Sandia National Laboratories: Implementation of and Experimenting with a Clustering Tool
|
|
|
- Alexandra Melton
- 5 years ago
- Views:
Transcription
1 Sandia National Laboratories: Implementation of and Experimenting with a Clustering Tool Team: Avani Gadani, Daniel Lowd, Brian Roney, Eric Wu Harvey Mudd College Dr. Belinda Thom (Advisor) Dr. Kevin Boyack (Liaison) May 8, 2003
2 Contents 1 Introduction 1 2 Background Problem Definition Approach Goals Management Timeline Team Member Contributions Tool Development Approach Development Environment Interface Description Code Structure Future Plans Proposed GUI changes Validity A Clustering Algorithms 11 A.1 kmeans A.2 SingleLinkage B Screenshots 13 C Source Code 17 C.1 controller.m C.2 plotwindow.m C.3 clustercontrol.m C.4 selectcontrol.m i
3 Chapter 1 Introduction Clustering is a tool for better understanding a given set of data. It has uses in various informatics fields such as genomics and crosscitation analysis that t to generate data sets large enough that it s difficult for humans to visually correlate. We have implemented a tool that allows side by side comparison of different dimension reductions of a given data set, as well as the ability to cluster points using a kmeans clustering and a singlelink clustering algorithm. The code is generic, so more clustering algorithms may be added in the future. The tool is written in MATLAB for its rapid prototyping ability and was designed to function on data sets of 500 to 10,000 twodimensional data points as requested by our sponsor. The two currently implemented clustering algorithms were chosen for their simplicity, speed, and ability to handle varying types of data sets. Kmeans clustering is a partitional algorithm which runs quickly but ts to result in clusters with nonconvex shapes [3]. Singlelink clustering, on the other hand, is a hierarchical algorithm that can deal with arbitrary shapes, potentially at the expense of simple clusters[3]. 1
4 Chapter 2 Background Sandia works in various fields that require careful analysis and correlation of data, e.g. they have data points in infant leukemia research, and they d like to determine similar genes to use medicines on. Correlation of titles and abstracts to institutions in research papers published from various sources would show trs in development. Researchers at Sandia desire a tool that would allow them to visually compare multiple ordinations, analyze them, and that would calculate various validity metrics for each ordination. We re approaching this problem through clustering. In clustering, the object is to place elements into the same cluster when they are similar enough according to some predefined metric. The predefined metric is one aspect that makes clustering a subjective process. There are two types of clustering: 1) partitional clustering, where every point is assigned to exactly one cluster, and 2) hierarchical clustering, where every cluster of size greater than one circumscribes a smaller group inside of it [2]. 2.1 Problem Definition Sandia needs to be able to view scatter plots of different dimensionally reduced data sets. Additionally, Sandia would like to be able to apply a clustering on different scatter plots in order to aid in comparing how similar various ordinations are. Also, user intervention on automatic clustering is desired in order to see how the clustering applies to scatterplots generated using other algorithms. Finally, we wish to provide a variety of clustering metrics in order to aid users in assessing a particular datasets clusterability. Ease of use and generality is considered important in the design. We have chosen to provide a GUI interface, as well as for many of the key functions to be accessible through the MATLAB command line. 2
5 Chapter 3 Approach Sandia s current approach to analyzing data begins with a data set with some arbitrary number of features. Each data point carries a unique ID and values for various features defining the data. The features dep of the type of data being studied, but may include data such as abstracts, citations, and keywords in a title for a data set consisting of technical articles, or pitch and tempo of a song over time in the case of a data set of musical pieces. In this form, a data set is not amenable to automatic clustering, nor to human insight. The dimensionality of the data (i.e., number of features) is likely to be too high to be easily visualized by a person. However, the data also cannot be automatically explored yet, due to problems in determining how to compare various features. Defining distance metrics between features is necessary for clustering to take place. Sandia deals with the problem by defining several different similarity metrics. These metrics may be as simple as Euclidean distance between numerical data, or as complex as necessary, e.g., number of papers cited in common between two articles. These similarity metrics produce a set of pairwise distances in a.sim file. Each row in a.sim file consists of two unique ID s and the pairwise similarity between them. Note that not all possible pairs of data points may be represented, due to incomplete information in the original data set or lack of similarity. The various.sim files can then be sent to VxOrd, a graph layout program. The similarity metric in conjunction with VxOrd acts as a dimensional reduction of the original data to a 2dimensional plane. Based on various parameters, VxOrd stochastically converts the.sim file to a 2D scattergraph, with each point representing one of the unique ID s in the original data set, and distances between points roughly corresponding to the inverse of the similarity between them. A given layout of points on the xy plane is referred to as an ordination. Different values for VxOrd s parameters will result in different scattergraphs with various levels of clumping. An ordination provided by VxOrd can be output as a.coord file, consisting of each unique ID and its x and ycoordinates. The.coord files can either be viewed via VxInsight, a visualization tool discussed later, or sent to our MATLAB clustering tool. Our tool can run various clustering algorithms (currently kmeans and singlelink) to cluster the ordination from VxOrd. We can also open multiple.coord files based on different similarity measures or different ordinations based on the same similarity measure sidebyside. With multiple ordinations open, a user can choose points in one plot either manually or from an automatic clustering and view where the corresponding points in the other plots are placed. This functionality allows us to identify points in other plots that appear together as a single cluster in a particular ordination. This is useful for human validation, since if a cluster ts to 3
6 CHAPTER 3. APPROACH 4 Data ID0001 Feature1 Feature2 ID0002 Feature1 Feature2... Similarity Computation Pairwise Similarities ID0001 ID ID0001 ID vxord Ordination ID ID vxinsight Figure 3.1: Sandia s current process. Starting with a database of identifiers and information about them (features), a pairwise similarity file (.sim file) is generated using some similarity computation algorithm operating on a subset of the features. Using vxord, each identifier is then mapped to an xy coordinate so that the Cartesian distance between two points roughly reflects their similarity. Finally, vxinsight is used to visualize the data. remain together in ordinations based on keywords in titles and cocitations, for example, then we know that articles with similar keywords in titles are likely to cite the same papers. In addition, the tool should support various validity metrics in the future. The sidebyside comparison of clusterings in addition to the validity metrics should suggest ordinations that should be studied in further detail in VxInsight. As shown in Figure 3.1, once the task of reducing data to a easily visualized state is completed, and one of many ordinations is chosen for further study, VxInsight can be used to manually explore the data. VxInsight is a visualization tool that links the.coord files to the original data set (in the form of a Microsoft Access database) and shows a 3d plot of the ordination. At this point, Figure 3.2 human investigation of clusters in detail can begin. 3.1 Goals The main goals of the tool are To allow a user to visually compare different coordinate files produced using different features or random seeds. To provide functionality for clustering the data using several algorithms, and to automatically compare clusterings using known validation methods. To ultimately allow the user and the clustering algorithm to work together to produce a better clustering solution than either could individually.
7 CHAPTER 3. APPROACH 5 Data ID01 Feature1 Feature2... FeatureM ID02 Feature1 Feature2... FeatureM... IDN Feature1 Feature2... FeatureM Similarity Computation 1 Similarity Computation 2... Similarity Computation K Sim File 1 ID01 ID ID01 ID Sim File 2 ID01 ID ID01 ID Sim File K ID01 ID ID01 ID vxord vxord... vxord Ordination 1 ID ID Ordination 2.1 ID ID Ordination 2.2 ID ID Ordination 2.H..... ID ID Ordination K ID ID Clustering Comparison Tool Figure 3.2: Extension to process, for comparing different similarity metrics. The tool we propose to develop will read multiple ordinations generated from different similarity files, which in turn were generated using different similarity measures. To allow a user to experiment with different validity metrics over clusterings. Our tool is currently MATLAB based and for Microsoft Windows c that will have filelevel interaction with vxord and possibly vxinsight. The actual layout of the tool will be described later, but essentially the tool displays scatterplots of data sets under different clustering algorithms or different similarity metrics from vxord. A user may manually recluster portions of a data set when automatic clustering algorithms perform poorly. Also, a user may apply metrics to the clustered plots to automatically determine the relative quality of a given clustering. To keep the transition from vxinsight to our tool simple, we need to to be able to identify and refer back to clusterings of points of the plot from vxinsight. One way to do this would be the use of a clustering algorithm such as kmeans. Currently, both kmeans and singlelinkage clustering algorithms (See Appix A) are implemented in our tool. We hope to add clustering algorithms and optimize them in C before delivering our product.
8 Chapter 4 Management 4.1 Timeline Our current plan is to meet the following deadlines: Late Dec Early January Implement nearestneighbor validity. January More clustering metrics (e.g. Gaussians). January Make necessary GUI changes. FebruaryMarch Do more experiments and begin work on a paper. Apr. nd Code Freeze Apr. nd Poster Due May th Final Report/CD 4.2 Team Member Contributions We ve currently divided the project so that Daniel and Brian are working on the coding of the problem while Eric and Avani are exploring the research aspect. As soon as the tool reaches a stable feature set, the entire team ints to become involved in the research aspect of this project. In particular, Daniel is going to focus on tool optimizations and integrations, Brian is going to explore data sets, and Eric and Avani are going to look into cluster validity (Chapter 6) 6
9 Chapter 5 Tool Development Approach 5.1 Development Environment To quickly develop an extensible solution, we used MATLAB as our programming language and development platform. The main disadvantages of MATLAB are that it makes some restrictions on GUI development and may be somewhat slower than an implementation in C or C++. In particular, singlelink is very slow, so we hope to optimize the builtin MATLAB algorithm using C. However, there s a large amount of functionality builtin to MATLAB, including several clustering algorithms and plotting tools. 5.2 Interface Description In our proposal we had the following Project Design Specifications: We shall implement a function in MATLAB that will generate a GUI. This GUI will include: A file browsing system to select data sets to examine, each of which will be placed in a separate plot window. The ability to zoom, scale, and/or rotate any of the plots individually. The ability to select a subset of a plot, which will highlight the equivalent points in the other plot windows. User can select an unlimited number of.coord files for comparison. All of these features, except for plot rotation, have already been implemented. In addition, users may cluster any ordination with either the kmeans or the single linkage clustering algorithm. The user also specifies the number of clusters to produce. Each cluster in the ordination is plotted in a different color and symbol combination. MATLAB will plot points in any of 7 colors and 13 symbols, yielding 91 potential combinations. When one ordination is clustered, all ordination plots are recolored to reflect the clustering. Users may also perform a manual clustering by specifying a color, a symbol, and a set of points in one of the plots. 7
10 CHAPTER 5. TOOL DEVELOPMENT APPROACH 8 Screeenshots of our tool can be found in Appix B. 5.3 Code Structure Our tool consists of four classes, each one responsible for the operations of one window type in the tool. The Controller manages the other windows, including windows for ordination plots and dialogs for clustering and point selection. The PlotWindow class colors and displays a single ordination. It communicates with the Controller through a number of callbacks. The ClusterControl class allows the user to specify the ordination to cluster and the algorithm and parameters to use for the clustering. This information is passed onto the Controller in a callback, which calls the actual clustering algorithm and tells each PlotWindow to recolor itself according to the new clustering. The SelectControl class lets the user specify the color and symbol type to color selected regions. None of PlotWindow, ClusterControl, and SelectControl ever references any class but Controller. This structure, with the Controller responsible for managing the other objects, simplifies design and implementation and results in code that is easier to maintain. This will be helpful for implementing additional requirements, such as the validity metrics.
11 Chapter 6 Future Plans 6.1 Proposed GUI changes A function that allows you to pick a point in a clustered plot and zoom onto the cluster that point is in. A Command Line Interface so that we can script our experiments. The ability to save and load results. Incorporate Validity. Cleaner and more optimized code. 6.2 Validity Another goal for next semester is to research and implement a variety of validity metrics. It may be difficult to determine whether an ordination has any clusterings, or to decide which of several clusterings is more meaningful. To determine whether an ordination can be clustered usefully, we int to implement geometric comparisons such as nearest neighbor and Hubert s statistic on the ordination. While this cannot solve the problem entirely, it will help to differentiate ordinations from each other and from randomly generated data [5]. Geometric validity measures focus on whether a representation of a data set is sufficiently nonrandom to have meaningful clusters at all, while clusterbased metrics look at the clusters generated [2]. Using Monte Carlo methods to create random data sets, we can use geometric measures to statistically compare ordinations to randomly distributed data. If the difference between the values of the validity measure for the random data and the ordination is too small, then the ordination may be too random to cluster effectively [4]. Clusterbased metrics such as the BaileyDubes index attempt to determine how well the data fits the pattern expected by a given clustering algorithm, indicating whether or not that algorithm provides a good clustering for a given data set. 9
12 CHAPTER 6. FUTURE PLANS Figure 6.1: A possible clustering There is no conclusive answer to the clustering problem, as everyone has a different idea of what a good clustering is. Validity metrics are one method of of attempting to objectively determine the quality of a clustering [5]. Ideally, given many different ordinations of a given data set, using many different similarity computations, validity metrics would help automatically suggest similarities and ordinations that are worthy of further study using vxinsight. Fig 6.1 demonstrates why the clustering problem is hard, as it is difficult to judge whether the bulges on the side of the plot are meant to be clusters or not. If one algorithm places those points in clusters and another does not, a validity metric may assist in determining which clustering fits the data best. However, cluster validity is subjective and domain specific, so a given metric may or may not work on a given algorithm. The final answer can only be found through human interpretation based on the similarities in the original data set.
13 Appix A Clustering Algorithms A.1 kmeans kmeans clustering is a commonly used partitional algorithm. The standard kmeans clustering requires choosing a number of clusters, k, and assigns each data point a label corresponding with its cluster membership. The algorithm works as follows: First, choose k cluster centers. Determine each data point s cluster membership according to the cluster center closest to it by a distance, d. Recompute the cluster centers based on the actual cluster membership. Then repeat until some criteria is met, such as a lack of data points being reassigned to new clusters. The underlying kmeans process minimizes, where! (A.1) is the center of cluster ". The runtime for the kmeans algorithm is # for each iteration. However, the number of iterations necessary is unknown since standard %$'&)( kmeans is not guaranteed to converge. In addition, clusterings produced by kmeans are depent on the starting points of the clusters. Variants of kmeans exist which are guaranteed to produce an optimal clustering from any starting points [1]. The advantages of this algorithm are that it runs in time linear to the cardinality of the data set, and modifications to the algorithm can be made to guarantee an optimal partitioning (Jain/Murty/Flynn, page 18). An inherent issue in kmeans clustering is that convex clusters t to be produced when Euclidean distance is used as the distance measure. A.2 SingleLinkage Singlelink clustering can broaden one s search to include nonconvex clusters, though the algorithm introduces other issues. Singlelink is an agglomerative hierarchical algorithm, 11
14 APPENDIX A. CLUSTERING ALGORITHMS 12 x x x x x x x x x x x x x x x x x x x x x x x x x x x x x x x x x x x x Figure A.1: Example of a scatterplot with nonconvex clusters which assigns each data point to an individual cluster, then merges clusters together according to the minimum of all pairwise distances between clusters members. The advantage of merging clusters based on distances between distinct members of each cluster is that nonconvex clusters may be formed, such as clusters consisting of concentric circles. However, this tency may also produce clusters that are chained, or noncompact. In addition, the process of determining the minimum of distances is relatively computational expensive; the algorithm runs in O( (*,+.0/1( ).
15 Appix B Screenshots 13
16 APPENDIX B. SCREENSHOTS 14 Figure B.1: The Controller GUI. The Open button is used for loading ordinations. The Cluster and Select buttons open separate dialogs for automatically or manually clustering ordinations, respectively. The Zoom button toggles zoom mode on all plots. When zoom mode is on, leftclicking on a plot zooms in and rightclicking zooms out. Figure B.2: The Cluster dialog. Once the user has specified the ordination to cluster (from the popup menu), the algorithm to use, and any additional algorithm parameters, clicking on the Cluster button runs the algorithm. Though the clustering is done according to the selected ordination, all plots are recolored accordingly.
17 APPENDIX B. SCREENSHOTS 15 Figure B.3: A single plot window. Specifically, this shows an ordination from the ARIST data set clustered using kmeans and 30 clusters. Different colors and symbols are used to distinguish one cluster from another. Figure B.4: Another plot window. This shows a different ordination form the ARIST data set clustered using the single linkage algorithm with 30 clusters.
18 APPENDIX B. SCREENSHOTS 16 Figure B.5: The Select dialog. Permits the user to choose the symbol and color for manually changing selected points. When this dialog is open, clicking and dragging a box on an ordination plot will redraw all contained points using the color and symbol specified here. Corresponding points in other plots will be redrawn with this new color and symbol as well.
19 Appix C Source Code C.1 controller.m function varargout = controller(varargin) % CONTROLLER Application Mfile for controller.fig % FIG = CONTROLLER launch controller GUI. % CONTROLLER( callback_name,...) invoke the named callback. % Last Modified by GUIDE v2.0 10Nov :11:02 if nargin == 0 % LAUNCH GUI fig = openfig(mfilename, reuse ); % Use system color scheme for figure: set(fig, Color,get(0, defaultuicontrolbackgroundcolor )); % Generate a structure of handles to pass to callbacks, and store it. handles = guihandles(fig); handles.controlmode = 0; % what the controller thinks clicks on plots should do % 0 = nothing (default) % 1 = user selection of points % 2 = zoom % 3 = cluster handles.basecolor = 1; handles.highlightcolor = 2; 17
20 APPENDIX C. SOURCE CODE 18 % array of windows handles.plots = []; handles.plotnames = []; handles.plothandles = []; % temporary manipulation variables handles.points = []; handles.names = []; handles.colors = []; handles.filename = 0; % Current path, for opening new files handles.pathname = 0; % FIX THIS non implemented function % changeselectmode(fig, handles, 0); % Update handles structure guidata(fig, handles); if nargout > 0 varargout{1} = fig; elseif ischar(varargin{1}) % INVOKE NAMED SUBFUNCTION OR CALLBACK try if (nargout) [varargout{1:nargout}] = feval(varargin{:}); % FEVAL switchyard else feval(varargin{:}); % FEVAL switchyard catch disp(lasterr); ABOUT CALLBACKS: GUIDE automatically apps subfunction prototypes to this file, and sets objects callback properties to call them through the FEVAL switchyard above. This comment describes that mechanism. Each callback subfunction declaration has the following form: <SUBFUNCTION_NAME>(H, EVENTDATA, HANDLES, VARARGIN) The subfunction name is composed using the object s Tag and the
21 APPENDIX C. SOURCE CODE 19 callback type separated by _, e.g. slider2_callback, figure1_closerequestfcn, axis1_buttondownfcn. H is the callback object s handle (obtained using GCBO). EVENTDATA is empty, but reserved for future use. HANDLES is a structure containing handles of components in GUI using tags as fieldnames, e.g. handles.figure1, handles.slider2. This structure is created at GUI startup using GUIHANDLES and stored in the figure s application data using GUIDATA. A copy of the structure is passed to each callback. You can store additional information in this structure at GUI startup, and you can change the structure during callbacks. Call guidata(h, handles) after changing your copy to replace the stored original so that subsequent callbacks see the updates. Type "help guihandles" and "help guidata" for more information. VARARGIN contains any extra arguments you have passed to the callback. Specify the extra arguments by editing the callback property in the inspector. By default, GUIDE sets the property to: <MFILENAME>( <SUBFUNCTION_NAME>, gcbo, [], guidata(gcbo)) Add any extra arguments after the last argument, before the final closing parenthesis. function varargout = openbutton_callback(h, eventdata, handles, varargin) % Stub for Callback of the uicontrol handles.openbutton1. [handles.filename, handles.names, handles.points] = openbuttoninternal; len = length(handles.plots); done = 0; id = 1; % initial id value (increment until valid) while done done =1; for i=1:len, if(handles.plots(i)== id) % if we find the id already in the array id = id+1; % increment the id ; ; ; done=0; % and try again % we now have a unique id
22 APPENDIX C. SOURCE CODE 20 handles.plots = [handles.plots, id]; % so put it in the array handles.plotnames = strvcat(handles.plotnames, handles.filename); % keep track of the name of this plot % Initial coloring: all the same color [height, width] = size(handles.points); handles.colors = zeros(height, 1) + handles.basecolor; if(handles.filename =0) f = plotwindow; plotwindow( init, f, h, handles.filename, handles.names, handles.points,... handles.colors, id); handles.plothandles= [handles.plothandles, f]; ; guidata(h, handles); function [filename, names, points] = openbuttoninternal % Stub for Callback of the uicontrol handles.openbutton1. names = 0; points = 0; % Get filename from the user [filename, pathname] =... uigetfile({ *.coord, Ordinations (*.coord) ; *.*, All Files (*.*) },... Open File ); if (isequal(filename,0)) return; % Save the directory cd(pathname); % Get points [names, xcoords, ycoords] = textread(filename, %s%f%f ); points = [xcoords, ycoords];
23 APPENDIX C. SOURCE CODE 21 function varargout = clusterbutton_callback(h, eventdata, handles, varargin) handles = guidata(h); clustercontrol( init, handles.plotnames, h); function varargout = activecluster(h, name, iterations, numclusters, clusteralg) handles = guidata(h); len = length(handles.plots); for i = 1:len, if(strcmp(handles.plotnames(i, :), name)) if(clusteralg==0) [filename, names, points, colors, id] = plotwindow( getinfo, handles.pl % clusters = kmeans(names, points, numclusters, iterations); clusters = kmeans(points, numclusters, Maxiter, iterations,... EmptyAction, singleton ); for j = 1:len, plotwindow( recolor, handles.plothandles(j), names, clusters); ; ; if(clusteralg==1) [filename, names, points, colors, id] = plotwindow( getinfo, handles.pl linky = linkage(pdist(points), single ); clusters = cluster(linky, maxclust, numclusters); for j = 1:len, plotwindow( recolor, handles.plothandles(j), names, clusters); ; ; break; ; ; function varargout = togglebutton3_callback(h, eventdata, handles, varargin) if (get(handles.togglebutton4, Value ) == 1 &&... get(handles.togglebutton3, Value ) == 1) set(handles.togglebutton4, Value, 0); togglebutton4_callback(h, 0, handles);
24 APPENDIX C. SOURCE CODE 22 if (get(handles.togglebutton3, Value ) == 1) % Open selection GUI handles.selectfig = selectcontrol( init, h); handles.controlmode = 1; else close(handles.selectfig); handles.controlmode = 0; guidata(h, handles); function selectclosingcallback(h) % Switch out of selection mode when required GUI is closed handles = guidata(h); set(handles.togglebutton3, Value, 0); handles.controlmode = 0; guidata(h, handles); function setselectcolor(h, c) handles = guidata(h); handles.selectcolor = c; guidata(h, handles); function mode = getmode(h) handles = guidata(h); mode = handles.controlmode; function plotpointselectioncallback(h, names) handles = guidata(h); len = length(handles.plots); for i = 1:len, plotwindow( select, handles.plothandles(i), names, handles.selectcolor); ;
25 APPENDIX C. SOURCE CODE 23 function varargout = togglebutton4_callback(h, eventdata, handles, varargin) if (get(handles.togglebutton4, Value ) == 1 &&... get(handles.togglebutton3, Value ) == 1) set(handles.togglebutton3, Value, 0); togglebutton3_callback(h, 0, handles); len = length(handles.plots); h = gcf; for i = 1:len, if (get(handles.togglebutton4, Value ) == 1) zoom(handles.plots(i), ON ); else zoom(handles.plots(i), OFF ); ; figure(gcf); function varargout = plotclosingcallback(h, id) handles = guidata(h); % get handle data back again len = length(handles.plots); % how many plots we had open before n=1; for i = 1:len, if(handles.plots(i)== id) n=i; % find the index of the array that matches filename ; ; handles.plots = [handles.plots(1:n1), handles.plots(n+1:len)]; % and remove it from the array handles.plotnames = strvcat(handles.plotnames(1:n1,:), handles.plotnames(n+1:len,:) handles.plothandles = [handles.plothandles(1:n1), handles.plothandles(n+1:len)]; guidata(h, handles); % save changes to handles
26 APPENDIX C. SOURCE CODE 24 C.2 plotwindow.m function varargout = plotwindow(varargin) % PLOTWINDOW Application Mfile for plotwindow.fig % FIG = PLOTWINDOW launch plotwindow GUI. % PLOTWINDOW( callback_name,...) invoke the named callback. % Last Modified by GUIDE v2.0 10Nov :07:54 if nargin == 0 % LAUNCH GUI fig = openfig(mfilename, new ); % Use system color scheme for figure: set(fig, Color,get(0, defaultuicontrolbackgroundcolor )); % Generate a structure of handles to pass to callbacks, and store it. handles = guihandles(fig); handles.parent = 0; handles.filename = 0; handles.names = 0; handles.points = 0; handles.colors = 0; handles.id = 0; guidata(fig, handles); if nargout > 0 varargout{1} = fig; elseif ischar(varargin{1}) % INVOKE NAMED SUBFUNCTION OR CALLBACK try if (nargout) [varargout{1:nargout}] = feval(varargin{:}); % FEVAL switchyard else feval(varargin{:}); % FEVAL switchyard catch disp(lasterr);
27 APPENDIX C. SOURCE CODE 25 function init(h, parent, filename, names, points, colors, id) % Set the title bar to the filename set(h, Name, filename); % Set all the handles data handles = guidata(h); handles.parent = parent; handles.filename = filename; handles.names = names; handles.points = points; handles.colors = colors; handles.id = id; guidata(h, handles); % Plot the graph redraw(h); function [filename, names, points, colors, id] = getinfo(h) handles = guidata(h); filename = handles.filename; names = handles.names; points = handles.points; colors = handles.colors; id = handles.id; function recolor(h, names, colors) handles = guidata(h); basecolors = zeros(length(handles.names),1)1; handles.colors = fastcolor(handles.names, names, colors, basecolors); guidata(h, handles); redraw(h); function colors2 = mergecolor(names2, names1, colors1, basecolors) colors2 = 0; if (length(names2) == 0 length(names1) == 0)
28 APPENDIX C. SOURCE CODE 26 colors2 = basecolors; return; for idx1 = ceil(length(names1)/2):length(names1) for idx2 = 1:length(names2) if (strcmp(names1(idx1), names2(idx2))) before = mergecolor(names2(1:idx21), names1(1:idx11),... colors1(1:idx11), basecolors(1:idx21)); after = mergecolor(names2(idx2+1:length(names2)),... names1(idx1+1:length(names1)),... colors1(idx1+1:length(names1)), basecolors(idx2+1:length(names2))); colors2 = [before; colors1(idx1); after]; return; colors2 = mergecolor(names2, names1(1:(ceil(length(names1)/2) 1)), colors1, colors2); function colors2 = mergecolor2(names2, names1, colors1, basecolors) colors2 = mergecolor2rec(names2, names1, colors1, basecolors,... 1, length(names1), 1, length(names2)); function colors2 = mergecolor2rec(names2, names1, colors1, basecolors,... min1, max1, min2, max2) colors2 = basecolors; if (max1 min1 < 0 max2 min2 < 0) return; % Start halfway between min1 and max1 idx1 = ceil((min1 + max1)/2); while (idx1 <= max1) for idx2 = min2:max2 if (strcmp(names1(idx1), names2(idx2)))
29 APPENDIX C. SOURCE CODE 27 colors2(idx2) = colors1(idx1); colors2 = mergecolor2rec(names2, names1, colors1, colors2,... min1, idx11, min2, idx21); colors2 = mergecolor2rec(names2, names1, colors1, colors2,... idx1+1, max1, idx2+1, max2); return; idx1 = idx1 + 1; colors2 = mergecolor2rec(names2, names1, colors1, colors2,... min1, min1 + ceil((max1min1)/2) 1, min2, max2); function colors2 = slowcolor(names2, names1, colors1, basecolors) colors2 = basecolors; idx2 = 1; for idx1 = 1:length(names1) for off2 = idx2:length(names2) if (strcmp(names1(idx1), names2(off2))) colors2(off2) = colors1(idx1); idx2 = off2 + 1; idx1 = idx1 + 1; function colors2 = fastcolor(names2, names1, colors1, basecolors) MAX_OFFSET = 500; colors2 = basecolors; idx1 = 1; idx2 = 1; while (idx1 <= length(colors1) & idx2 <= length(colors2)) if (idx1 == length(colors1)) a = 1;
30 APPENDIX C. SOURCE CODE 28 for k = 0:MAX_OFFSET % Ensure that our experimental offsets don t overflow the arrays if (idx1 + k > length(colors1)) k1 = length(colors1) idx1; else k1 = k; if (idx2 + k > length(colors2)) k2 = length(colors2) idx1; else k2 = k; for off1 = 0:k1 if strcmp(names1(idx1 + off1), names2(idx2 + k2)) break; if strcmp(names1(idx1 + off1), names2(idx2 + k2)) idx1 = idx1 + off1; idx2 = idx2 + k2; break; for off2 = 0:k2 if strcmp(names1(idx1 + k1), names2(idx2 + off2)) break; if strcmp(names1(idx1 + k1), names2(idx2 + off2)) idx1 = idx1 + k1; idx2 = idx2 + off2; break; Abort if (k == MAX_OFFSET) disp( Encountered section of MAX_OFFSET ID s with no match. ing ); break;
31 APPENDIX C. SOURCE CODE 29 colors2(idx2) = colors1(idx1); idx1 = idx1 + 1; idx2 = idx2 + 1; % Update the plot in this window function redraw(h) handles = guidata(h); axes(handles.axes1); %scaling = axis; cla; colorstrs = [ b, g, r, c, m, y ]; numcolors = length(colorstrs); symbolstrs = [., o, x, +, *, s, d, v, ˆ, <, >, p, h ]; numsymbols = length(symbolstrs); xlists = cell(numcolors, numsymbols); ylists = cell(numcolors, numsymbols); % Treat black points specially xblack = cell(1, numsymbols); yblack = cell(1, numsymbols); colors = handles.colors; numpoints = size(handles.points); for i = 1:numpoints % A color of sym => black if (colors(i) < 0) sym = colors(i); xblack{1,sym} = [xblack{1,sym}, handles.points(i,1)]; yblack{1,sym} = [yblack{1,sym}, handles.points(i,2)]; else col = mod(colors(i)1,numcolors) + 1; sym = mod(floor((colors(i)1)/numcolors), numsymbols) + 1; xlists{col,sym} = [xlists{col,sym}, handles.points(i, 1)]; ylists{col,sym} = [ylists{col,sym}, handles.points(i, 2)]; for sym = 1:numSymbols for col = 1:numColors hold on
32 APPENDIX C. SOURCE CODE 30 plot(xlists{col,sym}, ylists{col,sym}, [colorstrs(col), symbolstrs(sym)]); % Plot the black points separately hold on plot(xblack{1,sym}, yblack{1,sym}, [ k, symbolstrs(sym)]); %axis(scaling); ABOUT CALLBACKS: GUIDE automatically apps subfunction prototypes to this file, and sets objects callback properties to call them through the FEVAL switchyard above. This comment describes that mechanism. Each callback subfunction declaration has the following form: <SUBFUNCTION_NAME>(H, EVENTDATA, HANDLES, VARARGIN) The subfunction name is composed using the object s Tag and the callback type separated by _, e.g. slider2_callback, figure1_closerequestfcn, axis1_buttondownfcn. H is the callback object s handle (obtained using GCBO). EVENTDATA is empty, but reserved for future use. HANDLES is a structure containing handles of components in GUI using tags as fieldnames, e.g. handles.figure1, handles.slider2. This structure is created at GUI startup using GUIHANDLES and stored in the figure s application data using GUIDATA. A copy of the structure is passed to each callback. You can store additional information in this structure at GUI startup, and you can change the structure during callbacks. Call guidata(h, handles) after changing your copy to replace the stored original so that subsequent callbacks see the updates. Type "help guihandles" and "help guidata" for more information. VARARGIN contains any extra arguments you have passed to the callback. Specify the extra arguments by editing the callback property in the inspector. By default, GUIDE sets the property to: <MFILENAME>( <SUBFUNCTION_NAME>, gcbo, [], guidata(gcbo)) Add any extra arguments after the last argument, before the final closing parenthesis.
33 APPENDIX C. SOURCE CODE 31 function varargout = axes1_buttondownfcn(h, eventdata, handles, varargin) axes(handles.axes1); if (controller( getmode, handles.parent) = 1) return; point1 = get(gca, CurrentPoint ); finalrect = rbbox; point2 = get(gca, CurrentPoint ); point1 = point1(1,1:2); point2 = point2(1,1:2); minpoint = min(point1,point2); maxpoint = max(point1,point2); % starting location % wait for user to drag box % ing location % extract x and y % calculate min, max selectedpoints = []; % Get the selected region numpoints = size(handles.points); for i = 1:numPoints if (minpoint < handles.points(i,:)... & handles.points(i,:) < maxpoint) selectedpoints = [selectedpoints, handles.names(i)]; controller( plotpointselectioncallback, handles.parent, selectedpoints); function select(h, names, color) handles = guidata(h); handles.colors = mergecolor(handles.names, names,... zeros(length(names)) + color, handles.colors); guidata(h, handles); redraw(h);
34 APPENDIX C. SOURCE CODE 32 function varargout = figure1_deletefcn(h, eventdata, handles, varargin) % Notify our parent that we no longer exist controller( plotclosingcallback, handles.parent, handles.id); function varargout = figure1_resizefcn(h, eventdata, handles, varargin) % TODO eventually we ll do the layout manually, to make it look nicer.
35 APPENDIX C. SOURCE CODE 33 C.3 clustercontrol.m function varargout = clustercontrol(varargin) % CLUSTERCONTROL Application Mfile for clustercontrol.fig % FIG = CLUSTERCONTROL launch clustercontrol GUI. % CLUSTERCONTROL( callback_name,...) invoke the named callback. % Last Modified by GUIDE v2.0 17Nov :05:25 if nargin == 0 % LAUNCH GUI % See INIT this should never be reached elseif ischar(varargin{1}) % INVOKE NAMED SUBFUNCTION OR CALLBACK try if (nargout) [varargout{1:nargout}] = feval(varargin{:}); % FEVAL switchyard else feval(varargin{:}); % FEVAL switchyard catch disp(lasterr); ABOUT CALLBACKS: GUIDE automatically apps subfunction prototypes to this file, and sets objects callback properties to call them through the FEVAL switchyard above. This comment describes that mechanism. Each callback subfunction declaration has the following form: <SUBFUNCTION_NAME>(H, EVENTDATA, HANDLES, VARARGIN) The subfunction name is composed using the object s Tag and the callback type separated by _, e.g. slider2_callback, figure1_closerequestfcn, axis1_buttondownfcn. H is the callback object s handle (obtained using GCBO). EVENTDATA is empty, but reserved for future use. HANDLES is a structure containing handles of components in GUI using
36 APPENDIX C. SOURCE CODE 34 tags as fieldnames, e.g. handles.figure1, handles.slider2. This structure is created at GUI startup using GUIHANDLES and stored in the figure s application data using GUIDATA. A copy of the structure is passed to each callback. You can store additional information in this structure at GUI startup, and you can change the structure during callbacks. Call guidata(h, handles) after changing your copy to replace the stored original so that subsequent callbacks see the updates. Type "help guihandles" and "help guidata" for more information. VARARGIN contains any extra arguments you have passed to the callback. Specify the extra arguments by editing the callback property in the inspector. By default, GUIDE sets the property to: <MFILENAME>( <SUBFUNCTION_NAME>, gcbo, [], guidata(gcbo)) Add any extra arguments after the last argument, before the final closing parenthesis. % function init(plotnames, h) % Set the title bar to the filename fig = openfig(mfilename, reuse ); % Use system color scheme for figure: set(fig, Color,get(0, defaultuicontrolbackgroundcolor )); % Generate a structure of handles to pass to callbacks, and store it. handles = guihandles(fig); handles.numclusters = 30; handles.numiterations = 100; handles.clusteralg = 0; handles.plotnames = plotnames; handles.parent = h; % 0 = kmeans % 1 = singlelink set(handles.popupmenu1, String, plotnames); guidata(fig, handles); if nargout > 0 varargout{1} = fig;
37 APPENDIX C. SOURCE CODE 35 function varargout = popupmenu1_callback(h, eventdata, handles, varargin) temp=get(handles.popupmenu1, String ); value= get(handles.popupmenu1, Value ); handles.nameplot=temp(value, :); function varargout = singlelink_callback(h, eventdata, handles, varargin) handles = guidata(h); set(handles.singlelink, Value, 1); set(handles.kmeans, Value, 0); handles.clusteralg = 1; guidata(h, handles); function varargout = kmeans_callback(h, eventdata, handles, varargin) handles = guidata(h); set(handles.kmeans, Value, 1); set(handles.singlelink, Value, 0); handles.clusteralg = 0; guidata(h, handles); function varargout = clust_callback(h, eventdata, handles, varargin) handles = guidata(h); handles.numclusters = str2num(get(handles.clust, String )); guidata(h, handles); function varargout = iter_callback(h, eventdata, handles, varargin) handles = guidata(h); handles.numiterations = str2num(get(handles.iter, String )); guidata(h, handles); function varargout = clustergo_callback(h, eventdata, handles, varargin) handles = guidata(h); temp = get(handles.popupmenu1, String ); value = get(handles.popupmenu1, Value );
38 APPENDIX C. SOURCE CODE 36 handles.nameplot = temp(value,:); controller( activecluster, handles.parent, handles.nameplot,... handles.numiterations, handles.numclusters, handles.clusteralg); guidata(h, handles);
39 APPENDIX C. SOURCE CODE 37 C.4 selectcontrol.m function varargout = selectcontrol(varargin) % SELECTCONTROL Application Mfile for selectcontrol.fig % FIG = SELECTCONTROL launch selectcontrol GUI. % SELECTCONTROL( callback_name,...) invoke the named callback. % Last Modified by GUIDE v2.5 20Nov :05:54 if nargin == 0 % LAUNCH GUI fig = selectcontrol( init, 0); if nargout > 0 varargout{1} = fig; elseif ischar(varargin{1}) % INVOKE NAMED SUBFUNCTION OR CALLBACK try if (nargout) [varargout{1:nargout}] = feval(varargin{:}); % FEVAL switchyard else feval(varargin{:}); % FEVAL switchyard catch disp(lasterr); function fig = init(parent) fig = openfig(mfilename, reuse ); % Use system color scheme for figure: set(fig, Color,get(0, defaultuicontrolbackgroundcolor )); % Generate a structure of handles to pass to callbacks, and store it. handles = guihandles(fig); handles.parent = parent; handles.color = 1; handles.symbol = 1; guidata(fig, handles); notifyparent(fig, handles);
40 APPENDIX C. SOURCE CODE 38 if nargout > 0 varargout{1} = fig; ABOUT CALLBACKS: GUIDE automatically apps subfunction prototypes to this file, and sets objects callback properties to call them through the FEVAL switchyard above. This comment describes that mechanism. Each callback subfunction declaration has the following form: <SUBFUNCTION_NAME>(H, EVENTDATA, HANDLES, VARARGIN) The subfunction name is composed using the object s Tag and the callback type separated by _, e.g. slider2_callback, figure1_closerequestfcn, axis1_buttondownfcn. H is the callback object s handle (obtained using GCBO). EVENTDATA is empty, but reserved for future use. HANDLES is a structure containing handles of components in GUI using tags as fieldnames, e.g. handles.figure1, handles.slider2. This structure is created at GUI startup using GUIHANDLES and stored in the figure s application data using GUIDATA. A copy of the structure is passed to each callback. You can store additional information in this structure at GUI startup, and you can change the structure during callbacks. Call guidata(h, handles) after changing your copy to replace the stored original so that subsequent callbacks see the updates. Type "help guihandles" and "help guidata" for more information. VARARGIN contains any extra arguments you have passed to the callback. Specify the extra arguments by editing the callback property in the inspector. By default, GUIDE sets the property to: <MFILENAME>( <SUBFUNCTION_NAME>, gcbo, [], guidata(gcbo)) Add any extra arguments after the last argument, before the final closing parenthesis. function varargout = colorpopup_callback(h, eventdata, handles, varargin)
41 APPENDIX C. SOURCE CODE 39 handles.color = get(handles.colorpopup, Value ); guidata(h, handles); notifyparent(h, handles); function varargout = symbolpopup_callback(h, eventdata, handles, varargin) handles.symbol = get(handles.symbolpopup, Value ); guidata(h, handles); notifyparent(h, handles); function varargout = notifyparent(h, handles) if (handles.color == 7) controller( setselectcolor, handles.parent, handles.symbol); else controller( setselectcolor, handles.parent,... (handles.symbol1) * 6 + (handles.color)); % Executes during object deletion, before destroying properties. function varargout = figure1_deletefcn(h, eventdata, handles, varargin) controller( selectclosingcallback, handles.parent);
42 Bibliography A.K. Jain et. al. Data Clustering: A Review. ACM. Epter, Scott et. al. Clusterability Dectection and Cluster Initialization. IBM Power Parellel Group: RPI M. Halkidi et. al. Clustering Algorithms and Validity Measures. IEEE Tunkelang, Daniel. Making the Nearest Neighbor Meaningful. Endeca. p Zhao, Ying and George Karypis. Comparison of Agglomerative and Partitional Document Clustering Algorithms. University of Minnesota. 40
Implementation of and Experimenting with a Clustering Tool
 Computer Science Clinic Final Report for Sandia National Laboratories Implementation of and Experimenting with a Clustering Tool February 21, 2006 Team Members Avani Gadani (Team Leader) Daniel Lowd Brian
Computer Science Clinic Final Report for Sandia National Laboratories Implementation of and Experimenting with a Clustering Tool February 21, 2006 Team Members Avani Gadani (Team Leader) Daniel Lowd Brian
% Edit the above text to modify the response to help Video_Player. % Last Modified by GUIDE v May :38:12
 FILE NAME: Video_Player DESCRIPTION: Video Player Name Date Reason Sahariyaz 28-May-2015 Basic GUI concepts function varargout = Video_Player(varargin) VIDEO_PLAYER MATLAB code for Video_Player.fig VIDEO_PLAYER,
FILE NAME: Video_Player DESCRIPTION: Video Player Name Date Reason Sahariyaz 28-May-2015 Basic GUI concepts function varargout = Video_Player(varargin) VIDEO_PLAYER MATLAB code for Video_Player.fig VIDEO_PLAYER,
Signal and Systems. Matlab GUI based analysis. XpertSolver.com
 Signal and Systems Matlab GUI based analysis Description: This Signal and Systems based Project takes a sound file in.wav format and performed a detailed analysis, as well as filtering of the signal. The
Signal and Systems Matlab GUI based analysis Description: This Signal and Systems based Project takes a sound file in.wav format and performed a detailed analysis, as well as filtering of the signal. The
Clustering CS 550: Machine Learning
 Clustering CS 550: Machine Learning This slide set mainly uses the slides given in the following links: http://www-users.cs.umn.edu/~kumar/dmbook/ch8.pdf http://www-users.cs.umn.edu/~kumar/dmbook/dmslides/chap8_basic_cluster_analysis.pdf
Clustering CS 550: Machine Learning This slide set mainly uses the slides given in the following links: http://www-users.cs.umn.edu/~kumar/dmbook/ch8.pdf http://www-users.cs.umn.edu/~kumar/dmbook/dmslides/chap8_basic_cluster_analysis.pdf
Homeworks on FFT Instr. and Meas. for Communication Systems- Gianfranco Miele. Name Surname
 Homeworks on FFT 90822- Instr. and Meas. for Communication Systems- Gianfranco Miele Name Surname October 15, 2014 1 Name Surname 90822 (Gianfranco Miele): Homeworks on FFT Contents Exercise 1 (Solution)............................................
Homeworks on FFT 90822- Instr. and Meas. for Communication Systems- Gianfranco Miele Name Surname October 15, 2014 1 Name Surname 90822 (Gianfranco Miele): Homeworks on FFT Contents Exercise 1 (Solution)............................................
10701 Machine Learning. Clustering
 171 Machine Learning Clustering What is Clustering? Organizing data into clusters such that there is high intra-cluster similarity low inter-cluster similarity Informally, finding natural groupings among
171 Machine Learning Clustering What is Clustering? Organizing data into clusters such that there is high intra-cluster similarity low inter-cluster similarity Informally, finding natural groupings among
Clustering and Visualisation of Data
 Clustering and Visualisation of Data Hiroshi Shimodaira January-March 28 Cluster analysis aims to partition a data set into meaningful or useful groups, based on distances between data points. In some
Clustering and Visualisation of Data Hiroshi Shimodaira January-March 28 Cluster analysis aims to partition a data set into meaningful or useful groups, based on distances between data points. In some
Flow Control and Functions
 Flow Control and Functions Script files If's and For's Basics of writing functions Checking input arguments Variable input arguments Output arguments Documenting functions Profiling and Debugging Introduction
Flow Control and Functions Script files If's and For's Basics of writing functions Checking input arguments Variable input arguments Output arguments Documenting functions Profiling and Debugging Introduction
Clustering. Chapter 10 in Introduction to statistical learning
 Clustering Chapter 10 in Introduction to statistical learning 16 14 12 10 8 6 4 2 0 2 4 6 8 10 12 14 1 Clustering ² Clustering is the art of finding groups in data (Kaufman and Rousseeuw, 1990). ² What
Clustering Chapter 10 in Introduction to statistical learning 16 14 12 10 8 6 4 2 0 2 4 6 8 10 12 14 1 Clustering ² Clustering is the art of finding groups in data (Kaufman and Rousseeuw, 1990). ² What
Cluster Analysis for Microarray Data
 Cluster Analysis for Microarray Data Seventh International Long Oligonucleotide Microarray Workshop Tucson, Arizona January 7-12, 2007 Dan Nettleton IOWA STATE UNIVERSITY 1 Clustering Group objects that
Cluster Analysis for Microarray Data Seventh International Long Oligonucleotide Microarray Workshop Tucson, Arizona January 7-12, 2007 Dan Nettleton IOWA STATE UNIVERSITY 1 Clustering Group objects that
SYDE Winter 2011 Introduction to Pattern Recognition. Clustering
 SYDE 372 - Winter 2011 Introduction to Pattern Recognition Clustering Alexander Wong Department of Systems Design Engineering University of Waterloo Outline 1 2 3 4 5 All the approaches we have learned
SYDE 372 - Winter 2011 Introduction to Pattern Recognition Clustering Alexander Wong Department of Systems Design Engineering University of Waterloo Outline 1 2 3 4 5 All the approaches we have learned
1. Peralatan LAMPIRAN
 1. Peralatan LAMPIRAN 2. Data Sampel a. Air murni 3ml Energy(mj) Probability Air Murni 0.07 0.001 0.15 0.003 0.22 0.006 0.3 0.028 0.37 0.045 0.39 0.049 0.82 0.053 0.89 0.065 1.28 0.065 1.42 0.106 1.7
1. Peralatan LAMPIRAN 2. Data Sampel a. Air murni 3ml Energy(mj) Probability Air Murni 0.07 0.001 0.15 0.003 0.22 0.006 0.3 0.028 0.37 0.045 0.39 0.049 0.82 0.053 0.89 0.065 1.28 0.065 1.42 0.106 1.7
INF4820. Clustering. Erik Velldal. Nov. 17, University of Oslo. Erik Velldal INF / 22
 INF4820 Clustering Erik Velldal University of Oslo Nov. 17, 2009 Erik Velldal INF4820 1 / 22 Topics for Today More on unsupervised machine learning for data-driven categorization: clustering. The task
INF4820 Clustering Erik Velldal University of Oslo Nov. 17, 2009 Erik Velldal INF4820 1 / 22 Topics for Today More on unsupervised machine learning for data-driven categorization: clustering. The task
Hierarchical clustering
 Hierarchical clustering Rebecca C. Steorts, Duke University STA 325, Chapter 10 ISL 1 / 63 Agenda K-means versus Hierarchical clustering Agglomerative vs divisive clustering Dendogram (tree) Hierarchical
Hierarchical clustering Rebecca C. Steorts, Duke University STA 325, Chapter 10 ISL 1 / 63 Agenda K-means versus Hierarchical clustering Agglomerative vs divisive clustering Dendogram (tree) Hierarchical
Unsupervised Learning
 Outline Unsupervised Learning Basic concepts K-means algorithm Representation of clusters Hierarchical clustering Distance functions Which clustering algorithm to use? NN Supervised learning vs. unsupervised
Outline Unsupervised Learning Basic concepts K-means algorithm Representation of clusters Hierarchical clustering Distance functions Which clustering algorithm to use? NN Supervised learning vs. unsupervised
LAMPIRAN 1. Percobaan
 LAMPIRAN 1 1. Larutan 15 ppm Polystyrene ENERGI Percobaan 1 2 3 PROBABILITY 0.07 0.116 0.113 0.152 0.127 0.15 0.256 0.143 0.212 0.203 0.22 0.24 0.234 0.23 0.234 0.3 0.239 0.35 0.201 0.263 0.37 0.389 0.382
LAMPIRAN 1 1. Larutan 15 ppm Polystyrene ENERGI Percobaan 1 2 3 PROBABILITY 0.07 0.116 0.113 0.152 0.127 0.15 0.256 0.143 0.212 0.203 0.22 0.24 0.234 0.23 0.234 0.3 0.239 0.35 0.201 0.263 0.37 0.389 0.382
Cluster Analysis. Ying Shen, SSE, Tongji University
 Cluster Analysis Ying Shen, SSE, Tongji University Cluster analysis Cluster analysis groups data objects based only on the attributes in the data. The main objective is that The objects within a group
Cluster Analysis Ying Shen, SSE, Tongji University Cluster analysis Cluster analysis groups data objects based only on the attributes in the data. The main objective is that The objects within a group
CompClustTk Manual & Tutorial
 CompClustTk Manual & Tutorial Brandon King Copyright c California Institute of Technology Version 0.1.10 May 13, 2004 Contents 1 Introduction 1 1.1 Purpose.............................................
CompClustTk Manual & Tutorial Brandon King Copyright c California Institute of Technology Version 0.1.10 May 13, 2004 Contents 1 Introduction 1 1.1 Purpose.............................................
Machine Learning and Data Mining. Clustering (1): Basics. Kalev Kask
 Machine Learning and Data Mining Clustering (1): Basics Kalev Kask Unsupervised learning Supervised learning Predict target value ( y ) given features ( x ) Unsupervised learning Understand patterns of
Machine Learning and Data Mining Clustering (1): Basics Kalev Kask Unsupervised learning Supervised learning Predict target value ( y ) given features ( x ) Unsupervised learning Understand patterns of
CS 2750 Machine Learning. Lecture 19. Clustering. CS 2750 Machine Learning. Clustering. Groups together similar instances in the data sample
 Lecture 9 Clustering Milos Hauskrecht milos@cs.pitt.edu 539 Sennott Square Clustering Groups together similar instances in the data sample Basic clustering problem: distribute data into k different groups
Lecture 9 Clustering Milos Hauskrecht milos@cs.pitt.edu 539 Sennott Square Clustering Groups together similar instances in the data sample Basic clustering problem: distribute data into k different groups
Exploratory data analysis for microarrays
 Exploratory data analysis for microarrays Jörg Rahnenführer Computational Biology and Applied Algorithmics Max Planck Institute for Informatics D-66123 Saarbrücken Germany NGFN - Courses in Practical DNA
Exploratory data analysis for microarrays Jörg Rahnenführer Computational Biology and Applied Algorithmics Max Planck Institute for Informatics D-66123 Saarbrücken Germany NGFN - Courses in Practical DNA
Clustering. CE-717: Machine Learning Sharif University of Technology Spring Soleymani
 Clustering CE-717: Machine Learning Sharif University of Technology Spring 2016 Soleymani Outline Clustering Definition Clustering main approaches Partitional (flat) Hierarchical Clustering validation
Clustering CE-717: Machine Learning Sharif University of Technology Spring 2016 Soleymani Outline Clustering Definition Clustering main approaches Partitional (flat) Hierarchical Clustering validation
Computational Methods of Scientific Programming
 12.010 Computational Methods of Scientific Programming Lecturers Thomas A Herring, Jim Elliot, Chris Hill, Summary of Today s class We will look at Matlab: History Getting help Variable definitions and
12.010 Computational Methods of Scientific Programming Lecturers Thomas A Herring, Jim Elliot, Chris Hill, Summary of Today s class We will look at Matlab: History Getting help Variable definitions and
Unsupervised Learning : Clustering
 Unsupervised Learning : Clustering Things to be Addressed Traditional Learning Models. Cluster Analysis K-means Clustering Algorithm Drawbacks of traditional clustering algorithms. Clustering as a complex
Unsupervised Learning : Clustering Things to be Addressed Traditional Learning Models. Cluster Analysis K-means Clustering Algorithm Drawbacks of traditional clustering algorithms. Clustering as a complex
Lithium-Ion Battery Data. Management
 Lithium-Ion Battery Data Management Frank Ferrato Dr. Jung-Hyun Kim April 2018 Abstract: Lithium Ion Battery research is growing due to the need for renewable resources. Since the amount of research is
Lithium-Ion Battery Data Management Frank Ferrato Dr. Jung-Hyun Kim April 2018 Abstract: Lithium Ion Battery research is growing due to the need for renewable resources. Since the amount of research is
University of Florida CISE department Gator Engineering. Clustering Part 2
 Clustering Part 2 Dr. Sanjay Ranka Professor Computer and Information Science and Engineering University of Florida, Gainesville Partitional Clustering Original Points A Partitional Clustering Hierarchical
Clustering Part 2 Dr. Sanjay Ranka Professor Computer and Information Science and Engineering University of Florida, Gainesville Partitional Clustering Original Points A Partitional Clustering Hierarchical
11/2/2017 MIST.6060 Business Intelligence and Data Mining 1. Clustering. Two widely used distance metrics to measure the distance between two records
 11/2/2017 MIST.6060 Business Intelligence and Data Mining 1 An Example Clustering X 2 X 1 Objective of Clustering The objective of clustering is to group the data into clusters such that the records within
11/2/2017 MIST.6060 Business Intelligence and Data Mining 1 An Example Clustering X 2 X 1 Objective of Clustering The objective of clustering is to group the data into clusters such that the records within
Information Retrieval and Web Search Engines
 Information Retrieval and Web Search Engines Lecture 7: Document Clustering December 4th, 2014 Wolf-Tilo Balke and José Pinto Institut für Informationssysteme Technische Universität Braunschweig The Cluster
Information Retrieval and Web Search Engines Lecture 7: Document Clustering December 4th, 2014 Wolf-Tilo Balke and José Pinto Institut für Informationssysteme Technische Universität Braunschweig The Cluster
% Edit the above text to modify the response to help Principal
 function varargout = Principal(varargin) % OPFGUI MATLAB code for Principal.fig % OPFGUI, by itself, creates a new OPFGUI or raises the existing % singleton*. % % H = OPFGUI returns the handle to a new
function varargout = Principal(varargin) % OPFGUI MATLAB code for Principal.fig % OPFGUI, by itself, creates a new OPFGUI or raises the existing % singleton*. % % H = OPFGUI returns the handle to a new
Geostatistics 2D GMS 7.0 TUTORIALS. 1 Introduction. 1.1 Contents
 GMS 7.0 TUTORIALS 1 Introduction Two-dimensional geostatistics (interpolation) can be performed in GMS using the 2D Scatter Point module. The module is used to interpolate from sets of 2D scatter points
GMS 7.0 TUTORIALS 1 Introduction Two-dimensional geostatistics (interpolation) can be performed in GMS using the 2D Scatter Point module. The module is used to interpolate from sets of 2D scatter points
10601 Machine Learning. Hierarchical clustering. Reading: Bishop: 9-9.2
 161 Machine Learning Hierarchical clustering Reading: Bishop: 9-9.2 Second half: Overview Clustering - Hierarchical, semi-supervised learning Graphical models - Bayesian networks, HMMs, Reasoning under
161 Machine Learning Hierarchical clustering Reading: Bishop: 9-9.2 Second half: Overview Clustering - Hierarchical, semi-supervised learning Graphical models - Bayesian networks, HMMs, Reasoning under
Text Documents clustering using K Means Algorithm
 Text Documents clustering using K Means Algorithm Mrs Sanjivani Tushar Deokar Assistant professor sanjivanideokar@gmail.com Abstract: With the advancement of technology and reduced storage costs, individuals
Text Documents clustering using K Means Algorithm Mrs Sanjivani Tushar Deokar Assistant professor sanjivanideokar@gmail.com Abstract: With the advancement of technology and reduced storage costs, individuals
Clustering: Overview and K-means algorithm
 Clustering: Overview and K-means algorithm Informal goal Given set of objects and measure of similarity between them, group similar objects together K-Means illustrations thanks to 2006 student Martin
Clustering: Overview and K-means algorithm Informal goal Given set of objects and measure of similarity between them, group similar objects together K-Means illustrations thanks to 2006 student Martin
Building a Graphical User Interface
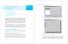 148 CHAPTER 9 Building a Graphical User Interface Building a Graphical User Interface CHAPTER 9 9.1 Getting started with GUIDE 9.2 Starting an action with a GUI element 9.3 Communicating with GUI elements
148 CHAPTER 9 Building a Graphical User Interface Building a Graphical User Interface CHAPTER 9 9.1 Getting started with GUIDE 9.2 Starting an action with a GUI element 9.3 Communicating with GUI elements
BBS654 Data Mining. Pinar Duygulu. Slides are adapted from Nazli Ikizler
 BBS654 Data Mining Pinar Duygulu Slides are adapted from Nazli Ikizler 1 Classification Classification systems: Supervised learning Make a rational prediction given evidence There are several methods for
BBS654 Data Mining Pinar Duygulu Slides are adapted from Nazli Ikizler 1 Classification Classification systems: Supervised learning Make a rational prediction given evidence There are several methods for
SECTION 2: PROGRAMMING WITH MATLAB. MAE 4020/5020 Numerical Methods with MATLAB
 SECTION 2: PROGRAMMING WITH MATLAB MAE 4020/5020 Numerical Methods with MATLAB 2 Functions and M Files M Files 3 Script file so called due to.m filename extension Contains a series of MATLAB commands The
SECTION 2: PROGRAMMING WITH MATLAB MAE 4020/5020 Numerical Methods with MATLAB 2 Functions and M Files M Files 3 Script file so called due to.m filename extension Contains a series of MATLAB commands The
Cluster Analysis. Mu-Chun Su. Department of Computer Science and Information Engineering National Central University 2003/3/11 1
 Cluster Analysis Mu-Chun Su Department of Computer Science and Information Engineering National Central University 2003/3/11 1 Introduction Cluster analysis is the formal study of algorithms and methods
Cluster Analysis Mu-Chun Su Department of Computer Science and Information Engineering National Central University 2003/3/11 1 Introduction Cluster analysis is the formal study of algorithms and methods
Lecture 10: Semantic Segmentation and Clustering
 Lecture 10: Semantic Segmentation and Clustering Vineet Kosaraju, Davy Ragland, Adrien Truong, Effie Nehoran, Maneekwan Toyungyernsub Department of Computer Science Stanford University Stanford, CA 94305
Lecture 10: Semantic Segmentation and Clustering Vineet Kosaraju, Davy Ragland, Adrien Truong, Effie Nehoran, Maneekwan Toyungyernsub Department of Computer Science Stanford University Stanford, CA 94305
Document Clustering: Comparison of Similarity Measures
 Document Clustering: Comparison of Similarity Measures Shouvik Sachdeva Bhupendra Kastore Indian Institute of Technology, Kanpur CS365 Project, 2014 Outline 1 Introduction The Problem and the Motivation
Document Clustering: Comparison of Similarity Measures Shouvik Sachdeva Bhupendra Kastore Indian Institute of Technology, Kanpur CS365 Project, 2014 Outline 1 Introduction The Problem and the Motivation
There are two ways to launch Graphical User Interface (GUI). You can either
 How to get started? There are two ways to launch Graphical User Interface (GUI). You can either 1. Click on the Guide icon 2. Type guide at the prompt Just follow the instruction below: To start GUI we
How to get started? There are two ways to launch Graphical User Interface (GUI). You can either 1. Click on the Guide icon 2. Type guide at the prompt Just follow the instruction below: To start GUI we
Information Retrieval and Web Search Engines
 Information Retrieval and Web Search Engines Lecture 7: Document Clustering May 25, 2011 Wolf-Tilo Balke and Joachim Selke Institut für Informationssysteme Technische Universität Braunschweig Homework
Information Retrieval and Web Search Engines Lecture 7: Document Clustering May 25, 2011 Wolf-Tilo Balke and Joachim Selke Institut für Informationssysteme Technische Universität Braunschweig Homework
MSA220 - Statistical Learning for Big Data
 MSA220 - Statistical Learning for Big Data Lecture 13 Rebecka Jörnsten Mathematical Sciences University of Gothenburg and Chalmers University of Technology Clustering Explorative analysis - finding groups
MSA220 - Statistical Learning for Big Data Lecture 13 Rebecka Jörnsten Mathematical Sciences University of Gothenburg and Chalmers University of Technology Clustering Explorative analysis - finding groups
Data Science and Statistics in Research: unlocking the power of your data Session 3.4: Clustering
 Data Science and Statistics in Research: unlocking the power of your data Session 3.4: Clustering 1/ 1 OUTLINE 2/ 1 Overview 3/ 1 CLUSTERING Clustering is a statistical technique which creates groupings
Data Science and Statistics in Research: unlocking the power of your data Session 3.4: Clustering 1/ 1 OUTLINE 2/ 1 Overview 3/ 1 CLUSTERING Clustering is a statistical technique which creates groupings
CSE 5243 INTRO. TO DATA MINING
 CSE 5243 INTRO. TO DATA MINING Cluster Analysis: Basic Concepts and Methods Huan Sun, CSE@The Ohio State University 09/25/2017 Slides adapted from UIUC CS412, Fall 2017, by Prof. Jiawei Han 2 Chapter 10.
CSE 5243 INTRO. TO DATA MINING Cluster Analysis: Basic Concepts and Methods Huan Sun, CSE@The Ohio State University 09/25/2017 Slides adapted from UIUC CS412, Fall 2017, by Prof. Jiawei Han 2 Chapter 10.
Understanding Clustering Supervising the unsupervised
 Understanding Clustering Supervising the unsupervised Janu Verma IBM T.J. Watson Research Center, New York http://jverma.github.io/ jverma@us.ibm.com @januverma Clustering Grouping together similar data
Understanding Clustering Supervising the unsupervised Janu Verma IBM T.J. Watson Research Center, New York http://jverma.github.io/ jverma@us.ibm.com @januverma Clustering Grouping together similar data
CS 1675 Introduction to Machine Learning Lecture 18. Clustering. Clustering. Groups together similar instances in the data sample
 CS 1675 Introduction to Machine Learning Lecture 18 Clustering Milos Hauskrecht milos@cs.pitt.edu 539 Sennott Square Clustering Groups together similar instances in the data sample Basic clustering problem:
CS 1675 Introduction to Machine Learning Lecture 18 Clustering Milos Hauskrecht milos@cs.pitt.edu 539 Sennott Square Clustering Groups together similar instances in the data sample Basic clustering problem:
CSE 5243 INTRO. TO DATA MINING
 CSE 5243 INTRO. TO DATA MINING Cluster Analysis: Basic Concepts and Methods Huan Sun, CSE@The Ohio State University Slides adapted from UIUC CS412, Fall 2017, by Prof. Jiawei Han 2 Chapter 10. Cluster
CSE 5243 INTRO. TO DATA MINING Cluster Analysis: Basic Concepts and Methods Huan Sun, CSE@The Ohio State University Slides adapted from UIUC CS412, Fall 2017, by Prof. Jiawei Han 2 Chapter 10. Cluster
Redefining and Enhancing K-means Algorithm
 Redefining and Enhancing K-means Algorithm Nimrat Kaur Sidhu 1, Rajneet kaur 2 Research Scholar, Department of Computer Science Engineering, SGGSWU, Fatehgarh Sahib, Punjab, India 1 Assistant Professor,
Redefining and Enhancing K-means Algorithm Nimrat Kaur Sidhu 1, Rajneet kaur 2 Research Scholar, Department of Computer Science Engineering, SGGSWU, Fatehgarh Sahib, Punjab, India 1 Assistant Professor,
ECLT 5810 Clustering
 ECLT 5810 Clustering What is Cluster Analysis? Cluster: a collection of data objects Similar to one another within the same cluster Dissimilar to the objects in other clusters Cluster analysis Grouping
ECLT 5810 Clustering What is Cluster Analysis? Cluster: a collection of data objects Similar to one another within the same cluster Dissimilar to the objects in other clusters Cluster analysis Grouping
Computational Methods of Scientific Programming. Matlab Lecture 3 Lecturers Thomas A Herring Chris Hill
 12.010 Computational Methods of Scientific Programming Matlab Lecture 3 Lecturers Thomas A Herring Chris Hill Summary of last class Continued examining Matlab operations path and addpath commands Variables
12.010 Computational Methods of Scientific Programming Matlab Lecture 3 Lecturers Thomas A Herring Chris Hill Summary of last class Continued examining Matlab operations path and addpath commands Variables
INTRODUCTION TO MATLAB INTERACTIVE GRAPHICS EXERCISES
 INTRODUCTION TO MATLAB INTERACTIVE GRAPHICS EXERCISES Eric Peasley, Department of Engineering Science, University of Oxford version 3.0, 2017 MATLAB Interactive Graphics Exercises In these exercises you
INTRODUCTION TO MATLAB INTERACTIVE GRAPHICS EXERCISES Eric Peasley, Department of Engineering Science, University of Oxford version 3.0, 2017 MATLAB Interactive Graphics Exercises In these exercises you
A NEW MACHINING COST CALCULATOR (MC 2 )
 A NEW MACHINING COST CALCULATOR (MC 2 ) By MATHEW RUSSELL JOHNSON A THESIS PRESENTED TO THE GRADUATE SCHOOL OF THE UNIVERSITY OF FLORIDA IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE OF MASTER
A NEW MACHINING COST CALCULATOR (MC 2 ) By MATHEW RUSSELL JOHNSON A THESIS PRESENTED TO THE GRADUATE SCHOOL OF THE UNIVERSITY OF FLORIDA IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE OF MASTER
Clustering to Reduce Spatial Data Set Size
 Clustering to Reduce Spatial Data Set Size Geoff Boeing arxiv:1803.08101v1 [cs.lg] 21 Mar 2018 1 Introduction Department of City and Regional Planning University of California, Berkeley March 2018 Traditionally
Clustering to Reduce Spatial Data Set Size Geoff Boeing arxiv:1803.08101v1 [cs.lg] 21 Mar 2018 1 Introduction Department of City and Regional Planning University of California, Berkeley March 2018 Traditionally
Unsupervised Learning: Clustering
 Unsupervised Learning: Clustering Vibhav Gogate The University of Texas at Dallas Slides adapted from Carlos Guestrin, Dan Klein & Luke Zettlemoyer Machine Learning Supervised Learning Unsupervised Learning
Unsupervised Learning: Clustering Vibhav Gogate The University of Texas at Dallas Slides adapted from Carlos Guestrin, Dan Klein & Luke Zettlemoyer Machine Learning Supervised Learning Unsupervised Learning
K-Means Clustering Using Localized Histogram Analysis
 K-Means Clustering Using Localized Histogram Analysis Michael Bryson University of South Carolina, Department of Computer Science Columbia, SC brysonm@cse.sc.edu Abstract. The first step required for many
K-Means Clustering Using Localized Histogram Analysis Michael Bryson University of South Carolina, Department of Computer Science Columbia, SC brysonm@cse.sc.edu Abstract. The first step required for many
CSE 547: Machine Learning for Big Data Spring Problem Set 2. Please read the homework submission policies.
 CSE 547: Machine Learning for Big Data Spring 2019 Problem Set 2 Please read the homework submission policies. 1 Principal Component Analysis and Reconstruction (25 points) Let s do PCA and reconstruct
CSE 547: Machine Learning for Big Data Spring 2019 Problem Set 2 Please read the homework submission policies. 1 Principal Component Analysis and Reconstruction (25 points) Let s do PCA and reconstruct
Clustering. CS294 Practical Machine Learning Junming Yin 10/09/06
 Clustering CS294 Practical Machine Learning Junming Yin 10/09/06 Outline Introduction Unsupervised learning What is clustering? Application Dissimilarity (similarity) of objects Clustering algorithm K-means,
Clustering CS294 Practical Machine Learning Junming Yin 10/09/06 Outline Introduction Unsupervised learning What is clustering? Application Dissimilarity (similarity) of objects Clustering algorithm K-means,
Supplementary Information
 Supplementary Information Retooling Laser Speckle Contrast Analysis Algorithm to Enhance Non-Invasive High Resolution Laser Speckle Functional Imaging of Cutaneous Microcirculation Surya C Gnyawali 1,
Supplementary Information Retooling Laser Speckle Contrast Analysis Algorithm to Enhance Non-Invasive High Resolution Laser Speckle Functional Imaging of Cutaneous Microcirculation Surya C Gnyawali 1,
Visual Profiler. User Guide
 Visual Profiler User Guide Version 3.0 Document No. 06-RM-1136 Revision: 4.B February 2008 Visual Profiler User Guide Table of contents Table of contents 1 Introduction................................................
Visual Profiler User Guide Version 3.0 Document No. 06-RM-1136 Revision: 4.B February 2008 Visual Profiler User Guide Table of contents Table of contents 1 Introduction................................................
The Language of Technical Computing. Computation. Visualization. Programming. Creating Graphical User Interfaces Version 1
 MATLAB The Language of Technical Computing Computation Visualization Programming Creating Graphical User Interfaces Version 1 How to Contact The MathWorks: 508-647-7000 Phone 508-647-7001 Fax The MathWorks,
MATLAB The Language of Technical Computing Computation Visualization Programming Creating Graphical User Interfaces Version 1 How to Contact The MathWorks: 508-647-7000 Phone 508-647-7001 Fax The MathWorks,
A Comparative study of Clustering Algorithms using MapReduce in Hadoop
 A Comparative study of Clustering Algorithms using MapReduce in Hadoop Dweepna Garg 1, Khushboo Trivedi 2, B.B.Panchal 3 1 Department of Computer Science and Engineering, Parul Institute of Engineering
A Comparative study of Clustering Algorithms using MapReduce in Hadoop Dweepna Garg 1, Khushboo Trivedi 2, B.B.Panchal 3 1 Department of Computer Science and Engineering, Parul Institute of Engineering
Ear Recognition. By: Zeyangyi Wang
 Ear Recognition By: Zeyangyi Wang Ear Recognition By: Zeyangyi Wang Online: < http://cnx.org/content/col11604/1.3/ > C O N N E X I O N S Rice University, Houston, Texas This selection and arrangement
Ear Recognition By: Zeyangyi Wang Ear Recognition By: Zeyangyi Wang Online: < http://cnx.org/content/col11604/1.3/ > C O N N E X I O N S Rice University, Houston, Texas This selection and arrangement
Clustering in R d. Clustering. Widely-used clustering methods. The k-means optimization problem CSE 250B
 Clustering in R d Clustering CSE 250B Two common uses of clustering: Vector quantization Find a finite set of representatives that provides good coverage of a complex, possibly infinite, high-dimensional
Clustering in R d Clustering CSE 250B Two common uses of clustering: Vector quantization Find a finite set of representatives that provides good coverage of a complex, possibly infinite, high-dimensional
Unsupervised Learning
 Unsupervised Learning Unsupervised learning Until now, we have assumed our training samples are labeled by their category membership. Methods that use labeled samples are said to be supervised. However,
Unsupervised Learning Unsupervised learning Until now, we have assumed our training samples are labeled by their category membership. Methods that use labeled samples are said to be supervised. However,
8. Control statements
 8. Control statements A simple C++ statement is each of the individual instructions of a program, like the variable declarations and expressions seen in previous sections. They always end with a semicolon
8. Control statements A simple C++ statement is each of the individual instructions of a program, like the variable declarations and expressions seen in previous sections. They always end with a semicolon
CHAPTER 4: CLUSTER ANALYSIS
 CHAPTER 4: CLUSTER ANALYSIS WHAT IS CLUSTER ANALYSIS? A cluster is a collection of data-objects similar to one another within the same group & dissimilar to the objects in other groups. Cluster analysis
CHAPTER 4: CLUSTER ANALYSIS WHAT IS CLUSTER ANALYSIS? A cluster is a collection of data-objects similar to one another within the same group & dissimilar to the objects in other groups. Cluster analysis
ECLT 5810 Clustering
 ECLT 5810 Clustering What is Cluster Analysis? Cluster: a collection of data objects Similar to one another within the same cluster Dissimilar to the objects in other clusters Cluster analysis Grouping
ECLT 5810 Clustering What is Cluster Analysis? Cluster: a collection of data objects Similar to one another within the same cluster Dissimilar to the objects in other clusters Cluster analysis Grouping
Breeding Guide. Customer Services PHENOME-NETWORKS 4Ben Gurion Street, 74032, Nes-Ziona, Israel
 Breeding Guide Customer Services PHENOME-NETWORKS 4Ben Gurion Street, 74032, Nes-Ziona, Israel www.phenome-netwoks.com Contents PHENOME ONE - INTRODUCTION... 3 THE PHENOME ONE LAYOUT... 4 THE JOBS ICON...
Breeding Guide Customer Services PHENOME-NETWORKS 4Ben Gurion Street, 74032, Nes-Ziona, Israel www.phenome-netwoks.com Contents PHENOME ONE - INTRODUCTION... 3 THE PHENOME ONE LAYOUT... 4 THE JOBS ICON...
University of Florida CISE department Gator Engineering. Clustering Part 4
 Clustering Part 4 Dr. Sanjay Ranka Professor Computer and Information Science and Engineering University of Florida, Gainesville DBSCAN DBSCAN is a density based clustering algorithm Density = number of
Clustering Part 4 Dr. Sanjay Ranka Professor Computer and Information Science and Engineering University of Florida, Gainesville DBSCAN DBSCAN is a density based clustering algorithm Density = number of
Gene expression & Clustering (Chapter 10)
 Gene expression & Clustering (Chapter 10) Determining gene function Sequence comparison tells us if a gene is similar to another gene, e.g., in a new species Dynamic programming Approximate pattern matching
Gene expression & Clustering (Chapter 10) Determining gene function Sequence comparison tells us if a gene is similar to another gene, e.g., in a new species Dynamic programming Approximate pattern matching
K-means Clustering & k-nn classification
 K-means Clustering & k-nn classification Andreas C. Kapourani (Credit: Hiroshi Shimodaira) 03 February 2016 1 Introduction In this lab session we will focus on K-means clustering and k-nearest Neighbour
K-means Clustering & k-nn classification Andreas C. Kapourani (Credit: Hiroshi Shimodaira) 03 February 2016 1 Introduction In this lab session we will focus on K-means clustering and k-nearest Neighbour
You are to turn in the following three graphs at the beginning of class on Wednesday, January 21.
 Computer Tools for Data Analysis & Presentation Graphs All public machines on campus are now equipped with Word 2010 and Excel 2010. Although fancier graphical and statistical analysis programs exist,
Computer Tools for Data Analysis & Presentation Graphs All public machines on campus are now equipped with Word 2010 and Excel 2010. Although fancier graphical and statistical analysis programs exist,
Copyright 2012, Oracle and/or its affiliates. All rights reserved.
 1 Oracle Partitioning für Einsteiger Hermann Bär Partitioning Produkt Management 2 Disclaimer The goal is to establish a basic understanding of what can be done with Partitioning I want you to start thinking
1 Oracle Partitioning für Einsteiger Hermann Bär Partitioning Produkt Management 2 Disclaimer The goal is to establish a basic understanding of what can be done with Partitioning I want you to start thinking
Ocean Data View. Getting Started
 Ocean Data View Getting Started January 20, 2011 1 Contents 1 General Information... 3 1.1 Installation... 3 1.2 ODV User Directory... 3 1.3 Application Window... 3 1.4 Data Model... 5 1.5 Data Import...
Ocean Data View Getting Started January 20, 2011 1 Contents 1 General Information... 3 1.1 Installation... 3 1.2 ODV User Directory... 3 1.3 Application Window... 3 1.4 Data Model... 5 1.5 Data Import...
v Overview SMS Tutorials Prerequisites Requirements Time Objectives
 v. 12.2 SMS 12.2 Tutorial Overview Objectives This tutorial describes the major components of the SMS interface and gives a brief introduction to the different SMS modules. Ideally, this tutorial should
v. 12.2 SMS 12.2 Tutorial Overview Objectives This tutorial describes the major components of the SMS interface and gives a brief introduction to the different SMS modules. Ideally, this tutorial should
Chapter 6 User-Defined Functions. dr.dcd.h CS 101 /SJC 5th Edition 1
 Chapter 6 User-Defined Functions dr.dcd.h CS 101 /SJC 5th Edition 1 MATLAB Functions M-files are collections of MATLAB statements that stored in a file, called a script file. Script files share the command
Chapter 6 User-Defined Functions dr.dcd.h CS 101 /SJC 5th Edition 1 MATLAB Functions M-files are collections of MATLAB statements that stored in a file, called a script file. Script files share the command
26 The closest pair problem
 The closest pair problem 1 26 The closest pair problem Sweep algorithms solve many kinds of proximity problems efficiently. We present a simple sweep that solves the two-dimensional closest pair problem
The closest pair problem 1 26 The closest pair problem Sweep algorithms solve many kinds of proximity problems efficiently. We present a simple sweep that solves the two-dimensional closest pair problem
Clustering Part 4 DBSCAN
 Clustering Part 4 Dr. Sanjay Ranka Professor Computer and Information Science and Engineering University of Florida, Gainesville DBSCAN DBSCAN is a density based clustering algorithm Density = number of
Clustering Part 4 Dr. Sanjay Ranka Professor Computer and Information Science and Engineering University of Florida, Gainesville DBSCAN DBSCAN is a density based clustering algorithm Density = number of
CS 223B Computer Vision Problem Set 3
 CS 223B Computer Vision Problem Set 3 Due: Feb. 22 nd, 2011 1 Probabilistic Recursion for Tracking In this problem you will derive a method for tracking a point of interest through a sequence of images.
CS 223B Computer Vision Problem Set 3 Due: Feb. 22 nd, 2011 1 Probabilistic Recursion for Tracking In this problem you will derive a method for tracking a point of interest through a sequence of images.
A New Online Clustering Approach for Data in Arbitrary Shaped Clusters
 A New Online Clustering Approach for Data in Arbitrary Shaped Clusters Richard Hyde, Plamen Angelov Data Science Group, School of Computing and Communications Lancaster University Lancaster, LA1 4WA, UK
A New Online Clustering Approach for Data in Arbitrary Shaped Clusters Richard Hyde, Plamen Angelov Data Science Group, School of Computing and Communications Lancaster University Lancaster, LA1 4WA, UK
Still More About Matlab GUI s (v. 1.3) Popup Menus. Popup Menu Exercise. Still More GUI Info - GE /29/2012. Copyright C. S. Tritt, Ph.D.
 Still More About Matlab GUI s (v. 1.3) Dr. C. S. Tritt with slides from Dr. J. LaMack January 24, 2012 Popup Menus User selects one from a mutually exclusive list of options The String property is typically
Still More About Matlab GUI s (v. 1.3) Dr. C. S. Tritt with slides from Dr. J. LaMack January 24, 2012 Popup Menus User selects one from a mutually exclusive list of options The String property is typically
Lesson 1 Parametric Modeling Fundamentals
 1-1 Lesson 1 Parametric Modeling Fundamentals Create Simple Parametric Models. Understand the Basic Parametric Modeling Process. Create and Profile Rough Sketches. Understand the "Shape before size" approach.
1-1 Lesson 1 Parametric Modeling Fundamentals Create Simple Parametric Models. Understand the Basic Parametric Modeling Process. Create and Profile Rough Sketches. Understand the "Shape before size" approach.
Chapter 6 Introduction to Defining Classes
 Introduction to Defining Classes Fundamentals of Java: AP Computer Science Essentials, 4th Edition 1 Objectives Design and implement a simple class from user requirements. Organize a program in terms of
Introduction to Defining Classes Fundamentals of Java: AP Computer Science Essentials, 4th Edition 1 Objectives Design and implement a simple class from user requirements. Organize a program in terms of
Unsupervised Learning and Clustering
 Unsupervised Learning and Clustering Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Spring 2009 CS 551, Spring 2009 c 2009, Selim Aksoy (Bilkent University)
Unsupervised Learning and Clustering Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Spring 2009 CS 551, Spring 2009 c 2009, Selim Aksoy (Bilkent University)
Function Graphing pdf (6/19/10)
 Specifications Quick Tour Tutorials Reference Forums Function Graphing pdf (6/19/10) Overview ND1 (v.1.1) can graph: single-valued real functions/programs, scatter plots, line and pie charts, images. In
Specifications Quick Tour Tutorials Reference Forums Function Graphing pdf (6/19/10) Overview ND1 (v.1.1) can graph: single-valued real functions/programs, scatter plots, line and pie charts, images. In
Artistic Text. Basics 1
 Basics 1 In this tutorial, we ll show you how to: Work with artistic text. Create, edit, and format text. Apply shadows, reflections, and other text effects. Create shaped text (or text-on-a-path). 2 Basics
Basics 1 In this tutorial, we ll show you how to: Work with artistic text. Create, edit, and format text. Apply shadows, reflections, and other text effects. Create shaped text (or text-on-a-path). 2 Basics
Survey of Math: Excel Spreadsheet Guide (for Excel 2016) Page 1 of 9
 Survey of Math: Excel Spreadsheet Guide (for Excel 2016) Page 1 of 9 Contents 1 Introduction to Using Excel Spreadsheets 2 1.1 A Serious Note About Data Security.................................... 2 1.2
Survey of Math: Excel Spreadsheet Guide (for Excel 2016) Page 1 of 9 Contents 1 Introduction to Using Excel Spreadsheets 2 1.1 A Serious Note About Data Security.................................... 2 1.2
Nearest Neighbor Predictors
 Nearest Neighbor Predictors September 2, 2018 Perhaps the simplest machine learning prediction method, from a conceptual point of view, and perhaps also the most unusual, is the nearest-neighbor method,
Nearest Neighbor Predictors September 2, 2018 Perhaps the simplest machine learning prediction method, from a conceptual point of view, and perhaps also the most unusual, is the nearest-neighbor method,
EL2310 Scientific Programming
 (pronobis@kth.se) Overview Overview Wrap Up More on Scripts and Functions Basic Programming Lecture 2 Lecture 3 Lecture 4 Wrap Up Last time Loading data from file: load( filename ) Graphical input and
(pronobis@kth.se) Overview Overview Wrap Up More on Scripts and Functions Basic Programming Lecture 2 Lecture 3 Lecture 4 Wrap Up Last time Loading data from file: load( filename ) Graphical input and
CLUSTERING IN BIOINFORMATICS
 CLUSTERING IN BIOINFORMATICS CSE/BIMM/BENG 8 MAY 4, 0 OVERVIEW Define the clustering problem Motivation: gene expression and microarrays Types of clustering Clustering algorithms Other applications of
CLUSTERING IN BIOINFORMATICS CSE/BIMM/BENG 8 MAY 4, 0 OVERVIEW Define the clustering problem Motivation: gene expression and microarrays Types of clustering Clustering algorithms Other applications of
Fast Stochastic Block Partition for Streaming Graphs
 Fast Stochastic Block Partition for Streaming Graphs Ahsen J. Uppal, and H. Howie Huang The George Washington University Abstract The graph partition problem continues to be challenging, particularly for
Fast Stochastic Block Partition for Streaming Graphs Ahsen J. Uppal, and H. Howie Huang The George Washington University Abstract The graph partition problem continues to be challenging, particularly for
142
 Scope Rules Thus, storage duration does not affect the scope of an identifier. The only identifiers with function-prototype scope are those used in the parameter list of a function prototype. As mentioned
Scope Rules Thus, storage duration does not affect the scope of an identifier. The only identifiers with function-prototype scope are those used in the parameter list of a function prototype. As mentioned
Reverse Engineering with a CASE Tool. Bret Johnson. Research advisors: Spencer Rugaber and Rich LeBlanc. October 6, Abstract
 Reverse Engineering with a CASE Tool Bret Johnson Research advisors: Spencer Rugaber and Rich LeBlanc October 6, 994 Abstract We examine using a CASE tool, Interactive Development Environment's Software
Reverse Engineering with a CASE Tool Bret Johnson Research advisors: Spencer Rugaber and Rich LeBlanc October 6, 994 Abstract We examine using a CASE tool, Interactive Development Environment's Software
FLUENT Secondary flow in a teacup Author: John M. Cimbala, Penn State University Latest revision: 26 January 2016
 FLUENT Secondary flow in a teacup Author: John M. Cimbala, Penn State University Latest revision: 26 January 2016 Note: These instructions are based on an older version of FLUENT, and some of the instructions
FLUENT Secondary flow in a teacup Author: John M. Cimbala, Penn State University Latest revision: 26 January 2016 Note: These instructions are based on an older version of FLUENT, and some of the instructions
Multivariate Analysis
 Multivariate Analysis Cluster Analysis Prof. Dr. Anselmo E de Oliveira anselmo.quimica.ufg.br anselmo.disciplinas@gmail.com Unsupervised Learning Cluster Analysis Natural grouping Patterns in the data
Multivariate Analysis Cluster Analysis Prof. Dr. Anselmo E de Oliveira anselmo.quimica.ufg.br anselmo.disciplinas@gmail.com Unsupervised Learning Cluster Analysis Natural grouping Patterns in the data
BoredGames Language Reference Manual A Language for Board Games. Brandon Kessler (bpk2107) and Kristen Wise (kew2132)
 BoredGames Language Reference Manual A Language for Board Games Brandon Kessler (bpk2107) and Kristen Wise (kew2132) 1 Table of Contents 1. Introduction... 4 2. Lexical Conventions... 4 2.A Comments...
BoredGames Language Reference Manual A Language for Board Games Brandon Kessler (bpk2107) and Kristen Wise (kew2132) 1 Table of Contents 1. Introduction... 4 2. Lexical Conventions... 4 2.A Comments...
K-means Clustering & PCA
 K-means Clustering & PCA Andreas C. Kapourani (Credit: Hiroshi Shimodaira) 02 February 2018 1 Introduction In this lab session we will focus on K-means clustering and Principal Component Analysis (PCA).
K-means Clustering & PCA Andreas C. Kapourani (Credit: Hiroshi Shimodaira) 02 February 2018 1 Introduction In this lab session we will focus on K-means clustering and Principal Component Analysis (PCA).
Accelerating Unique Strategy for Centroid Priming in K-Means Clustering
 IJIRST International Journal for Innovative Research in Science & Technology Volume 3 Issue 07 December 2016 ISSN (online): 2349-6010 Accelerating Unique Strategy for Centroid Priming in K-Means Clustering
IJIRST International Journal for Innovative Research in Science & Technology Volume 3 Issue 07 December 2016 ISSN (online): 2349-6010 Accelerating Unique Strategy for Centroid Priming in K-Means Clustering
Los Alamos National Laboratory I/O Visualization Tool
 Los Alamos National Laboratory I/O Visualization Tool Final Design Report Chaii Lee & Danny Morris June 18, 2008 Abstract Los Alamos National Laboratory is a leading national security research institution
Los Alamos National Laboratory I/O Visualization Tool Final Design Report Chaii Lee & Danny Morris June 18, 2008 Abstract Los Alamos National Laboratory is a leading national security research institution
Applications. Foreground / background segmentation Finding skin-colored regions. Finding the moving objects. Intelligent scissors
 Segmentation I Goal Separate image into coherent regions Berkeley segmentation database: http://www.eecs.berkeley.edu/research/projects/cs/vision/grouping/segbench/ Slide by L. Lazebnik Applications Intelligent
Segmentation I Goal Separate image into coherent regions Berkeley segmentation database: http://www.eecs.berkeley.edu/research/projects/cs/vision/grouping/segbench/ Slide by L. Lazebnik Applications Intelligent
