Computational Genomics and Molecular Biology, Fall
|
|
|
- Denis Harvey
- 5 years ago
- Views:
Transcription
1 Computational Genomics and Molecular Biology, Fall Sequence Alignment Dannie Durand Pairwise Sequence Alignment The goal of pairwise sequence alignment is to establish a correspondence between the elements in a pair of sequences that share a common property, such as common ancestry or a common structural or functional role. In computational biology, the sequences under consideration are typically nucleic acid or amino acid polymers. We will consider three variants of the pairwise sequence alignment problem: global alignment, semi-global alignment, and local alignment. Global alignment is used to compare sequences in cases where we have reason to believe that the sequences are related along their entire length. If, for example, sequences s and t are two independent sequencing runs of the same PCR product, then they should differ only in positions where there are sequencing errors. In order to find those sequencing errors, we align all of sequence s with all of sequence t. Other applications of global alignment include finding mutations in closely related gene or protein sequences and identification of single nucleotide polymorphisms (SNPs). Semi-global alignment is a variant of global alignment that allows for gaps at the beginning and/or the end of one of the sequences. Semi-global alignment should be used in cases where we believe that s and t are related along the entire length of the region where they overlap. For example, if s is a prokaryotic gene sequence and t is the spliced mrna transcript produced when s is expressed, every base in t should correspond to some base in s. Semi-global alignment jumps over the 5 untranslated region in s without exacting a penalty, but forces an alignment along the entire length of t. In contrast, local alignment addresses cases where we only expect to find isolated regions of similarity. One example is alignment of genomic DNA upstream from two co-expressed genes to find conserved regions that may correspond to transcription factor binding sites. Another application is identification of conserved domains 1 in two amino acid sequences that encode proteins that share one or more domains, but are otherwise unrelated. Prior to introducing algorithms for these pairwise alignment problems, we introduce some notation. Alphabet: An alphabet, denoted by Σ, is a finite, unordered set of symbols; e.g., DNA: Σ D = {A, C, G, T } 1 A domain is a peptide sequence that encodes a protein module that will fold into its characteristic shape independent of the surrounding amino acid context and that is found in many different proteins.
2 Computational Genomics and Molecular Biology, Fall RNA: Σ R = {A, C, G, U} Amino acids: Σ AA = {A, C, D, E, F, G, H, I, K, L, M, N, P, Q, R, S, T, V, W, Y } Sequences or Strings: A sequence or string, s, is a finite succession of the symbols in Σ. We say that s Σ, where Σ is the set of all sequences over alphabet Σ, including the empty sequence,. For example, Σ R = {, A, C, G, U, AA, AC, AG, AU, CA, CC, CG, CU,...}. Given a sequence s of length m, we use s[1]s[2] s[m] to denote the symbols in s. Sometimes, we will use s 1 s 2 s m for convenience. Subsequences: A subsequence of s is any sequence obtained by removing zero or more symbols from s. The sequences CATA and CTG are subsequences of CATTAG. AATTCG is not. A proper subsequence is a subsequence obtained by removing one or more symbols from s. Substrings: A substring of s is a subsequence of s consisting of consecutive symbols in s. Given a sequence, s, of length m, the substring that begins with s[i] and ends with s[j] is denoted s[i... j], 1 i j m. The sequence CAT is a substring of CATTAG. CATA is not. A prefix of s is denoted s[1... j], j m. A suffix of s is denoted s[i... m], 1 i. Global pairwise alignment Now that we have some notation to work with, we will introduce a formal definition of a global alignment. Next we will consider how to assign a numerical score to an alignment. The score represents an assessment of the quality of the alignment. Finally, we will introduce an efficient algorithm to find the alignment that is optimal with respect to a scoring function. Let Σ = Σ { } be the alphabet expanded to include a character to represent gaps. Given sequence s Σ of length m and sequence t Σ of length n, α = {s, t } is an alignment of s and t if and only if s, t (Σ ), s = t = l, where max(m, n) l m + n, s is the subsequence obtained by removing from s and t is the subsequence obtained by removing from t,
3 Computational Genomics and Molecular Biology, Fall there is no value of i for which s [i] = t [i] =. Our goal is to find the best global alignment; that is, we seek an alignment, α (s, t), that optimizes an objective criterion. Scoring an alignment Given sequences s and t and an alignment α(s, t) = {s, t }, it is convenient to assign a score to alignment α that quantifies how well α captures the relationship between s and t. This may be a minimization or a maximization criterion. Distance scoring: An alignment can be scored using a distance-based metric. This is a minimization criterion: a lower distance indicates a better alignment. We define the distance score of an alignment α k (s, t) = {s, t } to be D ( α k (s, t) ) = D(s, t ) l = d(s [i], t (1) [i]), where d(x, y) is the distance between a pair of symbols x and y and l is the length of the alignment. The optimal alignment is the alignment that minimizes the distance between s and t: i=1 α (s, t) = argmin D ( α k (s, t) ). α k (s,t) If d(x, y) = 1 and d(x, ) = 1, x, y, then D ( α (s, t) ) corresponds to the minimum number of operations required to transform s into t, where the operations are substitution, insertion, and deletion. This is called the edit distance. If d(x, y) > 1 or d(x, ) > 1 or both, then D[α (s, t)] is called the weighted edit distance. The function specifying the distance between pairs of symbols must obey the following properties, for all x, y, and z: 1. d(x, x) = 0 2. d(x, y) > 0 3. d(x, ) > 0 4. d(x, y) = d(y, x) 5. d(x, z) d(x, y) + d(y, z) (triangle inequality) Several properties of the distance scoring function are worth noting. First, D(s, t ) is a metric; that is, it satisfies the triangle inequality.second, the symmetric property, d(x, y) = d(y, x), implies that there is no directionality in the scoring system. This is because, given a column with symbols x and y in a pairwise alignment, we have no way of knowing whether the ancestor was an x, followed
4 Computational Genomics and Molecular Biology, Fall by an x y substitution, or vice versa. It is also possible that the ancestor was neither x nor y and a series of substitutions gave rise to x in one sequence and to y in the other. Similarly, when a symbol, x, in sequence s is aligned with a gap in sequence t (or vice versa), there is no way to know whether x was inserted in s or deleted from t. For this reason, gaps are also called indels. Similarity scoring: Alignments can also be scored with similarity measures. These are maximization criteria: a higher score indicates a better alignment. The similarity score of α k (s, t) = {s, t } is S(α k (s, t)) = l p(s [i], t [i]), (2) i=1 where p(x, y) is a score that reflects the similarity of x and y and p(x, ) is the gap score. The optimal alignment is the alignment that maximizes the similarity between s and t: α (s, t) = argmax S ( α k (s, t) ). α k (s,t) In general, amino acid alignments are scored with substitution matrices that assign a different similarity score to each pair of amino acid residues. Typically, pairs of amino acids with similar properties have higher scores than pairs with divergent properties. Examples of substitution matrices used to score alignments include the PAM and BLOSUM matrices. We will discuss how such substitution matrices are derived later in the semester. For now, we will consider a simple similarity scoring function that treats all symbols in Σ equally. This simple scoring function has just three values; a score for matching symbols (M), a score for a mismatch (m), and a gap score (g): p(x, x) = M, p(x, y) = m, p(x, ) = g. (3) In order to obtain alignments that make sense, we must impose several constraints on the values of M, m, and g. First, we require that M > m, since matches are preferred over mismatches. We further require that a substitution be preferred over two gaps (i.e., m > 2g). A scoring function with m < 2g would exclude the possibility of an alignment with substitutions, because the score of any substitution could be improved by replacing it with two gaps. Note that for both distance and similarity scoring, the score of an alignment is defined to be the sum of the scores for the individual positions in the alignment (Equations 1 and 2), which implies that each position in the alignment is independent of neighboring positions. This assumption is unrealistic: In real biomolecular sequences, there can be interactions between neighboring, or even distant, residues in the sequence. However, scoring functions that assume positional independence are widely used because they greatly simplify the calculation of alignment scores.
5 Computational Genomics and Molecular Biology, Fall A dynamic programming algorithm to align a pair of sequences We now have a formal definition of an alignment and a way of assigning a numerical score to any given alignment. How do we find the alignment with the optimal score? We could generate all possible alignments, score each one, and choose the alignment with the best score. However, the computational cost would be prohibitive, since the size of the space of all possible alignments of s and t is O(2 m+n ). (Convince yourself this is the case.) Instead, dynamic programming can be used to find the optimal alignment efficiently. The dynamic programs for sequence alignment compute a matrix a, where a[i, j] is the score of the optimal alignment of the prefixes s[1..i] and t[1..j], that is, the prefixes of s and t that end at positions i and j, respectively. Dynamic programming algorithms for sequence alignment have four components: Initialization of the first row and column of a[i, j]. A recurrence relation for a[i, j], i > 1, j > 1. Determination of the score of the optimal alignment from the matrix a[i, j]. Trace back through the matrix to obtain the optimal alignment. The details of each of these steps are what differentiate global, semi-global, and local alignment. In all cases, the algorithm proceeds by iteratively calculating the values of a[i, j] from the values of a[i 1, j], a[i, j 1], and a[i 1, j 1]. At each step, the indices of the entry in a that optimize the right hand side of the recurrence (Equations 4 and 5) are stored in a matrix, T. These pointers are used to reconstruct the alignment that gave the optimal score. The algorithm continues until all entries in the matrix a have been assigned values. The details of dynamic programming for global alignment are given below for both distance and similarity scoring functions. With distance scoring, all entries in a are non-negative, since d(x, y) 0, x, y. With a similarity scoring function, the entries in a may be positive or negative. This algorithm computes the scores of all pairs of prefixes in O(m n) time. The trace back through the alignment matrix to obtain the optimal alignment requires O(m + n) time. Note that there may be more than one optimal alignment.
6 Computational Genomics and Molecular Biology, Fall Global alignment with distance scoring Input: Sequences s and t of lengths m and n, respectively. Initialization: a[0, j] = a[0, j 1] + d(, t[j]) (top row) a[i, 0] = a[i 1, 0] + d(s[i], ) (left column) Recurrence: a[i, j] = min a[i 1, j] + d(s[i], ) a[i 1, j 1] + d(s[i], t[j]) a[i, j 1] + d(, t[j]) (4) Store the indices of the entry in a that minimize the right hand side of Equation 4 in a traceback matrix, T. Trace back: Output: Follow the pointers from T [m, n] to T [0, 0] to obtain the optimal alignment. The optimal alignment score, a[m, n]. The optimal global alignment of s and t with respect to distance function, D. Global alignment with similarity scoring The dynamic programming algorithm for global alignment with similarity scoring has the same general structure as the global alignment algorithm for distance scoring. However, the details of the initialization and recurrence differ. Initialization: a[0, i] = a[i 1, 0] + g a[j, 0] = a[0, j 1] + g Recurrence relation: a[i, j] = max a[i, j 1] + g a[i 1, j 1] + p(i, j) a[i 1, j] + g (5) Store the indices of the entry in a that minimize the right hand side of Equation 5 in a traceback matrix, T.
7 Computational Genomics and Molecular Biology, Fall Traceback: From T [m, n] to T [0, 0] to obtain the optimal alignment. Output: The optimal alignment score, a[m, n]. The optimal global alignment of s and t with respect to distance function, D. Semi-global alignment Global alignment seeks the best, full length alignment; that is, the best way to match up two sequences along their entire length. For some applications, it is desirable to relax this requirement and not penalize gaps at the beginning and/or end of an alignment. For example, for sequence assembly, we seek sequence fragments that overlap, that is we expect to be able to align the end of one fragment with the beginning of another. Very occasionally, we may find sequence fragments that start and end at the same position, but, in general, we expect some gaps at the beginning and end of the alignment. Another example is aligning cdna s with genomic DNA to identify gene structure. The cdna will be completely covered by the genomic DNA, so we expect gaps at both the beginning and end of the cdna fragment. Semi-global alignment is a modification of global alignment that allows the user to specify that gaps will be penalty-free at the beginning of one of the sequences and/or at the end of one of the sequences. Given sequences s and t, there are eight possible cases to consider: Gaps are penalty-free at the beginning of s; e.g., s: D O t: R E D O Gaps are penalty-free at the beginning of t; e.g., s: R E D O t: D O Gaps are penalty-free at the end of s; e.g., s: D O t: D O N E Gaps are penalty-free at the end of t; e.g.,
8 Computational Genomics and Molecular Biology, Fall s: D O N E t: D O Gaps are penalty-free at the beginning and end of s; e.g., s: D O t: R E D O N E Gaps are penalty-free at the beginning and end of t; e.g., s: R E D O N E t: D O Gaps are penalty-free at the beginning of s and at the end of t; e.g., s: D O N E t: R E D O Gaps are penalty-free at the beginning of t and at the end of s; e.g., s: R E D O t: D O N E In semi-global alignment, we do not allow gaps at the beginning of s and the beginning of t in the same alignment. Nor do we not allow gaps at the end of s and the end of t. Why not? Like global alignment, the optimal semi-global alignment can be found using dynamic programming using either distance or similarity scoring. Below, we describe the modifications required to adapt the dynamic program for global pairwise alignment with similarity scoring to the semi-global alignment problem. Similar modifications can be made to obtain a semi-global alignment algorithm that uses distances. Initialization: To allow gaps at the beginning of s, set a[0, j] = 0, j ; i.e., the first row is zero. The first column is initialized as in global alignment. To allow gaps at the beginning of t, set a[i, 0] = 0, i ; i.e., the first column is zero. The first row is initialized as in global alignment. Recurrence relation: Same as global.
9 Computational Genomics and Molecular Biology, Fall Optimal alignment score: Trace back: To avoid trailing gap penalties at the end of t, we define the optimal score to be max i a[i, n], the optimal score in the last column. To avoid trailing gap penalties at the end of s, we define the optimal score to be max j a[m, j], the optimal score in the bottom row. To avoid trailing gap penalties at the end of t, trace back from T [i, n], where i = argmax i a[i, n]. In other words, trace back from the cell(s) in the last column with optimal score. To avoid trailing gap penalties at the end of s, trace back from T [m, j ], where j = argmax j a[m, j]. In other words, trace back from the cell(s) in the last row with optimal score. Note that when the first row (or column) of the matrix is initialized to zero, the traceback will end in the first row (or column), but not necessarily in the cell a[0, 0]. Like global alignment, either distance or similarity scoring can be used for semiglobal alignment. There may be more than one optimal semiglobal alignment. Local pairwise alignment Global and semiglobal alignment are used in cases where we expect that s and t are related from end to end. Semi-global allows for some gaps at the beginning or end of one sequence, but the underlying assumption is the same: s and t share a relationship within the entire aligned region. In contrast, local alignment is used in cases where s and t share one or more local regions that are related, but are not related from end to end. The alignment of any substring s h,i = s[h... i] of s and any substring t j,k = t[j... k] of t is a local alignment of s and t. The optimal alignment of s h,i and t j,k is α (s h,i, t j,k ), the highest scoring global alignment of those substrings. The optimal local alignment of s and t is the highest scoring optimal alignment of all possible substrings of s and t, that is, α (s, t) = argmax S(α (s h,i, t j,k )), h,i,j,k where 1 h i m, and 1 j k n and S is the similarity scoring function defined in Equation 2. Note that there may be more than one optimal local alignment. High scoring sub-optimal alignments may also be of interest.
10 Computational Genomics and Molecular Biology, Fall For local alignment, the pairwise alignment dynamic programming algorithm must be modified to allow the alignment to start and stop anywhere in s and t. Unlike the dynamic programs for global alignment, the local alignment recurrence (Equation 6) has a fourth term that sets the score a[i, j] to zero whenever adding a substitution or a gap to the alignment results in a negative score. This is what allows the local alignment algorithm to consider all possible starting positions in s and in t. Initialization: Set the first row and column to zero: a[i, 0] = 0 and a[0, j] = 0 for all i and j. Recurrence: a[i, j] = max a[i, j 1] + g a[i 1, j 1] + p(s[i], t[j]) a[i, j 1] + g 0 (6) Optimal alignment score: Trace back: The score of the optimal alignment is max i,j a[i, j], where the maximum is taken over all i and all j. Trace back starting at T [i, j ], where (i, j ) = argmax i,j a[i, j] is the cell corresponding to the maximum score. End the trace back at the first cell with value zero. The point at which a[i, j] drops below zero depends on the scoring function and critically determines what the resulting alignment will look like. For this reason, scoring functions for local alignment are subject to more stringent constraints than scoring functions for global and semi-global alignment. In order to find biologically meaningful conserved regions, a scoring function for local pairwise alignment must satisfy the following requirements: M > m > 2g. The scoring function must be a similarity function. The local alignment that minimizes the edit distance (weighted or unweighted) is the empty alignment, which tells us nothing. There must be at least one pair of residues, x and y, for which the similarity score p(x,y) is positive. Otherwise, the optimal alignment is always the empty alignment.
11 Computational Genomics and Molecular Biology, Fall The expected alignment score of a pair of randomly generated sequences (i.e., sequences sampled from a background distribution) must be negative. These rules apply to general scoring functions, where the score for each pair of residues can have a different numerical value. For our simple similarity function, we require a positive score for matches (M > 0) and a negative score for mismatches (m < 0) and gaps (g < 0).
Computational Molecular Biology
 Computational Molecular Biology Erwin M. Bakker Lecture 2 Materials used from R. Shamir [2] and H.J. Hoogeboom [4]. 1 Molecular Biology Sequences DNA A, T, C, G RNA A, U, C, G Protein A, R, D, N, C E,
Computational Molecular Biology Erwin M. Bakker Lecture 2 Materials used from R. Shamir [2] and H.J. Hoogeboom [4]. 1 Molecular Biology Sequences DNA A, T, C, G RNA A, U, C, G Protein A, R, D, N, C E,
Computational Molecular Biology
 Computational Molecular Biology Erwin M. Bakker Lecture 3, mainly from material by R. Shamir [2] and H.J. Hoogeboom [4]. 1 Pairwise Sequence Alignment Biological Motivation Algorithmic Aspect Recursive
Computational Molecular Biology Erwin M. Bakker Lecture 3, mainly from material by R. Shamir [2] and H.J. Hoogeboom [4]. 1 Pairwise Sequence Alignment Biological Motivation Algorithmic Aspect Recursive
Special course in Computer Science: Advanced Text Algorithms
 Special course in Computer Science: Advanced Text Algorithms Lecture 6: Alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
Special course in Computer Science: Advanced Text Algorithms Lecture 6: Alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
As of August 15, 2008, GenBank contained bases from reported sequences. The search procedure should be
 48 Bioinformatics I, WS 09-10, S. Henz (script by D. Huson) November 26, 2009 4 BLAST and BLAT Outline of the chapter: 1. Heuristics for the pairwise local alignment of two sequences 2. BLAST: search and
48 Bioinformatics I, WS 09-10, S. Henz (script by D. Huson) November 26, 2009 4 BLAST and BLAT Outline of the chapter: 1. Heuristics for the pairwise local alignment of two sequences 2. BLAST: search and
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 04: Variations of sequence alignments http://www.pitt.edu/~mcs2/teaching/biocomp/tutorials/global.html Slides adapted from Dr. Shaojie Zhang (University
EECS730: Introduction to Bioinformatics Lecture 04: Variations of sequence alignments http://www.pitt.edu/~mcs2/teaching/biocomp/tutorials/global.html Slides adapted from Dr. Shaojie Zhang (University
Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014
 Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014 Dynamic programming is a group of mathematical methods used to sequentially split a complicated problem into
Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014 Dynamic programming is a group of mathematical methods used to sequentially split a complicated problem into
Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it.
 Sequence Alignments Overview Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it. Sequence alignment means arranging
Sequence Alignments Overview Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it. Sequence alignment means arranging
Lecture 9: Core String Edits and Alignments
 Biosequence Algorithms, Spring 2005 Lecture 9: Core String Edits and Alignments Pekka Kilpeläinen University of Kuopio Department of Computer Science BSA Lecture 9: String Edits and Alignments p.1/30 III:
Biosequence Algorithms, Spring 2005 Lecture 9: Core String Edits and Alignments Pekka Kilpeläinen University of Kuopio Department of Computer Science BSA Lecture 9: String Edits and Alignments p.1/30 III:
Computational Biology Lecture 4: Overlap detection, Local Alignment, Space Efficient Needleman-Wunsch Saad Mneimneh
 Computational Biology Lecture 4: Overlap detection, Local Alignment, Space Efficient Needleman-Wunsch Saad Mneimneh Overlap detection: Semi-Global Alignment An overlap of two sequences is considered an
Computational Biology Lecture 4: Overlap detection, Local Alignment, Space Efficient Needleman-Wunsch Saad Mneimneh Overlap detection: Semi-Global Alignment An overlap of two sequences is considered an
Biology 644: Bioinformatics
 Find the best alignment between 2 sequences with lengths n and m, respectively Best alignment is very dependent upon the substitution matrix and gap penalties The Global Alignment Problem tries to find
Find the best alignment between 2 sequences with lengths n and m, respectively Best alignment is very dependent upon the substitution matrix and gap penalties The Global Alignment Problem tries to find
Pairwise Sequence Alignment: Dynamic Programming Algorithms. COMP Spring 2015 Luay Nakhleh, Rice University
 Pairwise Sequence Alignment: Dynamic Programming Algorithms COMP 571 - Spring 2015 Luay Nakhleh, Rice University DP Algorithms for Pairwise Alignment The number of all possible pairwise alignments (if
Pairwise Sequence Alignment: Dynamic Programming Algorithms COMP 571 - Spring 2015 Luay Nakhleh, Rice University DP Algorithms for Pairwise Alignment The number of all possible pairwise alignments (if
Lecture 3: February Local Alignment: The Smith-Waterman Algorithm
 CSCI1820: Sequence Alignment Spring 2017 Lecture 3: February 7 Lecturer: Sorin Istrail Scribe: Pranavan Chanthrakumar Note: LaTeX template courtesy of UC Berkeley EECS dept. Notes are also adapted from
CSCI1820: Sequence Alignment Spring 2017 Lecture 3: February 7 Lecturer: Sorin Istrail Scribe: Pranavan Chanthrakumar Note: LaTeX template courtesy of UC Berkeley EECS dept. Notes are also adapted from
Global Alignment Scoring Matrices Local Alignment Alignment with Affine Gap Penalties
 Global Alignment Scoring Matrices Local Alignment Alignment with Affine Gap Penalties From LCS to Alignment: Change the Scoring The Longest Common Subsequence (LCS) problem the simplest form of sequence
Global Alignment Scoring Matrices Local Alignment Alignment with Affine Gap Penalties From LCS to Alignment: Change the Scoring The Longest Common Subsequence (LCS) problem the simplest form of sequence
Lecture 2 Pairwise sequence alignment. Principles Computational Biology Teresa Przytycka, PhD
 Lecture 2 Pairwise sequence alignment. Principles Computational Biology Teresa Przytycka, PhD Assumptions: Biological sequences evolved by evolution. Micro scale changes: For short sequences (e.g. one
Lecture 2 Pairwise sequence alignment. Principles Computational Biology Teresa Przytycka, PhD Assumptions: Biological sequences evolved by evolution. Micro scale changes: For short sequences (e.g. one
Lecture 10. Sequence alignments
 Lecture 10 Sequence alignments Alignment algorithms: Overview Given a scoring system, we need to have an algorithm for finding an optimal alignment for a pair of sequences. We want to maximize the score
Lecture 10 Sequence alignments Alignment algorithms: Overview Given a scoring system, we need to have an algorithm for finding an optimal alignment for a pair of sequences. We want to maximize the score
Principles of Bioinformatics. BIO540/STA569/CSI660 Fall 2010
 Principles of Bioinformatics BIO540/STA569/CSI660 Fall 2010 Lecture 11 Multiple Sequence Alignment I Administrivia Administrivia The midterm examination will be Monday, October 18 th, in class. Closed
Principles of Bioinformatics BIO540/STA569/CSI660 Fall 2010 Lecture 11 Multiple Sequence Alignment I Administrivia Administrivia The midterm examination will be Monday, October 18 th, in class. Closed
CISC 889 Bioinformatics (Spring 2003) Multiple Sequence Alignment
 CISC 889 Bioinformatics (Spring 2003) Multiple Sequence Alignment Courtesy of jalview 1 Motivations Collective statistic Protein families Identification and representation of conserved sequence features
CISC 889 Bioinformatics (Spring 2003) Multiple Sequence Alignment Courtesy of jalview 1 Motivations Collective statistic Protein families Identification and representation of conserved sequence features
Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA.
 Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA. Fasta is used to compare a protein or DNA sequence to all of the
Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA. Fasta is used to compare a protein or DNA sequence to all of the
BMI/CS 576 Fall 2015 Midterm Exam
 BMI/CS 576 Fall 2015 Midterm Exam Prof. Colin Dewey Tuesday, October 27th, 2015 11:00am-12:15pm Name: KEY Write your answers on these pages and show your work. You may use the back sides of pages as necessary.
BMI/CS 576 Fall 2015 Midterm Exam Prof. Colin Dewey Tuesday, October 27th, 2015 11:00am-12:15pm Name: KEY Write your answers on these pages and show your work. You may use the back sides of pages as necessary.
.. Fall 2011 CSC 570: Bioinformatics Alexander Dekhtyar..
 .. Fall 2011 CSC 570: Bioinformatics Alexander Dekhtyar.. PAM and BLOSUM Matrices Prepared by: Jason Banich and Chris Hoover Background As DNA sequences change and evolve, certain amino acids are more
.. Fall 2011 CSC 570: Bioinformatics Alexander Dekhtyar.. PAM and BLOSUM Matrices Prepared by: Jason Banich and Chris Hoover Background As DNA sequences change and evolve, certain amino acids are more
Lecture Overview. Sequence search & alignment. Searching sequence databases. Sequence Alignment & Search. Goals: Motivations:
 Lecture Overview Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating
Lecture Overview Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating
Sequence Alignment & Search
 Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating the first version
Sequence Alignment & Search Karin Verspoor, Ph.D. Faculty, Computational Bioscience Program University of Colorado School of Medicine With credit and thanks to Larry Hunter for creating the first version
Notes on Dynamic-Programming Sequence Alignment
 Notes on Dynamic-Programming Sequence Alignment Introduction. Following its introduction by Needleman and Wunsch (1970), dynamic programming has become the method of choice for rigorous alignment of DNA
Notes on Dynamic-Programming Sequence Alignment Introduction. Following its introduction by Needleman and Wunsch (1970), dynamic programming has become the method of choice for rigorous alignment of DNA
Algorithmic Approaches for Biological Data, Lecture #20
 Algorithmic Approaches for Biological Data, Lecture #20 Katherine St. John City University of New York American Museum of Natural History 20 April 2016 Outline Aligning with Gaps and Substitution Matrices
Algorithmic Approaches for Biological Data, Lecture #20 Katherine St. John City University of New York American Museum of Natural History 20 April 2016 Outline Aligning with Gaps and Substitution Matrices
BLAST: Basic Local Alignment Search Tool Altschul et al. J. Mol Bio CS 466 Saurabh Sinha
 BLAST: Basic Local Alignment Search Tool Altschul et al. J. Mol Bio. 1990. CS 466 Saurabh Sinha Motivation Sequence homology to a known protein suggest function of newly sequenced protein Bioinformatics
BLAST: Basic Local Alignment Search Tool Altschul et al. J. Mol Bio. 1990. CS 466 Saurabh Sinha Motivation Sequence homology to a known protein suggest function of newly sequenced protein Bioinformatics
Bioinformatics for Biologists
 Bioinformatics for Biologists Sequence Analysis: Part I. Pairwise alignment and database searching Fran Lewitter, Ph.D. Director Bioinformatics & Research Computing Whitehead Institute Topics to Cover
Bioinformatics for Biologists Sequence Analysis: Part I. Pairwise alignment and database searching Fran Lewitter, Ph.D. Director Bioinformatics & Research Computing Whitehead Institute Topics to Cover
1. R. Durbin, S. Eddy, A. Krogh und G. Mitchison: Biological sequence analysis, Cambridge, 1998
 7 Multiple Sequence Alignment The exposition was prepared by Clemens GrÃP pl, based on earlier versions by Daniel Huson, Knut Reinert, and Gunnar Klau. It is based on the following sources, which are all
7 Multiple Sequence Alignment The exposition was prepared by Clemens GrÃP pl, based on earlier versions by Daniel Huson, Knut Reinert, and Gunnar Klau. It is based on the following sources, which are all
FASTA. Besides that, FASTA package provides SSEARCH, an implementation of the optimal Smith- Waterman algorithm.
 FASTA INTRODUCTION Definition (by David J. Lipman and William R. Pearson in 1985) - Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence
FASTA INTRODUCTION Definition (by David J. Lipman and William R. Pearson in 1985) - Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence
Sequence alignment theory and applications Session 3: BLAST algorithm
 Sequence alignment theory and applications Session 3: BLAST algorithm Introduction to Bioinformatics online course : IBT Sonal Henson Learning Objectives Understand the principles of the BLAST algorithm
Sequence alignment theory and applications Session 3: BLAST algorithm Introduction to Bioinformatics online course : IBT Sonal Henson Learning Objectives Understand the principles of the BLAST algorithm
FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:
 FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:56 4001 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem
FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:56 4001 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem
An Analysis of Pairwise Sequence Alignment Algorithm Complexities: Needleman-Wunsch, Smith-Waterman, FASTA, BLAST and Gapped BLAST
 An Analysis of Pairwise Sequence Alignment Algorithm Complexities: Needleman-Wunsch, Smith-Waterman, FASTA, BLAST and Gapped BLAST Alexander Chan 5075504 Biochemistry 218 Final Project An Analysis of Pairwise
An Analysis of Pairwise Sequence Alignment Algorithm Complexities: Needleman-Wunsch, Smith-Waterman, FASTA, BLAST and Gapped BLAST Alexander Chan 5075504 Biochemistry 218 Final Project An Analysis of Pairwise
Programming assignment for the course Sequence Analysis (2006)
 Programming assignment for the course Sequence Analysis (2006) Original text by John W. Romein, adapted by Bart van Houte (bart@cs.vu.nl) Introduction Please note: This assignment is only obligatory for
Programming assignment for the course Sequence Analysis (2006) Original text by John W. Romein, adapted by Bart van Houte (bart@cs.vu.nl) Introduction Please note: This assignment is only obligatory for
Pairwise Sequence Alignment: Dynamic Programming Algorithms COMP 571 Luay Nakhleh, Rice University
 1 Pairwise Sequence Alignment: Dynamic Programming Algorithms COMP 571 Luay Nakhleh, Rice University DP Algorithms for Pairwise Alignment 2 The number of all possible pairwise alignments (if gaps are allowed)
1 Pairwise Sequence Alignment: Dynamic Programming Algorithms COMP 571 Luay Nakhleh, Rice University DP Algorithms for Pairwise Alignment 2 The number of all possible pairwise alignments (if gaps are allowed)
1. R. Durbin, S. Eddy, A. Krogh und G. Mitchison: Biological sequence analysis, Cambridge, 1998
 7 Multiple Sequence Alignment The exposition was prepared by Clemens Gröpl, based on earlier versions by Daniel Huson, Knut Reinert, and Gunnar Klau. It is based on the following sources, which are all
7 Multiple Sequence Alignment The exposition was prepared by Clemens Gröpl, based on earlier versions by Daniel Huson, Knut Reinert, and Gunnar Klau. It is based on the following sources, which are all
CS313 Exercise 4 Cover Page Fall 2017
 CS313 Exercise 4 Cover Page Fall 2017 Due by the start of class on Thursday, October 12, 2017. Name(s): In the TIME column, please estimate the time you spent on the parts of this exercise. Please try
CS313 Exercise 4 Cover Page Fall 2017 Due by the start of class on Thursday, October 12, 2017. Name(s): In the TIME column, please estimate the time you spent on the parts of this exercise. Please try
In this section we describe how to extend the match refinement to the multiple case and then use T-Coffee to heuristically compute a multiple trace.
 5 Multiple Match Refinement and T-Coffee In this section we describe how to extend the match refinement to the multiple case and then use T-Coffee to heuristically compute a multiple trace. This exposition
5 Multiple Match Refinement and T-Coffee In this section we describe how to extend the match refinement to the multiple case and then use T-Coffee to heuristically compute a multiple trace. This exposition
Mouse, Human, Chimpanzee
 More Alignments 1 Mouse, Human, Chimpanzee Mouse to Human Chimpanzee to Human 2 Mouse v.s. Human Chromosome X of Mouse to Human 3 Local Alignment Given: two sequences S and T Find: substrings of S and
More Alignments 1 Mouse, Human, Chimpanzee Mouse to Human Chimpanzee to Human 2 Mouse v.s. Human Chromosome X of Mouse to Human 3 Local Alignment Given: two sequences S and T Find: substrings of S and
FastA & the chaining problem
 FastA & the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 1 Sources for this lecture: Lectures by Volker Heun, Daniel Huson and Knut Reinert,
FastA & the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 1 Sources for this lecture: Lectures by Volker Heun, Daniel Huson and Knut Reinert,
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 06: Multiple Sequence Alignment https://upload.wikimedia.org/wikipedia/commons/thumb/7/79/rplp0_90_clustalw_aln.gif/575px-rplp0_90_clustalw_aln.gif Slides
EECS730: Introduction to Bioinformatics Lecture 06: Multiple Sequence Alignment https://upload.wikimedia.org/wikipedia/commons/thumb/7/79/rplp0_90_clustalw_aln.gif/575px-rplp0_90_clustalw_aln.gif Slides
Special course in Computer Science: Advanced Text Algorithms
 Special course in Computer Science: Advanced Text Algorithms Lecture 8: Multiple alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
Special course in Computer Science: Advanced Text Algorithms Lecture 8: Multiple alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
Dynamic Programming: Sequence alignment. CS 466 Saurabh Sinha
 Dynamic Programming: Sequence alignment CS 466 Saurabh Sinha DNA Sequence Comparison: First Success Story Finding sequence similarities with genes of known function is a common approach to infer a newly
Dynamic Programming: Sequence alignment CS 466 Saurabh Sinha DNA Sequence Comparison: First Success Story Finding sequence similarities with genes of known function is a common approach to infer a newly
Lectures by Volker Heun, Daniel Huson and Knut Reinert, in particular last years lectures
 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 4.1 Sources for this lecture Lectures by Volker Heun, Daniel Huson and Knut
4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 4.1 Sources for this lecture Lectures by Volker Heun, Daniel Huson and Knut
C E N T R. Introduction to bioinformatics 2007 E B I O I N F O R M A T I C S V U F O R I N T. Lecture 13 G R A T I V. Iterative homology searching,
 C E N T R E F O R I N T E G R A T I V E B I O I N F O R M A T I C S V U Introduction to bioinformatics 2007 Lecture 13 Iterative homology searching, PSI (Position Specific Iterated) BLAST basic idea use
C E N T R E F O R I N T E G R A T I V E B I O I N F O R M A T I C S V U Introduction to bioinformatics 2007 Lecture 13 Iterative homology searching, PSI (Position Specific Iterated) BLAST basic idea use
Sequence analysis Pairwise sequence alignment
 UMF11 Introduction to bioinformatics, 25 Sequence analysis Pairwise sequence alignment 1. Sequence alignment Lecturer: Marina lexandersson 12 September, 25 here are two types of sequence alignments, global
UMF11 Introduction to bioinformatics, 25 Sequence analysis Pairwise sequence alignment 1. Sequence alignment Lecturer: Marina lexandersson 12 September, 25 here are two types of sequence alignments, global
Reconstructing long sequences from overlapping sequence fragment. Searching databases for related sequences and subsequences
 SEQUENCE ALIGNMENT ALGORITHMS 1 Why compare sequences? Reconstructing long sequences from overlapping sequence fragment Searching databases for related sequences and subsequences Storing, retrieving and
SEQUENCE ALIGNMENT ALGORITHMS 1 Why compare sequences? Reconstructing long sequences from overlapping sequence fragment Searching databases for related sequences and subsequences Storing, retrieving and
24 Grundlagen der Bioinformatik, SS 10, D. Huson, April 26, This lecture is based on the following papers, which are all recommended reading:
 24 Grundlagen der Bioinformatik, SS 10, D. Huson, April 26, 2010 3 BLAST and FASTA This lecture is based on the following papers, which are all recommended reading: D.J. Lipman and W.R. Pearson, Rapid
24 Grundlagen der Bioinformatik, SS 10, D. Huson, April 26, 2010 3 BLAST and FASTA This lecture is based on the following papers, which are all recommended reading: D.J. Lipman and W.R. Pearson, Rapid
Shortest Path Algorithm
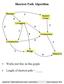 Shortest Path Algorithm C Works just fine on this graph. C Length of shortest path = Copyright 2005 DIMACS BioMath Connect Institute Robert Hochberg Dynamic Programming SP #1 Same Questions, Different
Shortest Path Algorithm C Works just fine on this graph. C Length of shortest path = Copyright 2005 DIMACS BioMath Connect Institute Robert Hochberg Dynamic Programming SP #1 Same Questions, Different
Multiple Sequence Alignment: Multidimensional. Biological Motivation
 Multiple Sequence Alignment: Multidimensional Dynamic Programming Boston University Biological Motivation Compare a new sequence with the sequences in a protein family. Proteins can be categorized into
Multiple Sequence Alignment: Multidimensional Dynamic Programming Boston University Biological Motivation Compare a new sequence with the sequences in a protein family. Proteins can be categorized into
Heuristic methods for pairwise alignment:
 Bi03c_1 Unit 03c: Heuristic methods for pairwise alignment: k-tuple-methods k-tuple-methods for alignment of pairs of sequences Bi03c_2 dynamic programming is too slow for large databases Use heuristic
Bi03c_1 Unit 03c: Heuristic methods for pairwise alignment: k-tuple-methods k-tuple-methods for alignment of pairs of sequences Bi03c_2 dynamic programming is too slow for large databases Use heuristic
Multiple Sequence Alignment. Mark Whitsitt - NCSA
 Multiple Sequence Alignment Mark Whitsitt - NCSA What is a Multiple Sequence Alignment (MA)? GMHGTVYANYAVDSSDLLLAFGVRFDDRVTGKLEAFASRAKIVHIDIDSAEIGKNKQPHV GMHGTVYANYAVEHSDLLLAFGVRFDDRVTGKLEAFASRAKIVHIDIDSAEIGKNKTPHV
Multiple Sequence Alignment Mark Whitsitt - NCSA What is a Multiple Sequence Alignment (MA)? GMHGTVYANYAVDSSDLLLAFGVRFDDRVTGKLEAFASRAKIVHIDIDSAEIGKNKQPHV GMHGTVYANYAVEHSDLLLAFGVRFDDRVTGKLEAFASRAKIVHIDIDSAEIGKNKTPHV
Sequence comparison: Local alignment
 Sequence comparison: Local alignment Genome 559: Introuction to Statistical an Computational Genomics Prof. James H. Thomas http://faculty.washington.eu/jht/gs559_217/ Review global alignment en traceback
Sequence comparison: Local alignment Genome 559: Introuction to Statistical an Computational Genomics Prof. James H. Thomas http://faculty.washington.eu/jht/gs559_217/ Review global alignment en traceback
EECS 4425: Introductory Computational Bioinformatics Fall Suprakash Datta
 EECS 4425: Introductory Computational Bioinformatics Fall 2018 Suprakash Datta datta [at] cse.yorku.ca Office: CSEB 3043 Phone: 416-736-2100 ext 77875 Course page: http://www.cse.yorku.ca/course/4425 Many
EECS 4425: Introductory Computational Bioinformatics Fall 2018 Suprakash Datta datta [at] cse.yorku.ca Office: CSEB 3043 Phone: 416-736-2100 ext 77875 Course page: http://www.cse.yorku.ca/course/4425 Many
1 Metric spaces. d(x, x) = 0 for all x M, d(x, y) = d(y, x) for all x, y M,
 1 Metric spaces For completeness, we recall the definition of metric spaces and the notions relating to measures on metric spaces. A metric space is a pair (M, d) where M is a set and d is a function from
1 Metric spaces For completeness, we recall the definition of metric spaces and the notions relating to measures on metric spaces. A metric space is a pair (M, d) where M is a set and d is a function from
Sequence alignment algorithms
 Sequence alignment algorithms Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 23 rd 27 After this lecture, you can decide when to use local and global sequence alignments
Sequence alignment algorithms Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 23 rd 27 After this lecture, you can decide when to use local and global sequence alignments
Central Issues in Biological Sequence Comparison
 Central Issues in Biological Sequence Comparison Definitions: What is one trying to find or optimize? Algorithms: Can one find the proposed object optimally or in reasonable time optimize? Statistics:
Central Issues in Biological Sequence Comparison Definitions: What is one trying to find or optimize? Algorithms: Can one find the proposed object optimally or in reasonable time optimize? Statistics:
Database Searching Using BLAST
 Mahidol University Objectives SCMI512 Molecular Sequence Analysis Database Searching Using BLAST Lecture 2B After class, students should be able to: explain the FASTA algorithm for database searching explain
Mahidol University Objectives SCMI512 Molecular Sequence Analysis Database Searching Using BLAST Lecture 2B After class, students should be able to: explain the FASTA algorithm for database searching explain
Profiles and Multiple Alignments. COMP 571 Luay Nakhleh, Rice University
 Profiles and Multiple Alignments COMP 571 Luay Nakhleh, Rice University Outline Profiles and sequence logos Profile hidden Markov models Aligning profiles Multiple sequence alignment by gradual sequence
Profiles and Multiple Alignments COMP 571 Luay Nakhleh, Rice University Outline Profiles and sequence logos Profile hidden Markov models Aligning profiles Multiple sequence alignment by gradual sequence
Today s Lecture. Edit graph & alignment algorithms. Local vs global Computational complexity of pairwise alignment Multiple sequence alignment
 Today s Lecture Edit graph & alignment algorithms Smith-Waterman algorithm Needleman-Wunsch algorithm Local vs global Computational complexity of pairwise alignment Multiple sequence alignment 1 Sequence
Today s Lecture Edit graph & alignment algorithms Smith-Waterman algorithm Needleman-Wunsch algorithm Local vs global Computational complexity of pairwise alignment Multiple sequence alignment 1 Sequence
GLOBEX Bioinformatics (Summer 2015) Multiple Sequence Alignment
 GLOBEX Bioinformatics (Summer 2015) Multiple Sequence Alignment Scoring Dynamic Programming algorithms Heuristic algorithms CLUSTAL W Courtesy of jalview Motivations Collective (or aggregate) statistic
GLOBEX Bioinformatics (Summer 2015) Multiple Sequence Alignment Scoring Dynamic Programming algorithms Heuristic algorithms CLUSTAL W Courtesy of jalview Motivations Collective (or aggregate) statistic
Data Mining Technologies for Bioinformatics Sequences
 Data Mining Technologies for Bioinformatics Sequences Deepak Garg Computer Science and Engineering Department Thapar Institute of Engineering & Tecnology, Patiala Abstract Main tool used for sequence alignment
Data Mining Technologies for Bioinformatics Sequences Deepak Garg Computer Science and Engineering Department Thapar Institute of Engineering & Tecnology, Patiala Abstract Main tool used for sequence alignment
BLAST. Basic Local Alignment Search Tool. Used to quickly compare a protein or DNA sequence to a database.
 BLAST Basic Local Alignment Search Tool Used to quickly compare a protein or DNA sequence to a database. There is no such thing as a free lunch BLAST is fast and highly sensitive compared to competitors.
BLAST Basic Local Alignment Search Tool Used to quickly compare a protein or DNA sequence to a database. There is no such thing as a free lunch BLAST is fast and highly sensitive compared to competitors.
COS 551: Introduction to Computational Molecular Biology Lecture: Oct 17, 2000 Lecturer: Mona Singh Scribe: Jacob Brenner 1. Database Searching
 COS 551: Introduction to Computational Molecular Biology Lecture: Oct 17, 2000 Lecturer: Mona Singh Scribe: Jacob Brenner 1 Database Searching In database search, we typically have a large sequence database
COS 551: Introduction to Computational Molecular Biology Lecture: Oct 17, 2000 Lecturer: Mona Singh Scribe: Jacob Brenner 1 Database Searching In database search, we typically have a large sequence database
Computational Biology Lecture 6: Affine gap penalty function, multiple sequence alignment Saad Mneimneh
 Computational Biology Lecture 6: Affine gap penalty function, multiple sequence alignment Saad Mneimneh We saw earlier how we can use a concave gap penalty function γ, i.e. one that satisfies γ(x+1) γ(x)
Computational Biology Lecture 6: Affine gap penalty function, multiple sequence alignment Saad Mneimneh We saw earlier how we can use a concave gap penalty function γ, i.e. one that satisfies γ(x+1) γ(x)
Alignment of Long Sequences
 Alignment of Long Sequences BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2009 Mark Craven craven@biostat.wisc.edu Pairwise Whole Genome Alignment: Task Definition Given a pair of genomes (or other large-scale
Alignment of Long Sequences BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2009 Mark Craven craven@biostat.wisc.edu Pairwise Whole Genome Alignment: Task Definition Given a pair of genomes (or other large-scale
Dynamic Programming (cont d) CS 466 Saurabh Sinha
 Dynamic Programming (cont d) CS 466 Saurabh Sinha Spliced Alignment Begins by selecting either all putative exons between potential acceptor and donor sites or by finding all substrings similar to the
Dynamic Programming (cont d) CS 466 Saurabh Sinha Spliced Alignment Begins by selecting either all putative exons between potential acceptor and donor sites or by finding all substrings similar to the
Gene regulation. DNA is merely the blueprint Shared spatially (among all tissues) and temporally But cells manage to differentiate
 Gene regulation DNA is merely the blueprint Shared spatially (among all tissues) and temporally But cells manage to differentiate Especially but not only during developmental stage And cells respond to
Gene regulation DNA is merely the blueprint Shared spatially (among all tissues) and temporally But cells manage to differentiate Especially but not only during developmental stage And cells respond to
MSCBIO 2070/02-710: Computational Genomics, Spring A4: spline, HMM, clustering, time-series data analysis, RNA-folding
 MSCBIO 2070/02-710:, Spring 2015 A4: spline, HMM, clustering, time-series data analysis, RNA-folding Due: April 13, 2015 by email to Silvia Liu (silvia.shuchang.liu@gmail.com) TA in charge: Silvia Liu
MSCBIO 2070/02-710:, Spring 2015 A4: spline, HMM, clustering, time-series data analysis, RNA-folding Due: April 13, 2015 by email to Silvia Liu (silvia.shuchang.liu@gmail.com) TA in charge: Silvia Liu
Sequence clustering. Introduction. Clustering basics. Hierarchical clustering
 Sequence clustering Introduction Data clustering is one of the key tools used in various incarnations of data-mining - trying to make sense of large datasets. It is, thus, natural to ask whether clustering
Sequence clustering Introduction Data clustering is one of the key tools used in various incarnations of data-mining - trying to make sense of large datasets. It is, thus, natural to ask whether clustering
Comparison of Sequence Similarity Measures for Distant Evolutionary Relationships
 Comparison of Sequence Similarity Measures for Distant Evolutionary Relationships Abhishek Majumdar, Peter Z. Revesz Department of Computer Science and Engineering, University of Nebraska-Lincoln, Lincoln,
Comparison of Sequence Similarity Measures for Distant Evolutionary Relationships Abhishek Majumdar, Peter Z. Revesz Department of Computer Science and Engineering, University of Nebraska-Lincoln, Lincoln,
Pairwise Sequence alignment Basic Algorithms
 Pairwise Sequence alignment Basic Algorithms Agenda - Previous Lesson: Minhala - + Biological Story on Biomolecular Sequences - + General Overview of Problems in Computational Biology - Reminder: Dynamic
Pairwise Sequence alignment Basic Algorithms Agenda - Previous Lesson: Minhala - + Biological Story on Biomolecular Sequences - + General Overview of Problems in Computational Biology - Reminder: Dynamic
Biology 644: Bioinformatics
 A statistical Markov model in which the system being modeled is assumed to be a Markov process with unobserved (hidden) states in the training data. First used in speech and handwriting recognition In
A statistical Markov model in which the system being modeled is assumed to be a Markov process with unobserved (hidden) states in the training data. First used in speech and handwriting recognition In
Dynamic Programming & Smith-Waterman algorithm
 m m Seminar: Classical Papers in Bioinformatics May 3rd, 2010 m m 1 2 3 m m Introduction m Definition is a method of solving problems by breaking them down into simpler steps problem need to contain overlapping
m m Seminar: Classical Papers in Bioinformatics May 3rd, 2010 m m 1 2 3 m m Introduction m Definition is a method of solving problems by breaking them down into simpler steps problem need to contain overlapping
Biostrings. Martin Morgan Bioconductor / Fred Hutchinson Cancer Research Center Seattle, WA, USA June 2009
 Biostrings Martin Morgan Bioconductor / Fred Hutchinson Cancer Research Center Seattle, WA, USA 15-19 June 2009 Biostrings Representation DNA, RNA, amino acid, and general biological strings Manipulation
Biostrings Martin Morgan Bioconductor / Fred Hutchinson Cancer Research Center Seattle, WA, USA 15-19 June 2009 Biostrings Representation DNA, RNA, amino acid, and general biological strings Manipulation
Bioinformatics explained: Smith-Waterman
 Bioinformatics Explained Bioinformatics explained: Smith-Waterman May 1, 2007 CLC bio Gustav Wieds Vej 10 8000 Aarhus C Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com info@clcbio.com
Bioinformatics Explained Bioinformatics explained: Smith-Waterman May 1, 2007 CLC bio Gustav Wieds Vej 10 8000 Aarhus C Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com info@clcbio.com
Similarity Searches on Sequence Databases
 Similarity Searches on Sequence Databases Lorenza Bordoli Swiss Institute of Bioinformatics EMBnet Course, Zürich, October 2004 Swiss Institute of Bioinformatics Swiss EMBnet node Outline Importance of
Similarity Searches on Sequence Databases Lorenza Bordoli Swiss Institute of Bioinformatics EMBnet Course, Zürich, October 2004 Swiss Institute of Bioinformatics Swiss EMBnet node Outline Importance of
HIDDEN MARKOV MODELS AND SEQUENCE ALIGNMENT
 HIDDEN MARKOV MODELS AND SEQUENCE ALIGNMENT - Swarbhanu Chatterjee. Hidden Markov models are a sophisticated and flexible statistical tool for the study of protein models. Using HMMs to analyze proteins
HIDDEN MARKOV MODELS AND SEQUENCE ALIGNMENT - Swarbhanu Chatterjee. Hidden Markov models are a sophisticated and flexible statistical tool for the study of protein models. Using HMMs to analyze proteins
Biological Sequence Matching Using Fuzzy Logic
 International Journal of Scientific & Engineering Research Volume 2, Issue 7, July-2011 1 Biological Sequence Matching Using Fuzzy Logic Nivit Gill, Shailendra Singh Abstract: Sequence alignment is the
International Journal of Scientific & Engineering Research Volume 2, Issue 7, July-2011 1 Biological Sequence Matching Using Fuzzy Logic Nivit Gill, Shailendra Singh Abstract: Sequence alignment is the
Long Read RNA-seq Mapper
 UNIVERSITY OF ZAGREB FACULTY OF ELECTRICAL ENGENEERING AND COMPUTING MASTER THESIS no. 1005 Long Read RNA-seq Mapper Josip Marić Zagreb, February 2015. Table of Contents 1. Introduction... 1 2. RNA Sequencing...
UNIVERSITY OF ZAGREB FACULTY OF ELECTRICAL ENGENEERING AND COMPUTING MASTER THESIS no. 1005 Long Read RNA-seq Mapper Josip Marić Zagreb, February 2015. Table of Contents 1. Introduction... 1 2. RNA Sequencing...
Multiple Sequence Alignment (MSA)
 I519 Introduction to Bioinformatics, Fall 2013 Multiple Sequence Alignment (MSA) Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Outline Multiple sequence alignment (MSA) Generalize
I519 Introduction to Bioinformatics, Fall 2013 Multiple Sequence Alignment (MSA) Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Outline Multiple sequence alignment (MSA) Generalize
Sequence Alignment (chapter 6) p The biological problem p Global alignment p Local alignment p Multiple alignment
 Sequence lignment (chapter 6) p The biological problem p lobal alignment p Local alignment p Multiple alignment Local alignment: rationale p Otherwise dissimilar proteins may have local regions of similarity
Sequence lignment (chapter 6) p The biological problem p lobal alignment p Local alignment p Multiple alignment Local alignment: rationale p Otherwise dissimilar proteins may have local regions of similarity
Sequencing Alignment I
 Sequencing Alignment I Lectures 16 Nov 21, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20 Johnson Hall (JHN) 022 1 Outline: Sequence
Sequencing Alignment I Lectures 16 Nov 21, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20 Johnson Hall (JHN) 022 1 Outline: Sequence
Salvador Capella-Gutiérrez, Jose M. Silla-Martínez and Toni Gabaldón
 trimal: a tool for automated alignment trimming in large-scale phylogenetics analyses Salvador Capella-Gutiérrez, Jose M. Silla-Martínez and Toni Gabaldón Version 1.2b Index of contents 1. General features
trimal: a tool for automated alignment trimming in large-scale phylogenetics analyses Salvador Capella-Gutiérrez, Jose M. Silla-Martínez and Toni Gabaldón Version 1.2b Index of contents 1. General features
Alignment of Pairs of Sequences
 Bi03a_1 Unit 03a: Alignment of Pairs of Sequences Partners for alignment Bi03a_2 Protein 1 Protein 2 =amino-acid sequences (20 letter alphabeth + gap) LGPSSKQTGKGS-SRIWDN LN-ITKSAGKGAIMRLGDA -------TGKG--------
Bi03a_1 Unit 03a: Alignment of Pairs of Sequences Partners for alignment Bi03a_2 Protein 1 Protein 2 =amino-acid sequences (20 letter alphabeth + gap) LGPSSKQTGKGS-SRIWDN LN-ITKSAGKGAIMRLGDA -------TGKG--------
TCCAGGTG-GAT TGCAAGTGCG-T. Local Sequence Alignment & Heuristic Local Aligners. Review: Probabilistic Interpretation. Chance or true homology?
 Local Sequence Alignment & Heuristic Local Aligners Lectures 18 Nov 28, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20 Johnson Hall
Local Sequence Alignment & Heuristic Local Aligners Lectures 18 Nov 28, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20 Johnson Hall
New Implementation for the Multi-sequence All-Against-All Substring Matching Problem
 New Implementation for the Multi-sequence All-Against-All Substring Matching Problem Oana Sandu Supervised by Ulrike Stege In collaboration with Chris Upton, Alex Thomo, and Marina Barsky University of
New Implementation for the Multi-sequence All-Against-All Substring Matching Problem Oana Sandu Supervised by Ulrike Stege In collaboration with Chris Upton, Alex Thomo, and Marina Barsky University of
Stephen Scott.
 1 / 33 sscott@cse.unl.edu 2 / 33 Start with a set of sequences In each column, residues are homolgous Residues occupy similar positions in 3D structure Residues diverge from a common ancestral residue
1 / 33 sscott@cse.unl.edu 2 / 33 Start with a set of sequences In each column, residues are homolgous Residues occupy similar positions in 3D structure Residues diverge from a common ancestral residue
PROTEIN MULTIPLE ALIGNMENT MOTIVATION: BACKGROUND: Marina Sirota
 Marina Sirota MOTIVATION: PROTEIN MULTIPLE ALIGNMENT To study evolution on the genetic level across a wide range of organisms, biologists need accurate tools for multiple sequence alignment of protein
Marina Sirota MOTIVATION: PROTEIN MULTIPLE ALIGNMENT To study evolution on the genetic level across a wide range of organisms, biologists need accurate tools for multiple sequence alignment of protein
Chapter 6 Continued: Partitioning Methods
 Chapter 6 Continued: Partitioning Methods Partitioning methods fix the number of clusters k and seek the best possible partition for that k. The goal is to choose the partition which gives the optimal
Chapter 6 Continued: Partitioning Methods Partitioning methods fix the number of clusters k and seek the best possible partition for that k. The goal is to choose the partition which gives the optimal
Inexact Matching, Alignment. See Gusfield, Chapter 9 Dasgupta et al. Chapter 6 (Dynamic Programming)
 Inexact Matching, Alignment See Gusfield, Chapter 9 Dasgupta et al. Chapter 6 (Dynamic Programming) Outline Yet more applications of generalized suffix trees, when combined with a least common ancestor
Inexact Matching, Alignment See Gusfield, Chapter 9 Dasgupta et al. Chapter 6 (Dynamic Programming) Outline Yet more applications of generalized suffix trees, when combined with a least common ancestor
Important Example: Gene Sequence Matching. Corrigiendum. Central Dogma of Modern Biology. Genetics. How Nucleotides code for Amino Acids
 Important Example: Gene Sequence Matching Century of Biology Two views of computer science s relationship to biology: Bioinformatics: computational methods to help discover new biology from lots of data
Important Example: Gene Sequence Matching Century of Biology Two views of computer science s relationship to biology: Bioinformatics: computational methods to help discover new biology from lots of data
Lab 4: Multiple Sequence Alignment (MSA)
 Lab 4: Multiple Sequence Alignment (MSA) The objective of this lab is to become familiar with the features of several multiple alignment and visualization tools, including the data input and output, basic
Lab 4: Multiple Sequence Alignment (MSA) The objective of this lab is to become familiar with the features of several multiple alignment and visualization tools, including the data input and output, basic
ON HEURISTIC METHODS IN NEXT-GENERATION SEQUENCING DATA ANALYSIS
 ON HEURISTIC METHODS IN NEXT-GENERATION SEQUENCING DATA ANALYSIS Ivan Vogel Doctoral Degree Programme (1), FIT BUT E-mail: xvogel01@stud.fit.vutbr.cz Supervised by: Jaroslav Zendulka E-mail: zendulka@fit.vutbr.cz
ON HEURISTIC METHODS IN NEXT-GENERATION SEQUENCING DATA ANALYSIS Ivan Vogel Doctoral Degree Programme (1), FIT BUT E-mail: xvogel01@stud.fit.vutbr.cz Supervised by: Jaroslav Zendulka E-mail: zendulka@fit.vutbr.cz
BGGN 213 Foundations of Bioinformatics Barry Grant
 BGGN 213 Foundations of Bioinformatics Barry Grant http://thegrantlab.org/bggn213 Recap From Last Time: 25 Responses: https://tinyurl.com/bggn213-02-f17 Why ALIGNMENT FOUNDATIONS Why compare biological
BGGN 213 Foundations of Bioinformatics Barry Grant http://thegrantlab.org/bggn213 Recap From Last Time: 25 Responses: https://tinyurl.com/bggn213-02-f17 Why ALIGNMENT FOUNDATIONS Why compare biological
Dynamic Programming Course: A structure based flexible search method for motifs in RNA. By: Veksler, I., Ziv-Ukelson, M., Barash, D.
 Dynamic Programming Course: A structure based flexible search method for motifs in RNA By: Veksler, I., Ziv-Ukelson, M., Barash, D., Kedem, K Outline Background Motivation RNA s structure representations
Dynamic Programming Course: A structure based flexible search method for motifs in RNA By: Veksler, I., Ziv-Ukelson, M., Barash, D., Kedem, K Outline Background Motivation RNA s structure representations
Regular Expression Constrained Sequence Alignment
 Regular Expression Constrained Sequence Alignment By Abdullah N. Arslan Department of Computer science University of Vermont Presented by Tomer Heber & Raz Nissim otivation When comparing two proteins,
Regular Expression Constrained Sequence Alignment By Abdullah N. Arslan Department of Computer science University of Vermont Presented by Tomer Heber & Raz Nissim otivation When comparing two proteins,
Multiple Sequence Alignment Sum-of-Pairs and ClustalW. Ulf Leser
 Multiple Sequence Alignment Sum-of-Pairs and ClustalW Ulf Leser This Lecture Multiple Sequence Alignment The problem Theoretical approach: Sum-of-Pairs scores Practical approach: ClustalW Ulf Leser: Bioinformatics,
Multiple Sequence Alignment Sum-of-Pairs and ClustalW Ulf Leser This Lecture Multiple Sequence Alignment The problem Theoretical approach: Sum-of-Pairs scores Practical approach: ClustalW Ulf Leser: Bioinformatics,
An improved algorithm for the regular expression constrained multiple sequence alignment problem
 An improved algorithm for the regular expression constrained multiple sequence alignment problem Abdullah N. Arslan and Dan He Department of Computer Science University of Vermont Burlington, VT 05405,
An improved algorithm for the regular expression constrained multiple sequence alignment problem Abdullah N. Arslan and Dan He Department of Computer Science University of Vermont Burlington, VT 05405,
FINDING APPROXIMATE REPEATS WITH MULTIPLE SPACED SEEDS
 FINDING APPROXIMATE REPEATS WITH MULTIPLE SPACED SEEDS FINDING APPROXIMATE REPEATS IN DNA SEQUENCES USING MULTIPLE SPACED SEEDS By SARAH BANYASSADY, B.S. A Thesis Submitted to the School of Graduate Studies
FINDING APPROXIMATE REPEATS WITH MULTIPLE SPACED SEEDS FINDING APPROXIMATE REPEATS IN DNA SEQUENCES USING MULTIPLE SPACED SEEDS By SARAH BANYASSADY, B.S. A Thesis Submitted to the School of Graduate Studies
6.047 / Computational Biology: Genomes, Networks, Evolution Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Sequence Alignment. part 2
 Sequence Alignment part 2 Dynamic programming with more realistic scoring scheme Using the same initial sequences, we ll look at a dynamic programming example with a scoring scheme that selects for matches
Sequence Alignment part 2 Dynamic programming with more realistic scoring scheme Using the same initial sequences, we ll look at a dynamic programming example with a scoring scheme that selects for matches
