Using Hoffman2 Cluster
|
|
|
- Jerome Miles
- 6 years ago
- Views:
Transcription
1 UCLA Institute for Digital Research & Education May 8 th, 2013
2 Hoffman2: largest and most powerful cluster in UC Total: > 1100 machine 11, 000 cores > 300 GPUs 102 Tflops Storage: > 1.5 Petabytes CPU: 8, 12, 16 cores, GHz Memory: 1G, 4G, 8G /core OS: CentOS Linux 6.4 MSA 400nodes (compute, gpu) (main storage) Hoffman2 Data Centers POD 500nodes (compute) IDRE 260nodes (compute) (login, vnc)
3 We have 1,200+ users, 175+ research groups Research Virtual Shared Cluster ~ 11,000 cores and increasing! Open to ALL campus users in shared base for <24 hour jobs Contributed researchers can use their own cores to run longer hour jobs
4 Before accessing Hoffman2 cluster Hoffman2 s official site: Need to have a log-in ID first via Grid Identity Manager: gim.ats.ucla.edu open to all current student, staff and faculty member with a valid UCLA log-on ID.
5 Two modes to use hoffman2 1 Web-service mode: UCLA Grid Portal: grid.ucla.edu Easiest way for windows users Slow, function-limited, depricating 2 Command-line mode : by logging into login nodes efficient, more control might need some basic unix knowledge + SGE commands
6 Two modes to use hoffman2 1 Web-service mode: UCLA Grid Portal: grid.ucla.edu Easiest way for windows users Slow, function-limited, depricating 2 Command-line mode : by logging into login nodes efficient, more control might need some basic unix knowledge + SGE commands
7 Outline via Command Line 1 via Command Line 2 A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs 3
8 Outline via Command Line 1 via Command Line 2 A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs 3
9 Outline via Command Line 1 via Command Line 2 A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs 3
10 For Unix, Linux or Mac users Using ssh command: ssh Yes for fingerprint for the first time: One of the physical login nodes will be randomly assigned login1 login4 Working with interactive GUI applications: Activate X11 forwarding: ssh -X Check Hoffman2 Login Node Fingerprints:
11 For Windows users via Command Line Getting ssh software: PuTTY, Xshell, Cygwin, Tunnelier,... Or just using web browser! Chrome Web Store Secure Shell Firefox Add-ons FireSSH Working with interactive GUI application: get an X server: PuTTY + XMing, Cygwin + Cygwin+X using NX client (not recommended for Mac user): 1 download and install NX client (NoMachine Player for Mac Lion) 2 get the key from /etc/nxserver/client.id_dsa.key 3 configure login info and input key Do not run computation on login node!
12 Additional commands from login nodes Change your password by command: passwd Configure your shell (command-line interpreter/scripts): Check your shell: echo $SHELL Text editor: vim, emacs, gedit... Tip: to keep ssh session alive: 1 vi /.ssh/config Add: ServerAliveInterval 5 2 vi /.bashrc Add: export TMOUT=36000
13 Data storage on Hoffman2 Home directory: cd $HOME 20 GB quota for general campus user backed up for 30 days Temporary use: on each node: cd $TMPDIR 100 GB, keep only during the job s run cd $SCRATCH 2 TB limit, keep for 7 days Remember to check your quota in your home directory 1 ATS scripts: get_pan_quotas $USER 2 Linux command: cd $HOME; du -sh
14 Major storage architecture change in June absolute home path will change distinguish home and sponsored directories bash profiles need to be copied into new home
15 Transferring files via Command Line from Linux/Mac terminal: Use dtn2 to transfer! ssh dtn2 scp [-r] source:path/file target:local_path from Windows/Mac/Linux GUI: Drag & drop FTP software (e.g. Cyberduck, Macfusion, FileZilla, etc.) For bigger files, use our high-speed GlobusOnline service!
16 UCLA GlobusOnline guidelines Multipoint, multi-stream data transfer fault-tolerant fire-and-forget for server-to-server transfer, 5x faster than scp 4 quick steps to use Web UI: 1 have your UCLA grid account ready 2 create a globus online account at Argonne National Lab 3 install and run GlobusConnect (if to your desktop) 4 sign in the globusonline.org and go! Command-line interface with restricted SSH Caveats: may not be appropriate for file transfers to/from a laptop need Cygwin on Windows XP
17 Don t forget Modules via Command Line Modules is for setting environmental variables $PATH $LD_LIBRARY_PATH other additional env variables. Common commands: current: module list available: module available load: module load matlab unload: module unload matlab show: module show matlab
18 Outline via Command Line 1 via Command Line 2 A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs 3
19 Taking advantage of Clusters Many Single Core Programs One Parallel Program Cluster Cluster Result Result
20 Before submission, you need to know: 1 the type of your job serial or multithreaded or distributed? 2 the group you belong to 3 the name of your executable or input file Your own code: Compile Link Executable Precompiled program (FSL, etc): Check the name of the executable Application: Matlab: m file R: R file LAMMPS, etc: input files
21 Job types on Hoffman2 Serial job: Shared memory job: MPI Parallel job: Hybrid job: Array job: single thread, single core multi-threaded, single node distributed, multiple node MPI and OpenMP serial or multi-thread same executable, different input
22 For code programmer: Intel compiler: 11.1(default), 12.0, 12.1, 13.0 GCC: 4.4(default), 4.3, 4.7 Python: 2.4, 2.6(default), 2.7, 3.1 Java: 1.6.0_23 matlab: 7.7, 7.11, 7.14(default) R: 2.9, , , (default), , 2.15 To check the software, libs installed on the cluster: module av ls /u/local/apps/ ask us!
23 Which group you are in Different groups, different policies 1 Research group: up to 14 days can use their own group s core 2 Others: up to 24 hours 4 GB memory/core up to a certain number of cores/user at a moment Use mygroup command to check! See "highp" for your own group s resource "queues": rules to jobs for requested resources.
24 Outline via Command Line 1 via Command Line 2 A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs 3
25 Of cores, nodes and tasks
26 Using ATS queue scripts for 5 types of jobs Serial Jobs job.q Serial Array Jobs jobarray.q Multi-threaded Jobs openmp.q Distributed Jobs mpi.q mpishm.q Application Jobs application.q for MPI for OpenMP with MPI
27 A quick transcript to submit a job 1 have the executable or control file ready 2 type scriptname.q and answer prompted questions: type b to build cmd file input executable or control file name input how much memory, how long time for parallel job: how many cpus... type y to submit 3 type myjobs to check the job status Tip Can submit cmd file later by qsub
28 Example: running Serial C++ program (1) Log into hoffman2: ssh hoffman2.idre... (2) Write C++ code: vi testserial.cpp (3) Compile and link: g++ -o target testserial.cpp (4) Submit & build jobs: just type job.q Type b to build Enter program name: target Enter memory request: 1024 MB for default Enter time limit: 24 hours for default Use your own group s cores: y for default Enter arguments: if needed Enter to submit
29 Example: running MPI C++ program (1) Log into hoffman2: ssh -l... (2) Write C++ code w/ MPI2 API: vi testmpi.cpp (3) Compile and link: Makefile make or mpicc -o[-c] target testmpi.c (4) Submit & build jobs in hoffman2: mpi.q Type b to build Enter program name: target Enter memory request: 1024 MB for default Enter time limit: 24 for default Use your own group s cores: y for default Enter task numbers: 8 for our example Enter arguments: if needed Enter to submit
30 Example: running Matlab using queue scripts (1) Log into login node: ssh -l... (2) Write.m file : vi test.m (3) Submit & build jobs in hoffman2: matlab.q Type b to build Enter control file name: test Enter a message option: bea for default Enter memory request: 1024 MB for default Enter time limit: 24 for default Use your own group s cores: y for default Enter arguments: if needed Enter to submit
31 Most-commonly-used SGE commands To submit a job: qsub myjob.cmd To determine the status of a job: myjobs (ATS scripts) To cancel a job: qdel [-f] jobnum For more information Use man command, for example: man qsub
32 Outline via Command Line A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs 1 via Command Line 2 A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs 3
33 Outline via Command Line A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs 1 via Command Line 2 A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs 3
34 mygroup command via Command Line A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs Group Job time Max slots/usr 0 14 days as purchased Contribute 0 24 hrs unlimited may change later 0 24 hrs 400 General may change Campus 0 2 hrs 600 Guaranteed start time < 24 hrs if grp not full none none none usually < 5 min Job type any any any serial, shm, jobarray SGE -l option highp express Interactive 0 24 hrs 8 immediately if available any i
35 More SGE commands A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs To changes the attributes of submitted but pending jobs: qalter To hold a queued job to prevent it running: qhold [jobid:taskid] To release a held job: qrls To display node information: qhost [-j]
36 More words on checkpoints A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs Job s running time on Hoffman2 is limited for campus group: as short as 24 hours. Application-level solution: (recommended) Save your data periodically in a restart file. Resubmit and restart from where it left off. System-level solution: Cluster has BLCR kernal library installed. Serial & shared-mem jobs OK. MPI job in test.
37 Outline via Command Line A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs 1 via Command Line 2 A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs 3
38 A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs Two steps in interactive-scheduler mode 1 Obtain an interactive session interactive session = reservation of resources. Reservation should correspond to your job requirements. Can request nodes up to 24 hours. Using SGE command: qrsh 2 Submit/run your job upon Do not use login node to run your program! Just use qrsh -l i to do test-running.
39 Using qrsh to request resources A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs -l options: commonly used parameters: i (or interactive): Request use the int-session nodes time (or h_rt): Wall-clock time limit (default = 2 hrs) mem (or h_data): Request memory size per core Max 1 GB/core for campus user parameters separated by commas without any space. Example: request a single core for 2 hours qrsh -l i,mem=1g,time=2:00:00 qrsh -l i,h_data=1024m,h_rt=2:00:00
40 A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs Using qrsh to request multiple cores -pe option OpenMP: -pe shared 8 MPI: -pe dc* 22 Hybrid: -pe 4threads* 20 To reserve an entire node: -l exclusive=true Examples: request an entire node for 4 hour qrsh -l i,mem=1g,time=4:00:00,exclusive=true
41 A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs Better understanding for PEs on Hoffman2 PE for MPI: dc* PE for OpenMP: shared Node 1 Node 2 Node n Slot 1 Slot 2 Slot 8 Slot 1 Slot 2 Slot 8 Slot 1 Slot 2 Slot 8 Thread Thread Thread Thread Thread Thread Thread Thread Thread PE for hybrid or double memory request: 2threads* PE for hybrid or high memory w/ whole node reservation: 8threads*
42 A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs After request, we can submit/run jobs: Serial, shared-mem, most app job: same as in a local machine. MPI job: extra work needed Example: MPI job (test) with 12 cores, 1GB/CPU, for 2 hours 1 Request CPUs from any nodes: qrsh -l i,mem=1g,time=2:00:00 -pe dc* 12 -now n 2 Obtain SGE environment variables: source /u/local/bin/set_qrsh_env.sh 3 Launch MPI job in OpenMPI: mpiexec -n $NSLOTS./test
43 Outline via Command Line A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs 1 via Command Line 2 A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs 3
44 R as a typical CLI application A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs Run R interactively: If running R serially: qrsh -l i If running R parallelly: qrsh -l i,exclusive module load R R Run R in batch: R.q About R in parallel: multithreading (R 2.14): multicore distributed (MPI) (R 2.12): snow(rmpi) or nprmpi both need to modify cmd file. Further info to check specific library docs
45 A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs If you want to run a Matlab application: Multithreading is default from v7.11 take either one slot or one whole node. We only have limited number of licenses. 6 for general, 4 for compiler 2 for statistical toolbox,... 2 ways to run matlab jobs 1 in matlab GUI, via interactive session 2 in batch, via matlab.q or matexe.q Special note on newly installed Matlab 7.14 Parallel toolbox: temporarily not working. Singlethread by compiler: temporarily not working.
46 Matlab GUI via Command Line A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs 1 enable X11-forwarding when login: ssh hoffman2.idre.ucla.edu -X -l... 2 request an interactive session: qrsh -l i,exclusive=true,time=4:00:00 3 load matlab module: module load matlab 4 run matlab: matlab To start matlab with single-thread mode matlab
47 Mablab batch mode via Command Line A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs matlab.q will do 2 steps: 1 compiling your m script to an executable 2 run the executable as a general job Can do 2 steps separately: 1 run matmcc.q 2 run matexe.q Parallel Compute Toolbox will need extra work. 1 export configuration file 2 add extra code and modify your m file Tip: sometimes hybriding does a better job! qrsh + mcc do the compilation. matexe.q do the job submission.
48 Outline via Command Line 1 via Command Line 2 A few more words about job running Computing in interactive-scheduler mode R Jobs and Matlab Jobs 3
49 Short answers for survey questions How to interactively run my application? Ans: use NX client + qrsh How to submit my job which will run in batch mode? Ans: use ATS queue scripts (job.q, mpi.q, etc) How to do HUGE data transferring? Ans: use GlobusOnline. How to let my job wait less? Ans: request short time and less mem. How to make my job running faster? Ans: request the whole node if possible. How to submit a bunch of jobs simultaneously? Ans: use job array or qsub -hold_jid [JobName]. How to use my group s contributed cores? Ans: use highp. How to run jobs with longer times (>24 hours)? Ans: add checkpoints.
50 IDRE is helping you! Hosting: resources clusters (Hoffman2, UC C 2 ) storages (high performance, archival) web services (RIM, Grid, GlobusOnline) Participating: research projects Consulting: supports, code clinics, helps Tutoring: classes, virtual summer school Contact HPC consultant hpc@ucla.edu
51 Backup slides via Command Line
52 UCLA GlobusOnline guidelines Data transfer with fault-tolerant, fire-and-forget, 5x faster than scp 4 quick steps to use Web UI: 1 have your UCLA grid account ready 2 create a globus online account at Argonne National Lab 3 install and run Globus Connect software 4 sign in the globusonline.org and go! Command-line interface with restricted SSH is available for client-side scripting. Caveats: may not be appropriate for file transfers to/from a laptop need Cygwin on Windows XP
53 Running array jobs with jobarray.queue Job array: same executable, different input (variables/files) Each core/task specific input file. Input files: names must include a sequence number. save in the same directory. Your code must identify the ID of tasks in Hoffman2. For compiled code: use getenv("sge_task_id") function For script program: use the $SGE_TASK_ID environment variable Run jobarray.q and submit
54 Example: writing C++ program for array jobs in1.dat 1 2 in2.dat 3 4 in3.dat 5 6 out1.dat i = 1 j = 2 out2.dat i = 3 j = 4 out3.dat i = 5 j = 6 testarray.cpp #include <iostream> #include <fstream> #include <sstream> #include <string> using namespace std; int main( int argc, char *argv[] ){ stringstream testid; testid << getenv( "SGE_TASK_ID" ); } string infilename="in"; string outfilename="out"; infilename += testid.str()+".dat"; outfilename += testid.str()+".dat"; ifstream infile( infilename.c_str() ); ofstream outfile( outfilename.c_str() ); int i, j; infile >> i >> j ; outfile << "i = " << i << endl; outfile << "j = " << j << endl; return 0;
55 Advanced job array submission More controls on job ids and task ids. 3 switches: -hold_jid: to specify dependency on a job_id -hold_jid_ad: to specify dependencies between tasks of different arrray jobs -tc: to define max number of concurrent jobs in the job array example: Qsub -N job1 -t 1:25 script1.cmd Qsub -hold_jid job1 -N job2 -t 26:50 script1.cmd Qsub -hold_jid job2 -N job3 -t 51:52 script1.cmd
56 Running high-memory jobs Serial: No need to combine the options of -pe and -l! Example: Serial job require 3GB memory #$ -l mem=3g MPI (e.g. 3 mpi procs, 8GB per proc): use mpi.q to allocate 3 core, 8G/core h_data or mem: still per core
Introduction to the SHARCNET Environment May-25 Pre-(summer)school webinar Speaker: Alex Razoumov University of Ontario Institute of Technology
 Introduction to the SHARCNET Environment 2010-May-25 Pre-(summer)school webinar Speaker: Alex Razoumov University of Ontario Institute of Technology available hardware and software resources our web portal
Introduction to the SHARCNET Environment 2010-May-25 Pre-(summer)school webinar Speaker: Alex Razoumov University of Ontario Institute of Technology available hardware and software resources our web portal
Introduction to HPC Using zcluster at GACRC
 Introduction to HPC Using zcluster at GACRC On-class PBIO/BINF8350 Georgia Advanced Computing Resource Center University of Georgia Zhuofei Hou, HPC Trainer zhuofei@uga.edu Outline What is GACRC? What
Introduction to HPC Using zcluster at GACRC On-class PBIO/BINF8350 Georgia Advanced Computing Resource Center University of Georgia Zhuofei Hou, HPC Trainer zhuofei@uga.edu Outline What is GACRC? What
High Performance Computing (HPC) Using zcluster at GACRC
 High Performance Computing (HPC) Using zcluster at GACRC On-class STAT8060 Georgia Advanced Computing Resource Center University of Georgia Zhuofei Hou, HPC Trainer zhuofei@uga.edu Outline What is GACRC?
High Performance Computing (HPC) Using zcluster at GACRC On-class STAT8060 Georgia Advanced Computing Resource Center University of Georgia Zhuofei Hou, HPC Trainer zhuofei@uga.edu Outline What is GACRC?
Introduction to HPC Using zcluster at GACRC
 Introduction to HPC Using zcluster at GACRC Georgia Advanced Computing Resource Center University of Georgia Zhuofei Hou, HPC Trainer zhuofei@uga.edu Outline What is GACRC? What is HPC Concept? What is
Introduction to HPC Using zcluster at GACRC Georgia Advanced Computing Resource Center University of Georgia Zhuofei Hou, HPC Trainer zhuofei@uga.edu Outline What is GACRC? What is HPC Concept? What is
Introduction to HPC Using zcluster at GACRC
 Introduction to HPC Using zcluster at GACRC On-class STAT8330 Georgia Advanced Computing Resource Center University of Georgia Suchitra Pakala pakala@uga.edu Slides courtesy: Zhoufei Hou 1 Outline What
Introduction to HPC Using zcluster at GACRC On-class STAT8330 Georgia Advanced Computing Resource Center University of Georgia Suchitra Pakala pakala@uga.edu Slides courtesy: Zhoufei Hou 1 Outline What
Introduction to HPC Using zcluster at GACRC On-Class GENE 4220
 Introduction to HPC Using zcluster at GACRC On-Class GENE 4220 Georgia Advanced Computing Resource Center University of Georgia Suchitra Pakala pakala@uga.edu Slides courtesy: Zhoufei Hou 1 OVERVIEW GACRC
Introduction to HPC Using zcluster at GACRC On-Class GENE 4220 Georgia Advanced Computing Resource Center University of Georgia Suchitra Pakala pakala@uga.edu Slides courtesy: Zhoufei Hou 1 OVERVIEW GACRC
Duke Compute Cluster Workshop. 3/28/2018 Tom Milledge rc.duke.edu
 Duke Compute Cluster Workshop 3/28/2018 Tom Milledge rc.duke.edu rescomputing@duke.edu Outline of talk Overview of Research Computing resources Duke Compute Cluster overview Running interactive and batch
Duke Compute Cluster Workshop 3/28/2018 Tom Milledge rc.duke.edu rescomputing@duke.edu Outline of talk Overview of Research Computing resources Duke Compute Cluster overview Running interactive and batch
UoW HPC Quick Start. Information Technology Services University of Wollongong. ( Last updated on October 10, 2011)
 UoW HPC Quick Start Information Technology Services University of Wollongong ( Last updated on October 10, 2011) 1 Contents 1 Logging into the HPC Cluster 3 1.1 From within the UoW campus.......................
UoW HPC Quick Start Information Technology Services University of Wollongong ( Last updated on October 10, 2011) 1 Contents 1 Logging into the HPC Cluster 3 1.1 From within the UoW campus.......................
Name Department/Research Area Have you used the Linux command line?
 Please log in with HawkID (IOWA domain) Macs are available at stations as marked To switch between the Windows and the Mac systems, press scroll lock twice 9/27/2018 1 Ben Rogers ITS-Research Services
Please log in with HawkID (IOWA domain) Macs are available at stations as marked To switch between the Windows and the Mac systems, press scroll lock twice 9/27/2018 1 Ben Rogers ITS-Research Services
Introduction to Discovery.
 Introduction to Discovery http://discovery.dartmouth.edu The Discovery Cluster 2 Agenda What is a cluster and why use it Overview of computer hardware in cluster Help Available to Discovery Users Logging
Introduction to Discovery http://discovery.dartmouth.edu The Discovery Cluster 2 Agenda What is a cluster and why use it Overview of computer hardware in cluster Help Available to Discovery Users Logging
Introduction to Discovery.
 Introduction to Discovery http://discovery.dartmouth.edu The Discovery Cluster 2 Agenda What is a cluster and why use it Overview of computer hardware in cluster Help Available to Discovery Users Logging
Introduction to Discovery http://discovery.dartmouth.edu The Discovery Cluster 2 Agenda What is a cluster and why use it Overview of computer hardware in cluster Help Available to Discovery Users Logging
Using the computational resources at the GACRC
 An introduction to zcluster Georgia Advanced Computing Resource Center (GACRC) University of Georgia Dr. Landau s PHYS4601/6601 course - Spring 2017 What is GACRC? Georgia Advanced Computing Resource Center
An introduction to zcluster Georgia Advanced Computing Resource Center (GACRC) University of Georgia Dr. Landau s PHYS4601/6601 course - Spring 2017 What is GACRC? Georgia Advanced Computing Resource Center
Introduction to HPC Using zcluster at GACRC
 Introduction to HPC Using zcluster at GACRC Georgia Advanced Computing Resource Center University of Georgia Zhuofei Hou, HPC Trainer zhuofei@uga.edu 1 Outline What is GACRC? What is HPC Concept? What
Introduction to HPC Using zcluster at GACRC Georgia Advanced Computing Resource Center University of Georgia Zhuofei Hou, HPC Trainer zhuofei@uga.edu 1 Outline What is GACRC? What is HPC Concept? What
MIGRATING TO THE SHARED COMPUTING CLUSTER (SCC) SCV Staff Boston University Scientific Computing and Visualization
 MIGRATING TO THE SHARED COMPUTING CLUSTER (SCC) SCV Staff Boston University Scientific Computing and Visualization 2 Glenn Bresnahan Director, SCV MGHPCC Buy-in Program Kadin Tseng HPC Programmer/Consultant
MIGRATING TO THE SHARED COMPUTING CLUSTER (SCC) SCV Staff Boston University Scientific Computing and Visualization 2 Glenn Bresnahan Director, SCV MGHPCC Buy-in Program Kadin Tseng HPC Programmer/Consultant
Introduction to High-Performance Computing (HPC)
 Introduction to High-Performance Computing (HPC) Computer components CPU : Central Processing Unit cores : individual processing units within a CPU Storage : Disk drives HDD : Hard Disk Drive SSD : Solid
Introduction to High-Performance Computing (HPC) Computer components CPU : Central Processing Unit cores : individual processing units within a CPU Storage : Disk drives HDD : Hard Disk Drive SSD : Solid
Introduction to HPC Using zcluster at GACRC
 Introduction to HPC Using zcluster at GACRC Georgia Advanced Computing Resource Center University of Georgia Suchitra Pakala pakala@uga.edu Slides courtesy: Zhoufei Hou OVERVIEW GACRC High Performance
Introduction to HPC Using zcluster at GACRC Georgia Advanced Computing Resource Center University of Georgia Suchitra Pakala pakala@uga.edu Slides courtesy: Zhoufei Hou OVERVIEW GACRC High Performance
Intel Manycore Testing Lab (MTL) - Linux Getting Started Guide
 Intel Manycore Testing Lab (MTL) - Linux Getting Started Guide Introduction What are the intended uses of the MTL? The MTL is prioritized for supporting the Intel Academic Community for the testing, validation
Intel Manycore Testing Lab (MTL) - Linux Getting Started Guide Introduction What are the intended uses of the MTL? The MTL is prioritized for supporting the Intel Academic Community for the testing, validation
Duke Compute Cluster Workshop. 11/10/2016 Tom Milledge h:ps://rc.duke.edu/
 Duke Compute Cluster Workshop 11/10/2016 Tom Milledge h:ps://rc.duke.edu/ rescompu>ng@duke.edu Outline of talk Overview of Research Compu>ng resources Duke Compute Cluster overview Running interac>ve and
Duke Compute Cluster Workshop 11/10/2016 Tom Milledge h:ps://rc.duke.edu/ rescompu>ng@duke.edu Outline of talk Overview of Research Compu>ng resources Duke Compute Cluster overview Running interac>ve and
Duke Compute Cluster Workshop. 10/04/2018 Tom Milledge rc.duke.edu
 Duke Compute Cluster Workshop 10/04/2018 Tom Milledge rc.duke.edu rescomputing@duke.edu Outline of talk Overview of Research Computing resources Duke Compute Cluster overview Running interactive and batch
Duke Compute Cluster Workshop 10/04/2018 Tom Milledge rc.duke.edu rescomputing@duke.edu Outline of talk Overview of Research Computing resources Duke Compute Cluster overview Running interactive and batch
Introduction to High Performance Computing Using Sapelo2 at GACRC
 Introduction to High Performance Computing Using Sapelo2 at GACRC Georgia Advanced Computing Resource Center University of Georgia Suchitra Pakala pakala@uga.edu 1 Outline High Performance Computing (HPC)
Introduction to High Performance Computing Using Sapelo2 at GACRC Georgia Advanced Computing Resource Center University of Georgia Suchitra Pakala pakala@uga.edu 1 Outline High Performance Computing (HPC)
Before We Start. Sign in hpcxx account slips Windows Users: Download PuTTY. Google PuTTY First result Save putty.exe to Desktop
 Before We Start Sign in hpcxx account slips Windows Users: Download PuTTY Google PuTTY First result Save putty.exe to Desktop Research Computing at Virginia Tech Advanced Research Computing Compute Resources
Before We Start Sign in hpcxx account slips Windows Users: Download PuTTY Google PuTTY First result Save putty.exe to Desktop Research Computing at Virginia Tech Advanced Research Computing Compute Resources
Introduction to Discovery.
 Introduction to Discovery http://discovery.dartmouth.edu March 2014 The Discovery Cluster 2 Agenda Resource overview Logging on to the cluster with ssh Transferring files to and from the cluster The Environment
Introduction to Discovery http://discovery.dartmouth.edu March 2014 The Discovery Cluster 2 Agenda Resource overview Logging on to the cluster with ssh Transferring files to and from the cluster The Environment
Our new HPC-Cluster An overview
 Our new HPC-Cluster An overview Christian Hagen Universität Regensburg Regensburg, 15.05.2009 Outline 1 Layout 2 Hardware 3 Software 4 Getting an account 5 Compiling 6 Queueing system 7 Parallelization
Our new HPC-Cluster An overview Christian Hagen Universität Regensburg Regensburg, 15.05.2009 Outline 1 Layout 2 Hardware 3 Software 4 Getting an account 5 Compiling 6 Queueing system 7 Parallelization
Grid Engine - A Batch System for DESY. Andreas Haupt, Peter Wegner DESY Zeuthen
 Grid Engine - A Batch System for DESY Andreas Haupt, Peter Wegner 15.6.2005 DESY Zeuthen Introduction Motivations for using a batch system more effective usage of available computers (e.g. reduce idle
Grid Engine - A Batch System for DESY Andreas Haupt, Peter Wegner 15.6.2005 DESY Zeuthen Introduction Motivations for using a batch system more effective usage of available computers (e.g. reduce idle
Introduction to GALILEO
 Introduction to GALILEO Parallel & production environment Mirko Cestari m.cestari@cineca.it Alessandro Marani a.marani@cineca.it Domenico Guida d.guida@cineca.it Maurizio Cremonesi m.cremonesi@cineca.it
Introduction to GALILEO Parallel & production environment Mirko Cestari m.cestari@cineca.it Alessandro Marani a.marani@cineca.it Domenico Guida d.guida@cineca.it Maurizio Cremonesi m.cremonesi@cineca.it
Introduction to High-Performance Computing (HPC)
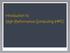 Introduction to High-Performance Computing (HPC) Computer components CPU : Central Processing Unit cores : individual processing units within a CPU Storage : Disk drives HDD : Hard Disk Drive SSD : Solid
Introduction to High-Performance Computing (HPC) Computer components CPU : Central Processing Unit cores : individual processing units within a CPU Storage : Disk drives HDD : Hard Disk Drive SSD : Solid
For Dr Landau s PHYS8602 course
 For Dr Landau s PHYS8602 course Shan-Ho Tsai (shtsai@uga.edu) Georgia Advanced Computing Resource Center - GACRC January 7, 2019 You will be given a student account on the GACRC s Teaching cluster. Your
For Dr Landau s PHYS8602 course Shan-Ho Tsai (shtsai@uga.edu) Georgia Advanced Computing Resource Center - GACRC January 7, 2019 You will be given a student account on the GACRC s Teaching cluster. Your
Introduction to the NCAR HPC Systems. 25 May 2018 Consulting Services Group Brian Vanderwende
 Introduction to the NCAR HPC Systems 25 May 2018 Consulting Services Group Brian Vanderwende Topics to cover Overview of the NCAR cluster resources Basic tasks in the HPC environment Accessing pre-built
Introduction to the NCAR HPC Systems 25 May 2018 Consulting Services Group Brian Vanderwende Topics to cover Overview of the NCAR cluster resources Basic tasks in the HPC environment Accessing pre-built
Working with Shell Scripting. Daniel Balagué
 Working with Shell Scripting Daniel Balagué Editing Text Files We offer many text editors in the HPC cluster. Command-Line Interface (CLI) editors: vi / vim nano (very intuitive and easy to use if you
Working with Shell Scripting Daniel Balagué Editing Text Files We offer many text editors in the HPC cluster. Command-Line Interface (CLI) editors: vi / vim nano (very intuitive and easy to use if you
HPC Introductory Course - Exercises
 HPC Introductory Course - Exercises The exercises in the following sections will guide you understand and become more familiar with how to use the Balena HPC service. Lines which start with $ are commands
HPC Introductory Course - Exercises The exercises in the following sections will guide you understand and become more familiar with how to use the Balena HPC service. Lines which start with $ are commands
Computing with the Moore Cluster
 Computing with the Moore Cluster Edward Walter An overview of data management and job processing in the Moore compute cluster. Overview Getting access to the cluster Data management Submitting jobs (MPI
Computing with the Moore Cluster Edward Walter An overview of data management and job processing in the Moore compute cluster. Overview Getting access to the cluster Data management Submitting jobs (MPI
HPC Workshop. Nov. 9, 2018 James Coyle, PhD Dir. Of High Perf. Computing
 HPC Workshop Nov. 9, 2018 James Coyle, PhD Dir. Of High Perf. Computing NEEDED EQUIPMENT 1. Laptop with Secure Shell (ssh) for login A. Windows: download/install putty from https://www.chiark.greenend.org.uk/~sgtatham/putty/latest.html
HPC Workshop Nov. 9, 2018 James Coyle, PhD Dir. Of High Perf. Computing NEEDED EQUIPMENT 1. Laptop with Secure Shell (ssh) for login A. Windows: download/install putty from https://www.chiark.greenend.org.uk/~sgtatham/putty/latest.html
A Hands-On Tutorial: RNA Sequencing Using High-Performance Computing
 A Hands-On Tutorial: RNA Sequencing Using Computing February 11th and 12th, 2016 1st session (Thursday) Preliminaries: Linux, HPC, command line interface Using HPC: modules, queuing system Presented by:
A Hands-On Tutorial: RNA Sequencing Using Computing February 11th and 12th, 2016 1st session (Thursday) Preliminaries: Linux, HPC, command line interface Using HPC: modules, queuing system Presented by:
XSEDE New User Tutorial
 April 2, 2014 XSEDE New User Tutorial Jay Alameda National Center for Supercomputing Applications XSEDE Training Survey Make sure you sign the sign in sheet! At the end of the module, I will ask you to
April 2, 2014 XSEDE New User Tutorial Jay Alameda National Center for Supercomputing Applications XSEDE Training Survey Make sure you sign the sign in sheet! At the end of the module, I will ask you to
Supercomputing environment TMA4280 Introduction to Supercomputing
 Supercomputing environment TMA4280 Introduction to Supercomputing NTNU, IMF February 21. 2018 1 Supercomputing environment Supercomputers use UNIX-type operating systems. Predominantly Linux. Using a shell
Supercomputing environment TMA4280 Introduction to Supercomputing NTNU, IMF February 21. 2018 1 Supercomputing environment Supercomputers use UNIX-type operating systems. Predominantly Linux. Using a shell
Image Sharpening. Practical Introduction to HPC Exercise. Instructions for Cirrus Tier-2 System
 Image Sharpening Practical Introduction to HPC Exercise Instructions for Cirrus Tier-2 System 2 1. Aims The aim of this exercise is to get you used to logging into an HPC resource, using the command line
Image Sharpening Practical Introduction to HPC Exercise Instructions for Cirrus Tier-2 System 2 1. Aims The aim of this exercise is to get you used to logging into an HPC resource, using the command line
Introduction to HPC Using zcluster at GACRC
 Introduction to HPC Using zcluster at GACRC Georgia Advanced Computing Resource Center University of Georgia Suchitra Pakala pakala@uga.edu Slides courtesy: Zhoufei Hou 1 OVERVIEW GACRC High Performance
Introduction to HPC Using zcluster at GACRC Georgia Advanced Computing Resource Center University of Georgia Suchitra Pakala pakala@uga.edu Slides courtesy: Zhoufei Hou 1 OVERVIEW GACRC High Performance
New User Seminar: Part 2 (best practices)
 New User Seminar: Part 2 (best practices) General Interest Seminar January 2015 Hugh Merz merz@sharcnet.ca Session Outline Submitting Jobs Minimizing queue waits Investigating jobs Checkpointing Efficiency
New User Seminar: Part 2 (best practices) General Interest Seminar January 2015 Hugh Merz merz@sharcnet.ca Session Outline Submitting Jobs Minimizing queue waits Investigating jobs Checkpointing Efficiency
CSE 351. Introduction & Course Tools
 CSE 351 Introduction & Course Tools Meet Your TA TA Name Interesting information examples: Where you are from Year in school Hobbies Unique talents Introductions Pick an interesting (but quick) ice breaker
CSE 351 Introduction & Course Tools Meet Your TA TA Name Interesting information examples: Where you are from Year in school Hobbies Unique talents Introductions Pick an interesting (but quick) ice breaker
Introduction to GALILEO
 November 27, 2016 Introduction to GALILEO Parallel & production environment Mirko Cestari m.cestari@cineca.it Alessandro Marani a.marani@cineca.it SuperComputing Applications and Innovation Department
November 27, 2016 Introduction to GALILEO Parallel & production environment Mirko Cestari m.cestari@cineca.it Alessandro Marani a.marani@cineca.it SuperComputing Applications and Innovation Department
XSEDE New User Tutorial
 June 12, 2015 XSEDE New User Tutorial Jay Alameda National Center for Supercomputing Applications XSEDE Training Survey Please remember to sign in for today s event: http://bit.ly/1fashvo Also, please
June 12, 2015 XSEDE New User Tutorial Jay Alameda National Center for Supercomputing Applications XSEDE Training Survey Please remember to sign in for today s event: http://bit.ly/1fashvo Also, please
Introduction to PICO Parallel & Production Enviroment
 Introduction to PICO Parallel & Production Enviroment Mirko Cestari m.cestari@cineca.it Alessandro Marani a.marani@cineca.it Domenico Guida d.guida@cineca.it Nicola Spallanzani n.spallanzani@cineca.it
Introduction to PICO Parallel & Production Enviroment Mirko Cestari m.cestari@cineca.it Alessandro Marani a.marani@cineca.it Domenico Guida d.guida@cineca.it Nicola Spallanzani n.spallanzani@cineca.it
Programming Environment on Ranger Cluster
 Programming Environment on Ranger Cluster Cornell Center for Advanced Computing December 8, 2010 12/8/2010 www.cac.cornell.edu 1 User Guides TACC Ranger (http://services.tacc.utexas.edu/index.php/ranger-user-guide)
Programming Environment on Ranger Cluster Cornell Center for Advanced Computing December 8, 2010 12/8/2010 www.cac.cornell.edu 1 User Guides TACC Ranger (http://services.tacc.utexas.edu/index.php/ranger-user-guide)
High Performance Computing (HPC) Club Training Session. Xinsheng (Shawn) Qin
 High Performance Computing (HPC) Club Training Session Xinsheng (Shawn) Qin Outline HPC Club The Hyak Supercomputer Logging in to Hyak Basic Linux Commands Transferring Files Between Your PC and Hyak Submitting
High Performance Computing (HPC) Club Training Session Xinsheng (Shawn) Qin Outline HPC Club The Hyak Supercomputer Logging in to Hyak Basic Linux Commands Transferring Files Between Your PC and Hyak Submitting
ACEnet for CS6702 Ross Dickson, Computational Research Consultant 29 Sep 2009
 ACEnet for CS6702 Ross Dickson, Computational Research Consultant 29 Sep 2009 What is ACEnet? Shared resource......for research computing... physics, chemistry, oceanography, biology, math, engineering,
ACEnet for CS6702 Ross Dickson, Computational Research Consultant 29 Sep 2009 What is ACEnet? Shared resource......for research computing... physics, chemistry, oceanography, biology, math, engineering,
An Introduction to Cluster Computing Using Newton
 An Introduction to Cluster Computing Using Newton Jason Harris and Dylan Storey March 25th, 2014 Jason Harris and Dylan Storey Introduction to Cluster Computing March 25th, 2014 1 / 26 Workshop design.
An Introduction to Cluster Computing Using Newton Jason Harris and Dylan Storey March 25th, 2014 Jason Harris and Dylan Storey Introduction to Cluster Computing March 25th, 2014 1 / 26 Workshop design.
XSEDE New User Tutorial
 May 13, 2016 XSEDE New User Tutorial Jay Alameda National Center for Supercomputing Applications XSEDE Training Survey Please complete a short on-line survey about this module at http://bit.ly/hamptonxsede.
May 13, 2016 XSEDE New User Tutorial Jay Alameda National Center for Supercomputing Applications XSEDE Training Survey Please complete a short on-line survey about this module at http://bit.ly/hamptonxsede.
Graham vs legacy systems
 New User Seminar Graham vs legacy systems This webinar only covers topics pertaining to graham. For the introduction to our legacy systems (Orca etc.), please check the following recorded webinar: SHARCNet
New User Seminar Graham vs legacy systems This webinar only covers topics pertaining to graham. For the introduction to our legacy systems (Orca etc.), please check the following recorded webinar: SHARCNet
Effective Use of CCV Resources
 Effective Use of CCV Resources Mark Howison User Services & Support This talk... Assumes you have some familiarity with a Unix shell Provides examples and best practices for typical usage of CCV systems
Effective Use of CCV Resources Mark Howison User Services & Support This talk... Assumes you have some familiarity with a Unix shell Provides examples and best practices for typical usage of CCV systems
Introduction to BioHPC
 Introduction to BioHPC New User Training [web] [email] portal.biohpc.swmed.edu biohpc-help@utsouthwestern.edu 1 Updated for 2015-06-03 Overview Today we re going to cover: What is BioHPC? How do I access
Introduction to BioHPC New User Training [web] [email] portal.biohpc.swmed.edu biohpc-help@utsouthwestern.edu 1 Updated for 2015-06-03 Overview Today we re going to cover: What is BioHPC? How do I access
Joint High Performance Computing Exchange (JHPCE) Cluster Orientation.
 Joint High Performance Computing Exchange (JHPCE) Cluster Orientation http://www.jhpce.jhu.edu/ Schedule - Introductions who are we, who are you? - Terminology - Logging in and account setup - Basics of
Joint High Performance Computing Exchange (JHPCE) Cluster Orientation http://www.jhpce.jhu.edu/ Schedule - Introductions who are we, who are you? - Terminology - Logging in and account setup - Basics of
High Performance Computing Cluster Basic course
 High Performance Computing Cluster Basic course Jeremie Vandenplas, Gwen Dawes 30 October 2017 Outline Introduction to the Agrogenomics HPC Connecting with Secure Shell to the HPC Introduction to the Unix/Linux
High Performance Computing Cluster Basic course Jeremie Vandenplas, Gwen Dawes 30 October 2017 Outline Introduction to the Agrogenomics HPC Connecting with Secure Shell to the HPC Introduction to the Unix/Linux
Using a Linux System 6
 Canaan User Guide Connecting to the Cluster 1 SSH (Secure Shell) 1 Starting an ssh session from a Mac or Linux system 1 Starting an ssh session from a Windows PC 1 Once you're connected... 1 Ending an
Canaan User Guide Connecting to the Cluster 1 SSH (Secure Shell) 1 Starting an ssh session from a Mac or Linux system 1 Starting an ssh session from a Windows PC 1 Once you're connected... 1 Ending an
PACE. Instructional Cluster Environment (ICE) Orientation. Research Scientist, PACE
 PACE Instructional Cluster Environment (ICE) Orientation Mehmet (Memo) Belgin, PhD Research Scientist, PACE www.pace.gatech.edu What is PACE A Partnership for an Advanced Computing Environment Provides
PACE Instructional Cluster Environment (ICE) Orientation Mehmet (Memo) Belgin, PhD Research Scientist, PACE www.pace.gatech.edu What is PACE A Partnership for an Advanced Computing Environment Provides
HPC DOCUMENTATION. 3. Node Names and IP addresses:- Node details with respect to their individual IP addresses are given below:-
 HPC DOCUMENTATION 1. Hardware Resource :- Our HPC consists of Blade chassis with 5 blade servers and one GPU rack server. a.total available cores for computing: - 96 cores. b.cores reserved and dedicated
HPC DOCUMENTATION 1. Hardware Resource :- Our HPC consists of Blade chassis with 5 blade servers and one GPU rack server. a.total available cores for computing: - 96 cores. b.cores reserved and dedicated
Kohinoor queuing document
 List of SGE Commands: qsub : Submit a job to SGE Kohinoor queuing document qstat : Determine the status of a job qdel : Delete a job qhost : Display Node information Some useful commands $qstat f -- Specifies
List of SGE Commands: qsub : Submit a job to SGE Kohinoor queuing document qstat : Determine the status of a job qdel : Delete a job qhost : Display Node information Some useful commands $qstat f -- Specifies
A Brief Introduction to The Center for Advanced Computing
 A Brief Introduction to The Center for Advanced Computing May 1, 2006 Hardware 324 Opteron nodes, over 700 cores 105 Athlon nodes, 210 cores 64 Apple nodes, 128 cores Gigabit networking, Myrinet networking,
A Brief Introduction to The Center for Advanced Computing May 1, 2006 Hardware 324 Opteron nodes, over 700 cores 105 Athlon nodes, 210 cores 64 Apple nodes, 128 cores Gigabit networking, Myrinet networking,
PACE. Instructional Cluster Environment (ICE) Orientation. Mehmet (Memo) Belgin, PhD Research Scientist, PACE
 PACE Instructional Cluster Environment (ICE) Orientation Mehmet (Memo) Belgin, PhD www.pace.gatech.edu Research Scientist, PACE What is PACE A Partnership for an Advanced Computing Environment Provides
PACE Instructional Cluster Environment (ICE) Orientation Mehmet (Memo) Belgin, PhD www.pace.gatech.edu Research Scientist, PACE What is PACE A Partnership for an Advanced Computing Environment Provides
Introduction to GALILEO
 Introduction to GALILEO Parallel & production environment Mirko Cestari m.cestari@cineca.it Alessandro Marani a.marani@cineca.it Alessandro Grottesi a.grottesi@cineca.it SuperComputing Applications and
Introduction to GALILEO Parallel & production environment Mirko Cestari m.cestari@cineca.it Alessandro Marani a.marani@cineca.it Alessandro Grottesi a.grottesi@cineca.it SuperComputing Applications and
Batch Systems. Running calculations on HPC resources
 Batch Systems Running calculations on HPC resources Outline What is a batch system? How do I interact with the batch system Job submission scripts Interactive jobs Common batch systems Converting between
Batch Systems Running calculations on HPC resources Outline What is a batch system? How do I interact with the batch system Job submission scripts Interactive jobs Common batch systems Converting between
NBIC TechTrack PBS Tutorial
 NBIC TechTrack PBS Tutorial by Marcel Kempenaar, NBIC Bioinformatics Research Support group, University Medical Center Groningen Visit our webpage at: http://www.nbic.nl/support/brs 1 NBIC PBS Tutorial
NBIC TechTrack PBS Tutorial by Marcel Kempenaar, NBIC Bioinformatics Research Support group, University Medical Center Groningen Visit our webpage at: http://www.nbic.nl/support/brs 1 NBIC PBS Tutorial
Working on the NewRiver Cluster
 Working on the NewRiver Cluster CMDA3634: Computer Science Foundations for Computational Modeling and Data Analytics 22 February 2018 NewRiver is a computing cluster provided by Virginia Tech s Advanced
Working on the NewRiver Cluster CMDA3634: Computer Science Foundations for Computational Modeling and Data Analytics 22 February 2018 NewRiver is a computing cluster provided by Virginia Tech s Advanced
ICS-ACI System Basics
 ICS-ACI System Basics Adam W. Lavely, Ph.D. Fall 2017 Slides available: goo.gl/ss9itf awl5173 ICS@PSU 1 Contents 1 Overview 2 HPC Overview 3 Getting Started on ACI 4 Moving On awl5173 ICS@PSU 2 Contents
ICS-ACI System Basics Adam W. Lavely, Ph.D. Fall 2017 Slides available: goo.gl/ss9itf awl5173 ICS@PSU 1 Contents 1 Overview 2 HPC Overview 3 Getting Started on ACI 4 Moving On awl5173 ICS@PSU 2 Contents
The cluster system. Introduction 22th February Jan Saalbach Scientific Computing Group
 The cluster system Introduction 22th February 2018 Jan Saalbach Scientific Computing Group cluster-help@luis.uni-hannover.de Contents 1 General information about the compute cluster 2 Available computing
The cluster system Introduction 22th February 2018 Jan Saalbach Scientific Computing Group cluster-help@luis.uni-hannover.de Contents 1 General information about the compute cluster 2 Available computing
Cluster Clonetroop: HowTo 2014
 2014/02/25 16:53 1/13 Cluster Clonetroop: HowTo 2014 Cluster Clonetroop: HowTo 2014 This section contains information about how to access, compile and execute jobs on Clonetroop, Laboratori de Càlcul Numeric's
2014/02/25 16:53 1/13 Cluster Clonetroop: HowTo 2014 Cluster Clonetroop: HowTo 2014 This section contains information about how to access, compile and execute jobs on Clonetroop, Laboratori de Càlcul Numeric's
Using WestGrid from the desktop Oct on Access Grid
 Using WestGrid from the desktop Oct 11 2007 on Access Grid Introduction Simon Sharpe, UCIT Client Services The best way to contact WestGrid support is to email support@westgrid.ca This seminar gives you
Using WestGrid from the desktop Oct 11 2007 on Access Grid Introduction Simon Sharpe, UCIT Client Services The best way to contact WestGrid support is to email support@westgrid.ca This seminar gives you
Quick Start Guide. by Burak Himmetoglu. Supercomputing Consultant. Enterprise Technology Services & Center for Scientific Computing
 Quick Start Guide by Burak Himmetoglu Supercomputing Consultant Enterprise Technology Services & Center for Scientific Computing E-mail: bhimmetoglu@ucsb.edu Contents User access, logging in Linux/Unix
Quick Start Guide by Burak Himmetoglu Supercomputing Consultant Enterprise Technology Services & Center for Scientific Computing E-mail: bhimmetoglu@ucsb.edu Contents User access, logging in Linux/Unix
High Performance Computing Cluster Advanced course
 High Performance Computing Cluster Advanced course Jeremie Vandenplas, Gwen Dawes 9 November 2017 Outline Introduction to the Agrogenomics HPC Submitting and monitoring jobs on the HPC Parallel jobs on
High Performance Computing Cluster Advanced course Jeremie Vandenplas, Gwen Dawes 9 November 2017 Outline Introduction to the Agrogenomics HPC Submitting and monitoring jobs on the HPC Parallel jobs on
Protected Environment at CHPC. Sean Igo Center for High Performance Computing September 11, 2014
 Protected Environment at CHPC Sean Igo Center for High Performance Computing Sean.Igo@utah.edu September 11, 2014 Purpose of Presentation Overview of CHPC environment / access Actually this is most of
Protected Environment at CHPC Sean Igo Center for High Performance Computing Sean.Igo@utah.edu September 11, 2014 Purpose of Presentation Overview of CHPC environment / access Actually this is most of
Choosing Resources Wisely Plamen Krastev Office: 38 Oxford, Room 117 FAS Research Computing
 Choosing Resources Wisely Plamen Krastev Office: 38 Oxford, Room 117 Email:plamenkrastev@fas.harvard.edu Objectives Inform you of available computational resources Help you choose appropriate computational
Choosing Resources Wisely Plamen Krastev Office: 38 Oxford, Room 117 Email:plamenkrastev@fas.harvard.edu Objectives Inform you of available computational resources Help you choose appropriate computational
Introduction to CINECA Computer Environment
 Introduction to CINECA Computer Environment Today you will learn... Basic commands for UNIX environment @ CINECA How to submitt your job to the PBS queueing system on Eurora Tutorial #1: Example: launch
Introduction to CINECA Computer Environment Today you will learn... Basic commands for UNIX environment @ CINECA How to submitt your job to the PBS queueing system on Eurora Tutorial #1: Example: launch
Getting started with the CEES Grid
 Getting started with the CEES Grid October, 2013 CEES HPC Manager: Dennis Michael, dennis@stanford.edu, 723-2014, Mitchell Building room 415. Please see our web site at http://cees.stanford.edu. Account
Getting started with the CEES Grid October, 2013 CEES HPC Manager: Dennis Michael, dennis@stanford.edu, 723-2014, Mitchell Building room 415. Please see our web site at http://cees.stanford.edu. Account
Batch system usage arm euthen F azo he Z J. B T
 Batch system usage 10.11.2010 General stuff Computing wikipage: http://dvinfo.ifh.de Central email address for questions & requests: uco-zn@desy.de Data storage: AFS ( /afs/ifh.de/group/amanda/scratch/
Batch system usage 10.11.2010 General stuff Computing wikipage: http://dvinfo.ifh.de Central email address for questions & requests: uco-zn@desy.de Data storage: AFS ( /afs/ifh.de/group/amanda/scratch/
Using ISMLL Cluster. Tutorial Lec 5. Mohsan Jameel, Information Systems and Machine Learning Lab, University of Hildesheim
 Using ISMLL Cluster Tutorial Lec 5 1 Agenda Hardware Useful command Submitting job 2 Computing Cluster http://www.admin-magazine.com/hpc/articles/building-an-hpc-cluster Any problem or query regarding
Using ISMLL Cluster Tutorial Lec 5 1 Agenda Hardware Useful command Submitting job 2 Computing Cluster http://www.admin-magazine.com/hpc/articles/building-an-hpc-cluster Any problem or query regarding
Introduction to CINECA HPC Environment
 Introduction to CINECA HPC Environment 23nd Summer School on Parallel Computing 19-30 May 2014 m.cestari@cineca.it, i.baccarelli@cineca.it Goals You will learn: The basic overview of CINECA HPC systems
Introduction to CINECA HPC Environment 23nd Summer School on Parallel Computing 19-30 May 2014 m.cestari@cineca.it, i.baccarelli@cineca.it Goals You will learn: The basic overview of CINECA HPC systems
XSEDE New User Tutorial
 October 20, 2017 XSEDE New User Tutorial Jay Alameda National Center for Supercomputing Applications XSEDE Training Survey Please complete a short on line survey about this module at http://bit.ly/xsedesurvey.
October 20, 2017 XSEDE New User Tutorial Jay Alameda National Center for Supercomputing Applications XSEDE Training Survey Please complete a short on line survey about this module at http://bit.ly/xsedesurvey.
Getting Started with XSEDE. Dan Stanzione
 November 3, 2011 Getting Started with XSEDE Dan Stanzione Welcome to XSEDE! XSEDE is an exciting cyberinfrastructure, providing large scale computing, data, and visualization resources. XSEDE is the evolution
November 3, 2011 Getting Started with XSEDE Dan Stanzione Welcome to XSEDE! XSEDE is an exciting cyberinfrastructure, providing large scale computing, data, and visualization resources. XSEDE is the evolution
Introduction to HPC Using the New Cluster at GACRC
 Introduction to HPC Using the New Cluster at GACRC Georgia Advanced Computing Resource Center University of Georgia Zhuofei Hou, HPC Trainer zhuofei@uga.edu Outline What is GACRC? What is the new cluster
Introduction to HPC Using the New Cluster at GACRC Georgia Advanced Computing Resource Center University of Georgia Zhuofei Hou, HPC Trainer zhuofei@uga.edu Outline What is GACRC? What is the new cluster
Using Sapelo2 Cluster at the GACRC
 Using Sapelo2 Cluster at the GACRC New User Training Workshop Georgia Advanced Computing Resource Center (GACRC) EITS/University of Georgia Zhuofei Hou zhuofei@uga.edu 1 Outline GACRC Sapelo2 Cluster Diagram
Using Sapelo2 Cluster at the GACRC New User Training Workshop Georgia Advanced Computing Resource Center (GACRC) EITS/University of Georgia Zhuofei Hou zhuofei@uga.edu 1 Outline GACRC Sapelo2 Cluster Diagram
Using Compute Canada. Masao Fujinaga Information Services and Technology University of Alberta
 Using Compute Canada Masao Fujinaga Information Services and Technology University of Alberta Introduction to cedar batch system jobs are queued priority depends on allocation and past usage Cedar Nodes
Using Compute Canada Masao Fujinaga Information Services and Technology University of Alberta Introduction to cedar batch system jobs are queued priority depends on allocation and past usage Cedar Nodes
Introduction to Unix Environment: modules, job scripts, PBS. N. Spallanzani (CINECA)
 Introduction to Unix Environment: modules, job scripts, PBS N. Spallanzani (CINECA) Bologna PATC 2016 In this tutorial you will learn... How to get familiar with UNIX environment @ CINECA How to submit
Introduction to Unix Environment: modules, job scripts, PBS N. Spallanzani (CINECA) Bologna PATC 2016 In this tutorial you will learn... How to get familiar with UNIX environment @ CINECA How to submit
STARTING THE DDT DEBUGGER ON MIO, AUN, & MC2. (Mouse over to the left to see thumbnails of all of the slides)
 STARTING THE DDT DEBUGGER ON MIO, AUN, & MC2 (Mouse over to the left to see thumbnails of all of the slides) ALLINEA DDT Allinea DDT is a powerful, easy-to-use graphical debugger capable of debugging a
STARTING THE DDT DEBUGGER ON MIO, AUN, & MC2 (Mouse over to the left to see thumbnails of all of the slides) ALLINEA DDT Allinea DDT is a powerful, easy-to-use graphical debugger capable of debugging a
Introduction to High-Performance Computing (HPC)
 Introduction to High-Performance Computing (HPC) Computer components CPU : Central Processing Unit CPU cores : individual processing units within a Storage : Disk drives HDD : Hard Disk Drive SSD : Solid
Introduction to High-Performance Computing (HPC) Computer components CPU : Central Processing Unit CPU cores : individual processing units within a Storage : Disk drives HDD : Hard Disk Drive SSD : Solid
Shark Cluster Overview
 Shark Cluster Overview 51 Execution Nodes 1 Head Node (shark) 1 Graphical login node (rivershark) 800 Cores = slots 714 TB Storage RAW Slide 1/14 Introduction What is a cluster? A cluster is a group of
Shark Cluster Overview 51 Execution Nodes 1 Head Node (shark) 1 Graphical login node (rivershark) 800 Cores = slots 714 TB Storage RAW Slide 1/14 Introduction What is a cluster? A cluster is a group of
Introduction to High Performance Computing (HPC) Resources at GACRC
 Introduction to High Performance Computing (HPC) Resources at GACRC Georgia Advanced Computing Resource Center University of Georgia Zhuofei Hou, HPC Trainer zhuofei@uga.edu Outline What is GACRC? Concept
Introduction to High Performance Computing (HPC) Resources at GACRC Georgia Advanced Computing Resource Center University of Georgia Zhuofei Hou, HPC Trainer zhuofei@uga.edu Outline What is GACRC? Concept
Parallel Computing with Matlab and R
 Parallel Computing with Matlab and R scsc@duke.edu https://wiki.duke.edu/display/scsc Tom Milledge tm103@duke.edu Overview Running Matlab and R interactively and in batch mode Introduction to Parallel
Parallel Computing with Matlab and R scsc@duke.edu https://wiki.duke.edu/display/scsc Tom Milledge tm103@duke.edu Overview Running Matlab and R interactively and in batch mode Introduction to Parallel
The RWTH Compute Cluster Environment
 The RWTH Compute Cluster Environment Tim Cramer 29.07.2013 Source: D. Both, Bull GmbH Rechen- und Kommunikationszentrum (RZ) The RWTH Compute Cluster (1/2) The Cluster provides ~300 TFlop/s No. 32 in TOP500
The RWTH Compute Cluster Environment Tim Cramer 29.07.2013 Source: D. Both, Bull GmbH Rechen- und Kommunikationszentrum (RZ) The RWTH Compute Cluster (1/2) The Cluster provides ~300 TFlop/s No. 32 in TOP500
User Guide of High Performance Computing Cluster in School of Physics
 User Guide of High Performance Computing Cluster in School of Physics Prepared by Sue Yang (xue.yang@sydney.edu.au) This document aims at helping users to quickly log into the cluster, set up the software
User Guide of High Performance Computing Cluster in School of Physics Prepared by Sue Yang (xue.yang@sydney.edu.au) This document aims at helping users to quickly log into the cluster, set up the software
Minnesota Supercomputing Institute Regents of the University of Minnesota. All rights reserved.
 Minnesota Supercomputing Institute Introduction to Job Submission and Scheduling Andrew Gustafson Interacting with MSI Systems Connecting to MSI SSH is the most reliable connection method Linux and Mac
Minnesota Supercomputing Institute Introduction to Job Submission and Scheduling Andrew Gustafson Interacting with MSI Systems Connecting to MSI SSH is the most reliable connection method Linux and Mac
Introduction to the Cluster
 Follow us on Twitter for important news and updates: @ACCREVandy Introduction to the Cluster Advanced Computing Center for Research and Education http://www.accre.vanderbilt.edu The Cluster We will be
Follow us on Twitter for important news and updates: @ACCREVandy Introduction to the Cluster Advanced Computing Center for Research and Education http://www.accre.vanderbilt.edu The Cluster We will be
Using Cartesius and Lisa. Zheng Meyer-Zhao - Consultant Clustercomputing
 Zheng Meyer-Zhao - zheng.meyer-zhao@surfsara.nl Consultant Clustercomputing Outline SURFsara About us What we do Cartesius and Lisa Architectures and Specifications File systems Funding Hands-on Logging
Zheng Meyer-Zhao - zheng.meyer-zhao@surfsara.nl Consultant Clustercomputing Outline SURFsara About us What we do Cartesius and Lisa Architectures and Specifications File systems Funding Hands-on Logging
New User Tutorial. OSU High Performance Computing Center
 New User Tutorial OSU High Performance Computing Center TABLE OF CONTENTS Logging In... 3-5 Windows... 3-4 Linux... 4 Mac... 4-5 Changing Password... 5 Using Linux Commands... 6 File Systems... 7 File
New User Tutorial OSU High Performance Computing Center TABLE OF CONTENTS Logging In... 3-5 Windows... 3-4 Linux... 4 Mac... 4-5 Changing Password... 5 Using Linux Commands... 6 File Systems... 7 File
INTRODUCTION TO THE CLUSTER
 INTRODUCTION TO THE CLUSTER WHAT IS A CLUSTER? A computer cluster consists of a group of interconnected servers (nodes) that work together to form a single logical system. COMPUTE NODES GATEWAYS SCHEDULER
INTRODUCTION TO THE CLUSTER WHAT IS A CLUSTER? A computer cluster consists of a group of interconnected servers (nodes) that work together to form a single logical system. COMPUTE NODES GATEWAYS SCHEDULER
HTC Brief Instructions
 HTC Brief Instructions Version 18.08.2018 University of Paderborn Paderborn Center for Parallel Computing Warburger Str. 100, D-33098 Paderborn http://pc2.uni-paderborn.de/ 2 HTC BRIEF INSTRUCTIONS Table
HTC Brief Instructions Version 18.08.2018 University of Paderborn Paderborn Center for Parallel Computing Warburger Str. 100, D-33098 Paderborn http://pc2.uni-paderborn.de/ 2 HTC BRIEF INSTRUCTIONS Table
National Biochemical Computational Research https://nbcr.net/accounts/apply.php. Familiarize yourself with the account policy
 Track 3: Molecular Visualization and Virtual Screening NBCR Summer Institute Session: NBCR clusters introduction August 11, 2006 Nadya Williams nadya@sdsc.edu Where to start National Biochemical Computational
Track 3: Molecular Visualization and Virtual Screening NBCR Summer Institute Session: NBCR clusters introduction August 11, 2006 Nadya Williams nadya@sdsc.edu Where to start National Biochemical Computational
The JANUS Computing Environment
 Research Computing UNIVERSITY OF COLORADO The JANUS Computing Environment Monte Lunacek monte.lunacek@colorado.edu rc-help@colorado.edu What is JANUS? November, 2011 1,368 Compute nodes 16,416 processors
Research Computing UNIVERSITY OF COLORADO The JANUS Computing Environment Monte Lunacek monte.lunacek@colorado.edu rc-help@colorado.edu What is JANUS? November, 2011 1,368 Compute nodes 16,416 processors
Sharpen Exercise: Using HPC resources and running parallel applications
 Sharpen Exercise: Using HPC resources and running parallel applications Andrew Turner, Dominic Sloan-Murphy, David Henty, Adrian Jackson Contents 1 Aims 2 2 Introduction 2 3 Instructions 3 3.1 Log into
Sharpen Exercise: Using HPC resources and running parallel applications Andrew Turner, Dominic Sloan-Murphy, David Henty, Adrian Jackson Contents 1 Aims 2 2 Introduction 2 3 Instructions 3 3.1 Log into
Migrating from Zcluster to Sapelo
 GACRC User Quick Guide: Migrating from Zcluster to Sapelo The GACRC Staff Version 1.0 8/4/17 1 Discussion Points I. Request Sapelo User Account II. III. IV. Systems Transfer Files Configure Software Environment
GACRC User Quick Guide: Migrating from Zcluster to Sapelo The GACRC Staff Version 1.0 8/4/17 1 Discussion Points I. Request Sapelo User Account II. III. IV. Systems Transfer Files Configure Software Environment
Lab 1 Introduction to UNIX and C
 Name: Lab 1 Introduction to UNIX and C This first lab is meant to be an introduction to computer environments we will be using this term. You must have a Pitt username to complete this lab. NOTE: Text
Name: Lab 1 Introduction to UNIX and C This first lab is meant to be an introduction to computer environments we will be using this term. You must have a Pitt username to complete this lab. NOTE: Text
Quick Start Guide. Table of Contents
 Quick Start Guide Table of Contents Account Registration... 2 Signup Request... 2 Account Activation... 4 Running FLOW-3D on POD... 9 Launching the GUI... 9 Running Simulations... 11 Collaborating with
Quick Start Guide Table of Contents Account Registration... 2 Signup Request... 2 Account Activation... 4 Running FLOW-3D on POD... 9 Launching the GUI... 9 Running Simulations... 11 Collaborating with
