Slurm and Abel job scripts. Katerina Michalickova The Research Computing Services Group SUF/USIT October 23, 2012
|
|
|
- Kerrie Sanders
- 6 years ago
- Views:
Transcription
1 Slurm and Abel job scripts Katerina Michalickova The Research Computing Services Group SUF/USIT October 23, 2012
2 Abel in numbers Nodes Cores (1 node->2 processors->16 cores) Total memory - ~40 TB (typically 64 GB per node) Total storage - ~400TB InfiniBand interconnect on all nodes (FDR) # 96 at top500.org Read more at
3 Topics queuing system job administration user administration software job scripts examples simple scratch arrayrun parallel jobs OpenMP MPI
4 Queuing system Lets you specify resources that your program needs. Keeps track of which resources are available on which nodes, and starts your job when the requested resources are available. On Abel, we use the Simple Linux Utility for Resource Management - SLURM A job is started by sending a shell-script to slurm with the command sbatch. Resources are requested by special comments in the shell-script (#SBATCH --).
5 Interactive use of Abel Abel is used through the queuing system. It is not allowed to run jobs directly on the login nodes (the nodes you are on when you do ssh abel.uio.no). The login nodes are just for logging in, copying files, editing, compiling, running short tests (no more than a couple of minutes), submitting jobs, checking job status, etc. If interactive login is needed, use qlogin.
6 Ask SLURM for the right resources Project Memory Time Queue Disk CPUS Nodes Combination thereof Constraints (communication and special features) Files
7 sbatch - project #SBATCH --account=project Specify the project to run under. Every Abel user is assigned a project. Use command projects to find out which project you belong to. UiO scientists/students can use the uio project It is recommended to seek additional resources if planning intensive work. Application for compute hours and data storage can be placed with the Norwegian metacenter for computational science (NOTUR) #SBATCH --job-name=jobname Job name
8 sbatch - memory #SBATCH --mem-per-cpu=size Memory required per allocated core (format: 2G or 2000M) How much memory should one specify? The maximum usage of RAM by your program (plus some). Exaggerated values might, delay the job start. Coming later #SBATCH --partition=hugemem If you need more than 64GB of RAM on a single node. Currently not many nodes available with this feature.
9 mem-per-cpu - top maximum usage of virtual RAM by your program
10 sbatch - time #SBATCH --time=hh:mm:ss Wall clock time limit on the job Some prior testing is necessary. One might, for example, test on smaller data sets and extrapolate. As with the memory, unnecessarily large values might delay the job start. #SBATCH --begin=hh:mm:ss Start the job at a given time (or later) Maximum time for a job is 1 week (168 hours). If more needed, use --partition=long
11 sbatch CPUs and nodes Does your program support more than one CPU? If so, do they have to be on a single node? How many CPUs will the program run efficiently on? #SBATCH --nodes=nodes Number of nodes to allocate #SBATCH --ntasks-per-node=cores Number of cores to allocate within each allocated node #SBATCH --ntasks=cores Number of cores to allocate
12 sbatch CPUs and nodes If you just need some cpus, no matter where: #SBATCH --ntasks=17 If you need a specific number of cpus on each node #SBATCH --nodes=8 --ntasks-per-node=4 If you need the cpu's on a single node #SBATCH --nodes=1 --ntasks-per-node=8
13 sbatch - interconnect #SBATCH --constraint=ib Run job on nodes with infiniband Gigabit Ethernet on all nodes All nodes on Abel are equipped with InfiniBand (56 Gbits/s) Selected automatically if you run MPI jobs
14 sbatch - contraints #SBATCH --constraint=feature Run job on nodes with a certain feature - ib, rackn. #SBATCH --constraint=ib&rack21 If you need more than one constraint in case of multiple specifications, the later overrides the earlier
15 sbatch - files #SBATCH --output=file Send 'stdout' (and stderr) to the specified file (instead of slurmxxx.out) #SBATCH --error=file Send 'stderr' to the specified file #SBATCH --input=file Read 'stdin' from the specified file
16 sbatch low priority #SBATCH --qos=lowpri Run a job in the lowpri queue Even if all of your project's cpus are busy, you may utilize other cpus Such a job may be terminated and put back into the queue at any time. If possible, your job should ensure its state is saved regularly, and should be prepared to pick up on where it left off.
17 sbatch - restart If for some reason you want your job to be restarted, you may use the following line in your script. touch $SCRATCH/.restart This will ensure your job is put back in the queue when it terminates.
18 Inside the job script All jobs must start with the bash-command: source /cluster/bin/jobsetup A job-specific scratch-directory is created for you on /work partition. The path is in the environment variable $SCRATCH. We recommend using this directory especially if your job is IO intensive. You can copy results back to your home-directory when the job exits using chkfile in your script.
19 Environment variables SLURM_JOBID job-id of the job SCRATCH name of job-specific scratch-area SLURM_NPROCS total number of cpus requested SLURM_CPUS_ON_NODE number of cpus allocated on node SUBMITDIR directory where sbatch was issued TASK_ID task number (for arrayrun-jobs)
20 Job administration cancel a job see job details see the queue see the projects
21 Cancel a job - scancel scancel jobid # scancel --user=me scancel --account=xxx Cancel a job Cancel all your jobs Cancel all jobs in project xxx
22 Job details - scontrol show job
23 See the queue - squeue [-j jobids] show only the specified jobs [-w nodes] show only jobs on the specified nodes [-A projects] show only jobs belonging to the specified projects [-t states] show only jobs in the specified states (pending, running, suspended, etc.) [-u users] show only jobs belonging to the specified users All specifications can be comma separated lists Examples: squeue j 4132,4133 shows jobs 4132 and 4133 squeue -w compute shows jobs running on compute squeue -u foo -t PD shows pending jobs belonging to user 'foo' squeue -A bar shows all jobs in the project 'bar'
24
25 --nonzero --pe --memory --group See the projects - qsumm only show accounts with at least one running or pending job show processor equivalents (PEs) instead of CPUs show memory usage instead of CPUs do not show the individual Notur and Grid accounts --user=username only count jobs belonging to username --help show all options
26
27 User administration - project and cost
28 User s disk space dusage (coming soon)
29 Interactive use of Abel - qlogin Send request for a resource Join the queue Work on command line when resource becomes available Book one node (or 16 cores) on Abel for your interactive use for 1 hour: qlogin --account=your_project --ntasks-per-node=16 --time=01:00:00 Run source /cluster/bin/jobsetup after receiving allocation For a more info, see:
30 Software on Abel Available on Abel: Software on Abel is organized in modules. List all software (and version) organized in modules: module avail Load software from a module: module load module_name If you cannot find what you looking for: ask us
31 Job script Your program joins the queue via a job script Job script - shell script with keywords in comments read by the queuing system Compulsory keywords: #SBATCH --account #SBATCH --time #SBATCH --mem-per-cpu Setting up a job environment source /cluster/bin/jobsetup
32 Minimal job script #!/bin/bash # Job name: #SBATCH --job-name=jobname # Project: #SBATCH --account=uio # Wall time: #SBATCH --time=hh:mm:ss # Max memory #SBATCH --mem-per-cpu=max_size_in_memory # Set up environment source /cluster/bin/jobsetup # Run command./executable > outfile
33 Use of the SCRATCH area #!/bin/sh #SBATCH --job-name=yourjobname #SBATCH --account=yourproject #SBATCH --time=hh:mm:ss #SBATCH --mem-per-cpu=max_size_in_memory source /cluster/bin/jobsetup ## Copy files to work directory: cp $SUBMITDIR/YourDatafile $SCRATCH ## Mark outfiles for automatic copying to $SUBMITDIR: chkfile YourOutputfile ## Run command cd $SCRATCH executable YourDatafile > YourOutputfile
34 Strength of cluster computing Large problems (or parts of) can be divided into smaller tasks and executed in parallel Types of parallel applications: Divide input data and execute your program on all subsets (array run) Execute parts of your program in parallel (MPI or OpenMP programming)
35 Arrayrun To run many instances of the same job, use an arrayrun command Special concern organizing input and output files so they do not overwrite each other
36 TASK_ID variable TASK_ID is an environment variable, it can be accessed by all scripts during the execution of arrayrun 1 st run TASK_ID = 1 2 nd run TASK_ID = 2 N th run TASK_ID = N TASK_ID is used to organize input and output files Accesing the value of TASK_ID variable: In shell script : $TASK_ID In perl script: ENV{TASK_ID}
37 Arrayrun job script worker script #!/bin/sh #SBATCH --account=yourproject #SBATCH --time=hh:mm:ss #SBATCH --mem-per-cpu=max_size_in_memory #SBATCH --partition=lowpri source /cluster/bin/jobsetup DATASET=dataset.$TASK_ID OUTFILE=result.$TASK_ID cp $SUBMITDIR/$DATASET $SCRATCH chkfile $OUTFILE cd $SCRATCH executable $DATASET > $OUTFILE
38 Arrayrun job script submit script #!/bin/sh #SBATCH --account=yourproject #SBATCH --time=hh:mm:ss (longer than worker script) #SBATCH --mem-per-cpu=max_size_in_memory (low) source /cluster/bin/jobsetup arrayrun workerscript 1,4,42 1, 4, , 2, 3, 4, :2 0, 2, 4, 6, 8, 10 32,56, , 56, 100, 101, 102,..., 200!no spaces, decimals, negative numbers
39 Array run example BLAST - sequence similarity search program Input biological sequences ftp://ftp.ncbi.nih.gov/genomes/influenza/influenza.faa Database of sequences ftp://ftp.ncbi.nih.gov/blast/db/
40 Array run example 2 Output sequence matches probabilistic scores sequence alignments
41 Parallelizing BLAST Split the query database into chunks Perl fasta splitter
42 Abel worker script
43 Abel submit script
44
45 In the process.
46
47 Parallel jobs on Abel start Two kinds of parallel jobs Single node OpenMP serial Init parallel env. Terminate parallel env. Multiple nodes MPI serial end
48 Single node Shared memory is possible Threads OpenMP Message passing MPI
49 OpenMP job script OpenMP]$ cat hello.run #!/bin/bash #SBATCH --account=staff #SBATCH --time=00:01:00 #SBATCH --mem-per-cpu=100m #SBATCH --ntasks-per-node=4 --nodes=1 source /cluster/bin/jobsetup export OMP_NUM_THREADS=$SLURM_CPUS_ON_NODE./hello.x
50 Multiple nodes Distributed memory Message passing, MPI
51 MPI on Abel we support Open MPI module load openmpi use mpicc and mpif90 as compilers use the same MPI module for compilation and execution read
52 MPI job script #!/bin/bash #SBATCH --account=staff #SBATCH --time=0:01:0 #SBATCH --mem-per-cpu=100m #SBATCH --ntasks-per-node=1 #SBATCH --nodes=4 source /cluster/bin/jobsetup module load openmpi mpirun./hello.x
53 Thank you.
Slurm and Abel job scripts. Katerina Michalickova The Research Computing Services Group SUF/USIT November 13, 2013
 Slurm and Abel job scripts Katerina Michalickova The Research Computing Services Group SUF/USIT November 13, 2013 Abel in numbers Nodes - 600+ Cores - 10000+ (1 node->2 processors->16 cores) Total memory
Slurm and Abel job scripts Katerina Michalickova The Research Computing Services Group SUF/USIT November 13, 2013 Abel in numbers Nodes - 600+ Cores - 10000+ (1 node->2 processors->16 cores) Total memory
Exercises: Abel/Colossus and SLURM
 Exercises: Abel/Colossus and SLURM November 08, 2016 Sabry Razick The Research Computing Services Group, USIT Topics Get access Running a simple job Job script Running a simple job -- qlogin Customize
Exercises: Abel/Colossus and SLURM November 08, 2016 Sabry Razick The Research Computing Services Group, USIT Topics Get access Running a simple job Job script Running a simple job -- qlogin Customize
Introduction to Abel/Colossus and the queuing system
 Introduction to Abel/Colossus and the queuing system November 14, 2018 Sabry Razick Research Infrastructure Services Group, USIT Topics First 7 slides are about us and links The Research Computing Services
Introduction to Abel/Colossus and the queuing system November 14, 2018 Sabry Razick Research Infrastructure Services Group, USIT Topics First 7 slides are about us and links The Research Computing Services
High Performance Computing Cluster Advanced course
 High Performance Computing Cluster Advanced course Jeremie Vandenplas, Gwen Dawes 9 November 2017 Outline Introduction to the Agrogenomics HPC Submitting and monitoring jobs on the HPC Parallel jobs on
High Performance Computing Cluster Advanced course Jeremie Vandenplas, Gwen Dawes 9 November 2017 Outline Introduction to the Agrogenomics HPC Submitting and monitoring jobs on the HPC Parallel jobs on
High Performance Computing Cluster Basic course
 High Performance Computing Cluster Basic course Jeremie Vandenplas, Gwen Dawes 30 October 2017 Outline Introduction to the Agrogenomics HPC Connecting with Secure Shell to the HPC Introduction to the Unix/Linux
High Performance Computing Cluster Basic course Jeremie Vandenplas, Gwen Dawes 30 October 2017 Outline Introduction to the Agrogenomics HPC Connecting with Secure Shell to the HPC Introduction to the Unix/Linux
Graham vs legacy systems
 New User Seminar Graham vs legacy systems This webinar only covers topics pertaining to graham. For the introduction to our legacy systems (Orca etc.), please check the following recorded webinar: SHARCNet
New User Seminar Graham vs legacy systems This webinar only covers topics pertaining to graham. For the introduction to our legacy systems (Orca etc.), please check the following recorded webinar: SHARCNet
Introduction to SLURM & SLURM batch scripts
 Introduction to SLURM & SLURM batch scripts Anita Orendt Assistant Director Research Consulting & Faculty Engagement anita.orendt@utah.edu 6 February 2018 Overview of Talk Basic SLURM commands SLURM batch
Introduction to SLURM & SLURM batch scripts Anita Orendt Assistant Director Research Consulting & Faculty Engagement anita.orendt@utah.edu 6 February 2018 Overview of Talk Basic SLURM commands SLURM batch
Introduction to SLURM & SLURM batch scripts
 Introduction to SLURM & SLURM batch scripts Anita Orendt Assistant Director Research Consulting & Faculty Engagement anita.orendt@utah.edu 16 Feb 2017 Overview of Talk Basic SLURM commands SLURM batch
Introduction to SLURM & SLURM batch scripts Anita Orendt Assistant Director Research Consulting & Faculty Engagement anita.orendt@utah.edu 16 Feb 2017 Overview of Talk Basic SLURM commands SLURM batch
Introduction to High-Performance Computing (HPC)
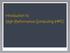 Introduction to High-Performance Computing (HPC) Computer components CPU : Central Processing Unit cores : individual processing units within a CPU Storage : Disk drives HDD : Hard Disk Drive SSD : Solid
Introduction to High-Performance Computing (HPC) Computer components CPU : Central Processing Unit cores : individual processing units within a CPU Storage : Disk drives HDD : Hard Disk Drive SSD : Solid
Introduction to SLURM on the High Performance Cluster at the Center for Computational Research
 Introduction to SLURM on the High Performance Cluster at the Center for Computational Research Cynthia Cornelius Center for Computational Research University at Buffalo, SUNY 701 Ellicott St Buffalo, NY
Introduction to SLURM on the High Performance Cluster at the Center for Computational Research Cynthia Cornelius Center for Computational Research University at Buffalo, SUNY 701 Ellicott St Buffalo, NY
Duke Compute Cluster Workshop. 3/28/2018 Tom Milledge rc.duke.edu
 Duke Compute Cluster Workshop 3/28/2018 Tom Milledge rc.duke.edu rescomputing@duke.edu Outline of talk Overview of Research Computing resources Duke Compute Cluster overview Running interactive and batch
Duke Compute Cluster Workshop 3/28/2018 Tom Milledge rc.duke.edu rescomputing@duke.edu Outline of talk Overview of Research Computing resources Duke Compute Cluster overview Running interactive and batch
Introduction to SLURM & SLURM batch scripts
 Introduction to SLURM & SLURM batch scripts Anita Orendt Assistant Director Research Consulting & Faculty Engagement anita.orendt@utah.edu 23 June 2016 Overview of Talk Basic SLURM commands SLURM batch
Introduction to SLURM & SLURM batch scripts Anita Orendt Assistant Director Research Consulting & Faculty Engagement anita.orendt@utah.edu 23 June 2016 Overview of Talk Basic SLURM commands SLURM batch
Duke Compute Cluster Workshop. 11/10/2016 Tom Milledge h:ps://rc.duke.edu/
 Duke Compute Cluster Workshop 11/10/2016 Tom Milledge h:ps://rc.duke.edu/ rescompu>ng@duke.edu Outline of talk Overview of Research Compu>ng resources Duke Compute Cluster overview Running interac>ve and
Duke Compute Cluster Workshop 11/10/2016 Tom Milledge h:ps://rc.duke.edu/ rescompu>ng@duke.edu Outline of talk Overview of Research Compu>ng resources Duke Compute Cluster overview Running interac>ve and
How to run a job on a Cluster?
 How to run a job on a Cluster? Cluster Training Workshop Dr Samuel Kortas Computational Scientist KAUST Supercomputing Laboratory Samuel.kortas@kaust.edu.sa 17 October 2017 Outline 1. Resources available
How to run a job on a Cluster? Cluster Training Workshop Dr Samuel Kortas Computational Scientist KAUST Supercomputing Laboratory Samuel.kortas@kaust.edu.sa 17 October 2017 Outline 1. Resources available
Introduction to the NCAR HPC Systems. 25 May 2018 Consulting Services Group Brian Vanderwende
 Introduction to the NCAR HPC Systems 25 May 2018 Consulting Services Group Brian Vanderwende Topics to cover Overview of the NCAR cluster resources Basic tasks in the HPC environment Accessing pre-built
Introduction to the NCAR HPC Systems 25 May 2018 Consulting Services Group Brian Vanderwende Topics to cover Overview of the NCAR cluster resources Basic tasks in the HPC environment Accessing pre-built
Submitting and running jobs on PlaFRIM2 Redouane Bouchouirbat
 Submitting and running jobs on PlaFRIM2 Redouane Bouchouirbat Summary 1. Submitting Jobs: Batch mode - Interactive mode 2. Partition 3. Jobs: Serial, Parallel 4. Using generic resources Gres : GPUs, MICs.
Submitting and running jobs on PlaFRIM2 Redouane Bouchouirbat Summary 1. Submitting Jobs: Batch mode - Interactive mode 2. Partition 3. Jobs: Serial, Parallel 4. Using generic resources Gres : GPUs, MICs.
June Workshop Series June 27th: All About SLURM University of Nebraska Lincoln Holland Computing Center. Carrie Brown, Adam Caprez
 June Workshop Series June 27th: All About SLURM University of Nebraska Lincoln Holland Computing Center Carrie Brown, Adam Caprez Setup Instructions Please complete these steps before the lessons start
June Workshop Series June 27th: All About SLURM University of Nebraska Lincoln Holland Computing Center Carrie Brown, Adam Caprez Setup Instructions Please complete these steps before the lessons start
An Introduction to Gauss. Paul D. Baines University of California, Davis November 20 th 2012
 An Introduction to Gauss Paul D. Baines University of California, Davis November 20 th 2012 What is Gauss? * http://wiki.cse.ucdavis.edu/support:systems:gauss * 12 node compute cluster (2 x 16 cores per
An Introduction to Gauss Paul D. Baines University of California, Davis November 20 th 2012 What is Gauss? * http://wiki.cse.ucdavis.edu/support:systems:gauss * 12 node compute cluster (2 x 16 cores per
Duke Compute Cluster Workshop. 10/04/2018 Tom Milledge rc.duke.edu
 Duke Compute Cluster Workshop 10/04/2018 Tom Milledge rc.duke.edu rescomputing@duke.edu Outline of talk Overview of Research Computing resources Duke Compute Cluster overview Running interactive and batch
Duke Compute Cluster Workshop 10/04/2018 Tom Milledge rc.duke.edu rescomputing@duke.edu Outline of talk Overview of Research Computing resources Duke Compute Cluster overview Running interactive and batch
Slurm basics. Summer Kickstart June slide 1 of 49
 Slurm basics Summer Kickstart 2017 June 2017 slide 1 of 49 Triton layers Triton is a powerful but complex machine. You have to consider: Connecting (ssh) Data storage (filesystems and Lustre) Resource
Slurm basics Summer Kickstart 2017 June 2017 slide 1 of 49 Triton layers Triton is a powerful but complex machine. You have to consider: Connecting (ssh) Data storage (filesystems and Lustre) Resource
Submitting batch jobs
 Submitting batch jobs SLURM on ECGATE Xavi Abellan Xavier.Abellan@ecmwf.int ECMWF February 20, 2017 Outline Interactive mode versus Batch mode Overview of the Slurm batch system on ecgate Batch basic concepts
Submitting batch jobs SLURM on ECGATE Xavi Abellan Xavier.Abellan@ecmwf.int ECMWF February 20, 2017 Outline Interactive mode versus Batch mode Overview of the Slurm batch system on ecgate Batch basic concepts
Using a Linux System 6
 Canaan User Guide Connecting to the Cluster 1 SSH (Secure Shell) 1 Starting an ssh session from a Mac or Linux system 1 Starting an ssh session from a Windows PC 1 Once you're connected... 1 Ending an
Canaan User Guide Connecting to the Cluster 1 SSH (Secure Shell) 1 Starting an ssh session from a Mac or Linux system 1 Starting an ssh session from a Windows PC 1 Once you're connected... 1 Ending an
OBTAINING AN ACCOUNT:
 HPC Usage Policies The IIA High Performance Computing (HPC) System is managed by the Computer Management Committee. The User Policies here were developed by the Committee. The user policies below aim to
HPC Usage Policies The IIA High Performance Computing (HPC) System is managed by the Computer Management Committee. The User Policies here were developed by the Committee. The user policies below aim to
Sherlock for IBIIS. William Law Stanford Research Computing
 Sherlock for IBIIS William Law Stanford Research Computing Overview How we can help System overview Tech specs Signing on Batch submission Software environment Interactive jobs Next steps We are here to
Sherlock for IBIIS William Law Stanford Research Computing Overview How we can help System overview Tech specs Signing on Batch submission Software environment Interactive jobs Next steps We are here to
Introduction to GACRC Teaching Cluster
 Introduction to GACRC Teaching Cluster Georgia Advanced Computing Resource Center (GACRC) EITS/University of Georgia Zhuofei Hou zhuofei@uga.edu 1 Outline GACRC Overview Computing Resources Three Folders
Introduction to GACRC Teaching Cluster Georgia Advanced Computing Resource Center (GACRC) EITS/University of Georgia Zhuofei Hou zhuofei@uga.edu 1 Outline GACRC Overview Computing Resources Three Folders
How to Use a Supercomputer - A Boot Camp
 How to Use a Supercomputer - A Boot Camp Shelley Knuth Peter Ruprecht shelley.knuth@colorado.edu peter.ruprecht@colorado.edu www.rc.colorado.edu Outline Today we will discuss: Who Research Computing is
How to Use a Supercomputer - A Boot Camp Shelley Knuth Peter Ruprecht shelley.knuth@colorado.edu peter.ruprecht@colorado.edu www.rc.colorado.edu Outline Today we will discuss: Who Research Computing is
Choosing Resources Wisely Plamen Krastev Office: 38 Oxford, Room 117 FAS Research Computing
 Choosing Resources Wisely Plamen Krastev Office: 38 Oxford, Room 117 Email:plamenkrastev@fas.harvard.edu Objectives Inform you of available computational resources Help you choose appropriate computational
Choosing Resources Wisely Plamen Krastev Office: 38 Oxford, Room 117 Email:plamenkrastev@fas.harvard.edu Objectives Inform you of available computational resources Help you choose appropriate computational
HPC Introductory Course - Exercises
 HPC Introductory Course - Exercises The exercises in the following sections will guide you understand and become more familiar with how to use the Balena HPC service. Lines which start with $ are commands
HPC Introductory Course - Exercises The exercises in the following sections will guide you understand and become more familiar with how to use the Balena HPC service. Lines which start with $ are commands
Batch Systems & Parallel Application Launchers Running your jobs on an HPC machine
 Batch Systems & Parallel Application Launchers Running your jobs on an HPC machine Partners Funding Reusing this material This work is licensed under a Creative Commons Attribution- NonCommercial-ShareAlike
Batch Systems & Parallel Application Launchers Running your jobs on an HPC machine Partners Funding Reusing this material This work is licensed under a Creative Commons Attribution- NonCommercial-ShareAlike
Submitting batch jobs Slurm on ecgate Solutions to the practicals
 Submitting batch jobs Slurm on ecgate Solutions to the practicals Xavi Abellan xavier.abellan@ecmwf.int User Support Section Com Intro 2015 Submitting batch jobs ECMWF 2015 Slide 1 Practical 1: Basic job
Submitting batch jobs Slurm on ecgate Solutions to the practicals Xavi Abellan xavier.abellan@ecmwf.int User Support Section Com Intro 2015 Submitting batch jobs ECMWF 2015 Slide 1 Practical 1: Basic job
Introduction to GACRC Teaching Cluster PHYS8602
 Introduction to GACRC Teaching Cluster PHYS8602 Georgia Advanced Computing Resource Center (GACRC) EITS/University of Georgia Zhuofei Hou zhuofei@uga.edu 1 Outline GACRC Overview Computing Resources Three
Introduction to GACRC Teaching Cluster PHYS8602 Georgia Advanced Computing Resource Center (GACRC) EITS/University of Georgia Zhuofei Hou zhuofei@uga.edu 1 Outline GACRC Overview Computing Resources Three
Introduction to GACRC Teaching Cluster
 Introduction to GACRC Teaching Cluster Georgia Advanced Computing Resource Center (GACRC) EITS/University of Georgia Zhuofei Hou zhuofei@uga.edu 1 Outline GACRC Overview Computing Resources Three Folders
Introduction to GACRC Teaching Cluster Georgia Advanced Computing Resource Center (GACRC) EITS/University of Georgia Zhuofei Hou zhuofei@uga.edu 1 Outline GACRC Overview Computing Resources Three Folders
P a g e 1. HPC Example for C with OpenMPI
 P a g e 1 HPC Example for C with OpenMPI Revision History Version Date Prepared By Summary of Changes 1.0 Jul 3, 2017 Raymond Tsang Initial release 1.1 Jul 24, 2018 Ray Cheung Minor change HPC Example
P a g e 1 HPC Example for C with OpenMPI Revision History Version Date Prepared By Summary of Changes 1.0 Jul 3, 2017 Raymond Tsang Initial release 1.1 Jul 24, 2018 Ray Cheung Minor change HPC Example
RHRK-Seminar. High Performance Computing with the Cluster Elwetritsch - II. Course instructor : Dr. Josef Schüle, RHRK
 RHRK-Seminar High Performance Computing with the Cluster Elwetritsch - II Course instructor : Dr. Josef Schüle, RHRK Overview Course I Login to cluster SSH RDP / NX Desktop Environments GNOME (default)
RHRK-Seminar High Performance Computing with the Cluster Elwetritsch - II Course instructor : Dr. Josef Schüle, RHRK Overview Course I Login to cluster SSH RDP / NX Desktop Environments GNOME (default)
Slurm at UPPMAX. How to submit jobs with our queueing system. Jessica Nettelblad sysadmin at UPPMAX
 Slurm at UPPMAX How to submit jobs with our queueing system Jessica Nettelblad sysadmin at UPPMAX Slurm at UPPMAX Intro Queueing with Slurm How to submit jobs Testing How to test your scripts before submission
Slurm at UPPMAX How to submit jobs with our queueing system Jessica Nettelblad sysadmin at UPPMAX Slurm at UPPMAX Intro Queueing with Slurm How to submit jobs Testing How to test your scripts before submission
Introduction to UBELIX
 Science IT Support (ScITS) Michael Rolli, Nico Färber Informatikdienste Universität Bern 06.06.2017, Introduction to UBELIX Agenda > Introduction to UBELIX (Overview only) Other topics spread in > Introducing
Science IT Support (ScITS) Michael Rolli, Nico Färber Informatikdienste Universität Bern 06.06.2017, Introduction to UBELIX Agenda > Introduction to UBELIX (Overview only) Other topics spread in > Introducing
Batch Usage on JURECA Introduction to Slurm. May 2016 Chrysovalantis Paschoulas HPS JSC
 Batch Usage on JURECA Introduction to Slurm May 2016 Chrysovalantis Paschoulas HPS group @ JSC Batch System Concepts Resource Manager is the software responsible for managing the resources of a cluster,
Batch Usage on JURECA Introduction to Slurm May 2016 Chrysovalantis Paschoulas HPS group @ JSC Batch System Concepts Resource Manager is the software responsible for managing the resources of a cluster,
Introduction to Joker Cyber Infrastructure Architecture Team CIA.NMSU.EDU
 Introduction to Joker Cyber Infrastructure Architecture Team CIA.NMSU.EDU What is Joker? NMSU s supercomputer. 238 core computer cluster. Intel E-5 Xeon CPUs and Nvidia K-40 GPUs. InfiniBand innerconnect.
Introduction to Joker Cyber Infrastructure Architecture Team CIA.NMSU.EDU What is Joker? NMSU s supercomputer. 238 core computer cluster. Intel E-5 Xeon CPUs and Nvidia K-40 GPUs. InfiniBand innerconnect.
Introduction to HPC Using zcluster at GACRC
 Introduction to HPC Using zcluster at GACRC On-class STAT8330 Georgia Advanced Computing Resource Center University of Georgia Suchitra Pakala pakala@uga.edu Slides courtesy: Zhoufei Hou 1 Outline What
Introduction to HPC Using zcluster at GACRC On-class STAT8330 Georgia Advanced Computing Resource Center University of Georgia Suchitra Pakala pakala@uga.edu Slides courtesy: Zhoufei Hou 1 Outline What
Choosing Resources Wisely. What is Research Computing?
 Choosing Resources Wisely Scott Yockel, PhD Harvard - Research Computing What is Research Computing? Faculty of Arts and Sciences (FAS) department that handles nonenterprise IT requests from researchers.
Choosing Resources Wisely Scott Yockel, PhD Harvard - Research Computing What is Research Computing? Faculty of Arts and Sciences (FAS) department that handles nonenterprise IT requests from researchers.
Submitting batch jobs Slurm on ecgate
 Submitting batch jobs Slurm on ecgate Xavi Abellan xavier.abellan@ecmwf.int User Support Section Com Intro 2015 Submitting batch jobs ECMWF 2015 Slide 1 Outline Interactive mode versus Batch mode Overview
Submitting batch jobs Slurm on ecgate Xavi Abellan xavier.abellan@ecmwf.int User Support Section Com Intro 2015 Submitting batch jobs ECMWF 2015 Slide 1 Outline Interactive mode versus Batch mode Overview
Introduction to GALILEO
 Introduction to GALILEO Parallel & production environment Mirko Cestari m.cestari@cineca.it Alessandro Marani a.marani@cineca.it Domenico Guida d.guida@cineca.it Maurizio Cremonesi m.cremonesi@cineca.it
Introduction to GALILEO Parallel & production environment Mirko Cestari m.cestari@cineca.it Alessandro Marani a.marani@cineca.it Domenico Guida d.guida@cineca.it Maurizio Cremonesi m.cremonesi@cineca.it
Using Compute Canada. Masao Fujinaga Information Services and Technology University of Alberta
 Using Compute Canada Masao Fujinaga Information Services and Technology University of Alberta Introduction to cedar batch system jobs are queued priority depends on allocation and past usage Cedar Nodes
Using Compute Canada Masao Fujinaga Information Services and Technology University of Alberta Introduction to cedar batch system jobs are queued priority depends on allocation and past usage Cedar Nodes
Heterogeneous Job Support
 Heterogeneous Job Support Tim Wickberg SchedMD SC17 Submitting Jobs Multiple independent job specifications identified in command line using : separator The job specifications are sent to slurmctld daemon
Heterogeneous Job Support Tim Wickberg SchedMD SC17 Submitting Jobs Multiple independent job specifications identified in command line using : separator The job specifications are sent to slurmctld daemon
Our new HPC-Cluster An overview
 Our new HPC-Cluster An overview Christian Hagen Universität Regensburg Regensburg, 15.05.2009 Outline 1 Layout 2 Hardware 3 Software 4 Getting an account 5 Compiling 6 Queueing system 7 Parallelization
Our new HPC-Cluster An overview Christian Hagen Universität Regensburg Regensburg, 15.05.2009 Outline 1 Layout 2 Hardware 3 Software 4 Getting an account 5 Compiling 6 Queueing system 7 Parallelization
Power9 CTE User s Guide
 Power9 CTE User s Guide Barcelona Supercomputing Center Copyright c 2017 BSC-CNS June 25, 2018 Contents 1 Introduction 2 2 System Overview 2 3 Compiling applications 2 4 Connecting to CTE-POWER 2 4.1 Password
Power9 CTE User s Guide Barcelona Supercomputing Center Copyright c 2017 BSC-CNS June 25, 2018 Contents 1 Introduction 2 2 System Overview 2 3 Compiling applications 2 4 Connecting to CTE-POWER 2 4.1 Password
Kimmo Mattila Ari-Matti Sarén. CSC Bioweek Computing intensive bioinformatics analysis on Taito
 Kimmo Mattila Ari-Matti Sarén CSC Bioweek 2018 Computing intensive bioinformatics analysis on Taito 7. 2. 2018 CSC Environment Sisu Cray XC40 Massively Parallel Processor (MPP) supercomputer 3376 12-core
Kimmo Mattila Ari-Matti Sarén CSC Bioweek 2018 Computing intensive bioinformatics analysis on Taito 7. 2. 2018 CSC Environment Sisu Cray XC40 Massively Parallel Processor (MPP) supercomputer 3376 12-core
Beginner's Guide for UK IBM systems
 Beginner's Guide for UK IBM systems This document is intended to provide some basic guidelines for those who already had certain programming knowledge with high level computer languages (e.g. Fortran,
Beginner's Guide for UK IBM systems This document is intended to provide some basic guidelines for those who already had certain programming knowledge with high level computer languages (e.g. Fortran,
Running Galaxy in an HPC environment requirements, challenges and some solutions : the LIFEPORTAL
 Running Galaxy in an HPC environment requirements, challenges and some solutions : the LIFEPORTAL Nikolay Vazov University Center for Information Technologies University of Oslo https://lifeportal.uio.no
Running Galaxy in an HPC environment requirements, challenges and some solutions : the LIFEPORTAL Nikolay Vazov University Center for Information Technologies University of Oslo https://lifeportal.uio.no
High Performance Computing (HPC) Using zcluster at GACRC
 High Performance Computing (HPC) Using zcluster at GACRC On-class STAT8060 Georgia Advanced Computing Resource Center University of Georgia Zhuofei Hou, HPC Trainer zhuofei@uga.edu Outline What is GACRC?
High Performance Computing (HPC) Using zcluster at GACRC On-class STAT8060 Georgia Advanced Computing Resource Center University of Georgia Zhuofei Hou, HPC Trainer zhuofei@uga.edu Outline What is GACRC?
For Dr Landau s PHYS8602 course
 For Dr Landau s PHYS8602 course Shan-Ho Tsai (shtsai@uga.edu) Georgia Advanced Computing Resource Center - GACRC January 7, 2019 You will be given a student account on the GACRC s Teaching cluster. Your
For Dr Landau s PHYS8602 course Shan-Ho Tsai (shtsai@uga.edu) Georgia Advanced Computing Resource Center - GACRC January 7, 2019 You will be given a student account on the GACRC s Teaching cluster. Your
ECE 574 Cluster Computing Lecture 4
 ECE 574 Cluster Computing Lecture 4 Vince Weaver http://web.eece.maine.edu/~vweaver vincent.weaver@maine.edu 31 January 2017 Announcements Don t forget about homework #3 I ran HPCG benchmark on Haswell-EP
ECE 574 Cluster Computing Lecture 4 Vince Weaver http://web.eece.maine.edu/~vweaver vincent.weaver@maine.edu 31 January 2017 Announcements Don t forget about homework #3 I ran HPCG benchmark on Haswell-EP
How to access Geyser and Caldera from Cheyenne. 19 December 2017 Consulting Services Group Brian Vanderwende
 How to access Geyser and Caldera from Cheyenne 19 December 2017 Consulting Services Group Brian Vanderwende Geyser nodes useful for large-scale data analysis and post-processing tasks 16 nodes with: 40
How to access Geyser and Caldera from Cheyenne 19 December 2017 Consulting Services Group Brian Vanderwende Geyser nodes useful for large-scale data analysis and post-processing tasks 16 nodes with: 40
Introduction to HPC Using zcluster at GACRC
 Introduction to HPC Using zcluster at GACRC Georgia Advanced Computing Resource Center University of Georgia Zhuofei Hou, HPC Trainer zhuofei@uga.edu Outline What is GACRC? What is HPC Concept? What is
Introduction to HPC Using zcluster at GACRC Georgia Advanced Computing Resource Center University of Georgia Zhuofei Hou, HPC Trainer zhuofei@uga.edu Outline What is GACRC? What is HPC Concept? What is
Using Cartesius and Lisa. Zheng Meyer-Zhao - Consultant Clustercomputing
 Zheng Meyer-Zhao - zheng.meyer-zhao@surfsara.nl Consultant Clustercomputing Outline SURFsara About us What we do Cartesius and Lisa Architectures and Specifications File systems Funding Hands-on Logging
Zheng Meyer-Zhao - zheng.meyer-zhao@surfsara.nl Consultant Clustercomputing Outline SURFsara About us What we do Cartesius and Lisa Architectures and Specifications File systems Funding Hands-on Logging
Name Department/Research Area Have you used the Linux command line?
 Please log in with HawkID (IOWA domain) Macs are available at stations as marked To switch between the Windows and the Mac systems, press scroll lock twice 9/27/2018 1 Ben Rogers ITS-Research Services
Please log in with HawkID (IOWA domain) Macs are available at stations as marked To switch between the Windows and the Mac systems, press scroll lock twice 9/27/2018 1 Ben Rogers ITS-Research Services
Working with Shell Scripting. Daniel Balagué
 Working with Shell Scripting Daniel Balagué Editing Text Files We offer many text editors in the HPC cluster. Command-Line Interface (CLI) editors: vi / vim nano (very intuitive and easy to use if you
Working with Shell Scripting Daniel Balagué Editing Text Files We offer many text editors in the HPC cluster. Command-Line Interface (CLI) editors: vi / vim nano (very intuitive and easy to use if you
Compilation and Parallel Start
 Compiling MPI Programs Programming with MPI Compiling and running MPI programs Type to enter text Jan Thorbecke Delft University of Technology 2 Challenge the future Compiling and Starting MPI Jobs Compiling:
Compiling MPI Programs Programming with MPI Compiling and running MPI programs Type to enter text Jan Thorbecke Delft University of Technology 2 Challenge the future Compiling and Starting MPI Jobs Compiling:
TITANI CLUSTER USER MANUAL V.1.3
 2016 TITANI CLUSTER USER MANUAL V.1.3 This document is intended to give some basic notes in order to work with the TITANI High Performance Green Computing Cluster of the Civil Engineering School (ETSECCPB)
2016 TITANI CLUSTER USER MANUAL V.1.3 This document is intended to give some basic notes in order to work with the TITANI High Performance Green Computing Cluster of the Civil Engineering School (ETSECCPB)
Kamiak Cheat Sheet. Display text file, one page at a time. Matches all files beginning with myfile See disk space on volume
 Kamiak Cheat Sheet Logging in to Kamiak ssh your.name@kamiak.wsu.edu ssh -X your.name@kamiak.wsu.edu X11 forwarding Transferring Files to and from Kamiak scp -r myfile your.name@kamiak.wsu.edu:~ Copy to
Kamiak Cheat Sheet Logging in to Kamiak ssh your.name@kamiak.wsu.edu ssh -X your.name@kamiak.wsu.edu X11 forwarding Transferring Files to and from Kamiak scp -r myfile your.name@kamiak.wsu.edu:~ Copy to
Introduction to High-Performance Computing (HPC)
 Introduction to High-Performance Computing (HPC) Computer components CPU : Central Processing Unit cores : individual processing units within a CPU Storage : Disk drives HDD : Hard Disk Drive SSD : Solid
Introduction to High-Performance Computing (HPC) Computer components CPU : Central Processing Unit cores : individual processing units within a CPU Storage : Disk drives HDD : Hard Disk Drive SSD : Solid
Bash for SLURM. Author: Wesley Schaal Pharmaceutical Bioinformatics, Uppsala University
 Bash for SLURM Author: Wesley Schaal Pharmaceutical Bioinformatics, Uppsala University wesley.schaal@farmbio.uu.se Lab session: Pavlin Mitev (pavlin.mitev@kemi.uu.se) it i slides at http://uppmax.uu.se/support/courses
Bash for SLURM Author: Wesley Schaal Pharmaceutical Bioinformatics, Uppsala University wesley.schaal@farmbio.uu.se Lab session: Pavlin Mitev (pavlin.mitev@kemi.uu.se) it i slides at http://uppmax.uu.se/support/courses
Introduction to HPC Using zcluster at GACRC
 Introduction to HPC Using zcluster at GACRC On-class PBIO/BINF8350 Georgia Advanced Computing Resource Center University of Georgia Zhuofei Hou, HPC Trainer zhuofei@uga.edu Outline What is GACRC? What
Introduction to HPC Using zcluster at GACRC On-class PBIO/BINF8350 Georgia Advanced Computing Resource Center University of Georgia Zhuofei Hou, HPC Trainer zhuofei@uga.edu Outline What is GACRC? What
LaPalma User's Guide
 LaPalma User's Guide Copyright 2018 Table of contents 1. Introduction...2 2. System Overview...2 3. Online Documentation...2 3.1. Man pages...2 4. Connecting to LaPalma...2 4.1. Login node...3 4.2. Transferring
LaPalma User's Guide Copyright 2018 Table of contents 1. Introduction...2 2. System Overview...2 3. Online Documentation...2 3.1. Man pages...2 4. Connecting to LaPalma...2 4.1. Login node...3 4.2. Transferring
Batch Systems. Running your jobs on an HPC machine
 Batch Systems Running your jobs on an HPC machine Reusing this material This work is licensed under a Creative Commons Attribution- NonCommercial-ShareAlike 4.0 International License. http://creativecommons.org/licenses/by-nc-sa/4.0/deed.en_us
Batch Systems Running your jobs on an HPC machine Reusing this material This work is licensed under a Creative Commons Attribution- NonCommercial-ShareAlike 4.0 International License. http://creativecommons.org/licenses/by-nc-sa/4.0/deed.en_us
LAB. Preparing for Stampede: Programming Heterogeneous Many-Core Supercomputers
 LAB Preparing for Stampede: Programming Heterogeneous Many-Core Supercomputers Dan Stanzione, Lars Koesterke, Bill Barth, Kent Milfeld dan/lars/bbarth/milfeld@tacc.utexas.edu XSEDE 12 July 16, 2012 1 Discovery
LAB Preparing for Stampede: Programming Heterogeneous Many-Core Supercomputers Dan Stanzione, Lars Koesterke, Bill Barth, Kent Milfeld dan/lars/bbarth/milfeld@tacc.utexas.edu XSEDE 12 July 16, 2012 1 Discovery
Slurm at UPPMAX. How to submit jobs with our queueing system. Jessica Nettelblad sysadmin at UPPMAX
 Slurm at UPPMAX How to submit jobs with our queueing system Jessica Nettelblad sysadmin at UPPMAX Free! Watch! Futurama S2 Ep.4 Fry and the Slurm factory Simple Linux Utility for Resource Management Open
Slurm at UPPMAX How to submit jobs with our queueing system Jessica Nettelblad sysadmin at UPPMAX Free! Watch! Futurama S2 Ep.4 Fry and the Slurm factory Simple Linux Utility for Resource Management Open
The cluster system. Introduction 22th February Jan Saalbach Scientific Computing Group
 The cluster system Introduction 22th February 2018 Jan Saalbach Scientific Computing Group cluster-help@luis.uni-hannover.de Contents 1 General information about the compute cluster 2 Available computing
The cluster system Introduction 22th February 2018 Jan Saalbach Scientific Computing Group cluster-help@luis.uni-hannover.de Contents 1 General information about the compute cluster 2 Available computing
Compiling applications for the Cray XC
 Compiling applications for the Cray XC Compiler Driver Wrappers (1) All applications that will run in parallel on the Cray XC should be compiled with the standard language wrappers. The compiler drivers
Compiling applications for the Cray XC Compiler Driver Wrappers (1) All applications that will run in parallel on the Cray XC should be compiled with the standard language wrappers. The compiler drivers
Introduction to PICO Parallel & Production Enviroment
 Introduction to PICO Parallel & Production Enviroment Mirko Cestari m.cestari@cineca.it Alessandro Marani a.marani@cineca.it Domenico Guida d.guida@cineca.it Nicola Spallanzani n.spallanzani@cineca.it
Introduction to PICO Parallel & Production Enviroment Mirko Cestari m.cestari@cineca.it Alessandro Marani a.marani@cineca.it Domenico Guida d.guida@cineca.it Nicola Spallanzani n.spallanzani@cineca.it
Training day SLURM cluster. Context. Context renewal strategy
 Training day cluster Context Infrastructure Environment Software usage Help section For further with Best practices Support Context PRE-REQUISITE : LINUX connect to «genologin» server Basic command line
Training day cluster Context Infrastructure Environment Software usage Help section For further with Best practices Support Context PRE-REQUISITE : LINUX connect to «genologin» server Basic command line
SCALABLE HYBRID PROTOTYPE
 SCALABLE HYBRID PROTOTYPE Scalable Hybrid Prototype Part of the PRACE Technology Evaluation Objectives Enabling key applications on new architectures Familiarizing users and providing a research platform
SCALABLE HYBRID PROTOTYPE Scalable Hybrid Prototype Part of the PRACE Technology Evaluation Objectives Enabling key applications on new architectures Familiarizing users and providing a research platform
Symmetric Computing. SC 14 Jerome VIENNE
 Symmetric Computing SC 14 Jerome VIENNE viennej@tacc.utexas.edu Symmetric Computing Run MPI tasks on both MIC and host Also called heterogeneous computing Two executables are required: CPU MIC Currently
Symmetric Computing SC 14 Jerome VIENNE viennej@tacc.utexas.edu Symmetric Computing Run MPI tasks on both MIC and host Also called heterogeneous computing Two executables are required: CPU MIC Currently
Before We Start. Sign in hpcxx account slips Windows Users: Download PuTTY. Google PuTTY First result Save putty.exe to Desktop
 Before We Start Sign in hpcxx account slips Windows Users: Download PuTTY Google PuTTY First result Save putty.exe to Desktop Research Computing at Virginia Tech Advanced Research Computing Compute Resources
Before We Start Sign in hpcxx account slips Windows Users: Download PuTTY Google PuTTY First result Save putty.exe to Desktop Research Computing at Virginia Tech Advanced Research Computing Compute Resources
CRUK cluster practical sessions (SLURM) Part I processes & scripts
 CRUK cluster practical sessions (SLURM) Part I processes & scripts login Log in to the head node, clust1-headnode, using ssh and your usual user name & password. SSH Secure Shell 3.2.9 (Build 283) Copyright
CRUK cluster practical sessions (SLURM) Part I processes & scripts login Log in to the head node, clust1-headnode, using ssh and your usual user name & password. SSH Secure Shell 3.2.9 (Build 283) Copyright
An Introduction to Cluster Computing Using Newton
 An Introduction to Cluster Computing Using Newton Jason Harris and Dylan Storey March 25th, 2014 Jason Harris and Dylan Storey Introduction to Cluster Computing March 25th, 2014 1 / 26 Workshop design.
An Introduction to Cluster Computing Using Newton Jason Harris and Dylan Storey March 25th, 2014 Jason Harris and Dylan Storey Introduction to Cluster Computing March 25th, 2014 1 / 26 Workshop design.
CNAG Advanced User Training
 www.bsc.es CNAG Advanced User Training Aníbal Moreno, CNAG System Administrator Pablo Ródenas, BSC HPC Support Rubén Ramos Horta, CNAG HPC Support Barcelona,May the 5th Aim Understand CNAG s cluster design
www.bsc.es CNAG Advanced User Training Aníbal Moreno, CNAG System Administrator Pablo Ródenas, BSC HPC Support Rubén Ramos Horta, CNAG HPC Support Barcelona,May the 5th Aim Understand CNAG s cluster design
Spark Programming at Comet. UCSB CS240A Tao Yang
 Spark Programming at Comet UCSB CS240A 2016. Tao Yang Comet Cluster Comet cluster has 1944 nodes and each node has 24 cores, built on two 12-core Intel Xeon E5-2680v3 2.5 GHz processors 128 GB memory and
Spark Programming at Comet UCSB CS240A 2016. Tao Yang Comet Cluster Comet cluster has 1944 nodes and each node has 24 cores, built on two 12-core Intel Xeon E5-2680v3 2.5 GHz processors 128 GB memory and
To connect to the cluster, simply use a SSH or SFTP client to connect to:
 RIT Computer Engineering Cluster The RIT Computer Engineering cluster contains 12 computers for parallel programming using MPI. One computer, cluster-head.ce.rit.edu, serves as the master controller or
RIT Computer Engineering Cluster The RIT Computer Engineering cluster contains 12 computers for parallel programming using MPI. One computer, cluster-head.ce.rit.edu, serves as the master controller or
Using Sapelo2 Cluster at the GACRC
 Using Sapelo2 Cluster at the GACRC New User Training Workshop Georgia Advanced Computing Resource Center (GACRC) EITS/University of Georgia Zhuofei Hou zhuofei@uga.edu 1 Outline GACRC Sapelo2 Cluster Diagram
Using Sapelo2 Cluster at the GACRC New User Training Workshop Georgia Advanced Computing Resource Center (GACRC) EITS/University of Georgia Zhuofei Hou zhuofei@uga.edu 1 Outline GACRC Sapelo2 Cluster Diagram
Introduction to GALILEO
 November 27, 2016 Introduction to GALILEO Parallel & production environment Mirko Cestari m.cestari@cineca.it Alessandro Marani a.marani@cineca.it SuperComputing Applications and Innovation Department
November 27, 2016 Introduction to GALILEO Parallel & production environment Mirko Cestari m.cestari@cineca.it Alessandro Marani a.marani@cineca.it SuperComputing Applications and Innovation Department
HPC Workshop. Nov. 9, 2018 James Coyle, PhD Dir. Of High Perf. Computing
 HPC Workshop Nov. 9, 2018 James Coyle, PhD Dir. Of High Perf. Computing NEEDED EQUIPMENT 1. Laptop with Secure Shell (ssh) for login A. Windows: download/install putty from https://www.chiark.greenend.org.uk/~sgtatham/putty/latest.html
HPC Workshop Nov. 9, 2018 James Coyle, PhD Dir. Of High Perf. Computing NEEDED EQUIPMENT 1. Laptop with Secure Shell (ssh) for login A. Windows: download/install putty from https://www.chiark.greenend.org.uk/~sgtatham/putty/latest.html
Introduction to HPC Using zcluster at GACRC On-Class GENE 4220
 Introduction to HPC Using zcluster at GACRC On-Class GENE 4220 Georgia Advanced Computing Resource Center University of Georgia Suchitra Pakala pakala@uga.edu Slides courtesy: Zhoufei Hou 1 OVERVIEW GACRC
Introduction to HPC Using zcluster at GACRC On-Class GENE 4220 Georgia Advanced Computing Resource Center University of Georgia Suchitra Pakala pakala@uga.edu Slides courtesy: Zhoufei Hou 1 OVERVIEW GACRC
Cluster Clonetroop: HowTo 2014
 2014/02/25 16:53 1/13 Cluster Clonetroop: HowTo 2014 Cluster Clonetroop: HowTo 2014 This section contains information about how to access, compile and execute jobs on Clonetroop, Laboratori de Càlcul Numeric's
2014/02/25 16:53 1/13 Cluster Clonetroop: HowTo 2014 Cluster Clonetroop: HowTo 2014 This section contains information about how to access, compile and execute jobs on Clonetroop, Laboratori de Càlcul Numeric's
Intel Manycore Testing Lab (MTL) - Linux Getting Started Guide
 Intel Manycore Testing Lab (MTL) - Linux Getting Started Guide Introduction What are the intended uses of the MTL? The MTL is prioritized for supporting the Intel Academic Community for the testing, validation
Intel Manycore Testing Lab (MTL) - Linux Getting Started Guide Introduction What are the intended uses of the MTL? The MTL is prioritized for supporting the Intel Academic Community for the testing, validation
User Guide of High Performance Computing Cluster in School of Physics
 User Guide of High Performance Computing Cluster in School of Physics Prepared by Sue Yang (xue.yang@sydney.edu.au) This document aims at helping users to quickly log into the cluster, set up the software
User Guide of High Performance Computing Cluster in School of Physics Prepared by Sue Yang (xue.yang@sydney.edu.au) This document aims at helping users to quickly log into the cluster, set up the software
Introduction to GALILEO
 Introduction to GALILEO Parallel & production environment Mirko Cestari m.cestari@cineca.it Alessandro Marani a.marani@cineca.it Alessandro Grottesi a.grottesi@cineca.it SuperComputing Applications and
Introduction to GALILEO Parallel & production environment Mirko Cestari m.cestari@cineca.it Alessandro Marani a.marani@cineca.it Alessandro Grottesi a.grottesi@cineca.it SuperComputing Applications and
Minnesota Supercomputing Institute Regents of the University of Minnesota. All rights reserved.
 Minnesota Supercomputing Institute Introduction to MSI Systems Andrew Gustafson The Machines at MSI Machine Type: Cluster Source: http://en.wikipedia.org/wiki/cluster_%28computing%29 Machine Type: Cluster
Minnesota Supercomputing Institute Introduction to MSI Systems Andrew Gustafson The Machines at MSI Machine Type: Cluster Source: http://en.wikipedia.org/wiki/cluster_%28computing%29 Machine Type: Cluster
Migrating from Zcluster to Sapelo
 GACRC User Quick Guide: Migrating from Zcluster to Sapelo The GACRC Staff Version 1.0 8/4/17 1 Discussion Points I. Request Sapelo User Account II. III. IV. Systems Transfer Files Configure Software Environment
GACRC User Quick Guide: Migrating from Zcluster to Sapelo The GACRC Staff Version 1.0 8/4/17 1 Discussion Points I. Request Sapelo User Account II. III. IV. Systems Transfer Files Configure Software Environment
Using CSC Environment Efficiently,
 Using CSC Environment Efficiently, 17.09.2018 1 Exercises a) Log in to Taito either with your training or CSC user account, either from a terminal (with X11 forwarding) or using NoMachine client b) Go
Using CSC Environment Efficiently, 17.09.2018 1 Exercises a) Log in to Taito either with your training or CSC user account, either from a terminal (with X11 forwarding) or using NoMachine client b) Go
Introduction to RCC. September 14, 2016 Research Computing Center
 Introduction to HPC @ RCC September 14, 2016 Research Computing Center What is HPC High Performance Computing most generally refers to the practice of aggregating computing power in a way that delivers
Introduction to HPC @ RCC September 14, 2016 Research Computing Center What is HPC High Performance Computing most generally refers to the practice of aggregating computing power in a way that delivers
Introduction to HPC2N
 Introduction to HPC2N Birgitte Brydsø HPC2N, Umeå University 4 May 2017 1 / 24 Overview Kebnekaise and Abisko Using our systems The File System The Module System Overview Compiler Tool Chains Examples
Introduction to HPC2N Birgitte Brydsø HPC2N, Umeå University 4 May 2017 1 / 24 Overview Kebnekaise and Abisko Using our systems The File System The Module System Overview Compiler Tool Chains Examples
Introduction to RCC. January 18, 2017 Research Computing Center
 Introduction to HPC @ RCC January 18, 2017 Research Computing Center What is HPC High Performance Computing most generally refers to the practice of aggregating computing power in a way that delivers much
Introduction to HPC @ RCC January 18, 2017 Research Computing Center What is HPC High Performance Computing most generally refers to the practice of aggregating computing power in a way that delivers much
Introduction to the Cluster
 Follow us on Twitter for important news and updates: @ACCREVandy Introduction to the Cluster Advanced Computing Center for Research and Education http://www.accre.vanderbilt.edu The Cluster We will be
Follow us on Twitter for important news and updates: @ACCREVandy Introduction to the Cluster Advanced Computing Center for Research and Education http://www.accre.vanderbilt.edu The Cluster We will be
Using the SLURM Job Scheduler
 Using the SLURM Job Scheduler [web] [email] portal.biohpc.swmed.edu biohpc-help@utsouthwestern.edu 1 Updated for 2015-05-13 Overview Today we re going to cover: Part I: What is SLURM? How to use a basic
Using the SLURM Job Scheduler [web] [email] portal.biohpc.swmed.edu biohpc-help@utsouthwestern.edu 1 Updated for 2015-05-13 Overview Today we re going to cover: Part I: What is SLURM? How to use a basic
UPPMAX Introduction Martin Dahlö Valentin Georgiev
 UPPMAX Introduction 2017-11-27 Martin Dahlö martin.dahlo@scilifelab.uu.se Valentin Georgiev valentin.georgiev@icm.uu.se Objectives What is UPPMAX what it provides Projects at UPPMAX How to access UPPMAX
UPPMAX Introduction 2017-11-27 Martin Dahlö martin.dahlo@scilifelab.uu.se Valentin Georgiev valentin.georgiev@icm.uu.se Objectives What is UPPMAX what it provides Projects at UPPMAX How to access UPPMAX
ACEnet for CS6702 Ross Dickson, Computational Research Consultant 29 Sep 2009
 ACEnet for CS6702 Ross Dickson, Computational Research Consultant 29 Sep 2009 What is ACEnet? Shared resource......for research computing... physics, chemistry, oceanography, biology, math, engineering,
ACEnet for CS6702 Ross Dickson, Computational Research Consultant 29 Sep 2009 What is ACEnet? Shared resource......for research computing... physics, chemistry, oceanography, biology, math, engineering,
Basic UNIX commands. HORT Lab 2 Instructor: Kranthi Varala
 Basic UNIX commands HORT 59000 Lab 2 Instructor: Kranthi Varala Client/Server architecture User1 User2 User3 Server (UNIX/ Web/ Database etc..) User4 High Performance Compute (HPC) cluster User1 Compute
Basic UNIX commands HORT 59000 Lab 2 Instructor: Kranthi Varala Client/Server architecture User1 User2 User3 Server (UNIX/ Web/ Database etc..) User4 High Performance Compute (HPC) cluster User1 Compute
MIC Lab Parallel Computing on Stampede
 MIC Lab Parallel Computing on Stampede Aaron Birkland and Steve Lantz Cornell Center for Advanced Computing June 11 & 18, 2013 1 Interactive Launching This exercise will walk through interactively launching
MIC Lab Parallel Computing on Stampede Aaron Birkland and Steve Lantz Cornell Center for Advanced Computing June 11 & 18, 2013 1 Interactive Launching This exercise will walk through interactively launching
Introduction to High Performance Computing and an Statistical Genetics Application on the Janus Supercomputer. Purpose
 Introduction to High Performance Computing and an Statistical Genetics Application on the Janus Supercomputer Daniel Yorgov Department of Mathematical & Statistical Sciences, University of Colorado Denver
Introduction to High Performance Computing and an Statistical Genetics Application on the Janus Supercomputer Daniel Yorgov Department of Mathematical & Statistical Sciences, University of Colorado Denver
