Preprocessing and Genotyping Illumina Arrays for Copy Number Analysis
|
|
|
- Doreen Gloria Cobb
- 6 years ago
- Views:
Transcription
1 Preprocessing and Genotyping Illumina Arrays for Copy Number Analysis Rob Scharpf September 18, 2012 Abstract This vignette illustrates the steps required prior to copy number analysis for Infinium platforms. Specifically, we require construction of a container to store processed forms of the raw data, preprocessing to normalize the arrays, and genotyping using the CRLMM algorithm. After completing these steps, users can refer to the copynumber vignette. 1 Set up The following codechunk declares a directory for saving ff files that will contain the normalized intensities and the genotype calls. > library(ff) > options(ffcaching="ffeachflush") > outdir <- paste("/local_data/r00/crlmm/", getrversion(), "/illumina_vignette", sep="") > ldpath(outdir) > dir.create(outdir, recursive=true, showwarnings=false) We will also store cached computations in the directory outdir. We declare that crlmm should process 150,000 markers at a time and/or 500 samples at a time (when possible) to reduce the memory footprint. As our example dataset in this vignette contains fewer than 500 samples, all samples will be processed simultaneously. > ocprobesets(150e3) > ocsamples(500) We begin by specifying the path containing the raw IDAT files for a set of samples from the Infinium 370k platform. > datadir <- "/thumper/ctsa/snpmicroarray/illumina/idats/370k" For Infinium platforms, an Illumina sample sheet containing information for reading the raw IDAT files is required. Please refer to the BeadStudio Genotyping guide, Appendix A, for additional information. The following code reads in the samplesheet for the IDAT files on our local server. > samplesheet = read.csv(file.path(datadir, "HumanHap370Duo_Sample_Map.csv"), header=true, as.is=true) For the purposes of this vignette, we indicate that we only wish to process a subset of the arrays. For our dataset, the file extensions are Grn.dat and Red.idat. We store the complete path to the filename without the file extension in the arraynames and check that all of the green and red IDAT files exists. > samplesheet <- samplesheet[-c(28:46,61:75,78:79), ] > arraynames <- file.path(datadir, unique(samplesheet[, "SentrixPosition"])) > all(file.exists(paste(arraynames, "_Grn.idat", sep=""))) 1
2 > all(file.exists(paste(arraynames, "_Red.idat", sep=""))) > arrayinfo <- list(barcode=null, position="sentrixposition") All supported platforms have a corresponding annotation package. The appropriate annotation package is specified by the platform identifier without the Crlmm postfix. > cdfname <- "human370v1c" Next, we construct a character vector that specifies the batch for each of the 43 arrays. Here, we have a small dataset and process the samples in a single batch. Processing the samples as a single batch is generally reasonable if the samples were processed at similar times (e.g., within a few weeks). > batch <- rep("1", nrow(samplesheet)) 2 Preprocessing and genotyping The raw intensities from the Infinium IDAT files are read and normalized using the function preprocessinf. The function preprocessinf returns a ff object containing the parameters for the mixture model used by the CRLMM genotyping algorithm. Following preprocessing, the genotypeinf genotypes the samples. The function genotype.illumina is a wrapper to the above functions and returns an object of class CNSet. > cnset <- genotype.illumina(samplesheet=samplesheet, arraynames=arraynames, arrayinfocolnames=arrayinfo, cdfname="human370v1c", batch=batch) Note, to fully remove the data associated with the cnset2 object, one should use the delete function in the ff package followed by the rm function. The following code is not evaluated is it would change the results of the cached computations in the previous code chunk. > lapply(assaydata(cnset), delete) > lapply(batchstatistics(cnset), delete) > delete(cnset$gender) > delete(cnset$snr) > delete(cnset$skw) > rm(cnset) Copy number routines in the crlmm package are available for Affymetrix 5.0 and 6.0 platforms, as well as several Illumina platforms. This vignette assumes that the arrays have already been successfully preprocessed and genotyped as per the instructions in the AffymetrixPreprocessCN and IlluminaPreprocessCN vignettes for the Affymetrix and Illumina platforms, respectively. While this vignette uses Affymetrix 6.0 arrays for illustration, the steps at this point are identical for both platforms. See [1] for details regarding the methodology implemented in crlmm for copy number analysis. In addition, a compendium describing copy number analysis using the crlmm package is available from the author s website: http: // Limitations: While a minimum number of samples is not required for preprocessing and genotyping, copy number estimation in the crlmm package currently requires at least 10 samples per batch. The parameter estimates for copy number and the corresponding estimates of raw copy number will tend to be more noisy for batches with small sample sizes (e.g., < 50). Chemistry plate or scan date are often useful surrogates for batch. Samples that were processed at similar times (e.g., in the same month) can be grouped together in the same batch. 2
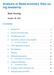
3 3 Quality control The signal to noise ratio (SNR) estimated by the CRLMM genotyping algorithm is an overall measure of the separation of the diallelic genotype clusters at polymorphic loci and can be a useful measure of array quality. Small SNR values can indicate possible problems with the DNA. Depending on the size of the dataset and the number of samples with low SNR, users may wish to rerun the preprocessing and genotyping steps after excluding samples with low SNR. The SNR is stored in the phenodata slot of the CNSet object and is available after preprocessing and genotyping. SNR values below 5 for Affymetrix or below 25 for Illumina may indicate poor sample quality. The following code chunk makes a histogram of the SNR values for the HapMap samples. > library(lattice) > invisible(open(cnset$snr)) > snr <- cnset$snr[] > close(cnset$snr) > print(histogram(~snr, panel=function(...){ panel.histogram(...)}, breaks=25, xlim=range(snr), xlab="snr")) 4 Copy number estimation As described in [1], the CRLMM-CopyNumber algorithm fits a linear model to the normalized intensities stratified by the diallic genotype call. The intercept and slope from the linear model are both SNP- and batch-specific. The implementation in the crlmm package is encapsulated by the function crlmmcopynumber that, using the default settings, can be called by passing a single object of class CNSet. See the appropriate preprocessing/genotyping vignette for the construction of an object of class CNSet. > (cnset.updated <- crlmmcopynumber(cnset)) The following steps were performed by the crlmmcopynumber function: ˆ sufficient statistics for the genotype clusters for each batch ˆ unobserved genotype centers imputed ˆ posterior summaries of sufficient statistics ˆ intercept and slope for linear model Depending on the value of ocprobesets(), these summaries are computed for subsets of the markers to reduce the required RAM. Note that the value returned by the crlmmcopynumber function in the above example is TRUE. The reason the function returns TRUE in the above example is that the elements of the batchstatistics slot have the class ff matrix. Rather than keep the statistical summaries in memory, the summaries are written to files on disk using protocols described in the ff package. Hence, while the cnset object itself is unchanged as a result of the crlmmcopynumber function, the data on disk is updated accordingly. Users that are interested in accessing these low-level summaries can refer to the Infrastructure vignette. Note that the data structure depends on whether the elements of the batchstatistics slot are ff objects or ordinary matrices. In this example, the elements of batchstatistics have the class ff matrix. > nms <- ls(batchstatistics(cnset)) > cls <- rep(na, length(nms)) > for(i in seq_along(nms)) cls[i] <- class(batchstatistics(cnset)[[nms[i]]])[1] > all(cls == "ff_matrix") 3
4 The batch-specific statistical summaries computed by crlmmcopynumber are written to files on disk using protocols described in the R package ff. The value returned by crlmmcopynumber is TRUE, indicating that the files on disk have been successfully updated. Note that while the cnset object is unchanged, the values on disk are different. On the other hand, subsetting the cnset with the [ method coerces all of the elements to class matrix. The batch-specific summaries are now ordinary matrices stored in RAM. The object returned by crlmmcopynumber is an object of class CNSet with the matrices in the batchstatistics slot updated. > chr1.index <- which(chromosome(cnset) == 1) > open(cnset) > cnset2 <- cnset[chr1.index, ] > close(cnset) > for(i in seq_along(nms)) cls[i] <- class(batchstatistics(cnset2)[[nms[i]]])[1] > all(cls == "matrix") > cnset3 <- crlmmcopynumber(cnset2) > class(cnset3) [1] "CNSet" attr(,"package") [1] "oligoclasses" 4.1 Marker-specific estimates Raw total copy number. Several functions are available that will compute relatively quickly the allelespecific, raw copy number estimates. At allele k, marker i, sample j, and batch p, the estimate of allelespecific copy number is computed by subtracting the estimated background from the normalized intensity and scaling by the slope coefficient. More formally, { } 1 ĉ k,ijp = max (I k,ijp ˆν k,ip ), 0 for k {A, B}. (1) ˆφ k,ip See [1] for details. The function totalcopynumber translates the normalized intensities to an estimate of raw copy number by adding the allele-specific summaries in Equation (1). For large datasets, the calculation will not be instantaneous as the I/O can be substantial. Users should specify either a subset of the markers or a subset of the samples to avoid using all of the available RAM. For example, in the following code chunk we compute the total copy number at all markers for the first 2 samples, and the total copy number for chromosome 20 for the first 50 samples. > tmp <- totalcopynumber(cnset, i=seq_len(nrow(cnset)), j=1:2) > dim(tmp) [1] > tmp2 <- totalcopynumber(cnset, i=which(chromosome(cnset) == 20), j=seq_len(ncol(cnset))) > dim(tmp2) [1] Alternatively, the functions CA and CB compute the allele-specific copy number. For instance, the following code chunk computes the allele-specific summaries at all polymorphic loci for the first 2 samples. > snp.index <- which(issnp(cnset) &!is.na(chromosome(cnset))) > ca <- CA(cnSet, i=snp.index, j=1:2) > cb <- CB(cnSet, i=snp.index, j=1:2) 4
5 4.2 Container for log R ratios and B allele frequencies A useful container for storing the crlmm genotypes, genotype confidence scores, and the total or relative copy number at each marker is the oligosetlist class. Coercion of a CNSet object to a oligosnpset object can be acheived by using the function constructoligosetfrom as illustrated below. Users should note that if the assaydata elements in the CNSet instance are ff objects, the assaydata elements of each element in the oligosetlist object will be ff-dervied objects (a new total_cn*.ff file will be created in the ldpath() directory). > library(vanillaice) > open(cnset3) > oligosetlist <- constructoligosetlistfrom(cnset3, batch.name=batch(cnset3)[1]) > close(cnset3) NULL > show(oligosetlist) oligosetlist of length 1 > class(oligosetlist) [1] "oligosetlist" attr(,"package") [1] "oligoclasses" > ## oligosnpset of first chromosome > oligosetlist[[1]] oligosnpset (storagemode: lockedenvironment) assaydata: features, 43 samples element names: baf, call, callprobability, copynumber protocoldata: none phenodata samplenames: _A _B _B (43 total) varlabels: gender SNR SKW varmetadata: labeldescription featuredata featurenames: rs rs rs (27125 total) fvarlabels: issnp position chromosome fvarmetadata: labeldescription experimentdata: use 'experimentdata(object)' Annotation: human370v1c genome: hg18 Note that log R ratios stored in the oligosnpset object can be retrieved by the copynumber accessor. B allele frequences are retrieved by the baf accessor. > lrrlist <- copynumber(oligosetlist) > class(lrrlist) [1] "list" 5
6 > dim(lrrlist[[1]]) ## log R ratios for chromosome 1. [1] > baflist <- baf(oligosetlist) > dim(baflist[[1]]) ## B allele frequencies for chromosome 1 [1] Session information > tolatex(sessioninfo()) ˆ R version Patched ( r59713), x86_64-unknown-linux-gnu ˆ Locale: LC_CTYPE=en_US.iso885915, LC_NUMERIC=C, LC_TIME=en_US.iso885915, LC_COLLATE=en_US.iso885915, LC_MONETARY=en_US.iso885915, LC_MESSAGES=en_US.iso885915, LC_PAPER=C, LC_NAME=C, LC_ADDRESS=C, LC_TELEPHONE=C, LC_MEASUREMENT=en_US.iso885915, LC_IDENTIFICATION=C ˆ Base packages: base, datasets, graphics, grdevices, methods, stats, tools, utils ˆ Other packages: Biobase , BiocGenerics 0.2.0, BiocInstaller 1.4.7, bit 1.1-8, cachesweave 0.6-1, crlmm , ff 2.2-7, filehash 2.2-1, hapmap370k 1.0.1, human370v1ccrlmm 1.0.2, lattice , oligoclasses , stashr 0.3-5, VanillaICE ˆ Loaded via a namespace (and not attached): affyio , annotate , AnnotationDbi , Biostrings , codetools 0.2-8, compiler , DBI 0.2-5, digest 0.5.2, ellipse 0.3-7, foreach 1.4.0, genefilter , GenomicRanges 1.8.7, grid , IRanges , iterators 1.0.6, msm 1.1.1, mvtnorm , preprocesscore , RSQLite , splines , stats , survival , XML 3.9-4, xtable 1.7-0, zlibbioc References [1] Robert B Scharpf, Ingo Ruczinski, Benilton Carvalho, Betty Doan, Aravinda Chakravarti, and Rafael A Irizarry. A multilevel model to address batch effects in copy number estimation using snp arrays. Biostatistics, 12(1):33 50, Jan
7 0.20 Density SNR Figure 1: The signal to noise ratio (SNR) for 180 HapMap samples. For Affymetrix platforms, SNR values below 5 can indicate possible problems with sample quality. In some circumstances, it may be more helpful to exclude samples with poor DNA quality. 7
Using crlmm for copy number estimation and genotype calling with Illumina platforms
 Using crlmm for copy number estimation and genotype calling with Illumina platforms Rob Scharpf November, Abstract This vignette illustrates the steps necessary for obtaining marker-level estimates of
Using crlmm for copy number estimation and genotype calling with Illumina platforms Rob Scharpf November, Abstract This vignette illustrates the steps necessary for obtaining marker-level estimates of
An Introduction to the methylumi package
 An Introduction to the methylumi package Sean Davis and Sven Bilke October 13, 2014 1 Introduction Gene expression patterns are very important in understanding any biologic system. The regulation of gene
An Introduction to the methylumi package Sean Davis and Sven Bilke October 13, 2014 1 Introduction Gene expression patterns are very important in understanding any biologic system. The regulation of gene
Analysis of screens with enhancer and suppressor controls
 Analysis of screens with enhancer and suppressor controls Lígia Brás, Michael Boutros and Wolfgang Huber April 16, 2015 Contents 1 Introduction 1 2 Assembling the data 2 2.1 Reading the raw intensity files..................
Analysis of screens with enhancer and suppressor controls Lígia Brás, Michael Boutros and Wolfgang Huber April 16, 2015 Contents 1 Introduction 1 2 Assembling the data 2 2.1 Reading the raw intensity files..................
crlmm to downstream data analysis
 crlmm to downstream data analysis VJ Carey, B Carvalho March, 2012 1 Running CRLMM on a nontrivial set of CEL files To use the crlmm algorithm, the user must load the crlmm package, as described below:
crlmm to downstream data analysis VJ Carey, B Carvalho March, 2012 1 Running CRLMM on a nontrivial set of CEL files To use the crlmm algorithm, the user must load the crlmm package, as described below:
Introduction to the Codelink package
 Introduction to the Codelink package Diego Diez October 30, 2018 1 Introduction This package implements methods to facilitate the preprocessing and analysis of Codelink microarrays. Codelink is a microarray
Introduction to the Codelink package Diego Diez October 30, 2018 1 Introduction This package implements methods to facilitate the preprocessing and analysis of Codelink microarrays. Codelink is a microarray
Djork-Arné Clevert and Andreas Mitterecker. Institute of Bioinformatics, Johannes Kepler University Linz
 Software Manual Institute of Bioinformatics, Johannes Kepler University Linz cn.farms: a latent variable model to detect copy number variations in microarray data with a low false discovery rate Manual
Software Manual Institute of Bioinformatics, Johannes Kepler University Linz cn.farms: a latent variable model to detect copy number variations in microarray data with a low false discovery rate Manual
Package crlmm. October 15, 2018
 Type Package Package crlmm October 15, 2018 Title Genotype Calling (CRLMM) and Copy Number Analysis tool for Affymetrix SNP 5.0 and 6.0 and Illumina arrays Version 1.38.0 Author Benilton S Carvalho, Robert
Type Package Package crlmm October 15, 2018 Title Genotype Calling (CRLMM) and Copy Number Analysis tool for Affymetrix SNP 5.0 and 6.0 and Illumina arrays Version 1.38.0 Author Benilton S Carvalho, Robert
Analysis of two-way cell-based assays
 Analysis of two-way cell-based assays Lígia Brás, Michael Boutros and Wolfgang Huber April 16, 2015 Contents 1 Introduction 1 2 Assembling the data 2 2.1 Reading the raw intensity files..................
Analysis of two-way cell-based assays Lígia Brás, Michael Boutros and Wolfgang Huber April 16, 2015 Contents 1 Introduction 1 2 Assembling the data 2 2.1 Reading the raw intensity files..................
An Introduction to the methylumi package
 An Introduction to the methylumi package Sean Davis and Sven Bilke April 30, 2018 1 Introduction Gene expression patterns are very important in understanding any biologic system. The regulation of gene
An Introduction to the methylumi package Sean Davis and Sven Bilke April 30, 2018 1 Introduction Gene expression patterns are very important in understanding any biologic system. The regulation of gene
SimBindProfiles: Similar Binding Profiles, identifies common and unique regions in array genome tiling array data
 SimBindProfiles: Similar Binding Profiles, identifies common and unique regions in array genome tiling array data Bettina Fischer October 30, 2018 Contents 1 Introduction 1 2 Reading data and normalisation
SimBindProfiles: Similar Binding Profiles, identifies common and unique regions in array genome tiling array data Bettina Fischer October 30, 2018 Contents 1 Introduction 1 2 Reading data and normalisation
Djork-Arné Clevert and Andreas Mitterecker. Institute of Bioinformatics, Johannes Kepler University Linz
 Software Manual Institute of Bioinformatics, Johannes Kepler University Linz cn.farms: a latent variable model to detect copy number variations in microarray data with a low false discovery rate Manual
Software Manual Institute of Bioinformatics, Johannes Kepler University Linz cn.farms: a latent variable model to detect copy number variations in microarray data with a low false discovery rate Manual
How to use CNTools. Overview. Algorithms. Jianhua Zhang. April 14, 2011
 How to use CNTools Jianhua Zhang April 14, 2011 Overview Studies have shown that genomic alterations measured as DNA copy number variations invariably occur across chromosomal regions that span over several
How to use CNTools Jianhua Zhang April 14, 2011 Overview Studies have shown that genomic alterations measured as DNA copy number variations invariably occur across chromosomal regions that span over several
Introduction: microarray quality assessment with arrayqualitymetrics
 Introduction: microarray quality assessment with arrayqualitymetrics Audrey Kauffmann, Wolfgang Huber April 4, 2013 Contents 1 Basic use 3 1.1 Affymetrix data - before preprocessing.......................
Introduction: microarray quality assessment with arrayqualitymetrics Audrey Kauffmann, Wolfgang Huber April 4, 2013 Contents 1 Basic use 3 1.1 Affymetrix data - before preprocessing.......................
Simulation of molecular regulatory networks with graphical models
 Simulation of molecular regulatory networks with graphical models Inma Tur 1 inma.tur@upf.edu Alberto Roverato 2 alberto.roverato@unibo.it Robert Castelo 1 robert.castelo@upf.edu 1 Universitat Pompeu Fabra,
Simulation of molecular regulatory networks with graphical models Inma Tur 1 inma.tur@upf.edu Alberto Roverato 2 alberto.roverato@unibo.it Robert Castelo 1 robert.castelo@upf.edu 1 Universitat Pompeu Fabra,
Introduction to QDNAseq
 Ilari Scheinin October 30, 2017 Contents 1 Running QDNAseq.......................... 1 1.1 Bin annotations.......................... 1 1.2 Processing bam files....................... 2 1.3 Downstream analyses......................
Ilari Scheinin October 30, 2017 Contents 1 Running QDNAseq.......................... 1 1.1 Bin annotations.......................... 1 1.2 Processing bam files....................... 2 1.3 Downstream analyses......................
Wave correction for arrays
 Wave correction for arrays Eitan Halper-Stromberg and Robert B. Scharpf April 30, 2018 Abstract ArrayTV is a GC correction algorithm for microarray data designed to mitigate the appearance of waves. This
Wave correction for arrays Eitan Halper-Stromberg and Robert B. Scharpf April 30, 2018 Abstract ArrayTV is a GC correction algorithm for microarray data designed to mitigate the appearance of waves. This
An Introduction to Bioconductor s ExpressionSet Class
 An Introduction to Bioconductor s ExpressionSet Class Seth Falcon, Martin Morgan, and Robert Gentleman 6 October, 2006; revised 9 February, 2007 1 Introduction Biobase is part of the Bioconductor project,
An Introduction to Bioconductor s ExpressionSet Class Seth Falcon, Martin Morgan, and Robert Gentleman 6 October, 2006; revised 9 February, 2007 1 Introduction Biobase is part of the Bioconductor project,
How to Use pkgdeptools
 How to Use pkgdeptools Seth Falcon February 10, 2018 1 Introduction The pkgdeptools package provides tools for computing and analyzing dependency relationships among R packages. With it, you can build
How to Use pkgdeptools Seth Falcon February 10, 2018 1 Introduction The pkgdeptools package provides tools for computing and analyzing dependency relationships among R packages. With it, you can build
RAPIDR. Kitty Lo. November 20, Intended use of RAPIDR 1. 2 Create binned counts file from BAMs Masking... 1
 RAPIDR Kitty Lo November 20, 2014 Contents 1 Intended use of RAPIDR 1 2 Create binned counts file from BAMs 1 2.1 Masking.................................................... 1 3 Build the reference 2 3.1
RAPIDR Kitty Lo November 20, 2014 Contents 1 Intended use of RAPIDR 1 2 Create binned counts file from BAMs 1 2.1 Masking.................................................... 1 3 Build the reference 2 3.1
Analyze Illumina Infinium methylation microarray data
 Analyze Illumina Infinium methylation microarray data Pan Du, Gang Feng, Spencer Huang, Warren A. Kibbe, Simon Lin May 3, 2016 Genentech Inc., South San Francisco, CA, 94080, USA Biomedical Informatics
Analyze Illumina Infinium methylation microarray data Pan Du, Gang Feng, Spencer Huang, Warren A. Kibbe, Simon Lin May 3, 2016 Genentech Inc., South San Francisco, CA, 94080, USA Biomedical Informatics
An Introduction to the genoset Package
 An Introduction to the genoset Package Peter M. Haverty April 4, 2013 Contents 1 Introduction 2 1.1 Creating Objects........................................... 2 1.2 Accessing Genome Information...................................
An Introduction to the genoset Package Peter M. Haverty April 4, 2013 Contents 1 Introduction 2 1.1 Creating Objects........................................... 2 1.2 Accessing Genome Information...................................
survsnp: Power and Sample Size Calculations for SNP Association Studies with Censored Time to Event Outcomes
 survsnp: Power and Sample Size Calculations for SNP Association Studies with Censored Time to Event Outcomes Kouros Owzar Zhiguo Li Nancy Cox Sin-Ho Jung Chanhee Yi June 29, 2016 1 Introduction This vignette
survsnp: Power and Sample Size Calculations for SNP Association Studies with Censored Time to Event Outcomes Kouros Owzar Zhiguo Li Nancy Cox Sin-Ho Jung Chanhee Yi June 29, 2016 1 Introduction This vignette
VanillaICE: Hidden Markov Models for the Assessment of Chromosomal Alterations using High-throughput SNP Arrays
 VanillaICE: Hidden Markov Models for the Assessment of Chromosomal Alterations using High-throughput SNP Arrays Robert Scharpf March 16, 2016 Abstract This package provides an implementation of a hidden
VanillaICE: Hidden Markov Models for the Assessment of Chromosomal Alterations using High-throughput SNP Arrays Robert Scharpf March 16, 2016 Abstract This package provides an implementation of a hidden
The bumphunter user s guide
 The bumphunter user s guide Kasper Daniel Hansen khansen@jhsph.edu Martin Aryee aryee.martin@mgh.harvard.edu Rafael A. Irizarry rafa@jhu.edu Modified: November 23, 2012. Compiled: April 30, 2018 Introduction
The bumphunter user s guide Kasper Daniel Hansen khansen@jhsph.edu Martin Aryee aryee.martin@mgh.harvard.edu Rafael A. Irizarry rafa@jhu.edu Modified: November 23, 2012. Compiled: April 30, 2018 Introduction
Introduction: microarray quality assessment with arrayqualitymetrics
 : microarray quality assessment with arrayqualitymetrics Audrey Kauffmann, Wolfgang Huber October 30, 2017 Contents 1 Basic use............................... 3 1.1 Affymetrix data - before preprocessing.............
: microarray quality assessment with arrayqualitymetrics Audrey Kauffmann, Wolfgang Huber October 30, 2017 Contents 1 Basic use............................... 3 1.1 Affymetrix data - before preprocessing.............
Biobase development and the new eset
 Biobase development and the new eset Martin T. Morgan * 7 August, 2006 Revised 4 September, 2006 featuredata slot. Revised 20 April 2007 minor wording changes; verbose and other arguments passed through
Biobase development and the new eset Martin T. Morgan * 7 August, 2006 Revised 4 September, 2006 featuredata slot. Revised 20 April 2007 minor wording changes; verbose and other arguments passed through
How to use cghmcr. October 30, 2017
 How to use cghmcr Jianhua Zhang Bin Feng October 30, 2017 1 Overview Copy number data (arraycgh or SNP) can be used to identify genomic regions (Regions Of Interest or ROI) showing gains or losses that
How to use cghmcr Jianhua Zhang Bin Feng October 30, 2017 1 Overview Copy number data (arraycgh or SNP) can be used to identify genomic regions (Regions Of Interest or ROI) showing gains or losses that
segmentseq: methods for detecting methylation loci and differential methylation
 segmentseq: methods for detecting methylation loci and differential methylation Thomas J. Hardcastle October 30, 2018 1 Introduction This vignette introduces analysis methods for data from high-throughput
segmentseq: methods for detecting methylation loci and differential methylation Thomas J. Hardcastle October 30, 2018 1 Introduction This vignette introduces analysis methods for data from high-throughput
CGHregions: Dimension reduction for array CGH data with minimal information loss.
 CGHregions: Dimension reduction for array CGH data with minimal information loss. Mark van de Wiel and Sjoerd Vosse April 30, 2018 Contents Department of Pathology VU University Medical Center mark.vdwiel@vumc.nl
CGHregions: Dimension reduction for array CGH data with minimal information loss. Mark van de Wiel and Sjoerd Vosse April 30, 2018 Contents Department of Pathology VU University Medical Center mark.vdwiel@vumc.nl
Package SNPchip. May 3, 2018
 Version 2.26.0 Title Visualizations for copy number alterations Package SNPchip May 3, 2018 Author Robert Scharpf and Ingo Ruczinski Maintainer Robert Scharpf Depends
Version 2.26.0 Title Visualizations for copy number alterations Package SNPchip May 3, 2018 Author Robert Scharpf and Ingo Ruczinski Maintainer Robert Scharpf Depends
segmentseq: methods for detecting methylation loci and differential methylation
 segmentseq: methods for detecting methylation loci and differential methylation Thomas J. Hardcastle October 13, 2015 1 Introduction This vignette introduces analysis methods for data from high-throughput
segmentseq: methods for detecting methylation loci and differential methylation Thomas J. Hardcastle October 13, 2015 1 Introduction This vignette introduces analysis methods for data from high-throughput
ReadqPCR: Functions to load RT-qPCR data into R
 ReadqPCR: Functions to load RT-qPCR data into R James Perkins University College London Matthias Kohl Furtwangen University April 30, 2018 Contents 1 Introduction 1 2 read.lc480 2 3 read.qpcr 7 4 read.taqman
ReadqPCR: Functions to load RT-qPCR data into R James Perkins University College London Matthias Kohl Furtwangen University April 30, 2018 Contents 1 Introduction 1 2 read.lc480 2 3 read.qpcr 7 4 read.taqman
Image Analysis with beadarray
 Mike Smith October 30, 2017 Introduction From version 2.0 beadarray provides more flexibility in the processing of array images and the extraction of bead intensities than its predecessor. In the past
Mike Smith October 30, 2017 Introduction From version 2.0 beadarray provides more flexibility in the processing of array images and the extraction of bead intensities than its predecessor. In the past
Example data for use with the beadarray package
 Example data for use with the beadarray package Mark Dunning November 1, 2018 Contents 1 Data Introduction 1 2 Loading the data 2 3 Data creation 4 1 Data Introduction This package provides a lightweight
Example data for use with the beadarray package Mark Dunning November 1, 2018 Contents 1 Data Introduction 1 2 Loading the data 2 3 Data creation 4 1 Data Introduction This package provides a lightweight
NacoStringQCPro. Dorothee Nickles, Thomas Sandmann, Robert Ziman, Richard Bourgon. Modified: April 10, Compiled: April 24, 2017.
 NacoStringQCPro Dorothee Nickles, Thomas Sandmann, Robert Ziman, Richard Bourgon Modified: April 10, 2017. Compiled: April 24, 2017 Contents 1 Scope 1 2 Quick start 1 3 RccSet loaded with NanoString data
NacoStringQCPro Dorothee Nickles, Thomas Sandmann, Robert Ziman, Richard Bourgon Modified: April 10, 2017. Compiled: April 24, 2017 Contents 1 Scope 1 2 Quick start 1 3 RccSet loaded with NanoString data
Shrinkage of logarithmic fold changes
 Shrinkage of logarithmic fold changes Michael Love August 9, 2014 1 Comparing the posterior distribution for two genes First, we run a DE analysis on the Bottomly et al. dataset, once with shrunken LFCs
Shrinkage of logarithmic fold changes Michael Love August 9, 2014 1 Comparing the posterior distribution for two genes First, we run a DE analysis on the Bottomly et al. dataset, once with shrunken LFCs
Cardinal design and development
 Kylie A. Bemis October 30, 2017 Contents 1 Introduction.............................. 2 2 Design overview........................... 2 3 iset: high-throughput imaging experiments............ 3 3.1 SImageSet:
Kylie A. Bemis October 30, 2017 Contents 1 Introduction.............................. 2 2 Design overview........................... 2 3 iset: high-throughput imaging experiments............ 3 3.1 SImageSet:
Count outlier detection using Cook s distance
 Count outlier detection using Cook s distance Michael Love August 9, 2014 1 Run DE analysis with and without outlier removal The following vignette produces the Supplemental Figure of the effect of replacing
Count outlier detection using Cook s distance Michael Love August 9, 2014 1 Run DE analysis with and without outlier removal The following vignette produces the Supplemental Figure of the effect of replacing
How To Use GOstats and Category to do Hypergeometric testing with unsupported model organisms
 How To Use GOstats and Category to do Hypergeometric testing with unsupported model organisms M. Carlson October 30, 2017 1 Introduction This vignette is meant as an extension of what already exists in
How To Use GOstats and Category to do Hypergeometric testing with unsupported model organisms M. Carlson October 30, 2017 1 Introduction This vignette is meant as an extension of what already exists in
Affymetrix Microarrays
 Affymetrix Microarrays Cavan Reilly November 3, 2017 Table of contents Overview The CLL data set Quality Assessment and Remediation Preprocessing Testing for Differential Expression Moderated Tests Volcano
Affymetrix Microarrays Cavan Reilly November 3, 2017 Table of contents Overview The CLL data set Quality Assessment and Remediation Preprocessing Testing for Differential Expression Moderated Tests Volcano
An introduction to Genomic Data Structures
 An introduction to Genomic Data Structures Cavan Reilly October 30, 2017 Table of contents Object Oriented Programming The ALL data set ExpressionSet Objects Environments More on ExpressionSet Objects
An introduction to Genomic Data Structures Cavan Reilly October 30, 2017 Table of contents Object Oriented Programming The ALL data set ExpressionSet Objects Environments More on ExpressionSet Objects
ReadqPCR: Functions to load RT-qPCR data into R
 ReadqPCR: Functions to load RT-qPCR data into R James Perkins University College London February 15, 2013 Contents 1 Introduction 1 2 read.qpcr 1 3 read.taqman 3 4 qpcrbatch 4 1 Introduction The package
ReadqPCR: Functions to load RT-qPCR data into R James Perkins University College London February 15, 2013 Contents 1 Introduction 1 2 read.qpcr 1 3 read.taqman 3 4 qpcrbatch 4 1 Introduction The package
HowTo: Querying online Data
 HowTo: Querying online Data Jeff Gentry and Robert Gentleman November 12, 2017 1 Overview This article demonstrates how you can make use of the tools that have been provided for on-line querying of data
HowTo: Querying online Data Jeff Gentry and Robert Gentleman November 12, 2017 1 Overview This article demonstrates how you can make use of the tools that have been provided for on-line querying of data
Starr: Simple Tiling ARRay analysis for Affymetrix ChIP-chip data
 Starr: Simple Tiling ARRay analysis for Affymetrix ChIP-chip data Benedikt Zacher, Achim Tresch October 30, 2017 1 Introduction Starr is an extension of the Ringo [6] package for the analysis of ChIP-chip
Starr: Simple Tiling ARRay analysis for Affymetrix ChIP-chip data Benedikt Zacher, Achim Tresch October 30, 2017 1 Introduction Starr is an extension of the Ringo [6] package for the analysis of ChIP-chip
Creating a New Annotation Package using SQLForge
 Creating a New Annotation Package using SQLForge Marc Carlson, Herve Pages, Nianhua Li November 19, 2013 1 Introduction The AnnotationForge package provides a series of functions that can be used to build
Creating a New Annotation Package using SQLForge Marc Carlson, Herve Pages, Nianhua Li November 19, 2013 1 Introduction The AnnotationForge package provides a series of functions that can be used to build
Bioconductor: Annotation Package Overview
 Bioconductor: Annotation Package Overview April 30, 2018 1 Overview In its current state the basic purpose of annotate is to supply interface routines that support user actions that rely on the different
Bioconductor: Annotation Package Overview April 30, 2018 1 Overview In its current state the basic purpose of annotate is to supply interface routines that support user actions that rely on the different
BioConductor Overviewr
 BioConductor Overviewr 2016-09-28 Contents Installing Bioconductor 1 Bioconductor basics 1 ExressionSet 2 assaydata (gene expression)........................................ 2 phenodata (sample annotations).....................................
BioConductor Overviewr 2016-09-28 Contents Installing Bioconductor 1 Bioconductor basics 1 ExressionSet 2 assaydata (gene expression)........................................ 2 phenodata (sample annotations).....................................
Analysis of Bead-level Data using beadarray
 Mark Dunning October 30, 2017 Introduction beadarray is a package for the pre-processing and analysis of Illumina BeadArray. The main advantage is being able to read raw data output by Illumina s scanning
Mark Dunning October 30, 2017 Introduction beadarray is a package for the pre-processing and analysis of Illumina BeadArray. The main advantage is being able to read raw data output by Illumina s scanning
Preparing the Final Data Set
 Preparing the Final Data Set Kevin R. Coombes 17 March 2011 Contents 1 Executive Summary 1 1.1 Introduction......................................... 1 1.1.1 Aims/Objectives..................................
Preparing the Final Data Set Kevin R. Coombes 17 March 2011 Contents 1 Executive Summary 1 1.1 Introduction......................................... 1 1.1.1 Aims/Objectives..................................
The rtracklayer package
 The rtracklayer package Michael Lawrence January 22, 2018 Contents 1 Introduction 2 2 Gene expression and microrna target sites 2 2.1 Creating a target site track..................... 2 2.1.1 Constructing
The rtracklayer package Michael Lawrence January 22, 2018 Contents 1 Introduction 2 2 Gene expression and microrna target sites 2 2.1 Creating a target site track..................... 2 2.1.1 Constructing
GxE.scan. October 30, 2018
 GxE.scan October 30, 2018 Overview GxE.scan can process a GWAS scan using the snp.logistic, additive.test, snp.score or snp.matched functions, whereas snp.scan.logistic only calls snp.logistic. GxE.scan
GxE.scan October 30, 2018 Overview GxE.scan can process a GWAS scan using the snp.logistic, additive.test, snp.score or snp.matched functions, whereas snp.scan.logistic only calls snp.logistic. GxE.scan
Preliminary Figures for Renormalizing Illumina SNP Cell Line Data
 Preliminary Figures for Renormalizing Illumina SNP Cell Line Data Kevin R. Coombes 17 March 2011 Contents 1 Executive Summary 1 1.1 Introduction......................................... 1 1.1.1 Aims/Objectives..................................
Preliminary Figures for Renormalizing Illumina SNP Cell Line Data Kevin R. Coombes 17 March 2011 Contents 1 Executive Summary 1 1.1 Introduction......................................... 1 1.1.1 Aims/Objectives..................................
Advanced analysis using bayseq; generic distribution definitions
 Advanced analysis using bayseq; generic distribution definitions Thomas J Hardcastle October 30, 2017 1 Generic Prior Distributions bayseq now offers complete user-specification of underlying distributions
Advanced analysis using bayseq; generic distribution definitions Thomas J Hardcastle October 30, 2017 1 Generic Prior Distributions bayseq now offers complete user-specification of underlying distributions
The analysis of rtpcr data
 The analysis of rtpcr data Jitao David Zhang, Markus Ruschhaupt October 30, 2017 With the help of this document, an analysis of rtpcr data can be performed. For this, the user has to specify several parameters
The analysis of rtpcr data Jitao David Zhang, Markus Ruschhaupt October 30, 2017 With the help of this document, an analysis of rtpcr data can be performed. For this, the user has to specify several parameters
MIGSA: Getting pbcmc datasets
 MIGSA: Getting pbcmc datasets Juan C Rodriguez Universidad Católica de Córdoba Universidad Nacional de Córdoba Cristóbal Fresno Instituto Nacional de Medicina Genómica Andrea S Llera Fundación Instituto
MIGSA: Getting pbcmc datasets Juan C Rodriguez Universidad Católica de Córdoba Universidad Nacional de Córdoba Cristóbal Fresno Instituto Nacional de Medicina Genómica Andrea S Llera Fundación Instituto
riboseqr Introduction Getting Data Workflow Example Thomas J. Hardcastle, Betty Y.W. Chung October 30, 2017
 riboseqr Thomas J. Hardcastle, Betty Y.W. Chung October 30, 2017 Introduction Ribosome profiling extracts those parts of a coding sequence currently bound by a ribosome (and thus, are likely to be undergoing
riboseqr Thomas J. Hardcastle, Betty Y.W. Chung October 30, 2017 Introduction Ribosome profiling extracts those parts of a coding sequence currently bound by a ribosome (and thus, are likely to be undergoing
Using FARMS for summarization Using I/NI-calls for gene filtering. Djork-Arné Clevert. Institute of Bioinformatics, Johannes Kepler University Linz
 Software Manual Institute of Bioinformatics, Johannes Kepler University Linz Using FARMS for summarization Using I/NI-calls for gene filtering Djork-Arné Clevert Institute of Bioinformatics, Johannes Kepler
Software Manual Institute of Bioinformatics, Johannes Kepler University Linz Using FARMS for summarization Using I/NI-calls for gene filtering Djork-Arné Clevert Institute of Bioinformatics, Johannes Kepler
Package frma. R topics documented: March 8, Version Date Title Frozen RMA and Barcode
 Version 1.34.0 Date 2017-11-08 Title Frozen RMA and Barcode Package frma March 8, 2019 Preprocessing and analysis for single microarrays and microarray batches. Author Matthew N. McCall ,
Version 1.34.0 Date 2017-11-08 Title Frozen RMA and Barcode Package frma March 8, 2019 Preprocessing and analysis for single microarrays and microarray batches. Author Matthew N. McCall ,
How to use the rbsurv Package
 How to use the rbsurv Package HyungJun Cho, Sukwoo Kim, Soo-heang Eo, and Jaewoo Kang April 30, 2018 Contents 1 Introduction 1 2 Robust likelihood-based survival modeling 2 3 Algorithm 2 4 Example: Glioma
How to use the rbsurv Package HyungJun Cho, Sukwoo Kim, Soo-heang Eo, and Jaewoo Kang April 30, 2018 Contents 1 Introduction 1 2 Robust likelihood-based survival modeling 2 3 Algorithm 2 4 Example: Glioma
Triform: peak finding in ChIP-Seq enrichment profiles for transcription factors
 Triform: peak finding in ChIP-Seq enrichment profiles for transcription factors Karl Kornacker * and Tony Håndstad October 30, 2018 A guide for using the Triform algorithm to predict transcription factor
Triform: peak finding in ChIP-Seq enrichment profiles for transcription factors Karl Kornacker * and Tony Håndstad October 30, 2018 A guide for using the Triform algorithm to predict transcription factor
Creating a New Annotation Package using SQLForge
 Creating a New Annotation Package using SQLForge Marc Carlson, HervÃľ PagÃĺs, Nianhua Li April 30, 2018 1 Introduction The AnnotationForge package provides a series of functions that can be used to build
Creating a New Annotation Package using SQLForge Marc Carlson, HervÃľ PagÃĺs, Nianhua Li April 30, 2018 1 Introduction The AnnotationForge package provides a series of functions that can be used to build
Labelled quantitative proteomics with MSnbase
 Labelled quantitative proteomics with MSnbase Laurent Gatto lg390@cam.ac.uk Cambridge Centre For Proteomics University of Cambridge European Bioinformatics Institute (EBI) 18 th November 2010 Plan 1 Introduction
Labelled quantitative proteomics with MSnbase Laurent Gatto lg390@cam.ac.uk Cambridge Centre For Proteomics University of Cambridge European Bioinformatics Institute (EBI) 18 th November 2010 Plan 1 Introduction
Evaluation of VST algorithm in lumi package
 Evaluation of VST algorithm in lumi package Pan Du 1, Simon Lin 1, Wolfgang Huber 2, Warrren A. Kibbe 1 December 22, 2010 1 Robert H. Lurie Comprehensive Cancer Center Northwestern University, Chicago,
Evaluation of VST algorithm in lumi package Pan Du 1, Simon Lin 1, Wolfgang Huber 2, Warrren A. Kibbe 1 December 22, 2010 1 Robert H. Lurie Comprehensive Cancer Center Northwestern University, Chicago,
puma User Guide R. D. Pearson, X. Liu, M. Rattray, M. Milo, N. D. Lawrence G. Sanguinetti, Li Zhang October 30, Abstract 2 2 Citing puma 2
 puma User Guide R. D. Pearson, X. Liu, M. Rattray, M. Milo, N. D. Lawrence G. Sanguinetti, Li Zhang October 30, 2017 Contents 1 Abstract 2 2 Citing puma 2 3 Introduction 3 4 Introductory example analysis
puma User Guide R. D. Pearson, X. Liu, M. Rattray, M. Milo, N. D. Lawrence G. Sanguinetti, Li Zhang October 30, 2017 Contents 1 Abstract 2 2 Citing puma 2 3 Introduction 3 4 Introductory example analysis
Design Issues in Matrix package Development
 Design Issues in Matrix package Development Martin Maechler and Douglas Bates R Core Development Team maechler@stat.math.ethz.ch, bates@r-project.org Spring 2008 (typeset on November 16, 2017) Abstract
Design Issues in Matrix package Development Martin Maechler and Douglas Bates R Core Development Team maechler@stat.math.ethz.ch, bates@r-project.org Spring 2008 (typeset on November 16, 2017) Abstract
Analysis of Pairwise Interaction Screens
 Analysis of Pairwise Interaction Screens Bernd Fischer April 11, 2014 Contents 1 Introduction 1 2 Installation of the RNAinteract package 1 3 Creating an RNAinteract object 2 4 Single Perturbation effects
Analysis of Pairwise Interaction Screens Bernd Fischer April 11, 2014 Contents 1 Introduction 1 2 Installation of the RNAinteract package 1 3 Creating an RNAinteract object 2 4 Single Perturbation effects
Reorganizing the data by sample
 Reorganizing the data by sample Kevin R. Coombes 23 March 2011 Contents 1 Executive Summary 1 1.1 Introduction......................................... 1 1.1.1 Aims/Objectives..................................
Reorganizing the data by sample Kevin R. Coombes 23 March 2011 Contents 1 Executive Summary 1 1.1 Introduction......................................... 1 1.1.1 Aims/Objectives..................................
Overview of ensemblvep
 Overview of ensemblvep Lori Shepherd and Valerie Obenchain October 30, 2017 Contents 1 Introduction 1 2 Results as R objects 1 3 Write results to a file 4 4 Configuring runtime options 5 5 sessioninfo()
Overview of ensemblvep Lori Shepherd and Valerie Obenchain October 30, 2017 Contents 1 Introduction 1 2 Results as R objects 1 3 Write results to a file 4 4 Configuring runtime options 5 5 sessioninfo()
R Basics. Laurent Gatto Computational Statistics for Genome Biology 2 nd July 2012
 R Basics Laurent Gatto lg390@cam.ac.uk Computational Statistics for Genome Biology 2 nd July 2012 Plan Introduction Data types and structures Basic data types Higher order objects Manipulating data Subsetting
R Basics Laurent Gatto lg390@cam.ac.uk Computational Statistics for Genome Biology 2 nd July 2012 Plan Introduction Data types and structures Basic data types Higher order objects Manipulating data Subsetting
Introduction to R. Educational Materials 2007 S. Falcon, R. Ihaka, and R. Gentleman
 Introduction to R Educational Materials 2007 S. Falcon, R. Ihaka, and R. Gentleman 1 Data Structures ˆ R has a rich set of self-describing data structures. > class(z) [1] "character" > class(x) [1] "data.frame"
Introduction to R Educational Materials 2007 S. Falcon, R. Ihaka, and R. Gentleman 1 Data Structures ˆ R has a rich set of self-describing data structures. > class(z) [1] "character" > class(x) [1] "data.frame"
Creating IGV HTML reports with tracktables
 Thomas Carroll 1 [1em] 1 Bioinformatics Core, MRC Clinical Sciences Centre; thomas.carroll (at)imperial.ac.uk June 13, 2018 Contents 1 The tracktables package.................... 1 2 Creating IGV sessions
Thomas Carroll 1 [1em] 1 Bioinformatics Core, MRC Clinical Sciences Centre; thomas.carroll (at)imperial.ac.uk June 13, 2018 Contents 1 The tracktables package.................... 1 2 Creating IGV sessions
An Overview of the S4Vectors package
 Patrick Aboyoun, Michael Lawrence, Hervé Pagès Edited: February 2018; Compiled: June 7, 2018 Contents 1 Introduction.............................. 1 2 Vector-like and list-like objects...................
Patrick Aboyoun, Michael Lawrence, Hervé Pagès Edited: February 2018; Compiled: June 7, 2018 Contents 1 Introduction.............................. 1 2 Vector-like and list-like objects...................
Starr: Simple Tiling ARRay analysis for Affymetrix ChIP-chip data
 Starr: Simple Tiling ARRay analysis for Affymetrix ChIP-chip data Benedikt Zacher, Achim Tresch January 20, 2010 1 Introduction Starr is an extension of the Ringo [6] package for the analysis of ChIP-chip
Starr: Simple Tiling ARRay analysis for Affymetrix ChIP-chip data Benedikt Zacher, Achim Tresch January 20, 2010 1 Introduction Starr is an extension of the Ringo [6] package for the analysis of ChIP-chip
Reorganizing the data by sample
 Reorganizing the data by sample Kevin R. Coombes 17 March 2011 Contents 1 Executive Summary 1 1.1 Introduction......................................... 1 1.1.1 Aims/Objectives..................................
Reorganizing the data by sample Kevin R. Coombes 17 March 2011 Contents 1 Executive Summary 1 1.1 Introduction......................................... 1 1.1.1 Aims/Objectives..................................
Design Issues in Matrix package Development
 Design Issues in Matrix package Development Martin Maechler and Douglas Bates R Core Development Team maechler@stat.math.ethz.ch, bates@r-project.org Spring 2008 (typeset on October 26, 2018) Abstract
Design Issues in Matrix package Development Martin Maechler and Douglas Bates R Core Development Team maechler@stat.math.ethz.ch, bates@r-project.org Spring 2008 (typeset on October 26, 2018) Abstract
Analysis of Bead-summary Data using beadarray
 Analysis of Bead-summary Data using beadarray Mark Dunning October 30, 2017 Contents 1 Introduction............................. 2 2 feature and pheno data..................... 4 3 Subsetting the data.......................
Analysis of Bead-summary Data using beadarray Mark Dunning October 30, 2017 Contents 1 Introduction............................. 2 2 feature and pheno data..................... 4 3 Subsetting the data.......................
Gene set enrichment analysis of RNA-Seq data with the SeqGSEA package
 Gene set enrichment analysis of RNA-Seq data with the SeqGSEA package Xi Wang 1,2 and Murray Cairns 1,2,3 April 30, 2018 1 School of Biomedical Sciences and Pharmacy, The University of Newcastle, Callaghan,
Gene set enrichment analysis of RNA-Seq data with the SeqGSEA package Xi Wang 1,2 and Murray Cairns 1,2,3 April 30, 2018 1 School of Biomedical Sciences and Pharmacy, The University of Newcastle, Callaghan,
Raman Spectra of Chondrocytes in Cartilage: hyperspec s chondro data set
 Raman Spectra of Chondrocytes in Cartilage: hyperspec s chondro data set Claudia Beleites CENMAT and DI3, University of Trieste Spectroscopy Imaging, IPHT Jena e.v. February 13,
Raman Spectra of Chondrocytes in Cartilage: hyperspec s chondro data set Claudia Beleites CENMAT and DI3, University of Trieste Spectroscopy Imaging, IPHT Jena e.v. February 13,
Visualisation, transformations and arithmetic operations for grouped genomic intervals
 ## Warning: replacing previous import ggplot2::position by BiocGenerics::Position when loading soggi Visualisation, transformations and arithmetic operations for grouped genomic intervals Thomas Carroll
## Warning: replacing previous import ggplot2::position by BiocGenerics::Position when loading soggi Visualisation, transformations and arithmetic operations for grouped genomic intervals Thomas Carroll
Overview of ensemblvep
 Overview of ensemblvep Lori Shepherd and Valerie Obenchain October 30, 2018 Contents 1 Introduction 1 2 Results as R objects 1 3 Write results to a file 4 4 Configuring runtime options 5 5 sessioninfo()
Overview of ensemblvep Lori Shepherd and Valerie Obenchain October 30, 2018 Contents 1 Introduction 1 2 Results as R objects 1 3 Write results to a file 4 4 Configuring runtime options 5 5 sessioninfo()
Package pdinfobuilder
 Package pdinfobuilder April 10, 2018 Title Platform Design Information Package Builder Builds platform design information packages. These consist of a SQLite database containing feature-level data such
Package pdinfobuilder April 10, 2018 Title Platform Design Information Package Builder Builds platform design information packages. These consist of a SQLite database containing feature-level data such
Calibration of Quinine Fluorescence Emission Vignette for the Data Set flu of the R package hyperspec
 Calibration of Quinine Fluorescence Emission Vignette for the Data Set flu of the R package hyperspec Claudia Beleites CENMAT and DI3, University of Trieste Spectroscopy Imaging,
Calibration of Quinine Fluorescence Emission Vignette for the Data Set flu of the R package hyperspec Claudia Beleites CENMAT and DI3, University of Trieste Spectroscopy Imaging,
Working with aligned nucleotides (WORK- IN-PROGRESS!)
 Working with aligned nucleotides (WORK- IN-PROGRESS!) Hervé Pagès Last modified: January 2014; Compiled: November 17, 2017 Contents 1 Introduction.............................. 1 2 Load the aligned reads
Working with aligned nucleotides (WORK- IN-PROGRESS!) Hervé Pagès Last modified: January 2014; Compiled: November 17, 2017 Contents 1 Introduction.............................. 1 2 Load the aligned reads
Course Introduction. Welcome! About us. Thanks to Cancer Research Uk! Mark Dunning 27/07/2015
 Course Introduction Mark Dunning 27/07/2015 Welcome! About us Admin at CRUK Victoria, Helen Genetics Paul, Gabry, Cathy Thanks to Cancer Research Uk! Admin Lunch, tea / coffee breaks provided Workshop
Course Introduction Mark Dunning 27/07/2015 Welcome! About us Admin at CRUK Victoria, Helen Genetics Paul, Gabry, Cathy Thanks to Cancer Research Uk! Admin Lunch, tea / coffee breaks provided Workshop
RMassBank for XCMS. Erik Müller. January 4, Introduction 2. 2 Input files LC/MS data Additional Workflow-Methods 2
 RMassBank for XCMS Erik Müller January 4, 2019 Contents 1 Introduction 2 2 Input files 2 2.1 LC/MS data........................... 2 3 Additional Workflow-Methods 2 3.1 Options..............................
RMassBank for XCMS Erik Müller January 4, 2019 Contents 1 Introduction 2 2 Input files 2 2.1 LC/MS data........................... 2 3 Additional Workflow-Methods 2 3.1 Options..............................
500K Data Analysis Workflow using BRLMM
 500K Data Analysis Workflow using BRLMM I. INTRODUCTION TO BRLMM ANALYSIS TOOL... 2 II. INSTALLATION AND SET-UP... 2 III. HARDWARE REQUIREMENTS... 3 IV. BRLMM ANALYSIS TOOL WORKFLOW... 3 V. RESULTS/OUTPUT
500K Data Analysis Workflow using BRLMM I. INTRODUCTION TO BRLMM ANALYSIS TOOL... 2 II. INSTALLATION AND SET-UP... 2 III. HARDWARE REQUIREMENTS... 3 IV. BRLMM ANALYSIS TOOL WORKFLOW... 3 V. RESULTS/OUTPUT
Sparse Model Matrices
 Sparse Model Matrices Martin Maechler R Core Development Team maechler@r-project.org July 2007, 2008 (typeset on April 6, 2018) Introduction Model matrices in the very widely used (generalized) linear
Sparse Model Matrices Martin Maechler R Core Development Team maechler@r-project.org July 2007, 2008 (typeset on April 6, 2018) Introduction Model matrices in the very widely used (generalized) linear
Fitting Baselines to Spectra
 Fitting Baselines to Spectra Claudia Beleites DIA Raman Spectroscopy Group, University of Trieste/Italy (2005 2008) Spectroscopy Imaging, IPHT, Jena/Germany (2008 2016) ÖPV, JKI,
Fitting Baselines to Spectra Claudia Beleites DIA Raman Spectroscopy Group, University of Trieste/Italy (2005 2008) Spectroscopy Imaging, IPHT, Jena/Germany (2008 2016) ÖPV, JKI,
Introduction to the TSSi package: Identification of Transcription Start Sites
 Introduction to the TSSi package: Identification of Transcription Start Sites Julian Gehring, Clemens Kreutz October 30, 2017 Along with the advances in high-throughput sequencing, the detection of transcription
Introduction to the TSSi package: Identification of Transcription Start Sites Julian Gehring, Clemens Kreutz October 30, 2017 Along with the advances in high-throughput sequencing, the detection of transcription
Programming with R. Educational Materials 2006 S. Falcon, R. Ihaka, and R. Gentleman
 Programming with R Educational Materials 2006 S. Falcon, R. Ihaka, and R. Gentleman 1 Data Structures ˆ R has a rich set of self-describing data structures. > class(z) [1] "character" > class(x) [1] "data.frame"
Programming with R Educational Materials 2006 S. Falcon, R. Ihaka, and R. Gentleman 1 Data Structures ˆ R has a rich set of self-describing data structures. > class(z) [1] "character" > class(x) [1] "data.frame"
A complete analysis of peptide microarray binding data using the pepstat framework
 A complete analysis of peptide microarray binding data using the pepstat framework Greg Imholte, Renan Sauteraud, Mike Jiang and Raphael Gottardo April 30, 2018 This document present a full analysis, from
A complete analysis of peptide microarray binding data using the pepstat framework Greg Imholte, Renan Sauteraud, Mike Jiang and Raphael Gottardo April 30, 2018 This document present a full analysis, from
genocn: integrated studies of copy number and genotype
 genocn: integrated studies of copy number and genotype Sun, W., Wright, F., Tang, Z., Nordgard, S.H., Van Loo, P., Yu, T., Kristensen, V., Perou, C. February 22, 2010 1 Overview > library(genocn) This
genocn: integrated studies of copy number and genotype Sun, W., Wright, F., Tang, Z., Nordgard, S.H., Van Loo, P., Yu, T., Kristensen, V., Perou, C. February 22, 2010 1 Overview > library(genocn) This
End-to-end analysis of cell-based screens: from raw intensity readings to the annotated hit list (Short version)
 End-to-end analysis of cell-based screens: from raw intensity readings to the annotated hit list (Short version) Michael Boutros, Lígia Brás, Florian Hahne and Wolfgang Huber October 30, 2018 Contents
End-to-end analysis of cell-based screens: from raw intensity readings to the annotated hit list (Short version) Michael Boutros, Lígia Brás, Florian Hahne and Wolfgang Huber October 30, 2018 Contents
An Introduction to ShortRead
 Martin Morgan Modified: 21 October, 2013. Compiled: April 30, 2018 > library("shortread") The ShortRead package provides functionality for working with FASTQ files from high throughput sequence analysis.
Martin Morgan Modified: 21 October, 2013. Compiled: April 30, 2018 > library("shortread") The ShortRead package provides functionality for working with FASTQ files from high throughput sequence analysis.
Manual for the R gaga package
 Manual for the R gaga package David Rossell Department of Bioinformatics & Biostatistics IRB Barcelona Barcelona, Spain. rosselldavid@gmail.com Here we illustrate several uses of the package gaga, including
Manual for the R gaga package David Rossell Department of Bioinformatics & Biostatistics IRB Barcelona Barcelona, Spain. rosselldavid@gmail.com Here we illustrate several uses of the package gaga, including
Package GEM. R topics documented: January 31, Type Package
 Type Package Package GEM January 31, 2018 Title GEM: fast association study for the interplay of Gene, Environment and Methylation Version 1.5.0 Date 2015-12-05 Author Hong Pan, Joanna D Holbrook, Neerja
Type Package Package GEM January 31, 2018 Title GEM: fast association study for the interplay of Gene, Environment and Methylation Version 1.5.0 Date 2015-12-05 Author Hong Pan, Joanna D Holbrook, Neerja
NOISeq: Differential Expression in RNA-seq
 NOISeq: Differential Expression in RNA-seq Sonia Tarazona (starazona@cipf.es) Pedro Furió-Tarí (pfurio@cipf.es) María José Nueda (mj.nueda@ua.es) Alberto Ferrer (aferrer@eio.upv.es) Ana Conesa (aconesa@cipf.es)
NOISeq: Differential Expression in RNA-seq Sonia Tarazona (starazona@cipf.es) Pedro Furió-Tarí (pfurio@cipf.es) María José Nueda (mj.nueda@ua.es) Alberto Ferrer (aferrer@eio.upv.es) Ana Conesa (aconesa@cipf.es)
Comparing Least Squares Calculations
 Comparing Least Squares Calculations Douglas Bates R Development Core Team Douglas.Bates@R-project.org November 16, 2017 Abstract Many statistics methods require one or more least squares problems to be
Comparing Least Squares Calculations Douglas Bates R Development Core Team Douglas.Bates@R-project.org November 16, 2017 Abstract Many statistics methods require one or more least squares problems to be
MMDiff2: Statistical Testing for ChIP-Seq Data Sets
 MMDiff2: Statistical Testing for ChIP-Seq Data Sets Gabriele Schwikert 1 and David Kuo 2 [1em] 1 The Informatics Forum, University of Edinburgh and The Wellcome
MMDiff2: Statistical Testing for ChIP-Seq Data Sets Gabriele Schwikert 1 and David Kuo 2 [1em] 1 The Informatics Forum, University of Edinburgh and The Wellcome
Analyse RT PCR data with ddct
 Analyse RT PCR data with ddct Jitao David Zhang, Rudolf Biczok and Markus Ruschhaupt October 30, 2017 Abstract Quantitative real time PCR (qrt PCR or RT PCR for short) is a laboratory technique based on
Analyse RT PCR data with ddct Jitao David Zhang, Rudolf Biczok and Markus Ruschhaupt October 30, 2017 Abstract Quantitative real time PCR (qrt PCR or RT PCR for short) is a laboratory technique based on
