Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging
|
|
|
- Phillip Johnson
- 5 years ago
- Views:
Transcription
1 1 CS 9 Final Project Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging Feiyu Chen Department of Electrical Engineering ABSTRACT Subject motion is a significant problem in magnetic resonance imaging (MRI). In this project, unsupervised learning using Kmeans clustering, and supervised learning using logistic regression () and support vector machine (SVM) are implemented and applied to solving (1) classification of MR images with/without motion, and () estimation of current motion state based on previous low frequency. 15 MR images with labeling of motion/no-motion and are collected as the training and testing data. Over 1 features were designed and manually selected for each learning approach based on principle component analysis (PCA). Results show that, for unsupervised learning, the K-means algorithm achieves an accuracy of ~3% for classification of MR images and a root-mean-squared error (RMSE) of 5.3% in prediction based on. For supervised learning, SVM using Gaussian kernels achieves the best accuracy of ~9% for classification of non-motion and motion images, and a RMSE of ~1.7% for estimation of motion states based on. These approaches provide a possibility for advanced reconstruction algorithms to perform more accurate motion corrections with or without a large training set, and may improve the quality of clinical images. INTRODUCTION Among all the medical imaging modalities, MRI is one of the best candidates for evaluation and diagnosis of diseases, because of its high spatial-resolution, good softtissue contrast and no ionizing radiation. However, MRI requires relatively long scan times, and therefore is more sensitive to subject motion during exams. Subject motion can create imaging artifacts such as imaging blurring, ghosting, and reduced SNR, and degrade image quality. This limits its wide use in clinical diagnoses. In order to overcome this disadvantage, several approaches have been developed for improving speed and reducing motion artifacts during reconstructions of acquired MRI signals. However, these approaches require classification and estimation of motion, which is usually difficult to achieve. The fundamental goal of this project is to classify the images with/without motion based on DICOM images. The second objective of this project is to estimate the current motion state based on previous information on patient positions. With the help of unsupervised and supervised learning models, hopefully MRI scanners can automatically determine the significance of motion artifacts, and perform appropriate corrections and reconstructions based on prior information of motion. has not been applied to prediction of motion states and subject positions, which is the main focus of this project. DATASET & FEATURES 1. Motion artifacts in MRI A good understanding of the motion artifacts in MRI is helpful for designing features for learning. As data are directly acquired in the frequency domain in MRI, motion artifacts of the displayed images are different from common motion artifacts in optical photos. According to the Fourier Transform theories, subject motion will cause phase shift in the frequency domain. Assuming several parts of the data in frequency domain are corrupted by motion, the image will show motion artifacts as blurring and ghosting, which contain shifted replicas of the original image without motion, as shown in Fig.1. Fig. 1 Illustration of motion artifacts in abdominal MRI RELATED WORK. Low frequency for motion Both unsupervised learning and supervised learning have been applied to solving MRI problems. However, only unsupervised learning has been used to classify MR images for cardiac perfusion applications[1]. Few attempts have been made to classify abdominal images and mixed images with different body parts. Supervised learning approaches, such as support vector machine (SVM), have been used to classify functional MRI data of brain states[][3]. However, it Low frequency signals are acquired before and after each image frame as navigators for the real-time motion state, as the distance of subject motion can be estimated directly from the phase of the low frequency signal (Fig. ). However, in contrast-enhanced dynamic MRI, many other factors contribute to the motion estimation, among which contrast change is the most significant one. This leads to wrong estimation of current motion state. In this project, this problem would be solved based on previous non-
2 CS 9 Final Project contaminated signals, which would be beneficial for time points before the current point are extracted in a clinical imaging. sliding-window pattern. Estimation of motion severity Fig. Illustration of low frequency in MRI 3. Samples of images and navigators For the first objective of the project, 57 images with motion and images without motion were collected in experiments conducted at Stanford, and online searches of public MR images. Two sets are formed based on these images. The first set contains images with similar regions of the abdomen. The second set contains images with different body parts, including the brain, the liver, the pelvis, and the heart. For the second objective, are acquired in a continuous scan with free breathing, where one breathing period is ~5 time points.. Pre-processing and features.1 Images Time/Per acquisition Pre-processing of images includes the following steps: (a) Resize images into the same size of 5*5; (b) Divide each image equally into blocks; (c) Compute the features for each block. The features being tested include: (1) The mean value of all the pixels in a block; () The rank of the image, estimated from the singular value decomposition (SVD); (3) The standard deviation of pixels in a block; () The number of points on edges; (5) The sum of intensity for the 5% center of images; () The sum of intensity for the edges of the image; (7) The ratio between 5 and ; () The total variation of the image; (9) The ratio between the center intensity and edge intensity of the frequency domain; (1) The top 1% eigenvalues of the image; (11) Phase of the Fourier transform of the image.. Low frequency Before extracting features of, they are first discretized into 1,, or states. The previous 5 The features being tested include: (1) A vector of previous signal values; () The range of previous signal values; (3) The mean of previous signal values; () The slope from the beginning to the end; (5) The center value of previous signals; () The coefficients of D polynomial fitting; (7) The index of points with max and min values; () The beginning slope and ending slope; (9) The coefficients of linear fitting..3 Normalizations and principle components analysis (PCA) After extracting the features, the values of each feature are normalized with the statistical average µ and standard deviation σ with x' = x µ. PCA is subsequently σ performed on all training data to project the features onto a subspace with maximized differences and efficiency. A threshold of 5% energy of the first principle component is used to determine the new feature space with reduced dimension. METHODS 1. Learning models 1.1 Unsupervised learning using K-means clustering The unsupervised learning model being implemented is the K-means method. This method uses the geometric distance between samples as the metric for classify and cluster a set of samples in an iterative way. For classification of images with motion and without motion, the number of clusters is set to be two. For estimations of the severity of motion, the number of clusters is set to the number of discretized states. 1. Supervised learning using and SVM uses the sigmoid function as the regression model to fit the labels with input features. It is implemented in MATLAB using the stochastic gradient ascent algorithm with coefficient α=.5. SVM uses support vectors to determine the boundary between classes with different labels. The kernels in the dual optimization problem of SVM determine the hyperspace of features. Linear kernels () and Gaussian kernels () are two common kernels and are implemented in this project. The LIBLINEAR and LIBSVM toolbox [] are used as implementations of and, respectively.. Evaluations.1 Evaluation of classifications based on MR images
3 CS 9 Final Project 3 Evaluations of the learning theories are conducted based 1. Supervised learning using and SVM on the true positive (TP), true negative (TN), false positive All the three supervised learning approaches perform well (FP), and false negative (FN) counts in leave-one-out for the first dataset. SVM with Gaussian kernels achieves crossing-validation experiments, as the size of training data slightly higher accuracy (% higher). For the second is relatively small. The accuracy, specificity, and sensitivity dataset, the three approaches achieve similar accuracy of of the learning theory are further computed based on these ~9%. However, achieves higher specificity (9%), indices. while achieves higher sensitivity (9%).. Evaluation of classifications based on lowfrequency Since the number of states is much greater than two, using true and false cannot comprehensively show the accuracy of each learning model. Therefore, the root-mean-squared error (RMSE) is calculated between the estimates and the reference labels in two-fold cross-validation experiments, due to the large size of training data. The RMSE is normalized with the total number of states to provide a fair comparison between different discretization of motion states. RMSE Normalized = (S est S true ) N Test N States where S means the signal, N test is the length of test data, and N states is the number of discretized states. RESULTS & DISCUSSION 1. Classification of images with and without motion The accuracy of each learning approach is evaluated for (1) a set of abdominal images, and () a set of images of different body parts. All learning approaches achieve similar performance for the first dataset. Compared with unsupervised learning using K-means, supervised learning approaches generally achieve higher accuracy of ~9% for the images of different body parts. 1.1 Unsupervised learning using K-means The K-means approach reaches an accuracy of 3% for both cases, as shown in Table 1. Compared with the first dataset, K-means for the second dataset has higher sensitivity (1%) but lower specificity (3%). Table 1 Unsupervised learning for classification of MR images with motion and without motion Images Dataset 1: Abdominal images only Dataset : images with variety of regions Features [1 9] [1 3 9] #Blocks [ ] [ ] TP 9 57 TN 3 3 FP FN 9 Sensitivity 7% 1% Specificity 9% 3% Accuracy 3% 3% Table Supervised learning for classification of MR images with motion and without motion Images Dataset 1: Abdominal images only Dataset : images with variety of regions Features [1 9] [ ] #Blocks [ ] [ ] Models TP TN FP FN Sensitivity 97% 97% 1% % % 9% Specificity 1% 1% 1% 9% 9% 1% Accuracy 9% 9% 1% 9% 9% 9% 1.3 Analysis of accuracy Errors in unsupervised learning with K-means The main error of K-means for classification of the second dataset comes from mislabeling of points on the boundary of the two classes, as shown in the red ellipsoid in Fig. 3. The scatter plot of correct labeling also reveals it is difficult to separate the two image classes using only these two features, because of the mixture of labels in the center. Feature component 1 Correct labeling for reference K-means clustering results Without motion With motion Feature component Feature component Fig. 3 Illustration of K-means clustering for MR images with/without motion in the second dataset. The x and y axes denote the two major feature components 1.3. Analysis of #blocks for supervised learning The size/total number of blocks influences the accuracy of all supervised learning methods. As shown in Fig., both and show fluctuations of accuracy with changes of the number of blocks, while the accuracy of remains stable for different number of blocks. However, all the three approaches show a trend of accuracy increase with increasing number of blocks. A possible reason is, the efficiency of information increases with increased number of blocks, which provides more features for training and classification.
4 CS 9 Final Project.9. Supervised learning using and SVM Accuracy of classification Number of groups in each side Fig. The accuracy of classification varies with changes in number of blocks in each side Training set size for supervised learning Training set size is another key factor for the accuracy of supervised learning. As shown in Fig. 5, all the three learning approaches demonstrate a tendency of accuracy increase with increased training set size. When the training set size is less than, there are fluctuations in the classification accuracy for all three methods. Therefore, it is necessary to maintain a training set size of over. Supervised learning approaches achieve higher accuracy of less than ~% compared with the K-means approach. achieves the lowest RMSE among all the three supervised learning approaches, while performs better than (Table 3). The signal evolution curves also show fits the curve better than. has higher accuracy and less noise than at the peaks and troughs of the curve (Fig. 5) K-means clustering for states K means clustering for States Logis&c regression for states Logistic Regression for States.95 Accuracy of classification SVM using linear kernels for states Support Vector Machine for States Training set size Fig. 5 The accuracy of classification varies with changes of the training set size. Estimation of the position of motion based on lowfrequency.1 Unsupervised learning using K-means As there are multiple classes in this classification problem, and the differences between difference classes are relatively small, unsupervised learning may encounter some difficulty in generating appropriate clusters. However, with appropriate features, the K-means approach achieves a RMSE of less than 1% for testing data points with feature set [1 7 9], as shown in Table 3. The curve in Fig.5 also shows K-means can follow the signal evolutions well, despite some minor delays in the decreasing periods. Table 3 RMSE for estimation of motion states based on and training data #States Features K-means [1 7 9] 5.% 3.51%.7% 1.7% [1 7 9].% 3.%.3% 1.% 1 [1 7 9] 1.7% 3.1%.% 1.5% SVM using Gaussian kernels for states Support Vector Machine for States Fig. 5 Illustration of estimation based on low frequency in MRI using K-means,,, and for signals with discretized states
5 CS 9 Final Project 5.3 Analysis of performance similar or fluctuates slightly as the number of states.3.1 Analysis of the size of prior information increases. This shows the number of discrete states has little impact on the estimation accuracy. The size of prior information, i.e., the length of time series being used to generate features, may have an effect on the accuracy of learning models. In specific cases with periodic motion, the period of breathing is ~5 time points, which makes the choice more important. As shown in Fig., only shows an improvement of accuracy (RMSE decrease) with increasing length of prior information. K-means and remains insensitive to the size of previous points, and shows significant fluctuations with this change. RMSE of estimation Fig. Changes of RMSE with increasing size of prior information (#points used for feature extractions) using a training set size of 5 for estimation of motion states based on.3. Analysis of training set size for supervised learning The relationship between the RMSE of testing data and training size is shown in Fig. 7. As the training size increases, the RMSE of,, and for supervised learning decrease. Among the three supervised learning methods, preserves the highest accuracy even with few training data. RMSE of testing data Size of previous points used K means K means CONCLUSION & FUTURE WORK 1. Choice of learning models for motion classification and estimation If a large amount of training data with correct labeling is available, supervised learning should be used for both classification of images with/without motion and estimation of motion states based on. Among the three supervised learning approaches, SVM with Gaussian kernels provides the most stable and accurate labeling for both cases. has slightly worse performance than, but it's also easier to understand and implement. If a reasonable amount of training data cannot be achieved, the K-means approach can give acceptable classification of images with/without motion, and good estimation of motion states. While its accuracy is not as good as supervised learning algorithms, it is not susceptible to the length of time points used to generate features when doing estimation based on.. Limitations of current approaches First, the size of training sets for classification of images with/without motion is limited, since no big public database is available online. This limits the performance of all the three supervised learning approaches (Fig. 5). Second, there are many kinds of subject motion during clinical studies. Therefore, a multi-level classification may be useful to improve the accuracy. Third, due to hardware limitations, the motion signal can only be discretized to states. More states would be beneficial to subsequent motion corrections and reconstructions. 3. Future work Future work can be done in (1) exploring more advanced learning models, () collecting more samples of images with/without motion, and (3) conducting motion correction based on the results of classification and estimation of motion to see how machine learning can help with motion corrections and reconstructions of MR images Training Size Fig. 7 RMSE of the testing data with increasing training size using 5 points of previous signals for estimation of motion states based on.3.3 Number of discretized states According to Table 3, for unsupervised learning with K- Means, the estimation accuracy increases with the number of discretized states. However, for all three supervised learning approaches, the estimation accuracy remains REFERENCES [1] Stegmann, Mikkel B., Hildur Ólafsdóttir, and Henrik BW Larsson. "Unsupervised motion-compensation of multi-slice cardiac perfusion MRI." Medical Image Analysis 9. (5): [] Mourão-Miranda, Janaina, et al. "Classifying brain states and determining the discriminating activation patterns: support vector machine on functional MRI data." NeuroImage. (5): [3] Wang, Ze, et al. "Assessment of functional development in normal infant brain using arterial spin labeled perfusion MRI." Neuroimage 39.3 (): [] Chih-Chung Chang and Chih-Jen Lin, LIBSVM : a library for support vector machines. ACM Transactions on Intelligent Systems and Technology, :7:1-- 7:7, 11. Software available at
Segmenting Lesions in Multiple Sclerosis Patients James Chen, Jason Su
 Segmenting Lesions in Multiple Sclerosis Patients James Chen, Jason Su Radiologists and researchers spend countless hours tediously segmenting white matter lesions to diagnose and study brain diseases.
Segmenting Lesions in Multiple Sclerosis Patients James Chen, Jason Su Radiologists and researchers spend countless hours tediously segmenting white matter lesions to diagnose and study brain diseases.
MEDICAL IMAGE ANALYSIS
 SECOND EDITION MEDICAL IMAGE ANALYSIS ATAM P. DHAWAN g, A B IEEE Engineering in Medicine and Biology Society, Sponsor IEEE Press Series in Biomedical Engineering Metin Akay, Series Editor +IEEE IEEE PRESS
SECOND EDITION MEDICAL IMAGE ANALYSIS ATAM P. DHAWAN g, A B IEEE Engineering in Medicine and Biology Society, Sponsor IEEE Press Series in Biomedical Engineering Metin Akay, Series Editor +IEEE IEEE PRESS
Functional MRI in Clinical Research and Practice Preprocessing
 Functional MRI in Clinical Research and Practice Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization
Functional MRI in Clinical Research and Practice Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization
SPM8 for Basic and Clinical Investigators. Preprocessing. fmri Preprocessing
 SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
Improving Positron Emission Tomography Imaging with Machine Learning David Fan-Chung Hsu CS 229 Fall
 Improving Positron Emission Tomography Imaging with Machine Learning David Fan-Chung Hsu (fcdh@stanford.edu), CS 229 Fall 2014-15 1. Introduction and Motivation High- resolution Positron Emission Tomography
Improving Positron Emission Tomography Imaging with Machine Learning David Fan-Chung Hsu (fcdh@stanford.edu), CS 229 Fall 2014-15 1. Introduction and Motivation High- resolution Positron Emission Tomography
CS 229 Final Project Report Learning to Decode Cognitive States of Rat using Functional Magnetic Resonance Imaging Time Series
 CS 229 Final Project Report Learning to Decode Cognitive States of Rat using Functional Magnetic Resonance Imaging Time Series Jingyuan Chen //Department of Electrical Engineering, cjy2010@stanford.edu//
CS 229 Final Project Report Learning to Decode Cognitive States of Rat using Functional Magnetic Resonance Imaging Time Series Jingyuan Chen //Department of Electrical Engineering, cjy2010@stanford.edu//
Enhao Gong, PhD Candidate, Electrical Engineering, Stanford University Dr. John Pauly, Professor in Electrical Engineering, Stanford University Dr.
 Enhao Gong, PhD Candidate, Electrical Engineering, Stanford University Dr. John Pauly, Professor in Electrical Engineering, Stanford University Dr. Greg Zaharchuk, Associate Professor in Radiology, Stanford
Enhao Gong, PhD Candidate, Electrical Engineering, Stanford University Dr. John Pauly, Professor in Electrical Engineering, Stanford University Dr. Greg Zaharchuk, Associate Professor in Radiology, Stanford
Basic fmri Design and Analysis. Preprocessing
 Basic fmri Design and Analysis Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial filtering
Basic fmri Design and Analysis Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial filtering
EPI Data Are Acquired Serially. EPI Data Are Acquired Serially 10/23/2011. Functional Connectivity Preprocessing. fmri Preprocessing
 Functional Connectivity Preprocessing Geometric distortion Head motion Geometric distortion Head motion EPI Data Are Acquired Serially EPI Data Are Acquired Serially descending 1 EPI Data Are Acquired
Functional Connectivity Preprocessing Geometric distortion Head motion Geometric distortion Head motion EPI Data Are Acquired Serially EPI Data Are Acquired Serially descending 1 EPI Data Are Acquired
SPM8 for Basic and Clinical Investigators. Preprocessing
 SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
Correction of Partial Volume Effects in Arterial Spin Labeling MRI
 Correction of Partial Volume Effects in Arterial Spin Labeling MRI By: Tracy Ssali Supervisors: Dr. Keith St. Lawrence and Udunna Anazodo Medical Biophysics 3970Z Six Week Project April 13 th 2012 Introduction
Correction of Partial Volume Effects in Arterial Spin Labeling MRI By: Tracy Ssali Supervisors: Dr. Keith St. Lawrence and Udunna Anazodo Medical Biophysics 3970Z Six Week Project April 13 th 2012 Introduction
Artificial Neural Networks (Feedforward Nets)
 Artificial Neural Networks (Feedforward Nets) y w 03-1 w 13 y 1 w 23 y 2 w 01 w 21 w 22 w 02-1 w 11 w 12-1 x 1 x 2 6.034 - Spring 1 Single Perceptron Unit y w 0 w 1 w n w 2 w 3 x 0 =1 x 1 x 2 x 3... x
Artificial Neural Networks (Feedforward Nets) y w 03-1 w 13 y 1 w 23 y 2 w 01 w 21 w 22 w 02-1 w 11 w 12-1 x 1 x 2 6.034 - Spring 1 Single Perceptron Unit y w 0 w 1 w n w 2 w 3 x 0 =1 x 1 x 2 x 3... x
Statistical Analysis of Neuroimaging Data. Phebe Kemmer BIOS 516 Sept 24, 2015
 Statistical Analysis of Neuroimaging Data Phebe Kemmer BIOS 516 Sept 24, 2015 Review from last time Structural Imaging modalities MRI, CAT, DTI (diffusion tensor imaging) Functional Imaging modalities
Statistical Analysis of Neuroimaging Data Phebe Kemmer BIOS 516 Sept 24, 2015 Review from last time Structural Imaging modalities MRI, CAT, DTI (diffusion tensor imaging) Functional Imaging modalities
INTRODUCTION TO MACHINE LEARNING. Measuring model performance or error
 INTRODUCTION TO MACHINE LEARNING Measuring model performance or error Is our model any good? Context of task Accuracy Computation time Interpretability 3 types of tasks Classification Regression Clustering
INTRODUCTION TO MACHINE LEARNING Measuring model performance or error Is our model any good? Context of task Accuracy Computation time Interpretability 3 types of tasks Classification Regression Clustering
Hybrid Approach for MRI Human Head Scans Classification using HTT based SFTA Texture Feature Extraction Technique
 Volume 118 No. 17 2018, 691-701 ISSN: 1311-8080 (printed version); ISSN: 1314-3395 (on-line version) url: http://www.ijpam.eu ijpam.eu Hybrid Approach for MRI Human Head Scans Classification using HTT
Volume 118 No. 17 2018, 691-701 ISSN: 1311-8080 (printed version); ISSN: 1314-3395 (on-line version) url: http://www.ijpam.eu ijpam.eu Hybrid Approach for MRI Human Head Scans Classification using HTT
Introduction to Neuroimaging Janaina Mourao-Miranda
 Introduction to Neuroimaging Janaina Mourao-Miranda Neuroimaging techniques have changed the way neuroscientists address questions about functional anatomy, especially in relation to behavior and clinical
Introduction to Neuroimaging Janaina Mourao-Miranda Neuroimaging techniques have changed the way neuroscientists address questions about functional anatomy, especially in relation to behavior and clinical
Louis Fourrier Fabien Gaie Thomas Rolf
 CS 229 Stay Alert! The Ford Challenge Louis Fourrier Fabien Gaie Thomas Rolf Louis Fourrier Fabien Gaie Thomas Rolf 1. Problem description a. Goal Our final project is a recent Kaggle competition submitted
CS 229 Stay Alert! The Ford Challenge Louis Fourrier Fabien Gaie Thomas Rolf Louis Fourrier Fabien Gaie Thomas Rolf 1. Problem description a. Goal Our final project is a recent Kaggle competition submitted
Dimension Reduction CS534
 Dimension Reduction CS534 Why dimension reduction? High dimensionality large number of features E.g., documents represented by thousands of words, millions of bigrams Images represented by thousands of
Dimension Reduction CS534 Why dimension reduction? High dimensionality large number of features E.g., documents represented by thousands of words, millions of bigrams Images represented by thousands of
FMRI Pre-Processing and Model- Based Statistics
 FMRI Pre-Processing and Model- Based Statistics Brief intro to FMRI experiments and analysis FMRI pre-stats image processing Simple Single-Subject Statistics Multi-Level FMRI Analysis Advanced FMRI Analysis
FMRI Pre-Processing and Model- Based Statistics Brief intro to FMRI experiments and analysis FMRI pre-stats image processing Simple Single-Subject Statistics Multi-Level FMRI Analysis Advanced FMRI Analysis
CPSC 340: Machine Learning and Data Mining. Principal Component Analysis Fall 2016
 CPSC 340: Machine Learning and Data Mining Principal Component Analysis Fall 2016 A2/Midterm: Admin Grades/solutions will be posted after class. Assignment 4: Posted, due November 14. Extra office hours:
CPSC 340: Machine Learning and Data Mining Principal Component Analysis Fall 2016 A2/Midterm: Admin Grades/solutions will be posted after class. Assignment 4: Posted, due November 14. Extra office hours:
Clinical Importance. Aortic Stenosis. Aortic Regurgitation. Ultrasound vs. MRI. Carotid Artery Stenosis
 Clinical Importance Rapid cardiovascular flow quantitation using sliceselective Fourier velocity encoding with spiral readouts Valve disease affects 10% of patients with heart disease in the U.S. Most
Clinical Importance Rapid cardiovascular flow quantitation using sliceselective Fourier velocity encoding with spiral readouts Valve disease affects 10% of patients with heart disease in the U.S. Most
Machine Learning for Medical Image Analysis. A. Criminisi
 Machine Learning for Medical Image Analysis A. Criminisi Overview Introduction to machine learning Decision forests Applications in medical image analysis Anatomy localization in CT Scans Spine Detection
Machine Learning for Medical Image Analysis A. Criminisi Overview Introduction to machine learning Decision forests Applications in medical image analysis Anatomy localization in CT Scans Spine Detection
Compressed Sensing Reconstructions for Dynamic Contrast Enhanced MRI
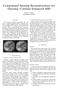 1 Compressed Sensing Reconstructions for Dynamic Contrast Enhanced MRI Kevin T. Looby klooby@stanford.edu ABSTRACT The temporal resolution necessary for dynamic contrast enhanced (DCE) magnetic resonance
1 Compressed Sensing Reconstructions for Dynamic Contrast Enhanced MRI Kevin T. Looby klooby@stanford.edu ABSTRACT The temporal resolution necessary for dynamic contrast enhanced (DCE) magnetic resonance
CS145: INTRODUCTION TO DATA MINING
 CS145: INTRODUCTION TO DATA MINING Clustering Evaluation and Practical Issues Instructor: Yizhou Sun yzsun@cs.ucla.edu November 7, 2017 Learnt Clustering Methods Vector Data Set Data Sequence Data Text
CS145: INTRODUCTION TO DATA MINING Clustering Evaluation and Practical Issues Instructor: Yizhou Sun yzsun@cs.ucla.edu November 7, 2017 Learnt Clustering Methods Vector Data Set Data Sequence Data Text
BME I5000: Biomedical Imaging
 1 Lucas Parra, CCNY BME I5000: Biomedical Imaging Lecture 4 Computed Tomography Lucas C. Parra, parra@ccny.cuny.edu some slides inspired by lecture notes of Andreas H. Hilscher at Columbia University.
1 Lucas Parra, CCNY BME I5000: Biomedical Imaging Lecture 4 Computed Tomography Lucas C. Parra, parra@ccny.cuny.edu some slides inspired by lecture notes of Andreas H. Hilscher at Columbia University.
CS249: ADVANCED DATA MINING
 CS249: ADVANCED DATA MINING Classification Evaluation and Practical Issues Instructor: Yizhou Sun yzsun@cs.ucla.edu April 24, 2017 Homework 2 out Announcements Due May 3 rd (11:59pm) Course project proposal
CS249: ADVANCED DATA MINING Classification Evaluation and Practical Issues Instructor: Yizhou Sun yzsun@cs.ucla.edu April 24, 2017 Homework 2 out Announcements Due May 3 rd (11:59pm) Course project proposal
White Pixel Artifact. Caused by a noise spike during acquisition Spike in K-space <--> sinusoid in image space
 White Pixel Artifact Caused by a noise spike during acquisition Spike in K-space sinusoid in image space Susceptibility Artifacts Off-resonance artifacts caused by adjacent regions with different
White Pixel Artifact Caused by a noise spike during acquisition Spike in K-space sinusoid in image space Susceptibility Artifacts Off-resonance artifacts caused by adjacent regions with different
Module 4. K-Space Symmetry. Review. K-Space Review. K-Space Symmetry. Partial or Fractional Echo. Half or Partial Fourier HASTE
 MRES 7005 - Fast Imaging Techniques Module 4 K-Space Symmetry Review K-Space Review K-Space Symmetry Partial or Fractional Echo Half or Partial Fourier HASTE Conditions for successful reconstruction Interpolation
MRES 7005 - Fast Imaging Techniques Module 4 K-Space Symmetry Review K-Space Review K-Space Symmetry Partial or Fractional Echo Half or Partial Fourier HASTE Conditions for successful reconstruction Interpolation
I How does the formulation (5) serve the purpose of the composite parameterization
 Supplemental Material to Identifying Alzheimer s Disease-Related Brain Regions from Multi-Modality Neuroimaging Data using Sparse Composite Linear Discrimination Analysis I How does the formulation (5)
Supplemental Material to Identifying Alzheimer s Disease-Related Brain Regions from Multi-Modality Neuroimaging Data using Sparse Composite Linear Discrimination Analysis I How does the formulation (5)
Evaluation Measures. Sebastian Pölsterl. April 28, Computer Aided Medical Procedures Technische Universität München
 Evaluation Measures Sebastian Pölsterl Computer Aided Medical Procedures Technische Universität München April 28, 2015 Outline 1 Classification 1. Confusion Matrix 2. Receiver operating characteristics
Evaluation Measures Sebastian Pölsterl Computer Aided Medical Procedures Technische Universität München April 28, 2015 Outline 1 Classification 1. Confusion Matrix 2. Receiver operating characteristics
Contents Machine Learning concepts 4 Learning Algorithm 4 Predictive Model (Model) 4 Model, Classification 4 Model, Regression 4 Representation
 Contents Machine Learning concepts 4 Learning Algorithm 4 Predictive Model (Model) 4 Model, Classification 4 Model, Regression 4 Representation Learning 4 Supervised Learning 4 Unsupervised Learning 4
Contents Machine Learning concepts 4 Learning Algorithm 4 Predictive Model (Model) 4 Model, Classification 4 Model, Regression 4 Representation Learning 4 Supervised Learning 4 Unsupervised Learning 4
Computer-Aided Diagnosis in Abdominal and Cardiac Radiology Using Neural Networks
 Computer-Aided Diagnosis in Abdominal and Cardiac Radiology Using Neural Networks Du-Yih Tsai, Masaru Sekiya and Yongbum Lee Department of Radiological Technology, School of Health Sciences, Faculty of
Computer-Aided Diagnosis in Abdominal and Cardiac Radiology Using Neural Networks Du-Yih Tsai, Masaru Sekiya and Yongbum Lee Department of Radiological Technology, School of Health Sciences, Faculty of
CIS 520, Machine Learning, Fall 2015: Assignment 7 Due: Mon, Nov 16, :59pm, PDF to Canvas [100 points]
![CIS 520, Machine Learning, Fall 2015: Assignment 7 Due: Mon, Nov 16, :59pm, PDF to Canvas [100 points] CIS 520, Machine Learning, Fall 2015: Assignment 7 Due: Mon, Nov 16, :59pm, PDF to Canvas [100 points]](/thumbs/89/100746783.jpg) CIS 520, Machine Learning, Fall 2015: Assignment 7 Due: Mon, Nov 16, 2015. 11:59pm, PDF to Canvas [100 points] Instructions. Please write up your responses to the following problems clearly and concisely.
CIS 520, Machine Learning, Fall 2015: Assignment 7 Due: Mon, Nov 16, 2015. 11:59pm, PDF to Canvas [100 points] Instructions. Please write up your responses to the following problems clearly and concisely.
Redundancy Encoding for Fast Dynamic MR Imaging using Structured Sparsity
 Redundancy Encoding for Fast Dynamic MR Imaging using Structured Sparsity Vimal Singh and Ahmed H. Tewfik Electrical and Computer Engineering Dept., The University of Texas at Austin, USA Abstract. For
Redundancy Encoding for Fast Dynamic MR Imaging using Structured Sparsity Vimal Singh and Ahmed H. Tewfik Electrical and Computer Engineering Dept., The University of Texas at Austin, USA Abstract. For
Deviceless respiratory motion correction in PET imaging exploring the potential of novel data driven strategies
 g Deviceless respiratory motion correction in PET imaging exploring the potential of novel data driven strategies Presented by Adam Kesner, Ph.D., DABR Assistant Professor, Division of Radiological Sciences,
g Deviceless respiratory motion correction in PET imaging exploring the potential of novel data driven strategies Presented by Adam Kesner, Ph.D., DABR Assistant Professor, Division of Radiological Sciences,
BME I5000: Biomedical Imaging
 BME I5000: Biomedical Imaging Lecture 1 Introduction Lucas C. Parra, parra@ccny.cuny.edu 1 Content Topics: Physics of medial imaging modalities (blue) Digital Image Processing (black) Schedule: 1. Introduction,
BME I5000: Biomedical Imaging Lecture 1 Introduction Lucas C. Parra, parra@ccny.cuny.edu 1 Content Topics: Physics of medial imaging modalities (blue) Digital Image Processing (black) Schedule: 1. Introduction,
CS145: INTRODUCTION TO DATA MINING
 CS145: INTRODUCTION TO DATA MINING 08: Classification Evaluation and Practical Issues Instructor: Yizhou Sun yzsun@cs.ucla.edu October 24, 2017 Learnt Prediction and Classification Methods Vector Data
CS145: INTRODUCTION TO DATA MINING 08: Classification Evaluation and Practical Issues Instructor: Yizhou Sun yzsun@cs.ucla.edu October 24, 2017 Learnt Prediction and Classification Methods Vector Data
Segmentation of MR Images of a Beating Heart
 Segmentation of MR Images of a Beating Heart Avinash Ravichandran Abstract Heart Arrhythmia is currently treated using invasive procedures. In order use to non invasive procedures accurate imaging modalities
Segmentation of MR Images of a Beating Heart Avinash Ravichandran Abstract Heart Arrhythmia is currently treated using invasive procedures. In order use to non invasive procedures accurate imaging modalities
FMA901F: Machine Learning Lecture 3: Linear Models for Regression. Cristian Sminchisescu
 FMA901F: Machine Learning Lecture 3: Linear Models for Regression Cristian Sminchisescu Machine Learning: Frequentist vs. Bayesian In the frequentist setting, we seek a fixed parameter (vector), with value(s)
FMA901F: Machine Learning Lecture 3: Linear Models for Regression Cristian Sminchisescu Machine Learning: Frequentist vs. Bayesian In the frequentist setting, we seek a fixed parameter (vector), with value(s)
Sparse sampling in MRI: From basic theory to clinical application. R. Marc Lebel, PhD Department of Electrical Engineering Department of Radiology
 Sparse sampling in MRI: From basic theory to clinical application R. Marc Lebel, PhD Department of Electrical Engineering Department of Radiology Objective Provide an intuitive overview of compressed sensing
Sparse sampling in MRI: From basic theory to clinical application R. Marc Lebel, PhD Department of Electrical Engineering Department of Radiology Objective Provide an intuitive overview of compressed sensing
Ischemic Stroke Lesion Segmentation Proceedings 5th October 2015 Munich, Germany
 0111010001110001101000100101010111100111011100100011011101110101101012 Ischemic Stroke Lesion Segmentation www.isles-challenge.org Proceedings 5th October 2015 Munich, Germany Preface Stroke is the second
0111010001110001101000100101010111100111011100100011011101110101101012 Ischemic Stroke Lesion Segmentation www.isles-challenge.org Proceedings 5th October 2015 Munich, Germany Preface Stroke is the second
Super-resolution Reconstruction of Fetal Brain MRI
 Super-resolution Reconstruction of Fetal Brain MRI Ali Gholipour and Simon K. Warfield Computational Radiology Laboratory Children s Hospital Boston, Harvard Medical School Worshop on Image Analysis for
Super-resolution Reconstruction of Fetal Brain MRI Ali Gholipour and Simon K. Warfield Computational Radiology Laboratory Children s Hospital Boston, Harvard Medical School Worshop on Image Analysis for
Evaluating Classifiers
 Evaluating Classifiers Charles Elkan elkan@cs.ucsd.edu January 18, 2011 In a real-world application of supervised learning, we have a training set of examples with labels, and a test set of examples with
Evaluating Classifiers Charles Elkan elkan@cs.ucsd.edu January 18, 2011 In a real-world application of supervised learning, we have a training set of examples with labels, and a test set of examples with
Applying Supervised Learning
 Applying Supervised Learning When to Consider Supervised Learning A supervised learning algorithm takes a known set of input data (the training set) and known responses to the data (output), and trains
Applying Supervised Learning When to Consider Supervised Learning A supervised learning algorithm takes a known set of input data (the training set) and known responses to the data (output), and trains
Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques
 Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques Sea Chen Department of Biomedical Engineering Advisors: Dr. Charles A. Bouman and Dr. Mark J. Lowe S. Chen Final Exam October
Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques Sea Chen Department of Biomedical Engineering Advisors: Dr. Charles A. Bouman and Dr. Mark J. Lowe S. Chen Final Exam October
A Model-Independent, Multi-Image Approach to MR Inhomogeneity Correction
 Tina Memo No. 2007-003 Published in Proc. MIUA 2007 A Model-Independent, Multi-Image Approach to MR Inhomogeneity Correction P. A. Bromiley and N.A. Thacker Last updated 13 / 4 / 2007 Imaging Science and
Tina Memo No. 2007-003 Published in Proc. MIUA 2007 A Model-Independent, Multi-Image Approach to MR Inhomogeneity Correction P. A. Bromiley and N.A. Thacker Last updated 13 / 4 / 2007 Imaging Science and
Matlab Code For Mean Square Error Of Two Images
 Matlab Code For Mean Square Error Of Two Images Snr = 30:-5:-10. % Calculate and display MSE between the original signal and noisy signal? If you have the Image Processing Toolbox, use immse() or psnr().
Matlab Code For Mean Square Error Of Two Images Snr = 30:-5:-10. % Calculate and display MSE between the original signal and noisy signal? If you have the Image Processing Toolbox, use immse() or psnr().
Automated Canvas Analysis for Painting Conservation. By Brendan Tobin
 Automated Canvas Analysis for Painting Conservation By Brendan Tobin 1. Motivation Distinctive variations in the spacings between threads in a painting's canvas can be used to show that two sections of
Automated Canvas Analysis for Painting Conservation By Brendan Tobin 1. Motivation Distinctive variations in the spacings between threads in a painting's canvas can be used to show that two sections of
G Practical Magnetic Resonance Imaging II Sackler Institute of Biomedical Sciences New York University School of Medicine. Compressed Sensing
 G16.4428 Practical Magnetic Resonance Imaging II Sackler Institute of Biomedical Sciences New York University School of Medicine Compressed Sensing Ricardo Otazo, PhD ricardo.otazo@nyumc.org Compressed
G16.4428 Practical Magnetic Resonance Imaging II Sackler Institute of Biomedical Sciences New York University School of Medicine Compressed Sensing Ricardo Otazo, PhD ricardo.otazo@nyumc.org Compressed
CT NOISE POWER SPECTRUM FOR FILTERED BACKPROJECTION AND ITERATIVE RECONSTRUCTION
 CT NOISE POWER SPECTRUM FOR FILTERED BACKPROJECTION AND ITERATIVE RECONSTRUCTION Frank Dong, PhD, DABR Diagnostic Physicist, Imaging Institute Cleveland Clinic Foundation and Associate Professor of Radiology
CT NOISE POWER SPECTRUM FOR FILTERED BACKPROJECTION AND ITERATIVE RECONSTRUCTION Frank Dong, PhD, DABR Diagnostic Physicist, Imaging Institute Cleveland Clinic Foundation and Associate Professor of Radiology
CHAPTER 3 TUMOR DETECTION BASED ON NEURO-FUZZY TECHNIQUE
 32 CHAPTER 3 TUMOR DETECTION BASED ON NEURO-FUZZY TECHNIQUE 3.1 INTRODUCTION In this chapter we present the real time implementation of an artificial neural network based on fuzzy segmentation process
32 CHAPTER 3 TUMOR DETECTION BASED ON NEURO-FUZZY TECHNIQUE 3.1 INTRODUCTION In this chapter we present the real time implementation of an artificial neural network based on fuzzy segmentation process
Medical Image Segmentation
 Medical Image Segmentation Xin Yang, HUST *Collaborated with UCLA Medical School and UCSB Segmentation to Contouring ROI Aorta & Kidney 3D Brain MR Image 3D Abdominal CT Image Liver & Spleen Caudate Nucleus
Medical Image Segmentation Xin Yang, HUST *Collaborated with UCLA Medical School and UCSB Segmentation to Contouring ROI Aorta & Kidney 3D Brain MR Image 3D Abdominal CT Image Liver & Spleen Caudate Nucleus
Image Registration. Prof. Dr. Lucas Ferrari de Oliveira UFPR Informatics Department
 Image Registration Prof. Dr. Lucas Ferrari de Oliveira UFPR Informatics Department Introduction Visualize objects inside the human body Advances in CS methods to diagnosis, treatment planning and medical
Image Registration Prof. Dr. Lucas Ferrari de Oliveira UFPR Informatics Department Introduction Visualize objects inside the human body Advances in CS methods to diagnosis, treatment planning and medical
Efficient Image Compression of Medical Images Using the Wavelet Transform and Fuzzy c-means Clustering on Regions of Interest.
 Efficient Image Compression of Medical Images Using the Wavelet Transform and Fuzzy c-means Clustering on Regions of Interest. D.A. Karras, S.A. Karkanis and D. E. Maroulis University of Piraeus, Dept.
Efficient Image Compression of Medical Images Using the Wavelet Transform and Fuzzy c-means Clustering on Regions of Interest. D.A. Karras, S.A. Karkanis and D. E. Maroulis University of Piraeus, Dept.
Unsupervised Learning
 Unsupervised Learning Learning without Class Labels (or correct outputs) Density Estimation Learn P(X) given training data for X Clustering Partition data into clusters Dimensionality Reduction Discover
Unsupervised Learning Learning without Class Labels (or correct outputs) Density Estimation Learn P(X) given training data for X Clustering Partition data into clusters Dimensionality Reduction Discover
8/3/2017. Contour Assessment for Quality Assurance and Data Mining. Objective. Outline. Tom Purdie, PhD, MCCPM
 Contour Assessment for Quality Assurance and Data Mining Tom Purdie, PhD, MCCPM Objective Understand the state-of-the-art in contour assessment for quality assurance including data mining-based techniques
Contour Assessment for Quality Assurance and Data Mining Tom Purdie, PhD, MCCPM Objective Understand the state-of-the-art in contour assessment for quality assurance including data mining-based techniques
Performance Evaluation of the TINA Medical Image Segmentation Algorithm on Brainweb Simulated Images
 Tina Memo No. 2008-003 Internal Memo Performance Evaluation of the TINA Medical Image Segmentation Algorithm on Brainweb Simulated Images P. A. Bromiley Last updated 20 / 12 / 2007 Imaging Science and
Tina Memo No. 2008-003 Internal Memo Performance Evaluation of the TINA Medical Image Segmentation Algorithm on Brainweb Simulated Images P. A. Bromiley Last updated 20 / 12 / 2007 Imaging Science and
SCIENCE & TECHNOLOGY
 Pertanika J. Sci. & Technol. 26 (1): 309-316 (2018) SCIENCE & TECHNOLOGY Journal homepage: http://www.pertanika.upm.edu.my/ Application of Active Contours Driven by Local Gaussian Distribution Fitting
Pertanika J. Sci. & Technol. 26 (1): 309-316 (2018) SCIENCE & TECHNOLOGY Journal homepage: http://www.pertanika.upm.edu.my/ Application of Active Contours Driven by Local Gaussian Distribution Fitting
Blind Sparse Motion MRI with Linear Subpixel Interpolation
 Blind Sparse Motion MRI with Linear Subpixel Interpolation Anita Möller, Marco Maaß, Alfred Mertins Institute for Signal Processing, University of Lübeck moeller@isip.uni-luebeck.de Abstract. Vital and
Blind Sparse Motion MRI with Linear Subpixel Interpolation Anita Möller, Marco Maaß, Alfred Mertins Institute for Signal Processing, University of Lübeck moeller@isip.uni-luebeck.de Abstract. Vital and
INF 4300 Classification III Anne Solberg The agenda today:
 INF 4300 Classification III Anne Solberg 28.10.15 The agenda today: More on estimating classifier accuracy Curse of dimensionality and simple feature selection knn-classification K-means clustering 28.10.15
INF 4300 Classification III Anne Solberg 28.10.15 The agenda today: More on estimating classifier accuracy Curse of dimensionality and simple feature selection knn-classification K-means clustering 28.10.15
ADVANCED IMAGE PROCESSING METHODS FOR ULTRASONIC NDE RESEARCH C. H. Chen, University of Massachusetts Dartmouth, N.
 ADVANCED IMAGE PROCESSING METHODS FOR ULTRASONIC NDE RESEARCH C. H. Chen, University of Massachusetts Dartmouth, N. Dartmouth, MA USA Abstract: The significant progress in ultrasonic NDE systems has now
ADVANCED IMAGE PROCESSING METHODS FOR ULTRASONIC NDE RESEARCH C. H. Chen, University of Massachusetts Dartmouth, N. Dartmouth, MA USA Abstract: The significant progress in ultrasonic NDE systems has now
COSC160: Detection and Classification. Jeremy Bolton, PhD Assistant Teaching Professor
 COSC160: Detection and Classification Jeremy Bolton, PhD Assistant Teaching Professor Outline I. Problem I. Strategies II. Features for training III. Using spatial information? IV. Reducing dimensionality
COSC160: Detection and Classification Jeremy Bolton, PhD Assistant Teaching Professor Outline I. Problem I. Strategies II. Features for training III. Using spatial information? IV. Reducing dimensionality
Shape-based Diffeomorphic Registration on Hippocampal Surfaces Using Beltrami Holomorphic Flow
 Shape-based Diffeomorphic Registration on Hippocampal Surfaces Using Beltrami Holomorphic Flow Abstract. Finding meaningful 1-1 correspondences between hippocampal (HP) surfaces is an important but difficult
Shape-based Diffeomorphic Registration on Hippocampal Surfaces Using Beltrami Holomorphic Flow Abstract. Finding meaningful 1-1 correspondences between hippocampal (HP) surfaces is an important but difficult
The exam is closed book, closed notes except your one-page (two-sided) cheat sheet.
 CS 189 Spring 2015 Introduction to Machine Learning Final You have 2 hours 50 minutes for the exam. The exam is closed book, closed notes except your one-page (two-sided) cheat sheet. No calculators or
CS 189 Spring 2015 Introduction to Machine Learning Final You have 2 hours 50 minutes for the exam. The exam is closed book, closed notes except your one-page (two-sided) cheat sheet. No calculators or
3D Surface Reconstruction of the Brain based on Level Set Method
 3D Surface Reconstruction of the Brain based on Level Set Method Shijun Tang, Bill P. Buckles, and Kamesh Namuduri Department of Computer Science & Engineering Department of Electrical Engineering University
3D Surface Reconstruction of the Brain based on Level Set Method Shijun Tang, Bill P. Buckles, and Kamesh Namuduri Department of Computer Science & Engineering Department of Electrical Engineering University
Joint Reconstruction of Multi-contrast MR Images for Multiple Sclerosis Lesion Segmentation
 Joint Reconstruction of Multi-contrast MR Images for Multiple Sclerosis Lesion Segmentation Pedro A Gómez 1,2,3, Jonathan I Sperl 3, Tim Sprenger 2,3, Claudia Metzler-Baddeley 4, Derek K Jones 4, Philipp
Joint Reconstruction of Multi-contrast MR Images for Multiple Sclerosis Lesion Segmentation Pedro A Gómez 1,2,3, Jonathan I Sperl 3, Tim Sprenger 2,3, Claudia Metzler-Baddeley 4, Derek K Jones 4, Philipp
Digital Image Processing COSC 6380/4393
 Digital Image Processing COSC 6380/4393 Lecture 21 Nov 16 th, 2017 Pranav Mantini Ack: Shah. M Image Processing Geometric Transformation Point Operations Filtering (spatial, Frequency) Input Restoration/
Digital Image Processing COSC 6380/4393 Lecture 21 Nov 16 th, 2017 Pranav Mantini Ack: Shah. M Image Processing Geometric Transformation Point Operations Filtering (spatial, Frequency) Input Restoration/
COMPREHENSIVE QUALITY CONTROL OF NMR TOMOGRAPHY USING 3D PRINTED PHANTOM
 COMPREHENSIVE QUALITY CONTROL OF NMR TOMOGRAPHY USING 3D PRINTED PHANTOM Mažena MACIUSOVIČ *, Marius BURKANAS *, Jonas VENIUS *, ** * Medical Physics Department, National Cancer Institute, Vilnius, Lithuania
COMPREHENSIVE QUALITY CONTROL OF NMR TOMOGRAPHY USING 3D PRINTED PHANTOM Mažena MACIUSOVIČ *, Marius BURKANAS *, Jonas VENIUS *, ** * Medical Physics Department, National Cancer Institute, Vilnius, Lithuania
Lab 9. Julia Janicki. Introduction
 Lab 9 Julia Janicki Introduction My goal for this project is to map a general land cover in the area of Alexandria in Egypt using supervised classification, specifically the Maximum Likelihood and Support
Lab 9 Julia Janicki Introduction My goal for this project is to map a general land cover in the area of Alexandria in Egypt using supervised classification, specifically the Maximum Likelihood and Support
Machine Learning / Jan 27, 2010
 Revisiting Logistic Regression & Naïve Bayes Aarti Singh Machine Learning 10-701/15-781 Jan 27, 2010 Generative and Discriminative Classifiers Training classifiers involves learning a mapping f: X -> Y,
Revisiting Logistic Regression & Naïve Bayes Aarti Singh Machine Learning 10-701/15-781 Jan 27, 2010 Generative and Discriminative Classifiers Training classifiers involves learning a mapping f: X -> Y,
MEDICAL IMAGE COMPUTING (CAP 5937) LECTURE 4: Pre-Processing Medical Images (II)
 SPRING 2016 1 MEDICAL IMAGE COMPUTING (CAP 5937) LECTURE 4: Pre-Processing Medical Images (II) Dr. Ulas Bagci HEC 221, Center for Research in Computer Vision (CRCV), University of Central Florida (UCF),
SPRING 2016 1 MEDICAL IMAGE COMPUTING (CAP 5937) LECTURE 4: Pre-Processing Medical Images (II) Dr. Ulas Bagci HEC 221, Center for Research in Computer Vision (CRCV), University of Central Florida (UCF),
Unsupervised learning in Vision
 Chapter 7 Unsupervised learning in Vision The fields of Computer Vision and Machine Learning complement each other in a very natural way: the aim of the former is to extract useful information from visual
Chapter 7 Unsupervised learning in Vision The fields of Computer Vision and Machine Learning complement each other in a very natural way: the aim of the former is to extract useful information from visual
All lecture slides will be available at CSC2515_Winter15.html
 CSC2515 Fall 2015 Introduc3on to Machine Learning Lecture 9: Support Vector Machines All lecture slides will be available at http://www.cs.toronto.edu/~urtasun/courses/csc2515/ CSC2515_Winter15.html Many
CSC2515 Fall 2015 Introduc3on to Machine Learning Lecture 9: Support Vector Machines All lecture slides will be available at http://www.cs.toronto.edu/~urtasun/courses/csc2515/ CSC2515_Winter15.html Many
Presented at the FIG Congress 2018, May 6-11, 2018 in Istanbul, Turkey
 Presented at the FIG Congress 2018, May 6-11, 2018 in Istanbul, Turkey Evangelos MALTEZOS, Charalabos IOANNIDIS, Anastasios DOULAMIS and Nikolaos DOULAMIS Laboratory of Photogrammetry, School of Rural
Presented at the FIG Congress 2018, May 6-11, 2018 in Istanbul, Turkey Evangelos MALTEZOS, Charalabos IOANNIDIS, Anastasios DOULAMIS and Nikolaos DOULAMIS Laboratory of Photogrammetry, School of Rural
CHAPTER 6 DETECTION OF MASS USING NOVEL SEGMENTATION, GLCM AND NEURAL NETWORKS
 130 CHAPTER 6 DETECTION OF MASS USING NOVEL SEGMENTATION, GLCM AND NEURAL NETWORKS A mass is defined as a space-occupying lesion seen in more than one projection and it is described by its shapes and margin
130 CHAPTER 6 DETECTION OF MASS USING NOVEL SEGMENTATION, GLCM AND NEURAL NETWORKS A mass is defined as a space-occupying lesion seen in more than one projection and it is described by its shapes and margin
Nonrigid Motion Compensation of Free Breathing Acquired Myocardial Perfusion Data
 Nonrigid Motion Compensation of Free Breathing Acquired Myocardial Perfusion Data Gert Wollny 1, Peter Kellman 2, Andrés Santos 1,3, María-Jesus Ledesma 1,3 1 Biomedical Imaging Technologies, Department
Nonrigid Motion Compensation of Free Breathing Acquired Myocardial Perfusion Data Gert Wollny 1, Peter Kellman 2, Andrés Santos 1,3, María-Jesus Ledesma 1,3 1 Biomedical Imaging Technologies, Department
Medical images, segmentation and analysis
 Medical images, segmentation and analysis ImageLab group http://imagelab.ing.unimo.it Università degli Studi di Modena e Reggio Emilia Medical Images Macroscopic Dermoscopic ELM enhance the features of
Medical images, segmentation and analysis ImageLab group http://imagelab.ing.unimo.it Università degli Studi di Modena e Reggio Emilia Medical Images Macroscopic Dermoscopic ELM enhance the features of
Recognition of Changes in SAR Images Based on Gauss-Log Ratio and MRFFCM
 Recognition of Changes in SAR Images Based on Gauss-Log Ratio and MRFFCM Jismy Alphonse M.Tech Scholar Computer Science and Engineering Department College of Engineering Munnar, Kerala, India Biju V. G.
Recognition of Changes in SAR Images Based on Gauss-Log Ratio and MRFFCM Jismy Alphonse M.Tech Scholar Computer Science and Engineering Department College of Engineering Munnar, Kerala, India Biju V. G.
Slide 1. Technical Aspects of Quality Control in Magnetic Resonance Imaging. Slide 2. Annual Compliance Testing. of MRI Systems.
 Slide 1 Technical Aspects of Quality Control in Magnetic Resonance Imaging Slide 2 Compliance Testing of MRI Systems, Ph.D. Department of Radiology Henry Ford Hospital, Detroit, MI Slide 3 Compliance Testing
Slide 1 Technical Aspects of Quality Control in Magnetic Resonance Imaging Slide 2 Compliance Testing of MRI Systems, Ph.D. Department of Radiology Henry Ford Hospital, Detroit, MI Slide 3 Compliance Testing
Norbert Schuff VA Medical Center and UCSF
 Norbert Schuff Medical Center and UCSF Norbert.schuff@ucsf.edu Medical Imaging Informatics N.Schuff Course # 170.03 Slide 1/67 Objective Learn the principle segmentation techniques Understand the role
Norbert Schuff Medical Center and UCSF Norbert.schuff@ucsf.edu Medical Imaging Informatics N.Schuff Course # 170.03 Slide 1/67 Objective Learn the principle segmentation techniques Understand the role
Learning to Recognize Faces in Realistic Conditions
 000 001 002 003 004 005 006 007 008 009 010 011 012 013 014 015 016 017 018 019 020 021 022 023 024 025 026 027 028 029 030 031 032 033 034 035 036 037 038 039 040 041 042 043 044 045 046 047 048 049 050
000 001 002 003 004 005 006 007 008 009 010 011 012 013 014 015 016 017 018 019 020 021 022 023 024 025 026 027 028 029 030 031 032 033 034 035 036 037 038 039 040 041 042 043 044 045 046 047 048 049 050
Role of Parallel Imaging in High Field Functional MRI
 Role of Parallel Imaging in High Field Functional MRI Douglas C. Noll & Bradley P. Sutton Department of Biomedical Engineering, University of Michigan Supported by NIH Grant DA15410 & The Whitaker Foundation
Role of Parallel Imaging in High Field Functional MRI Douglas C. Noll & Bradley P. Sutton Department of Biomedical Engineering, University of Michigan Supported by NIH Grant DA15410 & The Whitaker Foundation
MR IMAGE SEGMENTATION
 MR IMAGE SEGMENTATION Prepared by : Monil Shah What is Segmentation? Partitioning a region or regions of interest in images such that each region corresponds to one or more anatomic structures Classification
MR IMAGE SEGMENTATION Prepared by : Monil Shah What is Segmentation? Partitioning a region or regions of interest in images such that each region corresponds to one or more anatomic structures Classification
Analyzing Vocal Patterns to Determine Emotion Maisy Wieman, Andy Sun
 Analyzing Vocal Patterns to Determine Emotion Maisy Wieman, Andy Sun 1. Introduction The human voice is very versatile and carries a multitude of emotions. Emotion in speech carries extra insight about
Analyzing Vocal Patterns to Determine Emotion Maisy Wieman, Andy Sun 1. Introduction The human voice is very versatile and carries a multitude of emotions. Emotion in speech carries extra insight about
Classification of Abdominal Tissues by k-means Clustering for 3D Acoustic and Shear-Wave Modeling
 1 Classification of Abdominal Tissues by k-means Clustering for 3D Acoustic and Shear-Wave Modeling Kevin T. Looby klooby@stanford.edu I. ABSTRACT Clutter is an effect that degrades the quality of medical
1 Classification of Abdominal Tissues by k-means Clustering for 3D Acoustic and Shear-Wave Modeling Kevin T. Looby klooby@stanford.edu I. ABSTRACT Clutter is an effect that degrades the quality of medical
Bagging for One-Class Learning
 Bagging for One-Class Learning David Kamm December 13, 2008 1 Introduction Consider the following outlier detection problem: suppose you are given an unlabeled data set and make the assumptions that one
Bagging for One-Class Learning David Kamm December 13, 2008 1 Introduction Consider the following outlier detection problem: suppose you are given an unlabeled data set and make the assumptions that one
5 Learning hypothesis classes (16 points)
 5 Learning hypothesis classes (16 points) Consider a classification problem with two real valued inputs. For each of the following algorithms, specify all of the separators below that it could have generated
5 Learning hypothesis classes (16 points) Consider a classification problem with two real valued inputs. For each of the following algorithms, specify all of the separators below that it could have generated
More on Learning. Neural Nets Support Vectors Machines Unsupervised Learning (Clustering) K-Means Expectation-Maximization
 More on Learning Neural Nets Support Vectors Machines Unsupervised Learning (Clustering) K-Means Expectation-Maximization Neural Net Learning Motivated by studies of the brain. A network of artificial
More on Learning Neural Nets Support Vectors Machines Unsupervised Learning (Clustering) K-Means Expectation-Maximization Neural Net Learning Motivated by studies of the brain. A network of artificial
Cognitive States Detection in fmri Data Analysis using incremental PCA
 Department of Computer Engineering Cognitive States Detection in fmri Data Analysis using incremental PCA Hoang Trong Minh Tuan, Yonggwan Won*, Hyung-Jeong Yang International Conference on Computational
Department of Computer Engineering Cognitive States Detection in fmri Data Analysis using incremental PCA Hoang Trong Minh Tuan, Yonggwan Won*, Hyung-Jeong Yang International Conference on Computational
Supervised Learning with Neural Networks. We now look at how an agent might learn to solve a general problem by seeing examples.
 Supervised Learning with Neural Networks We now look at how an agent might learn to solve a general problem by seeing examples. Aims: to present an outline of supervised learning as part of AI; to introduce
Supervised Learning with Neural Networks We now look at how an agent might learn to solve a general problem by seeing examples. Aims: to present an outline of supervised learning as part of AI; to introduce
CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS
 CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS CHAPTER 4 CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS 4.1 Introduction Optical character recognition is one of
CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS CHAPTER 4 CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS 4.1 Introduction Optical character recognition is one of
Evaluating Classifiers
 Evaluating Classifiers Reading for this topic: T. Fawcett, An introduction to ROC analysis, Sections 1-4, 7 (linked from class website) Evaluating Classifiers What we want: Classifier that best predicts
Evaluating Classifiers Reading for this topic: T. Fawcett, An introduction to ROC analysis, Sections 1-4, 7 (linked from class website) Evaluating Classifiers What we want: Classifier that best predicts
Storage Efficient NL-Means Burst Denoising for Programmable Cameras
 Storage Efficient NL-Means Burst Denoising for Programmable Cameras Brendan Duncan Stanford University brendand@stanford.edu Miroslav Kukla Stanford University mkukla@stanford.edu Abstract An effective
Storage Efficient NL-Means Burst Denoising for Programmable Cameras Brendan Duncan Stanford University brendand@stanford.edu Miroslav Kukla Stanford University mkukla@stanford.edu Abstract An effective
IMAGE DE-NOISING IN WAVELET DOMAIN
 IMAGE DE-NOISING IN WAVELET DOMAIN Aaditya Verma a, Shrey Agarwal a a Department of Civil Engineering, Indian Institute of Technology, Kanpur, India - (aaditya, ashrey)@iitk.ac.in KEY WORDS: Wavelets,
IMAGE DE-NOISING IN WAVELET DOMAIN Aaditya Verma a, Shrey Agarwal a a Department of Civil Engineering, Indian Institute of Technology, Kanpur, India - (aaditya, ashrey)@iitk.ac.in KEY WORDS: Wavelets,
Weka ( )
 Weka ( http://www.cs.waikato.ac.nz/ml/weka/ ) The phases in which classifier s design can be divided are reflected in WEKA s Explorer structure: Data pre-processing (filtering) and representation Supervised
Weka ( http://www.cs.waikato.ac.nz/ml/weka/ ) The phases in which classifier s design can be divided are reflected in WEKA s Explorer structure: Data pre-processing (filtering) and representation Supervised
Statistical Analysis of Metabolomics Data. Xiuxia Du Department of Bioinformatics & Genomics University of North Carolina at Charlotte
 Statistical Analysis of Metabolomics Data Xiuxia Du Department of Bioinformatics & Genomics University of North Carolina at Charlotte Outline Introduction Data pre-treatment 1. Normalization 2. Centering,
Statistical Analysis of Metabolomics Data Xiuxia Du Department of Bioinformatics & Genomics University of North Carolina at Charlotte Outline Introduction Data pre-treatment 1. Normalization 2. Centering,
Evaluation Metrics. (Classifiers) CS229 Section Anand Avati
 Evaluation Metrics (Classifiers) CS Section Anand Avati Topics Why? Binary classifiers Metrics Rank view Thresholding Confusion Matrix Point metrics: Accuracy, Precision, Recall / Sensitivity, Specificity,
Evaluation Metrics (Classifiers) CS Section Anand Avati Topics Why? Binary classifiers Metrics Rank view Thresholding Confusion Matrix Point metrics: Accuracy, Precision, Recall / Sensitivity, Specificity,
Edge-Preserving Denoising for Segmentation in CT-Images
 Edge-Preserving Denoising for Segmentation in CT-Images Eva Eibenberger, Anja Borsdorf, Andreas Wimmer, Joachim Hornegger Lehrstuhl für Mustererkennung, Friedrich-Alexander-Universität Erlangen-Nürnberg
Edge-Preserving Denoising for Segmentation in CT-Images Eva Eibenberger, Anja Borsdorf, Andreas Wimmer, Joachim Hornegger Lehrstuhl für Mustererkennung, Friedrich-Alexander-Universität Erlangen-Nürnberg
Network Traffic Measurements and Analysis
 DEIB - Politecnico di Milano Fall, 2017 Sources Hastie, Tibshirani, Friedman: The Elements of Statistical Learning James, Witten, Hastie, Tibshirani: An Introduction to Statistical Learning Andrew Ng:
DEIB - Politecnico di Milano Fall, 2017 Sources Hastie, Tibshirani, Friedman: The Elements of Statistical Learning James, Witten, Hastie, Tibshirani: An Introduction to Statistical Learning Andrew Ng:
Facial Expression Classification with Random Filters Feature Extraction
 Facial Expression Classification with Random Filters Feature Extraction Mengye Ren Facial Monkey mren@cs.toronto.edu Zhi Hao Luo It s Me lzh@cs.toronto.edu I. ABSTRACT In our work, we attempted to tackle
Facial Expression Classification with Random Filters Feature Extraction Mengye Ren Facial Monkey mren@cs.toronto.edu Zhi Hao Luo It s Me lzh@cs.toronto.edu I. ABSTRACT In our work, we attempted to tackle
