RESEARCH TOPIC IN BIOINFORMANTIC
|
|
|
- Delphia Cook
- 6 years ago
- Views:
Transcription
1 RESEARCH TOPIC IN BIOINFORMANTIC GENOME ASSEMBLY Instructor: Dr. Yufeng Wu Noted by: February 25, 2012 Genome Assembly is a kind of string sequencing problems. As we all know, the human genome is very long. It is about 3Gb for each. Now a days, the second generation of genome assembly technique creates multiple copies of the genomes. Then, shears the genomes into fragments. After that, the short pieces is gained from these fragments as showed in Fig. 1. the researchers try to assemble these huge amount of pieces back to the original genome which process is called genome assembly. Fig. 1: the process to get the reads. Definition # 1 Reads refer to the short pieces obtained from the fragment. Definition # 2 Paired reads refer to wo reads obtained from two end-sides of one fragment. Now the genome assembly problem becomes: Page 1
2 Input: Goal: short sequencing reads or paired reads. reconstruct the reference string from the reads. But to reconstruct the reference sequence is still hard. There are several challenges remaining here. Challenges: 1. The length of reference sequence is long. There are huge amount of short reads. 2. The errors are contained in the reads. 3. Complex structure of the genome repeats. To aim the goal of genome assembly. One idea is constructing a completed weighted directed graph or namely overlap graph. Completed weighted directed graph (overlap graph): The weights of edges denote the length of the over lapping between two reads. The nodes denote the reads. An example showed in Fig. 2 illuminate the overlap graph. Input: Output: the reads: ACG, CGA, CGC,CGT, GAC, GCG, GTA, TCG construct the overlap graph. Fig. 2: the example of overlap graph. Not all the edges are drawn in the graph. Here we just draw some of the edges. To reduce the complexity of the problem. We make some assumptions and criterion here. Page 2
3 Assume: 1. There are no errors in the reads. 2. Ignore the genome repeats. i.e. each read only occur once in the reference sequence. Criterion: Assembly tries to find the parsimony simplest solution. In other words, the assembly tries to find the shortest sequence from the reads. Though the shortest sequence can not be the correct one, it is not far away from the truth. By the assumptions and criterion, we can observe that: What is the reference genome? = the path in the overlap graph. How to enforce that each read only occur once? = find the Hamilton path in the graph. i.e. find the path travelling each node once and only once. How to find the parsimony simplest solution? = Maximize the weight of path. Definition # 3 a Hamilton path is a path in the graph that visit each node exactly once. Thus, the problem approaches to the travelling salesman path (TSP) problem to minimize the negative weight of the OG. Unfortunately, this problem is NP-hard problem. The researchers try to get the approximate solutions by applying the greedy algorithms. One of the algorithms is the heuristic overlapping-layoutconsensus (OLC). 1. Overlap: construct the overlap graph. 2. Layout: find the maximum weighted path in the overlap graph by the greedy algorithm. 3. Consensus: obtain the consensus sequence. For example, Tab. 1 depicts how the consensus works. Definition # 4 Consensus sequence refers to the most common nucleotide or amino acid at a particular position after multiple sequences are aligned. Then, the researchers find another efficient way to assemble the genomes later. The method is called Eulerian path approach. Similarly, to implement the Eulerian path approach, we shall make some assumptions first. Page 3
4 ACGAC T ATGGAG Aligned sequences ACCTCC ACGGT T ACCGGC T G C A Consensus ACGGC T Tab. 1: consensus sequence from five aligned sequences. Assume: 1. All the reads are the same k length. we call k-mers. 2. Each read is distinct. i.e. each read only occur once. 3. read is sheared one nucleobase by onenucleobase. So, the reads start from every position of sequence. Definition # 5 k-mer refers to a specific n-tuple or n-gram of nucleic acid or amino acid sequences. The graph include Eulerian path is called de Bruijn graph. In genome assembly, the de bruijn graph is constructed as following. de Bruijn graph: The nodes denote (k 1) length prefix and suffix of the reads. The edge connect two nodes if they are from the prefix and suffix of the same read. In other words, the edges represent the reads. the graph is un-weighted and directed. Definition # 6 an Eulerian path is a path in the graph which visit every edge exactly once. Theorem # 6.1 a directed graph has the Eulerian path if and only if at most one node has one more in-degree than out-degree and at most one node has one more out-degree than in-degree and the graph is connected. For example, Fig. 3 illuminate the de Bruijn graph for the same reads set as the overlap graph example. Page 4
5 Input: Output: the reads: ACG, CGA, CGC,CGT, GAC, GCG, GTA, TCG construct the de Bruijn graph. Fig. 3: the example of the de Bruijn graph. According to the de Bruijn graph, a possible Eulerian path is TC CG GC CG GA AC CG GT TA Thus, one possible genome sequence is : TCGCGACGTA. The simple method to find the Eulerian path from one graph: 1. Start from one node that still has unused degrees. 2. Pick one edge not using before. Travel to next node along this edge. 3. Repeat Step 2 until close one cycle. 4. Repeat Step 1 through Step 3 to find the new cycle. 5. Combine the cycles by alternate the paths of two cycles that pass the conjoint node. But in the real world, some assumptions are hardly achieved. There are some challenges in applying the Eulerian path to the genome assembly. Challenges: 1. It is hard to get the read that start from every position of the genome sequence. 2. It is still an issue that the errors are contained in the reads. Thus, to get the continues k-mers, the Euler assembler provides an idea that for a given read, breaks it into k-mers. Then, the problem of the genome assembly becomes an Eulerian super-path problem. Page 5
6 Eulerian super-path problem: Given the reads and the k. Trained the paths of k-mers of each read, i, as the sub-path, SP i. Find an Eulerian super-path P s.t. P contain each SP i exactly once. If we relax the condition that pass each sub-path exactly once as that pass each edge in the graph at least once, the problem becomes the Chinese postman problem. Chinese postman problem: Given the directed graph. Find shortest path that visit each edge at least once. The Chinese postman problem can be solved in polynomial time. However, the Chinese postman problem constrained by the sub-path is NP-hard. In Euler assembler, the author provided several method to approach the expected result such as the x, y-detachment and x-cut (Fig. 4). (a) x, y-detachment (b) x-cut Fig. 4: Equivalent transformations: (a)x, y-detachment and (b)x-cut. Definition # 7 The x, y-detachment is a transformation that adds a new edge z = (v in, v out ) and delete the edges x and y from G (Fig. 4a). Page 6
7 Definition # 8 an x-cut is a transformation by simply removing x from all the paths that start from x or end at x without affecting the graph G itself (Fig. 4b). Some paths can be merged due to consistent with each other. Fig. 5 illuminate two possible consistent. P that is consistent only with P x,y1 (Fig. 5a) is resolvable while the P that is consistent both with P x,y1 and P x,y2 (Fig. 5b) is unresolvable. (a) P is consistent only with P x,y1. (b) P is consistent both with P x,y1 and P x,y2. Fig. 5: two possible consistent. The unambiguous paths are constructed as the contigs and the paired reads help us to construct the scaffoldings (the superior level of contigs) by the contigs. Definition # 9 a contig is a set of overlapping DNA segments that together represent a consensus region of DNA. Fig. 6: cycle graph before merging. Fig. 7: de Bruijn graph after merging. But due to the huge amount of the reads the de Bruijn graph is still very complex. For example, consider the cycle string, S = ATCAGATAGGAC. (1) Page 7
8 The k-mers with k = 2 can be formed as a cycle graph (Fig. 6). If we merge the identical nodes, this cycle graph will become the de Bruijn graph (Fig. 7) and the merging will cost the graph complex. Thus, is that a way to reduce the merging? Here we introduce the paired de Bruijn graph. paired de Bruijn graph: Each node denotes (k 1) length prefixes or suffixes of both paired reads. The edge connect two nodes if they are from the prefixes and suffixes of the same paired read. the graph is un-weighted and directed. Also, there are some assumptions as following. Assume: 1. The paired reads have same insert size. 2. d denotes the distance between the starting points of the paired reads. For example, consider the same string, Eq. (1), with d = 4 and k = 2. As showed in Fig. 8, only two nodes can be merged due to the identity. Thus, the paired de Bruijn graph reduces the merging efficiently. Fig. 8: the cycle graph with the paired reads. We can release the first assumption a little bit. If the first parts of two paired nodes are matched and the distance between two second parts of the paired nodes are close, these two nodes can be merged. For example, one paired node is AT/GA and another is AT/GG. The first parts of two nodes are the same as AT. So, if the distance between second pairs of two nodes, GA GG, is small, then these two nodes, AT/GA and AT/GG, can be merged. Page 8
Algorithms for Bioinformatics
 Adapted from slides by Alexandru Tomescu, Leena Salmela and Veli Mäkinen, which are partly from http://bix.ucsd.edu/bioalgorithms/slides.php 582670 Algorithms for Bioinformatics Lecture 3: Graph Algorithms
Adapted from slides by Alexandru Tomescu, Leena Salmela and Veli Mäkinen, which are partly from http://bix.ucsd.edu/bioalgorithms/slides.php 582670 Algorithms for Bioinformatics Lecture 3: Graph Algorithms
Sequence Assembly. BMI/CS 576 Mark Craven Some sequencing successes
 Sequence Assembly BMI/CS 576 www.biostat.wisc.edu/bmi576/ Mark Craven craven@biostat.wisc.edu Some sequencing successes Yersinia pestis Cannabis sativa The sequencing problem We want to determine the identity
Sequence Assembly BMI/CS 576 www.biostat.wisc.edu/bmi576/ Mark Craven craven@biostat.wisc.edu Some sequencing successes Yersinia pestis Cannabis sativa The sequencing problem We want to determine the identity
Genome 373: Genome Assembly. Doug Fowler
 Genome 373: Genome Assembly Doug Fowler What are some of the things we ve seen we can do with HTS data? We ve seen that HTS can enable a wide variety of analyses ranging from ID ing variants to genome-
Genome 373: Genome Assembly Doug Fowler What are some of the things we ve seen we can do with HTS data? We ve seen that HTS can enable a wide variety of analyses ranging from ID ing variants to genome-
DNA Sequencing The Shortest Superstring & Traveling Salesman Problems Sequencing by Hybridization
 Eulerian & Hamiltonian Cycle Problems DNA Sequencing The Shortest Superstring & Traveling Salesman Problems Sequencing by Hybridization The Bridge Obsession Problem Find a tour crossing every bridge just
Eulerian & Hamiltonian Cycle Problems DNA Sequencing The Shortest Superstring & Traveling Salesman Problems Sequencing by Hybridization The Bridge Obsession Problem Find a tour crossing every bridge just
Sequencing. Computational Biology IST Ana Teresa Freitas 2011/2012. (BACs) Whole-genome shotgun sequencing Celera Genomics
 Computational Biology IST Ana Teresa Freitas 2011/2012 Sequencing Clone-by-clone shotgun sequencing Human Genome Project Whole-genome shotgun sequencing Celera Genomics (BACs) 1 Must take the fragments
Computational Biology IST Ana Teresa Freitas 2011/2012 Sequencing Clone-by-clone shotgun sequencing Human Genome Project Whole-genome shotgun sequencing Celera Genomics (BACs) 1 Must take the fragments
DNA Sequencing. Overview
 BINF 3350, Genomics and Bioinformatics DNA Sequencing Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Backgrounds Eulerian Cycles Problem Hamiltonian Cycles
BINF 3350, Genomics and Bioinformatics DNA Sequencing Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Backgrounds Eulerian Cycles Problem Hamiltonian Cycles
Algorithms for Bioinformatics
 Adapted from slides by Alexandru Tomescu, Leena Salmela and Veli Mäkinen, which are partly from http://bix.ucsd.edu/bioalgorithms/slides.php 58670 Algorithms for Bioinformatics Lecture 5: Graph Algorithms
Adapted from slides by Alexandru Tomescu, Leena Salmela and Veli Mäkinen, which are partly from http://bix.ucsd.edu/bioalgorithms/slides.php 58670 Algorithms for Bioinformatics Lecture 5: Graph Algorithms
10/15/2009 Comp 590/Comp Fall
 Lecture 13: Graph Algorithms Study Chapter 8.1 8.8 10/15/2009 Comp 590/Comp 790-90 Fall 2009 1 The Bridge Obsession Problem Find a tour crossing every bridge just once Leonhard Euler, 1735 Bridges of Königsberg
Lecture 13: Graph Algorithms Study Chapter 8.1 8.8 10/15/2009 Comp 590/Comp 790-90 Fall 2009 1 The Bridge Obsession Problem Find a tour crossing every bridge just once Leonhard Euler, 1735 Bridges of Königsberg
Graph Algorithms in Bioinformatics
 Graph Algorithms in Bioinformatics Computational Biology IST Ana Teresa Freitas 2015/2016 Sequencing Clone-by-clone shotgun sequencing Human Genome Project Whole-genome shotgun sequencing Celera Genomics
Graph Algorithms in Bioinformatics Computational Biology IST Ana Teresa Freitas 2015/2016 Sequencing Clone-by-clone shotgun sequencing Human Genome Project Whole-genome shotgun sequencing Celera Genomics
Sequence Assembly Required!
 Sequence Assembly Required! 1 October 3, ISMB 20172007 1 Sequence Assembly Genome Sequenced Fragments (reads) Assembled Contigs Finished Genome 2 Greedy solution is bounded 3 Typical assembly strategy
Sequence Assembly Required! 1 October 3, ISMB 20172007 1 Sequence Assembly Genome Sequenced Fragments (reads) Assembled Contigs Finished Genome 2 Greedy solution is bounded 3 Typical assembly strategy
CSCI2950-C Lecture 4 DNA Sequencing and Fragment Assembly
 CSCI2950-C Lecture 4 DNA Sequencing and Fragment Assembly Ben Raphael Sept. 22, 2009 http://cs.brown.edu/courses/csci2950-c/ l-mer composition Def: Given string s, the Spectrum ( s, l ) is unordered multiset
CSCI2950-C Lecture 4 DNA Sequencing and Fragment Assembly Ben Raphael Sept. 22, 2009 http://cs.brown.edu/courses/csci2950-c/ l-mer composition Def: Given string s, the Spectrum ( s, l ) is unordered multiset
DNA Fragment Assembly
 Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri DNA Fragment Assembly Overlap
Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri DNA Fragment Assembly Overlap
(for more info see:
 Genome assembly (for more info see: http://www.cbcb.umd.edu/research/assembly_primer.shtml) Introduction Sequencing technologies can only "read" short fragments from a genome. Reconstructing the entire
Genome assembly (for more info see: http://www.cbcb.umd.edu/research/assembly_primer.shtml) Introduction Sequencing technologies can only "read" short fragments from a genome. Reconstructing the entire
10/8/13 Comp 555 Fall
 10/8/13 Comp 555 Fall 2013 1 Find a tour crossing every bridge just once Leonhard Euler, 1735 Bridges of Königsberg 10/8/13 Comp 555 Fall 2013 2 Find a cycle that visits every edge exactly once Linear
10/8/13 Comp 555 Fall 2013 1 Find a tour crossing every bridge just once Leonhard Euler, 1735 Bridges of Königsberg 10/8/13 Comp 555 Fall 2013 2 Find a cycle that visits every edge exactly once Linear
Genome Reconstruction: A Puzzle with a Billion Pieces Phillip E. C. Compeau and Pavel A. Pevzner
 Genome Reconstruction: A Puzzle with a Billion Pieces Phillip E. C. Compeau and Pavel A. Pevzner Outline I. Problem II. Two Historical Detours III.Example IV.The Mathematics of DNA Sequencing V.Complications
Genome Reconstruction: A Puzzle with a Billion Pieces Phillip E. C. Compeau and Pavel A. Pevzner Outline I. Problem II. Two Historical Detours III.Example IV.The Mathematics of DNA Sequencing V.Complications
de novo assembly Simon Rasmussen 36626: Next Generation Sequencing analysis DTU Bioinformatics Next Generation Sequencing Analysis
 de novo assembly Simon Rasmussen 36626: Next Generation Sequencing analysis DTU Bioinformatics 27626 - Next Generation Sequencing Analysis Generalized NGS analysis Data size Application Assembly: Compare
de novo assembly Simon Rasmussen 36626: Next Generation Sequencing analysis DTU Bioinformatics 27626 - Next Generation Sequencing Analysis Generalized NGS analysis Data size Application Assembly: Compare
Purpose of sequence assembly
 Sequence Assembly Purpose of sequence assembly Reconstruct long DNA/RNA sequences from short sequence reads Genome sequencing RNA sequencing for gene discovery Amplicon sequencing But not for transcript
Sequence Assembly Purpose of sequence assembly Reconstruct long DNA/RNA sequences from short sequence reads Genome sequencing RNA sequencing for gene discovery Amplicon sequencing But not for transcript
Introduction to Genome Assembly. Tandy Warnow
 Introduction to Genome Assembly Tandy Warnow 2 Shotgun DNA Sequencing DNA target sample SHEAR & SIZE End Reads / Mate Pairs 550bp 10,000bp Not all sequencing technologies produce mate-pairs. Different
Introduction to Genome Assembly Tandy Warnow 2 Shotgun DNA Sequencing DNA target sample SHEAR & SIZE End Reads / Mate Pairs 550bp 10,000bp Not all sequencing technologies produce mate-pairs. Different
Read Mapping. de Novo Assembly. Genomics: Lecture #2 WS 2014/2015
 Mapping de Novo Assembly Institut für Medizinische Genetik und Humangenetik Charité Universitätsmedizin Berlin Genomics: Lecture #2 WS 2014/2015 Today Genome assembly: the basics Hamiltonian and Eulerian
Mapping de Novo Assembly Institut für Medizinische Genetik und Humangenetik Charité Universitätsmedizin Berlin Genomics: Lecture #2 WS 2014/2015 Today Genome assembly: the basics Hamiltonian and Eulerian
BLAST & Genome assembly
 BLAST & Genome assembly Solon P. Pissis Tomáš Flouri Heidelberg Institute for Theoretical Studies May 15, 2014 1 BLAST What is BLAST? The algorithm 2 Genome assembly De novo assembly Mapping assembly 3
BLAST & Genome assembly Solon P. Pissis Tomáš Flouri Heidelberg Institute for Theoretical Studies May 15, 2014 1 BLAST What is BLAST? The algorithm 2 Genome assembly De novo assembly Mapping assembly 3
Genome Sequencing Algorithms
 Genome Sequencing Algorithms Phillip Compaeu and Pavel Pevzner Bioinformatics Algorithms: an Active Learning Approach Leonhard Euler (1707 1783) William Hamilton (1805 1865) Nicolaas Govert de Bruijn (1918
Genome Sequencing Algorithms Phillip Compaeu and Pavel Pevzner Bioinformatics Algorithms: an Active Learning Approach Leonhard Euler (1707 1783) William Hamilton (1805 1865) Nicolaas Govert de Bruijn (1918
BLAST & Genome assembly
 BLAST & Genome assembly Solon P. Pissis Tomáš Flouri Heidelberg Institute for Theoretical Studies November 17, 2012 1 Introduction Introduction 2 BLAST What is BLAST? The algorithm 3 Genome assembly De
BLAST & Genome assembly Solon P. Pissis Tomáš Flouri Heidelberg Institute for Theoretical Studies November 17, 2012 1 Introduction Introduction 2 BLAST What is BLAST? The algorithm 3 Genome assembly De
02-711/ Computational Genomics and Molecular Biology Fall 2016
 Literature assignment 2 Due: Nov. 3 rd, 2016 at 4:00pm Your name: Article: Phillip E C Compeau, Pavel A. Pevzner, Glenn Tesler. How to apply de Bruijn graphs to genome assembly. Nature Biotechnology 29,
Literature assignment 2 Due: Nov. 3 rd, 2016 at 4:00pm Your name: Article: Phillip E C Compeau, Pavel A. Pevzner, Glenn Tesler. How to apply de Bruijn graphs to genome assembly. Nature Biotechnology 29,
Reducing Genome Assembly Complexity with Optical Maps
 Reducing Genome Assembly Complexity with Optical Maps Lee Mendelowitz LMendelo@math.umd.edu Advisor: Dr. Mihai Pop Computer Science Department Center for Bioinformatics and Computational Biology mpop@umiacs.umd.edu
Reducing Genome Assembly Complexity with Optical Maps Lee Mendelowitz LMendelo@math.umd.edu Advisor: Dr. Mihai Pop Computer Science Department Center for Bioinformatics and Computational Biology mpop@umiacs.umd.edu
Genome Reconstruction: A Puzzle with a Billion Pieces. Phillip Compeau Carnegie Mellon University Computational Biology Department
 http://cbd.cmu.edu Genome Reconstruction: A Puzzle with a Billion Pieces Phillip Compeau Carnegie Mellon University Computational Biology Department Eternity II: The Highest-Stakes Puzzle in History Courtesy:
http://cbd.cmu.edu Genome Reconstruction: A Puzzle with a Billion Pieces Phillip Compeau Carnegie Mellon University Computational Biology Department Eternity II: The Highest-Stakes Puzzle in History Courtesy:
Graph Algorithms in Bioinformatics
 Graph Algorithms in Bioinformatics Bioinformatics: Issues and Algorithms CSE 308-408 Fall 2007 Lecture 13 Lopresti Fall 2007 Lecture 13-1 - Outline Introduction to graph theory Eulerian & Hamiltonian Cycle
Graph Algorithms in Bioinformatics Bioinformatics: Issues and Algorithms CSE 308-408 Fall 2007 Lecture 13 Lopresti Fall 2007 Lecture 13-1 - Outline Introduction to graph theory Eulerian & Hamiltonian Cycle
CS 68: BIOINFORMATICS. Prof. Sara Mathieson Swarthmore College Spring 2018
 CS 68: BIOINFORMATICS Prof. Sara Mathieson Swarthmore College Spring 2018 Outline: Jan 31 DBG assembly in practice Velvet assembler Evaluation of assemblies (if time) Start: string alignment Candidate
CS 68: BIOINFORMATICS Prof. Sara Mathieson Swarthmore College Spring 2018 Outline: Jan 31 DBG assembly in practice Velvet assembler Evaluation of assemblies (if time) Start: string alignment Candidate
DNA Fragment Assembly
 SIGCSE 009 Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri DNA Fragment Assembly
SIGCSE 009 Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri DNA Fragment Assembly
1. Sorting (assuming sorting into ascending order) a) BUBBLE SORT
 DECISION 1 Revision Notes 1. Sorting (assuming sorting into ascending order) a) BUBBLE SORT Make sure you show comparisons clearly and label each pass First Pass 8 4 3 6 1 4 8 3 6 1 4 3 8 6 1 4 3 6 8 1
DECISION 1 Revision Notes 1. Sorting (assuming sorting into ascending order) a) BUBBLE SORT Make sure you show comparisons clearly and label each pass First Pass 8 4 3 6 1 4 8 3 6 1 4 3 8 6 1 4 3 6 8 1
Omega: an Overlap-graph de novo Assembler for Metagenomics
 Omega: an Overlap-graph de novo Assembler for Metagenomics B a h l e l H a i d e r, Ta e - H y u k A h n, B r i a n B u s h n e l l, J u a n j u a n C h a i, A l e x C o p e l a n d, C h o n g l e Pa n
Omega: an Overlap-graph de novo Assembler for Metagenomics B a h l e l H a i d e r, Ta e - H y u k A h n, B r i a n B u s h n e l l, J u a n j u a n C h a i, A l e x C o p e l a n d, C h o n g l e Pa n
A THEORETICAL ANALYSIS OF SCALABILITY OF THE PARALLEL GENOME ASSEMBLY ALGORITHMS
 A THEORETICAL ANALYSIS OF SCALABILITY OF THE PARALLEL GENOME ASSEMBLY ALGORITHMS Munib Ahmed, Ishfaq Ahmad Department of Computer Science and Engineering, University of Texas At Arlington, Arlington, Texas
A THEORETICAL ANALYSIS OF SCALABILITY OF THE PARALLEL GENOME ASSEMBLY ALGORITHMS Munib Ahmed, Ishfaq Ahmad Department of Computer Science and Engineering, University of Texas At Arlington, Arlington, Texas
IDBA - A Practical Iterative de Bruijn Graph De Novo Assembler
 IDBA - A Practical Iterative de Bruijn Graph De Novo Assembler Yu Peng, Henry Leung, S.M. Yiu, Francis Y.L. Chin Department of Computer Science, The University of Hong Kong Pokfulam Road, Hong Kong {ypeng,
IDBA - A Practical Iterative de Bruijn Graph De Novo Assembler Yu Peng, Henry Leung, S.M. Yiu, Francis Y.L. Chin Department of Computer Science, The University of Hong Kong Pokfulam Road, Hong Kong {ypeng,
BMI/CS 576 Fall 2015 Midterm Exam
 BMI/CS 576 Fall 2015 Midterm Exam Prof. Colin Dewey Tuesday, October 27th, 2015 11:00am-12:15pm Name: KEY Write your answers on these pages and show your work. You may use the back sides of pages as necessary.
BMI/CS 576 Fall 2015 Midterm Exam Prof. Colin Dewey Tuesday, October 27th, 2015 11:00am-12:15pm Name: KEY Write your answers on these pages and show your work. You may use the back sides of pages as necessary.
Bioinformatics-themed projects in Discrete Mathematics
 Bioinformatics-themed projects in Discrete Mathematics Art Duval University of Texas at El Paso Joint Mathematics Meeting MAA Contributed Paper Session on Discrete Mathematics in the Undergraduate Curriculum
Bioinformatics-themed projects in Discrete Mathematics Art Duval University of Texas at El Paso Joint Mathematics Meeting MAA Contributed Paper Session on Discrete Mathematics in the Undergraduate Curriculum
Chapter 3: Paths and Cycles
 Chapter 3: Paths and Cycles 5 Connectivity 1. Definitions: Walk: finite sequence of edges in which any two consecutive edges are adjacent or identical. (Initial vertex, Final vertex, length) Trail: walk
Chapter 3: Paths and Cycles 5 Connectivity 1. Definitions: Walk: finite sequence of edges in which any two consecutive edges are adjacent or identical. (Initial vertex, Final vertex, length) Trail: walk
CS 173, Lecture B Introduction to Genome Assembly (using Eulerian Graphs) Tandy Warnow
 CS 173, Lecture B Introduction to Genome Assembly (using Eulerian Graphs) Tandy Warnow 2 Shotgun DNA Sequencing DNA target sample SHEAR & SIZE End Reads / Mate Pairs 550bp 10,000bp Not all sequencing technologies
CS 173, Lecture B Introduction to Genome Assembly (using Eulerian Graphs) Tandy Warnow 2 Shotgun DNA Sequencing DNA target sample SHEAR & SIZE End Reads / Mate Pairs 550bp 10,000bp Not all sequencing technologies
I519 Introduction to Bioinformatics, Genome assembly. Yuzhen Ye School of Informatics & Computing, IUB
 I519 Introduction to Bioinformatics, 2014 Genome assembly Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Genome assembly problem Approaches Comparative assembly The string
I519 Introduction to Bioinformatics, 2014 Genome assembly Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Genome assembly problem Approaches Comparative assembly The string
Computational models for bionformatics
 Computational models for bionformatics De-novo assembly and alignment-free measures Michele Schimd Department of Information Engineering July 8th, 2015 Michele Schimd (DEI) PostDoc @ DEI July 8th, 2015
Computational models for bionformatics De-novo assembly and alignment-free measures Michele Schimd Department of Information Engineering July 8th, 2015 Michele Schimd (DEI) PostDoc @ DEI July 8th, 2015
CS681: Advanced Topics in Computational Biology
 CS681: Advanced Topics in Computational Biology Can Alkan EA224 calkan@cs.bilkent.edu.tr Week 7 Lectures 2-3 http://www.cs.bilkent.edu.tr/~calkan/teaching/cs681/ Genome Assembly Test genome Random shearing
CS681: Advanced Topics in Computational Biology Can Alkan EA224 calkan@cs.bilkent.edu.tr Week 7 Lectures 2-3 http://www.cs.bilkent.edu.tr/~calkan/teaching/cs681/ Genome Assembly Test genome Random shearing
Computational Genomics and Molecular Biology, Fall
 Computational Genomics and Molecular Biology, Fall 2015 1 Sequence Alignment Dannie Durand Pairwise Sequence Alignment The goal of pairwise sequence alignment is to establish a correspondence between the
Computational Genomics and Molecular Biology, Fall 2015 1 Sequence Alignment Dannie Durand Pairwise Sequence Alignment The goal of pairwise sequence alignment is to establish a correspondence between the
DNA Fragment Assembly Algorithms: Toward a Solution for Long Repeats
 San Jose State University SJSU ScholarWorks Master's Projects Master's Theses and Graduate Research 2008 DNA Fragment Assembly Algorithms: Toward a Solution for Long Repeats Ching Li San Jose State University
San Jose State University SJSU ScholarWorks Master's Projects Master's Theses and Graduate Research 2008 DNA Fragment Assembly Algorithms: Toward a Solution for Long Repeats Ching Li San Jose State University
Reducing Genome Assembly Complexity with Optical Maps Mid-year Progress Report
 Reducing Genome Assembly Complexity with Optical Maps Mid-year Progress Report Lee Mendelowitz LMendelo@math.umd.edu Advisor: Dr. Mihai Pop Computer Science Department Center for Bioinformatics and Computational
Reducing Genome Assembly Complexity with Optical Maps Mid-year Progress Report Lee Mendelowitz LMendelo@math.umd.edu Advisor: Dr. Mihai Pop Computer Science Department Center for Bioinformatics and Computational
Shortest Path Algorithm
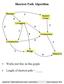 Shortest Path Algorithm C Works just fine on this graph. C Length of shortest path = Copyright 2005 DIMACS BioMath Connect Institute Robert Hochberg Dynamic Programming SP #1 Same Questions, Different
Shortest Path Algorithm C Works just fine on this graph. C Length of shortest path = Copyright 2005 DIMACS BioMath Connect Institute Robert Hochberg Dynamic Programming SP #1 Same Questions, Different
6 ROUTING PROBLEMS VEHICLE ROUTING PROBLEMS. Vehicle Routing Problem, VRP:
 6 ROUTING PROBLEMS VEHICLE ROUTING PROBLEMS Vehicle Routing Problem, VRP: Customers i=1,...,n with demands of a product must be served using a fleet of vehicles for the deliveries. The vehicles, with given
6 ROUTING PROBLEMS VEHICLE ROUTING PROBLEMS Vehicle Routing Problem, VRP: Customers i=1,...,n with demands of a product must be served using a fleet of vehicles for the deliveries. The vehicles, with given
CSCI 1820 Notes. Scribes: tl40. February 26 - March 02, Estimating size of graphs used to build the assembly.
 CSCI 1820 Notes Scribes: tl40 February 26 - March 02, 2018 Chapter 2. Genome Assembly Algorithms 2.1. Statistical Theory 2.2. Algorithmic Theory Idury-Waterman Algorithm Estimating size of graphs used
CSCI 1820 Notes Scribes: tl40 February 26 - March 02, 2018 Chapter 2. Genome Assembly Algorithms 2.1. Statistical Theory 2.2. Algorithmic Theory Idury-Waterman Algorithm Estimating size of graphs used
Modules. 6 Hamilton Graphs (4-8 lectures) Introduction Necessary conditions and sufficient conditions Exercises...
 Modules 6 Hamilton Graphs (4-8 lectures) 135 6.1 Introduction................................ 136 6.2 Necessary conditions and sufficient conditions............. 137 Exercises..................................
Modules 6 Hamilton Graphs (4-8 lectures) 135 6.1 Introduction................................ 136 6.2 Necessary conditions and sufficient conditions............. 137 Exercises..................................
Genome Sequencing & Assembly. Slides by Carl Kingsford
 Genome Sequencing & Assembly Slides by Carl Kingsford Genome Sequencing ACCGTCCAATTGG...! TGGCAGGTTAACC... E.g. human: 3 billion bases split into 23 chromosomes Main tool of traditional sequencing: DNA
Genome Sequencing & Assembly Slides by Carl Kingsford Genome Sequencing ACCGTCCAATTGG...! TGGCAGGTTAACC... E.g. human: 3 billion bases split into 23 chromosomes Main tool of traditional sequencing: DNA
Lecture 1. 2 Motivation: Fast. Reliable. Cheap. Choose two.
 Approximation Algorithms and Hardness of Approximation February 19, 2013 Lecture 1 Lecturer: Ola Svensson Scribes: Alantha Newman 1 Class Information 4 credits Lecturers: Ola Svensson (ola.svensson@epfl.ch)
Approximation Algorithms and Hardness of Approximation February 19, 2013 Lecture 1 Lecturer: Ola Svensson Scribes: Alantha Newman 1 Class Information 4 credits Lecturers: Ola Svensson (ola.svensson@epfl.ch)
Introduction to Graph Theory
 Introduction to Graph Theory Tandy Warnow January 20, 2017 Graphs Tandy Warnow Graphs A graph G = (V, E) is an object that contains a vertex set V and an edge set E. We also write V (G) to denote the vertex
Introduction to Graph Theory Tandy Warnow January 20, 2017 Graphs Tandy Warnow Graphs A graph G = (V, E) is an object that contains a vertex set V and an edge set E. We also write V (G) to denote the vertex
IDBA A Practical Iterative de Bruijn Graph De Novo Assembler
 IDBA A Practical Iterative de Bruijn Graph De Novo Assembler Yu Peng, Henry C.M. Leung, S.M. Yiu, and Francis Y.L. Chin Department of Computer Science, The University of Hong Kong Pokfulam Road, Hong Kong
IDBA A Practical Iterative de Bruijn Graph De Novo Assembler Yu Peng, Henry C.M. Leung, S.M. Yiu, and Francis Y.L. Chin Department of Computer Science, The University of Hong Kong Pokfulam Road, Hong Kong
3 Euler Tours, Hamilton Cycles, and Their Applications
 3 Euler Tours, Hamilton Cycles, and Their Applications 3.1 Euler Tours and Applications 3.1.1 Euler tours Carefully review the definition of (closed) walks, trails, and paths from Section 1... Definition
3 Euler Tours, Hamilton Cycles, and Their Applications 3.1 Euler Tours and Applications 3.1.1 Euler tours Carefully review the definition of (closed) walks, trails, and paths from Section 1... Definition
Alignment of Long Sequences
 Alignment of Long Sequences BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2009 Mark Craven craven@biostat.wisc.edu Pairwise Whole Genome Alignment: Task Definition Given a pair of genomes (or other large-scale
Alignment of Long Sequences BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2009 Mark Craven craven@biostat.wisc.edu Pairwise Whole Genome Alignment: Task Definition Given a pair of genomes (or other large-scale
CS270 Combinatorial Algorithms & Data Structures Spring Lecture 19:
 CS270 Combinatorial Algorithms & Data Structures Spring 2003 Lecture 19: 4.1.03 Lecturer: Satish Rao Scribes: Kevin Lacker and Bill Kramer Disclaimer: These notes have not been subjected to the usual scrutiny
CS270 Combinatorial Algorithms & Data Structures Spring 2003 Lecture 19: 4.1.03 Lecturer: Satish Rao Scribes: Kevin Lacker and Bill Kramer Disclaimer: These notes have not been subjected to the usual scrutiny
Sequencing. Short Read Alignment. Sequencing. Paired-End Sequencing 6/10/2010. Tobias Rausch 7 th June 2010 WGS. ChIP-Seq. Applied Biosystems.
 Sequencing Short Alignment Tobias Rausch 7 th June 2010 WGS RNA-Seq Exon Capture ChIP-Seq Sequencing Paired-End Sequencing Target genome Fragments Roche GS FLX Titanium Illumina Applied Biosystems SOLiD
Sequencing Short Alignment Tobias Rausch 7 th June 2010 WGS RNA-Seq Exon Capture ChIP-Seq Sequencing Paired-End Sequencing Target genome Fragments Roche GS FLX Titanium Illumina Applied Biosystems SOLiD
Description of a genome assembler: CABOG
 Theo Zimmermann Description of a genome assembler: CABOG CABOG (Celera Assembler with the Best Overlap Graph) is an assembler built upon the Celera Assembler, which, at first, was designed for Sanger sequencing,
Theo Zimmermann Description of a genome assembler: CABOG CABOG (Celera Assembler with the Best Overlap Graph) is an assembler built upon the Celera Assembler, which, at first, was designed for Sanger sequencing,
Genome Assembly and De Novo RNAseq
 Genome Assembly and De Novo RNAseq BMI 7830 Kun Huang Department of Biomedical Informatics The Ohio State University Outline Problem formulation Hamiltonian path formulation Euler path and de Bruijin graph
Genome Assembly and De Novo RNAseq BMI 7830 Kun Huang Department of Biomedical Informatics The Ohio State University Outline Problem formulation Hamiltonian path formulation Euler path and de Bruijin graph
Performance analysis of parallel de novo genome assembly in shared memory system
 IOP Conference Series: Earth and Environmental Science PAPER OPEN ACCESS Performance analysis of parallel de novo genome assembly in shared memory system To cite this article: Syam Budi Iryanto et al 2018
IOP Conference Series: Earth and Environmental Science PAPER OPEN ACCESS Performance analysis of parallel de novo genome assembly in shared memory system To cite this article: Syam Budi Iryanto et al 2018
Module 6 NP-Complete Problems and Heuristics
 Module 6 NP-Complete Problems and Heuristics Dr. Natarajan Meghanathan Professor of Computer Science Jackson State University Jackson, MS 39217 E-mail: natarajan.meghanathan@jsums.edu P, NP-Problems Class
Module 6 NP-Complete Problems and Heuristics Dr. Natarajan Meghanathan Professor of Computer Science Jackson State University Jackson, MS 39217 E-mail: natarajan.meghanathan@jsums.edu P, NP-Problems Class
Walking with Euler through Ostpreußen and RNA
 Walking with Euler through Ostpreußen and RNA Mark Muldoon February 4, 2010 Königsberg (1652) Kaliningrad (2007)? The Königsberg Bridge problem asks whether it is possible to walk around the old city in
Walking with Euler through Ostpreußen and RNA Mark Muldoon February 4, 2010 Königsberg (1652) Kaliningrad (2007)? The Königsberg Bridge problem asks whether it is possible to walk around the old city in
Advanced Operations Research Prof. G. Srinivasan Department of Management Studies Indian Institute of Technology, Madras
 Advanced Operations Research Prof. G. Srinivasan Department of Management Studies Indian Institute of Technology, Madras Lecture 28 Chinese Postman Problem In this lecture we study the Chinese postman
Advanced Operations Research Prof. G. Srinivasan Department of Management Studies Indian Institute of Technology, Madras Lecture 28 Chinese Postman Problem In this lecture we study the Chinese postman
5.1 Min-Max Theorem for General Matching
 CSC5160: Combinatorial Optimization and Approximation Algorithms Topic: General Matching Date: 4/01/008 Lecturer: Lap Chi Lau Scribe: Jennifer X.M. WU In this lecture, we discuss matchings in general graph.
CSC5160: Combinatorial Optimization and Approximation Algorithms Topic: General Matching Date: 4/01/008 Lecturer: Lap Chi Lau Scribe: Jennifer X.M. WU In this lecture, we discuss matchings in general graph.
Finishing Circular Assemblies. J Fass UCD Genome Center Bioinformatics Core Thursday April 16, 2015
 Finishing Circular Assemblies J Fass UCD Genome Center Bioinformatics Core Thursday April 16, 2015 Assembly Strategies de Bruijn graph Velvet, ABySS earlier, basic assemblers IDBA, SPAdes later, multi-k
Finishing Circular Assemblies J Fass UCD Genome Center Bioinformatics Core Thursday April 16, 2015 Assembly Strategies de Bruijn graph Velvet, ABySS earlier, basic assemblers IDBA, SPAdes later, multi-k
val(y, I) α (9.0.2) α (9.0.3)
 CS787: Advanced Algorithms Lecture 9: Approximation Algorithms In this lecture we will discuss some NP-complete optimization problems and give algorithms for solving them that produce a nearly optimal,
CS787: Advanced Algorithms Lecture 9: Approximation Algorithms In this lecture we will discuss some NP-complete optimization problems and give algorithms for solving them that produce a nearly optimal,
7.36/7.91 recitation. DG Lectures 5 & 6 2/26/14
 7.36/7.91 recitation DG Lectures 5 & 6 2/26/14 1 Announcements project specific aims due in a little more than a week (March 7) Pset #2 due March 13, start early! Today: library complexity BWT and read
7.36/7.91 recitation DG Lectures 5 & 6 2/26/14 1 Announcements project specific aims due in a little more than a week (March 7) Pset #2 due March 13, start early! Today: library complexity BWT and read
CMSC 451: Lecture 22 Approximation Algorithms: Vertex Cover and TSP Tuesday, Dec 5, 2017
 CMSC 451: Lecture 22 Approximation Algorithms: Vertex Cover and TSP Tuesday, Dec 5, 2017 Reading: Section 9.2 of DPV. Section 11.3 of KT presents a different approximation algorithm for Vertex Cover. Coping
CMSC 451: Lecture 22 Approximation Algorithms: Vertex Cover and TSP Tuesday, Dec 5, 2017 Reading: Section 9.2 of DPV. Section 11.3 of KT presents a different approximation algorithm for Vertex Cover. Coping
Optimal tour along pubs in the UK
 1 From Facebook Optimal tour along 24727 pubs in the UK Road distance (by google maps) see also http://www.math.uwaterloo.ca/tsp/pubs/index.html (part of TSP homepage http://www.math.uwaterloo.ca/tsp/
1 From Facebook Optimal tour along 24727 pubs in the UK Road distance (by google maps) see also http://www.math.uwaterloo.ca/tsp/pubs/index.html (part of TSP homepage http://www.math.uwaterloo.ca/tsp/
Classic Graph Theory Problems
 Classic Graph Theory Problems Hiroki Sayama sayama@binghamton.edu The Origin Königsberg bridge problem Pregel River (Solved negatively by Euler in 176) Representation in a graph Can all the seven edges
Classic Graph Theory Problems Hiroki Sayama sayama@binghamton.edu The Origin Königsberg bridge problem Pregel River (Solved negatively by Euler in 176) Representation in a graph Can all the seven edges
Reducing Genome Assembly Complexity with Optical Maps
 Reducing Genome Assembly Complexity with Optical Maps AMSC 663 Mid-Year Progress Report 12/13/2011 Lee Mendelowitz Lmendelo@math.umd.edu Advisor: Mihai Pop mpop@umiacs.umd.edu Computer Science Department
Reducing Genome Assembly Complexity with Optical Maps AMSC 663 Mid-Year Progress Report 12/13/2011 Lee Mendelowitz Lmendelo@math.umd.edu Advisor: Mihai Pop mpop@umiacs.umd.edu Computer Science Department
As of August 15, 2008, GenBank contained bases from reported sequences. The search procedure should be
 48 Bioinformatics I, WS 09-10, S. Henz (script by D. Huson) November 26, 2009 4 BLAST and BLAT Outline of the chapter: 1. Heuristics for the pairwise local alignment of two sequences 2. BLAST: search and
48 Bioinformatics I, WS 09-10, S. Henz (script by D. Huson) November 26, 2009 4 BLAST and BLAT Outline of the chapter: 1. Heuristics for the pairwise local alignment of two sequences 2. BLAST: search and
Manual of SOAPdenovo-Trans-v1.03. Yinlong Xie, Gengxiong Wu, Jingbo Tang,
 Manual of SOAPdenovo-Trans-v1.03 Yinlong Xie, 2013-07-19 Gengxiong Wu, 2013-07-19 Jingbo Tang, 2013-07-19 ********** Introduction SOAPdenovo-Trans is a de novo transcriptome assembler basing on the SOAPdenovo
Manual of SOAPdenovo-Trans-v1.03 Yinlong Xie, 2013-07-19 Gengxiong Wu, 2013-07-19 Jingbo Tang, 2013-07-19 ********** Introduction SOAPdenovo-Trans is a de novo transcriptome assembler basing on the SOAPdenovo
Lecture 5: Markov models
 Master s course Bioinformatics Data Analysis and Tools Lecture 5: Markov models Centre for Integrative Bioinformatics Problem in biology Data and patterns are often not clear cut When we want to make a
Master s course Bioinformatics Data Analysis and Tools Lecture 5: Markov models Centre for Integrative Bioinformatics Problem in biology Data and patterns are often not clear cut When we want to make a
Graphs and Puzzles. Eulerian and Hamiltonian Tours.
 Graphs and Puzzles. Eulerian and Hamiltonian Tours. CSE21 Winter 2017, Day 11 (B00), Day 7 (A00) February 3, 2017 http://vlsicad.ucsd.edu/courses/cse21-w17 Exam Announcements Seating Chart on Website Good
Graphs and Puzzles. Eulerian and Hamiltonian Tours. CSE21 Winter 2017, Day 11 (B00), Day 7 (A00) February 3, 2017 http://vlsicad.ucsd.edu/courses/cse21-w17 Exam Announcements Seating Chart on Website Good
GLOBEX Bioinformatics (Summer 2015) Multiple Sequence Alignment
 GLOBEX Bioinformatics (Summer 2015) Multiple Sequence Alignment Scoring Dynamic Programming algorithms Heuristic algorithms CLUSTAL W Courtesy of jalview Motivations Collective (or aggregate) statistic
GLOBEX Bioinformatics (Summer 2015) Multiple Sequence Alignment Scoring Dynamic Programming algorithms Heuristic algorithms CLUSTAL W Courtesy of jalview Motivations Collective (or aggregate) statistic
Eulerian tours. Russell Impagliazzo and Miles Jones Thanks to Janine Tiefenbruck. April 20, 2016
 Eulerian tours Russell Impagliazzo and Miles Jones Thanks to Janine Tiefenbruck http://cseweb.ucsd.edu/classes/sp16/cse21-bd/ April 20, 2016 Seven Bridges of Konigsberg Is there a path that crosses each
Eulerian tours Russell Impagliazzo and Miles Jones Thanks to Janine Tiefenbruck http://cseweb.ucsd.edu/classes/sp16/cse21-bd/ April 20, 2016 Seven Bridges of Konigsberg Is there a path that crosses each
Biology 644: Bioinformatics
 Find the best alignment between 2 sequences with lengths n and m, respectively Best alignment is very dependent upon the substitution matrix and gap penalties The Global Alignment Problem tries to find
Find the best alignment between 2 sequences with lengths n and m, respectively Best alignment is very dependent upon the substitution matrix and gap penalties The Global Alignment Problem tries to find
IE 102 Spring Routing Through Networks - 1
 IE 102 Spring 2017 Routing Through Networks - 1 The Bridges of Koenigsberg: Euler 1735 Graph Theory began in 1735 Leonard Eüler Visited Koenigsberg People wondered whether it is possible to take a walk,
IE 102 Spring 2017 Routing Through Networks - 1 The Bridges of Koenigsberg: Euler 1735 Graph Theory began in 1735 Leonard Eüler Visited Koenigsberg People wondered whether it is possible to take a walk,
3 No-Wait Job Shops with Variable Processing Times
 3 No-Wait Job Shops with Variable Processing Times In this chapter we assume that, on top of the classical no-wait job shop setting, we are given a set of processing times for each operation. We may select
3 No-Wait Job Shops with Variable Processing Times In this chapter we assume that, on top of the classical no-wait job shop setting, we are given a set of processing times for each operation. We may select
Graph theory. Po-Shen Loh. June We begin by collecting some basic facts which can be proved via bare-hands techniques.
 Graph theory Po-Shen Loh June 013 1 Basic results We begin by collecting some basic facts which can be proved via bare-hands techniques. 1. The sum of all of the degrees is equal to twice the number of
Graph theory Po-Shen Loh June 013 1 Basic results We begin by collecting some basic facts which can be proved via bare-hands techniques. 1. The sum of all of the degrees is equal to twice the number of
Number Theory and Graph Theory
 1 Number Theory and Graph Theory Chapter 7 Graph properties By A. Satyanarayana Reddy Department of Mathematics Shiv Nadar University Uttar Pradesh, India E-mail: satya8118@gmail.com 2 Module-2: Eulerian
1 Number Theory and Graph Theory Chapter 7 Graph properties By A. Satyanarayana Reddy Department of Mathematics Shiv Nadar University Uttar Pradesh, India E-mail: satya8118@gmail.com 2 Module-2: Eulerian
Hybrid Parallel Programming
 Hybrid Parallel Programming for Massive Graph Analysis KameshMdd Madduri KMadduri@lbl.gov ComputationalResearch Division Lawrence Berkeley National Laboratory SIAM Annual Meeting 2010 July 12, 2010 Hybrid
Hybrid Parallel Programming for Massive Graph Analysis KameshMdd Madduri KMadduri@lbl.gov ComputationalResearch Division Lawrence Berkeley National Laboratory SIAM Annual Meeting 2010 July 12, 2010 Hybrid
Reducing Genome Assembly Complexity with Optical Maps Final Report
 Reducing Genome Assembly Complexity with Optical Maps Final Report Lee Mendelowitz LMendelo@math.umd.edu Advisor: Dr. Mihai Pop Computer Science Department Center for Bioinformatics and Computational Biology
Reducing Genome Assembly Complexity with Optical Maps Final Report Lee Mendelowitz LMendelo@math.umd.edu Advisor: Dr. Mihai Pop Computer Science Department Center for Bioinformatics and Computational Biology
De-Novo Genome Assembly and its Current State
 De-Novo Genome Assembly and its Current State Anne-Katrin Emde April 17, 2013 Freie Universität Berlin, Algorithmische Bioinformatik Max Planck Institut für Molekulare Genetik, Computational Molecular
De-Novo Genome Assembly and its Current State Anne-Katrin Emde April 17, 2013 Freie Universität Berlin, Algorithmische Bioinformatik Max Planck Institut für Molekulare Genetik, Computational Molecular
Discrete Mathematics and Probability Theory Fall 2013 Vazirani Note 7
 CS 70 Discrete Mathematics and Probability Theory Fall 2013 Vazirani Note 7 An Introduction to Graphs A few centuries ago, residents of the city of Königsberg, Prussia were interested in a certain problem.
CS 70 Discrete Mathematics and Probability Theory Fall 2013 Vazirani Note 7 An Introduction to Graphs A few centuries ago, residents of the city of Königsberg, Prussia were interested in a certain problem.
Genomic Finishing & Consed
 Genomic Finishing & Consed SEA stages of genomic analysis Draft vs Finished Draft Sequence Single sequencing approach Limited human intervention Cheap, Fast Finished sequence Multiple approaches Human
Genomic Finishing & Consed SEA stages of genomic analysis Draft vs Finished Draft Sequence Single sequencing approach Limited human intervention Cheap, Fast Finished sequence Multiple approaches Human
Computational Molecular Biology
 Computational Molecular Biology Erwin M. Bakker Lecture 3, mainly from material by R. Shamir [2] and H.J. Hoogeboom [4]. 1 Pairwise Sequence Alignment Biological Motivation Algorithmic Aspect Recursive
Computational Molecular Biology Erwin M. Bakker Lecture 3, mainly from material by R. Shamir [2] and H.J. Hoogeboom [4]. 1 Pairwise Sequence Alignment Biological Motivation Algorithmic Aspect Recursive
Path Finding in Graphs. Problem Set #2 will be posted by tonight
 Path Finding in Graphs Problem Set #2 will be posted by tonight 1 From Last Time Two graphs representing 5-mers from the sequence "GACGGCGGCGCACGGCGCAA" Hamiltonian Path: Eulerian Path: Each k-mer is a
Path Finding in Graphs Problem Set #2 will be posted by tonight 1 From Last Time Two graphs representing 5-mers from the sequence "GACGGCGGCGCACGGCGCAA" Hamiltonian Path: Eulerian Path: Each k-mer is a
ON HEURISTIC METHODS IN NEXT-GENERATION SEQUENCING DATA ANALYSIS
 ON HEURISTIC METHODS IN NEXT-GENERATION SEQUENCING DATA ANALYSIS Ivan Vogel Doctoral Degree Programme (1), FIT BUT E-mail: xvogel01@stud.fit.vutbr.cz Supervised by: Jaroslav Zendulka E-mail: zendulka@fit.vutbr.cz
ON HEURISTIC METHODS IN NEXT-GENERATION SEQUENCING DATA ANALYSIS Ivan Vogel Doctoral Degree Programme (1), FIT BUT E-mail: xvogel01@stud.fit.vutbr.cz Supervised by: Jaroslav Zendulka E-mail: zendulka@fit.vutbr.cz
Michał Kierzynka et al. Poznan University of Technology. 17 March 2015, San Jose
 Michał Kierzynka et al. Poznan University of Technology 17 March 2015, San Jose The research has been supported by grant No. 2012/05/B/ST6/03026 from the National Science Centre, Poland. DNA de novo assembly
Michał Kierzynka et al. Poznan University of Technology 17 March 2015, San Jose The research has been supported by grant No. 2012/05/B/ST6/03026 from the National Science Centre, Poland. DNA de novo assembly
1 Abstract. 2 Introduction. 3 Requirements
 1 Abstract 2 Introduction This SOP describes the HMP Whole- Metagenome Annotation Pipeline run at CBCB. This pipeline generates a 'Pretty Good Assembly' - a reasonable attempt at reconstructing pieces
1 Abstract 2 Introduction This SOP describes the HMP Whole- Metagenome Annotation Pipeline run at CBCB. This pipeline generates a 'Pretty Good Assembly' - a reasonable attempt at reconstructing pieces
Introduction and tutorial for SOAPdenovo. Xiaodong Fang Department of Science and BGI May, 2012
 Introduction and tutorial for SOAPdenovo Xiaodong Fang fangxd@genomics.org.cn Department of Science and Technology @ BGI May, 2012 Why de novo assembly? Genome is the genetic basis for different phenotypes
Introduction and tutorial for SOAPdenovo Xiaodong Fang fangxd@genomics.org.cn Department of Science and Technology @ BGI May, 2012 Why de novo assembly? Genome is the genetic basis for different phenotypes
Traveling Salesman Problem (TSP) Input: undirected graph G=(V,E), c: E R + Goal: find a tour (Hamiltonian cycle) of minimum cost
 Traveling Salesman Problem (TSP) Input: undirected graph G=(V,E), c: E R + Goal: find a tour (Hamiltonian cycle) of minimum cost Traveling Salesman Problem (TSP) Input: undirected graph G=(V,E), c: E R
Traveling Salesman Problem (TSP) Input: undirected graph G=(V,E), c: E R + Goal: find a tour (Hamiltonian cycle) of minimum cost Traveling Salesman Problem (TSP) Input: undirected graph G=(V,E), c: E R
How to apply de Bruijn graphs to genome assembly
 PRIMER How to apply de Bruijn graphs to genome assembly Phillip E C Compeau, Pavel A Pevzner & lenn Tesler A mathematical concept known as a de Bruijn graph turns the formidable challenge of assembling
PRIMER How to apply de Bruijn graphs to genome assembly Phillip E C Compeau, Pavel A Pevzner & lenn Tesler A mathematical concept known as a de Bruijn graph turns the formidable challenge of assembling
Euler and Hamilton paths. Jorge A. Cobb The University of Texas at Dallas
 Euler and Hamilton paths Jorge A. Cobb The University of Texas at Dallas 1 Paths and the adjacency matrix The powers of the adjacency matrix A r (with normal, not boolean multiplication) contain the number
Euler and Hamilton paths Jorge A. Cobb The University of Texas at Dallas 1 Paths and the adjacency matrix The powers of the adjacency matrix A r (with normal, not boolean multiplication) contain the number
Chapter 6. The Traveling-Salesman Problem. Section 1. Hamilton circuits and Hamilton paths.
 Chapter 6. The Traveling-Salesman Problem Section 1. Hamilton circuits and Hamilton paths. Recall: an Euler path is a path that travels through every edge of a graph once and only once; an Euler circuit
Chapter 6. The Traveling-Salesman Problem Section 1. Hamilton circuits and Hamilton paths. Recall: an Euler path is a path that travels through every edge of a graph once and only once; an Euler circuit
EULERIAN GRAPHS AND ITS APPLICATIONS
 EULERIAN GRAPHS AND ITS APPLICATIONS Aruna R 1, Madhu N.R 2 & Shashidhar S.N 3 1.2&3 Assistant Professor, Department of Mathematics. R.L.Jalappa Institute of Technology, Doddaballapur, B lore Rural Dist
EULERIAN GRAPHS AND ITS APPLICATIONS Aruna R 1, Madhu N.R 2 & Shashidhar S.N 3 1.2&3 Assistant Professor, Department of Mathematics. R.L.Jalappa Institute of Technology, Doddaballapur, B lore Rural Dist
CSE 549: Genome Assembly Intro & OLC. All slides in this lecture not marked with * courtesy of Ben Langmead.
 CSE 9: Genome Assembly Intro & OLC All slides in this lecture not marked with * courtesy of Ben Langmead. Shotgun Sequencing Many copies of the DNA Shear it, randomly breaking them into many small pieces,
CSE 9: Genome Assembly Intro & OLC All slides in this lecture not marked with * courtesy of Ben Langmead. Shotgun Sequencing Many copies of the DNA Shear it, randomly breaking them into many small pieces,
1 The Traveling Salesperson Problem (TSP)
 CS 598CSC: Approximation Algorithms Lecture date: January 23, 2009 Instructor: Chandra Chekuri Scribe: Sungjin Im In the previous lecture, we had a quick overview of several basic aspects of approximation
CS 598CSC: Approximation Algorithms Lecture date: January 23, 2009 Instructor: Chandra Chekuri Scribe: Sungjin Im In the previous lecture, we had a quick overview of several basic aspects of approximation
February 19, Integer programming. Outline. Problem formulation. Branch-andbound
 Olga Galinina olga.galinina@tut.fi ELT-53656 Network Analysis and Dimensioning II Department of Electronics and Communications Engineering Tampere University of Technology, Tampere, Finland February 19,
Olga Galinina olga.galinina@tut.fi ELT-53656 Network Analysis and Dimensioning II Department of Electronics and Communications Engineering Tampere University of Technology, Tampere, Finland February 19,
Finding homologous sequences in databases
 Finding homologous sequences in databases There are multiple algorithms to search sequences databases BLAST (EMBL, NCBI, DDBJ, local) FASTA (EMBL, local) For protein only databases scan via Smith-Waterman
Finding homologous sequences in databases There are multiple algorithms to search sequences databases BLAST (EMBL, NCBI, DDBJ, local) FASTA (EMBL, local) For protein only databases scan via Smith-Waterman
Midterm 2 Solutions. CS70 Discrete Mathematics for Computer Science, Fall 2007
 CS70 Discrete Mathematics for Computer Science, Fall 007 Midterm Solutions Note: These solutions are not necessarily model answers Rather, they are designed to be tutorial in nature, and sometimes contain
CS70 Discrete Mathematics for Computer Science, Fall 007 Midterm Solutions Note: These solutions are not necessarily model answers Rather, they are designed to be tutorial in nature, and sometimes contain
