which the shape was irregular or the lesion had multiple large clusters. Fuzzy c-means (FCM) clustering is used in segmenting breast lesions in DCE- MRI [15; 80; 98]. Chen et al. [15] applied a FCM clustering method to segment a lesion in a small rectangular region that contained the lesion. One limitation of this FCM based algorithm is that it can successfully segment a lesion if the ROI is 73
96 small enough, however, the accuracy can be affected if the ROI includes many other surrounding enhancing tissues. Another limitation is that the spatial information about the voxels is not utilized. Therefore, it is difficult to compute an accurate contour for lesions with heterogeneous or rim enhancements that are common in malignant lesions. Shi et al. [98] refined the initial FCM segmentation method by using a 3D level set method. The level-set method was able to improve the lesion segmentation accuracy in their evaluation. Pang et al. [80] also applied FCM clustering in their initial breast lesion segmentation method. The gradient vector flow snake model was employed to refine the segmentation result by deforming a curve (lesion contour) until it reaches a balance between the internal (lesion) and external (normal tissue) force. However, only highly enhanced lesions were included in the evaluation of their study, which limits the generality of the evaluation performance. In this study, we model a ROI in a DCE-MR image as a graph with local Markov properties, of which each voxel is a node. The node potentials of the graph are defined as posterior probabilities of the class membership of the voxels using naive Bayes method. Three edge potentials are designed to encourage the continuity and smoothness of the segmented lesions. Ashraf et al. [5] have conducted work that is similar to the above, in which a multi-channel Markov random field (MRF) method is employed. The node potentials are computed using a Gaussian mixture model (GMM) and a binary function is used to compute edge potentials. Our approach differs fundamentally from their work as we have different definitions for both node and edge potentials. We also present and apply more comprehensive features in our GMM. Feature extraction plays an essential role in any classification problem. In DCE- MR images, the feature values can vary significantly amongst patients. It can affect the classification accuracy if the features extracted from some patients are used in estimating the tissue types in other patients. We propose a robust normalization of intensity and kinetic features across all MR images such that their values are within approximately the same scale and range. The proposed normalization method is shown to be superior to a widely used fixed range normalization method when applied to the features extracted in the DCE-MR images in our experiments. We evaluate our method by comparing the segmentation results against the groundtruth. We measure the performance of the proposed segmentation method by using three approaches to compute the edge potentials. The results are compared with a 74
97 widely used FCM method and a recently proposed GMM-MRF method. Both the GMM-MRF and proposed methods have a high AUC value that suggests that both methods generally perform well in separating lesion and non-lesion voxels. However, it is worth noting that the lesion voxels are only 2.89% of the total voxels. Therefore, a more robust validation method is required to measure the overlap between the segmented lesions with the ground-truth. The proposed method is demonstrated to be the most accurate in the region-wise evaluations on 72 lesions, with the highest overlap rate of 47%. 5.2 Methodology Our segmentation method is composed of six steps: (1) selecting a region of interest (ROI); (2) extracting features on each voxel in the ROI; (3) creating a naive Bayes classifier using a training set; (4) modeling the ROI as a Markov random field (MRF); (5) making inferences from the MRF using loopy belief prorogation; (6) connectedcomponent labeling and object selection. The system requires a user to select a 3D ROI that includes a lesion. Each ROI contained a whole lesion with a reasonable margin from the boundary of the ROI. There does not need to be a particular shape or a minimum bounding box for the lesion. As a demonstration, Figure 5.1 shows some selected slices in a rectangular ROI that contains a lesion. The figure represents a 4D ROI of which each row shows a lesion slice and each column represents the time series of the lesion Feature extraction Various features are extracted for each voxel in a ROI, including peak intensity, peak enhancement, time-to-peak, wash-in rate, wash-out rate and integral as illustrated in Figure 5.2. Let T be the set of all time steps included in a DCE-MR ROI, I t i be the intensity of the i-th voxel at the time step t T, the peak intensity (PI) of the voxel is defined as: PI i = max t T It i. (5.1) 75
98 slice 25 slice 27 slice 29 slice 31 slice 33 slice 35 slice 37 slice 39 slice 41 slice 43 slice 45 t = 0 t = 1 t = 2 t = 3 t = 4 t = 5 Figure 5.1: A 4D ROI (3D spatial + time dimensions). Each row represents a full dynamic sequence of a slice. Each column represents a time point of different slices in the lesion acquisition. 76
99 Enhancement Peak enhancement PE E WOR t WIR E PE TTP Time-to-peak TTP t t0 t1 t2 t3 4 t5 t Time Figure 5.2: Kinetic features that are defined based on time-intensity curve: peak enhancement (PE), time-to-peak (TTP), wash-out rate (WOR) and wash-in rate (WIR) of a voxel. The TTP in this example is t 3. The enhancement (E) at each time step is computed by dividing the voxel intensities in each post-contrast image by the corresponding intensities of pre-contrast image. The enhancement of the i-th voxel at the time step t is defined as: E t i = It i I 0 i I 0 i, (5.2) where I 0 i stands for the intensity of the i-th voxel in the pre-contrast image. To make all tissue enhancements (E) positive, the negative intensities are assigned to a default positive number. In the computation of the enhancements, Equation (5.2) can be regarded as the first-order approximation of the contrast agent concentration in breast tissues. However, a shortcoming of this method is that the enhancements can be sensitive to noise and motions. Therefore, a Gaussian filter (σ = 3 voxels) is applied on the enhancement map to reduce the noise. Peak enhancement (PE) of the i-th voxel is defined as the maximum enhancement 77
100 at all time steps: PE i = max t T I t i I 0 i I 0 i. (5.3) Time-to-peak (TTP) is the time step when a voxel reaches its maximum intensity. Typically malignant lesion tissues reach their maximum values at an earlier time [49]. Wash-in rate (WIR) measures the concentration rate in a tissue before it reaches its maximum intensity, defined as: WIR i = PE i TTP i. (5.4) Wash-out rate (WOR) measures the rate of a contrast agent losing the concentration in breast tissues, defined as: where t end is the last time step. WOR i = PE i E t end i, (5.5) t end TTP i The integral (area under the kinetic curve) at i-th voxel is defined as: Integral i = n E t i. (5.6) t T Some demonstrative slices of 3D feature maps of PI, PE, WIR, WOR and enhancement (E) at all time points are shown in Figure 5.3. These are extracted from the slices shown in Figure Feature normalization Based on the Breast Imaging and Reporting Data System (BIRADS) [4], the patterns of the enhancing parenchymal are classified as minimal/mild, moderate and marked. Normal tissue enhancement (especially moderate and marked) may cause false positive diagnosis of lesions. The cases with moderate and marked enhancement amount to 20% of patients as recorded in a study conducted by Jansen et al. [44]. Therefore, it is important that a segmentation method can perform well in cases with moderate or marked background enhancement. 78
101 Slice 25 Slice 27 Slice 29 Slice 31 Slice 33 Slice 35 Slice 37 Slice 39 Slice 41 Slice 43 Slice 45 PI PE WIR WOR E 1 E 2 E 3 E 4 E 5 Figure 5.3: Demonstrative slices showing the extracted features. Each row shows a collection of all features for a slice. Columns from left to right: peak intensity (PI), peak enhancement (PE), wash-in rate (WIR), wash-out rate (WOR) and enhancement (E) in all post-contrast images. Another challenge in a learning based segmentation method in DCE-MR breast im- 79
102 ages is that different patients can have very different enhancements for many reasons, such as variations in body size and menstrual cycles. Therefore, the enhancement level of the lesion tissues in one patient can be similar to that in another patient s normal parenchymal tissues. Therefore, tissue classifications require reducing the overlap of the feature values. In supervised classifications, normalization is usually applied to the features extracted from different patients. The most common method is to map the original value of each feature x to a fixed range between α and β, (β > α), by applying the following linear transformation: x = β α (x min) + α, (5.7) max min where max and min are the maximum and minimum value. However, this can be very unreliable when applied to the enhancement features as the voxels of very low intensity in a pre-contrast image can have large enhancement values due to noise or motion artifacts. Therefore, a safer upper bound is more a reliable method of scaling the feature ranges than the maximum value. We transform the feature values using the following normalization method: x = x µ + 3σ, (5.8) where µ and σ are the mean and standard deviation of a feature, and µ + 3σ is larger than roughly 99.7% of feature values. This normalization method is more reliable than the fixed range method as it ignores the outliers that cause most of the feature values that are scaled to have small values. By doing so, the kinetic features of the same type in different patients have an approximately similar value Markov random field The segmentation method proposed in this study is based on a probabilistic framework that is constructed using graph models, implemented by Schmidt [95]. A ROI in a DCE-MR image is modeled as a graph where the i-th voxel is represented by a class node y i and a feature vector node x i. The graph connects the neighboring class and/or feature vector nodes to form a Markov random field (MRF). There are two class mem- 80
103 berships for each node, lesion and non-lesion. All features in each class are assumed to have Gaussian distributions. The class membership y in graph Θ can be computed by solving the following problem: ŷ = arg max y φ i (y i, x i ) i Θ j N{i} where N{i} is the set of neighboring nodes to the i-th node. potentials are denoted as φ( ) and ψ( ) respectively. ψ i,j (y i, x i, y j, x j ) (5.9) The node and edge The node potential φ i ( ) shows the compatibility between the voxel feature x i with the label y i. In this study, the node potential is defined as the posterior probability of the node based on naive Bayes (NB) classification method, written as φ i (y i, x i ) = p(y i x i ). (5.10) In order to compute the posterior probability above, we compute all parameters in a naive Bayes classifier using a training ROI set where the same group of features is extracted for each voxel. In Markov random models, the edge potential ψ( ) measures the compatibility of neighboring nodes. We propose three approaches to computing the edge potentials. The first approach connects neighboring features nodes, as shown in Figure 5.4a. It determines the class of a node by its features as well as those of its neighbors. The edge potential function is defined as, ψ i,j (x i, x j ) = α e x i x j, (5.11) where the i-th node and j-th node are neighbors, is the Euclidean distance and α determines the weight. The edge potential is then normalized using Equation (5.8). The segmentation scheme using the Markov random model with the edge potential by connecting feature nodes is referred to as NB-MRF-F. In the second approach, the graph connects neighboring class and feature nodes, as shown in Figure 5.4b. In this way, both neighboring features and classes contribute to the node labeling in the MRF. The edge potential function is defined as ψ i,j (x i, y i, x j, y j ) = α e φ i(y i,x i ) φ j (y j,x j ), (5.12) 81
104 X X X X X X y y y y y y X X X X X X y y y y y y X X X X X X y y y y y y (a) NB-MRF-F (b) NB-MRF-NP Figure 5.4: Circles with y represent class nodes and circles with x represent feature nodes. (a) NB-MRF-F where neighboring features are connected. (b) NB-MRF-NP where neighboring features and classes are connected respectively. where the i-th node and j-th node are neighbors, represents the Euclidean distance and α determines the weight. The segmentation scheme using the edge potential defined in Equation (5.12) is denoted as NB-MRF-NP where NP is short for the neighboring node potentials. In the third approach, the graph connects the neighboring voxel intensities in the subtraction images of the second post-contrast image from the pre-contrast image. The edge potential is weighted by the mean peak intensity (PI) of the voxel and its neighbors, ψ i,j (PI i, PI j ) = α ( ) 2 PIi PI j + 1. (5.13) 2 PI max The segmentation scheme using Equation (5.13) as edge potential is denoted as NB- MRF-I. The edge potentials have a higher value of which the i-th and j-th nodes have similar feature values in the NB-MRF-F, similar node potentials in the NB-MRF-NP or high peak intensity in NB-MRF-I. All of these are designed to encourage the continuity of segmented lesions but with a different focus. The optimal label y in this study (Equation (5.9)) is determined using a loopy be- 82
105 lief propagation (LBP) [76] that is usually used to make inference in MRF models. Figure 5.5 presents some demonstrative results in the same slices in Figure 5.1. The prototype enhancement curves of lesion and non-lesion voxels for the ROI are shown in Figure 5.6. The curves are computed by taking the average of all the enhancements of the corresponding voxel at each post-contrast time point. In the post-processing method, a 3D connect-component labeling operation is applied to the binary ROI. The voxels in the component with the largest voxel number are labeled as lesion tissues and the remaining are labeled as non-lesion tissues. 5.3 Experiments Materials The imaging protocols were as specified in Chapter 2. There are 72 lesions included in the evaluation of various segmentation schemes, incorporating 28 malignant and 44 benign lesions, as shown in Table 5.1. There are 3 foci and 8 lesions with unknown types excluded from the database. Type Number Breast carcinoma 16 DCIS 6 IDC/DCIS 2 IDC 1 ILC 3 Cyst 2 Fibroadenoma 40 Benign papillary lesion 1 Adenoma 1 Total 72 Table 5.1: Lesion composition. *: the type of the breast carcinoma is not provided in the report. The contour of each lesion is manually extracted from the subtraction image of the second post-contrast images from the corresponding pre-contrast images. An experi- 83
106 slice 25 slice 27 slice 29 slice 31 slice 33 slice 35 slice 37 slice 39 slice 41 slice 43 slice 45 PI GT Label Figure 5.5: The segmentation result using NB-MRF-F. Each row shows (left to right) the peak intensity (PI), segmentation ground-truth (GT) and NB-MRF-F labeling results for a slice. enced radiologist then reviews this initial lesion segmentation. The regions of interest are selected by including 10 voxels in each direction from the minimum bounding box of each lesion. 84
107 Enhancement Lesion Non lesion Time Figure 5.6: Prototype enhancement curves found by segmenting lesion voxels and non-lesion voxels within the ROI. The curve of lesion/non-lesion voxels is computed by taking the average of the enhancement of all the corresponding voxels Segmentation schemes We use an MRF to model regions of interest in the breast DCE-MRI. The same posterior probability that is computed based on naive Bayes (NB) method is applied as node potential. We apply different edge potentials in the MRFs, categorized by connecting neighboring features, node potentials and peak intensities, known as NB-MRF-F (Equation (5.11)), NB-MRF-NP (Equation (5.12)) and NB-MRF-I (Equation (5.13)) respectively. These are compared with the segmentation methods using a naive Bayes (NB) classification, FCM clustering and the Gaussian mixture model (GMM) and GMM-MRF methods. The FCM based method serves as a baseline for performance comparison purpose. In the NB and FCM-based segmentation methods, a connected component labeling is followed by a hole filling as there might be some necrotic area in the lesion that may show low enhancement. In the GMM-MRF method, the node potentials are computed using GMM and the edge potentials are defined as: ψ i,j (y i, y j ) = e βi(y i y j ) (5.14) where I(y i y j ) is an indicator function that equals to 1 when y i y j and 0 otherwise. It is the same as used by Ashraf et al. [5]. 85
108 5.3.3 Feature selection and normalization We applied a linear regression test on all the features to the ground-truth. Time-topeak and washout rate are found to be not significant with p > 0.05, while wash-in rate (WIR), peak intensity (PI), peak enhancement (PE) and enhancement in all postcontrast images (E t, t 2, 3, 4, 5) are significant. However only WIR, PI, PE and E t, t 2, 5 are applied in all the segmentation methods. The reason for not using E t, t 1, 3, 4 is to avoid too many similar features. Most feature values normalized by the proposed method fall between 0 and 1.5, therefore, for purpose of comparison, the parameters in the fixed range method are set to be α = 0, β = 1.5 in Equation (5.7). These two methods are compared on all the selected features grouped by lesion and non-lesion voxels. Table 5.2 shows the mean and standard deviation of the feature values using the two normalization methods in the lesion and non-lesion groups. By using the fixed range method, the value ranges of different features are larger in both lesion and non-lesion groups compared to our method. By using our method, the mean values of the features in the lesion group ranges from whereas the means of those computed by the fixed range method falls between Similarly, the mean values in the non-lesion group range from in the proposed method and between in the fixed range method. In both lesion and non-lesion groups, the standard deviation of all features normalized by our method are smaller, except for the wash-in-rate. As shown in Table 5.2, the differences of mean values between the lesion and non-lesion groups are larger using our method in the feature E t 2, E t 5, PI and WIR. The fixed range method shows a larger difference (0.63) in the feature of the PE but the value in the proposed method itself is also large enough (0.54). We compute the Jaccard index to examine the overlap of normalized feature histogram in lesion and non-lesion groups using the proposed and fixed range method. The results are shown in Table 5.3. For each feature, we illustrate the normalized histogram in Figure Based on the above observations, it is more reasonable to apply the proposed method to normalize the features in the voxel classification based segmentation in DCE-MR images. In terms of the edge potential function in equation (5.11) that requires the values in different features lie in a similar range, the pro- 86
109 Proposed method Fixed range method Lesion Non-lesion Diff Lesion Non-lesion Diff E t ± ± ± ± E t ± ± ± ± PI 0.71± ± ± ± PE 0.93± ± ± ± WIR 0.60± ± ± ± Table 5.2: Feature values (mean ± standard variation) computed using the fixed range and proposed normalization methods on the lesion and non-lesion voxels. The third and sixth columns (Diff) show the differences of mean values in the two methods. posed normalization method is shown to be superior. Therefore, our method is applied on all the features in the experiments. All segmentation schemes utilize the same feature values. the proposed fixed range E t E t PE WIR PI Table 5.3: The Jaccard index of features: enhancement at the 2 and 5 time phrase (E t 2, E t 5 ), peak intensity (PI), (peak enhancement) PE and wash-in rate (WIR) on lesion and non-lesion groups using the proposed and fixed range normalization methods Evaluation method The segmentation performance of different methods is evaluated by a region-wise approach and voxel-wise approaches. We employ a leave-one-out cross validation scheme in all the segmentation schemes and AUC is computed to measure the segmentation performance at voxel level. In addition to the voxel-based evaluation, we also measure the overlap between the segmented lesion region (A) and the ground-truth 87
110 Lesion Non lesion (a) Proposed method Lesion Non Lesion (b) Fixed range method Figure 5.7: The normalized histogram of E t 2 in the lesion and non-lesion groups using the proposed (top) and fixed range (bottom) normalization methods. E t 2 : enhancement in the second post-contrast image. (B), defined as, Overlap = A B A B Results Figure 5.12 shows the ROC curves of various voxel classification results of which the NB-MRF-F and NB-MRF-NP overlap with each other. The area under curve (AUC) is computed to evaluate the segmentation performance at voxel level, as shown in the first column of Table 5.4. Our implementation of the GMM-MRF has produced an AUC value of 0.97, which is consistent with Ashraf et al. [5], who reported a similar AUC value of The AUC of the proposed NB-MRF-F and NB-MRF-NP schemes have a similar score of Note that no small connected component removal or hole filling methods are performed in computing the AUC for the NB (0.95) or FCM (0.74) 88
111 Lesion Non lesion (a) Proposed method Lesion Non lesion (b) Fixed range method Figure 5.8: The normalized histogram of E t 5 in the lesion and non-lesion groups using the proposed (top) and fixed range (bottom) normalization methods. methods. It is worth noting that only 2.89% breast tissues are lesions in our dataset. Therefore, a high AUC value only suggests a general segmentation performance over the whole ROI regions. We hereby measure the overlap of the lesion segmentation results with the ground-truth ( A B ) which is more valuable. A B As a result, the proposed methods (NB-MRF-F, NB-MRF-NP and NB-MRF-I) have consistently performed better than the existing methods on all the four groups of regions of interest by having higher lesion overlap rates. It is worth noting that the FCM based method can achieve a much higher overlap rate of 53.31% if the minimum bounding box of an enhancing lesion is the region of interest in our mixed group. However, its accuracy rapidly decreases to 9% when 10 surrounding voxels in all directions are included. Therefore FCM is not a good choice for lesion segmentation when a small enough bounding box cannot be obtained. 89
112 Lesion Non lesion (a) Proposed method Lesion Non lesion (b) Fixed range method Figure 5.9: The normalized histogram of peak enhancement in the lesion and nonlesion groups using the proposed (top) and fixed range (bottom) normalization methods. Method AUC Overlap FCM ±17 GMM ±27 GMM-MRF ±30 NB ±31 NB-MRF-F ±30 NB-MRF-NP ±30 NB-MRF-I ±31 Table 5.4: Segmentation results of AUC and overlap rate with the ground-truth. We label a lesion as successfully segmented if the overlap with the ground-truth is no less than 40%. Table 5.5 shows the statistics of the lesion volumes that are successfully segmented and misclassified by using different segmentation schemes. 90
113 Lesion Non lesion (a) Proposed method 8 x 10 3 Lesion 6 Non lesion (b) Fixed range method Figure 5.10: The normalized histogram of peak intensity in the lesion and non-lesion groups using the proposed (top) and fixed range (bottom) normalization methods. The proposed NB-MRF-F, NB-MRF-NP and NB-MRF-I methods show similar results and they have segmented more lesions than that using FCM, GMM, GMM-MRF- or NB methods. Method Number Mean Max Min Y N Y N Y N Y N FCM GMM GMM-MRF NB NB-MRF-F NB-MRF-NP NB-MRF-I Table 5.5: The statistics on the successfully segmented (Y) and misclassified (N) lesions, including the number of lesions, mean and maximum and minimum volumes. 91
114 0.04 Lesion Non lesion (a) Proposed method Lesion Non lesion (b) Fixed range method Figure 5.11: The normalized histogram of wash-in rate in the lesion and non-lesion groups using the proposed (top) and fixed range (bottom) normalization methods. The three proposed segmentation schemes detect more lesions (55/72) than both FCM (6/72) and GMM-MRF method (46/72). Our methods can also segment smaller lesions with the mean volume of 1.23 to 1.24 cm 3 when compared to those segmented FCM of 4.56 cm 3 and by GMM-MRF of 1.30 cm 3. Our methods are able to segment lesions with a volume that is larger than 2.95 cm 3. Both the GMM-MRF and our methods can segment lesions of a volume as small as 0.04 cm 3. To date, there are no methods that are able to segment lesions of size 0.01 cm 3. Figure 5.13 shows the lesion volume distributions of segmented and missed lesions using the existing schemes and our methods. 92
115 1 True positive rate FCM NB NB MRF F NB MRF NP NB MRF I GMM GMM MRF False positive rate Figure 5.12: ROC curves of all the segmentation methods. Note that the ROC curves of the NB-MRF-F and NB-MRF-NP methods overlap with each other. 5.4 Summary In this chapter, we proposed three edge potential functions in Markov random field based method to segmentation lesions in a user-defined region of interest in DCE-MR images. Among these three edge potential functions, the node potential based function showed the best results by having the largest number of successfully segmented lesions. We also proposed a normalization method applied to the intensity and kinetic features extracted in tissue classifications. The proposed normalization method showed better segmentation performance by having a significantly higher lesion overlap rate, compared to the FCM and recently proposed GMM-MRF methods. A future work could be to evaluate the performance by varying the size of regions of interest to observing the dependency between the segmentation results and the ROI size. 93
116 number * overlap >= 40% overlap < 40% volume (cm 3 ) (a) FCM number overlap >= 40% overlap < 40% volume (cm 3 ) (b) GMM number overlap >= 40% overlap < 40% volume (cm 3 ) (c) GMM-MRF number overlap >= 40% overlap < 40% volume (cm 3 ) (d) NB Figure 5.13: Lesion volume distributions of successfully segmented (overlap 40%) and missed lesions by using (a) FCM, (b) GMM, (c) GMM-MRF and (d) NB. 94
117 number * overlap >= 40% overlap < 40% volume (cm 3 ) (e) NB-MRF-F number * overlap >= 40% overlap < 40% volume (cm 3 ) (f) NB-MRF-I number * overlap >= 40% overlap < 40% volume (cm 3 ) (g) NB-MRF-NP Figure 5.12: Lesion volume distributions, mean and standard deviation of successfully segmented (overlap 40%) and missed lesions (overlap 40%) by using (e) NB- MRF-F, (f) NB-MRF-I and (g) NB-MRF-NP. 95
118 Chapter 6 Feature analysis and lesion classification In the previous chapters, we have introduced our image registration, lesion detection and lesion segmentation methods. This chapter focuses on the morphological, textural and kinetic feature analysis and classification of lesions. Our method of morphological analysis was published in the paper below 1. The main difference in this chapter from the findings of that paper is in the evaluation, where we test our method using a large dataset. 1 Xi Liang, Kotagiri Ramamohanarao, Helen Frazer, and Qing Yang. A lesion shape and margin characterization method in dynamic contrast enhanced magnetic resonance imaging of breast. In the 9th IEEE International Symposium on Biomedical Imaging (ISBI), pages IEEE,
119 6.1 Introduction In image processing, feature extraction in a ROI is to represent the ROI by mathematical measurements. In the context of the Breast Imaging-Reporting Data System (BI-RADS) of DCE-MRI published by the American College of Radiology (ACR) in 2003, a tumor mass is represented by morphological (shape, margin), textural (internal enhancement) and kinetic (dynamic enhancement) descriptors. There are efforts being made to loosely link mathematical measurements to the BI-RADS lexicons or descriptors. In this section, we review the existing lesion feature analysis and classifications Kinetic features Kinetic features have been extensively investigated either by fitting them within the parameters of pharmacokinetic models or applying pattern recognition to extract tissue enhancement patterns. Kinetic features have been applied in breast lesion detection [8; 21; 22; 23; 30; 108], segmentation [5; 15; 80; 98], classification [6; 16; 20; 105; 111] and visualization [107] Pharmacokinetic model Pharmacokinetic models (PKM) are used to recover the origin of enhanced MR signals in breast lesions by modeling vascular and tissue-specific parameters that influence the perfusion of contrast agents. PKM represents a human body as compartments through which contrast agents flow, such as in the blood plasma and surrounding tissues. A widely used PKM is the two-compartment minimal model [105] as shown in Figure 6.1. After the contrast agents are injected into the plasma, they can reversibly flow between the plasma and surrounding body tissues and eventually filtered by the kidney and cleared out in the urine. The extravascular extracellular space (EES) is part of the tissue of the human body that can absorb and release the contrast agents. The volume fraction of the EES per unit volume of the tissue is denoted as v e, and different tissues usually have different v e values. The concentration (C t ) of the contrast agents in tissue C t is defined as, C t = v e C e, (6.1) 97
120 Contrast agent injection Plasma C ( ) p t trans K Tissue extravascular extracellular space (EES) C ( t e ) C t ( t) Kidney Figure 6.1: Two-compartment pharmacokinetic model. After contrast agents are injected into blood circulation, they are rapidly distributed into intravascular volume. There is a reversible perfusion between plasma and the extravascular extracellular space (EES) of tissues, with a transfer constant K trans that is tissue dependent. The contrast agent is slowly filtered by the kidney and eventually cleared in the urine. where C e is the concentration in EES. The flow of contrast agent from plasma into the EES per unit tissue is v e dc e (t) dt = P S(C p C e ), (6.2) where C p is the concentration in plasma, P is the permeability coefficient, S is the surface area of the EES. Assuming the flow is proportional to the area of EES and to the concentration difference between the plasma and EES, the concentration of the contrast agent in the tissue C t can be rewritten as below by combining Equation (6.1) 98
121 and (6.2), dc t dt = Ktrans (C p C e ) = K trans (C p C t v e ) = K trans C p k ep C t, (6.3) where K trans is the permeability surface area product per unit volume of the tissue, or the volume transfer constant between the plasma and EES, k ep is the flux rate constant between the plasma and EES that is equal to the ratio of the transfer constant to the EES volume fraction (k ep = K trans /v e ). The solution to Equation (6.3) is written as below [105] with the initial conditions that C p = C t = 0 at t = 0, dc t (t) dt = K trans C p (t) k ep C t (t), (6.4) where the parameters K trans and k ep can be computed by fitting the time course of the concentration of the contrast agents at each voxel in equation (6.4). There are several limitations to the kinetic analysis using PKM. Firstly, it assumes that the time intensity curve (TIC) can be represented by certain mathematical formulation that could be very approximate. The second is that there is no ground-truth or gold standard to confirm the validity of the estimated parameters. Thirdly, PKM has a strong dependency on imaging protocols and/or machines [25] which requires a new model to be built for a different imaging setting. Some studies have showed large uncertainty [12; 24] in computing these parameters. Fourthly, as this model-based equations become more complex, the accuracy of fitting data to equations requires higher temporal resolution than the capability of current MRI technology. Therefore, we choose not to apply PKM to compute kinetic features Three-time points (3TP) model In the three-time points (3TP) model [20; 111], only three MR volumes acquired at three selected times are used to characterize the lesion heterogeneity of microvascular permeability and extracellular fraction. The 3TP model is used to compute washin rate and wash-out patterns. The wash-in rate is associate with the extracellular 99
122 Relative enhancement extravascular volume fraction v e whereas the wash-out patterns suggest microvascular permeability (K trans ). Early phase Later phase Persistent ±20% Plateau Washout t t t Time Figure 6.2: Color-coding scheme for creating the color hue and color intensity of the three-time point (3TP) model. The time between the pre-contrast image acquisition at t 0 and the second post-contrast image at t 2 is the early phase. The relative enhancement in the early phase is coded by color intensity from dark to bright. The enhancement patterns in the later phase are coded as green for persistent, blue for plateau, and red for washout patterns. The 3TP model allows convenient visualization of the kinetic features (wash-in and wash-out rates) using the color code as shown in Figure 6.2. Analysis of the wash-in and wash-out patterns are able to differentiate benign and malignant lesions [39]. The early phase (first four minutes after the injection of contrast agents) of signal intensity enhancement, namely the initial enhancement phase, is coded by color intensity. The enhancement patterns in the later phase (from four minutes to the end time of dynamic sequences) are coded by color hue (red for washout, blue for plateau and green for a persistent pattern). The limitation of the 3TP model is that it computes the kinetic 100
123 features using only 3 MR volumes where the information in the remaining volumes is missed. It also assumes that the all type of lesions reach their peak intensity at the same time points which is not always the case Time-intensity curve based kinetic features Due to the experimental limitations and inherent shortcomings of the PKM, model-free algorithms are generated to decompose enhancement patterns in order to segment and classify different breast tissues. Model-free algorithms (supervised and unsupervised) use various pattern recognition techniques to extract a prototypical curve to represent the time intensity curve of each tissue class (malignant, benign and normal). In this way, it has the potential to uncover intrinsic properties from images without any assumption of any physical models. Besides the kinetic features extracted using the two-compartment PKM (K trans and k ep ) and 3TP model (wash-in rate and washout patterns), there are various other kinetic features proposed to quantify the tissue kinetic behavior, including intensities, enhancements, peak enhancement (Equation (5.3)), time-to-peak, wash-in rate (Equation (5.4)), wash-out rate (Equation (5.5)) and integral (Equation (5.6)) as shown in Figure 5.2 in Chapter 5. In the 3TP model, the wash-in rate (WIR) is measured at a fixed time point. In the following section of this thesis, the WIRs are all computed by dividing the peak enhancement from the time-to-peak Extract representative region to generate curve It is suggested that kinetic features should be evaluated in the most suspicious area of a lesion [49; 50] that contains at least 3 voxels [48]. Therefore, the kinetic features of a lesion can be defined as a mean of the selected sub-region of the lesion. Newell et al. [77] automatically extracted 3 3 voxels in a lesion to compute the kinetic features. These voxels showed the strongest enhancement in the subtraction image at 1 minute after contrast injection. Schlossbauer et al. [94] employed unsupervised vector-quantization to partition a lesion into 4 regions. The region with the highest ranking was used to compute the kinetic features of the lesion. The FCM clustering method was used to partition time intensity curves in a lesion region into a number of clusters [13; 16]. A curve with the highest initial enhancement was chosen as the rep- 101
124 resentative curve. Figure 6.3 and 6.4 show the representative curves for malignant and benign lesions. Lucht et al. [68] applied an artificial neural networks (ANN) method to compute different TICs for carcinoma, benign lesion and parenchymal tissues. This method was further extended to combine an unsupervised vector quantization algorithm to include the spatial information [106]. Mean shift clustering [101; 102] subdivided each lesion into clusters based on pharmacokinetic parameters. The number of clusters was not fixed beforehand and depended on the heterogeneity of the feature space. In addition, the clustering result produced spatially spatially coherent clusters enhancement time Figure 6.3: TIC of 9 clustered in a malignant lesion computed by FCM. The TIC in red with the highest initial enhancement is selected as the representative curve for the lesion. 102
125 3.5 3 enhancement time Figure 6.4: TIC of 9 clusters in a benign lesion computed by FCM method. The TIC in red with the highest initial enhancement is selected as the representative curve for the lesion Morphological features In early lesion analysis and diagnosis, the focus was mainly on kinetic analysis. There is increasingly more attention being paid to both shape and margin analysis [1; 2; 63; 73; 77; 78]. Kuhl et al. [50] showed that increasing the spatial resolution can significantly improve lesion diagnosis accuracy, even at the cost of reducing temporal resolution. Shape analysis in the domain of medical image processing is commonly used in feature extraction of regions of interest for the purpose of lesion classification [14] or retrieval [47] or in the evaluation of the status or development of the regions of interest [11]. In the characterization of morphological features in breast lesions in DCE-MR im- 103
TUMOR DETECTION IN MRI IMAGES
 TUMOR DETECTION IN MRI IMAGES Prof. Pravin P. Adivarekar, 2 Priyanka P. Khatate, 3 Punam N. Pawar Prof. Pravin P. Adivarekar, 2 Priyanka P. Khatate, 3 Punam N. Pawar Asst. Professor, 2,3 BE Student,,2,3
TUMOR DETECTION IN MRI IMAGES Prof. Pravin P. Adivarekar, 2 Priyanka P. Khatate, 3 Punam N. Pawar Prof. Pravin P. Adivarekar, 2 Priyanka P. Khatate, 3 Punam N. Pawar Asst. Professor, 2,3 BE Student,,2,3
CHAPTER 6 DETECTION OF MASS USING NOVEL SEGMENTATION, GLCM AND NEURAL NETWORKS
 130 CHAPTER 6 DETECTION OF MASS USING NOVEL SEGMENTATION, GLCM AND NEURAL NETWORKS A mass is defined as a space-occupying lesion seen in more than one projection and it is described by its shapes and margin
130 CHAPTER 6 DETECTION OF MASS USING NOVEL SEGMENTATION, GLCM AND NEURAL NETWORKS A mass is defined as a space-occupying lesion seen in more than one projection and it is described by its shapes and margin
Automated segmentation methods for liver analysis in oncology applications
 University of Szeged Department of Image Processing and Computer Graphics Automated segmentation methods for liver analysis in oncology applications Ph. D. Thesis László Ruskó Thesis Advisor Dr. Antal
University of Szeged Department of Image Processing and Computer Graphics Automated segmentation methods for liver analysis in oncology applications Ph. D. Thesis László Ruskó Thesis Advisor Dr. Antal
Chapter 7 UNSUPERVISED LEARNING TECHNIQUES FOR MAMMOGRAM CLASSIFICATION
 UNSUPERVISED LEARNING TECHNIQUES FOR MAMMOGRAM CLASSIFICATION Supervised and unsupervised learning are the two prominent machine learning algorithms used in pattern recognition and classification. In this
UNSUPERVISED LEARNING TECHNIQUES FOR MAMMOGRAM CLASSIFICATION Supervised and unsupervised learning are the two prominent machine learning algorithms used in pattern recognition and classification. In this
Improving Positron Emission Tomography Imaging with Machine Learning David Fan-Chung Hsu CS 229 Fall
 Improving Positron Emission Tomography Imaging with Machine Learning David Fan-Chung Hsu (fcdh@stanford.edu), CS 229 Fall 2014-15 1. Introduction and Motivation High- resolution Positron Emission Tomography
Improving Positron Emission Tomography Imaging with Machine Learning David Fan-Chung Hsu (fcdh@stanford.edu), CS 229 Fall 2014-15 1. Introduction and Motivation High- resolution Positron Emission Tomography
MEDICAL IMAGE ANALYSIS
 SECOND EDITION MEDICAL IMAGE ANALYSIS ATAM P. DHAWAN g, A B IEEE Engineering in Medicine and Biology Society, Sponsor IEEE Press Series in Biomedical Engineering Metin Akay, Series Editor +IEEE IEEE PRESS
SECOND EDITION MEDICAL IMAGE ANALYSIS ATAM P. DHAWAN g, A B IEEE Engineering in Medicine and Biology Society, Sponsor IEEE Press Series in Biomedical Engineering Metin Akay, Series Editor +IEEE IEEE PRESS
New Technology Allows Multiple Image Contrasts in a Single Scan
 These images were acquired with an investigational device. PD T2 T2 FLAIR T1 MAP T1 FLAIR PSIR T1 New Technology Allows Multiple Image Contrasts in a Single Scan MR exams can be time consuming. A typical
These images were acquired with an investigational device. PD T2 T2 FLAIR T1 MAP T1 FLAIR PSIR T1 New Technology Allows Multiple Image Contrasts in a Single Scan MR exams can be time consuming. A typical
Slide 1. Technical Aspects of Quality Control in Magnetic Resonance Imaging. Slide 2. Annual Compliance Testing. of MRI Systems.
 Slide 1 Technical Aspects of Quality Control in Magnetic Resonance Imaging Slide 2 Compliance Testing of MRI Systems, Ph.D. Department of Radiology Henry Ford Hospital, Detroit, MI Slide 3 Compliance Testing
Slide 1 Technical Aspects of Quality Control in Magnetic Resonance Imaging Slide 2 Compliance Testing of MRI Systems, Ph.D. Department of Radiology Henry Ford Hospital, Detroit, MI Slide 3 Compliance Testing
Refraction Corrected Transmission Ultrasound Computed Tomography for Application in Breast Imaging
 Refraction Corrected Transmission Ultrasound Computed Tomography for Application in Breast Imaging Joint Research With Trond Varslot Marcel Jackowski Shengying Li and Klaus Mueller Ultrasound Detection
Refraction Corrected Transmission Ultrasound Computed Tomography for Application in Breast Imaging Joint Research With Trond Varslot Marcel Jackowski Shengying Li and Klaus Mueller Ultrasound Detection
8/3/2017. Contour Assessment for Quality Assurance and Data Mining. Objective. Outline. Tom Purdie, PhD, MCCPM
 Contour Assessment for Quality Assurance and Data Mining Tom Purdie, PhD, MCCPM Objective Understand the state-of-the-art in contour assessment for quality assurance including data mining-based techniques
Contour Assessment for Quality Assurance and Data Mining Tom Purdie, PhD, MCCPM Objective Understand the state-of-the-art in contour assessment for quality assurance including data mining-based techniques
Automated Lesion Detection Methods for 2D and 3D Chest X-Ray Images
 Automated Lesion Detection Methods for 2D and 3D Chest X-Ray Images Takeshi Hara, Hiroshi Fujita,Yongbum Lee, Hitoshi Yoshimura* and Shoji Kido** Department of Information Science, Gifu University Yanagido
Automated Lesion Detection Methods for 2D and 3D Chest X-Ray Images Takeshi Hara, Hiroshi Fujita,Yongbum Lee, Hitoshi Yoshimura* and Shoji Kido** Department of Information Science, Gifu University Yanagido
Breast MRI Accreditation Program Clinical Image Quality Guide
 Breast MRI Accreditation Program Clinical Image Quality Guide Introduction This document provides guidance on breast MRI clinical image quality and describes the criteria used by the ACR Breast MRI Accreditation
Breast MRI Accreditation Program Clinical Image Quality Guide Introduction This document provides guidance on breast MRI clinical image quality and describes the criteria used by the ACR Breast MRI Accreditation
doi: /
 Yiting Xie ; Anthony P. Reeves; Single 3D cell segmentation from optical CT microscope images. Proc. SPIE 934, Medical Imaging 214: Image Processing, 9343B (March 21, 214); doi:1.1117/12.243852. (214)
Yiting Xie ; Anthony P. Reeves; Single 3D cell segmentation from optical CT microscope images. Proc. SPIE 934, Medical Imaging 214: Image Processing, 9343B (March 21, 214); doi:1.1117/12.243852. (214)
CHAPTER-1 INTRODUCTION
 CHAPTER-1 INTRODUCTION 1.1 Fuzzy concept, digital image processing and application in medicine With the advancement of digital computers, it has become easy to store large amount of data and carry out
CHAPTER-1 INTRODUCTION 1.1 Fuzzy concept, digital image processing and application in medicine With the advancement of digital computers, it has become easy to store large amount of data and carry out
Global Journal of Engineering Science and Research Management
 ADVANCED K-MEANS ALGORITHM FOR BRAIN TUMOR DETECTION USING NAIVE BAYES CLASSIFIER Veena Bai K*, Dr. Niharika Kumar * MTech CSE, Department of Computer Science and Engineering, B.N.M. Institute of Technology,
ADVANCED K-MEANS ALGORITHM FOR BRAIN TUMOR DETECTION USING NAIVE BAYES CLASSIFIER Veena Bai K*, Dr. Niharika Kumar * MTech CSE, Department of Computer Science and Engineering, B.N.M. Institute of Technology,
Mass Classification Method in Mammogram Using Fuzzy K-Nearest Neighbour Equality
 Mass Classification Method in Mammogram Using Fuzzy K-Nearest Neighbour Equality Abstract: Mass classification of objects is an important area of research and application in a variety of fields. In this
Mass Classification Method in Mammogram Using Fuzzy K-Nearest Neighbour Equality Abstract: Mass classification of objects is an important area of research and application in a variety of fields. In this
Clinical Importance. Aortic Stenosis. Aortic Regurgitation. Ultrasound vs. MRI. Carotid Artery Stenosis
 Clinical Importance Rapid cardiovascular flow quantitation using sliceselective Fourier velocity encoding with spiral readouts Valve disease affects 10% of patients with heart disease in the U.S. Most
Clinical Importance Rapid cardiovascular flow quantitation using sliceselective Fourier velocity encoding with spiral readouts Valve disease affects 10% of patients with heart disease in the U.S. Most
Magnetic Resonance Elastography (MRE) of Liver Disease
 Magnetic Resonance Elastography (MRE) of Liver Disease Authored by: Jennifer Dolan Fox, PhD VirtualScopics Inc. jennifer_fox@virtualscopics.com 1-585-249-6231 1. Overview of MRE Imaging MRE is a magnetic
Magnetic Resonance Elastography (MRE) of Liver Disease Authored by: Jennifer Dolan Fox, PhD VirtualScopics Inc. jennifer_fox@virtualscopics.com 1-585-249-6231 1. Overview of MRE Imaging MRE is a magnetic
SVM-based CBIR of Breast Masses on Mammograms
 SVM-based CBIR of Breast Masses on Mammograms Lazaros Tsochatzidis, Konstantinos Zagoris, Michalis Savelonas, and Ioannis Pratikakis Visual Computing Group, Dept. of Electrical and Computer Engineering,
SVM-based CBIR of Breast Masses on Mammograms Lazaros Tsochatzidis, Konstantinos Zagoris, Michalis Savelonas, and Ioannis Pratikakis Visual Computing Group, Dept. of Electrical and Computer Engineering,
Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging
 1 CS 9 Final Project Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging Feiyu Chen Department of Electrical Engineering ABSTRACT Subject motion is a significant
1 CS 9 Final Project Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging Feiyu Chen Department of Electrical Engineering ABSTRACT Subject motion is a significant
Computer-aided detection of clustered microcalcifications in digital mammograms.
 Computer-aided detection of clustered microcalcifications in digital mammograms. Stelios Halkiotis, John Mantas & Taxiarchis Botsis. University of Athens Faculty of Nursing- Health Informatics Laboratory.
Computer-aided detection of clustered microcalcifications in digital mammograms. Stelios Halkiotis, John Mantas & Taxiarchis Botsis. University of Athens Faculty of Nursing- Health Informatics Laboratory.
The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy
 The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy Sokratis K. Makrogiannis, PhD From post-doctoral research at SBIA lab, Department of Radiology,
The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy Sokratis K. Makrogiannis, PhD From post-doctoral research at SBIA lab, Department of Radiology,
RADIOMICS: potential role in the clinics and challenges
 27 giugno 2018 Dipartimento di Fisica Università degli Studi di Milano RADIOMICS: potential role in the clinics and challenges Dr. Francesca Botta Medical Physicist Istituto Europeo di Oncologia (Milano)
27 giugno 2018 Dipartimento di Fisica Università degli Studi di Milano RADIOMICS: potential role in the clinics and challenges Dr. Francesca Botta Medical Physicist Istituto Europeo di Oncologia (Milano)
Computer-Aided Detection system for Hemorrhage contained region
 Computer-Aided Detection system for Hemorrhage contained region Myat Mon Kyaw Faculty of Information and Communication Technology University of Technology (Yatanarpon Cybercity), Pyin Oo Lwin, Myanmar
Computer-Aided Detection system for Hemorrhage contained region Myat Mon Kyaw Faculty of Information and Communication Technology University of Technology (Yatanarpon Cybercity), Pyin Oo Lwin, Myanmar
Enhanced material contrast by dual-energy microct imaging
 Enhanced material contrast by dual-energy microct imaging Method note Page 1 of 12 2 Method note: Dual-energy microct analysis 1. Introduction 1.1. The basis for dual energy imaging Micro-computed tomography
Enhanced material contrast by dual-energy microct imaging Method note Page 1 of 12 2 Method note: Dual-energy microct analysis 1. Introduction 1.1. The basis for dual energy imaging Micro-computed tomography
MRI Physics II: Gradients, Imaging
 MRI Physics II: Gradients, Imaging Douglas C., Ph.D. Dept. of Biomedical Engineering University of Michigan, Ann Arbor Magnetic Fields in MRI B 0 The main magnetic field. Always on (0.5-7 T) Magnetizes
MRI Physics II: Gradients, Imaging Douglas C., Ph.D. Dept. of Biomedical Engineering University of Michigan, Ann Arbor Magnetic Fields in MRI B 0 The main magnetic field. Always on (0.5-7 T) Magnetizes
Exploration and Study of MultiVolume Image Data using 3D Slicer
 Exploration and Study of MultiVolume Image Data using 3D Slicer Meysam Torabi and Andriy Fedorov torabi@bwh.harvard.edu, fedorov@bwh.harvard.edu Surgical Navigation and Robotics Lab and Surgical Planning
Exploration and Study of MultiVolume Image Data using 3D Slicer Meysam Torabi and Andriy Fedorov torabi@bwh.harvard.edu, fedorov@bwh.harvard.edu Surgical Navigation and Robotics Lab and Surgical Planning
Exam 8N080 - Introduction MRI
 Exam 8N080 - Introduction MRI Friday January 23 rd 2015, 13.30-16.30h For this exam you may use an ordinary calculator (not a graphical one). In total there are 6 assignments and a total of 65 points can
Exam 8N080 - Introduction MRI Friday January 23 rd 2015, 13.30-16.30h For this exam you may use an ordinary calculator (not a graphical one). In total there are 6 assignments and a total of 65 points can
Abbie M. Diak, PhD Loyola University Medical Center Dept. of Radiation Oncology
 Abbie M. Diak, PhD Loyola University Medical Center Dept. of Radiation Oncology Outline High Spectral and Spatial Resolution MR Imaging (HiSS) What it is How to do it Ways to use it HiSS for Radiation
Abbie M. Diak, PhD Loyola University Medical Center Dept. of Radiation Oncology Outline High Spectral and Spatial Resolution MR Imaging (HiSS) What it is How to do it Ways to use it HiSS for Radiation
Deviceless respiratory motion correction in PET imaging exploring the potential of novel data driven strategies
 g Deviceless respiratory motion correction in PET imaging exploring the potential of novel data driven strategies Presented by Adam Kesner, Ph.D., DABR Assistant Professor, Division of Radiological Sciences,
g Deviceless respiratory motion correction in PET imaging exploring the potential of novel data driven strategies Presented by Adam Kesner, Ph.D., DABR Assistant Professor, Division of Radiological Sciences,
ISSN: (Online) Volume 3, Issue 6, June 2015 International Journal of Advance Research in Computer Science and Management Studies
 ISSN: 2321-7782 (Online) Volume 3, Issue 6, June 2015 International Journal of Advance Research in Computer Science and Management Studies Research Article / Survey Paper / Case Study Available online
ISSN: 2321-7782 (Online) Volume 3, Issue 6, June 2015 International Journal of Advance Research in Computer Science and Management Studies Research Article / Survey Paper / Case Study Available online
Limitations of Projection Radiography. Stereoscopic Breast Imaging. Limitations of Projection Radiography. 3-D Breast Imaging Methods
 Stereoscopic Breast Imaging Andrew D. A. Maidment, Ph.D. Chief, Physics Section Department of Radiology University of Pennsylvania Limitations of Projection Radiography Mammography is a projection imaging
Stereoscopic Breast Imaging Andrew D. A. Maidment, Ph.D. Chief, Physics Section Department of Radiology University of Pennsylvania Limitations of Projection Radiography Mammography is a projection imaging
Image Processing Techniques for Brain Tumor Extraction from MRI Images using SVM Classifier
 Image Processing Techniques for Brain Tumor Extraction from MRI Images using SVM Classifier Mr. Ajaj Khan M. Tech (CSE) Scholar Central India Institute of Technology Indore, India ajajkhan72@gmail.com
Image Processing Techniques for Brain Tumor Extraction from MRI Images using SVM Classifier Mr. Ajaj Khan M. Tech (CSE) Scholar Central India Institute of Technology Indore, India ajajkhan72@gmail.com
Optimizing Flip Angle Selection in Breast MRI for Accurate Extraction and Visualization of T1 Tissue Relaxation Time
 Optimizing Flip Angle Selection in Breast MRI for Accurate Extraction and Visualization of T1 Tissue Relaxation Time GEORGIOS KETSETZIS AND MICHAEL BRADY Medical Vision Laboratory Department of Engineering
Optimizing Flip Angle Selection in Breast MRI for Accurate Extraction and Visualization of T1 Tissue Relaxation Time GEORGIOS KETSETZIS AND MICHAEL BRADY Medical Vision Laboratory Department of Engineering
Tumor Detection and classification of Medical MRI UsingAdvance ROIPropANN Algorithm
 International Journal of Engineering Research and Advanced Technology (IJERAT) DOI:http://dx.doi.org/10.31695/IJERAT.2018.3273 E-ISSN : 2454-6135 Volume.4, Issue 6 June -2018 Tumor Detection and classification
International Journal of Engineering Research and Advanced Technology (IJERAT) DOI:http://dx.doi.org/10.31695/IJERAT.2018.3273 E-ISSN : 2454-6135 Volume.4, Issue 6 June -2018 Tumor Detection and classification
MR IMAGE SEGMENTATION
 MR IMAGE SEGMENTATION Prepared by : Monil Shah What is Segmentation? Partitioning a region or regions of interest in images such that each region corresponds to one or more anatomic structures Classification
MR IMAGE SEGMENTATION Prepared by : Monil Shah What is Segmentation? Partitioning a region or regions of interest in images such that each region corresponds to one or more anatomic structures Classification
Lesion Segmentation and Bias Correction in Breast Ultrasound B-mode Images Including Elastography Information
 Lesion Segmentation and Bias Correction in Breast Ultrasound B-mode Images Including Elastography Information Gerard Pons a, Joan Martí a, Robert Martí a, Mariano Cabezas a, Andrew di Battista b, and J.
Lesion Segmentation and Bias Correction in Breast Ultrasound B-mode Images Including Elastography Information Gerard Pons a, Joan Martí a, Robert Martí a, Mariano Cabezas a, Andrew di Battista b, and J.
ANALYSIS OF PULMONARY FIBROSIS IN MRI, USING AN ELASTIC REGISTRATION TECHNIQUE IN A MODEL OF FIBROSIS: Scleroderma
 ANALYSIS OF PULMONARY FIBROSIS IN MRI, USING AN ELASTIC REGISTRATION TECHNIQUE IN A MODEL OF FIBROSIS: Scleroderma ORAL DEFENSE 8 th of September 2017 Charlotte MARTIN Supervisor: Pr. MP REVEL M2 Bio Medical
ANALYSIS OF PULMONARY FIBROSIS IN MRI, USING AN ELASTIC REGISTRATION TECHNIQUE IN A MODEL OF FIBROSIS: Scleroderma ORAL DEFENSE 8 th of September 2017 Charlotte MARTIN Supervisor: Pr. MP REVEL M2 Bio Medical
Segmentation and Classification of Breast Tumor Using Dynamic Contrast-Enhanced MR Images
 Segmentation and Classification of Breast Tumor Using Dynamic Contrast-Enhanced MR Images Yuanjie Zheng, Sajjad Baloch, Sarah Englander, Mitchell D. Schnall, and Dinggang Shen Department of Radiology,
Segmentation and Classification of Breast Tumor Using Dynamic Contrast-Enhanced MR Images Yuanjie Zheng, Sajjad Baloch, Sarah Englander, Mitchell D. Schnall, and Dinggang Shen Department of Radiology,
CP Generalize Concepts in Abstract Multi-dimensional Image Model Component Semantics. David Clunie.
 CP-1390 - Generalize Concepts in Abstract Multi-dimensional Image Model Semantics Page 1 STATUS Date of Last Update Person Assigned Submitter Name Submission Date Assigned 2014/06/09 David Clunie mailto:dclunie@dclunie.com
CP-1390 - Generalize Concepts in Abstract Multi-dimensional Image Model Semantics Page 1 STATUS Date of Last Update Person Assigned Submitter Name Submission Date Assigned 2014/06/09 David Clunie mailto:dclunie@dclunie.com
Automatic Determination of Arterial Input Function for Dynamic Contrast Enhanced MRI in Tumor Assessment
 Automatic Determination of Arterial Input Function for Dynamic Contrast Enhanced MRI in Tumor Assessment Jeremy Chen, Jianhua Yao, and David Thomasson Diagnostic Radiology Department, Clinical Center,
Automatic Determination of Arterial Input Function for Dynamic Contrast Enhanced MRI in Tumor Assessment Jeremy Chen, Jianhua Yao, and David Thomasson Diagnostic Radiology Department, Clinical Center,
White Pixel Artifact. Caused by a noise spike during acquisition Spike in K-space <--> sinusoid in image space
 White Pixel Artifact Caused by a noise spike during acquisition Spike in K-space sinusoid in image space Susceptibility Artifacts Off-resonance artifacts caused by adjacent regions with different
White Pixel Artifact Caused by a noise spike during acquisition Spike in K-space sinusoid in image space Susceptibility Artifacts Off-resonance artifacts caused by adjacent regions with different
Content Based Medical Image Retrieval Using Fuzzy C- Means Clustering With RF
 Content Based Medical Image Retrieval Using Fuzzy C- Means Clustering With RF Jasmine Samraj #1, NazreenBee. M *2 # Associate Professor, Department of Computer Science, Quaid-E-Millath Government college
Content Based Medical Image Retrieval Using Fuzzy C- Means Clustering With RF Jasmine Samraj #1, NazreenBee. M *2 # Associate Professor, Department of Computer Science, Quaid-E-Millath Government college
Development of computer-aided diagnostic system for breast MRI lesion classification
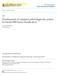 University of Iowa Iowa Research Online Theses and Dissertations 2005 Development of computer-aided diagnostic system for breast MRI lesion classification Lina Arbash Meinel University of Iowa Copyright
University of Iowa Iowa Research Online Theses and Dissertations 2005 Development of computer-aided diagnostic system for breast MRI lesion classification Lina Arbash Meinel University of Iowa Copyright
Introduction to Medical Image Processing
 Introduction to Medical Image Processing Δ Essential environments of a medical imaging system Subject Image Analysis Energy Imaging System Images Image Processing Feature Images Image processing may be
Introduction to Medical Image Processing Δ Essential environments of a medical imaging system Subject Image Analysis Energy Imaging System Images Image Processing Feature Images Image processing may be
CHAPTER 3 RETINAL OPTIC DISC SEGMENTATION
 60 CHAPTER 3 RETINAL OPTIC DISC SEGMENTATION 3.1 IMPORTANCE OF OPTIC DISC Ocular fundus images provide information about ophthalmic, retinal and even systemic diseases such as hypertension, diabetes, macular
60 CHAPTER 3 RETINAL OPTIC DISC SEGMENTATION 3.1 IMPORTANCE OF OPTIC DISC Ocular fundus images provide information about ophthalmic, retinal and even systemic diseases such as hypertension, diabetes, macular
Image Acquisition Systems
 Image Acquisition Systems Goals and Terminology Conventional Radiography Axial Tomography Computer Axial Tomography (CAT) Magnetic Resonance Imaging (MRI) PET, SPECT Ultrasound Microscopy Imaging ITCS
Image Acquisition Systems Goals and Terminology Conventional Radiography Axial Tomography Computer Axial Tomography (CAT) Magnetic Resonance Imaging (MRI) PET, SPECT Ultrasound Microscopy Imaging ITCS
Chapter 3 Set Redundancy in Magnetic Resonance Brain Images
 16 Chapter 3 Set Redundancy in Magnetic Resonance Brain Images 3.1 MRI (magnetic resonance imaging) MRI is a technique of measuring physical structure within the human anatomy. Our proposed research focuses
16 Chapter 3 Set Redundancy in Magnetic Resonance Brain Images 3.1 MRI (magnetic resonance imaging) MRI is a technique of measuring physical structure within the human anatomy. Our proposed research focuses
Available Online through
 Available Online through www.ijptonline.com ISSN: 0975-766X CODEN: IJPTFI Research Article ANALYSIS OF CT LIVER IMAGES FOR TUMOUR DIAGNOSIS BASED ON CLUSTERING TECHNIQUE AND TEXTURE FEATURES M.Krithika
Available Online through www.ijptonline.com ISSN: 0975-766X CODEN: IJPTFI Research Article ANALYSIS OF CT LIVER IMAGES FOR TUMOUR DIAGNOSIS BASED ON CLUSTERING TECHNIQUE AND TEXTURE FEATURES M.Krithika
Segmenting Lesions in Multiple Sclerosis Patients James Chen, Jason Su
 Segmenting Lesions in Multiple Sclerosis Patients James Chen, Jason Su Radiologists and researchers spend countless hours tediously segmenting white matter lesions to diagnose and study brain diseases.
Segmenting Lesions in Multiple Sclerosis Patients James Chen, Jason Su Radiologists and researchers spend countless hours tediously segmenting white matter lesions to diagnose and study brain diseases.
A ranklet-based CAD for digital mammography
 A ranklet-based CAD for digital mammography Enrico Angelini 1, Renato Campanini 1, Emiro Iampieri 1, Nico Lanconelli 1, Matteo Masotti 1, Todor Petkov 1, and Matteo Roffilli 2 1 Physics Department, University
A ranklet-based CAD for digital mammography Enrico Angelini 1, Renato Campanini 1, Emiro Iampieri 1, Nico Lanconelli 1, Matteo Masotti 1, Todor Petkov 1, and Matteo Roffilli 2 1 Physics Department, University
MEDICAL IMAGE NOISE REDUCTION AND REGION CONTRAST ENHANCEMENT USING PARTIAL DIFFERENTIAL EQUATIONS
 MEDICAL IMAGE NOISE REDUCTION AND REGION CONTRAST ENHANCEMENT USING PARTIAL DIFFERENTIAL EQUATIONS Miguel Alemán-Flores, Luis Álvarez-León Departamento de Informática y Sistemas, Universidad de Las Palmas
MEDICAL IMAGE NOISE REDUCTION AND REGION CONTRAST ENHANCEMENT USING PARTIAL DIFFERENTIAL EQUATIONS Miguel Alemán-Flores, Luis Álvarez-León Departamento de Informática y Sistemas, Universidad de Las Palmas
Nonrigid Motion Compensation of Free Breathing Acquired Myocardial Perfusion Data
 Nonrigid Motion Compensation of Free Breathing Acquired Myocardial Perfusion Data Gert Wollny 1, Peter Kellman 2, Andrés Santos 1,3, María-Jesus Ledesma 1,3 1 Biomedical Imaging Technologies, Department
Nonrigid Motion Compensation of Free Breathing Acquired Myocardial Perfusion Data Gert Wollny 1, Peter Kellman 2, Andrés Santos 1,3, María-Jesus Ledesma 1,3 1 Biomedical Imaging Technologies, Department
Computational Medical Imaging Analysis
 Computational Medical Imaging Analysis Chapter 2: Image Acquisition Systems Jun Zhang Laboratory for Computational Medical Imaging & Data Analysis Department of Computer Science University of Kentucky
Computational Medical Imaging Analysis Chapter 2: Image Acquisition Systems Jun Zhang Laboratory for Computational Medical Imaging & Data Analysis Department of Computer Science University of Kentucky
Brilliance CT Big Bore.
 1 2 2 There are two methods of RCCT acquisition in widespread clinical use: cine axial and helical. In RCCT with cine axial acquisition, repeat CT images are taken each couch position while recording respiration.
1 2 2 There are two methods of RCCT acquisition in widespread clinical use: cine axial and helical. In RCCT with cine axial acquisition, repeat CT images are taken each couch position while recording respiration.
Ch. 4 Physical Principles of CT
 Ch. 4 Physical Principles of CT CLRS 408: Intro to CT Department of Radiation Sciences Review: Why CT? Solution for radiography/tomography limitations Superimposition of structures Distinguishing between
Ch. 4 Physical Principles of CT CLRS 408: Intro to CT Department of Radiation Sciences Review: Why CT? Solution for radiography/tomography limitations Superimposition of structures Distinguishing between
Diffusion MRI Acquisition. Karla Miller FMRIB Centre, University of Oxford
 Diffusion MRI Acquisition Karla Miller FMRIB Centre, University of Oxford karla@fmrib.ox.ac.uk Diffusion Imaging How is diffusion weighting achieved? How is the image acquired? What are the limitations,
Diffusion MRI Acquisition Karla Miller FMRIB Centre, University of Oxford karla@fmrib.ox.ac.uk Diffusion Imaging How is diffusion weighting achieved? How is the image acquired? What are the limitations,
Lab Location: MRI, B2, Cardinal Carter Wing, St. Michael s Hospital, 30 Bond Street
 Lab Location: MRI, B2, Cardinal Carter Wing, St. Michael s Hospital, 30 Bond Street MRI is located in the sub basement of CC wing. From Queen or Victoria, follow the baby blue arrows and ride the CC south
Lab Location: MRI, B2, Cardinal Carter Wing, St. Michael s Hospital, 30 Bond Street MRI is located in the sub basement of CC wing. From Queen or Victoria, follow the baby blue arrows and ride the CC south
Can we Distinguish Between Benign and Malignant Breast Tumors in DCE-MRI by Studying a Tumor s Most Suspect Region Only?
 Can we Distinguish Between Benign and Malignant Breast Tumors in DCE-MRI by Studying a Tumor s Most Suspect Region Only? Sylvia Glaßer, Uli Niemann, Bernhard Preim Department for Simulation and Graphics
Can we Distinguish Between Benign and Malignant Breast Tumors in DCE-MRI by Studying a Tumor s Most Suspect Region Only? Sylvia Glaßer, Uli Niemann, Bernhard Preim Department for Simulation and Graphics
Computer-Aided Diagnosis in Abdominal and Cardiac Radiology Using Neural Networks
 Computer-Aided Diagnosis in Abdominal and Cardiac Radiology Using Neural Networks Du-Yih Tsai, Masaru Sekiya and Yongbum Lee Department of Radiological Technology, School of Health Sciences, Faculty of
Computer-Aided Diagnosis in Abdominal and Cardiac Radiology Using Neural Networks Du-Yih Tsai, Masaru Sekiya and Yongbum Lee Department of Radiological Technology, School of Health Sciences, Faculty of
ISSN: X Impact factor: 4.295
 ISSN: 2454-132X Impact factor: 4.295 (Volume3, Issue1) Available online at: www.ijariit.com Performance Analysis of Image Clustering Algorithm Applied to Brain MRI Kalyani R.Mandlik 1, Dr. Suresh S. Salankar
ISSN: 2454-132X Impact factor: 4.295 (Volume3, Issue1) Available online at: www.ijariit.com Performance Analysis of Image Clustering Algorithm Applied to Brain MRI Kalyani R.Mandlik 1, Dr. Suresh S. Salankar
algorithms ISSN
 Algorithms 2009, 2, 828-849; doi:10.3390/a2020828 OPEN ACCESS algorithms ISSN 1999-4893 www.mdpi.com/journal/algorithms Review Computer-Aided Diagnosis in Mammography Using Content- Based Image Retrieval
Algorithms 2009, 2, 828-849; doi:10.3390/a2020828 OPEN ACCESS algorithms ISSN 1999-4893 www.mdpi.com/journal/algorithms Review Computer-Aided Diagnosis in Mammography Using Content- Based Image Retrieval
Role of Parallel Imaging in High Field Functional MRI
 Role of Parallel Imaging in High Field Functional MRI Douglas C. Noll & Bradley P. Sutton Department of Biomedical Engineering, University of Michigan Supported by NIH Grant DA15410 & The Whitaker Foundation
Role of Parallel Imaging in High Field Functional MRI Douglas C. Noll & Bradley P. Sutton Department of Biomedical Engineering, University of Michigan Supported by NIH Grant DA15410 & The Whitaker Foundation
COSC160: Detection and Classification. Jeremy Bolton, PhD Assistant Teaching Professor
 COSC160: Detection and Classification Jeremy Bolton, PhD Assistant Teaching Professor Outline I. Problem I. Strategies II. Features for training III. Using spatial information? IV. Reducing dimensionality
COSC160: Detection and Classification Jeremy Bolton, PhD Assistant Teaching Professor Outline I. Problem I. Strategies II. Features for training III. Using spatial information? IV. Reducing dimensionality
Colonic Polyp Characterization and Detection based on Both Morphological and Texture Features
 Colonic Polyp Characterization and Detection based on Both Morphological and Texture Features Zigang Wang 1, Lihong Li 1,4, Joseph Anderson 2, Donald Harrington 1 and Zhengrong Liang 1,3 Departments of
Colonic Polyp Characterization and Detection based on Both Morphological and Texture Features Zigang Wang 1, Lihong Li 1,4, Joseph Anderson 2, Donald Harrington 1 and Zhengrong Liang 1,3 Departments of
MRI Imaging Options. Frank R. Korosec, Ph.D. Departments of Radiology and Medical Physics University of Wisconsin Madison
 MRI Imaging Options Frank R. Korosec, Ph.D. Departments of Radiolog and Medical Phsics Universit of Wisconsin Madison f.korosec@hosp.wisc.edu As MR imaging becomes more developed, more imaging options
MRI Imaging Options Frank R. Korosec, Ph.D. Departments of Radiolog and Medical Phsics Universit of Wisconsin Madison f.korosec@hosp.wisc.edu As MR imaging becomes more developed, more imaging options
Detection & Classification of Lung Nodules Using multi resolution MTANN in Chest Radiography Images
 The International Journal Of Engineering And Science (IJES) ISSN (e): 2319 1813 ISSN (p): 2319 1805 Pages 98-104 March - 2015 Detection & Classification of Lung Nodules Using multi resolution MTANN in
The International Journal Of Engineering And Science (IJES) ISSN (e): 2319 1813 ISSN (p): 2319 1805 Pages 98-104 March - 2015 Detection & Classification of Lung Nodules Using multi resolution MTANN in
A fast breast nonlinear elastography reconstruction technique using the Veronda-Westman model
 A fast breast nonlinear elastography reconstruction technique using the Veronda-Westman model Mohammadhosein Amooshahi a and Abbas Samani abc a Department of Electrical & Computer Engineering, University
A fast breast nonlinear elastography reconstruction technique using the Veronda-Westman model Mohammadhosein Amooshahi a and Abbas Samani abc a Department of Electrical & Computer Engineering, University
Content based Image Retrievals for Brain Related Diseases
 Content based Image Retrievals for Brain Related Diseases T.V. Madhusudhana Rao Department of CSE, T.P.I.S.T., Bobbili, Andhra Pradesh, INDIA S. Pallam Setty Department of CS&SE, Andhra University, Visakhapatnam,
Content based Image Retrievals for Brain Related Diseases T.V. Madhusudhana Rao Department of CSE, T.P.I.S.T., Bobbili, Andhra Pradesh, INDIA S. Pallam Setty Department of CS&SE, Andhra University, Visakhapatnam,
Toward Automated Cancer Diagnosis: An Interactive System for Cell Feature Extraction
 Toward Automated Cancer Diagnosis: An Interactive System for Cell Feature Extraction Nick Street Computer Sciences Department University of Wisconsin-Madison street@cs.wisc.edu Abstract Oncologists at
Toward Automated Cancer Diagnosis: An Interactive System for Cell Feature Extraction Nick Street Computer Sciences Department University of Wisconsin-Madison street@cs.wisc.edu Abstract Oncologists at
COMPREHENSIVE QUALITY CONTROL OF NMR TOMOGRAPHY USING 3D PRINTED PHANTOM
 COMPREHENSIVE QUALITY CONTROL OF NMR TOMOGRAPHY USING 3D PRINTED PHANTOM Mažena MACIUSOVIČ *, Marius BURKANAS *, Jonas VENIUS *, ** * Medical Physics Department, National Cancer Institute, Vilnius, Lithuania
COMPREHENSIVE QUALITY CONTROL OF NMR TOMOGRAPHY USING 3D PRINTED PHANTOM Mažena MACIUSOVIČ *, Marius BURKANAS *, Jonas VENIUS *, ** * Medical Physics Department, National Cancer Institute, Vilnius, Lithuania
REAL-TIME ADAPTIVITY IN HEAD-AND-NECK AND LUNG CANCER RADIOTHERAPY IN A GPU ENVIRONMENT
 REAL-TIME ADAPTIVITY IN HEAD-AND-NECK AND LUNG CANCER RADIOTHERAPY IN A GPU ENVIRONMENT Anand P Santhanam Assistant Professor, Department of Radiation Oncology OUTLINE Adaptive radiotherapy for head and
REAL-TIME ADAPTIVITY IN HEAD-AND-NECK AND LUNG CANCER RADIOTHERAPY IN A GPU ENVIRONMENT Anand P Santhanam Assistant Professor, Department of Radiation Oncology OUTLINE Adaptive radiotherapy for head and
Computer aided analysis of inflammatory muscle disease using magnetic resonance imaging
 Loughborough University Institutional Repository Computer aided analysis of inflammatory muscle disease using magnetic resonance imaging This item was submitted to Loughborough University's Institutional
Loughborough University Institutional Repository Computer aided analysis of inflammatory muscle disease using magnetic resonance imaging This item was submitted to Loughborough University's Institutional
Intraoperative Prostate Tracking with Slice-to-Volume Registration in MR
 Intraoperative Prostate Tracking with Slice-to-Volume Registration in MR Sean Gill a, Purang Abolmaesumi a,b, Siddharth Vikal a, Parvin Mousavi a and Gabor Fichtinger a,b,* (a) School of Computing, Queen
Intraoperative Prostate Tracking with Slice-to-Volume Registration in MR Sean Gill a, Purang Abolmaesumi a,b, Siddharth Vikal a, Parvin Mousavi a and Gabor Fichtinger a,b,* (a) School of Computing, Queen
Development and Evaluation of Computerized Segmentation. Algorithm for 3D Multimodality Breast Images
 Development and Evaluation of Computerized Segmentation Algorithm for 3D Multimodality Breast Images BY HSIEN-CHI KUO B.S., National Taiwan University, 2006 THESIS Submitted as partial fulfillment of the
Development and Evaluation of Computerized Segmentation Algorithm for 3D Multimodality Breast Images BY HSIEN-CHI KUO B.S., National Taiwan University, 2006 THESIS Submitted as partial fulfillment of the
Parameter Optimization for Mammogram Image Classification with Support Vector Machine
 , March 15-17, 2017, Hong Kong Parameter Optimization for Mammogram Image Classification with Support Vector Machine Keerachart Suksut, Ratiporn Chanklan, Nuntawut Kaoungku, Kedkard Chaiyakhan, Nittaya
, March 15-17, 2017, Hong Kong Parameter Optimization for Mammogram Image Classification with Support Vector Machine Keerachart Suksut, Ratiporn Chanklan, Nuntawut Kaoungku, Kedkard Chaiyakhan, Nittaya
The Discovery and Retrieval of Temporal Rules in Interval Sequence Data
 The Discovery and Retrieval of Temporal Rules in Interval Sequence Data by Edi Winarko, B.Sc., M.Sc. School of Informatics and Engineering, Faculty of Science and Engineering March 19, 2007 A thesis presented
The Discovery and Retrieval of Temporal Rules in Interval Sequence Data by Edi Winarko, B.Sc., M.Sc. School of Informatics and Engineering, Faculty of Science and Engineering March 19, 2007 A thesis presented
CT NOISE POWER SPECTRUM FOR FILTERED BACKPROJECTION AND ITERATIVE RECONSTRUCTION
 CT NOISE POWER SPECTRUM FOR FILTERED BACKPROJECTION AND ITERATIVE RECONSTRUCTION Frank Dong, PhD, DABR Diagnostic Physicist, Imaging Institute Cleveland Clinic Foundation and Associate Professor of Radiology
CT NOISE POWER SPECTRUM FOR FILTERED BACKPROJECTION AND ITERATIVE RECONSTRUCTION Frank Dong, PhD, DABR Diagnostic Physicist, Imaging Institute Cleveland Clinic Foundation and Associate Professor of Radiology
Imaging-based Classification Algorithms on Clinical Trial Data with Injected Tumour Responses
 Imaging-based Classification Algorithms on Clinical Trial Data with Injected Tumour Responses Yunpeng Li, Emily Porter, Mark Coates Dept. of Electrical and Computer Engineering, McGill University, Montréal,
Imaging-based Classification Algorithms on Clinical Trial Data with Injected Tumour Responses Yunpeng Li, Emily Porter, Mark Coates Dept. of Electrical and Computer Engineering, McGill University, Montréal,
MR-Guided Mixed Reality for Breast Conserving Surgical Planning
 MR-Guided Mixed Reality for Breast Conserving Surgical Planning Suba Srinivasan (subashini7@gmail.com) March 30 th 2017 Mentors: Prof. Brian A. Hargreaves, Prof. Bruce L. Daniel MEDICINE MRI Guided Mixed
MR-Guided Mixed Reality for Breast Conserving Surgical Planning Suba Srinivasan (subashini7@gmail.com) March 30 th 2017 Mentors: Prof. Brian A. Hargreaves, Prof. Bruce L. Daniel MEDICINE MRI Guided Mixed
Medical Image Feature, Extraction, Selection And Classification
 Medical Image Feature, Extraction, Selection And Classification 1 M.VASANTHA *, 2 DR.V.SUBBIAH BHARATHI, 3 R.DHAMODHARAN 1. Research Scholar, Mother Teresa Women s University, KodaiKanal, and Asst.Professor,
Medical Image Feature, Extraction, Selection And Classification 1 M.VASANTHA *, 2 DR.V.SUBBIAH BHARATHI, 3 R.DHAMODHARAN 1. Research Scholar, Mother Teresa Women s University, KodaiKanal, and Asst.Professor,
Comparison of Vessel Segmentations Using STAPLE
 Comparison of Vessel Segmentations Using STAPLE Julien Jomier, Vincent LeDigarcher, and Stephen R. Aylward Computer-Aided Diagnosis and Display Lab, The University of North Carolina at Chapel Hill, Department
Comparison of Vessel Segmentations Using STAPLE Julien Jomier, Vincent LeDigarcher, and Stephen R. Aylward Computer-Aided Diagnosis and Display Lab, The University of North Carolina at Chapel Hill, Department
Imaging Guide Pancreatic Imaging
 Imaging Guide Guide to Small Animal Pancreatic Imaging using the Vevo 2100 Imaging System Ver 1.0 Guide to Small Animal Pancreatic Imaging using the Vevo 2100 Imaging System Course Objectives: This guide
Imaging Guide Guide to Small Animal Pancreatic Imaging using the Vevo 2100 Imaging System Ver 1.0 Guide to Small Animal Pancreatic Imaging using the Vevo 2100 Imaging System Course Objectives: This guide
MR Advance Techniques. Vascular Imaging. Class III
 MR Advance Techniques Vascular Imaging Class III 1 Vascular Imaging There are several methods that can be used to evaluate the cardiovascular systems with the use of MRI. MRI will aloud to evaluate morphology
MR Advance Techniques Vascular Imaging Class III 1 Vascular Imaging There are several methods that can be used to evaluate the cardiovascular systems with the use of MRI. MRI will aloud to evaluate morphology
Fast methods for magnetic resonance angiography (MRA)
 Fast methods for magnetic resonance angiography (MRA) Bahareh Vafadar Department of Electrical and Computer Engineering A thesis presented for the degree of Doctor of Philosophy University of Canterbury,
Fast methods for magnetic resonance angiography (MRA) Bahareh Vafadar Department of Electrical and Computer Engineering A thesis presented for the degree of Doctor of Philosophy University of Canterbury,
A Virtual MR Scanner for Education
 A Virtual MR Scanner for Education Hackländer T, Schalla C, Trümper A, Mertens H, Hiltner J, Cramer BM Hospitals of the University Witten/Herdecke, Department of Radiology Wuppertal, Germany Purpose A
A Virtual MR Scanner for Education Hackländer T, Schalla C, Trümper A, Mertens H, Hiltner J, Cramer BM Hospitals of the University Witten/Herdecke, Department of Radiology Wuppertal, Germany Purpose A
Extraction and recognition of the thoracic organs based on 3D CT images and its application
 1 Extraction and recognition of the thoracic organs based on 3D CT images and its application Xiangrong Zhou, PhD a, Takeshi Hara, PhD b, Hiroshi Fujita, PhD b, Yoshihiro Ida, RT c, Kazuhiro Katada, MD
1 Extraction and recognition of the thoracic organs based on 3D CT images and its application Xiangrong Zhou, PhD a, Takeshi Hara, PhD b, Hiroshi Fujita, PhD b, Yoshihiro Ida, RT c, Kazuhiro Katada, MD
Outline: Contrast-enhanced MRA
 Outline: Contrast-enhanced MRA Background Technique Clinical Indications Future Directions Disclosures: GE Health Care: Research support Consultant: Bracco, Bayer The Basics During rapid IV infusion, Gadolinium
Outline: Contrast-enhanced MRA Background Technique Clinical Indications Future Directions Disclosures: GE Health Care: Research support Consultant: Bracco, Bayer The Basics During rapid IV infusion, Gadolinium
(a Scrhon5 R2iwd b. P)jc%z 5. ivcr3. 1. I. ZOms Xn,s. 1E IDrAS boms. EE225E/BIOE265 Spring 2013 Principles of MRI. Assignment 8 Solutions
 EE225E/BIOE265 Spring 2013 Principles of MRI Miki Lustig Assignment 8 Solutions 1. Nishimura 7.1 P)jc%z 5 ivcr3. 1. I Due Wednesday April 10th, 2013 (a Scrhon5 R2iwd b 0 ZOms Xn,s r cx > qs 4-4 8ni6 4
EE225E/BIOE265 Spring 2013 Principles of MRI Miki Lustig Assignment 8 Solutions 1. Nishimura 7.1 P)jc%z 5 ivcr3. 1. I Due Wednesday April 10th, 2013 (a Scrhon5 R2iwd b 0 ZOms Xn,s r cx > qs 4-4 8ni6 4
4/19/2016. Deborah Thames R.T. (R)(M)(QM) Theory & Technology and advancement in 3D imaging DBT
 Deborah Thames R.T. (R)(M)(QM) Theory & Technology and advancement in 3D imaging DBT 1 Three manufacturers approved for Tomo Hologic and GE, and Siemens Why 2D Digital Mammography 2D FFDM it appears to
Deborah Thames R.T. (R)(M)(QM) Theory & Technology and advancement in 3D imaging DBT 1 Three manufacturers approved for Tomo Hologic and GE, and Siemens Why 2D Digital Mammography 2D FFDM it appears to
MIXTURE MODELING FOR DIGITAL MAMMOGRAM DISPLAY AND ANALYSIS 1
 MIXTURE MODELING FOR DIGITAL MAMMOGRAM DISPLAY AND ANALYSIS 1 Stephen R. Aylward, Bradley M. Hemminger, and Etta D. Pisano Department of Radiology Medical Image Display and Analysis Group University of
MIXTURE MODELING FOR DIGITAL MAMMOGRAM DISPLAY AND ANALYSIS 1 Stephen R. Aylward, Bradley M. Hemminger, and Etta D. Pisano Department of Radiology Medical Image Display and Analysis Group University of
UNIVERSITY OF SOUTHAMPTON
 UNIVERSITY OF SOUTHAMPTON PHYS2007W1 SEMESTER 2 EXAMINATION 2014-2015 MEDICAL PHYSICS Duration: 120 MINS (2 hours) This paper contains 10 questions. Answer all questions in Section A and only two questions
UNIVERSITY OF SOUTHAMPTON PHYS2007W1 SEMESTER 2 EXAMINATION 2014-2015 MEDICAL PHYSICS Duration: 120 MINS (2 hours) This paper contains 10 questions. Answer all questions in Section A and only two questions
Diffusion Mapping with FireVoxel Quick Start Guide
 Diffusion Mapping with FireVoxel Quick Start Guide Medical image analysis tool developed by Artem Mikheev and Henry Rusinek Radiology Department, NYU School of Medicine Original version prepared by Jinyu
Diffusion Mapping with FireVoxel Quick Start Guide Medical image analysis tool developed by Artem Mikheev and Henry Rusinek Radiology Department, NYU School of Medicine Original version prepared by Jinyu
Similarity Measurement of Breast Cancer Mammographic Images Using Combination of Mesh Distance Fourier Transform and Global Features
 South Dakota State University Open PRAIRIE: Open Public Research Access Institutional Repository and Information Exchange Theses and Dissertations 2016 Similarity Measurement of Breast Cancer Mammographic
South Dakota State University Open PRAIRIE: Open Public Research Access Institutional Repository and Information Exchange Theses and Dissertations 2016 Similarity Measurement of Breast Cancer Mammographic
Comparison of Vessel Segmentations using STAPLE
 Comparison of Vessel Segmentations using STAPLE Julien Jomier, Vincent LeDigarcher, and Stephen R. Aylward Computer-Aided Diagnosis and Display Lab The University of North Carolina at Chapel Hill, Department
Comparison of Vessel Segmentations using STAPLE Julien Jomier, Vincent LeDigarcher, and Stephen R. Aylward Computer-Aided Diagnosis and Display Lab The University of North Carolina at Chapel Hill, Department
Tomographic Reconstruction
 Tomographic Reconstruction 3D Image Processing Torsten Möller Reading Gonzales + Woods, Chapter 5.11 2 Overview Physics History Reconstruction basic idea Radon transform Fourier-Slice theorem (Parallel-beam)
Tomographic Reconstruction 3D Image Processing Torsten Möller Reading Gonzales + Woods, Chapter 5.11 2 Overview Physics History Reconstruction basic idea Radon transform Fourier-Slice theorem (Parallel-beam)
Enhao Gong, PhD Candidate, Electrical Engineering, Stanford University Dr. John Pauly, Professor in Electrical Engineering, Stanford University Dr.
 Enhao Gong, PhD Candidate, Electrical Engineering, Stanford University Dr. John Pauly, Professor in Electrical Engineering, Stanford University Dr. Greg Zaharchuk, Associate Professor in Radiology, Stanford
Enhao Gong, PhD Candidate, Electrical Engineering, Stanford University Dr. John Pauly, Professor in Electrical Engineering, Stanford University Dr. Greg Zaharchuk, Associate Professor in Radiology, Stanford
CHAPTER 3 WAVELET DECOMPOSITION USING HAAR WAVELET
 69 CHAPTER 3 WAVELET DECOMPOSITION USING HAAR WAVELET 3.1 WAVELET Wavelet as a subject is highly interdisciplinary and it draws in crucial ways on ideas from the outside world. The working of wavelet in
69 CHAPTER 3 WAVELET DECOMPOSITION USING HAAR WAVELET 3.1 WAVELET Wavelet as a subject is highly interdisciplinary and it draws in crucial ways on ideas from the outside world. The working of wavelet in
Module 5: Dynamic Imaging and Phase Sharing. (true-fisp, TRICKS, CAPR, DISTAL, DISCO, HYPR) Review. Improving Temporal Resolution.
 MRES 7005 - Fast Imaging Techniques Module 5: Dynamic Imaging and Phase Sharing (true-fisp, TRICKS, CAPR, DISTAL, DISCO, HYPR) Review Improving Temporal Resolution True-FISP (I) True-FISP (II) Keyhole
MRES 7005 - Fast Imaging Techniques Module 5: Dynamic Imaging and Phase Sharing (true-fisp, TRICKS, CAPR, DISTAL, DISCO, HYPR) Review Improving Temporal Resolution True-FISP (I) True-FISP (II) Keyhole
RIGID IMAGE REGISTRATION
 RIGID IMAGE REGISTRATION Duygu Tosun-Turgut, Ph.D. Center for Imaging of Neurodegenerative Diseases Department of Radiology and Biomedical Imaging duygu.tosun@ucsf.edu What is registration? Image registration
RIGID IMAGE REGISTRATION Duygu Tosun-Turgut, Ph.D. Center for Imaging of Neurodegenerative Diseases Department of Radiology and Biomedical Imaging duygu.tosun@ucsf.edu What is registration? Image registration
