Lectures by Volker Heun, Daniel Huson and Knut Reinert, in particular last years lectures
|
|
|
- Clarissa Black
- 6 years ago
- Views:
Transcription
1 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 4.1 Sources for this lecture Lectures by Volker Heun, Daniel Huson and Knut Reinert, in particular last years lectures Dan Gusfield: Algorithms on Strings, Trees, and Sequences. Computer Science and Computational Biology. Cambridge University Press, Cambridge, ISBN David B. Mount: Bioinformatics. Sequence and Genome Analysis. Cold Spring Harbor Laboratory Press, New York, ISBN File fasta3x.doc from the FastA distribution 4.2 Motivation A local alignment of two sequences x, y can be computed by the Smith-Waterman algorithm, which requires O( x y ) time and space in its original form. While the space requirement can be pushed down to O( x + y ) using a divide-and-conquer algorithm, the running time remains O( x y ), which is not feasible for many applications. Therefore, heuristic approaches are of great importance in practice. Heuristics A heuristic is an often problem-specific algorithm that will yield reasonable results, even if it is not provably optimal or even lacks a performance guarantee. In most cases, it is intuitively clear that the method works / is fast / is space efficient (in all / many / typical / special cases) but we do not (yet?) have a sound analysis for it. 4.3 FastA FastA is a program for heuristic pairwise local alignment of nucleic and amino acid sequences. FastA is much faster than the exact algorithms based on dynamic programming, and can be used to search a database of sequences for homologs to a query sequence. However its speed is based on heuristic considerations and it may fail to detect similar sequences in certain cases.
2 FastA and the chaining problem, by Clemens Gröpl, Gunnar Klau, November 22, 2010, 09: The name FastA is an abbreviation for fast all. (There was a version for nucleic acids before.) The official pronounciation is fast-ay. First version by Pearson and Lipman, 1988; recent version (3.4) August Main steps of the FastA algorithm 1. Find hot-spots. A hot-spot is a short, exact match between the two sequences. 2. Find diagonal runs. A diagonal run is a collection of hot-spots on the same diagonal within a short distance from each other. 3. Rescore the best diagonal runs. This is done using a substitution matrix. The best initial region is INIT1. 4. Chain several initial regions. This is where the chaining problem comes up. The result is INITN. 5. Moreover, compute an optimal local alignment in a band around INIT1. The result is called OPT. 6. Use Smith-Waterman alignments to display final results. 4.5 Hot-spots Essentially, a hot-spot is a dot in a dot plot. Definition. Let the input sequences be x = x[1... m] and y = y[1... n]. Then a hot-spot is a pair (i, j) such that the substrings of length k starting at the positions i and j are equal, i. e., x[i... i + k 1] = y[j... j + k 1]. Here k = ktup (for k-tuple ) is a user-specified parameter. The idea behind hot-spots is that in most cases, homologous sequences will contain at least a few identical segments. By default, FastA uses ktup = 2 for amino acids and ktup = 6 for nucleic acids. Larger values of ktup increase speed (because fewer hot-spots have to be considered in subsequent steps) and precision, but decrease recall. 4.6 Precision and recall Precision and recall are concepts from information retrieval and can be defined as follows: Precision: The probability that a retrieved item is correct. precision = Recall: The probability that a correct item is retrieved. recall = TP TP + FP TP TP + FN (where TP is true positives, etc.). A perfect precision score of 1.0 means that every result retrieved by a search was relevant (but says nothing about whether all relevant items were retrieved) whereas a perfect recall score of 1.0 means that all relevant items were retrieved by the search (but says nothing about how many irrelevant items were also retrieved).
3 4002 FastA and the chaining problem, by Clemens Gröpl, Gunnar Klau, November 22, 2010, 09: Diagonal runs If there are similar regions, they will show up in the dot plot as a number of hot-spots which are roughly on a diagonal line. FastA gains efficiency by relying on the heuristic assumption that a local alignment will contain several hot-spots along the same diagonal line. Definition. A diagonal run is a set of hot-spots {(i 1, j 1 ),..., (i l, j l )} such that i p j p = i q j q for all 1 p, q l. (They are on exactly the same diagonal.) A diagonal can contain more than one diagonal run, and a diagonal run need not contain all hot-spots on the same diagonal. Each diagonal run is assigned a score. Hot-spots score positive. The gaps between them score negative, depending on their length. At this stage, FastA does not yet use a substitution matrix; it just considers whether the sequence positions are identical or not. For each diagonal, the hot-spots on it are joined by FastA into diagonal runs. The crucial observation is that diagonal runs can be found without looking at the whole matrix. FastA does not create a dot plot! Since ktup is small, we can hash all positions i of x into the bins of a lookup-table, using the substring x[i... i + k 1] as a key. (There are 20 2 = 400 bins for AA and 4 6 = 4096 bins for NA, using the default values for ktup.) Collisions in the hash table can be resolved, e.g., by linked lists. We can fill the hash table in a single pass over x. This way the collision lists are already in sorted order. Example. We use a two-letter alphabet {a, b} and ktup = 2 for simplicity. sequence x aababbaaab key collision list aa 0, 6, 7 ab 1, 3, 8 ba 2, 5 bb 4 Using the hash we can compute the diagonal runs as we walk along sequence y: When we are at position j of y, we look at the positions i 1, i 2,... of x which we find in the hash bin indexed by y[j... j + k 1]. Each pair (i 1, j), (i 2, j),... is a hot-spot, and we can update the corresponding diagonal runs (and their scores) accordingly. Two hot-spots (i, j), (i, j ) are on the same diagonal iff i j = i j. Hence the growing diagonal runs (along the diagonals) can be adressed by an array of size m + n 1. Thus the time needed to compute the diagonal runs is proportional to the number of hot-spots which is usually much smaller than x y. Figure A shows the diagonal runs and the hot-spots which are found using the hash table.
4 FastA and the chaining problem, by Clemens Gröpl, Gunnar Klau, November 22, 2010, 09: j sequence y i_1 i_2 sequence x i_3 4.8 Rescoring the best diagonal runs Next, the 10 highest-scoring diagonal runs are identified and rescored using a substitution matrix. Note that thus far, FastA still does not allow any indels in the alignment; it just counts matches and mismatches. Definition. The best diagonal run is called INIT1. It is marked with a * in Figure B below. 4.9 Chaining of initial regions To account for indels, FastA tries to chain several initial regions into a better alignment. Definition. The resulting alignment is called INITN. The chaining problem can be modeled as a purely graph theoretical problem...
5 4004 FastA and the chaining problem, by Clemens Gröpl, Gunnar Klau, November 22, 2010, 09: The chaining problem Chaining problem or heaviest-path-in-dag problem Find a heaviest path in a directed acyclic graph G = (V, E) with weights w : V E Z on the edges and vertices. Two diagonal runs can be chained iff they do not overlap in one of the sequences. This is the case iff the lower right end of one diagonal run is to the left and top of the upper left end of the other one. and Formally, let be diagonal runs, then we define r = ( [i... i + p 1], [j... j + p 1] ) r = ( [i... i + q 1], [j... j + q 1] ) r < r : i + p 1 < i j + p 1 < j. Let G = (V, E) be the directed graph whose vertex set V are all diagonal runs and whose edge set is E := {(r, r ) r < r }. Clearly, G is acyclic, so we can find a topological ordering of G in O( V + E ) time. That is, we can write V = {r 1,..., r N } such that r i < r j i < j. Moreover, FastA assigns a score s(r) > 0 to every diagonal run r V and a score s(e) < 0 to every edge e E. Note that Dijkstra s algorithm is not applicable, because we have negative edge weights. However, we can use that the graph is acyclic. Example. For simplicity, each letter scores 2 and all other columns of the (resulting) alignment score 1 (diagonal shortcuts allowed). i i g g g d a c d a c a h a k h b k h b f h b f j b f e j b e j b k / 4 a / 8 c / 4-1 g / f / 6-2 j / 6-7 h / 8-1 e / 4 i / 4 d / 4-4 b / 12
6 FastA and the chaining problem, by Clemens Gröpl, Gunnar Klau, November 22, 2010, 09: Formally, we have to compute a path r π(1) < r π(2) <... < r π(l) that maximizes l 1 ( s(rπ(i) ) + s(r π(i), r π(i+1) ) ) + s(r π(l) ). i=1 In the following, we show how to use the algorithm DAG shortest paths for this task. (1) DAG shortest paths(g, w, s, d, π) (2) { (3) G = (V, E) : directed acyclic graph; (4) Adj : V P(V) : set of successor nodes, this implements E; (5) w : E Z : edge weights; (6) d : V Z : distance from s as computed by algorithm; (7) π : V V : predecessor; (8) V = {s = v 1, v 2,..., v n } : topological ordering starting with s. (9) d[s] 0; (10) π[s] nil; (11) for i = 1 to n do (12) foreach ( v Adj(v i ))do (13) c = d[v i ] + w(v i, v); (14) if (c < d[v]) (15) d[v] c; (16) π[v] v i ; (17) fi (18) od (19) od (20) } Theorem Let G = (V, E) be a directed acyclic graph, let s V be a vertex without predecessors and let w : E R be a weight function on the edges (which may take negative values). Let v 1,..., v n be the topological ordering used by Algorithm DAG shortest paths. Then the following holds: After the loop for v i has been run through, d[v i ] equals dist w (s, v i ), the weight of a shortest path from s to v i, and one such path is given by s,..., π(π(v i )), π(v i ), v i. Proof. We apply induction on i = 1,..., n. For v i = s there is nothing to be shown. Next we derive two inequalities from which together the claim will follow. Since backtracking from v i yields a path s,..., π(π(v i )), π(v i ), v i of length d[v i ], we have d[v i ] dist w (s, v i ). Now let P = (s = u 0, u 1,..., u k = v i ) be a shortest path. Since u k 1 is a predecessor of v i, and the loop runs through the vertices in a topological ordering, u k 1 was processed before v i and by our inductive assumption, d[u k 1 ] = dist w (s, u k 1 ). Due to the relaxation step we have d[v i ] = d[u k ] d[u k 1 ] + w(u k 1, u k ) = dist w (s, u k 1 ) + w(u k 1, u k ) = dist w (s, v i ). By combination of these inequalities, it follows that d[v i ] = dist w (s, v i ).
7 4006 FastA and the chaining problem, by Clemens Gröpl, Gunnar Klau, November 22, 2010, 09: Reduction of the problem The chaining problem in FastA appears in a slightly different form from that expected by the DAG shortest path algorithm, so we need to transform the input before we can apply it. There are three issues: 1. Allow any vertex to play the role of s and t 2. Eliminate the vertex weights 3. Reverse the optimization goal Assume that G = (V, E), w : V E Z is an instance of the heaviest-path-in-dag problem. Arbitrary source and target. Add a new vertex s with outgoing edges to all other vertices. Let w(s, x) = 0, w(s) = 0. Add a new vertex t with incoming edges from all other vertices. Let w(x, t) = 0, w(t) = 0. Eliminating the vertex weights. Replace w(u, v) with w (u, v) := w(u, v) + w(v), for all edges (u, v). Then any path s = v 1, v 2,..., v k = t has weight in both problems. w(v 1 ) + w(v 1, v 2 ) + w(v 2 ) + w(v 2, v 3 ) w(v k ) Reversing the optimization goal. Replace w with w := w. Replace max with min. Example. (Finished on blackboard)
8 FastA and the chaining problem, by Clemens Gröpl, Gunnar Klau, November 22, 2010, 09: c / 4 i / 4 b / 12-1 k / 4 g / 6-7 d / a / 8 f / 6 h / j / 6 e / 4 To illustrate the idea, consider the path c k f. total weight: path before reduction: c k f weights: path after reduction: s c k f t weights: Figure C shows the result (INITN) of chaining the initial regions in Figure B Banded alignment Apart from solving the chaining problem, FastA makes another heuristic attempt to improve INIT1.
9 4008 FastA and the chaining problem, by Clemens Gröpl, Gunnar Klau, November 22, 2010, 09:09 Definition. A banded alignment is an alignment that stays within a certain diagonal band of the DP matrix. FastA computes a banded alignment around the diagonal containing INIT1. Definition. This alignment is called OPT. Note that OPT is not necessarily optimal! We use the (confusing) terminology from the original paper. %-) To compute a banded alignment, we simply ignore entries of the DP matrix which are outside the band. Of course, the recurrence has to be modified in the obvious way for cells at the boundary of the band. This reduces running time and space requirement. A band can be specified by an interval I and the condition that computed entries (i, j) satisfy i j I. An example with I = { 1, 0, 1}: 4.13 Final application of Smith-Waterman As is usual practice with heuristic alignment programs, FastA computes an exact local alignment using the Smith-Waterman algorithm (or some variant thereof) before it displays the final result. This is possible in reasonable time, because the search has been narrowed down to a small region Concluding remarks on FastA FastA is actually a collection of programs which are customized for various tasks (DNA, proteins, peptide collections, mixed input types). If FastA is used to search through a large database, one will often find a relatively good hit simply by chance. Therefore, FastA also computes a statistical significance score. This information is very important to assess the results however, the underlying theory is beyond this lecture...
FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:
 FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:56 4001 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem
FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:56 4001 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem
FastA & the chaining problem
 FastA & the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 1 Sources for this lecture: Lectures by Volker Heun, Daniel Huson and Knut Reinert,
FastA & the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 1 Sources for this lecture: Lectures by Volker Heun, Daniel Huson and Knut Reinert,
As of August 15, 2008, GenBank contained bases from reported sequences. The search procedure should be
 48 Bioinformatics I, WS 09-10, S. Henz (script by D. Huson) November 26, 2009 4 BLAST and BLAT Outline of the chapter: 1. Heuristics for the pairwise local alignment of two sequences 2. BLAST: search and
48 Bioinformatics I, WS 09-10, S. Henz (script by D. Huson) November 26, 2009 4 BLAST and BLAT Outline of the chapter: 1. Heuristics for the pairwise local alignment of two sequences 2. BLAST: search and
24 Grundlagen der Bioinformatik, SS 10, D. Huson, April 26, This lecture is based on the following papers, which are all recommended reading:
 24 Grundlagen der Bioinformatik, SS 10, D. Huson, April 26, 2010 3 BLAST and FASTA This lecture is based on the following papers, which are all recommended reading: D.J. Lipman and W.R. Pearson, Rapid
24 Grundlagen der Bioinformatik, SS 10, D. Huson, April 26, 2010 3 BLAST and FASTA This lecture is based on the following papers, which are all recommended reading: D.J. Lipman and W.R. Pearson, Rapid
Computational Molecular Biology
 Computational Molecular Biology Erwin M. Bakker Lecture 3, mainly from material by R. Shamir [2] and H.J. Hoogeboom [4]. 1 Pairwise Sequence Alignment Biological Motivation Algorithmic Aspect Recursive
Computational Molecular Biology Erwin M. Bakker Lecture 3, mainly from material by R. Shamir [2] and H.J. Hoogeboom [4]. 1 Pairwise Sequence Alignment Biological Motivation Algorithmic Aspect Recursive
Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA.
 Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA. Fasta is used to compare a protein or DNA sequence to all of the
Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA. Fasta is used to compare a protein or DNA sequence to all of the
FASTA. Besides that, FASTA package provides SSEARCH, an implementation of the optimal Smith- Waterman algorithm.
 FASTA INTRODUCTION Definition (by David J. Lipman and William R. Pearson in 1985) - Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence
FASTA INTRODUCTION Definition (by David J. Lipman and William R. Pearson in 1985) - Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence
COS 551: Introduction to Computational Molecular Biology Lecture: Oct 17, 2000 Lecturer: Mona Singh Scribe: Jacob Brenner 1. Database Searching
 COS 551: Introduction to Computational Molecular Biology Lecture: Oct 17, 2000 Lecturer: Mona Singh Scribe: Jacob Brenner 1 Database Searching In database search, we typically have a large sequence database
COS 551: Introduction to Computational Molecular Biology Lecture: Oct 17, 2000 Lecturer: Mona Singh Scribe: Jacob Brenner 1 Database Searching In database search, we typically have a large sequence database
Computer Science & Engineering 423/823 Design and Analysis of Algorithms
 Computer Science & Engineering 423/823 Design and Analysis of Algorithms Lecture 07 Single-Source Shortest Paths (Chapter 24) Stephen Scott and Vinodchandran N. Variyam sscott@cse.unl.edu 1/36 Introduction
Computer Science & Engineering 423/823 Design and Analysis of Algorithms Lecture 07 Single-Source Shortest Paths (Chapter 24) Stephen Scott and Vinodchandran N. Variyam sscott@cse.unl.edu 1/36 Introduction
Sequence Alignment (chapter 6) p The biological problem p Global alignment p Local alignment p Multiple alignment
 Sequence lignment (chapter 6) p The biological problem p lobal alignment p Local alignment p Multiple alignment Local alignment: rationale p Otherwise dissimilar proteins may have local regions of similarity
Sequence lignment (chapter 6) p The biological problem p lobal alignment p Local alignment p Multiple alignment Local alignment: rationale p Otherwise dissimilar proteins may have local regions of similarity
Sequence analysis Pairwise sequence alignment
 UMF11 Introduction to bioinformatics, 25 Sequence analysis Pairwise sequence alignment 1. Sequence alignment Lecturer: Marina lexandersson 12 September, 25 here are two types of sequence alignments, global
UMF11 Introduction to bioinformatics, 25 Sequence analysis Pairwise sequence alignment 1. Sequence alignment Lecturer: Marina lexandersson 12 September, 25 here are two types of sequence alignments, global
Pairwise alignment using shortest path algorithms, Gunnar Klau, November 29, 2005, 11:
 airwise alignment using shortest path algorithms, Gunnar Klau, November 9,, : 3 3 airwise alignment using shortest path algorithms e will iscuss: it graph Dijkstra s algorithm algorithm (GDU) 3. References
airwise alignment using shortest path algorithms, Gunnar Klau, November 9,, : 3 3 airwise alignment using shortest path algorithms e will iscuss: it graph Dijkstra s algorithm algorithm (GDU) 3. References
1. R. Durbin, S. Eddy, A. Krogh und G. Mitchison: Biological sequence analysis, Cambridge, 1998
 7 Multiple Sequence Alignment The exposition was prepared by Clemens Gröpl, based on earlier versions by Daniel Huson, Knut Reinert, and Gunnar Klau. It is based on the following sources, which are all
7 Multiple Sequence Alignment The exposition was prepared by Clemens Gröpl, based on earlier versions by Daniel Huson, Knut Reinert, and Gunnar Klau. It is based on the following sources, which are all
Biology 644: Bioinformatics
 Find the best alignment between 2 sequences with lengths n and m, respectively Best alignment is very dependent upon the substitution matrix and gap penalties The Global Alignment Problem tries to find
Find the best alignment between 2 sequences with lengths n and m, respectively Best alignment is very dependent upon the substitution matrix and gap penalties The Global Alignment Problem tries to find
An Analysis of Pairwise Sequence Alignment Algorithm Complexities: Needleman-Wunsch, Smith-Waterman, FASTA, BLAST and Gapped BLAST
 An Analysis of Pairwise Sequence Alignment Algorithm Complexities: Needleman-Wunsch, Smith-Waterman, FASTA, BLAST and Gapped BLAST Alexander Chan 5075504 Biochemistry 218 Final Project An Analysis of Pairwise
An Analysis of Pairwise Sequence Alignment Algorithm Complexities: Needleman-Wunsch, Smith-Waterman, FASTA, BLAST and Gapped BLAST Alexander Chan 5075504 Biochemistry 218 Final Project An Analysis of Pairwise
Bioinformatics for Biologists
 Bioinformatics for Biologists Sequence Analysis: Part I. Pairwise alignment and database searching Fran Lewitter, Ph.D. Director Bioinformatics & Research Computing Whitehead Institute Topics to Cover
Bioinformatics for Biologists Sequence Analysis: Part I. Pairwise alignment and database searching Fran Lewitter, Ph.D. Director Bioinformatics & Research Computing Whitehead Institute Topics to Cover
Lecture 2 Pairwise sequence alignment. Principles Computational Biology Teresa Przytycka, PhD
 Lecture 2 Pairwise sequence alignment. Principles Computational Biology Teresa Przytycka, PhD Assumptions: Biological sequences evolved by evolution. Micro scale changes: For short sequences (e.g. one
Lecture 2 Pairwise sequence alignment. Principles Computational Biology Teresa Przytycka, PhD Assumptions: Biological sequences evolved by evolution. Micro scale changes: For short sequences (e.g. one
Single Source Shortest Path
 Single Source Shortest Path A directed graph G = (V, E) and a pair of nodes s, d is given. The edges have a real-valued weight W i. This time we are looking for the weight and the shortest path from s
Single Source Shortest Path A directed graph G = (V, E) and a pair of nodes s, d is given. The edges have a real-valued weight W i. This time we are looking for the weight and the shortest path from s
Scoring and heuristic methods for sequence alignment CG 17
 Scoring and heuristic methods for sequence alignment CG 17 Amino Acid Substitution Matrices Used to score alignments. Reflect evolution of sequences. Unitary Matrix: M ij = 1 i=j { 0 o/w Genetic Code Matrix:
Scoring and heuristic methods for sequence alignment CG 17 Amino Acid Substitution Matrices Used to score alignments. Reflect evolution of sequences. Unitary Matrix: M ij = 1 i=j { 0 o/w Genetic Code Matrix:
CS 270 Algorithms. Oliver Kullmann. Analysing BFS. Depth-first search. Analysing DFS. Dags and topological sorting.
 General remarks Week 5 2 We finish, by analysing it. Then we consider the second main graph- algorithm, depth-first (). And we consider one application of, of graphs. Reading from CLRS for week 5 Chapter
General remarks Week 5 2 We finish, by analysing it. Then we consider the second main graph- algorithm, depth-first (). And we consider one application of, of graphs. Reading from CLRS for week 5 Chapter
CS 341: Algorithms. Douglas R. Stinson. David R. Cheriton School of Computer Science University of Waterloo. February 26, 2019
 CS 341: Algorithms Douglas R. Stinson David R. Cheriton School of Computer Science University of Waterloo February 26, 2019 D.R. Stinson (SCS) CS 341 February 26, 2019 1 / 296 1 Course Information 2 Introduction
CS 341: Algorithms Douglas R. Stinson David R. Cheriton School of Computer Science University of Waterloo February 26, 2019 D.R. Stinson (SCS) CS 341 February 26, 2019 1 / 296 1 Course Information 2 Introduction
CISC 636 Computational Biology & Bioinformatics (Fall 2016)
 CISC 636 Computational Biology & Bioinformatics (Fall 2016) Sequence pairwise alignment Score statistics: E-value and p-value Heuristic algorithms: BLAST and FASTA Database search: gene finding and annotations
CISC 636 Computational Biology & Bioinformatics (Fall 2016) Sequence pairwise alignment Score statistics: E-value and p-value Heuristic algorithms: BLAST and FASTA Database search: gene finding and annotations
Alignment of Long Sequences
 Alignment of Long Sequences BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2009 Mark Craven craven@biostat.wisc.edu Pairwise Whole Genome Alignment: Task Definition Given a pair of genomes (or other large-scale
Alignment of Long Sequences BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2009 Mark Craven craven@biostat.wisc.edu Pairwise Whole Genome Alignment: Task Definition Given a pair of genomes (or other large-scale
Shortest Path Problem
 Shortest Path Problem CLRS Chapters 24.1 3, 24.5, 25.2 Shortest path problem Shortest path problem (and variants) Properties of shortest paths Algorithmic framework Bellman-Ford algorithm Shortest paths
Shortest Path Problem CLRS Chapters 24.1 3, 24.5, 25.2 Shortest path problem Shortest path problem (and variants) Properties of shortest paths Algorithmic framework Bellman-Ford algorithm Shortest paths
CS 270 Algorithms. Oliver Kullmann. Breadth-first search. Analysing BFS. Depth-first. search. Analysing DFS. Dags and topological sorting.
 Week 5 General remarks and 2 We consider the simplest graph- algorithm, breadth-first (). We apply to compute shortest paths. Then we consider the second main graph- algorithm, depth-first (). And we consider
Week 5 General remarks and 2 We consider the simplest graph- algorithm, breadth-first (). We apply to compute shortest paths. Then we consider the second main graph- algorithm, depth-first (). And we consider
Computational Genomics and Molecular Biology, Fall
 Computational Genomics and Molecular Biology, Fall 2015 1 Sequence Alignment Dannie Durand Pairwise Sequence Alignment The goal of pairwise sequence alignment is to establish a correspondence between the
Computational Genomics and Molecular Biology, Fall 2015 1 Sequence Alignment Dannie Durand Pairwise Sequence Alignment The goal of pairwise sequence alignment is to establish a correspondence between the
USING AN EXTENDED SUFFIX TREE TO SPEED-UP SEQUENCE ALIGNMENT
 IADIS International Conference Applied Computing 2006 USING AN EXTENDED SUFFIX TREE TO SPEED-UP SEQUENCE ALIGNMENT Divya R. Singh Software Engineer Microsoft Corporation, Redmond, WA 98052, USA Abdullah
IADIS International Conference Applied Computing 2006 USING AN EXTENDED SUFFIX TREE TO SPEED-UP SEQUENCE ALIGNMENT Divya R. Singh Software Engineer Microsoft Corporation, Redmond, WA 98052, USA Abdullah
Week 5. 1 Analysing BFS. 2 Depth-first search. 3 Analysing DFS. 4 Dags and topological sorting. 5 Detecting cycles. CS 270 Algorithms.
 1 2 Week 5 3 4 5 General remarks We finish, by analysing it. Then we consider the second main graph- algorithm, depth-first (). And we consider one application of, of graphs. Reading from CLRS for week
1 2 Week 5 3 4 5 General remarks We finish, by analysing it. Then we consider the second main graph- algorithm, depth-first (). And we consider one application of, of graphs. Reading from CLRS for week
Introduction. I Given a weighted, directed graph G =(V, E) with weight function
 ntroduction Computer Science & Engineering 2/82 Design and Analysis of Algorithms Lecture 06 Single-Source Shortest Paths (Chapter 2) Stephen Scott (Adapted from Vinodchandran N. Variyam) sscott@cse.unl.edu
ntroduction Computer Science & Engineering 2/82 Design and Analysis of Algorithms Lecture 06 Single-Source Shortest Paths (Chapter 2) Stephen Scott (Adapted from Vinodchandran N. Variyam) sscott@cse.unl.edu
Utility of Sliding Window FASTA in Predicting Cross- Reactivity with Allergenic Proteins. Bob Cressman Pioneer Crop Genetics
 Utility of Sliding Window FASTA in Predicting Cross- Reactivity with Allergenic Proteins Bob Cressman Pioneer Crop Genetics The issue FAO/WHO 2001 Step 2: prepare a complete set of 80-amino acid length
Utility of Sliding Window FASTA in Predicting Cross- Reactivity with Allergenic Proteins Bob Cressman Pioneer Crop Genetics The issue FAO/WHO 2001 Step 2: prepare a complete set of 80-amino acid length
Minimum Spanning Trees
 Minimum Spanning Trees Problem A town has a set of houses and a set of roads. A road connects 2 and only 2 houses. A road connecting houses u and v has a repair cost w(u, v). Goal: Repair enough (and no
Minimum Spanning Trees Problem A town has a set of houses and a set of roads. A road connects 2 and only 2 houses. A road connecting houses u and v has a repair cost w(u, v). Goal: Repair enough (and no
Finding homologous sequences in databases
 Finding homologous sequences in databases There are multiple algorithms to search sequences databases BLAST (EMBL, NCBI, DDBJ, local) FASTA (EMBL, local) For protein only databases scan via Smith-Waterman
Finding homologous sequences in databases There are multiple algorithms to search sequences databases BLAST (EMBL, NCBI, DDBJ, local) FASTA (EMBL, local) For protein only databases scan via Smith-Waterman
The Encoding Complexity of Network Coding
 The Encoding Complexity of Network Coding Michael Langberg Alexander Sprintson Jehoshua Bruck California Institute of Technology Email: mikel,spalex,bruck @caltech.edu Abstract In the multicast network
The Encoding Complexity of Network Coding Michael Langberg Alexander Sprintson Jehoshua Bruck California Institute of Technology Email: mikel,spalex,bruck @caltech.edu Abstract In the multicast network
Sequence Alignment Heuristics
 Sequence Alignment Heuristics Some slides from: Iosif Vaisman, GMU mason.gmu.edu/~mmasso/binf630alignment.ppt Serafim Batzoglu, Stanford http://ai.stanford.edu/~serafim/ Geoffrey J. Barton, Oxford Protein
Sequence Alignment Heuristics Some slides from: Iosif Vaisman, GMU mason.gmu.edu/~mmasso/binf630alignment.ppt Serafim Batzoglu, Stanford http://ai.stanford.edu/~serafim/ Geoffrey J. Barton, Oxford Protein
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 04: Variations of sequence alignments http://www.pitt.edu/~mcs2/teaching/biocomp/tutorials/global.html Slides adapted from Dr. Shaojie Zhang (University
EECS730: Introduction to Bioinformatics Lecture 04: Variations of sequence alignments http://www.pitt.edu/~mcs2/teaching/biocomp/tutorials/global.html Slides adapted from Dr. Shaojie Zhang (University
Dynamic Programming Part I: Examples. Bioinfo I (Institut Pasteur de Montevideo) Dynamic Programming -class4- July 25th, / 77
 Dynamic Programming Part I: Examples Bioinfo I (Institut Pasteur de Montevideo) Dynamic Programming -class4- July 25th, 2011 1 / 77 Dynamic Programming Recall: the Change Problem Other problems: Manhattan
Dynamic Programming Part I: Examples Bioinfo I (Institut Pasteur de Montevideo) Dynamic Programming -class4- July 25th, 2011 1 / 77 Dynamic Programming Recall: the Change Problem Other problems: Manhattan
Algorithms and Data Structures 2014 Exercises and Solutions Week 9
 Algorithms and Data Structures 2014 Exercises and Solutions Week 9 November 26, 2014 1 Directed acyclic graphs We are given a sequence (array) of numbers, and we would like to find the longest increasing
Algorithms and Data Structures 2014 Exercises and Solutions Week 9 November 26, 2014 1 Directed acyclic graphs We are given a sequence (array) of numbers, and we would like to find the longest increasing
Lecture Overview. Sequence search & alignment. Searching sequence databases. Sequence Alignment & Search. Goals: Motivations:
 Lecture Overview Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating
Lecture Overview Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating
TCCAGGTG-GAT TGCAAGTGCG-T. Local Sequence Alignment & Heuristic Local Aligners. Review: Probabilistic Interpretation. Chance or true homology?
 Local Sequence Alignment & Heuristic Local Aligners Lectures 18 Nov 28, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20 Johnson Hall
Local Sequence Alignment & Heuristic Local Aligners Lectures 18 Nov 28, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20 Johnson Hall
1. R. Durbin, S. Eddy, A. Krogh und G. Mitchison: Biological sequence analysis, Cambridge, 1998
 7 Multiple Sequence Alignment The exposition was prepared by Clemens GrÃP pl, based on earlier versions by Daniel Huson, Knut Reinert, and Gunnar Klau. It is based on the following sources, which are all
7 Multiple Sequence Alignment The exposition was prepared by Clemens GrÃP pl, based on earlier versions by Daniel Huson, Knut Reinert, and Gunnar Klau. It is based on the following sources, which are all
Sequence alignment theory and applications Session 3: BLAST algorithm
 Sequence alignment theory and applications Session 3: BLAST algorithm Introduction to Bioinformatics online course : IBT Sonal Henson Learning Objectives Understand the principles of the BLAST algorithm
Sequence alignment theory and applications Session 3: BLAST algorithm Introduction to Bioinformatics online course : IBT Sonal Henson Learning Objectives Understand the principles of the BLAST algorithm
Sequence Alignment & Search
 Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating the first version
Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating the first version
Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it.
 Sequence Alignments Overview Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it. Sequence alignment means arranging
Sequence Alignments Overview Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it. Sequence alignment means arranging
V1.0: Seth Gilbert, V1.1: Steven Halim August 30, Abstract. d(e), and we assume that the distance function is non-negative (i.e., d(x, y) 0).
 CS4234: Optimisation Algorithms Lecture 4 TRAVELLING-SALESMAN-PROBLEM (4 variants) V1.0: Seth Gilbert, V1.1: Steven Halim August 30, 2016 Abstract The goal of the TRAVELLING-SALESMAN-PROBLEM is to find
CS4234: Optimisation Algorithms Lecture 4 TRAVELLING-SALESMAN-PROBLEM (4 variants) V1.0: Seth Gilbert, V1.1: Steven Halim August 30, 2016 Abstract The goal of the TRAVELLING-SALESMAN-PROBLEM is to find
1 Dijkstra s Algorithm
 Lecture 11 Dijkstra s Algorithm Scribes: Himanshu Bhandoh (2015), Virginia Williams, and Date: November 1, 2017 Anthony Kim (2016), G. Valiant (2017), M. Wootters (2017) (Adapted from Virginia Williams
Lecture 11 Dijkstra s Algorithm Scribes: Himanshu Bhandoh (2015), Virginia Williams, and Date: November 1, 2017 Anthony Kim (2016), G. Valiant (2017), M. Wootters (2017) (Adapted from Virginia Williams
From Smith-Waterman to BLAST
 From Smith-Waterman to BLAST Jeremy Buhler July 23, 2015 Smith-Waterman is the fundamental tool that we use to decide how similar two sequences are. Isn t that all that BLAST does? In principle, it is
From Smith-Waterman to BLAST Jeremy Buhler July 23, 2015 Smith-Waterman is the fundamental tool that we use to decide how similar two sequences are. Isn t that all that BLAST does? In principle, it is
Lecture 22 Tuesday, April 10
 CIS 160 - Spring 2018 (instructor Val Tannen) Lecture 22 Tuesday, April 10 GRAPH THEORY Directed Graphs Directed graphs (a.k.a. digraphs) are an important mathematical modeling tool in Computer Science,
CIS 160 - Spring 2018 (instructor Val Tannen) Lecture 22 Tuesday, April 10 GRAPH THEORY Directed Graphs Directed graphs (a.k.a. digraphs) are an important mathematical modeling tool in Computer Science,
Algorithms in Bioinformatics: A Practical Introduction. Database Search
 Algorithms in Bioinformatics: A Practical Introduction Database Search Biological databases Biological data is double in size every 15 or 16 months Increasing in number of queries: 40,000 queries per day
Algorithms in Bioinformatics: A Practical Introduction Database Search Biological databases Biological data is double in size every 15 or 16 months Increasing in number of queries: 40,000 queries per day
Computer Science & Engineering 423/823 Design and Analysis of Algorithms
 s of s Computer Science & Engineering 423/823 Design and Analysis of Lecture 03 (Chapter 22) Stephen Scott (Adapted from Vinodchandran N. Variyam) 1 / 29 s of s s are abstract data types that are applicable
s of s Computer Science & Engineering 423/823 Design and Analysis of Lecture 03 (Chapter 22) Stephen Scott (Adapted from Vinodchandran N. Variyam) 1 / 29 s of s s are abstract data types that are applicable
BLAST: Basic Local Alignment Search Tool Altschul et al. J. Mol Bio CS 466 Saurabh Sinha
 BLAST: Basic Local Alignment Search Tool Altschul et al. J. Mol Bio. 1990. CS 466 Saurabh Sinha Motivation Sequence homology to a known protein suggest function of newly sequenced protein Bioinformatics
BLAST: Basic Local Alignment Search Tool Altschul et al. J. Mol Bio. 1990. CS 466 Saurabh Sinha Motivation Sequence homology to a known protein suggest function of newly sequenced protein Bioinformatics
Algorithm Design, Anal. & Imp., Homework 4 Solution
 Algorithm Design, Anal. & Imp., Homework 4 Solution Note: The solution is for your personal use for this course. You are not allowed to post the solution in public place. There could be mistakes in the
Algorithm Design, Anal. & Imp., Homework 4 Solution Note: The solution is for your personal use for this course. You are not allowed to post the solution in public place. There could be mistakes in the
Lecture 10. Sequence alignments
 Lecture 10 Sequence alignments Alignment algorithms: Overview Given a scoring system, we need to have an algorithm for finding an optimal alignment for a pair of sequences. We want to maximize the score
Lecture 10 Sequence alignments Alignment algorithms: Overview Given a scoring system, we need to have an algorithm for finding an optimal alignment for a pair of sequences. We want to maximize the score
CSE 331: Introduction to Algorithm Analysis and Design Graphs
 CSE 331: Introduction to Algorithm Analysis and Design Graphs 1 Graph Definitions Graph: A graph consists of a set of verticies V and a set of edges E such that: G = (V, E) V = {v 0, v 1,..., v n 1 } E
CSE 331: Introduction to Algorithm Analysis and Design Graphs 1 Graph Definitions Graph: A graph consists of a set of verticies V and a set of edges E such that: G = (V, E) V = {v 0, v 1,..., v n 1 } E
Sequence alignment algorithms
 Sequence alignment algorithms Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 23 rd 27 After this lecture, you can decide when to use local and global sequence alignments
Sequence alignment algorithms Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 23 rd 27 After this lecture, you can decide when to use local and global sequence alignments
Introduction. I Given a weighted, directed graph G =(V, E) with weight function
 ntroduction Computer Science & Engineering 2/82 Design and Analysis of Algorithms Lecture 05 Single-Source Shortest Paths (Chapter 2) Stephen Scott (Adapted from Vinodchandran N. Variyam) sscott@cse.unl.edu
ntroduction Computer Science & Engineering 2/82 Design and Analysis of Algorithms Lecture 05 Single-Source Shortest Paths (Chapter 2) Stephen Scott (Adapted from Vinodchandran N. Variyam) sscott@cse.unl.edu
Chapter 22. Elementary Graph Algorithms
 Graph Algorithms - Spring 2011 Set 7. Lecturer: Huilan Chang Reference: (1) Cormen, Leiserson, Rivest, and Stein, Introduction to Algorithms, 2nd Edition, The MIT Press. (2) Lecture notes from C. Y. Chen
Graph Algorithms - Spring 2011 Set 7. Lecturer: Huilan Chang Reference: (1) Cormen, Leiserson, Rivest, and Stein, Introduction to Algorithms, 2nd Edition, The MIT Press. (2) Lecture notes from C. Y. Chen
Graph Search. Adnan Aziz
 Graph Search Adnan Aziz Based on CLRS, Ch 22. Recall encountered graphs several weeks ago (CLRS B.4) restricted our attention to definitions, terminology, properties Now we ll see how to perform basic
Graph Search Adnan Aziz Based on CLRS, Ch 22. Recall encountered graphs several weeks ago (CLRS B.4) restricted our attention to definitions, terminology, properties Now we ll see how to perform basic
Graph Algorithms. Definition
 Graph Algorithms Many problems in CS can be modeled as graph problems. Algorithms for solving graph problems are fundamental to the field of algorithm design. Definition A graph G = (V, E) consists of
Graph Algorithms Many problems in CS can be modeled as graph problems. Algorithms for solving graph problems are fundamental to the field of algorithm design. Definition A graph G = (V, E) consists of
Bioinformatics explained: Smith-Waterman
 Bioinformatics Explained Bioinformatics explained: Smith-Waterman May 1, 2007 CLC bio Gustav Wieds Vej 10 8000 Aarhus C Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com info@clcbio.com
Bioinformatics Explained Bioinformatics explained: Smith-Waterman May 1, 2007 CLC bio Gustav Wieds Vej 10 8000 Aarhus C Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com info@clcbio.com
In this section we describe how to extend the match refinement to the multiple case and then use T-Coffee to heuristically compute a multiple trace.
 5 Multiple Match Refinement and T-Coffee In this section we describe how to extend the match refinement to the multiple case and then use T-Coffee to heuristically compute a multiple trace. This exposition
5 Multiple Match Refinement and T-Coffee In this section we describe how to extend the match refinement to the multiple case and then use T-Coffee to heuristically compute a multiple trace. This exposition
Special course in Computer Science: Advanced Text Algorithms
 Special course in Computer Science: Advanced Text Algorithms Lecture 8: Multiple alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
Special course in Computer Science: Advanced Text Algorithms Lecture 8: Multiple alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
Jyoti Lakhani 1, Ajay Khunteta 2, Dharmesh Harwani *3 1 Poornima University, Jaipur & Maharaja Ganga Singh University, Bikaner, Rajasthan, India
 International Journal of Scientific Research in Computer Science, Engineering and Information Technology 2017 IJSRCSEIT Volume 2 Issue 6 ISSN : 2456-3307 Improvisation of Global Pairwise Sequence Alignment
International Journal of Scientific Research in Computer Science, Engineering and Information Technology 2017 IJSRCSEIT Volume 2 Issue 6 ISSN : 2456-3307 Improvisation of Global Pairwise Sequence Alignment
Research Article International Journals of Advanced Research in Computer Science and Software Engineering ISSN: X (Volume-7, Issue-6)
 International Journals of Advanced Research in Computer Science and Software Engineering ISSN: 77-18X (Volume-7, Issue-6) Research Article June 017 DDGARM: Dotlet Driven Global Alignment with Reduced Matrix
International Journals of Advanced Research in Computer Science and Software Engineering ISSN: 77-18X (Volume-7, Issue-6) Research Article June 017 DDGARM: Dotlet Driven Global Alignment with Reduced Matrix
Lecture 10. Elementary Graph Algorithm Minimum Spanning Trees
 Lecture 10. Elementary Graph Algorithm Minimum Spanning Trees T. H. Cormen, C. E. Leiserson and R. L. Rivest Introduction to Algorithms, 3rd Edition, MIT Press, 2009 Sungkyunkwan University Hyunseung Choo
Lecture 10. Elementary Graph Algorithm Minimum Spanning Trees T. H. Cormen, C. E. Leiserson and R. L. Rivest Introduction to Algorithms, 3rd Edition, MIT Press, 2009 Sungkyunkwan University Hyunseung Choo
Single Source Shortest Paths
 Single Source Shortest Paths Given a connected weighted directed graph G(V, E), associated with each edge u, v E, there is a weight w(u, v). The single source shortest paths (SSSP) problem is to find a
Single Source Shortest Paths Given a connected weighted directed graph G(V, E), associated with each edge u, v E, there is a weight w(u, v). The single source shortest paths (SSSP) problem is to find a
Chapter 4: Blast. Chaochun Wei Fall 2014
 Course organization Introduction ( Week 1-2) Course introduction A brief introduction to molecular biology A brief introduction to sequence comparison Part I: Algorithms for Sequence Analysis (Week 3-11)
Course organization Introduction ( Week 1-2) Course introduction A brief introduction to molecular biology A brief introduction to sequence comparison Part I: Algorithms for Sequence Analysis (Week 3-11)
Lecture 5: Multiple sequence alignment
 Lecture 5: Multiple sequence alignment Introduction to Computational Biology Teresa Przytycka, PhD (with some additions by Martin Vingron) Why do we need multiple sequence alignment Pairwise sequence alignment
Lecture 5: Multiple sequence alignment Introduction to Computational Biology Teresa Przytycka, PhD (with some additions by Martin Vingron) Why do we need multiple sequence alignment Pairwise sequence alignment
Unit 2: Algorithmic Graph Theory
 Unit 2: Algorithmic Graph Theory Course contents: Introduction to graph theory Basic graph algorithms Reading Chapter 3 Reference: Cormen, Leiserson, and Rivest, Introduction to Algorithms, 2 nd Ed., McGraw
Unit 2: Algorithmic Graph Theory Course contents: Introduction to graph theory Basic graph algorithms Reading Chapter 3 Reference: Cormen, Leiserson, and Rivest, Introduction to Algorithms, 2 nd Ed., McGraw
Sample Solutions to Homework #4
 National Taiwan University Handout #25 Department of Electrical Engineering January 02, 207 Algorithms, Fall 206 TA: Zhi-Wen Lin and Yen-Chun Liu Sample Solutions to Homework #4. (0) (a) Both of the answers
National Taiwan University Handout #25 Department of Electrical Engineering January 02, 207 Algorithms, Fall 206 TA: Zhi-Wen Lin and Yen-Chun Liu Sample Solutions to Homework #4. (0) (a) Both of the answers
Bioinformatics explained: BLAST. March 8, 2007
 Bioinformatics Explained Bioinformatics explained: BLAST March 8, 2007 CLC bio Gustav Wieds Vej 10 8000 Aarhus C Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com info@clcbio.com Bioinformatics
Bioinformatics Explained Bioinformatics explained: BLAST March 8, 2007 CLC bio Gustav Wieds Vej 10 8000 Aarhus C Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com info@clcbio.com Bioinformatics
Computer Science & Engineering 423/823 Design and Analysis of Algorithms
 Computer Science & Engineering 423/823 Design and Analysis of Algorithms Lecture 04 Elementary Graph Algorithms (Chapter 22) Stephen Scott (Adapted from Vinodchandran N. Variyam) sscott@cse.unl.edu Introduction
Computer Science & Engineering 423/823 Design and Analysis of Algorithms Lecture 04 Elementary Graph Algorithms (Chapter 22) Stephen Scott (Adapted from Vinodchandran N. Variyam) sscott@cse.unl.edu Introduction
6.006 Introduction to Algorithms Spring 2008
 MIT OpenCourseWare http://ocw.mit.edu 6.006 Introduction to Algorithms Spring 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms. Introduction to Algorithms:
MIT OpenCourseWare http://ocw.mit.edu 6.006 Introduction to Algorithms Spring 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms. Introduction to Algorithms:
CISC 889 Bioinformatics (Spring 2003) Multiple Sequence Alignment
 CISC 889 Bioinformatics (Spring 2003) Multiple Sequence Alignment Courtesy of jalview 1 Motivations Collective statistic Protein families Identification and representation of conserved sequence features
CISC 889 Bioinformatics (Spring 2003) Multiple Sequence Alignment Courtesy of jalview 1 Motivations Collective statistic Protein families Identification and representation of conserved sequence features
Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014
 Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014 Dynamic programming is a group of mathematical methods used to sequentially split a complicated problem into
Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014 Dynamic programming is a group of mathematical methods used to sequentially split a complicated problem into
Pairwise Sequence Alignment. Zhongming Zhao, PhD
 Pairwise Sequence Alignment Zhongming Zhao, PhD Email: zhongming.zhao@vanderbilt.edu http://bioinfo.mc.vanderbilt.edu/ Sequence Similarity match mismatch A T T A C G C G T A C C A T A T T A T G C G A T
Pairwise Sequence Alignment Zhongming Zhao, PhD Email: zhongming.zhao@vanderbilt.edu http://bioinfo.mc.vanderbilt.edu/ Sequence Similarity match mismatch A T T A C G C G T A C C A T A T T A T G C G A T
Lecture 11: Analysis of Algorithms (CS ) 1
 Lecture 11: Analysis of Algorithms (CS583-002) 1 Amarda Shehu November 12, 2014 1 Some material adapted from Kevin Wayne s Algorithm Class @ Princeton 1 2 Dynamic Programming Approach Floyd-Warshall Shortest
Lecture 11: Analysis of Algorithms (CS583-002) 1 Amarda Shehu November 12, 2014 1 Some material adapted from Kevin Wayne s Algorithm Class @ Princeton 1 2 Dynamic Programming Approach Floyd-Warshall Shortest
CSC 373: Algorithm Design and Analysis Lecture 8
 CSC 373: Algorithm Design and Analysis Lecture 8 Allan Borodin January 23, 2013 1 / 19 Lecture 8: Announcements and Outline Announcements No lecture (or tutorial) this Friday. Lecture and tutorials as
CSC 373: Algorithm Design and Analysis Lecture 8 Allan Borodin January 23, 2013 1 / 19 Lecture 8: Announcements and Outline Announcements No lecture (or tutorial) this Friday. Lecture and tutorials as
CS521 \ Notes for the Final Exam
 CS521 \ Notes for final exam 1 Ariel Stolerman Asymptotic Notations: CS521 \ Notes for the Final Exam Notation Definition Limit Big-O ( ) Small-o ( ) Big- ( ) Small- ( ) Big- ( ) Notes: ( ) ( ) ( ) ( )
CS521 \ Notes for final exam 1 Ariel Stolerman Asymptotic Notations: CS521 \ Notes for the Final Exam Notation Definition Limit Big-O ( ) Small-o ( ) Big- ( ) Small- ( ) Big- ( ) Notes: ( ) ( ) ( ) ( )
Brief review from last class
 Sequence Alignment Brief review from last class DNA is has direction, we will use only one (5 -> 3 ) and generate the opposite strand as needed. DNA is a 3D object (see lecture 1) but we will model it
Sequence Alignment Brief review from last class DNA is has direction, we will use only one (5 -> 3 ) and generate the opposite strand as needed. DNA is a 3D object (see lecture 1) but we will model it
CSE 431/531: Algorithm Analysis and Design (Spring 2018) Greedy Algorithms. Lecturer: Shi Li
 CSE 431/531: Algorithm Analysis and Design (Spring 2018) Greedy Algorithms Lecturer: Shi Li Department of Computer Science and Engineering University at Buffalo Main Goal of Algorithm Design Design fast
CSE 431/531: Algorithm Analysis and Design (Spring 2018) Greedy Algorithms Lecturer: Shi Li Department of Computer Science and Engineering University at Buffalo Main Goal of Algorithm Design Design fast
Pairwise Sequence alignment Basic Algorithms
 Pairwise Sequence alignment Basic Algorithms Agenda - Previous Lesson: Minhala - + Biological Story on Biomolecular Sequences - + General Overview of Problems in Computational Biology - Reminder: Dynamic
Pairwise Sequence alignment Basic Algorithms Agenda - Previous Lesson: Minhala - + Biological Story on Biomolecular Sequences - + General Overview of Problems in Computational Biology - Reminder: Dynamic
1 The Shortest Path Problem
 CS161 Lecture 11 Single Source Shortest Paths Scribed by: Romil Verma (2015), Virginia Date: May 8, 2016 1 The Shortest Path Problem In this lecture, we ll discuss the shortest path problem. Assume we
CS161 Lecture 11 Single Source Shortest Paths Scribed by: Romil Verma (2015), Virginia Date: May 8, 2016 1 The Shortest Path Problem In this lecture, we ll discuss the shortest path problem. Assume we
BIOL 7020 Special Topics Cell/Molecular: Molecular Phylogenetics. Spring 2010 Section A
 BIOL 7020 Special Topics Cell/Molecular: Molecular Phylogenetics. Spring 2010 Section A Steve Thompson: stthompson@valdosta.edu http://www.bioinfo4u.net 1 Similarity searching and homology First, just
BIOL 7020 Special Topics Cell/Molecular: Molecular Phylogenetics. Spring 2010 Section A Steve Thompson: stthompson@valdosta.edu http://www.bioinfo4u.net 1 Similarity searching and homology First, just
Data Mining Technologies for Bioinformatics Sequences
 Data Mining Technologies for Bioinformatics Sequences Deepak Garg Computer Science and Engineering Department Thapar Institute of Engineering & Tecnology, Patiala Abstract Main tool used for sequence alignment
Data Mining Technologies for Bioinformatics Sequences Deepak Garg Computer Science and Engineering Department Thapar Institute of Engineering & Tecnology, Patiala Abstract Main tool used for sequence alignment
Database Searching Using BLAST
 Mahidol University Objectives SCMI512 Molecular Sequence Analysis Database Searching Using BLAST Lecture 2B After class, students should be able to: explain the FASTA algorithm for database searching explain
Mahidol University Objectives SCMI512 Molecular Sequence Analysis Database Searching Using BLAST Lecture 2B After class, students should be able to: explain the FASTA algorithm for database searching explain
2386 IEEE TRANSACTIONS ON INFORMATION THEORY, VOL. 52, NO. 6, JUNE 2006
 2386 IEEE TRANSACTIONS ON INFORMATION THEORY, VOL. 52, NO. 6, JUNE 2006 The Encoding Complexity of Network Coding Michael Langberg, Member, IEEE, Alexander Sprintson, Member, IEEE, and Jehoshua Bruck,
2386 IEEE TRANSACTIONS ON INFORMATION THEORY, VOL. 52, NO. 6, JUNE 2006 The Encoding Complexity of Network Coding Michael Langberg, Member, IEEE, Alexander Sprintson, Member, IEEE, and Jehoshua Bruck,
Problem Set 7 Solutions
 " [ Introduction to Algorithms November 29, 200 Massachusetts Institute of Technology 6046J/18410J Professors Shafi Goldwasser and Silvio Micali Handout 27 Problem Set 7 Solutions Problem 7-1 Shortest
" [ Introduction to Algorithms November 29, 200 Massachusetts Institute of Technology 6046J/18410J Professors Shafi Goldwasser and Silvio Micali Handout 27 Problem Set 7 Solutions Problem 7-1 Shortest
BLAST, Profile, and PSI-BLAST
 BLAST, Profile, and PSI-BLAST Jianlin Cheng, PhD School of Electrical Engineering and Computer Science University of Central Florida 26 Free for academic use Copyright @ Jianlin Cheng & original sources
BLAST, Profile, and PSI-BLAST Jianlin Cheng, PhD School of Electrical Engineering and Computer Science University of Central Florida 26 Free for academic use Copyright @ Jianlin Cheng & original sources
Computer Science & Engineering 423/823 Design and Analysis of Algorithms
 s of s Computer Science & Engineering 423/823 Design and Analysis of Lecture 03 (Chapter 22) Stephen Scott (Adapted from Vinodchandran N. Variyam) 1 / 29 Spring 2010 s of s s are abstract data types that
s of s Computer Science & Engineering 423/823 Design and Analysis of Lecture 03 (Chapter 22) Stephen Scott (Adapted from Vinodchandran N. Variyam) 1 / 29 Spring 2010 s of s s are abstract data types that
Chapter 9. Greedy Technique. Copyright 2007 Pearson Addison-Wesley. All rights reserved.
 Chapter 9 Greedy Technique Copyright 2007 Pearson Addison-Wesley. All rights reserved. Greedy Technique Constructs a solution to an optimization problem piece by piece through a sequence of choices that
Chapter 9 Greedy Technique Copyright 2007 Pearson Addison-Wesley. All rights reserved. Greedy Technique Constructs a solution to an optimization problem piece by piece through a sequence of choices that
Algorithm Design and Analysis
 Algorithm Design and Analysis LECTURE 6 Greedy Graph Algorithms Shortest paths Adam Smith 9/8/14 The (Algorithm) Design Process 1. Work out the answer for some examples. Look for a general principle Does
Algorithm Design and Analysis LECTURE 6 Greedy Graph Algorithms Shortest paths Adam Smith 9/8/14 The (Algorithm) Design Process 1. Work out the answer for some examples. Look for a general principle Does
Computational Biology Lecture 4: Overlap detection, Local Alignment, Space Efficient Needleman-Wunsch Saad Mneimneh
 Computational Biology Lecture 4: Overlap detection, Local Alignment, Space Efficient Needleman-Wunsch Saad Mneimneh Overlap detection: Semi-Global Alignment An overlap of two sequences is considered an
Computational Biology Lecture 4: Overlap detection, Local Alignment, Space Efficient Needleman-Wunsch Saad Mneimneh Overlap detection: Semi-Global Alignment An overlap of two sequences is considered an
Shortest Path Algorithm
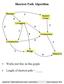 Shortest Path Algorithm C Works just fine on this graph. C Length of shortest path = Copyright 2005 DIMACS BioMath Connect Institute Robert Hochberg Dynamic Programming SP #1 Same Questions, Different
Shortest Path Algorithm C Works just fine on this graph. C Length of shortest path = Copyright 2005 DIMACS BioMath Connect Institute Robert Hochberg Dynamic Programming SP #1 Same Questions, Different
BLAST & Genome assembly
 BLAST & Genome assembly Solon P. Pissis Tomáš Flouri Heidelberg Institute for Theoretical Studies May 15, 2014 1 BLAST What is BLAST? The algorithm 2 Genome assembly De novo assembly Mapping assembly 3
BLAST & Genome assembly Solon P. Pissis Tomáš Flouri Heidelberg Institute for Theoretical Studies May 15, 2014 1 BLAST What is BLAST? The algorithm 2 Genome assembly De novo assembly Mapping assembly 3
Taking Stock. IE170: Algorithms in Systems Engineering: Lecture 20. Example. Shortest Paths Definitions
 Taking Stock IE170: Algorithms in Systems Engineering: Lecture 20 Jeff Linderoth Department of Industrial and Systems Engineering Lehigh University March 19, 2007 Last Time Minimum Spanning Trees Strongly
Taking Stock IE170: Algorithms in Systems Engineering: Lecture 20 Jeff Linderoth Department of Industrial and Systems Engineering Lehigh University March 19, 2007 Last Time Minimum Spanning Trees Strongly
Computational Molecular Biology
 Computational Molecular Biology Erwin M. Bakker Lecture 2 Materials used from R. Shamir [2] and H.J. Hoogeboom [4]. 1 Molecular Biology Sequences DNA A, T, C, G RNA A, U, C, G Protein A, R, D, N, C E,
Computational Molecular Biology Erwin M. Bakker Lecture 2 Materials used from R. Shamir [2] and H.J. Hoogeboom [4]. 1 Molecular Biology Sequences DNA A, T, C, G RNA A, U, C, G Protein A, R, D, N, C E,
Today s Lecture. Multiple sequence alignment. Improved scoring of pairwise alignments. Affine gap penalties Profiles
 Today s Lecture Multiple sequence alignment Improved scoring of pairwise alignments Affine gap penalties Profiles 1 The Edit Graph for a Pair of Sequences G A C G T T G A A T G A C C C A C A T G A C G
Today s Lecture Multiple sequence alignment Improved scoring of pairwise alignments Affine gap penalties Profiles 1 The Edit Graph for a Pair of Sequences G A C G T T G A A T G A C C C A C A T G A C G
New Implementation for the Multi-sequence All-Against-All Substring Matching Problem
 New Implementation for the Multi-sequence All-Against-All Substring Matching Problem Oana Sandu Supervised by Ulrike Stege In collaboration with Chris Upton, Alex Thomo, and Marina Barsky University of
New Implementation for the Multi-sequence All-Against-All Substring Matching Problem Oana Sandu Supervised by Ulrike Stege In collaboration with Chris Upton, Alex Thomo, and Marina Barsky University of
Algorithm Design and Analysis
 Algorithm Design and Analysis LECTURE 4 Graphs Definitions Traversals Adam Smith 9/8/10 Exercise How can you simulate an array with two unbounded stacks and a small amount of memory? (Hint: think of a
Algorithm Design and Analysis LECTURE 4 Graphs Definitions Traversals Adam Smith 9/8/10 Exercise How can you simulate an array with two unbounded stacks and a small amount of memory? (Hint: think of a
Design and Analysis of Algorithms
 Design and Analysis of Algorithms CSE 5311 Lecture 18 Graph Algorithm Junzhou Huang, Ph.D. Department of Computer Science and Engineering CSE5311 Design and Analysis of Algorithms 1 Graphs Graph G = (V,
Design and Analysis of Algorithms CSE 5311 Lecture 18 Graph Algorithm Junzhou Huang, Ph.D. Department of Computer Science and Engineering CSE5311 Design and Analysis of Algorithms 1 Graphs Graph G = (V,
