Contents 1 Admin 2 Testing hypotheses tests 4 Simulation 5 Parallelization Admin
|
|
|
- Katrina Lindsay Harrington
- 5 years ago
- Views:
Transcription
1 magrittr t F F
2 .. NA library(pacman) p_load(dplyr) x <- data.frame(a = c(1:4, NA), b = c("..", 1, 2, 3, "..")) x %>% as_tibble() ## # A tibble: 5 x 2 ## a b ## <int> <chr> ## ## ## ## ## 5 NA.. x <- data.frame(a = c(1:4, NA), b = c("..", 1, 2, 3, "..")) x[x == ".."] <- NA x %>% as_tibble() ## # A tibble: 5 x 2 ## a b ## <int> <chr> ## 1 1 <NA> ## ## ## ## 5 NA <NA> b x <- mutate(x, b = as.numeric(b)) replace mutate replace() dplyr mutate() replace() replace() as.numeric() # Define our function rep_na <- function(y) replace(y, which(y == ".."), NA) # Make some data x <- data.frame(a = c(1:4, NA), b = c("..", 1, 2, 3, "..")) %>% as_tibble()
3 # Replace.. with NA x %>% mutate(b = rep_na(b) %>% as.numeric()) ## # A tibble: 5 x 2 ## a b ## <int> <dbl> ## 1 1 NA ## ## ## ## 5 NA NA mutate_all() # Make some data x <- data.frame(a = c(1:4, NA), b = c("..", 1, 2, 3, "..")) # Replace.. with NA x %>% mutate_all(rep_na) %>% mutate_all(as.numeric) %>% as_tibble() ## # A tibble: 5 x 2 ## a b ## <dbl> <dbl> ## 1 1 NA ## ## ## ## 5 NA NA mutate_all() mutate_at() mutate_each() mutate_if() dplyr NA ".." "" %in% x[x %in% c("..", "")] <- NA I() x + I(xˆ2) + I(xˆ3) : x1:x2 * x1*x2 x1 + x2 + x1:x2
4 dplyr lfe readr magrittr parallel parallel auto.csv y x y x y x y x # Settings options(stringsasfactors = F) # Packages library(pacman) p_load(dplyr, lfe, magrittr, readr)
5 # Directories dir_data <- "/Users/edwardarubin/Dropbox/Teaching/ARE212/Section05/" # Load the dataset from CSV cars <- paste0(dir_data, "auto.csv") %>% read_csv() # Functions ---- # Function to convert tibble, data.frame, or tbl_df to matrix to_matrix <- function(the_df, vars) { # Create a matrix from variables in var new_mat <- the_df %>% # Select the columns given in 'vars' select_(.dots = vars) %>% # Convert to matrix as.matrix() # Return 'new_mat' return(new_mat) } # Function for OLS coefficient estimates b_ols <- function(data, y_var, X_vars, intercept = TRUE) { # Require the 'dplyr' package require(dplyr) # Create the y matrix y <- to_matrix(the_df = data, vars = y_var) # Create the X matrix X <- to_matrix(the_df = data, vars = X_vars) # If 'intercept' is TRUE, then add a column of ones if (intercept == T) { # Bind a column of ones to X X <- cbind(1, X) # Name the new column "intercept" colnames(x) <- c("intercept", X_vars) } # Calculate beta hat beta_hat <- solve(t(x) %*% X) %*% t(x) %*% y # Return beta_hat return(beta_hat) } magrittr magrittr magrittr dplyr %>%
6 t β j γ α t j = s 2 b j γ { (X X) 1} jj = b j γ (b j ) b j β j s 2 σ 2 (X X) 1 t_stat <- function(data, y_var, X_vars, gamma, intercept = T) { } # Turn data into matrices y <- to_matrix(data, y_var) X <- to_matrix(data, X_vars) # Add intercept if requested if (intercept == T) X <- cbind(1, X) # Calculate n and k for degrees of freedom n <- nrow(x) k <- ncol(x) # Estimate coefficients b <- b_ols(data, y_var, X_vars, intercept) # Calculate OLS residuals e <- y - X %*% b # Calculate s^2 s2 <- (t(e) %*% e) / (n-k) # Force s2 to numeric s2 %<>% as.numeric() # Inverse of X'X XX_inv <- solve(t(x) %*% X) # Standard error se <- sqrt(s2 * diag(xx_inv)) # Vector of _t_ statistics t_stats <- (b - gamma) / se # Return the _t_ statistics return(t_stats) { (X X) 1} jj
7 t_stat() γ = [0 0 0] felm() coefficients summary() felm() coefficients summary() %$% magrittr %$% %$% %$% $ cor(cars$price, cars$weight) cars %$% cor(price, weight) cor(cars$price, cars$weight) ## [1] cars %$% cor(price, weight) ## [1] magrittr vignette("magrittr") browsevignettes() t felm() t_stat(cars, y_var = "price", X_vars = c("mpg", "weight"), gamma = 0, intercept = T) ## price ## intercept ## mpg ## weight felm() felm(price ~ mpg + weight, cars) %>% summary() %$% (coefficients)[,3] ## (Intercept) mpg weight ## summary() summary() felm() summary.felm() summary() (coefficients)[,3] coefficients [,3]
8 n k p t j p = (t > t j ) 2 pt(q, df) q df pt() lower.tail lower.tail TRUE pt() q lower.tail FALSE # The default: lower.tail = TRUE pt(q = 2, df = 15) ## [1] # Setting lower.tail = TRUE pt(q = 2, df = 15, lower.tail = T) ## [1] # Setting lower.tail = FALSE pt(q = 2, df = 15, lower.tail = F) ## [1] t_stats pt(q = abs(t_stats), df = n-k, lower.tail = F) * 2 felm() ols() as.vector() 1 k k 1 round() ols <- function(data, y_var, X_vars, intercept = T) { # Turn data into matrices y <- to_matrix(data, y_var) gamma
9 } X <- to_matrix(data, X_vars) # Add intercept if requested if (intercept == T) X <- cbind(1, X) # Calculate n and k for degrees of freedom n <- nrow(x) k <- ncol(x) # Estimate coefficients b <- b_ols(data, y_var, X_vars, intercept) # Calculate OLS residuals e <- y - X %*% b # Calculate s^2 s2 <- (t(e) %*% e) / (n-k) # Update s2 to numeric s2 %<>% as.numeric() # Inverse of X'X XX_inv <- solve(t(x) %*% X) # Standard error se <- sqrt(s2 * diag(xx_inv)) # Vector of _t_ statistics t_stats <- (b - 0) / se # Calculate the p-values p_values = pt(q = abs(t_stats), df = n-k, lower.tail = F) * 2 # Nice table (data.frame) of results results <- data.frame( # The rows have the coef. names effect = rownames(b), # Estimated coefficients coef = as.vector(b) %>% round(3), # Standard errors std_error = as.vector(se) %>% round(3), # t statistics t_stat = as.vector(t_stats) %>% round(3), # p-values p_value = as.vector(p_values) %>% round(4) ) # Return the results return(results) felm() summary() ols(data = cars, y_var = "price", X_vars = c("mpg", "weight"), intercept = T) ## effect coef std_error t_stat p_value ## 1 intercept ## 2 mpg
10 ## 3 weight felm(price ~ mpg + weight, cars) %>% summary() %$% coefficients ## Estimate Std. Error t value Pr(> t ) ## (Intercept) ## mpg ## weight F β 1 β 2 F y = β 0 + x 1 β 1 + x 2 β 2 + x 3 β 3 + ε β 1 β 2 β 3 R r R j k j k R j 1 r β 1 = 1 β 2 = 0 β 3 = 7 R = r = β 1 β 2 β 3 R = r = F R r Rβ = r β
11 [ (Rb r) R (X X) 1] (Rb r)/j F = s 2 [ = (Rb) R (X X) 1] (Rb)/J s 2 r = 0 J = 3 F β i = 0 R r joint_test <- function(data, y_var, X_vars) { # Turn data into matrices y <- to_matrix(data, y_var) X <- to_matrix(data, X_vars) # Add intercept X <- cbind(1, X) # Name the new column "intercept" colnames(x) <- c("intercept", X_vars) # Calculate n and k for degrees of freedom n <- nrow(x) k <- ncol(x) # J is k-1 J <- k - 1 # Create the R matrix: bind a column of zeros # onto a J-by-J identity matrix R <- cbind(0, diag(j)) # Estimate coefficients b <- b_ols(data, y_var, X_vars) # Calculate OLS residuals e <- y - X %*% b # Calculate s^2 s2 <- (t(e) %*% e) / (n-k) # Force s2 to numeric s2 %<>% as.numeric() # Create the inner matrix R(X'X)^(-1)R' RXXR <- R %*% solve(t(x) %*% X) %*% t(r) # Calculate the F stat f_stat <- t(r %*% b) %*% solve(rxxr) %*% (R %*% b) / J / s2 # Calculate the p-value p_value <- pf(q = f_stat, df1 = J, df2 = n-k, lower.tail = F) # Create a data.frame of the f stat. and p-value results <- data.frame( f_stat = f_stat %>% as.vector(),
12 } p_value = p_value %>% as.vector()) return(results) F joint_test(data = cars, y_var = "price", X_vars = c("mpg", "weight")) ## f_stat p_value ## e-06 felm() felm(price ~ mpg + weight, cars) %>% summary() %$% F.fstat ## F df1 df2 p ## e e e e-06 F ols() ols() F F ols() F if intercept = F intercept FALSE if (intercept == F) stop("no intercept!") intercept = F warning() F ols_joint() ols_joint <- function(data, y_var, X_vars, intercept = T) { # Run the ols() function ols_results <- ols(data, y_var, X_vars, intercept) # If intercept is T, run the joint_test() function # Otherwise, define joint_results to be NA and # issue a warning if (intercept == T) {
13 } joint_results <- joint_test(data, y_var, X_vars) } else { warning("no intercept: will not perform F test.") joint_results <- data.frame( f_stat = NA, p_value = NA) } # Create the results list results <- list(ols_results, joint_results) # Return the results list return(results) ols_joint(data = cars, y_var = "price", X_vars = c("mpg", "weight"), intercept = T) ## [[1]] ## effect coef std_error t_stat p_value ## 1 intercept ## 2 mpg ## 3 weight ## ## [[2]] ## f_stat p_value ## e-06 ols_joint(data = cars, y_var = "price", X_vars = c("mpg", "weight"), intercept = F) ## Warning in ols_joint(data = cars, y_var = "price", X_vars = c("mpg", ## "weight"), : No intercept: will not perform F test. ## [[1]] ## effect coef std_error t_stat p_value ## 1 mpg ## 2 weight ## ## [[2]] ## f_stat p_value ## 1 NA NA
14 sample_size y = β 0 + β 1 x + ε β 0 = 7 β 1 = 0.5 x ε # Function to generate data gen_data <- function(sample_size) { } # Create data.frame with random x and error data_df <- data.frame( x = rnorm(sample_size), e = rnorm(sample_size)) # Calculate y = x + e; drop 'e' data_df %<>% mutate(y = * x + e) %>% select(-e) # Return data_df return(data_df) gen_data() ols() one_sim <- function(sample_size) { # Estimate via OLS ols_est <- ols(data = gen_data(sample_size), y_var = "y", X_vars = "x") # Grab the estimated coefficient on x # (the second element of 'coef') b1 <- ols_est %$% coef[2] # Grab the second p-value # (the first p-value is for the intercept) p_value <- ols_est %$% p_value[2] # Return a data.frame with b1 and p_value %<>% magrittr %>% %<>% tmp %<>% mean() tmp tmp %<>% tmp <- tmp %>% mean() tmp <- mean(tmp) %<>% %>%
15 } return(data.frame(b1, p_value)) n_sims lapply() replicate() replicate() n expr replicate() simplify simplify = F bind_rows() n_sims n seed seed ols_sim <- function(n_sims, sample_size, seed = 12345) { } # Set the seed set.seed(seed) # Run one_sim n_sims times; convert results to data.frame sim_df <- replicate( n = n_sims, expr = one_sim(sample_size), simplify = F ) %>% bind_rows() # Return sim_df return(sim_df) ols_sim() # Run ols_sim for sample size of 10 sim10 <- ols_sim(n_sims = 1e3, sample_size = 10) # Run ols_sim for sample size of 100 sim100 <- ols_sim(n_sims = 1e3, sample_size = 100) ggplot2 ggplot2 p_load(ggplot2, ggthemes) ggplot(data = sim10, aes(x = p_value)) + stat_density(fill = "grey20", alpha = 0.9) + replicate() lapply() lapply() lapply(x = rep(sample_size, n_sims), FUN = one_sim) ggplot2 ggthemes
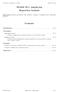
16 geom_vline(xintercept = 0.05, color = "deeppink2", size = 1) + theme_pander() + xlab(expression(paste(italic("p"), "-Value"))) + ylab("density") + ggtitle(expression(paste("distribution of ", italic(p), "-Values from 1,000 simulations with sample size 10"))) Distribution of p Values from 1,000 simulations with sample size 10 2 Density p Value ggplot(data = sim100, aes(x = p_value)) + stat_density(fill = "grey20", alpha = 0.9) + geom_vline(xintercept = 0.05, color = "deeppink2", size = 1) + xlim(0, 0.1) + theme_pander() + xlab(expression(paste(italic("p"), "-Value"))) + ylab("density") + ggtitle(expression(paste("distribution of ", italic(p), "-Values from 1,000 simulations with sample size 100")))
17 Distribution of p Values from 1,000 simulations with sample size Density p Value β 1 β 1 ggplot(data = sim10, aes(x = b1)) + stat_density(fill = "grey70") + geom_vline(xintercept = 0.5, color = "darkviolet", size = 1) + theme_pander() + xlab(expression(paste(beta[1], " estimate"))) + ylab("density") + ggtitle(expression(paste("distribution of ", beta[1], " estimates from 1,000 simulations with sample size 10")))
18 Distribution of β 1 estimates from 1,000 simulations with sample size Density β 1 estimate β 1 ggplot(data = sim100, aes(x = b1)) + stat_density(fill = "grey70") + geom_vline(xintercept = 0.5, color = "darkviolet", size = 1) + theme_pander() + xlab(expression(paste(beta[1], " estimate"))) + ylab("density") + ggtitle(expression(paste("distribution of ", beta[1], " estimates from 1,000 simulations with sample size 100")))
19 Distribution of β 1 estimates from 1,000 simulations with sample size Density β 1 estimate β 1 parallel parallel lapply() mclapply() parallel?mclapply
20 mclapply() lapply()x FUN mc.cores p_load(parallel) mc.cores detectcores() detectcores() ## [1] 4 ols_sim() replicate() lapply() lapply() mclapply() ols_sim_mc <- function(n_sims, sample_size, seed = 12345) { } # Require the parallel package require(parallel) # Set the seed set.seed(seed) # Run one_sim n_sims times; convert results to data.frame sim_df <- mclapply( X = rep(x = sample_size, times = n_sims), FUN = one_sim, # Specify that we want 4 cores mc.cores = 4 ) %>% bind_rows() # Return sim_df return(sim_df) proc.time() # Run ols_sim for sample size of 10 start1 <- proc.time() sim10 <- ols_sim(n_sims = 1e4, sample_size = 10) stop1 <- proc.time() # Run ols_sim for sample size of 100 start2 <- proc.time() sim100 <- ols_sim(n_sims = 1e4, sample_size = 100) stop2 <- proc.time()
21 # Run ols_sim_mc for sample size of 10 start3 <- proc.time() sim10_mc <- ols_sim_mc(n_sims = 1e4, sample_size = 10) stop3 <- proc.time() # Run ols_sim_mc for sample size of 100 start4 <- proc.time() sim100_mc <- ols_sim_mc(n_sims = 1e4, sample_size = 100) stop4 <- proc.time() stop1 - start1 ## user system elapsed ## stop2 - start2 ## user system elapsed ## stop3 - start3 ## user system elapsed ## stop4 - start4 ## user system elapsed ## elapsed parallel parlapply() mclapply() detectcores() parallel mclapply() mc.cores
22 p_load(parallel) detectcores() ## [1] 4 makecluster() clusterexport() clusterevalq() makecluster() makecluster() # Make the cluster cl <- makecluster(4) clusterexport() clusterevalq() ls() dplyr magrittr clusterevalq() dplyr magrittr # Load packages to cluster clusterevalq(cl = cl, { # Load packages library(dplyr) library(magrittr) }) clusterevalq() clusterexport() # Load custom functions clusterexport(cl = cl, c("b_ols", "gen_data", "one_sim", "ols", "to_matrix")) clusterexport() clusterexport(cl, "cars") clusterexport(cl, ls()) ols_sim_par() replicate() parlapply() parlapply() cl fun lapply() ols_sim_par
23 # Define the function ols_sim_par <- function(n_sims, sample_size, seed = 12345) { } # Require the parallel package require(parallel) # Set the seed set.seed(seed) # Run one_sim n_sims times; convert results to data.frame sim_df <- parlapply( cl = cl, X = rep(x = sample_size, times = n_sims), fun = one_sim ) %>% bind_rows() # Return sim_df return(sim_df) # Send it to the cluster clusterexport(cl, "ols_sim_par") # Run ols_sim_par for sample size of 10 start5 <- proc.time() sim10_par <- ols_sim_par(n_sims = 1e4, sample_size = 10) stop5 <- proc.time() # Run ols_sim_par for sample size of 100 start6 <- proc.time() sim100_par <- ols_sim_par(n_sims = 1e4, sample_size = 100) stop6 <- proc.time() # Run ols_sim for sample size of 10 start7 <- proc.time() sim10 <- ols_sim(n_sims = 1e4, sample_size = 10) stop7 <- proc.time() # Run ols_sim for sample size of 100 start8 <- proc.time() sim100 <- ols_sim(n_sims = 1e4, sample_size = 100) stop8 <- proc.time() # Stop the cluster stopcluster(cl) stop5 - start5
24 ## user system elapsed ## stop6 - start6 ## user system elapsed ## stop7 - start7 ## user system elapsed ## stop8 - start8 ## user system elapsed ## ols_sim_par()
Contents 1 Admin 2 Testing hypotheses tests 4 Simulation 5 Parallelization Admin
 magrittr t F F dplyr lfe readr magrittr parallel parallel auto.csv y x y x y x y x # Setup ---- # Settings options(stringsasfactors = F) # Packages library(dplyr) library(lfe) library(magrittr) library(readr)
magrittr t F F dplyr lfe readr magrittr parallel parallel auto.csv y x y x y x y x # Setup ---- # Settings options(stringsasfactors = F) # Packages library(dplyr) library(lfe) library(magrittr) library(readr)
Section 4: FWL and model fit
 Section 4: FWL and model fit Ed Rubin Contents 1 Admin 1 1.1 What you will need........................................ 1 1.2 Last week............................................. 2 1.3 This week.............................................
Section 4: FWL and model fit Ed Rubin Contents 1 Admin 1 1.1 What you will need........................................ 1 1.2 Last week............................................. 2 1.3 This week.............................................
An Introduction to R. Ed D. J. Berry 9th January 2017
 An Introduction to R Ed D. J. Berry 9th January 2017 Overview Why now? Why R? General tips Recommended packages Recommended resources 2/48 Why now? Efficiency Pointandclick software just isn't time efficient
An Introduction to R Ed D. J. Berry 9th January 2017 Overview Why now? Why R? General tips Recommended packages Recommended resources 2/48 Why now? Efficiency Pointandclick software just isn't time efficient
First steps in Parallel Computing
 MilanoR 4th meeting October 24, 2013 First steps in Parallel Computing Anna Longari anna.longari@quantide.com Outline Parallel Computing Implicit Parallelism Explicit Parallelism Example on Amazon Servers
MilanoR 4th meeting October 24, 2013 First steps in Parallel Computing Anna Longari anna.longari@quantide.com Outline Parallel Computing Implicit Parallelism Explicit Parallelism Example on Amazon Servers
Standard Errors in OLS Luke Sonnet
 Standard Errors in OLS Luke Sonnet Contents Variance-Covariance of ˆβ 1 Standard Estimation (Spherical Errors) 2 Robust Estimation (Heteroskedasticity Constistent Errors) 4 Cluster Robust Estimation 7
Standard Errors in OLS Luke Sonnet Contents Variance-Covariance of ˆβ 1 Standard Estimation (Spherical Errors) 2 Robust Estimation (Heteroskedasticity Constistent Errors) 4 Cluster Robust Estimation 7
CSSS 510: Lab 2. Introduction to Maximum Likelihood Estimation
 CSSS 510: Lab 2 Introduction to Maximum Likelihood Estimation 2018-10-12 0. Agenda 1. Housekeeping: simcf, tile 2. Questions about Homework 1 or lecture 3. Simulating heteroskedastic normal data 4. Fitting
CSSS 510: Lab 2 Introduction to Maximum Likelihood Estimation 2018-10-12 0. Agenda 1. Housekeeping: simcf, tile 2. Questions about Homework 1 or lecture 3. Simulating heteroskedastic normal data 4. Fitting
Package pbapply. R topics documented: January 10, Type Package Title Adding Progress Bar to '*apply' Functions Version Date
 Type Package Title Adding Progress Bar to '*apply' Functions Version 1.3-4 Date 2018-01-09 Package pbapply January 10, 2018 Author Peter Solymos [aut, cre], Zygmunt Zawadzki [aut] Maintainer Peter Solymos
Type Package Title Adding Progress Bar to '*apply' Functions Version 1.3-4 Date 2018-01-09 Package pbapply January 10, 2018 Author Peter Solymos [aut, cre], Zygmunt Zawadzki [aut] Maintainer Peter Solymos
Subsetting, dplyr, magrittr Author: Lloyd Low; add:
 Subsetting, dplyr, magrittr Author: Lloyd Low; Email add: wai.low@adelaide.edu.au Introduction So you have got a table with data that might be a mixed of categorical, integer, numeric, etc variables? And
Subsetting, dplyr, magrittr Author: Lloyd Low; Email add: wai.low@adelaide.edu.au Introduction So you have got a table with data that might be a mixed of categorical, integer, numeric, etc variables? And
Problem Set #8. Econ 103
 Problem Set #8 Econ 103 Part I Problems from the Textbook No problems from the textbook on this assignment. Part II Additional Problems 1. For this question assume that we have a random sample from a normal
Problem Set #8 Econ 103 Part I Problems from the Textbook No problems from the textbook on this assignment. Part II Additional Problems 1. For this question assume that we have a random sample from a normal
1 Lab 1. Graphics and Checking Residuals
 R is an object oriented language. We will use R for statistical analysis in FIN 504/ORF 504. To download R, go to CRAN (the Comprehensive R Archive Network) at http://cran.r-project.org Versions for Windows
R is an object oriented language. We will use R for statistical analysis in FIN 504/ORF 504. To download R, go to CRAN (the Comprehensive R Archive Network) at http://cran.r-project.org Versions for Windows
Exercise 2.23 Villanova MAT 8406 September 7, 2015
 Exercise 2.23 Villanova MAT 8406 September 7, 2015 Step 1: Understand the Question Consider the simple linear regression model y = 50 + 10x + ε where ε is NID(0, 16). Suppose that n = 20 pairs of observations
Exercise 2.23 Villanova MAT 8406 September 7, 2015 Step 1: Understand the Question Consider the simple linear regression model y = 50 + 10x + ε where ε is NID(0, 16). Suppose that n = 20 pairs of observations
EXST 7014, Lab 1: Review of R Programming Basics and Simple Linear Regression
 EXST 7014, Lab 1: Review of R Programming Basics and Simple Linear Regression OBJECTIVES 1. Prepare a scatter plot of the dependent variable on the independent variable 2. Do a simple linear regression
EXST 7014, Lab 1: Review of R Programming Basics and Simple Linear Regression OBJECTIVES 1. Prepare a scatter plot of the dependent variable on the independent variable 2. Do a simple linear regression
Dplyr Introduction Matthew Flickinger July 12, 2017
 Dplyr Introduction Matthew Flickinger July 12, 2017 Introduction to Dplyr This document gives an overview of many of the features of the dplyr library include in the tidyverse of related R pacakges. First
Dplyr Introduction Matthew Flickinger July 12, 2017 Introduction to Dplyr This document gives an overview of many of the features of the dplyr library include in the tidyverse of related R pacakges. First
INSTRUCTOR: KLAUS MOELTNER
 SCRIPT MOD1S3: SMALL-SAMPLE PROPERTIES OF OLS ESTIMATOR INSTRUCTOR: KLAUS MOELTNER Set basic R-options upfront and load all required R packages: Set parameters for sampling. 1. Data Generation R> N
SCRIPT MOD1S3: SMALL-SAMPLE PROPERTIES OF OLS ESTIMATOR INSTRUCTOR: KLAUS MOELTNER Set basic R-options upfront and load all required R packages: Set parameters for sampling. 1. Data Generation R> N
Lab #13 - Resampling Methods Econ 224 October 23rd, 2018
 Lab #13 - Resampling Methods Econ 224 October 23rd, 2018 Introduction In this lab you will work through Section 5.3 of ISL and record your code and results in an RMarkdown document. I have added section
Lab #13 - Resampling Methods Econ 224 October 23rd, 2018 Introduction In this lab you will work through Section 5.3 of ISL and record your code and results in an RMarkdown document. I have added section
Calibration of Quinine Fluorescence Emission Vignette for the Data Set flu of the R package hyperspec
 Calibration of Quinine Fluorescence Emission Vignette for the Data Set flu of the R package hyperspec Claudia Beleites CENMAT and DI3, University of Trieste Spectroscopy Imaging,
Calibration of Quinine Fluorescence Emission Vignette for the Data Set flu of the R package hyperspec Claudia Beleites CENMAT and DI3, University of Trieste Spectroscopy Imaging,
Package dotwhisker. R topics documented: June 28, Type Package
 Type Package Package dotwhisker June 28, 2017 Title Dot-and-Whisker Plots of Regression Results Version 0.3.0 Date 2017-06-28 Maintainer Yue Hu Quick and easy dot-and-whisker plots
Type Package Package dotwhisker June 28, 2017 Title Dot-and-Whisker Plots of Regression Results Version 0.3.0 Date 2017-06-28 Maintainer Yue Hu Quick and easy dot-and-whisker plots
R Workshop Guide. 1 Some Programming Basics. 1.1 Writing and executing code in R
 R Workshop Guide This guide reviews the examples we will cover in today s workshop. It should be a helpful introduction to R, but for more details, you can access a more extensive user guide for R on the
R Workshop Guide This guide reviews the examples we will cover in today s workshop. It should be a helpful introduction to R, but for more details, you can access a more extensive user guide for R on the
Package snow. R topics documented: February 20, 2015
 Package snow February 20, 2015 Title Simple Network of Workstations Version 0.3-13 Author Luke Tierney, A. J. Rossini, Na Li, H. Sevcikova Support for simple parallel computing in R. Maintainer Luke Tierney
Package snow February 20, 2015 Title Simple Network of Workstations Version 0.3-13 Author Luke Tierney, A. J. Rossini, Na Li, H. Sevcikova Support for simple parallel computing in R. Maintainer Luke Tierney
Compare Linear Regression Lines for the HP-67
 Compare Linear Regression Lines for the HP-67 by Namir Shammas This article presents an HP-67 program that calculates the linear regression statistics for two data sets and then compares their slopes and
Compare Linear Regression Lines for the HP-67 by Namir Shammas This article presents an HP-67 program that calculates the linear regression statistics for two data sets and then compares their slopes and
The grplasso Package
 The grplasso Package June 27, 2007 Type Package Title Fitting user specified models with Group Lasso penalty Version 0.2-1 Date 2007-06-27 Author Lukas Meier Maintainer Lukas Meier
The grplasso Package June 27, 2007 Type Package Title Fitting user specified models with Group Lasso penalty Version 0.2-1 Date 2007-06-27 Author Lukas Meier Maintainer Lukas Meier
Performing Cluster Bootstrapped Regressions in R
 Performing Cluster Bootstrapped Regressions in R Francis L. Huang / October 6, 2016 Supplementary material for: Using Cluster Bootstrapping to Analyze Nested Data with a Few Clusters in Educational and
Performing Cluster Bootstrapped Regressions in R Francis L. Huang / October 6, 2016 Supplementary material for: Using Cluster Bootstrapping to Analyze Nested Data with a Few Clusters in Educational and
Simple Parallel Statistical Computing in R
 Simple Parallel Statistical Computing in R Luke Tierney Department of Statistics & Actuarial Science University of Iowa December 7, 2007 Luke Tierney (U. of Iowa) Simple Parallel Statistical Computing
Simple Parallel Statistical Computing in R Luke Tierney Department of Statistics & Actuarial Science University of Iowa December 7, 2007 Luke Tierney (U. of Iowa) Simple Parallel Statistical Computing
Package dat. January 20, 2018
 Package dat Type Package Title Tools for Data Manipulation Version 0.4.0 January 20, 2018 BugReports https://github.com/wahani/dat/issues An implementation of common higher order functions with syntactic
Package dat Type Package Title Tools for Data Manipulation Version 0.4.0 January 20, 2018 BugReports https://github.com/wahani/dat/issues An implementation of common higher order functions with syntactic
Knitr. Introduction to R for Public Health Researchers
 Knitr Introduction to R for Public Health Researchers Introduction Exploratory Analysis Plots of bike length Multiple Facets Means by type Linear Models Grabbing coefficients Broom package Testing Nested
Knitr Introduction to R for Public Health Researchers Introduction Exploratory Analysis Plots of bike length Multiple Facets Means by type Linear Models Grabbing coefficients Broom package Testing Nested
Data Manipulation. Module 5
 Data Manipulation http://datascience.tntlab.org Module 5 Today s Agenda A couple of base-r notes Advanced data typing Relabeling text In depth with dplyr (part of tidyverse) tbl class dplyr grammar Grouping
Data Manipulation http://datascience.tntlab.org Module 5 Today s Agenda A couple of base-r notes Advanced data typing Relabeling text In depth with dplyr (part of tidyverse) tbl class dplyr grammar Grouping
Financial Econometrics Practical
 Financial Econometrics Practical Practical 3: Plotting in R NF Katzke Table of Contents 1 Introduction 1 1.0.1 Install ggplot2................................................. 2 1.1 Get data Tidy.....................................................
Financial Econometrics Practical Practical 3: Plotting in R NF Katzke Table of Contents 1 Introduction 1 1.0.1 Install ggplot2................................................. 2 1.1 Get data Tidy.....................................................
Biostatistics 615/815 Lecture 22: Matrix with C++
 Biostatistics 615/815 Lecture 22: Matrix with C++ Hyun Min Kang December 1st, 2011 Hyun Min Kang Biostatistics 615/815 - Lecture 22 December 1st, 2011 1 / 33 Recap - A case for simple linear regression
Biostatistics 615/815 Lecture 22: Matrix with C++ Hyun Min Kang December 1st, 2011 Hyun Min Kang Biostatistics 615/815 - Lecture 22 December 1st, 2011 1 / 33 Recap - A case for simple linear regression
2 Calculation of the within-class covariance matrix
 1 Topic Parallel programming in R. Using the «parallel» and «doparallel» packages. Personal computers become more and more efficient. They are mostly equipped with multi-core processors. At the same time,
1 Topic Parallel programming in R. Using the «parallel» and «doparallel» packages. Personal computers become more and more efficient. They are mostly equipped with multi-core processors. At the same time,
Week 1 R Warm-Ups for Finance
 Week 1 R Warm-Ups for Finance Copyright 2016, William G. Foote. All rights reserved. Copyright 2016, William G. Foote. All rights reserved. Week 1 R Warm-Ups for Finance 1 / 97 Imagine this... You work
Week 1 R Warm-Ups for Finance Copyright 2016, William G. Foote. All rights reserved. Copyright 2016, William G. Foote. All rights reserved. Week 1 R Warm-Ups for Finance 1 / 97 Imagine this... You work
Package rcpphmmparclip
 Package rcpphmmparclip October 15, 2013 Type Package Title Rcpp functions for analysing PAR-CLIP using the HMM Version 0.1.0 Date 2013-10-15 Author Jonghyun Yun Maintainer Jonghyun Yun
Package rcpphmmparclip October 15, 2013 Type Package Title Rcpp functions for analysing PAR-CLIP using the HMM Version 0.1.0 Date 2013-10-15 Author Jonghyun Yun Maintainer Jonghyun Yun
Simulating power in practice
 Simulating power in practice Author: Nicholas G Reich This material is part of the statsteachr project Made available under the Creative Commons Attribution-ShareAlike 3.0 Unported License: http://creativecommons.org/licenses/by-sa/3.0/deed.en
Simulating power in practice Author: Nicholas G Reich This material is part of the statsteachr project Made available under the Creative Commons Attribution-ShareAlike 3.0 Unported License: http://creativecommons.org/licenses/by-sa/3.0/deed.en
Parallel programming in R
 Parallel programming in R Bjørn-Helge Mevik Research Infrastructure Services Group, USIT, UiO RIS Course Week, spring 2014 Bjørn-Helge Mevik (RIS) Parallel programming in R RIS Course Week 1 / 13 Introduction
Parallel programming in R Bjørn-Helge Mevik Research Infrastructure Services Group, USIT, UiO RIS Course Week, spring 2014 Bjørn-Helge Mevik (RIS) Parallel programming in R RIS Course Week 1 / 13 Introduction
Model Selection and Inference
 Model Selection and Inference Merlise Clyde January 29, 2017 Last Class Model for brain weight as a function of body weight In the model with both response and predictor log transformed, are dinosaurs
Model Selection and Inference Merlise Clyde January 29, 2017 Last Class Model for brain weight as a function of body weight In the model with both response and predictor log transformed, are dinosaurs
Introduction to R and the tidyverse. Paolo Crosetto
 Introduction to R and the tidyverse Paolo Crosetto Lecture 1: plotting Before we start: Rstudio Interactive console Object explorer Script window Plot window Before we start: R concatenate: c() assign:
Introduction to R and the tidyverse Paolo Crosetto Lecture 1: plotting Before we start: Rstudio Interactive console Object explorer Script window Plot window Before we start: R concatenate: c() assign:
Practice for Learning R and Learning Latex
 Practice for Learning R and Learning Latex Jennifer Pan August, 2011 Latex Environments A) Try to create the following equations: 1. 5+6 α = β2 2. P r( 1.96 Z 1.96) = 0.95 ( ) ( ) sy 1 r 2 3. ˆβx = r xy
Practice for Learning R and Learning Latex Jennifer Pan August, 2011 Latex Environments A) Try to create the following equations: 1. 5+6 α = β2 2. P r( 1.96 Z 1.96) = 0.95 ( ) ( ) sy 1 r 2 3. ˆβx = r xy
Assignment 5.5. Nothing here to hand in
 Assignment 5.5 Nothing here to hand in Load the tidyverse before we start: library(tidyverse) ## Loading tidyverse: ggplot2 ## Loading tidyverse: tibble ## Loading tidyverse: tidyr ## Loading tidyverse:
Assignment 5.5 Nothing here to hand in Load the tidyverse before we start: library(tidyverse) ## Loading tidyverse: ggplot2 ## Loading tidyverse: tibble ## Loading tidyverse: tidyr ## Loading tidyverse:
Package bayescl. April 14, 2017
 Package bayescl April 14, 2017 Version 0.0.1 Date 2017-04-10 Title Bayesian Inference on a GPU using OpenCL Author Rok Cesnovar, Erik Strumbelj Maintainer Rok Cesnovar Description
Package bayescl April 14, 2017 Version 0.0.1 Date 2017-04-10 Title Bayesian Inference on a GPU using OpenCL Author Rok Cesnovar, Erik Strumbelj Maintainer Rok Cesnovar Description
The Tidyverse BIOF 339 9/25/2018
 The Tidyverse BIOF 339 9/25/2018 What is the Tidyverse? The tidyverse is an opinionated collection of R packages designed for data science. All packages share an underlying design philosophy, grammar,
The Tidyverse BIOF 339 9/25/2018 What is the Tidyverse? The tidyverse is an opinionated collection of R packages designed for data science. All packages share an underlying design philosophy, grammar,
Module 25.1: nag lin reg Regression Analysis. Contents
 Correlation and Regression Analysis Module Contents Module 25.1: nag lin reg Regression Analysis nag lin reg contains procedures that perform a simple or multiple linear regression analysis. Contents Introduction...
Correlation and Regression Analysis Module Contents Module 25.1: nag lin reg Regression Analysis nag lin reg contains procedures that perform a simple or multiple linear regression analysis. Contents Introduction...
Reliability Coefficients
 Reliability Coefficients Introductory notes That data used for these computations is the pre-treatment scores of all subjects. There are three items in the SIAS (5, 9 and 11) that require reverse-scoring.
Reliability Coefficients Introductory notes That data used for these computations is the pre-treatment scores of all subjects. There are three items in the SIAS (5, 9 and 11) that require reverse-scoring.
Parallel Computing with R and How to Use it on High Performance Computing Cluster
 UNIVERSITY OF TEXAS AT SAN ANTONIO Parallel Computing with R and How to Use it on High Performance Computing Cluster Liang Jing Nov. 2010 1 1 ABSTRACT Methodological advances have led to much more computationally
UNIVERSITY OF TEXAS AT SAN ANTONIO Parallel Computing with R and How to Use it on High Performance Computing Cluster Liang Jing Nov. 2010 1 1 ABSTRACT Methodological advances have led to much more computationally
Journal of Avian Biology
 Journal of Avian Biology JAV-01710 Capilla-Lasheras, P. 2018. incr: a new R package to analyse incubation behaviour. J. Avian Biol. 2018: e01710 Supplementary material incr: a new R package to analyse
Journal of Avian Biology JAV-01710 Capilla-Lasheras, P. 2018. incr: a new R package to analyse incubation behaviour. J. Avian Biol. 2018: e01710 Supplementary material incr: a new R package to analyse
Informative Priors for Regularization in Bayesian Predictive Modeling
 Informative Priors for Regularization in Bayesian Predictive Modeling Kyle M. Lang Institute for Measurement, Methodology, Analysis & Policy Texas Tech University Lubbock, TX November 23, 2016 Outline
Informative Priors for Regularization in Bayesian Predictive Modeling Kyle M. Lang Institute for Measurement, Methodology, Analysis & Policy Texas Tech University Lubbock, TX November 23, 2016 Outline
1 Simple Linear Regression
 Math 158 Jo Hardin R code 1 Simple Linear Regression Consider a dataset from ISLR on credit scores. Because we don t know the sampling mechanism used to collect the data, we are unable to generalize the
Math 158 Jo Hardin R code 1 Simple Linear Regression Consider a dataset from ISLR on credit scores. Because we don t know the sampling mechanism used to collect the data, we are unable to generalize the
Speeding up R code using Rcpp and foreach packages.
 Speeding up R code using Rcpp and foreach packages. Pedro Albuquerque Universidade de Brasília February, 8th 2017 Speeding up R code using Rcpp and foreach packages. 1 1 Speeding up R. 2 foreach package.
Speeding up R code using Rcpp and foreach packages. Pedro Albuquerque Universidade de Brasília February, 8th 2017 Speeding up R code using Rcpp and foreach packages. 1 1 Speeding up R. 2 foreach package.
STAT 5200 Handout #25. R-Square & Design Matrix in Mixed Models
 STAT 5200 Handout #25 R-Square & Design Matrix in Mixed Models I. R-Square in Mixed Models (with Example from Handout #20): For mixed models, the concept of R 2 is a little complicated (and neither PROC
STAT 5200 Handout #25 R-Square & Design Matrix in Mixed Models I. R-Square in Mixed Models (with Example from Handout #20): For mixed models, the concept of R 2 is a little complicated (and neither PROC
OVERVIEW OF ESTIMATION FRAMEWORKS AND ESTIMATORS
 OVERVIEW OF ESTIMATION FRAMEWORKS AND ESTIMATORS Set basic R-options upfront and load all required R packages: > options(prompt = " ", digits = 4) setwd('c:/klaus/aaec5126/test/')#r Sweaves to the default
OVERVIEW OF ESTIMATION FRAMEWORKS AND ESTIMATORS Set basic R-options upfront and load all required R packages: > options(prompt = " ", digits = 4) setwd('c:/klaus/aaec5126/test/')#r Sweaves to the default
Risk Management Using R, SoSe 2013
 1. Problem (vectors and factors) a) Create a vector containing the numbers 1 to 10. In this vector, replace all numbers greater than 4 with 5. b) Create a sequence of length 5 starting at 0 with an increment
1. Problem (vectors and factors) a) Create a vector containing the numbers 1 to 10. In this vector, replace all numbers greater than 4 with 5. b) Create a sequence of length 5 starting at 0 with an increment
EXPLORATORY DATA ANALYSIS. Introducing the data
 EXPLORATORY DATA ANALYSIS Introducing the data Email data set > email # A tibble: 3,921 21 spam to_multiple from cc sent_email time image 1 not-spam 0 1 0 0
EXPLORATORY DATA ANALYSIS Introducing the data Email data set > email # A tibble: 3,921 21 spam to_multiple from cc sent_email time image 1 not-spam 0 1 0 0
Regression on the trees data with R
 > trees Girth Height Volume 1 8.3 70 10.3 2 8.6 65 10.3 3 8.8 63 10.2 4 10.5 72 16.4 5 10.7 81 18.8 6 10.8 83 19.7 7 11.0 66 15.6 8 11.0 75 18.2 9 11.1 80 22.6 10 11.2 75 19.9 11 11.3 79 24.2 12 11.4 76
> trees Girth Height Volume 1 8.3 70 10.3 2 8.6 65 10.3 3 8.8 63 10.2 4 10.5 72 16.4 5 10.7 81 18.8 6 10.8 83 19.7 7 11.0 66 15.6 8 11.0 75 18.2 9 11.1 80 22.6 10 11.2 75 19.9 11 11.3 79 24.2 12 11.4 76
Model selection. Peter Hoff STAT 423. Applied Regression and Analysis of Variance. University of Washington /53
 /53 Model selection Peter Hoff STAT 423 Applied Regression and Analysis of Variance University of Washington Diabetes example: y = diabetes progression x 1 = age x 2 = sex. dim(x) ## [1] 442 64 colnames(x)
/53 Model selection Peter Hoff STAT 423 Applied Regression and Analysis of Variance University of Washington Diabetes example: y = diabetes progression x 1 = age x 2 = sex. dim(x) ## [1] 442 64 colnames(x)
INTRODUCTION TO DATA. Welcome to the course!
 INTRODUCTION TO DATA Welcome to the course! High School and Beyond id gender race socst 70 male white 57 121 female white 61 86 male white 31 137 female white 61 Loading data > # Load package > library(openintro)
INTRODUCTION TO DATA Welcome to the course! High School and Beyond id gender race socst 70 male white 57 121 female white 61 86 male white 31 137 female white 61 Loading data > # Load package > library(openintro)
Package ChIPseeker. July 12, 2018
 Type Package Package ChIPseeker July 12, 2018 Title ChIPseeker for ChIP peak Annotation, Comparison, and Visualization Version 1.16.0 Encoding UTF-8 Maintainer Guangchuang Yu
Type Package Package ChIPseeker July 12, 2018 Title ChIPseeker for ChIP peak Annotation, Comparison, and Visualization Version 1.16.0 Encoding UTF-8 Maintainer Guangchuang Yu
S CHAPTER return.data S CHAPTER.Data S CHAPTER
 1 S CHAPTER return.data S CHAPTER.Data MySwork S CHAPTER.Data 2 S e > return ; return + # 3 setenv S_CLEDITOR emacs 4 > 4 + 5 / 3 ## addition & divison [1] 5.666667 > (4 + 5) / 3 ## using parentheses [1]
1 S CHAPTER return.data S CHAPTER.Data MySwork S CHAPTER.Data 2 S e > return ; return + # 3 setenv S_CLEDITOR emacs 4 > 4 + 5 / 3 ## addition & divison [1] 5.666667 > (4 + 5) / 3 ## using parentheses [1]
References R's single biggest strenght is it online community. There are tons of free tutorials on R.
 Introduction to R Syllabus Instructor Grant Cavanaugh Department of Agricultural Economics University of Kentucky E-mail: gcavanugh@uky.edu Course description Introduction to R is a short course intended
Introduction to R Syllabus Instructor Grant Cavanaugh Department of Agricultural Economics University of Kentucky E-mail: gcavanugh@uky.edu Course description Introduction to R is a short course intended
Review Stats after TOS Change
 Review Stats after TOS Change library(knitr) library(ggplot2) library(plotrix) # for pie3d suppressmessages(library(dplyr)) Description of dataset #sum(dfremovals$perc == 100) #dfremovals$perc == 100 df
Review Stats after TOS Change library(knitr) library(ggplot2) library(plotrix) # for pie3d suppressmessages(library(dplyr)) Description of dataset #sum(dfremovals$perc == 100) #dfremovals$perc == 100 df
Statistical Analysis of Series of N-of-1 Trials Using R. Artur Araujo
 Statistical Analysis of Series of N-of-1 Trials Using R Artur Araujo March 2016 Acknowledgements I would like to thank Boehringer Ingelheim GmbH for having paid my tuition fees at the University of Sheffield
Statistical Analysis of Series of N-of-1 Trials Using R Artur Araujo March 2016 Acknowledgements I would like to thank Boehringer Ingelheim GmbH for having paid my tuition fees at the University of Sheffield
Practice in R. 1 Sivan s practice. 2 Hetroskadasticity. January 28, (pdf version)
 Practice in R January 28, 2010 (pdf version) 1 Sivan s practice Her practice file should be (here), or check the web for a more useful pointer. 2 Hetroskadasticity ˆ Let s make some hetroskadastic data:
Practice in R January 28, 2010 (pdf version) 1 Sivan s practice Her practice file should be (here), or check the web for a more useful pointer. 2 Hetroskadasticity ˆ Let s make some hetroskadastic data:
Getting started with simulating data in R: some helpful functions and how to use them Ariel Muldoon August 28, 2018
 Getting started with simulating data in R: some helpful functions and how to use them Ariel Muldoon August 28, 2018 Contents Overview 2 Generating random numbers 2 rnorm() to generate random numbers from
Getting started with simulating data in R: some helpful functions and how to use them Ariel Muldoon August 28, 2018 Contents Overview 2 Generating random numbers 2 rnorm() to generate random numbers from
Section 2.3: Simple Linear Regression: Predictions and Inference
 Section 2.3: Simple Linear Regression: Predictions and Inference Jared S. Murray The University of Texas at Austin McCombs School of Business Suggested reading: OpenIntro Statistics, Chapter 7.4 1 Simple
Section 2.3: Simple Linear Regression: Predictions and Inference Jared S. Murray The University of Texas at Austin McCombs School of Business Suggested reading: OpenIntro Statistics, Chapter 7.4 1 Simple
Package mvprobit. November 2, 2015
 Version 0.1-8 Date 2015-11-02 Title Multivariate Probit Models Package mvprobit November 2, 2015 Author Arne Henningsen Maintainer Arne Henningsen
Version 0.1-8 Date 2015-11-02 Title Multivariate Probit Models Package mvprobit November 2, 2015 Author Arne Henningsen Maintainer Arne Henningsen
Introduction to Functions. Biostatistics
 Introduction to Functions Biostatistics 140.776 Functions The development of a functions in R represents the next level of R programming, beyond writing code at the console or in a script. 1. Code 2. Functions
Introduction to Functions Biostatistics 140.776 Functions The development of a functions in R represents the next level of R programming, beyond writing code at the console or in a script. 1. Code 2. Functions
Package postal. July 27, 2018
 Type Package Title United States Postal Service API Interface Version 0.1.0 Package postal July 27, 2018 Author Amanda Dobbyn Maintainer Amanda Dobbyn
Type Package Title United States Postal Service API Interface Version 0.1.0 Package postal July 27, 2018 Author Amanda Dobbyn Maintainer Amanda Dobbyn
Supplementary tutorial for A Practical Guide and Power Analysis for GLMMs: Detecting Among Treatment Variation in Random Effects using R
 Supplementary tutorial for A Practical Guide and Power Analysis for GLMMs: Detecting Among Treatment Variation in Random Effects using R Kain, Morgan, Ben M. Bolker, and Michael W. McCoy Introduction The
Supplementary tutorial for A Practical Guide and Power Analysis for GLMMs: Detecting Among Treatment Variation in Random Effects using R Kain, Morgan, Ben M. Bolker, and Michael W. McCoy Introduction The
More advanced use of mgcv. Simon Wood Mathematical Sciences, University of Bath, U.K.
 More advanced use of mgcv Simon Wood Mathematical Sciences, University of Bath, U.K. Fine control of smoothness: gamma Suppose that we fit a model but a component is too wiggly. For GCV/AIC we can increase
More advanced use of mgcv Simon Wood Mathematical Sciences, University of Bath, U.K. Fine control of smoothness: gamma Suppose that we fit a model but a component is too wiggly. For GCV/AIC we can increase
Compare Linear Regression Lines for the HP-41C
 Compare Linear Regression Lines for the HP-41C by Namir Shammas This article presents an HP-41C program that calculates the linear regression statistics for two data sets and then compares their slopes
Compare Linear Regression Lines for the HP-41C by Namir Shammas This article presents an HP-41C program that calculates the linear regression statistics for two data sets and then compares their slopes
Package infer. July 11, Type Package Title Tidy Statistical Inference Version 0.3.0
 Type Package Title Tidy Statistical Inference Version 0.3.0 Package infer July 11, 2018 The objective of this package is to perform inference using an epressive statistical grammar that coheres with the
Type Package Title Tidy Statistical Inference Version 0.3.0 Package infer July 11, 2018 The objective of this package is to perform inference using an epressive statistical grammar that coheres with the
STENO Introductory R-Workshop: Loading a Data Set Tommi Suvitaival, Steno Diabetes Center June 11, 2015
 STENO Introductory R-Workshop: Loading a Data Set Tommi Suvitaival, tsvv@steno.dk, Steno Diabetes Center June 11, 2015 Contents 1 Introduction 1 2 Recap: Variables 2 3 Data Containers 2 3.1 Vectors................................................
STENO Introductory R-Workshop: Loading a Data Set Tommi Suvitaival, tsvv@steno.dk, Steno Diabetes Center June 11, 2015 Contents 1 Introduction 1 2 Recap: Variables 2 3 Data Containers 2 3.1 Vectors................................................
The "R" Statistics library: Research Applications
 Edith Cowan University Research Online ECU Research Week Conferences, Symposia and Campus Events 2012 The "R" Statistics library: Research Applications David Allen Edith Cowan University Abhay Singh Edith
Edith Cowan University Research Online ECU Research Week Conferences, Symposia and Campus Events 2012 The "R" Statistics library: Research Applications David Allen Edith Cowan University Abhay Singh Edith
Facets and Continuous graphs
 Facets and Continuous graphs One way to add additional variables is with aesthetics. Another way, particularly useful for categorical variables, is to split your plot into facets, subplots that each display
Facets and Continuous graphs One way to add additional variables is with aesthetics. Another way, particularly useful for categorical variables, is to split your plot into facets, subplots that each display
Among those 14 potential explanatory variables,non-dummy variables are:
 Among those 14 potential explanatory variables,non-dummy variables are: Size: 2nd column in the dataset Land: 14th column in the dataset Bed.Rooms: 5th column in the dataset Fireplace: 7th column in the
Among those 14 potential explanatory variables,non-dummy variables are: Size: 2nd column in the dataset Land: 14th column in the dataset Bed.Rooms: 5th column in the dataset Fireplace: 7th column in the
Package visdat. February 15, 2019
 Title Preliminary Visualisation of Data Version 0.5.3 Package visdat February 15, 2019 Create preliminary eploratory data visualisations of an entire dataset to identify problems or unepected features
Title Preliminary Visualisation of Data Version 0.5.3 Package visdat February 15, 2019 Create preliminary eploratory data visualisations of an entire dataset to identify problems or unepected features
file:///d:/r/stateofther/bigdata/slides/bigdatapresentation.html#1 1 sur 44 06/07/2018 à 16:56
 1 sur 44 06/07/2018 à 16:56 2 sur 44 06/07/2018 à 16:56 Arabidopsis[1:5,1:10] ## L1 L2 L3 L4 L5 L6 L7 L8 L9 L10 ## M1 1 0 1 1 0 1 0 1 1 1 ## M2 1 0 1 1 0 1 1 1 1 1 ## M3 1 0 1 1 0 1 1 1 1 1 ## M4 0 0 0
1 sur 44 06/07/2018 à 16:56 2 sur 44 06/07/2018 à 16:56 Arabidopsis[1:5,1:10] ## L1 L2 L3 L4 L5 L6 L7 L8 L9 L10 ## M1 1 0 1 1 0 1 0 1 1 1 ## M2 1 0 1 1 0 1 1 1 1 1 ## M3 1 0 1 1 0 1 1 1 1 1 ## M4 0 0 0
Package tidystats. May 6, 2018
 Title Create a Tidy Statistics Output File Version 0.2 Package tidystats May 6, 2018 Produce a data file containing the output of statistical models and assist with a workflow aimed at writing scientific
Title Create a Tidy Statistics Output File Version 0.2 Package tidystats May 6, 2018 Produce a data file containing the output of statistical models and assist with a workflow aimed at writing scientific
Introduction to R for Beginners, Level II. Jeon Lee Bio-Informatics Core Facility (BICF), UTSW
 Introduction to R for Beginners, Level II Jeon Lee Bio-Informatics Core Facility (BICF), UTSW Basics of R Powerful programming language and environment for statistical computing Useful for very basic analysis
Introduction to R for Beginners, Level II Jeon Lee Bio-Informatics Core Facility (BICF), UTSW Basics of R Powerful programming language and environment for statistical computing Useful for very basic analysis
Lab #7 - More on Regression in R Econ 224 September 18th, 2018
 Lab #7 - More on Regression in R Econ 224 September 18th, 2018 Robust Standard Errors Your reading assignment from Chapter 3 of ISL briefly discussed two ways that the standard regression inference formulas
Lab #7 - More on Regression in R Econ 224 September 18th, 2018 Robust Standard Errors Your reading assignment from Chapter 3 of ISL briefly discussed two ways that the standard regression inference formulas
Handling Missing Values
 Handling Missing Values STAT 133 Gaston Sanchez Department of Statistics, UC Berkeley gastonsanchez.com github.com/gastonstat/stat133 Course web: gastonsanchez.com/stat133 Missing Values 2 Introduction
Handling Missing Values STAT 133 Gaston Sanchez Department of Statistics, UC Berkeley gastonsanchez.com github.com/gastonstat/stat133 Course web: gastonsanchez.com/stat133 Missing Values 2 Introduction
Data Input/Output. Introduction to R for Public Health Researchers
 Data Input/Output Introduction to R for Public Health Researchers Common new user mistakes we have seen 1. Working directory problems: trying to read files that R can t find RStudio can help, and so do
Data Input/Output Introduction to R for Public Health Researchers Common new user mistakes we have seen 1. Working directory problems: trying to read files that R can t find RStudio can help, and so do
Stat 3701 Lecture Notes: Parallel Computing in R Charles J. Geyer September 18, 2018
 Stat 3701 Lecture Notes: Parallel Computing in R Charles J. Geyer September 18, 2018 1 License This work is licensed under a Creative Commons Attribution-ShareAlike 4.0 International License (http: //creativecommons.org/licenses/by-sa/4.0/).
Stat 3701 Lecture Notes: Parallel Computing in R Charles J. Geyer September 18, 2018 1 License This work is licensed under a Creative Commons Attribution-ShareAlike 4.0 International License (http: //creativecommons.org/licenses/by-sa/4.0/).
Package sfc. August 29, 2016
 Type Package Title Substance Flow Computation Version 0.1.0 Package sfc August 29, 2016 Description Provides a function sfc() to compute the substance flow with the input files --- ``data'' and ``model''.
Type Package Title Substance Flow Computation Version 0.1.0 Package sfc August 29, 2016 Description Provides a function sfc() to compute the substance flow with the input files --- ``data'' and ``model''.
Data visualization with ggplot2
 Data visualization with ggplot2 Visualizing data in R with the ggplot2 package Authors: Mateusz Kuzak, Diana Marek, Hedi Peterson, Dmytro Fishman Disclaimer We will be using the functions in the ggplot2
Data visualization with ggplot2 Visualizing data in R with the ggplot2 package Authors: Mateusz Kuzak, Diana Marek, Hedi Peterson, Dmytro Fishman Disclaimer We will be using the functions in the ggplot2
Package crossword.r. January 19, 2018
 Date 2018-01-13 Type Package Title Generating s from Word Lists Version 0.3.5 Author Peter Meissner Package crossword.r January 19, 2018 Maintainer Peter Meissner Generate crosswords
Date 2018-01-13 Type Package Title Generating s from Word Lists Version 0.3.5 Author Peter Meissner Package crossword.r January 19, 2018 Maintainer Peter Meissner Generate crosswords
Data Input/Output. Introduction to R for Public Health Researchers
 Data Input/Output Introduction to R for Public Health Researchers Common new user mistakes we have seen 1. Working directory problems: trying to read files that R "can't find" RStudio can help, and so
Data Input/Output Introduction to R for Public Health Researchers Common new user mistakes we have seen 1. Working directory problems: trying to read files that R "can't find" RStudio can help, and so
Package msgps. February 20, 2015
 Type Package Package msgps February 20, 2015 Title Degrees of freedom of elastic net, adaptive lasso and generalized elastic net Version 1.3 Date 2012-5-17 Author Kei Hirose Maintainer Kei Hirose
Type Package Package msgps February 20, 2015 Title Degrees of freedom of elastic net, adaptive lasso and generalized elastic net Version 1.3 Date 2012-5-17 Author Kei Hirose Maintainer Kei Hirose
Data types and structures
 An introduc+on to Data types and structures Noémie Becker & Benedikt Holtmann Winter Semester 16/17 Course outline Day 3 Review GeFng started with R Crea+ng Objects Data types in R Data structures in R
An introduc+on to Data types and structures Noémie Becker & Benedikt Holtmann Winter Semester 16/17 Course outline Day 3 Review GeFng started with R Crea+ng Objects Data types in R Data structures in R
Introduction to R. Introduction to Econometrics W
 Introduction to R Introduction to Econometrics W3412 Begin Download R from the Comprehensive R Archive Network (CRAN) by choosing a location close to you. Students are also recommended to download RStudio,
Introduction to R Introduction to Econometrics W3412 Begin Download R from the Comprehensive R Archive Network (CRAN) by choosing a location close to you. Students are also recommended to download RStudio,
Session 3 Nick Hathaway;
 Session 3 Nick Hathaway; nicholas.hathaway@umassmed.edu Contents Manipulating Data frames and matrices 1 Converting to long vs wide formats.................................... 2 Manipulating data in table........................................
Session 3 Nick Hathaway; nicholas.hathaway@umassmed.edu Contents Manipulating Data frames and matrices 1 Converting to long vs wide formats.................................... 2 Manipulating data in table........................................
R. Muralikrishnan Max Planck Institute for Empirical Aesthetics Frankfurt. 08 June 2017
 R R. Muralikrishnan Max Planck Institute for Empirical Aesthetics Frankfurt 08 June 2017 Introduction What is R?! R is a programming language for statistical computing and graphics R is free and open-source
R R. Muralikrishnan Max Planck Institute for Empirical Aesthetics Frankfurt 08 June 2017 Introduction What is R?! R is a programming language for statistical computing and graphics R is free and open-source
S Basics. Statistics 135. Autumn Copyright c 2005 by Mark E. Irwin
 S Basics Statistics 135 Autumn 2005 Copyright c 2005 by Mark E. Irwin S Basics When discussing the S environment, I will, or at least try to, make the following distinctions. S will be used when what is
S Basics Statistics 135 Autumn 2005 Copyright c 2005 by Mark E. Irwin S Basics When discussing the S environment, I will, or at least try to, make the following distinctions. S will be used when what is
An Introduction to Model-Fitting with the R package glmm
 An Introduction to Model-Fitting with the R package glmm Christina Knudson November 7, 2017 Contents 1 Introduction 2 2 Formatting the Data 2 3 Fitting the Model 4 4 Reading the Model Summary 6 5 Isolating
An Introduction to Model-Fitting with the R package glmm Christina Knudson November 7, 2017 Contents 1 Introduction 2 2 Formatting the Data 2 3 Fitting the Model 4 4 Reading the Model Summary 6 5 Isolating
Package ruv. March 12, 2018
 Package ruv March 12, 2018 Title Detect and Remove Unwanted Variation using Negative Controls Implements the 'RUV' (Remove Unwanted Variation) algorithms. These algorithms attempt to adjust for systematic
Package ruv March 12, 2018 Title Detect and Remove Unwanted Variation using Negative Controls Implements the 'RUV' (Remove Unwanted Variation) algorithms. These algorithms attempt to adjust for systematic
MACAU User Manual. Xiang Zhou. March 15, 2017
 MACAU User Manual Xiang Zhou March 15, 2017 Contents 1 Introduction 2 1.1 What is MACAU...................................... 2 1.2 How to Cite MACAU................................... 2 1.3 The Model.........................................
MACAU User Manual Xiang Zhou March 15, 2017 Contents 1 Introduction 2 1.1 What is MACAU...................................... 2 1.2 How to Cite MACAU................................... 2 1.3 The Model.........................................
Package descriptr. March 20, 2018
 Type Package Package descriptr March 20, 2018 Title Generate Descriptive Statistics & Eplore Statistical Distributions Version 0.4.1 Generate descriptive statistics such as measures of location, dispersion,
Type Package Package descriptr March 20, 2018 Title Generate Descriptive Statistics & Eplore Statistical Distributions Version 0.4.1 Generate descriptive statistics such as measures of location, dispersion,
Introduction to the R Statistical Computing Environment R Programming: Exercises
 Introduction to the R Statistical Computing Environment R Programming: Exercises John Fox (McMaster University) ICPSR 2014 1. A straightforward problem: Write an R function for linear least-squares regression.
Introduction to the R Statistical Computing Environment R Programming: Exercises John Fox (McMaster University) ICPSR 2014 1. A straightforward problem: Write an R function for linear least-squares regression.
Old Faithful Chris Parrish
 Old Faithful Chris Parrish 17-4-27 Contents Old Faithful eruptions 1 data.................................................. 1 duration................................................ 1 waiting time..............................................
Old Faithful Chris Parrish 17-4-27 Contents Old Faithful eruptions 1 data.................................................. 1 duration................................................ 1 waiting time..............................................
Introduction to R 21/11/2016
 Introduction to R 21/11/2016 C3BI Vincent Guillemot & Anne Biton R: presentation and installation Where? https://cran.r-project.org/ How to install and use it? Follow the steps: you don t need advanced
Introduction to R 21/11/2016 C3BI Vincent Guillemot & Anne Biton R: presentation and installation Where? https://cran.r-project.org/ How to install and use it? Follow the steps: you don t need advanced
Package addhaz. September 26, 2018
 Package addhaz September 26, 2018 Title Binomial and Multinomial Additive Hazard Models Version 0.5 Description Functions to fit the binomial and multinomial additive hazard models and to estimate the
Package addhaz September 26, 2018 Title Binomial and Multinomial Additive Hazard Models Version 0.5 Description Functions to fit the binomial and multinomial additive hazard models and to estimate the
Package lucid. August 24, 2018
 Version 1.6 Package lucid August 24, 2018 Title Printing Floating Point Numbers in a Human-Friendly Format Description Print vectors (and data frames) of floating point numbers using a non-scientific format
Version 1.6 Package lucid August 24, 2018 Title Printing Floating Point Numbers in a Human-Friendly Format Description Print vectors (and data frames) of floating point numbers using a non-scientific format
Multinomial Logit Models with R
 Multinomial Logit Models with R > rm(list=ls()); options(scipen=999) # To avoid scientific notation > # install.packages("mlogit", dependencies=true) # Only need to do this once > library(mlogit) # Load
Multinomial Logit Models with R > rm(list=ls()); options(scipen=999) # To avoid scientific notation > # install.packages("mlogit", dependencies=true) # Only need to do this once > library(mlogit) # Load
