A Learning Based Algorithm for Automatic Extraction of the Cortical Sulci
|
|
|
- Rhoda Douglas
- 6 years ago
- Views:
Transcription
1 A Learning Based Algorithm for Automatic Extraction of the Cortical Sulci Songfeng Zheng 1, Zhuowen Tu 2, Alan L. Yuille 1, Allan L. Reiss 3, Rebecca A. Dutton 2, Agatha D. Lee 2, Albert M. Galaburda 4, Paul M. Thompson 2, Ivo Dinov 1, 2, Arthur W. Toga 2 1 Department of Statistics, UCLA, Los Angeles, CA, USA {sfzheng, ztu, yuille}@stat.ucla.edu 2 Laboratory of Neuro Imaging, UCLA Medical School, Los Angeles, CA, USA 3 School of Medicine, Stanford University, Stanford, CA, USA 4 School of Medical, Harvard University, Cambridge, MA, USA Abstract. This paper presents a learning based method for automatic extraction of the major cortical sulci from MRI volumes or extracted surfaces. Instead of using a few pre-defined rules such as the mean curvature properties, to detect the major sulci, the algorithm learns a discriminative model by selecting and combining features from a large pool of candidates. We used the Probabilistic Boosting Tree algorithm [16] to learn the model, which implicitly discovers and combines rules based on manually annotated sulci traced by neuroanatomists. The algorithm almost has no parameters to tune and is fast because of the adoption of integral volume and 3D Haar filters. For a given approximately registered MRI volume, the algorithm computes the probability of how likely it is that each voxel lies on a major sulcus curve. Dynamic programming is then applied to extract the curve based on the probability map and a shape prior. Because the algorithm can be applied to MRI volumes directly, there is no need to perform preprocessing such as tissue segmentation or mapping to a canonical space. The learning aspect makes the approach flexible and it also works on extracted cortical surfaces. 1 Introduction Cortical sulci are important structures of the brain. Reliably extracting major cortical sulci from MR images helps us to better understand the functionalities of the brain [7], facilitates many neuro studies for comparing different subjects[12], and assists other brain mapping tasks such as registration [5]. Fig. 1 shows several major cortical sulci: the Central Sulcus, the Postcentral Sulcus, the Superior Temporal Sulcus, the Intraparietal Sulcus, the Middle Frontal Sulcus and the Intraparietal Sulcus. Cortical sulci lie on the valleys of cortical folds, and can be characterized by mathematical measures such as the mean curvatures [6, 1]. However, major cortical sulci have complicated geometric and photometric patterns and one often needs to use many protocols and high-level knowledge [9] to guide the manually labelling process. In general, automatic extraction of cortical sulci is a challenging task [12] due to their large intra-class variation and inter-class similarity across different subjects.
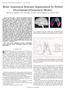
2 2 Most existing approaches for automatic cortical sulci detection work on the extracted cortical surfaces, which require a pre-processing stage to segment the tissue. Tao et al. [14] used global and local shape priors of sulcus curves on a canonical unit sphere to guide the extraction of the curves, but the method involved mapping the cortical surface to the unit sphere. Khaneja [6] used a dynamic programming approach to find the curves by minimizing an energy function, but this algorithm was not fully automatic since it required the starting and ending points of the sulci to be specified by hand. In [11], the patterns of different local folds are learned and they form a random graph using neural networks. The work by Rettmann [10] extracted sulcal regions using a watershed transformation method applied to cortical surfaces. Middle Frontal Sulcus Precentral Sulcus Central Sulcus Postcentra l Sulcus Superior Temporal Sulcus Main Body (a) (b) Fig. 1. Examples of cortical sulci: (a) shows an MRI volume overlayed with several major cortical sulci, such as the Central Sulcus and the Postcentral Sulcus; (b) illustrates a corresponding extracted cortical surface with the same set of manually labelled sulcus curves. Our method is different from the above methods and it can be directly applied to either MRI volumes or extracted cortical surfaces. Instead of using a number of pre-defined rules/features, we learn/compute the likelihood of each voxel being on a sulcus curve based on a sub-volume ( ) centered at this voxel. The probabilistic boosting tree (PBT) algorithm [16] is employed to select and combine hundreds of features from a large set of candidates to make an overall decision using a hierarchical structure. The candidate pool consists of around 5,000 features at three scales including intensity, gradients, curvatures, shape indices, locations, and 3D Haar filters etc.. A data-dependent prior is used to put geometric constraints on the curve. Similar to [6], we use a dynamic programming approach to combine the likelihood and shape prior. The algorithm is fully automatic, very general, and has almost no parameter to tune for different major sulcus curves. Moreover, it can be applied in other curve/object detection tasks in medical imaging. We have a dataset of 40 volumes with several major cortical sulci manually labelled. We split them randomly into 15 training volumes and 25 testing volumes. Training is done both on the MRI volume and extracted cortical surfaces, and we compare the results on the testing images. 2 Problem Formulation In this section, we give the problem formulation for 3D curve detection. For the rest of this paper, we use V to represent a 3D MRI volume. When a surface Intraparietal Sulcus
3 3 has been extracted, the algorithm works almost the same except for some minor modifications in computing gradients and curvature features. For an input volume, V, the task of cortical sulcus curve detection is to extract the 3D curve C: C = {r i, i = 0,, L} where L is the length of the curve, and r i is the coordinates of the i th point on the curve. We define variable B as the background, i.e. the part of the volume not on the curve C. Clearly, C B = and Λ = C B is the domain of the whole volume. We define a label y for each voxel r, so that y = +1 if the voxel is on the curve C and y = 1 if it is on the background B. We then define a discriminative model ˆp(C V) to be of form: log ˆp(C V) = E 1 (C) + E 2 (C) + E 3 where E 3 is a constant which does not depend on C (corresponding to the normalization constant for the distribution ˆp(C V)). From the Bayesian point of view, the optimal curve C is the one maximizes ˆp(C V). The term E 1 is given by: E 1 (C) = r B = r Λ log p(v(r), y = 1 V(N(r)/r)) r C log p(v(r), y = 1 V(N(r)/r)) r C log p(v(r), y = +1 V(N(r)/r)) log p(y = +1 V(N(r))) p(y = 1 V(N(r))). where N(r) is the sub-volume centered at voxel r; V( ) is the intensity value(s) at the given voxel(s); N(r)/r includes all the voxels in the sub-volume except r; p(v(r), y V(N(r)/r)) is a conditional joint probability like pseudo-likelihood [2] in spirit. The first term in the above equation does not depend on C, therefore can be ignored. The probabilities p(y = +1 V(N(r))) and p(y = 1 V(N(r))) will be learned by PBT from manually labelled data. The term E 2 is defined as: L 1 E 2 (C) = αl + β V(r i+1 ) V(r i ) i=0 where α and β are positive parameters to balance the importance of the corresponding terms. The motivation of introducing the first term is the observation that the sulcus curves are not very smooth, while the first term favors long curves, therefore, it discourages a curve being too straight. The second term of E 2 accumulates data dependent cues, i.e., we prefer that the intensity along the detected curve does not change too much. In summary, maximizing the probability ˆp(C V) is equivalent to minimizing the energy function: E(C) = r C log p(y = +1 V(N(r))) p(y = 1 V(N(r))) + E 2(C) (1)
4 4 Here, the models capturing the appearances of foreground and background are combined in the discriminative probability ratio. Note that p(y = +1 V(N(r))) is the posterior probability of a voxel r belonging to the foreground (sulcus curve) given the sub-volume centered at r. The second column in Fig. 3 shows such probability maps. The optimal curve C is the one that minimizes the above energy function E(C). We use dynamic programming (DP) to minimize E(C) given by equation (1). DP is guaranteed to find the global minimum of E(C), but requires starting and ending points (we refer to both as end points). We propose an adaptive method for selecting the end points, instead of choosing the end points by hand [6]. Our method uses the training data to measure the mean and covariance of the positions of the end points and then constrains the end points to lie within boxes centered on the means and with sides equal to twice the variance. We further localize the end points by requiring that p(y = +1 V(N(r)) > T where T is a learned threshold. Of these remaining points, we select the one that has the smallest value of p(y = +1 V(N(r)). 3 The Discriminative Models Now the task is to learn and compute the discriminative probability p(y V(N(r))) for each voxel r. We adopt a new learning framework, probabilistic boosting tree [16], to learn complex discriminative models based on boosting algorithms [4]. For learning, we design a pool of around 5,000 features at three scales, including intensity, gradients, curvatures, shape indices, locations, and 3D Haar filters etc.. We used integral volume to compute the response of 3D Haar filters. At each y1 z1 0 0 location (x 1, y 1, z 1 ), the integral volume is computed as x 1 V (x, y, z)dxdydz. 0 Thus the computational cost for computing the response of 3D Haar filters is reduced significantly since every time we only need to sum up the values at the corners of the 3D Haar filter in the integral volume. boosting boosting Fig. 2. Illustration of a boosting tree on a training volume: The left branch shows the probability of each voxel is not on the sulcus curve and the right branch shows the probability that each voxel is on the sulcus curve. The tree is trained recursively: At the current node, the empirical distribution ˆq(y) of the data is calculated, if the node is not pure (i.e., ˆq(y) is not close to 0 or 1), a strong classifier is trained on the data. Each sample is then passed to the left and right subtree, weighted by q( 1 V(N(r))) and q(+1 V(N(r))),
5 5 respectively, where q(+1 V(N(r))) is the probability that V(N(r)) is a positive sample, according to the strong classifier. Thus, the strong classifier at each node is not used to return the class of the sample but rather to assign the sample to the left or right subtree. Training proceeds recursively(see [16] for information about the basic algorithm). Fig. 2 shows the first two levels of a tree. PBT does training and testing in a divide-and-conquer manner and outputs the overall discriminative probability as: p(y V(N(r))) = l 1 p(y l 1, V(N(r)))q(l 1 V(N(r))) = l 1,l 2 p(y l 2, l 1, V(N(r)))q(l 2 l 1, V(N(r)))q(l 1 V(N(r))) = l 1,,l n p(y l n,, l 1, V(N(r))),, q(l 2 l 1, V(N(r)))q(l 1 V(N(r))) where l i {+1, 1} is augmented hidden variable as shown in Fig. 2, indicating which branch is for this node: l i = 1 and l i = +1 point to the left and right branch, respectively. q(l i l i 1,, l 1, V(N(r))) is the discriminative probability computed according to the boosting strong classifier learned at each node, and ˆq(y l n,, l 1, V(N(r))) is the empirical distribution at the leaf node. 4 Experiments We have tested the proposed approach for extracting the Central and the Middle Frontal Sulci. In both cases, we used ground truth estimates of the Sulci location to train and test the algorithm. We also compared the performance for extracting the Central Sulcus when the cortical surface is available. The MR images, cortical surfaces and manually labelled landmark data are exactly the same as those in [13]. Learning the PBT took approximately eight hours (it is a function of the size of the training dataset); computing the posterior probability by PBT took about one minute per image; running the dynamic programming took around twenty seconds per image. The computer used has 2.4 GHz CPU and 1.0GB memory. Standard code optimization techniques can reduce these times significantly. We have a dataset of 40 volumes of which we use 15 for training and the remained 25 for testing. In these volumes, the length of the sulci varies from voxels. The PBT is learned on the 15 training volumes and the majority of features it selected to use were 3D Haar filters, though some curvature features were also selected. Fig. 3 shows the results on some of the testing volumes for detecting the Central Sulcus and the Middle Frontal Sulcus on MRI volumes. The first column shows the input with manual labels superimposed and the second column shows the posterior probability of the sulci output by the PBT. Note that the probability maps have large responses around the correct position, but the maps are blurred, sometimes disconnected, and therefore are not sufficient to localize the sulcus directly. We then applied dynamic programming to minimize the full energy function, equation (1). This resulted in a very clear estimate of the sulcus location, which are compared to the ground truth in Fig. 3.
6 6 Our method can also be applied to cases where only the cortical surface is available. This requires removing certain features from those considered in the case of volume data (i.e. we keep the 3D Haar filters, but remove intensity). We repeated the experiment on the Central Sulcus with the same training and testing dataset as before, and the result is also shown in Fig. 3. Fig. 3. Detection results on some of the testing images for the Central Sulcus on MRI, the Middle Frontal Sulcus on MRI, and the Central Sulcus on the cortical surfaces. The ground truth (left), the PBT posterior p(y = +1 V(N(r))) (middle), and the result of DP superimposed on the image (right). To quantitatively evaluate the performance of our approach, we measured the distances between the estimated curves and the ground truth. We used the
7 7 following distance measures: H wor (C, G) = max c C min g G c g, H av(c, G) = 1 C c C min g G c g. (2) Here C represents the curve found by DP and G is the ground truth. H av (C, G) gives the average distances from curve C to their closest points on curve G. By contrast, H wor (C, G) measures the worst case fit from curve C to curve G. For symmetry, we also consider H av (G, C). Our evaluation results, as shown in table 1, show very good performance. In the table, < > denotes the average over the dataset, for example, < H av (C, G) >= (1/N) N i=1 H av(c i, G i ), where N is the number of examples in the dataset. All the values of H av are in the range of 3-5 voxels for both training and testing datasets. Observe that although the worst case measures are relatively big, the average distance is small, which suggests that some points occasionally have big offset while the overall curves still can be detected fairly accurately. The biggest errors occurred at the starting and ending points of the sulci. The testing errors are only slightly bigger than the training errors, which implies that our algorithm generalizes well. Dataset < H av(c, G) > < H av(g, C) > < H wor(c, G) > Testing (Central on volume) Central (Central on volume) Testing (Middle Frontal on volume) Training (Middle Frontal on volume) Testing (Central on surface) Training (Central on surface) Table 1. Error measures on 15 training and 25 testing images for the extracting of the Central Sulcus on volume, the Middle Frontal Sulcus on volume, and the Central Sulcus on surfaces. See text for notations. The unit of distance is the voxel. 5 Conclusions and Future Work We presented a method for extracting cortical sulci, which uses a discriminative model including a discriminative term and a data dependent prior. The discriminative term was learned in a supervised way by the probabilistic boosting tree algorithm, and dynamic programming was used to find the optimal solution by minimizing the discriminative energy function. A procedure was given for adaptively selecting the end points for dynamic programming. The method was applied to extracting the Central Sulcus and the Middle Frontal Sulcus from raw MRI volume data. In both cases, the performance was very good when evaluated on both the training and testing data. The method was also applied to extracting the Central Sulcus when the cortical surface has already been extracted and the results were similar to those obtained directly from raw volume data. The biggest errors occurred at the end points of the sulci and might be due to ambiguity in the precise positions of these points (i.e. errors arising in the ground truth), or the procedure for estimating the starting and ending points. This is a topic for future research.
8 8 In this work, we designed a feature pool consists of about 5,000 candidates, most of which are 3D Haar filters. If effective features are used, we expect that the algorithm will achieve similar performance with fewer features. Our next step is to explore more sophisticated features for this particular medical image task. Acknowledgement This work was funded by the National Institutes of Health through the NIH Roadmap for Medical Research, Grant U54 RR entitled Center for Computational Biology (CCB). References 1. A. Bartesaghi and G. Sapiro, A System for the Generation of Curves on 3D Brain Images, Human Brain Map. 14: J. Besag, Efficiency of pseudo-likelihood estimation for simple Gaussian fields, Biometrika 64: , A. Caunce and C.J. Taylor, Building 3D Sulcal Models Using Local geometry, Medical Image Analysis, , Y. Freund and R. Schapire, A Decision-Theoretic Generalization of On-line Learning and An Application to Boosting, J. of Computer and Sys. Sci., P. Hellier and C. Barillot, Coupling Dense and Landmark-Based Approaches for Nonrigid Registration, IEEE Trans. Med. Imaging, vol. 22, no. 2, Feb N. Khaneja, M.I. Miller, and U. Grenander, Dynamic Programming Generation of Curves on Brain Surfaces, PAMI, vo.20, no. 11, G. LeGoualher, E. Procyk, D.L. Collins, R. Venugopal, C. Barillot, A.C. Evans, Automated extraction and variability analysis of sulcal neuroanatomy, IEEE Trans. Med. Imaging vo.18, no. 3, March G. Lohmann, Extracting Line Representations of Sulcal and Gyral Patterns in MR Images of the Hunam Brain, IEEE Trans. on Medical Imag., vol.17, no.6, Dec M. Ono, S. Kubik, and S.D. Abernathey, Atlas of the Cerebral Sulci, New York:Thieme Medical, M.E. Rettmann, X. Han, C. Xu, J.L. Prince, Automated sulcal segmentation using watersheds on the cortical surface, Neuroimage, 15(2):329-44, Feb D. Riviere, J.F. Mangin, D. Papadopoulos-Orfanos, J.M. Martinez, V. Frouin, and J. Regis, Automatic Recognition of Cortical Sulci of the Human Brain Using A Congregation of Neural Networks, Medical Image Analysis, 77-92, P.M. Thompson et al. Mapping Cortical Change in Alzheimer s Disease, Brain Development, and Schizophrenia, NeuroImage, 23 Suppl 1:S2-18, September P.M. Thompson, A.D. Lee, R.A. Dutton, J.A. Geaga,K.M. Hayashi KM, U. Bellugi, A.M. Galaburda, J.R. Korenberg, D.L. Mills, A.W. Toga, A.L. Reiss, Abnormal Cortical Complexity and Thickness Profiles Mapped in Williams Syndrome, J. of Neuroscience, 25(18):, April X. Tao, J.L. Prince, and C. Davatzikos, Using a Statistical Shape Model to Extract Sulcal Curves on the Outer Cortex of the Human Brain, IEEE Tran. on Medical Imag., vol.21, no. 5, May J.P. Thirion and A. Gourdon, Computing the Differential Characteristics of Isointensity Surfaces, Computer Vision and Image Understanding, vol. 61, no. 2, Z. Tu, Probabilistic Boosting-Tree: Learning Discriminative Models for Classification, Recognition, and Clustering, Proc. of ICCV, 2005.
CORTICAL sulci are important structures of the brain
 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 26, NO. 4, APRIL 2007 541 Automated Extraction of the Cortical Sulci Based on a Supervised Learning Approach Zhuowen Tu, Songfeng Zheng, Alan L. Yuille, Allan
IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 26, NO. 4, APRIL 2007 541 Automated Extraction of the Cortical Sulci Based on a Supervised Learning Approach Zhuowen Tu, Songfeng Zheng, Alan L. Yuille, Allan
Joint Sulci Detection Using Graphical Models and Boosted Priors
 Joint Sulci Detection Using Graphical Models and Boosted Priors Yonggang Shi 1,ZhuowenTu 1, Allan L. Reiss 2, Rebecca A. Dutton 1, Agatha D. Lee 1, Albert M. Galaburda 3,IvoDinov 1, Paul M. Thompson 1,
Joint Sulci Detection Using Graphical Models and Boosted Priors Yonggang Shi 1,ZhuowenTu 1, Allan L. Reiss 2, Rebecca A. Dutton 1, Agatha D. Lee 1, Albert M. Galaburda 3,IvoDinov 1, Paul M. Thompson 1,
Towards Learning-Based Holistic Brain Image Segmentation
 Towards Learning-Based Holistic Brain Image Segmentation Zhuowen Tu Lab of Neuro Imaging University of California, Los Angeles in collaboration with (A. Toga, P. Thompson et al.) Supported by (NIH CCB
Towards Learning-Based Holistic Brain Image Segmentation Zhuowen Tu Lab of Neuro Imaging University of California, Los Angeles in collaboration with (A. Toga, P. Thompson et al.) Supported by (NIH CCB
Simultaneous Cortical Surface Labeling and Sulcal Curve Extraction
 Simultaneous Cortical Surface Labeling and Sulcal Curve Extraction Zhen Yang a, Aaron Carass a, Chen Chen a, Jerry L Prince a,b a Electrical and Computer Engineering, b Biomedical Engineering, Johns Hopkins
Simultaneous Cortical Surface Labeling and Sulcal Curve Extraction Zhen Yang a, Aaron Carass a, Chen Chen a, Jerry L Prince a,b a Electrical and Computer Engineering, b Biomedical Engineering, Johns Hopkins
Towards Whole Brain Segmentation by a Hybrid Model
 Towards Whole Brain Segmentation by a Hybrid Model Zhuowen Tu and Arthur W. Toga Lab of Neuro Imaging, School of Medicine University of California, Los Angeles, USA Abstract. Segmenting cortical and sub-cortical
Towards Whole Brain Segmentation by a Hybrid Model Zhuowen Tu and Arthur W. Toga Lab of Neuro Imaging, School of Medicine University of California, Los Angeles, USA Abstract. Segmenting cortical and sub-cortical
SEGMENTING subcortical structures from 3-D brain images
 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 27, NO. 4, APRIL 2008 495 Brain Anatomical Structure Segmentation by Hybrid Discriminative/Generative Models Zhuowen Tu, Katherine L. Narr, Piotr Dollár, Ivo
IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 27, NO. 4, APRIL 2008 495 Brain Anatomical Structure Segmentation by Hybrid Discriminative/Generative Models Zhuowen Tu, Katherine L. Narr, Piotr Dollár, Ivo
IMA Preprint Series # 2078
 A GEOMETRIC METHOD FOR AUTOMATIC EXTRACTION OF SULCAL FUNDI By C.-Y. Kao M. Hofer G. Sapiro J. Stern and D. Rottenberg IMA Preprint Series # 2078 ( November 2005 ) INSTITUTE FOR MATHEMATICS AND ITS APPLICATIONS
A GEOMETRIC METHOD FOR AUTOMATIC EXTRACTION OF SULCAL FUNDI By C.-Y. Kao M. Hofer G. Sapiro J. Stern and D. Rottenberg IMA Preprint Series # 2078 ( November 2005 ) INSTITUTE FOR MATHEMATICS AND ITS APPLICATIONS
Labeling the Brain Surface Using a Deformable Multiresolution Mesh
 Labeling the Brain Surface Using a Deformable Multiresolution Mesh Sylvain Jaume 1, Benoît Macq 2, and Simon K. Warfield 3 1 Multiresolution Modeling Group, California Institute of Technology, 256-80,
Labeling the Brain Surface Using a Deformable Multiresolution Mesh Sylvain Jaume 1, Benoît Macq 2, and Simon K. Warfield 3 1 Multiresolution Modeling Group, California Institute of Technology, 256-80,
Using a Statistical Shape Model to Extract Sulcal Curves on the Outer Cortex of the Human Brain
 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 21, NO. 5, MAY 2002 513 Using a Statistical Shape Model to Extract Sulcal Curves on the Outer Cortex of the Human Brain Xiaodong Tao, Jerry L. Prince, and Christos
IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 21, NO. 5, MAY 2002 513 Using a Statistical Shape Model to Extract Sulcal Curves on the Outer Cortex of the Human Brain Xiaodong Tao, Jerry L. Prince, and Christos
Retrospective Evaluation of Inter-subject Brain Registration
 Retrospective Evaluation of Inter-subject Brain Registration P. Hellier 1, C. Barillot 1, I. Corouge 1,B.Gibaud 2, G. Le Goualher 2,3, L. Collins 3, A. Evans 3, G. Malandain 4, and N. Ayache 4 1 Projet
Retrospective Evaluation of Inter-subject Brain Registration P. Hellier 1, C. Barillot 1, I. Corouge 1,B.Gibaud 2, G. Le Goualher 2,3, L. Collins 3, A. Evans 3, G. Malandain 4, and N. Ayache 4 1 Projet
Retrospective evaluation of inter-subject brain registration
 Retrospective evaluation of inter-subject brain registration P. Hellier 1, C. Barillot 1, I. Corouge 1,B.Gibaud 2, G. Le Goualher 2,3, L. Collins 3, A. Evans 3, G. Malandain 4, N. Ayache 4 1 Projet Vista,
Retrospective evaluation of inter-subject brain registration P. Hellier 1, C. Barillot 1, I. Corouge 1,B.Gibaud 2, G. Le Goualher 2,3, L. Collins 3, A. Evans 3, G. Malandain 4, N. Ayache 4 1 Projet Vista,
Multivariate Tensor-based Brain Anatomical Surface Morphometry via Holomorphic One-Forms
 Multivariate Tensor-based Brain Anatomical Surface Morphometry via Holomorphic One-Forms Yalin Wang 1,2, Tony F. Chan 2, Arthur W. Toga 1, and Paul M. Thompson 1 1 Lab. of Neuro Imaging, UCLA School of
Multivariate Tensor-based Brain Anatomical Surface Morphometry via Holomorphic One-Forms Yalin Wang 1,2, Tony F. Chan 2, Arthur W. Toga 1, and Paul M. Thompson 1 1 Lab. of Neuro Imaging, UCLA School of
Hybrid Spline-based Multimodal Registration using a Local Measure for Mutual Information
 Hybrid Spline-based Multimodal Registration using a Local Measure for Mutual Information Andreas Biesdorf 1, Stefan Wörz 1, Hans-Jürgen Kaiser 2, Karl Rohr 1 1 University of Heidelberg, BIOQUANT, IPMB,
Hybrid Spline-based Multimodal Registration using a Local Measure for Mutual Information Andreas Biesdorf 1, Stefan Wörz 1, Hans-Jürgen Kaiser 2, Karl Rohr 1 1 University of Heidelberg, BIOQUANT, IPMB,
Brain Warping Via Landmark Points and Curves with a Level Set Representation
 Brain Warping Via Landmark Points and Curves with a Level Set Representation Andrew Y. Wang, Alex D. Leow, 2 Hillary D. Protas, Arthur W. Toga, Paul M. Thompson UCLA Laboratory of Neuro Imaging, Los Angeles,
Brain Warping Via Landmark Points and Curves with a Level Set Representation Andrew Y. Wang, Alex D. Leow, 2 Hillary D. Protas, Arthur W. Toga, Paul M. Thompson UCLA Laboratory of Neuro Imaging, Los Angeles,
A Fast and Accurate Method for Automated Cortical Surface Registration and Labeling
 A Fast and Accurate Method for Automated Cortical Surface Registration and Labeling Anand A Joshi 1, David W Shattuck 2 and Richard M Leahy 1 1 Signal and Image Processing Institute, University of Southern
A Fast and Accurate Method for Automated Cortical Surface Registration and Labeling Anand A Joshi 1, David W Shattuck 2 and Richard M Leahy 1 1 Signal and Image Processing Institute, University of Southern
Detecting Object Boundaries Using Low-, Mid-, and High-level Information
 Detecting Object Boundaries Using Low-, Mid-, and High-level Information Songfeng Zheng Dept. of Statistics, UCLA sfzheng@stat.ucla.edu Zhuowen Tu Lab of Neuro Imaging, UCLA zhuowen.tu@loni.ucla.edu Alan
Detecting Object Boundaries Using Low-, Mid-, and High-level Information Songfeng Zheng Dept. of Statistics, UCLA sfzheng@stat.ucla.edu Zhuowen Tu Lab of Neuro Imaging, UCLA zhuowen.tu@loni.ucla.edu Alan
Brain Surface Conformal Parameterization with Algebraic Functions
 Brain Surface Conformal Parameterization with Algebraic Functions Yalin Wang 1,2, Xianfeng Gu 3, Tony F. Chan 1, Paul M. Thompson 2, and Shing-Tung Yau 4 1 Mathematics Department, UCLA, Los Angeles, CA
Brain Surface Conformal Parameterization with Algebraic Functions Yalin Wang 1,2, Xianfeng Gu 3, Tony F. Chan 1, Paul M. Thompson 2, and Shing-Tung Yau 4 1 Mathematics Department, UCLA, Los Angeles, CA
Automatic Subcortical Segmentation Using a Contextual Model
 Automatic Subcortical Segmentation Using a Contextual Model Jonathan H. Morra 1, Zhuowen Tu 1, Liana G. Apostolova 1,2, Amity E. Green 1,2, Arthur W. Toga 1, and Paul M. Thompson 1 1 Laboratory of Neuro
Automatic Subcortical Segmentation Using a Contextual Model Jonathan H. Morra 1, Zhuowen Tu 1, Liana G. Apostolova 1,2, Amity E. Green 1,2, Arthur W. Toga 1, and Paul M. Thompson 1 1 Laboratory of Neuro
Morphological Analysis of Brain Structures Using Spatial Normalization
 Morphological Analysis of Brain Structures Using Spatial Normalization C. Davatzikos 1, M. Vaillant 1, S. Resnick 2, J.L. Prince 3;1, S. Letovsky 1, and R.N. Bryan 1 1 Department of Radiology, Johns Hopkins
Morphological Analysis of Brain Structures Using Spatial Normalization C. Davatzikos 1, M. Vaillant 1, S. Resnick 2, J.L. Prince 3;1, S. Letovsky 1, and R.N. Bryan 1 1 Department of Radiology, Johns Hopkins
Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge
 Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge Christian Wasserthal 1, Karin Engel 1, Karsten Rink 1 und André Brechmann
Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge Christian Wasserthal 1, Karin Engel 1, Karsten Rink 1 und André Brechmann
Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm
 Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm Lin Chen and Gudrun Wagenknecht Central Institute for Electronics, Research Center Jülich, Jülich, Germany Email: l.chen@fz-juelich.de
Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm Lin Chen and Gudrun Wagenknecht Central Institute for Electronics, Research Center Jülich, Jülich, Germany Email: l.chen@fz-juelich.de
Automatic Caudate Segmentation by Hybrid Generative/Discriminative Models
 Automatic Caudate Segmentation by Hybrid Generative/Discriminative Models Zhuowen Tu, Mubeena Mirza, Ivo Dinov, and Arthur W. Toga Lab of Neuro Imaging, School of Medicine University of California, Los
Automatic Caudate Segmentation by Hybrid Generative/Discriminative Models Zhuowen Tu, Mubeena Mirza, Ivo Dinov, and Arthur W. Toga Lab of Neuro Imaging, School of Medicine University of California, Los
Segmentation of a Human Brain Cortical Surface Mesh Using Watersheds
 Segmentation of a Human Brain Cortical Surface Mesh Using Watersheds 1.0 Abstract A 3D cortical surface mesh is a convoluted structure. The purpose of segmenting such a mesh is to impose a higher level
Segmentation of a Human Brain Cortical Surface Mesh Using Watersheds 1.0 Abstract A 3D cortical surface mesh is a convoluted structure. The purpose of segmenting such a mesh is to impose a higher level
Multiple Cortical Surface Correspondence Using Pairwise Shape Similarity
 Multiple Cortical Surface Correspondence Using Pairwise Shape Similarity Pahal Dalal 1, Feng Shi 2, Dinggang Shen 2, and Song Wang 1 1 Department of Computer Science and Engineering, University of South
Multiple Cortical Surface Correspondence Using Pairwise Shape Similarity Pahal Dalal 1, Feng Shi 2, Dinggang Shen 2, and Song Wang 1 1 Department of Computer Science and Engineering, University of South
Automatic Optimization of Segmentation Algorithms Through Simultaneous Truth and Performance Level Estimation (STAPLE)
 Automatic Optimization of Segmentation Algorithms Through Simultaneous Truth and Performance Level Estimation (STAPLE) Mahnaz Maddah, Kelly H. Zou, William M. Wells, Ron Kikinis, and Simon K. Warfield
Automatic Optimization of Segmentation Algorithms Through Simultaneous Truth and Performance Level Estimation (STAPLE) Mahnaz Maddah, Kelly H. Zou, William M. Wells, Ron Kikinis, and Simon K. Warfield
A Genus Bound for Digital Image Boundaries
 A Genus Bound for Digital Image Boundaries Lowell Abrams and Donniell E. Fishkind March 9, 2005 Abstract Shattuck and Leahy [4] conjectured and Abrams, Fishkind, and Priebe [1],[2] proved that the boundary
A Genus Bound for Digital Image Boundaries Lowell Abrams and Donniell E. Fishkind March 9, 2005 Abstract Shattuck and Leahy [4] conjectured and Abrams, Fishkind, and Priebe [1],[2] proved that the boundary
Watersheds on the Cortical Surface for Automated Sulcal Segmentation
 Watersheds on the Cortical Surface for Automated Sulcal Segmentation Maryam E. Rettmann,XiaoHan y, and Jerry L. Prince y Department of Biomedical Engineering y Department of Electrical and Computer Engineering
Watersheds on the Cortical Surface for Automated Sulcal Segmentation Maryam E. Rettmann,XiaoHan y, and Jerry L. Prince y Department of Biomedical Engineering y Department of Electrical and Computer Engineering
Estimating Human Pose in Images. Navraj Singh December 11, 2009
 Estimating Human Pose in Images Navraj Singh December 11, 2009 Introduction This project attempts to improve the performance of an existing method of estimating the pose of humans in still images. Tasks
Estimating Human Pose in Images Navraj Singh December 11, 2009 Introduction This project attempts to improve the performance of an existing method of estimating the pose of humans in still images. Tasks
Wavelet-Based Representation of Biological Shapes
 Wavelet-Based Representation of Biological Shapes Bin Dong 1,YuMao 2, Ivo D. Dinov 3,ZhuowenTu 3, Yonggang Shi 3, Yalin Wang 3, and Arthur W. Toga 3 1 Department of Mathematics, University of California,
Wavelet-Based Representation of Biological Shapes Bin Dong 1,YuMao 2, Ivo D. Dinov 3,ZhuowenTu 3, Yonggang Shi 3, Yalin Wang 3, and Arthur W. Toga 3 1 Department of Mathematics, University of California,
ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION
 ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION Abstract: MIP Project Report Spring 2013 Gaurav Mittal 201232644 This is a detailed report about the course project, which was to implement
ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION Abstract: MIP Project Report Spring 2013 Gaurav Mittal 201232644 This is a detailed report about the course project, which was to implement
Surface-based Analysis: Inter-subject Registration and Smoothing
 Surface-based Analysis: Inter-subject Registration and Smoothing Outline Exploratory Spatial Analysis Coordinate Systems 3D (Volumetric) 2D (Surface-based) Inter-subject registration Volume-based Surface-based
Surface-based Analysis: Inter-subject Registration and Smoothing Outline Exploratory Spatial Analysis Coordinate Systems 3D (Volumetric) 2D (Surface-based) Inter-subject registration Volume-based Surface-based
Wavelet-Based Representation of Biological Shapes
 Wavelet-Based Representation of Biological Shapes Bin Dong*, Yu Mao*, Ivo D. Dinov**, Zhuowen Tu**, Yonggang Shi**, Yalin Wang**, Arthur W. Toga** * Department of Mathematics, University of California,
Wavelet-Based Representation of Biological Shapes Bin Dong*, Yu Mao*, Ivo D. Dinov**, Zhuowen Tu**, Yonggang Shi**, Yalin Wang**, Arthur W. Toga** * Department of Mathematics, University of California,
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit
 Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
Correspondence Detection Using Wavelet-Based Attribute Vectors
 Correspondence Detection Using Wavelet-Based Attribute Vectors Zhong Xue, Dinggang Shen, and Christos Davatzikos Section of Biomedical Image Analysis, Department of Radiology University of Pennsylvania,
Correspondence Detection Using Wavelet-Based Attribute Vectors Zhong Xue, Dinggang Shen, and Christos Davatzikos Section of Biomedical Image Analysis, Department of Radiology University of Pennsylvania,
A Nonparametric Bayesian Approach to Detecting Spatial Activation Patterns in fmri Data
 A Nonparametric Bayesian Approach to Detecting Spatial Activation Patterns in fmri Data Seyoung Kim, Padhraic Smyth, and Hal Stern Bren School of Information and Computer Sciences University of California,
A Nonparametric Bayesian Approach to Detecting Spatial Activation Patterns in fmri Data Seyoung Kim, Padhraic Smyth, and Hal Stern Bren School of Information and Computer Sciences University of California,
Subvoxel Segmentation and Representation of Brain Cortex Using Fuzzy Clustering and Gradient Vector Diffusion
 Subvoxel Segmentation and Representation of Brain Cortex Using Fuzzy Clustering and Gradient Vector Diffusion Ming-Ching Chang Xiaodong Tao GE Global Research Center {changm, taox} @ research.ge.com SPIE
Subvoxel Segmentation and Representation of Brain Cortex Using Fuzzy Clustering and Gradient Vector Diffusion Ming-Ching Chang Xiaodong Tao GE Global Research Center {changm, taox} @ research.ge.com SPIE
Learning-based Neuroimage Registration
 Learning-based Neuroimage Registration Leonid Teverovskiy and Yanxi Liu 1 October 2004 CMU-CALD-04-108, CMU-RI-TR-04-59 School of Computer Science Carnegie Mellon University Pittsburgh, PA 15213 Abstract
Learning-based Neuroimage Registration Leonid Teverovskiy and Yanxi Liu 1 October 2004 CMU-CALD-04-108, CMU-RI-TR-04-59 School of Computer Science Carnegie Mellon University Pittsburgh, PA 15213 Abstract
Locating Salient Object Features
 Locating Salient Object Features K.N.Walker, T.F.Cootes and C.J.Taylor Dept. Medical Biophysics, Manchester University, UK knw@sv1.smb.man.ac.uk Abstract We present a method for locating salient object
Locating Salient Object Features K.N.Walker, T.F.Cootes and C.J.Taylor Dept. Medical Biophysics, Manchester University, UK knw@sv1.smb.man.ac.uk Abstract We present a method for locating salient object
Semantic Context Forests for Learning- Based Knee Cartilage Segmentation in 3D MR Images
 Semantic Context Forests for Learning- Based Knee Cartilage Segmentation in 3D MR Images MICCAI 2013: Workshop on Medical Computer Vision Authors: Quan Wang, Dijia Wu, Le Lu, Meizhu Liu, Kim L. Boyer,
Semantic Context Forests for Learning- Based Knee Cartilage Segmentation in 3D MR Images MICCAI 2013: Workshop on Medical Computer Vision Authors: Quan Wang, Dijia Wu, Le Lu, Meizhu Liu, Kim L. Boyer,
Cost-alleviative Learning for Deep Convolutional Neural Network-based Facial Part Labeling
 [DOI: 10.2197/ipsjtcva.7.99] Express Paper Cost-alleviative Learning for Deep Convolutional Neural Network-based Facial Part Labeling Takayoshi Yamashita 1,a) Takaya Nakamura 1 Hiroshi Fukui 1,b) Yuji
[DOI: 10.2197/ipsjtcva.7.99] Express Paper Cost-alleviative Learning for Deep Convolutional Neural Network-based Facial Part Labeling Takayoshi Yamashita 1,a) Takaya Nakamura 1 Hiroshi Fukui 1,b) Yuji
Non-rigid Image Registration using Electric Current Flow
 Non-rigid Image Registration using Electric Current Flow Shu Liao, Max W. K. Law and Albert C. S. Chung Lo Kwee-Seong Medical Image Analysis Laboratory, Department of Computer Science and Engineering,
Non-rigid Image Registration using Electric Current Flow Shu Liao, Max W. K. Law and Albert C. S. Chung Lo Kwee-Seong Medical Image Analysis Laboratory, Department of Computer Science and Engineering,
Statistical Study on Cortical Sulci of Human Brains
 Statistical Study on Cortical Sulci of Human Brains Xiaodong Tao 1,3,XiaoHan 1, Maryam E. Rettmann 2, Jerry L. Prince 1,2,3,and Christos Davatzikos 3 1 Electrical and Computer Engineering Johns Hopkins
Statistical Study on Cortical Sulci of Human Brains Xiaodong Tao 1,3,XiaoHan 1, Maryam E. Rettmann 2, Jerry L. Prince 1,2,3,and Christos Davatzikos 3 1 Electrical and Computer Engineering Johns Hopkins
A Hierarchical Compositional System for Rapid Object Detection
 A Hierarchical Compositional System for Rapid Object Detection Long Zhu and Alan Yuille Department of Statistics University of California at Los Angeles Los Angeles, CA 90095 {lzhu,yuille}@stat.ucla.edu
A Hierarchical Compositional System for Rapid Object Detection Long Zhu and Alan Yuille Department of Statistics University of California at Los Angeles Los Angeles, CA 90095 {lzhu,yuille}@stat.ucla.edu
Automatic Generation of Training Data for Brain Tissue Classification from MRI
 Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. COCOSCO, Alex P. ZIJDENBOS, and Alan C. EVANS http://www.bic.mni.mcgill.ca/users/crisco/ McConnell Brain Imaging
Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. COCOSCO, Alex P. ZIJDENBOS, and Alan C. EVANS http://www.bic.mni.mcgill.ca/users/crisco/ McConnell Brain Imaging
Learning Based Coarse-to-fine Image Registration
 Learning Based Coarse-to-fine Image Registration Jiayan Jiang LONI, UCLA jet.jiang@loni.ucla.edu Songfeng Zheng Dept. of Statistics, UCLA sfzheng@stat.ucla.edu Arthur W. Toga LONI, UCLA toga@loni.ucla.edu
Learning Based Coarse-to-fine Image Registration Jiayan Jiang LONI, UCLA jet.jiang@loni.ucla.edu Songfeng Zheng Dept. of Statistics, UCLA sfzheng@stat.ucla.edu Arthur W. Toga LONI, UCLA toga@loni.ucla.edu
Fast Interactive Region of Interest Selection for Volume Visualization
 Fast Interactive Region of Interest Selection for Volume Visualization Dominik Sibbing and Leif Kobbelt Lehrstuhl für Informatik 8, RWTH Aachen, 20 Aachen Email: {sibbing,kobbelt}@informatik.rwth-aachen.de
Fast Interactive Region of Interest Selection for Volume Visualization Dominik Sibbing and Leif Kobbelt Lehrstuhl für Informatik 8, RWTH Aachen, 20 Aachen Email: {sibbing,kobbelt}@informatik.rwth-aachen.de
The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy
 The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy Sokratis K. Makrogiannis, PhD From post-doctoral research at SBIA lab, Department of Radiology,
The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy Sokratis K. Makrogiannis, PhD From post-doctoral research at SBIA lab, Department of Radiology,
MR IMAGE SEGMENTATION
 MR IMAGE SEGMENTATION Prepared by : Monil Shah What is Segmentation? Partitioning a region or regions of interest in images such that each region corresponds to one or more anatomic structures Classification
MR IMAGE SEGMENTATION Prepared by : Monil Shah What is Segmentation? Partitioning a region or regions of interest in images such that each region corresponds to one or more anatomic structures Classification
Open Topology: A Toolkit for Brain Isosurface Correction
 Open Topology: A Toolkit for Brain Isosurface Correction Sylvain Jaume 1, Patrice Rondao 2, and Benoît Macq 2 1 National Institute of Research in Computer Science and Control, INRIA, France, sylvain@mit.edu,
Open Topology: A Toolkit for Brain Isosurface Correction Sylvain Jaume 1, Patrice Rondao 2, and Benoît Macq 2 1 National Institute of Research in Computer Science and Control, INRIA, France, sylvain@mit.edu,
STATISTICAL ATLAS-BASED SUB-VOXEL SEGMENTATION OF 3D BRAIN MRI
 STATISTICA ATAS-BASED SUB-VOXE SEGMENTATION OF 3D BRAIN MRI Marcel Bosc 1,2, Fabrice Heitz 1, Jean-Paul Armspach 2 (1) SIIT UMR-7005 CNRS / Strasbourg I University, 67400 Illkirch, France (2) IPB UMR-7004
STATISTICA ATAS-BASED SUB-VOXE SEGMENTATION OF 3D BRAIN MRI Marcel Bosc 1,2, Fabrice Heitz 1, Jean-Paul Armspach 2 (1) SIIT UMR-7005 CNRS / Strasbourg I University, 67400 Illkirch, France (2) IPB UMR-7004
Medical Image Segmentation
 Medical Image Segmentation Xin Yang, HUST *Collaborated with UCLA Medical School and UCSB Segmentation to Contouring ROI Aorta & Kidney 3D Brain MR Image 3D Abdominal CT Image Liver & Spleen Caudate Nucleus
Medical Image Segmentation Xin Yang, HUST *Collaborated with UCLA Medical School and UCSB Segmentation to Contouring ROI Aorta & Kidney 3D Brain MR Image 3D Abdominal CT Image Liver & Spleen Caudate Nucleus
Comparison of Vessel Segmentations Using STAPLE
 Comparison of Vessel Segmentations Using STAPLE Julien Jomier, Vincent LeDigarcher, and Stephen R. Aylward Computer-Aided Diagnosis and Display Lab, The University of North Carolina at Chapel Hill, Department
Comparison of Vessel Segmentations Using STAPLE Julien Jomier, Vincent LeDigarcher, and Stephen R. Aylward Computer-Aided Diagnosis and Display Lab, The University of North Carolina at Chapel Hill, Department
CS 534: Computer Vision Segmentation and Perceptual Grouping
 CS 534: Computer Vision Segmentation and Perceptual Grouping Ahmed Elgammal Dept of Computer Science CS 534 Segmentation - 1 Outlines Mid-level vision What is segmentation Perceptual Grouping Segmentation
CS 534: Computer Vision Segmentation and Perceptual Grouping Ahmed Elgammal Dept of Computer Science CS 534 Segmentation - 1 Outlines Mid-level vision What is segmentation Perceptual Grouping Segmentation
A Novel Method for Enhanced Needle Localization Using Ultrasound-Guidance
 A Novel Method for Enhanced Needle Localization Using Ultrasound-Guidance Bin Dong 1,EricSavitsky 2, and Stanley Osher 3 1 Department of Mathematics, University of California, San Diego, 9500 Gilman Drive,
A Novel Method for Enhanced Needle Localization Using Ultrasound-Guidance Bin Dong 1,EricSavitsky 2, and Stanley Osher 3 1 Department of Mathematics, University of California, San Diego, 9500 Gilman Drive,
COMBINATIONOF BRAIN CONFORMAL MAPPING AND LANDMARKS: A VARIATIONALAPPROACH
 COMBINATIONOF BRAIN CONFORMAL MAPPING AND LANDMARKS: A VARIATIONALAPPROACH Yalin Wang Mathematics Department, UCLA email: ylwang@math.ucla.edu Lok Ming Lui Mathematics Department, UCLA email: malmlui@math.ucla.edu
COMBINATIONOF BRAIN CONFORMAL MAPPING AND LANDMARKS: A VARIATIONALAPPROACH Yalin Wang Mathematics Department, UCLA email: ylwang@math.ucla.edu Lok Ming Lui Mathematics Department, UCLA email: malmlui@math.ucla.edu
Supervised Learning for Image Segmentation
 Supervised Learning for Image Segmentation Raphael Meier 06.10.2016 Raphael Meier MIA 2016 06.10.2016 1 / 52 References A. Ng, Machine Learning lecture, Stanford University. A. Criminisi, J. Shotton, E.
Supervised Learning for Image Segmentation Raphael Meier 06.10.2016 Raphael Meier MIA 2016 06.10.2016 1 / 52 References A. Ng, Machine Learning lecture, Stanford University. A. Criminisi, J. Shotton, E.
ABSTRACT 1. INTRODUCTION 2. METHODS
 Finding Seeds for Segmentation Using Statistical Fusion Fangxu Xing *a, Andrew J. Asman b, Jerry L. Prince a,c, Bennett A. Landman b,c,d a Department of Electrical and Computer Engineering, Johns Hopkins
Finding Seeds for Segmentation Using Statistical Fusion Fangxu Xing *a, Andrew J. Asman b, Jerry L. Prince a,c, Bennett A. Landman b,c,d a Department of Electrical and Computer Engineering, Johns Hopkins
Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques
 Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques Sea Chen Department of Biomedical Engineering Advisors: Dr. Charles A. Bouman and Dr. Mark J. Lowe S. Chen Final Exam October
Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques Sea Chen Department of Biomedical Engineering Advisors: Dr. Charles A. Bouman and Dr. Mark J. Lowe S. Chen Final Exam October
The organization of the human cerebral cortex estimated by intrinsic functional connectivity
 1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
SURFACE RECONSTRUCTION OF EX-VIVO HUMAN V1 THROUGH IDENTIFICATION OF THE STRIA OF GENNARI USING MRI AT 7T
 SURFACE RECONSTRUCTION OF EX-VIVO HUMAN V1 THROUGH IDENTIFICATION OF THE STRIA OF GENNARI USING MRI AT 7T Oliver P. Hinds 1, Jonathan R. Polimeni 2, Megan L. Blackwell 3, Christopher J. Wiggins 3, Graham
SURFACE RECONSTRUCTION OF EX-VIVO HUMAN V1 THROUGH IDENTIFICATION OF THE STRIA OF GENNARI USING MRI AT 7T Oliver P. Hinds 1, Jonathan R. Polimeni 2, Megan L. Blackwell 3, Christopher J. Wiggins 3, Graham
Retrospective Evaluation of Intersubject Brain Registration
 1120 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 22, NO. 9, SEPTEMBER 2003 Retrospective Evaluation of Intersubject Brain Registration P. Hellier*, C. Barillot, I. Corouge, B. Gibaud, G. Le Goualher, D.
1120 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 22, NO. 9, SEPTEMBER 2003 Retrospective Evaluation of Intersubject Brain Registration P. Hellier*, C. Barillot, I. Corouge, B. Gibaud, G. Le Goualher, D.
CAP 6412 Advanced Computer Vision
 CAP 6412 Advanced Computer Vision http://www.cs.ucf.edu/~bgong/cap6412.html Boqing Gong April 21st, 2016 Today Administrivia Free parameters in an approach, model, or algorithm? Egocentric videos by Aisha
CAP 6412 Advanced Computer Vision http://www.cs.ucf.edu/~bgong/cap6412.html Boqing Gong April 21st, 2016 Today Administrivia Free parameters in an approach, model, or algorithm? Egocentric videos by Aisha
MARS: Multiple Atlases Robust Segmentation
 Software Release (1.0.1) Last updated: April 30, 2014. MARS: Multiple Atlases Robust Segmentation Guorong Wu, Minjeong Kim, Gerard Sanroma, and Dinggang Shen {grwu, mjkim, gerard_sanroma, dgshen}@med.unc.edu
Software Release (1.0.1) Last updated: April 30, 2014. MARS: Multiple Atlases Robust Segmentation Guorong Wu, Minjeong Kim, Gerard Sanroma, and Dinggang Shen {grwu, mjkim, gerard_sanroma, dgshen}@med.unc.edu
doi: /
 Yiting Xie ; Anthony P. Reeves; Single 3D cell segmentation from optical CT microscope images. Proc. SPIE 934, Medical Imaging 214: Image Processing, 9343B (March 21, 214); doi:1.1117/12.243852. (214)
Yiting Xie ; Anthony P. Reeves; Single 3D cell segmentation from optical CT microscope images. Proc. SPIE 934, Medical Imaging 214: Image Processing, 9343B (March 21, 214); doi:1.1117/12.243852. (214)
Machine Learning for Medical Image Analysis. A. Criminisi
 Machine Learning for Medical Image Analysis A. Criminisi Overview Introduction to machine learning Decision forests Applications in medical image analysis Anatomy localization in CT Scans Spine Detection
Machine Learning for Medical Image Analysis A. Criminisi Overview Introduction to machine learning Decision forests Applications in medical image analysis Anatomy localization in CT Scans Spine Detection
A Design Toolbox to Generate Complex Phantoms for the Evaluation of Medical Image Processing Algorithms
 A Design Toolbox to Generate Complex Phantoms for the Evaluation of Medical Image Processing Algorithms Omar Hamo, Georg Nelles, Gudrun Wagenknecht Central Institute for Electronics, Research Center Juelich,
A Design Toolbox to Generate Complex Phantoms for the Evaluation of Medical Image Processing Algorithms Omar Hamo, Georg Nelles, Gudrun Wagenknecht Central Institute for Electronics, Research Center Juelich,
Mapping the Cerebral Sulci: Application to Morphological Analysis of the Cortex and to Non-rigid Registration
 Mapping the Cerebral Sulci: Application to Morphological Analysis of the Cortex and to Non-rigid Registration Marc Vaillant and Christos Davatzikos Neuroimaging Laboratory, Department of Radiology, Johns
Mapping the Cerebral Sulci: Application to Morphological Analysis of the Cortex and to Non-rigid Registration Marc Vaillant and Christos Davatzikos Neuroimaging Laboratory, Department of Radiology, Johns
Towards the completion of assignment 1
 Towards the completion of assignment 1 What to do for calibration What to do for point matching What to do for tracking What to do for GUI COMPSCI 773 Feature Point Detection Why study feature point detection?
Towards the completion of assignment 1 What to do for calibration What to do for point matching What to do for tracking What to do for GUI COMPSCI 773 Feature Point Detection Why study feature point detection?
NIH Public Access Author Manuscript Proc SPIE. Author manuscript; available in PMC 2013 December 30.
 NIH Public Access Author Manuscript Published in final edited form as: Proc SPIE. 2013 March 12; 8669:. doi:10.1117/12.2006682. Longitudinal Intensity Normalization of Magnetic Resonance Images using Patches
NIH Public Access Author Manuscript Published in final edited form as: Proc SPIE. 2013 March 12; 8669:. doi:10.1117/12.2006682. Longitudinal Intensity Normalization of Magnetic Resonance Images using Patches
Image Segmentation and Registration
 Image Segmentation and Registration Dr. Christine Tanner (tanner@vision.ee.ethz.ch) Computer Vision Laboratory, ETH Zürich Dr. Verena Kaynig, Machine Learning Laboratory, ETH Zürich Outline Segmentation
Image Segmentation and Registration Dr. Christine Tanner (tanner@vision.ee.ethz.ch) Computer Vision Laboratory, ETH Zürich Dr. Verena Kaynig, Machine Learning Laboratory, ETH Zürich Outline Segmentation
Comparison Study of Clinical 3D MRI Brain Segmentation Evaluation
 Comparison Study of Clinical 3D MRI Brain Segmentation Evaluation Ting Song 1, Elsa D. Angelini 2, Brett D. Mensh 3, Andrew Laine 1 1 Heffner Biomedical Imaging Laboratory Department of Biomedical Engineering,
Comparison Study of Clinical 3D MRI Brain Segmentation Evaluation Ting Song 1, Elsa D. Angelini 2, Brett D. Mensh 3, Andrew Laine 1 1 Heffner Biomedical Imaging Laboratory Department of Biomedical Engineering,
MARS: Multiple Atlases Robust Segmentation
 Software Release (1.0.1) Last updated: April 30, 2014. MARS: Multiple Atlases Robust Segmentation Guorong Wu, Minjeong Kim, Gerard Sanroma, and Dinggang Shen {grwu, mjkim, gerard_sanroma, dgshen}@med.unc.edu
Software Release (1.0.1) Last updated: April 30, 2014. MARS: Multiple Atlases Robust Segmentation Guorong Wu, Minjeong Kim, Gerard Sanroma, and Dinggang Shen {grwu, mjkim, gerard_sanroma, dgshen}@med.unc.edu
Integrated Approaches to Non-Rigid Registration in Medical Images
 Work. on Appl. of Comp. Vision, pg 102-108. 1 Integrated Approaches to Non-Rigid Registration in Medical Images Yongmei Wang and Lawrence H. Staib + Departments of Electrical Engineering and Diagnostic
Work. on Appl. of Comp. Vision, pg 102-108. 1 Integrated Approaches to Non-Rigid Registration in Medical Images Yongmei Wang and Lawrence H. Staib + Departments of Electrical Engineering and Diagnostic
Interactive Differential Segmentation of the Prostate using Graph-Cuts with a Feature Detector-based Boundary Term
 MOSCHIDIS, GRAHAM: GRAPH-CUTS WITH FEATURE DETECTORS 1 Interactive Differential Segmentation of the Prostate using Graph-Cuts with a Feature Detector-based Boundary Term Emmanouil Moschidis emmanouil.moschidis@postgrad.manchester.ac.uk
MOSCHIDIS, GRAHAM: GRAPH-CUTS WITH FEATURE DETECTORS 1 Interactive Differential Segmentation of the Prostate using Graph-Cuts with a Feature Detector-based Boundary Term Emmanouil Moschidis emmanouil.moschidis@postgrad.manchester.ac.uk
Categorization by Learning and Combining Object Parts
 Categorization by Learning and Combining Object Parts Bernd Heisele yz Thomas Serre y Massimiliano Pontil x Thomas Vetter Λ Tomaso Poggio y y Center for Biological and Computational Learning, M.I.T., Cambridge,
Categorization by Learning and Combining Object Parts Bernd Heisele yz Thomas Serre y Massimiliano Pontil x Thomas Vetter Λ Tomaso Poggio y y Center for Biological and Computational Learning, M.I.T., Cambridge,
A MORPHOLOGY-BASED FILTER STRUCTURE FOR EDGE-ENHANCING SMOOTHING
 Proceedings of the 1994 IEEE International Conference on Image Processing (ICIP-94), pp. 530-534. (Austin, Texas, 13-16 November 1994.) A MORPHOLOGY-BASED FILTER STRUCTURE FOR EDGE-ENHANCING SMOOTHING
Proceedings of the 1994 IEEE International Conference on Image Processing (ICIP-94), pp. 530-534. (Austin, Texas, 13-16 November 1994.) A MORPHOLOGY-BASED FILTER STRUCTURE FOR EDGE-ENHANCING SMOOTHING
Shape-based Diffeomorphic Registration on Hippocampal Surfaces Using Beltrami Holomorphic Flow
 Shape-based Diffeomorphic Registration on Hippocampal Surfaces Using Beltrami Holomorphic Flow Abstract. Finding meaningful 1-1 correspondences between hippocampal (HP) surfaces is an important but difficult
Shape-based Diffeomorphic Registration on Hippocampal Surfaces Using Beltrami Holomorphic Flow Abstract. Finding meaningful 1-1 correspondences between hippocampal (HP) surfaces is an important but difficult
A Multiple-Layer Flexible Mesh Template Matching Method for Nonrigid Registration between a Pelvis Model and CT Images
 A Multiple-Layer Flexible Mesh Template Matching Method for Nonrigid Registration between a Pelvis Model and CT Images Jianhua Yao 1, Russell Taylor 2 1. Diagnostic Radiology Department, Clinical Center,
A Multiple-Layer Flexible Mesh Template Matching Method for Nonrigid Registration between a Pelvis Model and CT Images Jianhua Yao 1, Russell Taylor 2 1. Diagnostic Radiology Department, Clinical Center,
Automatic Registration-Based Segmentation for Neonatal Brains Using ANTs and Atropos
 Automatic Registration-Based Segmentation for Neonatal Brains Using ANTs and Atropos Jue Wu and Brian Avants Penn Image Computing and Science Lab, University of Pennsylvania, Philadelphia, USA Abstract.
Automatic Registration-Based Segmentation for Neonatal Brains Using ANTs and Atropos Jue Wu and Brian Avants Penn Image Computing and Science Lab, University of Pennsylvania, Philadelphia, USA Abstract.
NIH Public Access Author Manuscript Conf Proc IEEE Eng Med Biol Soc. Author manuscript; available in PMC 2009 December 11.
 NIH Public Access Author Manuscript Published in final edited form as: Conf Proc IEEE Eng Med Biol Soc. 2004 ; 2: 1521 1524. doi:10.1109/iembs.2004.1403466. An Elliptic PDE Approach for Shape Characterization
NIH Public Access Author Manuscript Published in final edited form as: Conf Proc IEEE Eng Med Biol Soc. 2004 ; 2: 1521 1524. doi:10.1109/iembs.2004.1403466. An Elliptic PDE Approach for Shape Characterization
ON-LINE BOOSTING BASED REAL-TIME TRACKING WITH EFFICIENT HOG
 ON-LINE BOOSTING BASED REAL-TIME TRACKING WITH EFFICIENT HOG Shuifa Sun **, Qing Guo *, Fangmin Dong, Bangjun Lei Institute of Intelligent Vision and Image Information, China Three Gorges University, Yichang
ON-LINE BOOSTING BASED REAL-TIME TRACKING WITH EFFICIENT HOG Shuifa Sun **, Qing Guo *, Fangmin Dong, Bangjun Lei Institute of Intelligent Vision and Image Information, China Three Gorges University, Yichang
Smart point landmark distribution for thin-plate splines
 Smart point landmark distribution for thin-plate splines John Lewis a, Hea-Juen Hwang a, Ulrich Neumann a, and Reyes Enciso b a Integrated Media Systems Center, University of Southern California, 3740
Smart point landmark distribution for thin-plate splines John Lewis a, Hea-Juen Hwang a, Ulrich Neumann a, and Reyes Enciso b a Integrated Media Systems Center, University of Southern California, 3740
Brain Anatomical Structure Segmentation by Hybrid Discriminative/Generative Models
 Brain Anatomical Structure Segmentation by Hybrid Discriminative/Generative Models Zhuowen Tu, Katherine L. Narr, Piotr Dollár, Ivo Dinov, Paul M. Thompson, and Arthur W. Toga Abstract In this paper, a
Brain Anatomical Structure Segmentation by Hybrid Discriminative/Generative Models Zhuowen Tu, Katherine L. Narr, Piotr Dollár, Ivo Dinov, Paul M. Thompson, and Arthur W. Toga Abstract In this paper, a
GENERAL AUTOMATED FLAW DETECTION SCHEME FOR NDE X-RAY IMAGES
 GENERAL AUTOMATED FLAW DETECTION SCHEME FOR NDE X-RAY IMAGES Karl W. Ulmer and John P. Basart Center for Nondestructive Evaluation Department of Electrical and Computer Engineering Iowa State University
GENERAL AUTOMATED FLAW DETECTION SCHEME FOR NDE X-RAY IMAGES Karl W. Ulmer and John P. Basart Center for Nondestructive Evaluation Department of Electrical and Computer Engineering Iowa State University
FACE RECOGNITION USING SUPPORT VECTOR MACHINES
 FACE RECOGNITION USING SUPPORT VECTOR MACHINES Ashwin Swaminathan ashwins@umd.edu ENEE633: Statistical and Neural Pattern Recognition Instructor : Prof. Rama Chellappa Project 2, Part (b) 1. INTRODUCTION
FACE RECOGNITION USING SUPPORT VECTOR MACHINES Ashwin Swaminathan ashwins@umd.edu ENEE633: Statistical and Neural Pattern Recognition Instructor : Prof. Rama Chellappa Project 2, Part (b) 1. INTRODUCTION
NeuroImage 49 (2010) Contents lists available at ScienceDirect. NeuroImage. journal homepage:
 NeuroImage 49 (2010) 2479 2493 Contents lists available at ScienceDirect NeuroImage journal homepage: www.elsevier.com/locate/ynimg Comparison of landmark-based and automatic methods for cortical surface
NeuroImage 49 (2010) 2479 2493 Contents lists available at ScienceDirect NeuroImage journal homepage: www.elsevier.com/locate/ynimg Comparison of landmark-based and automatic methods for cortical surface
IEEE TRANSACTIONS ON IMAGE PROCESSING, VOL. 26, NO. 10, OCTOBER
 IEEE TRANSACTIONS ON IMAGE PROCESSING, VOL. 26, NO. 10, OCTOBER 2017 4753 Detecting Anatomical Landmarks From Limited Medical Imaging Data Using Two-Stage Task-Oriented Deep Neural Networks Jun Zhang,
IEEE TRANSACTIONS ON IMAGE PROCESSING, VOL. 26, NO. 10, OCTOBER 2017 4753 Detecting Anatomical Landmarks From Limited Medical Imaging Data Using Two-Stage Task-Oriented Deep Neural Networks Jun Zhang,
Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging
 1 CS 9 Final Project Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging Feiyu Chen Department of Electrical Engineering ABSTRACT Subject motion is a significant
1 CS 9 Final Project Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging Feiyu Chen Department of Electrical Engineering ABSTRACT Subject motion is a significant
Skull Segmentation of MR images based on texture features for attenuation correction in PET/MR
 Skull Segmentation of MR images based on texture features for attenuation correction in PET/MR CHAIBI HASSEN, NOURINE RACHID ITIO Laboratory, Oran University Algeriachaibih@yahoo.fr, nourine@yahoo.com
Skull Segmentation of MR images based on texture features for attenuation correction in PET/MR CHAIBI HASSEN, NOURINE RACHID ITIO Laboratory, Oran University Algeriachaibih@yahoo.fr, nourine@yahoo.com
Comparison of Vessel Segmentations using STAPLE
 Comparison of Vessel Segmentations using STAPLE Julien Jomier, Vincent LeDigarcher, and Stephen R. Aylward Computer-Aided Diagnosis and Display Lab The University of North Carolina at Chapel Hill, Department
Comparison of Vessel Segmentations using STAPLE Julien Jomier, Vincent LeDigarcher, and Stephen R. Aylward Computer-Aided Diagnosis and Display Lab The University of North Carolina at Chapel Hill, Department
Segmenting Glioma in Multi-Modal Images using a Generative Model for Brain Lesion Segmentation
 Segmenting Glioma in Multi-Modal Images using a Generative Model for Brain Lesion Segmentation Bjoern H. Menze 1,2, Koen Van Leemput 3, Danial Lashkari 4 Marc-André Weber 5, Nicholas Ayache 2, and Polina
Segmenting Glioma in Multi-Modal Images using a Generative Model for Brain Lesion Segmentation Bjoern H. Menze 1,2, Koen Van Leemput 3, Danial Lashkari 4 Marc-André Weber 5, Nicholas Ayache 2, and Polina
Manifold Learning-based Data Sampling for Model Training
 Manifold Learning-based Data Sampling for Model Training Shuqing Chen 1, Sabrina Dorn 2, Michael Lell 3, Marc Kachelrieß 2,Andreas Maier 1 1 Pattern Recognition Lab, FAU Erlangen-Nürnberg 2 German Cancer
Manifold Learning-based Data Sampling for Model Training Shuqing Chen 1, Sabrina Dorn 2, Michael Lell 3, Marc Kachelrieß 2,Andreas Maier 1 1 Pattern Recognition Lab, FAU Erlangen-Nürnberg 2 German Cancer
CHAPTER 6 MODIFIED FUZZY TECHNIQUES BASED IMAGE SEGMENTATION
 CHAPTER 6 MODIFIED FUZZY TECHNIQUES BASED IMAGE SEGMENTATION 6.1 INTRODUCTION Fuzzy logic based computational techniques are becoming increasingly important in the medical image analysis arena. The significant
CHAPTER 6 MODIFIED FUZZY TECHNIQUES BASED IMAGE SEGMENTATION 6.1 INTRODUCTION Fuzzy logic based computational techniques are becoming increasingly important in the medical image analysis arena. The significant
An ITK Filter for Bayesian Segmentation: itkbayesianclassifierimagefilter
 An ITK Filter for Bayesian Segmentation: itkbayesianclassifierimagefilter John Melonakos 1, Karthik Krishnan 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332, USA {jmelonak,
An ITK Filter for Bayesian Segmentation: itkbayesianclassifierimagefilter John Melonakos 1, Karthik Krishnan 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332, USA {jmelonak,
RESEARCH STATEMENT Ronald Lok Ming Lui
 Ronald Lok Ming Lui 1/11 RESEARCH STATEMENT Ronald Lok Ming Lui (malmlui@math.harvard.edu) The main focus of my research has been on problems in computational conformal geometry and human brain mapping.
Ronald Lok Ming Lui 1/11 RESEARCH STATEMENT Ronald Lok Ming Lui (malmlui@math.harvard.edu) The main focus of my research has been on problems in computational conformal geometry and human brain mapping.
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit
 Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
Non-rigid Image Registration
 Overview Non-rigid Image Registration Introduction to image registration - he goal of image registration - Motivation for medical image registration - Classification of image registration - Nonrigid registration
Overview Non-rigid Image Registration Introduction to image registration - he goal of image registration - Motivation for medical image registration - Classification of image registration - Nonrigid registration
Discrete Optimization of Ray Potentials for Semantic 3D Reconstruction
 Discrete Optimization of Ray Potentials for Semantic 3D Reconstruction Marc Pollefeys Joined work with Nikolay Savinov, Christian Haene, Lubor Ladicky 2 Comparison to Volumetric Fusion Higher-order ray
Discrete Optimization of Ray Potentials for Semantic 3D Reconstruction Marc Pollefeys Joined work with Nikolay Savinov, Christian Haene, Lubor Ladicky 2 Comparison to Volumetric Fusion Higher-order ray
Whole Body MRI Intensity Standardization
 Whole Body MRI Intensity Standardization Florian Jäger 1, László Nyúl 1, Bernd Frericks 2, Frank Wacker 2 and Joachim Hornegger 1 1 Institute of Pattern Recognition, University of Erlangen, {jaeger,nyul,hornegger}@informatik.uni-erlangen.de
Whole Body MRI Intensity Standardization Florian Jäger 1, László Nyúl 1, Bernd Frericks 2, Frank Wacker 2 and Joachim Hornegger 1 1 Institute of Pattern Recognition, University of Erlangen, {jaeger,nyul,hornegger}@informatik.uni-erlangen.de
NIH Public Access Author Manuscript Proc IEEE Int Symp Biomed Imaging. Author manuscript; available in PMC 2014 November 15.
 NIH Public Access Author Manuscript Published in final edited form as: Proc IEEE Int Symp Biomed Imaging. 2013 April ; 2013: 748 751. doi:10.1109/isbi.2013.6556583. BRAIN TUMOR SEGMENTATION WITH SYMMETRIC
NIH Public Access Author Manuscript Published in final edited form as: Proc IEEE Int Symp Biomed Imaging. 2013 April ; 2013: 748 751. doi:10.1109/isbi.2013.6556583. BRAIN TUMOR SEGMENTATION WITH SYMMETRIC
