Merge Conflicts p. 92 More GitHub Workflows: Forking and Pull Requests p. 97 Using Git to Make Life Easier: Working with Past Commits p.
|
|
|
- Byron White
- 5 years ago
- Views:
Transcription
1 Preface p. xiii Ideology: Data Skills for Robust and Reproducible Bioinformatics How to Learn Bioinformatics p. 1 Why Bioinformatics? Biology's Growing Data p. 1 Learning Data Skills to Learn Bioinformatics p. 4 New Challenges for Reproducible and Robust Research p. 5 Reproducible Research p. 6 Robust Research and the Golden Rule of Bioinformatics p. 8 Adopting Robust and Reproducible Practices Will Make Your Life Easier, Too p. 9 Recommendations for Robust Research p. 10 Pay Attention to Experimental Design p. 10 Write Code for Humans, Write Data for Computers p. 11 Let Your Computer Do the Work For You p. 12 Make Assertions and Be Loud, in Code and in Your Methods p. 12 Test Code, or Better Yet, Let Code Test Code p. 13 Use Existing Libraries Whenever Possible p. 14 Treat Data as Read-Only p. 14 Spend Time Developing Frequently Used Scripts into Tools p. 15 Let Data Prove That It's High Quality p. 15 Recommendations for Reproducible Research p. 16 Release Your Code and Data p. 16 Document Everything p. 16 Make Figures and Statistics the Results of Scripts p. 17 Use Code as Documentation p. 17 Continually Improving Your Bioinformatics Data Skills p. 17 Prerequisites: Essential Skills for Getting Started with a Bioinformatics Project Setting Up and Managing a Bioinformatics Project p. 21 Project Directories and Directory Structures p. 21 Project Documentation p. 24 Use Directories to Divide Up Your Project into Subprojects p. 26 Organizing Data to Automate File Processing Tasks p. 26 Markdown for Project Notebooks p. 31 Markdown Formatting Basics p. 31 Using Pandoc to Render Markdown to HTML p. 35 Remedial Unix Shell p. 37 Why Do We Use Unix in Bioinformatics? Modularity and the Unix Philosophy p. 37 Working with Streams and Redirection p. 41 Redirecting Standard Out to a File p. 41 Redirecting Standard Error p. 43 Using Standard Input Redirection p. 45 The Almighty Unix Pipe: Speed and Beauty in One p. 45
2 Pipes in Action: Creating Simple Programs with Grep and Pipes p. 47 Combining Pipes and Redirection p. 48 Even More Redirection: A tee in Your Pipe p. 49 Managing and Interacting with Processes p. 50 Background Processes p. 50 Killing Processes p. 51 Exit Status: How to Programmatically Tell Whether Your Command Worked p. 52 Command Substitution p. 54 Working with Remote Machines p. 57 Connecting to Remote Machines with SSH p. 57 Quick Authentication with SSH Keys p. 59 Maintaining Long-Running Jobs with nohup and tmux p. 61 Nohup p. 61 Working with Remote Machines Through Tmux p. 61 Installing and Configuring Tmux p. 62 Creating, Detaching, and Attaching Tmux Sessions p. 62 Working with Tmux Windows p. 64 Git for Scientists p. 67 Why Git Is Necessary in Bioinformatics Projects p. 68 Git Allows You to Keep Snapshots of Your Project p. 68 Git Helps You Keep Track of Important Changes to Code p. 69 Git Helps Keep Software Organized and Available After People Leave p. 69 Installing Git p. 70 Basic Git: Creating Repositories, Tracking Files, and Staging and Committing Changes p. 70 Git Setup: Telling Git Who You Are p. 70 Git init and git clone: Creating Repositories p. 70 Tracking Files in Git: git add and git status Part I p. 72 Staging Files in Git: git add and git status Part II p. 73 Git commit: Taking a Snapshot of Your Project p. 76 Seeing File Differences: git diff p. 77 Seeing Your Commit History: git log p. 79 Moving and Removing Files; git mv and git rm p. 80 Telling Git What to Ignore:.gitignore p. 81 Undoing a Stage: git reset p. 83 Collaborating with Git: Git Remotes, git push, and git pull p. 83 Creating a Shared Central Repository with GitHub p. 86 Authenticating with Git Remotes p. 87 Connecting with Git Remotes: git remote p. 87 Pushing Commits to a Remote Repository with git push p. 88 Pulling Commits from a Remote Repository with git pull p. 89 Working with Your Collaborators: Pushing and Pulling p. 90
3 Merge Conflicts p. 92 More GitHub Workflows: Forking and Pull Requests p. 97 Using Git to Make Life Easier: Working with Past Commits p. 97 Getting Files from the Past: git checkout p. 97 Stashing Your Changes: git stash p. 99 More git diff: Comparing Commits and Files p. 100 Undoing and Editing Commits: git commit -amend p. 102 Working with Branches p. 102 Creating and Working with Branches; git branch and git checkout p. 103 Merging Branches: git merge p. 105 Branches and Remotes p. 106 Continuing Your Git Education p. 108 Bioinformatics Data p. 109 Retrieving Bioinformatics Data p. 110 Downloading Data with wget and curl p. 110 Rsync and Secure Copy (scp) p. 113 Data Integrity p. 114 SHA and MD5 Checksums p. 115 Looking at Differences Between Data p. 116 Compressing Data and Working with Compressed Data p. 118 Gzip p. 119 Working with Gzipped Compressed Files p. 120 Case Study: Reproducibly Downloading Data p. 120 Practice: Bioinformatics Data Skills Unix Data Tools p. 125 Unix Data Tools and the Unix One-Liner Approach: Lessons from Programming Pearls p. 125 When to Use the Unix Pipeline Approach and How to Use It Safely p. 127 Inspecting and Manipulating Text Data with Unix Tools p. 128 Inspecting Data with Head and Tail p. 129 Less p. 131 Plain-Text Data Summary Information with wc, Is, and awk p. 134 Working with Column Data with cut and Columns p. 138 Formatting Tabular Data with column p. 139 The All-powerful Grep p. 140 Decoding Plain-Text Data: hexdump p. 145 Sorting Plain-Text Data with Sort p. 147 Finding Unique Values in Uniq p. 152 Join p. 155 Text Processing with Awk p. 157 Bioawk: An Awk for Biological Formats p. 163 Stream Editing with Sed p. 165
4 Advanced Shell Tricks p. 169 Subshells p. 169 Named Pipes and Process Substitution p. 171 The Unix Philosophy Revisited p. 173 A Rapid Introduction to the R Language p. 175 Getting Started with R and RStudio p. 176 R Language Basics p. 178 Simple Calculations in R, Calling Functions, and Getting Help in R p. 178 Variables and Assignment p. 182 Vectors, Vectorization, and Indexing p. 183 Working with and Visualizing Data in R p. 193 Loading Data into R p. 194 Exploring and Transforming Dataframes p. 199 Exploring Data Through Slicing and Dicing: Subsetting Dataframes p. 203 Exploring Data Visually with ggplot2 I: Scatterplots and Densities p. 207 Exploring Data Visually with ggplot2 II: Smoothing p. 213 Binning Data with cut() and Bar Plots with ggplot2 p. 215 Merging and Combining Data: Matching Vectors and Merging Dataframes p. 219 Using ggplot2 Facets p. 224 More R Data Structures: Lists p. 228 Writing and Applying Functions to Lists with lapply() and sapply() p. 231 Working with the Split-Apply-Combine Pattern p. 239 Exploring Dataframes with dplyr p. 243 Working with Strings p. 248 Developing Workflows with R Scripts p. 253 Control Flow: if, for, and while p. 253 Working with R Scripts p. 254 Workflows for Loading and Combining Multiple Files p. 257 Exporting Data p. 260 Further R Directions and Resources p. 261 Working with Range Data p. 263 A Crash Course in Genomic Ranges and Coordinate Systems p. 264 An Interactive Introduction to Range Data with GenomicRanges p. 269 Installing and Working with Bioconductor Packages p. 269 Storing Generic Ranges with IRanges p. 270 Basic Range Operations: Arithmetic, Transformations, and Set Operations p. 275 Finding Overlapping Ranges p. 281 Finding Nearest Ranges and Calculating Distance p. 290 Run Length Encoding and Views p. 292 Storing Genomic Ranges with GenomicRanges p. 299 Grouping Data with GRangesList p. 303
5 Working with Annotation Data: GenomicFeatures and rtracklayer p. 308 Retrieving Promoter Regions: Flank and Promoters p. 314 Retrieving Promoter Sequence: Connection GenomicRanges with Sequence Data p. 316 Getting Intergenic and Intronic Regions: Gaps, Reduce, and Setdiffs in Practice p. 319 Finding and Working with Overlapping Ranges p. 324 Calculating Coverage of GRanges Objects p. 328 Working with Ranges Data on the Command Line with BEDTools p. 329 Computing Overlaps with BEDTools Intersect p. 330 BEDTools Slop and Flank p. 333 Coverage with BEDTools p. 335 Other BEDTools Subcommands and pybedtools p. 336 Working with Sequence Data p. 339 The FASTA Format p. 339 The FASTQ Format p. 341 Nucleotide Codes p. 343 Base Qualities p. 344 Example: Inspecting and Trimming Low-Quality Bases p. 346 A FASTA/FASTQ Parsing Example-. Counting Nucleotides p. 349 Indexed FASTA Files p. 352 Working with Alignment Data p. 355 Getting to Know Alignment Formats: SAM and BAM p. 356 The SAM Header p. 356 The SAM Alignment Section p. 359 Bitwise Flags p. 360 CIGAR Strings p. 363 Mapping Qualities p. 365 Command-Line Tools for Working with Alignments in the SAM Format p. 365 Using samtools view to Convert between SAM and BAM p. 365 Samtools Sort and Index p. 367 Extracting and Filtering Alignments with samtools view p. 368 Visualizing Alignments with samtools tview and the Integrated Genomics Viewer p. 372 Pileups with samtools pileup, Variant Calling, and Base Alignment Quality p. 378 Creating Your Own SAM/BAM Processing Tools with Pysam p. 384 Opening BAM Files, Fetching Alignments from a Region, and Iterating Across Reads p. 384 Extracting SAM/BAM Header Information from an AlignmentFile Object p. 387 Working with AlignedSegment Objects p. 388 Writing a Program to Record Alignment Statistics p. 391 Additional Pysam Features and Other SAM/BAM APIs p. 394 Bioinformatics Shell Scripting, Writing Pipelines, and Parallelizing Tasks p. 395 Basic Bash Scripting p. 396 Writing and Running Robust Bash Scripts p. 396
6 Variables and Command Arguments p. 398 Conditionals in a Bash Script: if Statements p. 401 Processing Files with Bash Using for Loops and Globbing p. 405 Automating File-Processing with find and xargs p. 411 Using find and xargs p. 411 Finding Files with find p. 412 Find's Expressions p. 413 Find's -exec: Running Commands on finds Results p. 415 Xargs: A Unix Powertool p. 416 Using xargs with Replacement Strings to Apply Commands to Files p. 418 Xargs and Parallelization p. 419 Make and Makefiles: Another Option for Pipelines p. 421 Out-of-Memory Approaches: Tabix and SQLite p. 425 Fast Access to Indexed Tab-Delimited Files with BGZF and Tabix p. 425 Compressing Files for Tabix with Bgzip p. 426 Indexing Files with Tabix p. 427 Using Tabix p. 427 Introducing Relational Databases Through SQLite p. 428 When to Use Relational Databases in Bioinformatics p. 429 Installing SQLite p. 431 Exploring SQLite Databases with the Command-Line Interface p. 431 Querying Out Data: The Almighty SELECT Command p. 434 SQLite Functions p. 441 SQLite Aggregate Functions p. 442 Subqueries p. 447 Organizing Relational Databases and Joins p. 448 Writing to Databases p. 455 Dropping Tables and Deleting Databases p. 458 Interacting with SQLite from Python p. 459 Dumping Databases p. 465 Conclusion p. 467 Where to Go From Here? p. 468 Glossary p. 471 Bibliography p. 479 Index p. 483 Table of Contents provided by Blackwell's Book Services and R.R. Bowker. Used with permission.
Lecture 3. Essential skills for bioinformatics: Unix/Linux
 Lecture 3 Essential skills for bioinformatics: Unix/Linux RETRIEVING DATA Overview Whether downloading large sequencing datasets or accessing a web application hundreds of times to download specific files,
Lecture 3 Essential skills for bioinformatics: Unix/Linux RETRIEVING DATA Overview Whether downloading large sequencing datasets or accessing a web application hundreds of times to download specific files,
Essential Skills for Bioinformatics: Unix/Linux
 Essential Skills for Bioinformatics: Unix/Linux WORKING WITH COMPRESSED DATA Overview Data compression, the process of condensing data so that it takes up less space (on disk drives, in memory, or across
Essential Skills for Bioinformatics: Unix/Linux WORKING WITH COMPRESSED DATA Overview Data compression, the process of condensing data so that it takes up less space (on disk drives, in memory, or across
Programming for Data Science Syllabus
 Programming for Data Science Syllabus Learn to use Python and SQL to solve problems with data Before You Start Prerequisites: There are no prerequisites for this program, aside from basic computer skills.
Programming for Data Science Syllabus Learn to use Python and SQL to solve problems with data Before You Start Prerequisites: There are no prerequisites for this program, aside from basic computer skills.
Recap From Last Time:
 Recap From Last Time: BGGN 213 Working with UNIX Barry Grant http://thegrantlab.org/bggn213 Motivation: Why we use UNIX for bioinformatics. Modularity, Programmability, Infrastructure, Reliability and
Recap From Last Time: BGGN 213 Working with UNIX Barry Grant http://thegrantlab.org/bggn213 Motivation: Why we use UNIX for bioinformatics. Modularity, Programmability, Infrastructure, Reliability and
BGGN 213 Working with UNIX Barry Grant
 BGGN 213 Working with UNIX Barry Grant http://thegrantlab.org/bggn213 Recap From Last Time: Motivation: Why we use UNIX for bioinformatics. Modularity, Programmability, Infrastructure, Reliability and
BGGN 213 Working with UNIX Barry Grant http://thegrantlab.org/bggn213 Recap From Last Time: Motivation: Why we use UNIX for bioinformatics. Modularity, Programmability, Infrastructure, Reliability and
Recap From Last Time: Setup Checklist BGGN 213. Todays Menu. Introduction to UNIX.
 Recap From Last Time: BGGN 213 Introduction to UNIX Barry Grant http://thegrantlab.org/bggn213 Substitution matrices: Where our alignment match and mis-match scores typically come from Comparing methods:
Recap From Last Time: BGGN 213 Introduction to UNIX Barry Grant http://thegrantlab.org/bggn213 Substitution matrices: Where our alignment match and mis-match scores typically come from Comparing methods:
Practical Linux examples: Exercises
 Practical Linux examples: Exercises 1. Login (ssh) to the machine that you are assigned for this workshop (assigned machines: https://cbsu.tc.cornell.edu/ww/machines.aspx?i=87 ). Prepare working directory,
Practical Linux examples: Exercises 1. Login (ssh) to the machine that you are assigned for this workshop (assigned machines: https://cbsu.tc.cornell.edu/ww/machines.aspx?i=87 ). Prepare working directory,
Introduction p. 1 Who Should Read This Book? p. 1 What You Need to Know Before Reading This Book p. 2 How This Book Is Organized p.
 Introduction p. 1 Who Should Read This Book? p. 1 What You Need to Know Before Reading This Book p. 2 How This Book Is Organized p. 2 Conventions Used in This Book p. 2 Introduction to UNIX p. 5 An Overview
Introduction p. 1 Who Should Read This Book? p. 1 What You Need to Know Before Reading This Book p. 2 How This Book Is Organized p. 2 Conventions Used in This Book p. 2 Introduction to UNIX p. 5 An Overview
Git. Charles J. Geyer School of Statistics University of Minnesota. Stat 8054 Lecture Notes
 Git Charles J. Geyer School of Statistics University of Minnesota Stat 8054 Lecture Notes 1 Before Anything Else Tell git who you are. git config --global user.name "Charles J. Geyer" git config --global
Git Charles J. Geyer School of Statistics University of Minnesota Stat 8054 Lecture Notes 1 Before Anything Else Tell git who you are. git config --global user.name "Charles J. Geyer" git config --global
Introduction To. Barry Grant
 Introduction To Barry Grant bjgrant@umich.edu http://thegrantlab.org Working with Unix How do we actually use Unix? Inspecting text files less - visualize a text file: use arrow keys page down/page up
Introduction To Barry Grant bjgrant@umich.edu http://thegrantlab.org Working with Unix How do we actually use Unix? Inspecting text files less - visualize a text file: use arrow keys page down/page up
Contents. Note: pay attention to where you are. Note: Plaintext version. Note: pay attention to where you are... 1 Note: Plaintext version...
 Contents Note: pay attention to where you are........................................... 1 Note: Plaintext version................................................... 1 Hello World of the Bash shell 2 Accessing
Contents Note: pay attention to where you are........................................... 1 Note: Plaintext version................................................... 1 Hello World of the Bash shell 2 Accessing
Lecture 5. Essential skills for bioinformatics: Unix/Linux
 Lecture 5 Essential skills for bioinformatics: Unix/Linux UNIX DATA TOOLS Text processing with awk We have illustrated two ways awk can come in handy: Filtering data using rules that can combine regular
Lecture 5 Essential skills for bioinformatics: Unix/Linux UNIX DATA TOOLS Text processing with awk We have illustrated two ways awk can come in handy: Filtering data using rules that can combine regular
Introduction to UNIX command-line II
 Introduction to UNIX command-line II Boyce Thompson Institute 2017 Prashant Hosmani Class Content Terminal file system navigation Wildcards, shortcuts and special characters File permissions Compression
Introduction to UNIX command-line II Boyce Thompson Institute 2017 Prashant Hosmani Class Content Terminal file system navigation Wildcards, shortcuts and special characters File permissions Compression
Genomic Files. University of Massachusetts Medical School. October, 2014
 .. Genomic Files University of Massachusetts Medical School October, 2014 2 / 39. A Typical Deep-Sequencing Workflow Samples Fastq Files Fastq Files Sam / Bam Files Various files Deep Sequencing Further
.. Genomic Files University of Massachusetts Medical School October, 2014 2 / 39. A Typical Deep-Sequencing Workflow Samples Fastq Files Fastq Files Sam / Bam Files Various files Deep Sequencing Further
Lecture 8. Sequence alignments
 Lecture 8 Sequence alignments DATA FORMATS bioawk bioawk is a program that extends awk s powerful processing of tabular data to processing tasks involving common bioinformatics formats like FASTA/FASTQ,
Lecture 8 Sequence alignments DATA FORMATS bioawk bioawk is a program that extends awk s powerful processing of tabular data to processing tasks involving common bioinformatics formats like FASTA/FASTQ,
Genomic Files. University of Massachusetts Medical School. October, 2015
 .. Genomic Files University of Massachusetts Medical School October, 2015 2 / 55. A Typical Deep-Sequencing Workflow Samples Fastq Files Fastq Files Sam / Bam Files Various files Deep Sequencing Further
.. Genomic Files University of Massachusetts Medical School October, 2015 2 / 55. A Typical Deep-Sequencing Workflow Samples Fastq Files Fastq Files Sam / Bam Files Various files Deep Sequencing Further
Introduction To. Barry Grant
 Introduction To Barry Grant bjgrant@umich.edu http://thegrantlab.org Introduction to Biocomputing http://bioboot.github.io/web-2016/ Monday Tuesday Wednesday Thursday Friday Introduction to UNIX* Introduction
Introduction To Barry Grant bjgrant@umich.edu http://thegrantlab.org Introduction to Biocomputing http://bioboot.github.io/web-2016/ Monday Tuesday Wednesday Thursday Friday Introduction to UNIX* Introduction
SAM / BAM Tutorial. EMBL Heidelberg. Course Materials. Tobias Rausch September 2012
 SAM / BAM Tutorial EMBL Heidelberg Course Materials Tobias Rausch September 2012 Contents 1 SAM / BAM 3 1.1 Introduction................................... 3 1.2 Tasks.......................................
SAM / BAM Tutorial EMBL Heidelberg Course Materials Tobias Rausch September 2012 Contents 1 SAM / BAM 3 1.1 Introduction................................... 3 1.2 Tasks.......................................
BASH SHELL SCRIPT 1- Introduction to Shell
 BASH SHELL SCRIPT 1- Introduction to Shell What is shell Installation of shell Shell features Bash Keywords Built-in Commands Linux Commands Specialized Navigation and History Commands Shell Aliases Bash
BASH SHELL SCRIPT 1- Introduction to Shell What is shell Installation of shell Shell features Bash Keywords Built-in Commands Linux Commands Specialized Navigation and History Commands Shell Aliases Bash
Linux Fundamentals (L-120)
 Linux Fundamentals (L-120) Modality: Virtual Classroom Duration: 5 Days SUBSCRIPTION: Master, Master Plus About this course: This is a challenging course that focuses on the fundamental tools and concepts
Linux Fundamentals (L-120) Modality: Virtual Classroom Duration: 5 Days SUBSCRIPTION: Master, Master Plus About this course: This is a challenging course that focuses on the fundamental tools and concepts
Practical Linux Examples
 Practical Linux Examples Processing large text file Parallelization of independent tasks Qi Sun & Robert Bukowski Bioinformatics Facility Cornell University http://cbsu.tc.cornell.edu/lab/doc/linux_examples_slides.pdf
Practical Linux Examples Processing large text file Parallelization of independent tasks Qi Sun & Robert Bukowski Bioinformatics Facility Cornell University http://cbsu.tc.cornell.edu/lab/doc/linux_examples_slides.pdf
Outline The three W s Overview of gits structure Using git Final stuff. Git. A fast distributed revision control system
 Git A fast distributed revision control system Nils Moschüring PhD Student (LMU) 1 The three W s What? Why? Workflow and nomenclature 2 Overview of gits structure Structure Branches 3 Using git Setting
Git A fast distributed revision control system Nils Moschüring PhD Student (LMU) 1 The three W s What? Why? Workflow and nomenclature 2 Overview of gits structure Structure Branches 3 Using git Setting
Lecture 12. Short read aligners
 Lecture 12 Short read aligners Ebola reference genome We will align ebola sequencing data against the 1976 Mayinga reference genome. We will hold the reference gnome and all indices: mkdir -p ~/reference/ebola
Lecture 12 Short read aligners Ebola reference genome We will align ebola sequencing data against the 1976 Mayinga reference genome. We will hold the reference gnome and all indices: mkdir -p ~/reference/ebola
Table of contents. Our goal. Notes. Notes. Notes. Summer June 29, Our goal is to see how we can use Unix as a tool for developing programs
 Summer 2010 Department of Computer Science and Engineering York University Toronto June 29, 2010 1 / 36 Table of contents 1 2 3 4 2 / 36 Our goal Our goal is to see how we can use Unix as a tool for developing
Summer 2010 Department of Computer Science and Engineering York University Toronto June 29, 2010 1 / 36 Table of contents 1 2 3 4 2 / 36 Our goal Our goal is to see how we can use Unix as a tool for developing
git Version: 2.0b Merge combines trees, and checks out the result Pull does a fetch, then a merge If you only can remember one command:
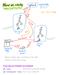 Merge combines trees, and checks out the result Pull does a fetch, then a merge If you only can remember one command: git --help Get common commands and help git --help How to use git
Merge combines trees, and checks out the result Pull does a fetch, then a merge If you only can remember one command: git --help Get common commands and help git --help How to use git
Git. A fast distributed revision control system. Nils Moschüring PhD Student (LMU)
 Git A fast distributed revision control system Nils Moschüring PhD Student (LMU) Nils Moschüring PhD Student (LMU), Git 1 1 The three W s What? Why? Workflow and nomenclature 2 Overview of gits structure
Git A fast distributed revision control system Nils Moschüring PhD Student (LMU) Nils Moschüring PhD Student (LMU), Git 1 1 The three W s What? Why? Workflow and nomenclature 2 Overview of gits structure
Git. Presenter: Haotao (Eric) Lai Contact:
 Git Presenter: Haotao (Eric) Lai Contact: haotao.lai@gmail.com 1 Acknowledge images with white background is from the following link: http://marklodato.github.io/visual-git-guide/index-en.html images with
Git Presenter: Haotao (Eric) Lai Contact: haotao.lai@gmail.com 1 Acknowledge images with white background is from the following link: http://marklodato.github.io/visual-git-guide/index-en.html images with
Useful Unix Commands Cheat Sheet
 Useful Unix Commands Cheat Sheet The Chinese University of Hong Kong SIGSC Training (Fall 2016) FILE AND DIRECTORY pwd Return path to current directory. ls List directories and files here. ls dir List
Useful Unix Commands Cheat Sheet The Chinese University of Hong Kong SIGSC Training (Fall 2016) FILE AND DIRECTORY pwd Return path to current directory. ls List directories and files here. ls dir List
Essential Skills for Bioinformatics: Unix/Linux
 Essential Skills for Bioinformatics: Unix/Linux SHELL SCRIPTING Overview Bash, the shell we have used interactively in this course, is a full-fledged scripting language. Unlike Python, Bash is not a general-purpose
Essential Skills for Bioinformatics: Unix/Linux SHELL SCRIPTING Overview Bash, the shell we have used interactively in this course, is a full-fledged scripting language. Unlike Python, Bash is not a general-purpose
About SJTUG. SJTU *nix User Group SJTU Joyful Techie User Group
 About SJTUG SJTU *nix User Group SJTU Joyful Techie User Group Homepage - https://sjtug.org/ SJTUG Mirrors - https://mirrors.sjtug.sjtu.edu.cn/ GitHub - https://github.com/sjtug Git Basic Tutorial Zhou
About SJTUG SJTU *nix User Group SJTU Joyful Techie User Group Homepage - https://sjtug.org/ SJTUG Mirrors - https://mirrors.sjtug.sjtu.edu.cn/ GitHub - https://github.com/sjtug Git Basic Tutorial Zhou
Managing Projects with Git
 Managing Projects with Git (and other command-line skills) Dr. Chris Mayfield Department of Computer Science James Madison University Feb 09, 2018 Part 1: Command Line Review as needed YouTube video tutorials
Managing Projects with Git (and other command-line skills) Dr. Chris Mayfield Department of Computer Science James Madison University Feb 09, 2018 Part 1: Command Line Review as needed YouTube video tutorials
An Introduction to Linux and Bowtie
 An Introduction to Linux and Bowtie Cavan Reilly November 10, 2017 Table of contents Introduction to UNIX-like operating systems Installing programs Bowtie SAMtools Introduction to Linux In order to use
An Introduction to Linux and Bowtie Cavan Reilly November 10, 2017 Table of contents Introduction to UNIX-like operating systems Installing programs Bowtie SAMtools Introduction to Linux In order to use
CS197U: A Hands on Introduction to Unix
 CS197U: A Hands on Introduction to Unix Lecture 11: WWW and Wrap up Tian Guo University of Massachusetts Amherst CICS 1 Reminders Assignment 4 was graded and scores on Moodle Assignment 5 was due and you
CS197U: A Hands on Introduction to Unix Lecture 11: WWW and Wrap up Tian Guo University of Massachusetts Amherst CICS 1 Reminders Assignment 4 was graded and scores on Moodle Assignment 5 was due and you
Section 1: Tools. Kaifei Chen, Luca Zuccarini. January 23, Make Motivation How... 2
 Kaifei Chen, Luca Zuccarini January 23, 2015 Contents 1 Make 2 1.1 Motivation............................................ 2 1.2 How................................................ 2 2 Git 2 2.1 Learn by
Kaifei Chen, Luca Zuccarini January 23, 2015 Contents 1 Make 2 1.1 Motivation............................................ 2 1.2 How................................................ 2 2 Git 2 2.1 Learn by
ls /data/atrnaseq/ egrep "(fastq fasta fq fa)\.gz" ls /data/atrnaseq/ egrep "(cn ts)[1-3]ln[^3a-za-z]\."
![ls /data/atrnaseq/ egrep (fastq fasta fq fa)\.gz ls /data/atrnaseq/ egrep (cn ts)[1-3]ln[^3a-za-z]\. ls /data/atrnaseq/ egrep (fastq fasta fq fa)\.gz ls /data/atrnaseq/ egrep (cn ts)[1-3]ln[^3a-za-z]\.](/thumbs/73/69027269.jpg) Command line tools - bash, awk and sed We can only explore a small fraction of the capabilities of the bash shell and command-line utilities in Linux during this course. An entire course could be taught
Command line tools - bash, awk and sed We can only explore a small fraction of the capabilities of the bash shell and command-line utilities in Linux during this course. An entire course could be taught
Useful software utilities for computational genomics. Shamith Samarajiwa CRUK Autumn School in Bioinformatics September 2017
 Useful software utilities for computational genomics Shamith Samarajiwa CRUK Autumn School in Bioinformatics September 2017 Overview Search and download genomic datasets: GEOquery, GEOsearch and GEOmetadb,
Useful software utilities for computational genomics Shamith Samarajiwa CRUK Autumn School in Bioinformatics September 2017 Overview Search and download genomic datasets: GEOquery, GEOsearch and GEOmetadb,
Preparation of alignments for variant calling with GATK: exercise instructions for BioHPC Lab computers
 Preparation of alignments for variant calling with GATK: exercise instructions for BioHPC Lab computers Data used in the exercise We will use D. melanogaster WGS paired-end Illumina data with NCBI accessions
Preparation of alignments for variant calling with GATK: exercise instructions for BioHPC Lab computers Data used in the exercise We will use D. melanogaster WGS paired-end Illumina data with NCBI accessions
Git, the magical version control
 Git, the magical version control Git is an open-source version control system (meaning, it s free!) that allows developers to track changes made on their code files throughout the lifetime of a project.
Git, the magical version control Git is an open-source version control system (meaning, it s free!) that allows developers to track changes made on their code files throughout the lifetime of a project.
Version Control with Git
 Version Control with Git Methods & Tools for Software Engineering (MTSE) Fall 2017 Prof. Arie Gurfinkel based on https://git-scm.com/book What is Version (Revision) Control A system for managing changes
Version Control with Git Methods & Tools for Software Engineering (MTSE) Fall 2017 Prof. Arie Gurfinkel based on https://git-scm.com/book What is Version (Revision) Control A system for managing changes
LINUX FUNDAMENTALS (5 Day)
 www.peaklearningllc.com LINUX FUNDAMENTALS (5 Day) Designed to provide the essential skills needed to be proficient at the Unix or Linux command line. This challenging course focuses on the fundamental
www.peaklearningllc.com LINUX FUNDAMENTALS (5 Day) Designed to provide the essential skills needed to be proficient at the Unix or Linux command line. This challenging course focuses on the fundamental
What is git? Distributed Version Control System (VCS); Created by Linus Torvalds, to help with Linux development;
 What is git? Distributed Version Control System (VCS); Created by Linus Torvalds, to help with Linux development; Why should I use a VCS? Repositories Types of repositories: Private - only you and the
What is git? Distributed Version Control System (VCS); Created by Linus Torvalds, to help with Linux development; Why should I use a VCS? Repositories Types of repositories: Private - only you and the
UNIX Shell Programming
 $!... 5:13 $$ and $!... 5:13.profile File... 7:4 /etc/bashrc... 10:13 /etc/profile... 10:12 /etc/profile File... 7:5 ~/.bash_login... 10:15 ~/.bash_logout... 10:18 ~/.bash_profile... 10:14 ~/.bashrc...
$!... 5:13 $$ and $!... 5:13.profile File... 7:4 /etc/bashrc... 10:13 /etc/profile... 10:12 /etc/profile File... 7:5 ~/.bash_login... 10:15 ~/.bash_logout... 10:18 ~/.bash_profile... 10:14 ~/.bashrc...
The software and data for the RNA-Seq exercise are already available on the USB system
 BIT815 Notes on R analysis of RNA-seq data The software and data for the RNA-Seq exercise are already available on the USB system The notes below regarding installation of R packages and other software
BIT815 Notes on R analysis of RNA-seq data The software and data for the RNA-Seq exercise are already available on the USB system The notes below regarding installation of R packages and other software
Variant calling using SAMtools
 Variant calling using SAMtools Calling variants - a trivial use of an Interactive Session We are going to conduct the variant calling exercises in an interactive idev session just so you can get a feel
Variant calling using SAMtools Calling variants - a trivial use of an Interactive Session We are going to conduct the variant calling exercises in an interactive idev session just so you can get a feel
Version Control with GIT
 Version Control with GIT Benjamin Roth CIS LMU München Benjamin Roth (CIS LMU München) Version Control with GIT 1 / 30 Version Control Version control [...] is the management of changes to documents, computer
Version Control with GIT Benjamin Roth CIS LMU München Benjamin Roth (CIS LMU München) Version Control with GIT 1 / 30 Version Control Version control [...] is the management of changes to documents, computer
HIPPIE User Manual. (v0.0.2-beta, 2015/4/26, Yih-Chii Hwang, yihhwang [at] mail.med.upenn.edu)
![HIPPIE User Manual. (v0.0.2-beta, 2015/4/26, Yih-Chii Hwang, yihhwang [at] mail.med.upenn.edu) HIPPIE User Manual. (v0.0.2-beta, 2015/4/26, Yih-Chii Hwang, yihhwang [at] mail.med.upenn.edu)](/thumbs/71/65752105.jpg) HIPPIE User Manual (v0.0.2-beta, 2015/4/26, Yih-Chii Hwang, yihhwang [at] mail.med.upenn.edu) OVERVIEW OF HIPPIE o Flowchart of HIPPIE o Requirements PREPARE DIRECTORY STRUCTURE FOR HIPPIE EXECUTION o
HIPPIE User Manual (v0.0.2-beta, 2015/4/26, Yih-Chii Hwang, yihhwang [at] mail.med.upenn.edu) OVERVIEW OF HIPPIE o Flowchart of HIPPIE o Requirements PREPARE DIRECTORY STRUCTURE FOR HIPPIE EXECUTION o
Version control with git and Rstudio. Remko Duursma
 Version control with git and Rstudio Remko Duursma November 14, 2017 Contents 1 Version control with git 2 1.1 Should I learn version control?...................................... 2 1.2 Basics of git..................................................
Version control with git and Rstudio Remko Duursma November 14, 2017 Contents 1 Version control with git 2 1.1 Should I learn version control?...................................... 2 1.2 Basics of git..................................................
Contents. xxvii. Preface
 Preface xxvii Chapter 1: Welcome to Linux 1 The GNU Linux Connection 2 The History of GNU Linux 2 The Code Is Free 4 Have Fun! 5 The Heritage of Linux: UNIX 5 What Is So Good About Linux? 6 Why Linux Is
Preface xxvii Chapter 1: Welcome to Linux 1 The GNU Linux Connection 2 The History of GNU Linux 2 The Code Is Free 4 Have Fun! 5 The Heritage of Linux: UNIX 5 What Is So Good About Linux? 6 Why Linux Is
CSE 15L Winter Midterm :) Review
 CSE 15L Winter 2015 Midterm :) Review Makefiles Makefiles - The Overview Questions you should be able to answer What is the point of a Makefile Why don t we just compile it again? Why don t we just use
CSE 15L Winter 2015 Midterm :) Review Makefiles Makefiles - The Overview Questions you should be able to answer What is the point of a Makefile Why don t we just compile it again? Why don t we just use
Index. Alias syntax, 31 Author and commit attributes, 334
 Index A Alias syntax, 31 Author and commit attributes, 334 B Bare repository, 19 Binary conflict creating conflicting changes, 218 during merging, 219 during rebasing, 221 Branches backup, 140 clone-with-branches
Index A Alias syntax, 31 Author and commit attributes, 334 B Bare repository, 19 Binary conflict creating conflicting changes, 218 during merging, 219 during rebasing, 221 Branches backup, 140 clone-with-branches
Versioning with git. Moritz August Git/Bash/Python-Course for MPE. Moritz August Versioning with Git
 Versioning with git Moritz August 13.03.2017 Git/Bash/Python-Course for MPE 1 Agenda What s git and why is it good? The general concept of git It s a graph! What is a commit? The different levels Remote
Versioning with git Moritz August 13.03.2017 Git/Bash/Python-Course for MPE 1 Agenda What s git and why is it good? The general concept of git It s a graph! What is a commit? The different levels Remote
Section 1: Tools. Contents CS162. January 19, Make More details about Make Git Commands to know... 3
 CS162 January 19, 2017 Contents 1 Make 2 1.1 More details about Make.................................... 2 2 Git 3 2.1 Commands to know....................................... 3 3 GDB: The GNU Debugger
CS162 January 19, 2017 Contents 1 Make 2 1.1 More details about Make.................................... 2 2 Git 3 2.1 Commands to know....................................... 3 3 GDB: The GNU Debugger
Unix as a Platform Exercises. Course Code: OS-01-UNXPLAT
 Unix as a Platform Exercises Course Code: OS-01-UNXPLAT Working with Unix 1. Use the on-line manual page to determine the option for cat, which causes nonprintable characters to be displayed. Run the command
Unix as a Platform Exercises Course Code: OS-01-UNXPLAT Working with Unix 1. Use the on-line manual page to determine the option for cat, which causes nonprintable characters to be displayed. Run the command
Reproducible Research with R and RStudio
 The R Series Reproducible Research with R and RStudio Christopher Gandrud C\ CRC Press cj* Taylor & Francis Croup Boca Raton London New York CRC Press is an imprint of the Taylor & Francis Group an informa
The R Series Reproducible Research with R and RStudio Christopher Gandrud C\ CRC Press cj* Taylor & Francis Croup Boca Raton London New York CRC Press is an imprint of the Taylor & Francis Group an informa
Welcome to MAPHiTS (Mapping Analysis Pipeline for High-Throughput Sequences) tutorial page.
 Welcome to MAPHiTS (Mapping Analysis Pipeline for High-Throughput Sequences) tutorial page. In this page you will learn to use the tools of the MAPHiTS suite. A little advice before starting : rename your
Welcome to MAPHiTS (Mapping Analysis Pipeline for High-Throughput Sequences) tutorial page. In this page you will learn to use the tools of the MAPHiTS suite. A little advice before starting : rename your
The student will have the essential skills needed to be proficient at the Unix or Linux command line.
 Table of Contents Introduction Audience At Course Completion Prerequisites Certified Professional Exams Student Materials Course Outline Introduction This challenging course focuses on the fundamental
Table of Contents Introduction Audience At Course Completion Prerequisites Certified Professional Exams Student Materials Course Outline Introduction This challenging course focuses on the fundamental
TangeloHub Documentation
 TangeloHub Documentation Release None Kitware, Inc. September 21, 2015 Contents 1 User s Guide 3 1.1 Managing Data.............................................. 3 1.2 Running an Analysis...........................................
TangeloHub Documentation Release None Kitware, Inc. September 21, 2015 Contents 1 User s Guide 3 1.1 Managing Data.............................................. 3 1.2 Running an Analysis...........................................
API RI. Application Programming Interface Reference Implementation. Policies and Procedures Discussion
 API Working Group Meeting, Harris County, TX March 22-23, 2016 Policies and Procedures Discussion Developing a Mission Statement What do we do? How do we do it? Whom do we do it for? What value are we
API Working Group Meeting, Harris County, TX March 22-23, 2016 Policies and Procedures Discussion Developing a Mission Statement What do we do? How do we do it? Whom do we do it for? What value are we
EECS 470 Lab 5. Linux Shell Scripting. Friday, 1 st February, 2018
 EECS 470 Lab 5 Linux Shell Scripting Department of Electrical Engineering and Computer Science College of Engineering University of Michigan Friday, 1 st February, 2018 (University of Michigan) Lab 5:
EECS 470 Lab 5 Linux Shell Scripting Department of Electrical Engineering and Computer Science College of Engineering University of Michigan Friday, 1 st February, 2018 (University of Michigan) Lab 5:
Sequence Analysis Pipeline
 Sequence Analysis Pipeline Transcript fragments 1. PREPROCESSING 2. ASSEMBLY (today) Removal of contaminants, vector, adaptors, etc Put overlapping sequence together and calculate bigger sequences 3. Analysis/Annotation
Sequence Analysis Pipeline Transcript fragments 1. PREPROCESSING 2. ASSEMBLY (today) Removal of contaminants, vector, adaptors, etc Put overlapping sequence together and calculate bigger sequences 3. Analysis/Annotation
GIT. CS 490MT/5555, Spring 2017, Yongjie Zheng
 GIT CS 490MT/5555, Spring 2017, Yongjie Zheng GIT Overview GIT Basics Highlights: snapshot, the three states Working with the Private (Local) Repository Creating a repository and making changes to it Working
GIT CS 490MT/5555, Spring 2017, Yongjie Zheng GIT Overview GIT Basics Highlights: snapshot, the three states Working with the Private (Local) Repository Creating a repository and making changes to it Working
5/8/2012. Exploring Utilities Chapter 5
 Exploring Utilities Chapter 5 Examining the contents of files. Working with the cut and paste feature. Formatting output with the column utility. Searching for lines containing a target string with grep.
Exploring Utilities Chapter 5 Examining the contents of files. Working with the cut and paste feature. Formatting output with the column utility. Searching for lines containing a target string with grep.
Whole genome assembly comparison of duplication originally described in Bailey et al
 WGAC Whole genome assembly comparison of duplication originally described in Bailey et al. 2001. Inputs species name path to FASTA sequence(s) to be processed either a directory of chromosomal FASTA files
WGAC Whole genome assembly comparison of duplication originally described in Bailey et al. 2001. Inputs species name path to FASTA sequence(s) to be processed either a directory of chromosomal FASTA files
Handling important NGS data formats in UNIX Prac8cal training course NGS Workshop in Nove Hrady 2014
 Handling important NGS data formats in UNIX Prac8cal training course NGS Workshop in Nove Hrady 2014 Vaclav Janousek, Libor Morkovsky hjp://ngs- course- nhrady.readthedocs.org (Exercises & Reference Manual)
Handling important NGS data formats in UNIX Prac8cal training course NGS Workshop in Nove Hrady 2014 Vaclav Janousek, Libor Morkovsky hjp://ngs- course- nhrady.readthedocs.org (Exercises & Reference Manual)
Github/Git Primer. Tyler Hague
 Github/Git Primer Tyler Hague Why Use Github? Github keeps all of our code up to date in one place Github tracks changes so we can see what is being worked on Github has issue tracking for keeping up with
Github/Git Primer Tyler Hague Why Use Github? Github keeps all of our code up to date in one place Github tracks changes so we can see what is being worked on Github has issue tracking for keeping up with
Linux command line basics III: piping commands for text processing. Yanbin Yin Fall 2015
 Linux command line basics III: piping commands for text processing Yanbin Yin Fall 2015 1 h.p://korflab.ucdavis.edu/unix_and_perl/unix_and_perl_v3.1.1.pdf 2 The beauty of Unix for bioinformagcs sort, cut,
Linux command line basics III: piping commands for text processing Yanbin Yin Fall 2015 1 h.p://korflab.ucdavis.edu/unix_and_perl/unix_and_perl_v3.1.1.pdf 2 The beauty of Unix for bioinformagcs sort, cut,
Submitting your Work using GIT
 Submitting your Work using GIT You will be using the git distributed source control system in order to manage and submit your assignments. Why? allows you to take snapshots of your project at safe points
Submitting your Work using GIT You will be using the git distributed source control system in order to manage and submit your assignments. Why? allows you to take snapshots of your project at safe points
Handling sam and vcf data, quality control
 Handling sam and vcf data, quality control We continue with the earlier analyses and get some new data: cd ~/session_3 wget http://wasabiapp.org/vbox/data/session_4/file3.tgz tar xzf file3.tgz wget http://wasabiapp.org/vbox/data/session_4/file4.tgz
Handling sam and vcf data, quality control We continue with the earlier analyses and get some new data: cd ~/session_3 wget http://wasabiapp.org/vbox/data/session_4/file3.tgz tar xzf file3.tgz wget http://wasabiapp.org/vbox/data/session_4/file4.tgz
Introduction to distributed version control with git
 Institut für theoretische Physik TU Clausthal 04.03.2013 Inhalt 1 Basics Differences to Subversion Translation of commands 2 Config Create and clone States and workflow Remote repos Branching and merging
Institut für theoretische Physik TU Clausthal 04.03.2013 Inhalt 1 Basics Differences to Subversion Translation of commands 2 Config Create and clone States and workflow Remote repos Branching and merging
The UNIX Shells. Computer Center, CS, NCTU. How shell works. Unix shells. Fetch command Analyze Execute
 Shells The UNIX Shells How shell works Fetch command Analyze Execute Unix shells Shell Originator System Name Prompt Bourne Shell S. R. Bourne /bin/sh $ Csh Bill Joy /bin/csh % Tcsh Ken Greer /bin/tcsh
Shells The UNIX Shells How shell works Fetch command Analyze Execute Unix shells Shell Originator System Name Prompt Bourne Shell S. R. Bourne /bin/sh $ Csh Bill Joy /bin/csh % Tcsh Ken Greer /bin/tcsh
Git Tutorial. André Sailer. ILD Technical Meeting April 24, 2017 CERN-EP-LCD. ILD Technical Meeting, Apr 24, 2017 A. Sailer: Git Tutorial 1/36
 ILD Technical Meeting, Apr 24, 2017 A. Sailer: Git Tutorial 1/36 Git Tutorial André Sailer CERN-EP-LCD ILD Technical Meeting April 24, 2017 LD Technical Meeting, Apr 24, 2017 A. Sailer: Git Tutorial 2/36
ILD Technical Meeting, Apr 24, 2017 A. Sailer: Git Tutorial 1/36 Git Tutorial André Sailer CERN-EP-LCD ILD Technical Meeting April 24, 2017 LD Technical Meeting, Apr 24, 2017 A. Sailer: Git Tutorial 2/36
A Brief Introduction to the Linux Shell for Data Science
 A Brief Introduction to the Linux Shell for Data Science Aris Anagnostopoulos 1 Introduction Here we will see a brief introduction of the Linux command line or shell as it is called. Linux is a Unix-like
A Brief Introduction to the Linux Shell for Data Science Aris Anagnostopoulos 1 Introduction Here we will see a brief introduction of the Linux command line or shell as it is called. Linux is a Unix-like
SAM : Sequence Alignment/Map format. A TAB-delimited text format storing the alignment information. A header section is optional.
 Alignment of NGS reads, samtools and visualization Hands-on Software used in this practical BWA MEM : Burrows-Wheeler Aligner. A software package for mapping low-divergent sequences against a large reference
Alignment of NGS reads, samtools and visualization Hands-on Software used in this practical BWA MEM : Burrows-Wheeler Aligner. A software package for mapping low-divergent sequences against a large reference
GIT FOR SYSTEM ADMINS JUSTIN ELLIOTT PENN STATE UNIVERSITY
 GIT FOR SYSTEM ADMINS JUSTIN ELLIOTT PENN STATE UNIVERSITY 1 WHAT IS VERSION CONTROL? Management of changes to documents like source code, scripts, text files Provides the ability to check documents in
GIT FOR SYSTEM ADMINS JUSTIN ELLIOTT PENN STATE UNIVERSITY 1 WHAT IS VERSION CONTROL? Management of changes to documents like source code, scripts, text files Provides the ability to check documents in
Mastering Linux. Paul S. Wang. CRC Press. Taylor & Francis Group. Taylor & Francis Croup an informa business. A CHAPMAN St HALL BOOK
 Mastering Linux Paul S. Wang CRC Press Taylor & Francis Group Boca Raton London New York CRC Press is an Imprint of the Taylor & Francis Croup an informa business A CHAPMAN St HALL BOOK Contents Preface
Mastering Linux Paul S. Wang CRC Press Taylor & Francis Group Boca Raton London New York CRC Press is an Imprint of the Taylor & Francis Croup an informa business A CHAPMAN St HALL BOOK Contents Preface
Unix Essentials. BaRC Hot Topics Bioinformatics and Research Computing Whitehead Institute October 12 th
 Unix Essentials BaRC Hot Topics Bioinformatics and Research Computing Whitehead Institute October 12 th 2016 http://barc.wi.mit.edu/hot_topics/ 1 Outline Unix overview Logging in to tak Directory structure
Unix Essentials BaRC Hot Topics Bioinformatics and Research Computing Whitehead Institute October 12 th 2016 http://barc.wi.mit.edu/hot_topics/ 1 Outline Unix overview Logging in to tak Directory structure
Fundamentals of Git 1
 Fundamentals of Git 1 Outline History of Git Distributed V.S Centralized Version Control Getting started Branching and Merging Working with remote Summary 2 A Brief History of Git Linus uses BitKeeper
Fundamentals of Git 1 Outline History of Git Distributed V.S Centralized Version Control Getting started Branching and Merging Working with remote Summary 2 A Brief History of Git Linus uses BitKeeper
Shells and Shell Programming
 Shells and Shell Programming 1 Shells A shell is a command line interpreter that is the interface between the user and the OS. The shell: analyzes each command determines what actions are to be performed
Shells and Shell Programming 1 Shells A shell is a command line interpreter that is the interface between the user and the OS. The shell: analyzes each command determines what actions are to be performed
NGS Data Visualization and Exploration Using IGV
 1 What is Galaxy Galaxy for Bioinformaticians Galaxy for Experimental Biologists Using Galaxy for NGS Analysis NGS Data Visualization and Exploration Using IGV 2 What is Galaxy Galaxy for Bioinformaticians
1 What is Galaxy Galaxy for Bioinformaticians Galaxy for Experimental Biologists Using Galaxy for NGS Analysis NGS Data Visualization and Exploration Using IGV 2 What is Galaxy Galaxy for Bioinformaticians
replace my_user_id in the commands with your actual user ID
 Exercise 1. Alignment with TOPHAT Part 1. Prepare the working directory. 1. Find out the name of the computer that has been reserved for you (https://cbsu.tc.cornell.edu/ww/machines.aspx?i=57 ). Everyone
Exercise 1. Alignment with TOPHAT Part 1. Prepare the working directory. 1. Find out the name of the computer that has been reserved for you (https://cbsu.tc.cornell.edu/ww/machines.aspx?i=57 ). Everyone
Git better. Collaborative project management using Git and GitHub. Matteo Sostero March 13, Sant Anna School of Advanced Studies
 Git better Collaborative project management using Git and GitHub Matteo Sostero March 13, 2018 Sant Anna School of Advanced Studies Let s Git it done! These slides are a brief primer to Git, and how it
Git better Collaborative project management using Git and GitHub Matteo Sostero March 13, 2018 Sant Anna School of Advanced Studies Let s Git it done! These slides are a brief primer to Git, and how it
Shells and Shell Programming
 Shells and Shell Programming Shells A shell is a command line interpreter that is the interface between the user and the OS. The shell: analyzes each command determines what actions are to be performed
Shells and Shell Programming Shells A shell is a command line interpreter that is the interface between the user and the OS. The shell: analyzes each command determines what actions are to be performed
Lab 2: Linux/Unix shell
 Lab 2: Linux/Unix shell Comp Sci 1585 Data Structures Lab: Tools for Computer Scientists Outline 1 2 3 4 5 6 7 What is a shell? What is a shell? login is a program that logs users in to a computer. When
Lab 2: Linux/Unix shell Comp Sci 1585 Data Structures Lab: Tools for Computer Scientists Outline 1 2 3 4 5 6 7 What is a shell? What is a shell? login is a program that logs users in to a computer. When
Git GitHub & secrets
 Git&GitHub secrets git branch -D local-branch git push origin :local-branch Much like James Cameron s Avatar, Git doesn t make any goddamn sense This talk throws a ton of stuff at you This talk throws
Git&GitHub secrets git branch -D local-branch git push origin :local-branch Much like James Cameron s Avatar, Git doesn t make any goddamn sense This talk throws a ton of stuff at you This talk throws
Introduction to the shell Part II
 Introduction to the shell Part II Graham Markall http://www.doc.ic.ac.uk/~grm08 grm08@doc.ic.ac.uk Civil Engineering Tech Talks 16 th November, 1pm Last week Covered applications and Windows compatibility
Introduction to the shell Part II Graham Markall http://www.doc.ic.ac.uk/~grm08 grm08@doc.ic.ac.uk Civil Engineering Tech Talks 16 th November, 1pm Last week Covered applications and Windows compatibility
CS 246 Winter Tutorial 2
 CS 246 Winter 2016 - Tutorial 2 Detailed Version January 14, 2016 1 Summary git Stuff File Properties Regular Expressions Output Redirection Piping Commands Basic Scripting Persistent Data Password-less
CS 246 Winter 2016 - Tutorial 2 Detailed Version January 14, 2016 1 Summary git Stuff File Properties Regular Expressions Output Redirection Piping Commands Basic Scripting Persistent Data Password-less
Intro to Git. Getting started with Version Control. Murray Anderegg February 9, 2018
 Intro to Git Getting started with Version Control Murray Anderegg February 9, 2018 What is Version Control? * It provides one method for an entire team to use; everybody operates under the same 'ground
Intro to Git Getting started with Version Control Murray Anderegg February 9, 2018 What is Version Control? * It provides one method for an entire team to use; everybody operates under the same 'ground
 http://xkcd.com/208/ 1. Review of pipes 2. Regular expressions 3. sed 4. awk 5. Editing Files 6. Shell loops 7. Shell scripts cat seqs.fa >0! TGCAGGTATATCTATTAGCAGGTTTAATTTTGCCTGCACTTGGTTGGGTACATTATTTTAAGTGTATTTGACAAG!
http://xkcd.com/208/ 1. Review of pipes 2. Regular expressions 3. sed 4. awk 5. Editing Files 6. Shell loops 7. Shell scripts cat seqs.fa >0! TGCAGGTATATCTATTAGCAGGTTTAATTTTGCCTGCACTTGGTTGGGTACATTATTTTAAGTGTATTTGACAAG!
Version Control with GIT: an introduction
 Version Control with GIT: an introduction Muzzamil LUQMAN (L3i) and Antoine FALAIZE (LaSIE) 23/11/2017 LaSIE Seminar Université de La Rochelle Version Control with GIT: an introduction - Why Git? - What
Version Control with GIT: an introduction Muzzamil LUQMAN (L3i) and Antoine FALAIZE (LaSIE) 23/11/2017 LaSIE Seminar Université de La Rochelle Version Control with GIT: an introduction - Why Git? - What
Git for Subversion users
 Git for Subversion users Zend webinar, 23-02-2012 Stefan who? Stefan who? Freelancer: Ingewikkeld Stefan who? Freelancer: Ingewikkeld Symfony Community Manager Stefan who? Freelancer: Ingewikkeld Symfony
Git for Subversion users Zend webinar, 23-02-2012 Stefan who? Stefan who? Freelancer: Ingewikkeld Stefan who? Freelancer: Ingewikkeld Symfony Community Manager Stefan who? Freelancer: Ingewikkeld Symfony
2 Initialize a git repository on your machine, add a README file, commit and push
 BioHPC Git Training Demo Script First, ensure that git is installed on your machine, and you have configured an ssh key. See the main slides for instructions. To follow this demo script open a terminal
BioHPC Git Training Demo Script First, ensure that git is installed on your machine, and you have configured an ssh key. See the main slides for instructions. To follow this demo script open a terminal
Review of Fundamentals
 Review of Fundamentals 1 The shell vi General shell review 2 http://teaching.idallen.com/cst8207/14f/notes/120_shell_basics.html The shell is a program that is executed for us automatically when we log
Review of Fundamentals 1 The shell vi General shell review 2 http://teaching.idallen.com/cst8207/14f/notes/120_shell_basics.html The shell is a program that is executed for us automatically when we log
Introduction to UNIX command-line
 Introduction to UNIX command-line Boyce Thompson Institute March 17, 2015 Lukas Mueller & Noe Fernandez Class Content Terminal file system navigation Wildcards, shortcuts and special characters File permissions
Introduction to UNIX command-line Boyce Thompson Institute March 17, 2015 Lukas Mueller & Noe Fernandez Class Content Terminal file system navigation Wildcards, shortcuts and special characters File permissions
Git AN INTRODUCTION. Introduction to Git as a version control system: concepts, main features and practical aspects.
 Git AN INTRODUCTION Introduction to Git as a version control system: concepts, main features and practical aspects. How do you share and save data? I m working solo and I only have one computer What I
Git AN INTRODUCTION Introduction to Git as a version control system: concepts, main features and practical aspects. How do you share and save data? I m working solo and I only have one computer What I
High-throughput sequencing: Alignment and related topic. Simon Anders EMBL Heidelberg
 High-throughput sequencing: Alignment and related topic Simon Anders EMBL Heidelberg Established platforms HTS Platforms Illumina HiSeq, ABI SOLiD, Roche 454 Newcomers: Benchtop machines 454 GS Junior,
High-throughput sequencing: Alignment and related topic Simon Anders EMBL Heidelberg Established platforms HTS Platforms Illumina HiSeq, ABI SOLiD, Roche 454 Newcomers: Benchtop machines 454 GS Junior,
Git Introduction CS 400. February 11, 2018
 Git Introduction CS 400 February 11, 2018 1 Introduction Git is one of the most popular version control system. It is a mature, actively maintained open source project originally developed in 2005 by Linus
Git Introduction CS 400 February 11, 2018 1 Introduction Git is one of the most popular version control system. It is a mature, actively maintained open source project originally developed in 2005 by Linus
git commit --amend git rebase <base> git reflog git checkout -b Create and check out a new branch named <branch>. Drop the -b
 Git Cheat Sheet Git Basics Rewriting Git History git init Create empty Git repo in specified directory. Run with no arguments to initialize the current directory as a git repository. git commit
Git Cheat Sheet Git Basics Rewriting Git History git init Create empty Git repo in specified directory. Run with no arguments to initialize the current directory as a git repository. git commit
CSE 391 Lecture 9. Version control with Git
 CSE 391 Lecture 9 Version control with Git slides created by Ruth Anderson & Marty Stepp, images from http://git-scm.com/book/en/ http://www.cs.washington.edu/391/ 1 Problems Working Alone Ever done one
CSE 391 Lecture 9 Version control with Git slides created by Ruth Anderson & Marty Stepp, images from http://git-scm.com/book/en/ http://www.cs.washington.edu/391/ 1 Problems Working Alone Ever done one
Windows. Everywhere else
 Git version control Enable native scrolling Git is a tool to manage sourcecode Never lose your coding progress again An empty folder 1/30 Windows Go to your programs overview and start Git Bash Everywhere
Git version control Enable native scrolling Git is a tool to manage sourcecode Never lose your coding progress again An empty folder 1/30 Windows Go to your programs overview and start Git Bash Everywhere
6 Git & Modularization
 6 Git & Modularization Bálint Aradi Course: Scientific Programming / Wissenchaftliches Programmieren (Python) Prerequisites Additional programs needed: Spyder3, Pylint3 Git, Gitk KDiff3 (non-kde (qt-only)
6 Git & Modularization Bálint Aradi Course: Scientific Programming / Wissenchaftliches Programmieren (Python) Prerequisites Additional programs needed: Spyder3, Pylint3 Git, Gitk KDiff3 (non-kde (qt-only)
