Central Issues in Biological Sequence Comparison
|
|
|
- Isabel Cunningham
- 5 years ago
- Views:
Transcription
1 Central Issues in Biological Sequence Comparison Definitions: What is one trying to find or optimize? Algorithms: Can one find the proposed object optimally or in reasonable time optimize? Statistics: Can one s result be explained by chance? In general there is a tension between questions. A definition that is too simple may allow efficient algorithms, but may not yield results of biological interest. However, a definition that includes most of the relevant biology may entail intractable algorithms and statistics. The most successful approaches find a balance between these considerations.
2 The Problem Given: Two protein or DNA sequences where the and are chosen from a finite alphabet, e.g.,,,. How can one define the distancebetween the sequences and, or alternatively their similarity? We shall adopt the somewhat more flexible formalism of similarity, with higher values considered better. Although there are other possibilities, similarity is generally defined with reference to a sequence alignment, in which individual letters from each sequence are placed into correspondence.
3 Examples of Sequence Alignment groan : grown colo-r colour theatre :: theater theatre theater elephant : eleg-ant vermiform :: ::::: formation vermiform----- about -----formation disestablishment dis s--ent disestablishment : dis sent
4 Applications Sequence alignment arises in many fields: Molecular biology Inexact text matching (e.g. spell checkers; web page search) Speech recognition In general: The precise definition of what constitutes an alignment may vary by field, and even within a field. Many different alignments of two sequences are possible, so to select among them one requires an objective (score) function on alignments. The number of possible alignments of two sequences grows exponentially with the length of the sequences. Good algorithms are required.
5 NCBI BLAST Protein Database Search BLAST is a widely-used program for searching DNA and protein sequence databases for sequences similar to a query sequence. Here is one alignment returned by a BLAST protein database search: >sp Q BARD1_HUMAN Length=777 GENE ID: 58 BARD1 BRCA1 associated RING domain 1 [Homo sapiens] Score = 53.1 bits (126), Expect = 3e-7, Method: Composition-based stats. Identities = 32/111 (29%), Positives = 55/111 (5%), Gaps = 15/111 (14%) Query 24 THVVMKTDAEFVCERTLKYFLGIAGGKWVVSYFWVTQSIKERKMLNEHDFEVRGDVVNGR 83 THVV+ DA + TLK LGI G W++ + WV ++ + E +E+ Sbjct 65 THVVVPGDA---VQSTLKCMLGILNGCWILKFEWVKACLRRKVCEQEEKYEIP Query 84 NHQGPKRARESQDR---KIFRGLEICCYGPFTNMPTDQLEWMVQLCGASVV 131 +GP+R+R ++++ K+F G +G F + P D L +V G ++ Sbjct EGPRRSRLNREQLLPKLFDGCYFYLWGTFKHHPKDNLIKLVTAGGGQIL 73
6 Elements of Global Sequence Alignment No crossings allowed. For algorithmic reasons, it is fortunate that, although there are natural mechanisms (mutations) that lead to amino acid or nucleotide substitutions, insertions and deletions, there are none that yield transpositions, unlike with keyboardproduced text. In contrast, when analyzing RNA folding, one may choose for algorithmic reasons to exclude pseudoknots, which do in fact occur naturally. Gaps. An arbitrary number of nullcharacters (represented by dashes) may be placed into either sequence, and aligned with letters in the other sequence. Two nulls may not be aligned. Depending upon one s perspective, the alignment of a letter with a null may be understood as the insertionof a letter into one sequence, or the deletionof a letter from the other. Therefore, a letter aligned with a null is sometimes called an indel. Alignment scores. The score for an alignment is taken to be the sum of scores for aligned pairs of letters, and scores for letters aligned with nulls. Each such pairing is called an alignment column. Substitution scores. Scores for aligned pairs of letters are called substitution scores, whether the letter aligned are identical or not. Most simply, substitution scores may take the form of match scores and mismatch scores. Gap scores. The score for a letter aligned with a null is called a gap score. Usually gap scores are letter-independent. Global alignment. All letters and nulls in each sequence must be aligned.
7 Sequence Similarity Define the similarityof two sequences as the score of their highest-scoring (optimal) alignment. How do we find the an optimal alignment of two sequence, and its score? Brute force enumeration is impractical, because the number of possible alignments becomes astronomically large for even fairly short sequences. Fortunately, the problem is soluble efficiently using a technique called dynamic programming.
8 Dynamic Programming and Global Alignment Dynamic programming is a method by which a larger problem may be solved by first solving smaller, partial versions of the problem. We demonstrate here how it may be applied to global sequence alignment, where at first we are interested only in the similarity of two sequences, and not the alignment that yields this score. Definitions:, the substitution score for aligning letters and the gap score for aligning any letter to a null the partial sequence consisting of the first letters of the partial sequence consisting of the first letters of (, ) the similarity of and Consider the last column of an optimal alignment of and. This column either aligns to, or to a null, or to a null. Because we do not allow crossing, there are no other possibilities. This observation yields the following recurrence:, + (, ), =, +, + and aligned aligned with a null aligned with a null In brief, we can solve for (, )by solving smaller versions of the problem first.
9 Path graphs A global alignment may be viewed as a path through a directed path graphwhich begins at the upper left corner and ends at the lower right. Diagonal steps correspond to substitutions, while horizontal or vertical steps correspond to indels. Scores are associated with each edge, and the score of an alignment is the sum of the scores of the edges it traverses. Each alignment corresponds to a unique path, and vice versa. Start End
10 Dynamic programming on path graphs One may associate a partial similarity with each node of a path graph. If the values of 1, 1, 1, and, 1 are known, the value of, may be calculated. (, ) (, ) (, ) (, ) (, )
11 A C G C An Example G A C T A C Scores: Match +1 Mismatch Gap -1
12 What is the optimal alignment? A C G C G A C T A C Record tracebackinformation: Which edge or edges led to the optimal score at each node?
13 Follow the traceback edges from the final node A C G C G A C T A C Optimal alignments: -ACG-C GACTAC and -AC-GC GACTAC
14 Pseudocode for Finding Sequence Similarity Similarity(X,Y): For i =,...,m: SIM[i,] = i*g For j = 1,...,n: SIM[,j] = j*g For i = 1,...,m: For j = 1,...,n: SIM[i,j] = max( SIM[i-1,j-1] + s(x[i],y[j]), SIM[i-1,j]+g, SIM[i,j-1]+g ) EndFor EndFor Return SIM[m,n] Exercise: Generalize the code to include tracebackinformation, and produce one optimal alignment. Note: This is generally known as the Needleman-Wunschalgorithm, after the first paper in the field of computational molecular biology to apply dynamic programming to the global alignment problem. However, the paper actually describes a somewhat different algorithm which is almost never used. Needleman, S.B. & Wunsch, C.D. (197) A general method applicable to the search for similarities in the amino acid sequences of two proteins. J. Mol. Biol. 48:
15 Observations and Generalizations The nodes can be expanded in a variety of orders, so long as all nodes that feed into a given node are expanded before that node. Possible expansion orders are: The time complexity of the algorithm is. If only the similarity is desired, the space complexity is min, ; if an optimal alignment is sought, the space complexity is, but as we shall see, this too can be reduced to [min, ]. It is possible to save time (but in general no more than a constant factor) by not expanding nodes that can not possibly participate in an optimal path. Fickett, J.W. (1984) Nucl. Acids Res. 12:175-18; Spouge, J.L. (1989) SIAM J. Appl. Math. 49:
16 Global Alignment Scores Multiplying all substitution and gap scores by a positive constant does not change the optimal alignment. Why? Adding a constant to all substitution scores, and /2to all gap scores, does not change the optimal alignment. Why? A global alignment scoring system with the three nominal parameters of match score, mismatch score, and gap score, in fact has a single free parameter. For example, assuming >, one can always construct an equivalent scoring system with =1and =. What is the scoring system of this form equivalent to ( =1, =, = 1)? Modifying global alignment scores so that =can speed up the inner loop of the dynamic programming algorithm.
17 Local Alignment: Motivation In the early days of protein sequence comparison, most known related proteins, were related over their whole lengths. However, soon proteins that shared only isolated regions of similarity were found. A schematic of a protein superfamilyis shown here, with related domains represented by similar boxes. The measure of global sequence similarity, and the Needleman-Wunsch alignment algorithm, was not welladapted to finding such domains. A new definition of local similarity was required, along with a new algorithm for finding locally optimal alignments.
18 Local Alignment: Definition During the 197s and early 198s, a variety of definitions for local alignment were proposed. The one that eventually gained the greatest popularity, along with an associated algorithm, is due to Smith & Waterman. Smith & Waterman proposed simply that a local alignment of two sequences allow arbitrary-length segments of each sequence to be aligned, with no penalty for the unaligned portions of the sequences. Otherwise, the score for a local alignment is calculated the same way as that for a global alignment. It would at first appear that the problem of finding an optimal local alignment should be significantly more complex than the problem of finding an optimal global alignment, because the start and stop positions of the alignment must be located as well. However, only a constant factor more calculation is necessary. Smith, T.F. & Waterman, M.S. (1981) Identification of common molecular subsequences. J. Mol. Biol. 147:
19 . The Smith-Waterman Algorithm Two modifications to Needleman-Wunsch: 1) Allow a node to start at. 2) Record the highestscoring node, and trace back from there. Why does this algorithm yield an optimal local alignment? A C G C G A T T G A Scores: Match +4 Mismatch -1 Gap -2
20 Pseudocode for Finding Local Sequence Similarity Local_Similarity(X,Y): S= For i =,...,m: SIM[i,] = i*g For j = 1,...,n: SIM[,j] = j*g For i = 1,...,m: For j = 1,...,n: SIM[i,j] = max(, SIM[i-1,j-1] + s(x[i],y[j]), SIM[i-1,j]+g, SIM[i,j-1]+g ) S=max(S,SIM[i,j]) EndFor EndFor Return S Exercise: Generalize the code to include tracebackinformation, and produce one optimal local alignment. Multiplying all substitution and gap scores by a positive constant does not change the optimal alignment. Why? Adding a constant to all substitution scores, and /2 to all gap scores, canchange the optimal alignment. Why?
21 . The Smith-Waterman Algorithm: Traceback Optimal local alignments, or subalignments: AC-G ATTG and Questions: A-CG ATTG Can one find other locally optimal subalignments? How can they be defined? A C G C G A T T G A Scores: Match +4 Mismatch -1 Gap -2
22 Local optimality: Definitions and Algorithms A definition of local optimality was proposed in 1984, along with an algorithm to find all locally optimal subalignments. [Sellers, P.H. (1984) Bull. Math. Biol. 46: ] A subalignmentis locally optimal if its score is greater than or equal to that of any subalignment it touches. A provably ( )algorithm for finding all locally optimal subalignmentswas subsequently described. [Altschul, S.F. & Erickson, B.W (1986) Bull. Math. Biol. 48: ] Problem: By Sellers definition, a strong subalignmentcan suppress, by means of intermediaries, subalignmentsit does not actually touch. This can be a particular problem if one is seeking internal approximate repeats. One may advance an alternative definition to address this problem: A subalignmentis weakly locally optimal if it touches no weakly locally optimal subalignmentthat has greater score (Altschul & Erickson, 1986). This definition is not circular, but recursive. No algorithm for finding all weakly locally optimal subalignmentsof two sequences has been described, although several incorrect ones have been published. > > ( )
23 . Locally Optimal Subalignments Optimal subalignments: AC-G ATTG and A-CG ATTG Additional, locally optimal subalignments: G G and A A A C G C G A T T G A Scores: Match +4 Mismatch -1 Gap -2
24 Semi-Global Alignment Biological problem: Find approximate matches to a given pattern within a large sequence. For example, seek promoters within a DNA sequence, or a copies of a domain within a protein sequence. Solution: Semi-global alignment. Needleman-Wunschalgorithm with two modifications: 1) Penalize end gaps in the pattern, but not in the long sequence; 2) Trace back from the highest scoring node along the long edge of the path graph Erickson, B.W. & Sellers, P.H. (1983) Recognition of patterns in genetic sequences. In Time Warps, String Edits and Macromolecules: The Theory and Practice of Sequence Comparison, D. Sankoff& J.B. Kruskal(eds.), pp , Addison-Wesley, Reading, MA.
Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014
 Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014 Dynamic programming is a group of mathematical methods used to sequentially split a complicated problem into
Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014 Dynamic programming is a group of mathematical methods used to sequentially split a complicated problem into
) I R L Press Limited, Oxford, England. The protein identification resource (PIR)
 Volume 14 Number 1 Volume 1986 Nucleic Acids Research 14 Number 1986 Nucleic Acids Research The protein identification resource (PIR) David G.George, Winona C.Barker and Lois T.Hunt National Biomedical
Volume 14 Number 1 Volume 1986 Nucleic Acids Research 14 Number 1986 Nucleic Acids Research The protein identification resource (PIR) David G.George, Winona C.Barker and Lois T.Hunt National Biomedical
Notes on Dynamic-Programming Sequence Alignment
 Notes on Dynamic-Programming Sequence Alignment Introduction. Following its introduction by Needleman and Wunsch (1970), dynamic programming has become the method of choice for rigorous alignment of DNA
Notes on Dynamic-Programming Sequence Alignment Introduction. Following its introduction by Needleman and Wunsch (1970), dynamic programming has become the method of choice for rigorous alignment of DNA
Computational Genomics and Molecular Biology, Fall
 Computational Genomics and Molecular Biology, Fall 2015 1 Sequence Alignment Dannie Durand Pairwise Sequence Alignment The goal of pairwise sequence alignment is to establish a correspondence between the
Computational Genomics and Molecular Biology, Fall 2015 1 Sequence Alignment Dannie Durand Pairwise Sequence Alignment The goal of pairwise sequence alignment is to establish a correspondence between the
Bioinformatics explained: Smith-Waterman
 Bioinformatics Explained Bioinformatics explained: Smith-Waterman May 1, 2007 CLC bio Gustav Wieds Vej 10 8000 Aarhus C Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com info@clcbio.com
Bioinformatics Explained Bioinformatics explained: Smith-Waterman May 1, 2007 CLC bio Gustav Wieds Vej 10 8000 Aarhus C Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com info@clcbio.com
Distributed Protein Sequence Alignment
 Distributed Protein Sequence Alignment ABSTRACT J. Michael Meehan meehan@wwu.edu James Hearne hearne@wwu.edu Given the explosive growth of biological sequence databases and the computational complexity
Distributed Protein Sequence Alignment ABSTRACT J. Michael Meehan meehan@wwu.edu James Hearne hearne@wwu.edu Given the explosive growth of biological sequence databases and the computational complexity
Biology 644: Bioinformatics
 Find the best alignment between 2 sequences with lengths n and m, respectively Best alignment is very dependent upon the substitution matrix and gap penalties The Global Alignment Problem tries to find
Find the best alignment between 2 sequences with lengths n and m, respectively Best alignment is very dependent upon the substitution matrix and gap penalties The Global Alignment Problem tries to find
Lecture 10. Sequence alignments
 Lecture 10 Sequence alignments Alignment algorithms: Overview Given a scoring system, we need to have an algorithm for finding an optimal alignment for a pair of sequences. We want to maximize the score
Lecture 10 Sequence alignments Alignment algorithms: Overview Given a scoring system, we need to have an algorithm for finding an optimal alignment for a pair of sequences. We want to maximize the score
An Efficient Algorithm to Locate All Locally Optimal Alignments Between Two Sequences Allowing for Gaps
 An Efficient Algorithm to Locate All Locally Optimal Alignments Between Two Sequences Allowing for Gaps Geoffrey J. Barton Laboratory of Molecular Biophysics University of Oxford Rex Richards Building
An Efficient Algorithm to Locate All Locally Optimal Alignments Between Two Sequences Allowing for Gaps Geoffrey J. Barton Laboratory of Molecular Biophysics University of Oxford Rex Richards Building
Programming assignment for the course Sequence Analysis (2006)
 Programming assignment for the course Sequence Analysis (2006) Original text by John W. Romein, adapted by Bart van Houte (bart@cs.vu.nl) Introduction Please note: This assignment is only obligatory for
Programming assignment for the course Sequence Analysis (2006) Original text by John W. Romein, adapted by Bart van Houte (bart@cs.vu.nl) Introduction Please note: This assignment is only obligatory for
A CAM(Content Addressable Memory)-based architecture for molecular sequence matching
 A CAM(Content Addressable Memory)-based architecture for molecular sequence matching P.K. Lala 1 and J.P. Parkerson 2 1 Department Electrical Engineering, Texas A&M University, Texarkana, Texas, USA 2
A CAM(Content Addressable Memory)-based architecture for molecular sequence matching P.K. Lala 1 and J.P. Parkerson 2 1 Department Electrical Engineering, Texas A&M University, Texarkana, Texas, USA 2
An Analysis of Pairwise Sequence Alignment Algorithm Complexities: Needleman-Wunsch, Smith-Waterman, FASTA, BLAST and Gapped BLAST
 An Analysis of Pairwise Sequence Alignment Algorithm Complexities: Needleman-Wunsch, Smith-Waterman, FASTA, BLAST and Gapped BLAST Alexander Chan 5075504 Biochemistry 218 Final Project An Analysis of Pairwise
An Analysis of Pairwise Sequence Alignment Algorithm Complexities: Needleman-Wunsch, Smith-Waterman, FASTA, BLAST and Gapped BLAST Alexander Chan 5075504 Biochemistry 218 Final Project An Analysis of Pairwise
Lecture Overview. Sequence search & alignment. Searching sequence databases. Sequence Alignment & Search. Goals: Motivations:
 Lecture Overview Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating
Lecture Overview Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating
Sequence Alignment & Search
 Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating the first version
Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating the first version
Pairwise Sequence Alignment: Dynamic Programming Algorithms COMP 571 Luay Nakhleh, Rice University
 1 Pairwise Sequence Alignment: Dynamic Programming Algorithms COMP 571 Luay Nakhleh, Rice University DP Algorithms for Pairwise Alignment 2 The number of all possible pairwise alignments (if gaps are allowed)
1 Pairwise Sequence Alignment: Dynamic Programming Algorithms COMP 571 Luay Nakhleh, Rice University DP Algorithms for Pairwise Alignment 2 The number of all possible pairwise alignments (if gaps are allowed)
PROTEIN MULTIPLE ALIGNMENT MOTIVATION: BACKGROUND: Marina Sirota
 Marina Sirota MOTIVATION: PROTEIN MULTIPLE ALIGNMENT To study evolution on the genetic level across a wide range of organisms, biologists need accurate tools for multiple sequence alignment of protein
Marina Sirota MOTIVATION: PROTEIN MULTIPLE ALIGNMENT To study evolution on the genetic level across a wide range of organisms, biologists need accurate tools for multiple sequence alignment of protein
BLAST: Basic Local Alignment Search Tool Altschul et al. J. Mol Bio CS 466 Saurabh Sinha
 BLAST: Basic Local Alignment Search Tool Altschul et al. J. Mol Bio. 1990. CS 466 Saurabh Sinha Motivation Sequence homology to a known protein suggest function of newly sequenced protein Bioinformatics
BLAST: Basic Local Alignment Search Tool Altschul et al. J. Mol Bio. 1990. CS 466 Saurabh Sinha Motivation Sequence homology to a known protein suggest function of newly sequenced protein Bioinformatics
Special course in Computer Science: Advanced Text Algorithms
 Special course in Computer Science: Advanced Text Algorithms Lecture 6: Alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
Special course in Computer Science: Advanced Text Algorithms Lecture 6: Alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
Today s Lecture. Multiple sequence alignment. Improved scoring of pairwise alignments. Affine gap penalties Profiles
 Today s Lecture Multiple sequence alignment Improved scoring of pairwise alignments Affine gap penalties Profiles 1 The Edit Graph for a Pair of Sequences G A C G T T G A A T G A C C C A C A T G A C G
Today s Lecture Multiple sequence alignment Improved scoring of pairwise alignments Affine gap penalties Profiles 1 The Edit Graph for a Pair of Sequences G A C G T T G A A T G A C C C A C A T G A C G
Lecture 3: February Local Alignment: The Smith-Waterman Algorithm
 CSCI1820: Sequence Alignment Spring 2017 Lecture 3: February 7 Lecturer: Sorin Istrail Scribe: Pranavan Chanthrakumar Note: LaTeX template courtesy of UC Berkeley EECS dept. Notes are also adapted from
CSCI1820: Sequence Alignment Spring 2017 Lecture 3: February 7 Lecturer: Sorin Istrail Scribe: Pranavan Chanthrakumar Note: LaTeX template courtesy of UC Berkeley EECS dept. Notes are also adapted from
24 Grundlagen der Bioinformatik, SS 10, D. Huson, April 26, This lecture is based on the following papers, which are all recommended reading:
 24 Grundlagen der Bioinformatik, SS 10, D. Huson, April 26, 2010 3 BLAST and FASTA This lecture is based on the following papers, which are all recommended reading: D.J. Lipman and W.R. Pearson, Rapid
24 Grundlagen der Bioinformatik, SS 10, D. Huson, April 26, 2010 3 BLAST and FASTA This lecture is based on the following papers, which are all recommended reading: D.J. Lipman and W.R. Pearson, Rapid
Pairwise Sequence Alignment: Dynamic Programming Algorithms. COMP Spring 2015 Luay Nakhleh, Rice University
 Pairwise Sequence Alignment: Dynamic Programming Algorithms COMP 571 - Spring 2015 Luay Nakhleh, Rice University DP Algorithms for Pairwise Alignment The number of all possible pairwise alignments (if
Pairwise Sequence Alignment: Dynamic Programming Algorithms COMP 571 - Spring 2015 Luay Nakhleh, Rice University DP Algorithms for Pairwise Alignment The number of all possible pairwise alignments (if
Bioinformatics for Biologists
 Bioinformatics for Biologists Sequence Analysis: Part I. Pairwise alignment and database searching Fran Lewitter, Ph.D. Director Bioinformatics & Research Computing Whitehead Institute Topics to Cover
Bioinformatics for Biologists Sequence Analysis: Part I. Pairwise alignment and database searching Fran Lewitter, Ph.D. Director Bioinformatics & Research Computing Whitehead Institute Topics to Cover
Pairwise Sequence Alignment. Zhongming Zhao, PhD
 Pairwise Sequence Alignment Zhongming Zhao, PhD Email: zhongming.zhao@vanderbilt.edu http://bioinfo.mc.vanderbilt.edu/ Sequence Similarity match mismatch A T T A C G C G T A C C A T A T T A T G C G A T
Pairwise Sequence Alignment Zhongming Zhao, PhD Email: zhongming.zhao@vanderbilt.edu http://bioinfo.mc.vanderbilt.edu/ Sequence Similarity match mismatch A T T A C G C G T A C C A T A T T A T G C G A T
OPEN MP-BASED PARALLEL AND SCALABLE GENETIC SEQUENCE ALIGNMENT
 OPEN MP-BASED PARALLEL AND SCALABLE GENETIC SEQUENCE ALIGNMENT Asif Ali Khan*, Laiq Hassan*, Salim Ullah* ABSTRACT: In bioinformatics, sequence alignment is a common and insistent task. Biologists align
OPEN MP-BASED PARALLEL AND SCALABLE GENETIC SEQUENCE ALIGNMENT Asif Ali Khan*, Laiq Hassan*, Salim Ullah* ABSTRACT: In bioinformatics, sequence alignment is a common and insistent task. Biologists align
Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it.
 Sequence Alignments Overview Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it. Sequence alignment means arranging
Sequence Alignments Overview Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it. Sequence alignment means arranging
BGGN 213 Foundations of Bioinformatics Barry Grant
 BGGN 213 Foundations of Bioinformatics Barry Grant http://thegrantlab.org/bggn213 Recap From Last Time: 25 Responses: https://tinyurl.com/bggn213-02-f17 Why ALIGNMENT FOUNDATIONS Why compare biological
BGGN 213 Foundations of Bioinformatics Barry Grant http://thegrantlab.org/bggn213 Recap From Last Time: 25 Responses: https://tinyurl.com/bggn213-02-f17 Why ALIGNMENT FOUNDATIONS Why compare biological
Alignment of Long Sequences
 Alignment of Long Sequences BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2009 Mark Craven craven@biostat.wisc.edu Pairwise Whole Genome Alignment: Task Definition Given a pair of genomes (or other large-scale
Alignment of Long Sequences BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2009 Mark Craven craven@biostat.wisc.edu Pairwise Whole Genome Alignment: Task Definition Given a pair of genomes (or other large-scale
Sequence alignment algorithms
 Sequence alignment algorithms Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 23 rd 27 After this lecture, you can decide when to use local and global sequence alignments
Sequence alignment algorithms Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 23 rd 27 After this lecture, you can decide when to use local and global sequence alignments
Research on Pairwise Sequence Alignment Needleman-Wunsch Algorithm
 5th International Conference on Mechatronics, Materials, Chemistry and Computer Engineering (ICMMCCE 2017) Research on Pairwise Sequence Alignment Needleman-Wunsch Algorithm Xiantao Jiang1, a,*,xueliang
5th International Conference on Mechatronics, Materials, Chemistry and Computer Engineering (ICMMCCE 2017) Research on Pairwise Sequence Alignment Needleman-Wunsch Algorithm Xiantao Jiang1, a,*,xueliang
INTRODUCTION TO BIOINFORMATICS
 Molecular Biology-2019 1 INTRODUCTION TO BIOINFORMATICS In this section, we want to provide a simple introduction to using the web site of the National Center for Biotechnology Information NCBI) to obtain
Molecular Biology-2019 1 INTRODUCTION TO BIOINFORMATICS In this section, we want to provide a simple introduction to using the web site of the National Center for Biotechnology Information NCBI) to obtain
Gibbs Sampling. Stephen F. Altschul. National Center for Biotechnology Information National Library of Medicine National Institutes of Health
 Gibbs Sampling Stephen F. Altschul National Center for Biotechnology Information National Library of Medicine National Institutes of Health Optimization in High-Dimensional Space Smooth and simple landscapes
Gibbs Sampling Stephen F. Altschul National Center for Biotechnology Information National Library of Medicine National Institutes of Health Optimization in High-Dimensional Space Smooth and simple landscapes
Today s Lecture. Edit graph & alignment algorithms. Local vs global Computational complexity of pairwise alignment Multiple sequence alignment
 Today s Lecture Edit graph & alignment algorithms Smith-Waterman algorithm Needleman-Wunsch algorithm Local vs global Computational complexity of pairwise alignment Multiple sequence alignment 1 Sequence
Today s Lecture Edit graph & alignment algorithms Smith-Waterman algorithm Needleman-Wunsch algorithm Local vs global Computational complexity of pairwise alignment Multiple sequence alignment 1 Sequence
The Effect of Inverse Document Frequency Weights on Indexed Sequence Retrieval. Kevin C. O'Kane. Department of Computer Science
 The Effect of Inverse Document Frequency Weights on Indexed Sequence Retrieval Kevin C. O'Kane Department of Computer Science The University of Northern Iowa Cedar Falls, Iowa okane@cs.uni.edu http://www.cs.uni.edu/~okane
The Effect of Inverse Document Frequency Weights on Indexed Sequence Retrieval Kevin C. O'Kane Department of Computer Science The University of Northern Iowa Cedar Falls, Iowa okane@cs.uni.edu http://www.cs.uni.edu/~okane
Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA.
 Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA. Fasta is used to compare a protein or DNA sequence to all of the
Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA. Fasta is used to compare a protein or DNA sequence to all of the
INTRODUCTION TO BIOINFORMATICS
 Molecular Biology-2017 1 INTRODUCTION TO BIOINFORMATICS In this section, we want to provide a simple introduction to using the web site of the National Center for Biotechnology Information NCBI) to obtain
Molecular Biology-2017 1 INTRODUCTION TO BIOINFORMATICS In this section, we want to provide a simple introduction to using the web site of the National Center for Biotechnology Information NCBI) to obtain
Biologically significant sequence alignments using Boltzmann probabilities
 Biologically significant sequence alignments using Boltzmann probabilities P Clote Department of Biology, Boston College Gasson Hall 16, Chestnut Hill MA 0267 clote@bcedu Abstract In this paper, we give
Biologically significant sequence alignments using Boltzmann probabilities P Clote Department of Biology, Boston College Gasson Hall 16, Chestnut Hill MA 0267 clote@bcedu Abstract In this paper, we give
.. Fall 2011 CSC 570: Bioinformatics Alexander Dekhtyar..
 .. Fall 2011 CSC 570: Bioinformatics Alexander Dekhtyar.. PAM and BLOSUM Matrices Prepared by: Jason Banich and Chris Hoover Background As DNA sequences change and evolve, certain amino acids are more
.. Fall 2011 CSC 570: Bioinformatics Alexander Dekhtyar.. PAM and BLOSUM Matrices Prepared by: Jason Banich and Chris Hoover Background As DNA sequences change and evolve, certain amino acids are more
Sequence alignment theory and applications Session 3: BLAST algorithm
 Sequence alignment theory and applications Session 3: BLAST algorithm Introduction to Bioinformatics online course : IBT Sonal Henson Learning Objectives Understand the principles of the BLAST algorithm
Sequence alignment theory and applications Session 3: BLAST algorithm Introduction to Bioinformatics online course : IBT Sonal Henson Learning Objectives Understand the principles of the BLAST algorithm
Bioinformatics explained: BLAST. March 8, 2007
 Bioinformatics Explained Bioinformatics explained: BLAST March 8, 2007 CLC bio Gustav Wieds Vej 10 8000 Aarhus C Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com info@clcbio.com Bioinformatics
Bioinformatics Explained Bioinformatics explained: BLAST March 8, 2007 CLC bio Gustav Wieds Vej 10 8000 Aarhus C Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com info@clcbio.com Bioinformatics
Acceleration of Algorithm of Smith-Waterman Using Recursive Variable Expansion.
 www.ijarcet.org 54 Acceleration of Algorithm of Smith-Waterman Using Recursive Variable Expansion. Hassan Kehinde Bello and Kazeem Alagbe Gbolagade Abstract Biological sequence alignment is becoming popular
www.ijarcet.org 54 Acceleration of Algorithm of Smith-Waterman Using Recursive Variable Expansion. Hassan Kehinde Bello and Kazeem Alagbe Gbolagade Abstract Biological sequence alignment is becoming popular
Chapter 4: Blast. Chaochun Wei Fall 2014
 Course organization Introduction ( Week 1-2) Course introduction A brief introduction to molecular biology A brief introduction to sequence comparison Part I: Algorithms for Sequence Analysis (Week 3-11)
Course organization Introduction ( Week 1-2) Course introduction A brief introduction to molecular biology A brief introduction to sequence comparison Part I: Algorithms for Sequence Analysis (Week 3-11)
Sequence analysis Pairwise sequence alignment
 UMF11 Introduction to bioinformatics, 25 Sequence analysis Pairwise sequence alignment 1. Sequence alignment Lecturer: Marina lexandersson 12 September, 25 here are two types of sequence alignments, global
UMF11 Introduction to bioinformatics, 25 Sequence analysis Pairwise sequence alignment 1. Sequence alignment Lecturer: Marina lexandersson 12 September, 25 here are two types of sequence alignments, global
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 04: Variations of sequence alignments http://www.pitt.edu/~mcs2/teaching/biocomp/tutorials/global.html Slides adapted from Dr. Shaojie Zhang (University
EECS730: Introduction to Bioinformatics Lecture 04: Variations of sequence alignments http://www.pitt.edu/~mcs2/teaching/biocomp/tutorials/global.html Slides adapted from Dr. Shaojie Zhang (University
Important Example: Gene Sequence Matching. Corrigiendum. Central Dogma of Modern Biology. Genetics. How Nucleotides code for Amino Acids
 Important Example: Gene Sequence Matching Century of Biology Two views of computer science s relationship to biology: Bioinformatics: computational methods to help discover new biology from lots of data
Important Example: Gene Sequence Matching Century of Biology Two views of computer science s relationship to biology: Bioinformatics: computational methods to help discover new biology from lots of data
CS 284A: Algorithms for Computational Biology Notes on Lecture: BLAST. The statistics of alignment scores.
 CS 284A: Algorithms for Computational Biology Notes on Lecture: BLAST. The statistics of alignment scores. prepared by Oleksii Kuchaiev, based on presentation by Xiaohui Xie on February 20th. 1 Introduction
CS 284A: Algorithms for Computational Biology Notes on Lecture: BLAST. The statistics of alignment scores. prepared by Oleksii Kuchaiev, based on presentation by Xiaohui Xie on February 20th. 1 Introduction
Lecture 2 Pairwise sequence alignment. Principles Computational Biology Teresa Przytycka, PhD
 Lecture 2 Pairwise sequence alignment. Principles Computational Biology Teresa Przytycka, PhD Assumptions: Biological sequences evolved by evolution. Micro scale changes: For short sequences (e.g. one
Lecture 2 Pairwise sequence alignment. Principles Computational Biology Teresa Przytycka, PhD Assumptions: Biological sequences evolved by evolution. Micro scale changes: For short sequences (e.g. one
Lecture 9: Core String Edits and Alignments
 Biosequence Algorithms, Spring 2005 Lecture 9: Core String Edits and Alignments Pekka Kilpeläinen University of Kuopio Department of Computer Science BSA Lecture 9: String Edits and Alignments p.1/30 III:
Biosequence Algorithms, Spring 2005 Lecture 9: Core String Edits and Alignments Pekka Kilpeläinen University of Kuopio Department of Computer Science BSA Lecture 9: String Edits and Alignments p.1/30 III:
Principles of Bioinformatics. BIO540/STA569/CSI660 Fall 2010
 Principles of Bioinformatics BIO540/STA569/CSI660 Fall 2010 Lecture 11 Multiple Sequence Alignment I Administrivia Administrivia The midterm examination will be Monday, October 18 th, in class. Closed
Principles of Bioinformatics BIO540/STA569/CSI660 Fall 2010 Lecture 11 Multiple Sequence Alignment I Administrivia Administrivia The midterm examination will be Monday, October 18 th, in class. Closed
Brief review from last class
 Sequence Alignment Brief review from last class DNA is has direction, we will use only one (5 -> 3 ) and generate the opposite strand as needed. DNA is a 3D object (see lecture 1) but we will model it
Sequence Alignment Brief review from last class DNA is has direction, we will use only one (5 -> 3 ) and generate the opposite strand as needed. DNA is a 3D object (see lecture 1) but we will model it
Computational Molecular Biology
 Computational Molecular Biology Erwin M. Bakker Lecture 3, mainly from material by R. Shamir [2] and H.J. Hoogeboom [4]. 1 Pairwise Sequence Alignment Biological Motivation Algorithmic Aspect Recursive
Computational Molecular Biology Erwin M. Bakker Lecture 3, mainly from material by R. Shamir [2] and H.J. Hoogeboom [4]. 1 Pairwise Sequence Alignment Biological Motivation Algorithmic Aspect Recursive
COS 551: Introduction to Computational Molecular Biology Lecture: Oct 17, 2000 Lecturer: Mona Singh Scribe: Jacob Brenner 1. Database Searching
 COS 551: Introduction to Computational Molecular Biology Lecture: Oct 17, 2000 Lecturer: Mona Singh Scribe: Jacob Brenner 1 Database Searching In database search, we typically have a large sequence database
COS 551: Introduction to Computational Molecular Biology Lecture: Oct 17, 2000 Lecturer: Mona Singh Scribe: Jacob Brenner 1 Database Searching In database search, we typically have a large sequence database
Parallel Characteristics of Sequence Alignment Algorithms
 Parallel Characteristics of Sequence Alignment Algorithms Arun I
Parallel Characteristics of Sequence Alignment Algorithms Arun I
Multiple Sequence Alignment: Multidimensional. Biological Motivation
 Multiple Sequence Alignment: Multidimensional Dynamic Programming Boston University Biological Motivation Compare a new sequence with the sequences in a protein family. Proteins can be categorized into
Multiple Sequence Alignment: Multidimensional Dynamic Programming Boston University Biological Motivation Compare a new sequence with the sequences in a protein family. Proteins can be categorized into
SimSearch: A new variant of dynamic programming based on distance series for optimal and near-optimal similarity discovery in biological sequences
 SimSearch: A new variant of dynamic programming based on distance series for optimal and near-optimal similarity discovery in biological sequences Sérgio A. D. Deusdado 1 and Paulo M. M. Carvalho 2 1 ESA,
SimSearch: A new variant of dynamic programming based on distance series for optimal and near-optimal similarity discovery in biological sequences Sérgio A. D. Deusdado 1 and Paulo M. M. Carvalho 2 1 ESA,
Sequence Alignment (chapter 6) p The biological problem p Global alignment p Local alignment p Multiple alignment
 Sequence lignment (chapter 6) p The biological problem p lobal alignment p Local alignment p Multiple alignment Local alignment: rationale p Otherwise dissimilar proteins may have local regions of similarity
Sequence lignment (chapter 6) p The biological problem p lobal alignment p Local alignment p Multiple alignment Local alignment: rationale p Otherwise dissimilar proteins may have local regions of similarity
Sequence comparison: Local alignment
 Sequence comparison: Local alignment Genome 559: Introuction to Statistical an Computational Genomics Prof. James H. Thomas http://faculty.washington.eu/jht/gs559_217/ Review global alignment en traceback
Sequence comparison: Local alignment Genome 559: Introuction to Statistical an Computational Genomics Prof. James H. Thomas http://faculty.washington.eu/jht/gs559_217/ Review global alignment en traceback
Data Mining Technologies for Bioinformatics Sequences
 Data Mining Technologies for Bioinformatics Sequences Deepak Garg Computer Science and Engineering Department Thapar Institute of Engineering & Tecnology, Patiala Abstract Main tool used for sequence alignment
Data Mining Technologies for Bioinformatics Sequences Deepak Garg Computer Science and Engineering Department Thapar Institute of Engineering & Tecnology, Patiala Abstract Main tool used for sequence alignment
FastA & the chaining problem
 FastA & the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 1 Sources for this lecture: Lectures by Volker Heun, Daniel Huson and Knut Reinert,
FastA & the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 1 Sources for this lecture: Lectures by Volker Heun, Daniel Huson and Knut Reinert,
FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:
 FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:56 4001 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem
FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:56 4001 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem
As of August 15, 2008, GenBank contained bases from reported sequences. The search procedure should be
 48 Bioinformatics I, WS 09-10, S. Henz (script by D. Huson) November 26, 2009 4 BLAST and BLAT Outline of the chapter: 1. Heuristics for the pairwise local alignment of two sequences 2. BLAST: search and
48 Bioinformatics I, WS 09-10, S. Henz (script by D. Huson) November 26, 2009 4 BLAST and BLAT Outline of the chapter: 1. Heuristics for the pairwise local alignment of two sequences 2. BLAST: search and
Dynamic Programming & Smith-Waterman algorithm
 m m Seminar: Classical Papers in Bioinformatics May 3rd, 2010 m m 1 2 3 m m Introduction m Definition is a method of solving problems by breaking them down into simpler steps problem need to contain overlapping
m m Seminar: Classical Papers in Bioinformatics May 3rd, 2010 m m 1 2 3 m m Introduction m Definition is a method of solving problems by breaking them down into simpler steps problem need to contain overlapping
Jyoti Lakhani 1, Ajay Khunteta 2, Dharmesh Harwani *3 1 Poornima University, Jaipur & Maharaja Ganga Singh University, Bikaner, Rajasthan, India
 International Journal of Scientific Research in Computer Science, Engineering and Information Technology 2017 IJSRCSEIT Volume 2 Issue 6 ISSN : 2456-3307 Improvisation of Global Pairwise Sequence Alignment
International Journal of Scientific Research in Computer Science, Engineering and Information Technology 2017 IJSRCSEIT Volume 2 Issue 6 ISSN : 2456-3307 Improvisation of Global Pairwise Sequence Alignment
Pairwise Sequence alignment Basic Algorithms
 Pairwise Sequence alignment Basic Algorithms Agenda - Previous Lesson: Minhala - + Biological Story on Biomolecular Sequences - + General Overview of Problems in Computational Biology - Reminder: Dynamic
Pairwise Sequence alignment Basic Algorithms Agenda - Previous Lesson: Minhala - + Biological Story on Biomolecular Sequences - + General Overview of Problems in Computational Biology - Reminder: Dynamic
BLAST MCDB 187. Friday, February 8, 13
 BLAST MCDB 187 BLAST Basic Local Alignment Sequence Tool Uses shortcut to compute alignments of a sequence against a database very quickly Typically takes about a minute to align a sequence against a database
BLAST MCDB 187 BLAST Basic Local Alignment Sequence Tool Uses shortcut to compute alignments of a sequence against a database very quickly Typically takes about a minute to align a sequence against a database
BLAST - Basic Local Alignment Search Tool
 Lecture for ic Bioinformatics (DD2450) April 11, 2013 Searching 1. Input: Query Sequence 2. Database of sequences 3. Subject Sequence(s) 4. Output: High Segment Pairs (HSPs) Sequence Similarity Measures:
Lecture for ic Bioinformatics (DD2450) April 11, 2013 Searching 1. Input: Query Sequence 2. Database of sequences 3. Subject Sequence(s) 4. Output: High Segment Pairs (HSPs) Sequence Similarity Measures:
Biological Sequence Analysis. CSEP 521: Applied Algorithms Final Project. Archie Russell ( ), Jason Hogg ( )
 Biological Sequence Analysis CSEP 521: Applied Algorithms Final Project Archie Russell (0638782), Jason Hogg (0641054) Introduction Background The schematic for every living organism is stored in long
Biological Sequence Analysis CSEP 521: Applied Algorithms Final Project Archie Russell (0638782), Jason Hogg (0641054) Introduction Background The schematic for every living organism is stored in long
Biostrings. Martin Morgan Bioconductor / Fred Hutchinson Cancer Research Center Seattle, WA, USA June 2009
 Biostrings Martin Morgan Bioconductor / Fred Hutchinson Cancer Research Center Seattle, WA, USA 15-19 June 2009 Biostrings Representation DNA, RNA, amino acid, and general biological strings Manipulation
Biostrings Martin Morgan Bioconductor / Fred Hutchinson Cancer Research Center Seattle, WA, USA 15-19 June 2009 Biostrings Representation DNA, RNA, amino acid, and general biological strings Manipulation
A BANDED SMITH-WATERMAN FPGA ACCELERATOR FOR MERCURY BLASTP
 A BANDED SITH-WATERAN FPGA ACCELERATOR FOR ERCURY BLASTP Brandon Harris*, Arpith C. Jacob*, Joseph. Lancaster*, Jeremy Buhler*, Roger D. Chamberlain* *Dept. of Computer Science and Engineering, Washington
A BANDED SITH-WATERAN FPGA ACCELERATOR FOR ERCURY BLASTP Brandon Harris*, Arpith C. Jacob*, Joseph. Lancaster*, Jeremy Buhler*, Roger D. Chamberlain* *Dept. of Computer Science and Engineering, Washington
BLAST, Profile, and PSI-BLAST
 BLAST, Profile, and PSI-BLAST Jianlin Cheng, PhD School of Electrical Engineering and Computer Science University of Central Florida 26 Free for academic use Copyright @ Jianlin Cheng & original sources
BLAST, Profile, and PSI-BLAST Jianlin Cheng, PhD School of Electrical Engineering and Computer Science University of Central Florida 26 Free for academic use Copyright @ Jianlin Cheng & original sources
Acceleration of the Smith-Waterman algorithm for DNA sequence alignment using an FPGA platform
 Acceleration of the Smith-Waterman algorithm for DNA sequence alignment using an FPGA platform Barry Strengholt Matthijs Brobbel Delft University of Technology Faculty of Electrical Engineering, Mathematics
Acceleration of the Smith-Waterman algorithm for DNA sequence alignment using an FPGA platform Barry Strengholt Matthijs Brobbel Delft University of Technology Faculty of Electrical Engineering, Mathematics
A Design of a Hybrid System for DNA Sequence Alignment
 IMECS 2008, 9-2 March, 2008, Hong Kong A Design of a Hybrid System for DNA Sequence Alignment Heba Khaled, Hossam M. Faheem, Tayseer Hasan, Saeed Ghoneimy Abstract This paper describes a parallel algorithm
IMECS 2008, 9-2 March, 2008, Hong Kong A Design of a Hybrid System for DNA Sequence Alignment Heba Khaled, Hossam M. Faheem, Tayseer Hasan, Saeed Ghoneimy Abstract This paper describes a parallel algorithm
6.047 / Computational Biology: Genomes, Networks, Evolution Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
FASTA. Besides that, FASTA package provides SSEARCH, an implementation of the optimal Smith- Waterman algorithm.
 FASTA INTRODUCTION Definition (by David J. Lipman and William R. Pearson in 1985) - Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence
FASTA INTRODUCTION Definition (by David J. Lipman and William R. Pearson in 1985) - Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence
CAP BLAST. BIOINFORMATICS Su-Shing Chen CISE. 8/20/2005 Su-Shing Chen, CISE 1
 CAP 5510-6 BLAST BIOINFORMATICS Su-Shing Chen CISE 8/20/2005 Su-Shing Chen, CISE 1 BLAST Basic Local Alignment Prof Search Su-Shing Chen Tool A Fast Pair-wise Alignment and Database Searching Tool 8/20/2005
CAP 5510-6 BLAST BIOINFORMATICS Su-Shing Chen CISE 8/20/2005 Su-Shing Chen, CISE 1 BLAST Basic Local Alignment Prof Search Su-Shing Chen Tool A Fast Pair-wise Alignment and Database Searching Tool 8/20/2005
Heuristic methods for pairwise alignment:
 Bi03c_1 Unit 03c: Heuristic methods for pairwise alignment: k-tuple-methods k-tuple-methods for alignment of pairs of sequences Bi03c_2 dynamic programming is too slow for large databases Use heuristic
Bi03c_1 Unit 03c: Heuristic methods for pairwise alignment: k-tuple-methods k-tuple-methods for alignment of pairs of sequences Bi03c_2 dynamic programming is too slow for large databases Use heuristic
Shortest Path Algorithm
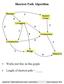 Shortest Path Algorithm C Works just fine on this graph. C Length of shortest path = Copyright 2005 DIMACS BioMath Connect Institute Robert Hochberg Dynamic Programming SP #1 Same Questions, Different
Shortest Path Algorithm C Works just fine on this graph. C Length of shortest path = Copyright 2005 DIMACS BioMath Connect Institute Robert Hochberg Dynamic Programming SP #1 Same Questions, Different
BLAST & Genome assembly
 BLAST & Genome assembly Solon P. Pissis Tomáš Flouri Heidelberg Institute for Theoretical Studies November 17, 2012 1 Introduction Introduction 2 BLAST What is BLAST? The algorithm 3 Genome assembly De
BLAST & Genome assembly Solon P. Pissis Tomáš Flouri Heidelberg Institute for Theoretical Studies November 17, 2012 1 Introduction Introduction 2 BLAST What is BLAST? The algorithm 3 Genome assembly De
Comparison of Sequence Similarity Measures for Distant Evolutionary Relationships
 Comparison of Sequence Similarity Measures for Distant Evolutionary Relationships Abhishek Majumdar, Peter Z. Revesz Department of Computer Science and Engineering, University of Nebraska-Lincoln, Lincoln,
Comparison of Sequence Similarity Measures for Distant Evolutionary Relationships Abhishek Majumdar, Peter Z. Revesz Department of Computer Science and Engineering, University of Nebraska-Lincoln, Lincoln,
Computational Molecular Biology
 Computational Molecular Biology Erwin M. Bakker Lecture 2 Materials used from R. Shamir [2] and H.J. Hoogeboom [4]. 1 Molecular Biology Sequences DNA A, T, C, G RNA A, U, C, G Protein A, R, D, N, C E,
Computational Molecular Biology Erwin M. Bakker Lecture 2 Materials used from R. Shamir [2] and H.J. Hoogeboom [4]. 1 Molecular Biology Sequences DNA A, T, C, G RNA A, U, C, G Protein A, R, D, N, C E,
Research Article International Journals of Advanced Research in Computer Science and Software Engineering ISSN: X (Volume-7, Issue-6)
 International Journals of Advanced Research in Computer Science and Software Engineering ISSN: 77-18X (Volume-7, Issue-6) Research Article June 017 DDGARM: Dotlet Driven Global Alignment with Reduced Matrix
International Journals of Advanced Research in Computer Science and Software Engineering ISSN: 77-18X (Volume-7, Issue-6) Research Article June 017 DDGARM: Dotlet Driven Global Alignment with Reduced Matrix
On the Efficacy of Haskell for High Performance Computational Biology
 On the Efficacy of Haskell for High Performance Computational Biology Jacqueline Addesa Academic Advisors: Jeremy Archuleta, Wu chun Feng 1. Problem and Motivation Biologists can leverage the power of
On the Efficacy of Haskell for High Performance Computational Biology Jacqueline Addesa Academic Advisors: Jeremy Archuleta, Wu chun Feng 1. Problem and Motivation Biologists can leverage the power of
Biological Sequence Matching Using Fuzzy Logic
 International Journal of Scientific & Engineering Research Volume 2, Issue 7, July-2011 1 Biological Sequence Matching Using Fuzzy Logic Nivit Gill, Shailendra Singh Abstract: Sequence alignment is the
International Journal of Scientific & Engineering Research Volume 2, Issue 7, July-2011 1 Biological Sequence Matching Using Fuzzy Logic Nivit Gill, Shailendra Singh Abstract: Sequence alignment is the
Introduction to BLAST with Protein Sequences. Utah State University Spring 2014 STAT 5570: Statistical Bioinformatics Notes 6.2
 Introduction to BLAST with Protein Sequences Utah State University Spring 2014 STAT 5570: Statistical Bioinformatics Notes 6.2 1 References Chapter 2 of Biological Sequence Analysis (Durbin et al., 2001)
Introduction to BLAST with Protein Sequences Utah State University Spring 2014 STAT 5570: Statistical Bioinformatics Notes 6.2 1 References Chapter 2 of Biological Sequence Analysis (Durbin et al., 2001)
BIOL 7020 Special Topics Cell/Molecular: Molecular Phylogenetics. Spring 2010 Section A
 BIOL 7020 Special Topics Cell/Molecular: Molecular Phylogenetics. Spring 2010 Section A Steve Thompson: stthompson@valdosta.edu http://www.bioinfo4u.net 1 Similarity searching and homology First, just
BIOL 7020 Special Topics Cell/Molecular: Molecular Phylogenetics. Spring 2010 Section A Steve Thompson: stthompson@valdosta.edu http://www.bioinfo4u.net 1 Similarity searching and homology First, just
Similarity Searches on Sequence Databases
 Similarity Searches on Sequence Databases Lorenza Bordoli Swiss Institute of Bioinformatics EMBnet Course, Zürich, October 2004 Swiss Institute of Bioinformatics Swiss EMBnet node Outline Importance of
Similarity Searches on Sequence Databases Lorenza Bordoli Swiss Institute of Bioinformatics EMBnet Course, Zürich, October 2004 Swiss Institute of Bioinformatics Swiss EMBnet node Outline Importance of
2. Take a few minutes to look around the site. The goal is to familiarize yourself with a few key components of the NCBI.
 2 Navigating the NCBI Instructions Aim: To become familiar with the resources available at the National Center for Bioinformatics (NCBI) and the search engine Entrez. Instructions: Write the answers to
2 Navigating the NCBI Instructions Aim: To become familiar with the resources available at the National Center for Bioinformatics (NCBI) and the search engine Entrez. Instructions: Write the answers to
CMSC423: Bioinformatic Algorithms, Databases and Tools Lecture 8. Note
 MS: Bioinformatic lgorithms, Databases and ools Lecture 8 Sequence alignment: inexact alignment dynamic programming, gapped alignment Note Lecture 7 suffix trees and suffix arrays will be rescheduled Exact
MS: Bioinformatic lgorithms, Databases and ools Lecture 8 Sequence alignment: inexact alignment dynamic programming, gapped alignment Note Lecture 7 suffix trees and suffix arrays will be rescheduled Exact
Sequence Alignment. part 2
 Sequence Alignment part 2 Dynamic programming with more realistic scoring scheme Using the same initial sequences, we ll look at a dynamic programming example with a scoring scheme that selects for matches
Sequence Alignment part 2 Dynamic programming with more realistic scoring scheme Using the same initial sequences, we ll look at a dynamic programming example with a scoring scheme that selects for matches
ICB Fall G4120: Introduction to Computational Biology. Oliver Jovanovic, Ph.D. Columbia University Department of Microbiology
 ICB Fall 2008 G4120: Computational Biology Oliver Jovanovic, Ph.D. Columbia University Department of Microbiology Copyright 2008 Oliver Jovanovic, All Rights Reserved. The Digital Language of Computers
ICB Fall 2008 G4120: Computational Biology Oliver Jovanovic, Ph.D. Columbia University Department of Microbiology Copyright 2008 Oliver Jovanovic, All Rights Reserved. The Digital Language of Computers
Lectures by Volker Heun, Daniel Huson and Knut Reinert, in particular last years lectures
 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 4.1 Sources for this lecture Lectures by Volker Heun, Daniel Huson and Knut
4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 4.1 Sources for this lecture Lectures by Volker Heun, Daniel Huson and Knut
HIDDEN MARKOV MODELS AND SEQUENCE ALIGNMENT
 HIDDEN MARKOV MODELS AND SEQUENCE ALIGNMENT - Swarbhanu Chatterjee. Hidden Markov models are a sophisticated and flexible statistical tool for the study of protein models. Using HMMs to analyze proteins
HIDDEN MARKOV MODELS AND SEQUENCE ALIGNMENT - Swarbhanu Chatterjee. Hidden Markov models are a sophisticated and flexible statistical tool for the study of protein models. Using HMMs to analyze proteins
BLAST & Genome assembly
 BLAST & Genome assembly Solon P. Pissis Tomáš Flouri Heidelberg Institute for Theoretical Studies May 15, 2014 1 BLAST What is BLAST? The algorithm 2 Genome assembly De novo assembly Mapping assembly 3
BLAST & Genome assembly Solon P. Pissis Tomáš Flouri Heidelberg Institute for Theoretical Studies May 15, 2014 1 BLAST What is BLAST? The algorithm 2 Genome assembly De novo assembly Mapping assembly 3
Comparison MICHAEL S. WATERMAN*
 ADVANCBS IN APPLIED MATHEMATICS 6,129-134 (1985) : Dynamic Programming Algorithms for Picture Comparison MICHAEL S. WATERMAN* Departments of Mathematics and Biological Sciences, Univeniry of Southern California.
ADVANCBS IN APPLIED MATHEMATICS 6,129-134 (1985) : Dynamic Programming Algorithms for Picture Comparison MICHAEL S. WATERMAN* Departments of Mathematics and Biological Sciences, Univeniry of Southern California.
Sequence Identification using BLAST
 Sequence Identification using BLAST Vivek Krishnakumar JCVI Genomic Science and Leadership Workshop Presented on: 05/26/2016 Overview Introduction Why compare sequences? Sequence alignment steps Causes
Sequence Identification using BLAST Vivek Krishnakumar JCVI Genomic Science and Leadership Workshop Presented on: 05/26/2016 Overview Introduction Why compare sequences? Sequence alignment steps Causes
CS313 Exercise 4 Cover Page Fall 2017
 CS313 Exercise 4 Cover Page Fall 2017 Due by the start of class on Thursday, October 12, 2017. Name(s): In the TIME column, please estimate the time you spent on the parts of this exercise. Please try
CS313 Exercise 4 Cover Page Fall 2017 Due by the start of class on Thursday, October 12, 2017. Name(s): In the TIME column, please estimate the time you spent on the parts of this exercise. Please try
Comparative Analysis of Protein Alignment Algorithms in Parallel environment using CUDA
 Comparative Analysis of Protein Alignment Algorithms in Parallel environment using BLAST versus Smith-Waterman Shadman Fahim shadmanbracu09@gmail.com Shehabul Hossain rudrozzal@gmail.com Gulshan Jubaed
Comparative Analysis of Protein Alignment Algorithms in Parallel environment using BLAST versus Smith-Waterman Shadman Fahim shadmanbracu09@gmail.com Shehabul Hossain rudrozzal@gmail.com Gulshan Jubaed
A Scalable Coprocessor for Bioinformatic Sequence Alignments
 A Scalable Coprocessor for Bioinformatic Sequence Alignments Scott F. Smith Department of Electrical and Computer Engineering Boise State University Boise, ID, U.S.A. Abstract A hardware coprocessor for
A Scalable Coprocessor for Bioinformatic Sequence Alignments Scott F. Smith Department of Electrical and Computer Engineering Boise State University Boise, ID, U.S.A. Abstract A hardware coprocessor for
USING AN EXTENDED SUFFIX TREE TO SPEED-UP SEQUENCE ALIGNMENT
 IADIS International Conference Applied Computing 2006 USING AN EXTENDED SUFFIX TREE TO SPEED-UP SEQUENCE ALIGNMENT Divya R. Singh Software Engineer Microsoft Corporation, Redmond, WA 98052, USA Abdullah
IADIS International Conference Applied Computing 2006 USING AN EXTENDED SUFFIX TREE TO SPEED-UP SEQUENCE ALIGNMENT Divya R. Singh Software Engineer Microsoft Corporation, Redmond, WA 98052, USA Abdullah
Scalable Hardware Accelerator for Comparing DNA and Protein Sequences
 Scalable Hardware Accelerator for Comparing DNA and Protein Sequences Philippe Faes, Bram Minnaert, Mark Christiaens, Eric Bonnet, Yvan Saeys, Dirk Stroobandt, Yves Van de Peer Abstract Comparing genetic
Scalable Hardware Accelerator for Comparing DNA and Protein Sequences Philippe Faes, Bram Minnaert, Mark Christiaens, Eric Bonnet, Yvan Saeys, Dirk Stroobandt, Yves Van de Peer Abstract Comparing genetic
