Segmentation Of Bone And Soft Tissue Regions In Digital Radiographic Images Of Extremities
|
|
|
- Anabel Higgins
- 5 years ago
- Views:
Transcription
1 Segmentation Of Bone And Soft Tissue Regions In Digital Radiographic Images Of Extremities S. Kubilay Pakina, Roger S. Gaborskib, Lori L. Barskic, Dave H. Foosd, and Kevin J. Parkere auniversity of Rochester, Electrical and Computer Engineering, 204 Hopeman Bldg., Rochester NY USA brochester Institute of Technology, Rochester NY USA c,deastman Kodak Co., Rochester NY USA euniversity of Rochester, Electrical and Computer Engineering, Rochester NY USA ABSTRACT This paper presents an algorithm for segmentation of computed radiography (CR) images of extremities into bone and soft tissue regions. The algorithm is a region-based one in which the regions are constructed using a growing procedure with two different statistical tests. Following the growing process, tissue classification procedure is employed. The purpose of the classification is to label each region as either bone or soft tissue. This binary classification goal is achieved by using a voting procedure that consists of clustering of regions in each neighborhood system into two classes. The voting procedure provides a crucial compromise between local and global analysis of the image, which is necessary due to strong exposure variations seen on the imaging plate. Also, the existence of regions whose size is large enough such that exposure variations can be observed through them makes it necessary to use overlapping blocks during the classification. After the classification step, resulting bone and soft tissue regions are refined by fitting a 2nd order surface to each tissue, and reevaluating the label of each region according to the distance between the region and surfaces. The performance of the algorithm is tested on a variety of extremity images using manually segmented images as "gold standard". The experiments showed that our algorithm provided a bone boundary with an average area overlap of 90% compared to the gold standard. Keywords: Segmentation, region growing, voting, surface fitting 1. INTRODUCTION Accurate diagnosis of disease states in medical radiographic images often depends on the detection of small, low contrast details in the image. The visibility of these details is dependent of the film's tonescale curve. The tonescale response curve of traditional silver halide films is determined by the film's built-in chemical emulsion and the development process. The manufacturer attempts to optimize the film for a range of exposure techniques and exam types. Digital radiographic imaging systems record raw image data without the built in tonescale curve and with greater dynamic range. The increased dynamic range of the digital image along with digital image processing techniques allow for manipulation of the image before it is viewed by the radiologist. Segmentation of the ROT (region of interest) in the code value histogram is an important first step in performing image enhancement processing. Once identified, the range of code values corresponding to the ROT can be optimally mapped, via a human brightness model, to the film density response of a laser printer or to the luminance response of a CRT for visual interpretation. Robust algorithms exist for segmentation of direct exposure and collimation regions in radiographic imagery. Previous methods have focused on segmenting the code value histogram based on knowledge of the body part being imaged and the characteristics of the corresponding histogram.' For a particular body part the histogram contains certain features (peaks, valleys, etc.) that can be used to map code values to anatomical features. This technique lacks robustness in certain exam types and exposure conditions. Another approach focused on segmenting Further author information: (Send correspondence to S.K.P.) S. K. P.: pakin ece.rochester.edu R.S.G.: rsg cs.rit.edu L.L.B.: lori.barski kodak.com D.H.F.: dfoos kodak.com K. J.P.: parker ece.rochester.edu 1296 Medical Imaging 2001: Image Processing, Milan Sonka, Kenneth M. Hanson, Editors, Proceedings of SPIE Vol (2001) 2001 SPIE /01/$15.00
2 the image into bone and tissue regions based on a neural network classifier using texture measures as features.2 The time required to obtain the texture features makes the approach impractical. The definition of an robust and effective method to refine the estimate of the ROT by segmenting the bone and soft tissue components of the image has remained elusive. The algorithm proposed in this paper can be considered as a hybrid technique that combines the features of regionbased and histogram-based techniques.3 Histogram-based techniques follow the Bayesian approach for estimation of the label field by modeling the image as a sample from a Gaussian distributed random field. Also, the intensity values of the pixels are assumed to be independent random variables whose parameters of their density function (mean and variance values since they are Gaussian distributed) depend on the label of the pixel. Assuming equally likely labels for every pixel results in well-known K-means algorithm.4 There are also numerous papers in the literature which impose spatial constraints by modeling the label field as a Gibbs or Markov random field.57 In general, these algorithms provide better results than K-means algorithm but they are also known for their significant computational burden. The main problem associated with the usage of histogram-based techniques in the context of the segmentation of CR images is that CR images do not follow the image model that histogram-based techniques rely on due to the exposure variations. Our algorithm attempts to address this problem in several steps. First, the image is divided into regions in which the pixels share certain statistical similarities. The feature extraction and region growing stages that perform this task will be covered in Section 2. After region growing stage, each region is labeled as bone or soft tissue. The voting procedure, which performs the labeling task, consists of clustering of regions using K-means algorithm with 2 classes. Note that performing region-based instead of pixel-based clustering effectively introduces spatial constraints to the label field. The effect of exposure variations is compensated by processing the image locally using the neighborhood systems and sub-images. The voting procedure will be explained in detail in Section 3. The last stage of the algorithm refines the labels of the regions by fitting a 2nd order bivariate polynomial to each tissue and investigating the average distance between each region and the surface of the label that they belong. The details of this stage is provided in Section 4. Finally, the experiments and the conclusions are stated in Sections 5 and 6, respectively. 2. CONSTRUCTION OF REGIONS 2.1. Feature Extraction In the first stage of the algorithm, two features are computed for each pixel. The features are denoted as,u and o_i, where i denotes the index of the pixel site, and they reflect the local characteristics around the pixel that they belong.. Iii : The median value of the intensities of the pixels that are inside the neighborhood of pixel x assuming 8-connectivity.4 The region growing algorithm depends on local differences which are suppressed to some extent in sample mean computation. Because of this fact, median value is used instead of sample mean to avoid the smoothing on region boundaries. More sophisticated schemes for feature extraction such as iterative methods that preserve line structures have been avoided due to their high computational cost.8. Ui : The standard deviation of the intensities of the pixels that are inside the neighborhood of pixel x assuming 8-connectivity Region Growing The two feature values computed in the previous step are utilized for investigation of the statistical similarity of each adjacent pixel pair where adjacency is defined in an 8-connectivity sense. For the pixel pair (xi, x2), the following statistical tests are utilized to determine whether they are sufficiently similar to be in the same region.. F-test : The pixels (xi, x3) and their corresponding neighbors are assumed to be the realizations of the random variables whose distributions are N(i, vi?), and N(i,v), respectively. The well-known F-test, which derives from the fact that if zi = v, then the RV has F-distribution with degrees of freedom (K 1, K 1), where K is the size of both ensemble,9 is used to investigate the similarity of the variances of the distributions in order to test the hypothesis that both ensemble are indeed governed by the same distribution. Note that, Proc. SPIE Vol
3 a and a are the sample variance values and have been computed in the feature extraction step. F-test is formulation as follows 0-i <7F, (1) 0-j where 7F 15 a predetermined threshold. Note that 'YF is always greater than 1, and the reciprocal of the Eqn 1 has to be considered if a < cr3.. Mahalanobis distance : We exploit the interpretation of Mahalanobis distance that is used for investigating the closeness (statistical) of a point to an ensemble whose distribution is known. Thus, we end up with the following two tests in which x and x3 are compared to the ensemble that consists of x3 and x and their neighbors, respectively. where 'YM 5 a predetermined threshold. /ii /iji <7M and IiiI <'YM (2) 0_i For a particular adjacent pixel pair (xi, x3), if all the above statistical tests are satisfied, that pixel pair is said to be connected. Following the investigation of the similarity of each adjacent pixel pair, regions that are defined as maximal set of pixels all belonging to the same connected components are formed. During the stages that follows region growing, the algorithm basically attempts to label each region as either bone or soft tissue. Hence, it is vital to adjust the thresholds 'YM and 'YF in order to construct regions whose pixels belong to only one type of tissue, otherwise some of the pixels will inevitably be assigned incorrect tissue labels. While high threshold values will cause leaking between regions that correspond to different tissue types, too low values will result in over segmentation which deteriorates the performance of the stages that follows growing. 0_j 3. VOTING PROCEDURE Following the region growing stage, each region is assigned a tissue label using the voting procedure. "Voting" is a broad term and it has been used under several contexts in the literature such as Hough transform related grouping techniques,' and majority voting schemes used for combining the decisions of several classifiers." The procedure we propose assigns bone or soft tissue votes to the regions that have been constructed in the previous step using local 2-class clustering. As mentioned in Section 1, the exposure variations seen on the image plate prevents the digital radiographic images from having a bimodal histogram. The significant overlap between the histograms of bone and soft tissue intensities makes the use of global clustering algorithms inefficient. The voting procedure is essentially governed by the following observation: Because of the relatively high x-ray absorption of bone, locally, bone pixels are darker than soft tissue pixels. Exposure variations makes this observation invalid globally. During the voting procedure, the image to be segmented is analyzed using overlapping blocks. The following notation is introduced in order to explain the voting procedure: R'm : mt region. Vm(B) : The votes for a bone label in the mthl region. Vm(S) : The votes for a soft tissue label in the mth region. : kt block. l?.ik : region in /çth block. : Neighborhood system of R-k. : th region in A/'ik. : The label of l?jik as the result of clustering of the regions in JVk Proc. SPIE Vol. 4322
4 The voting procedure can be summarized as follows: S For each!3k For each Rjk * Compute the average intensities of {Rjjk}. * Cluster {R-jk} using k-means algorithm with 2 clusters. * For each 'Ijik. Jf jik is bone, increment V(B) of corresponding l?'m.. Jf jik is soft tissue, increment V(S) of corresponding 7?m. Since overlapping blocks are used, a particular region, l?'m, which has been constructed in growing stage, could quite probably appear in different blocks. Furthermore, the borders of blocks can pass through 7m, thus different patches of m could appear in different blocks. Hence, according to the above formulation, two different symbols, R'jk and R,j'k', may correspond to different patches of a particular region, 'km. Even though, another symbol for this mapping between the regions in the image and the regions in the blocks is not reserved here for the sake of convenience, it is very crucial in the implementation. The reason for this importance is that only the votes of l?-'m '5 are kept, but Rjk'5 are processed. Since a particular region, l?'m, will appear in different blocks and neighborhood systems, it will acquire several votes. After all the blocks and regions are processed the tissue labels of the regions are determined as follows:. For each 'R-m If Vm(B) > Vm(S), 7?-m is a bone region. If Vm(B) < Vm(S), R'm is a soft tissue region. If Vm(B) = Vm(S), the label of 7Z is undecided. The regions whose labels are undecided are reconsidered and their labels are assigned in the next stage which will be discussed in the next section. 4. REFINEMENT OF LABELS VIA SURFACE FITTING Although the voting procedure results in correct tissue labels for most of the regions, the probability of misclassification increases in the areas of image where regions that belong to same type of tissue densely clustered. Such dense clusters tend to occur especially in the soft tissue close to tissue boundary. Since the thickness of the tissue varies greatly in such areas, the magnitude of the image gradient is usually significant compared to other areas of the images. Because of the reasons above, the tissue labels are reevaluated by the last stage of the algorithm in order to ameliorate the segmentation result provided by the voting procedure, and to assign tissue labels to the regions left undecided. The last stage essentially attempts to represent bone and soft tissue regions of the image with 2nd order bivariate polynomials whose coefficients are computed in a LS sense. After the computation of the surfaces, the labels of the regions whose RMS fitting error to the surface that corresponds to its tissue type is significant are reversed. The regions whose labels have been left undecided at the end of the voting procedure are assigned labels during the first iteration. The steps of the label refinement with surface fitting step can be stated as follows: Divide the image into 4 quadrants along the principal axes of tissue regions. Iterate the following steps until no label is changed. Compute the surfaces in each quadrant for bone, lb,k, and soft tissue, PS,k, using LS fitting, for k = For each R'm Proc. SPIE Vol
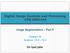
5 * Compute B,m MPB,k R'mURMS and S,m IIPS,k lmmrms * If the label of 'R'm IS bone and B,m > S,m, then change the label. * If the label of 7?'m js soft tissue and B,m < S,m, then change the label. One drawback of the surface fitting algorithm is that, since it depends on the LS data fitting concept, it inherently assumes that a label might be incorrect only if the size of its corresponding region is relatively small. A big region has the ability to bend the surface to be fitted towards itself strongly due to the well-known fact that LS data-fitting is very sensitive to noise. Hence, even if a relatively big region is misclassified during the voting procedure, its RMS fitting error will turn out to be small and this will make the label correction impossible. The surface fitting stage can also be conceptualized as a clustering scheme in which the regions are clustered into two classes using a different kind of metric than the usual ones (e.g. Mahalanobis distance) that are used in conventional clustering algorithms. From this point of view, the voting procedure provides the initial classification. 5. EXPERIMENTS The performance of the algorithm was tested on 14 extremity images which include wrist, knee, hand, ankle and foot exams. Images that are manually segmented under the supervision of a radiologist were used as the gold standard. The original images were in three different sizes, 2500 x 2048, 2048 x 2500, and 2392 x 1792, with 12-bits per pixel. For the experiments, 2md level approximation coefficients of the wavelet decomposition which exploits the wavelet filter Daubechies4 were used.12 After the computation of approximation coefficients, the resulting images, which are essentially the versions of the originals that are acquired by filtering and downsampling by 4, are cropped in order to feed the algorithm with an image that is free of artifacts that occur towards the image boundaries. For each image used in the experiment, blocks of size 64 x 64 which overlap 32 pixels in all directions were used. A common misclassification case in the images in which the bone tissue is clustered towards the center (such as knee) is that, some of relatively small regions that are close to the tissue boundary can be labeled erroneously. This is due to the fact that comparing the regions close to the tissue boundary with the bone tissue surface is effectively an extrapolation. In order to correct such misclassifications, binary opening operation is applied to both bone and soft tissue regions after surface fitting stage. In Figure 1, two different images that were used during the experiments and their corresponding segmentation results are displayed in l and 2nd row, respectively. The red regions in the images of the 2nd row show the bone tissue that is detected by the algorithm. Also, a visual comparison between the result of algorithm and manual segmentation is provided in the last row. In those images, red pixels correspond to ones that were classified as bone in both cases. In addition to that, blue region indicates the pixels that were classified as bone by the algorithm but as soft tissue during manual segmentation, and vice versa for green region. In Table 1, the summary of the experiments, which contains the type of extremities and the sizes (after cropping) of the images, and the computational time, is provided. In order to evaluate the performance of the algorithm statistically, the labeling process can be viewed as a binary hypothesis testing problem. If the events that a pixel belongs to bone, and to soft tissue are assumed as the null and alternate hypothesis, respectively, the performance of the algorithm can be evaluated by computing the ratio of correct detections to total number of decisions where being correct is defined with respect to the gold standard. In fact, this ratio is the well-known parameter "accuracy" which is equal to TP + TN Accuracy = TP + TN + FN + FP (3) where (TN+TP), FN, and FP are the number of correct decisions, false alarms (green regions in last row of Figure 1), and misses (blue regions in last row of Figure 1), respectively. The experiments were conducted on an SGI Indigo2 workstation. It can be seen from Table 1 that the algorithm provides over 90% accuracy (except an outlier image which is the case 6) with reasonable computational time. The observer variability has to be taken into consideration when the accuracy results are interpreted. For this purpose, a particular wrist image (case 5) is manually segmented 10 times and the accuracy values of those experiments are computed by assuming the original manual segmentation "gold standard". The manual segmentations resulted in an average accuracy value of 96.53% with a standard deviation of No attempt has been made to streamline or optimize the code for more rapid completion of the analysis Proc. SPIE Vol. 4322
6 CASE SIZE TIME (sec) ACCURACY. 13 (knee) 501 x % 18 (wrist) 381 x % 20 (knee) 606 x % 22 (knee) 526 x % 23 (knee) 566 x % 25 (foot) 441 x % 38 (foot) 596 x % 47 (knee) 596 x % 5 (wrist) 426 x % 52 (ankle) 551 x % 53 (hand) 451 x % 55 (ankle) 606 x % 6 (foot) 381 x % 9 (ankle) 321 x % Table 1. Summary of the experiments 6. CONCLUSION In this paper, we presented a novel algorithm for segmentation of CR images of extremities. The algorithm requires no supervision and performs the segmentation task with reasonable computational complexity. The experiments showed that the algorithm is robust in parameter space. In other words, the optimal parameter values, which are found experimentally, for the images that share common characteristics are close to each other. This algorithm can be useful for a variety of applications, including image enhancement and automatic recognition of x-ray exam type. ACKNOWLED GMENTS We would like to thank Thomas Gaborski and Dr. Saara Totterman for their help in acquisition of manual segmentations which have been used in segmentation evaluation and observer variability studies. REFERENCES 1. J. Capozzi and R. Schaetzing, "Method and apparatus for automatic tonescale generation in digital radiographic images," U.S. Patent 5,164,998, L. Barski, R. Gaborski, and P. Anderson, "A neural network approach to the segmentation of digital radiographic images," in ANNIE 93 Artificial Neural Networks in Engineering, St. Louis, MO, K. Hans, S. N. Efstratiadis, N. Maglaveras, and A. K. Katsaggelos, "Hybrid image segmentation using watersheds and fast region merging," IEEE Trans. on Image Proc. 7, pp , M. Nadler and E. P. Smith, Pattern Recognition Engineering, John Wiley & Sons, Geman and D. Geman, "Stochastic relaxation, gibbs distribution, and the bayesian restoration of images," IEEE Trans. Patt. Anal. Machine Intell. PAMI-6, pp , J. Besag, "On the statistical analysis of dirty pictures," J. Roy. Stat. Soc. B 48, pp , T. N. Pappas, "An adaptive clustering algorithm for image segmentation," IEEE Trans on Signal Proc. 40, pp , J. G. Tamez-Pena, Four-Dimensional Reconstruction and Visualization of Complex Musculos/celetal Structures, Ph.D. Thesis, Univ. of Rochester, V. K. Rohatgi, An Introduction to Probability Theory and Mathematical Statistics, John Wiley & Sons, G. L. Foresti, V. Murino, C. S. Regazzoni, and A. Trucco, "A voing-based approach for fast object recognition in underwater acoustic images," IEEE Journal of Oceanic Eng. 22, pp , L. Lam and C. Y. Suen, "Application of majority voting to pattern recognition: An analysis of ots behavior and performance," IEEE Trans. on Sys., Man, and Cyber. - Part A: Sys and Humans 27, pp , I. Daubechies, "Orthonormal bases of compactly supported wavelets," Comm. in Pure and Applied Math. 41, pp , Proc. SPIE Vol
7 1 Figure 1. From top to bottom, left to right: Knee image; bone region of knee at the end of segmentation; comparison of the result of the algorithm with manual segmentation; wrist image; bone region of wrist at the end of segmentation; comparison of the result of the algorithm with manual segmentation Proc. SPIE Vol. 4322
AN EFFICIENT BINARIZATION TECHNIQUE FOR FINGERPRINT IMAGES S. B. SRIDEVI M.Tech., Department of ECE
 AN EFFICIENT BINARIZATION TECHNIQUE FOR FINGERPRINT IMAGES S. B. SRIDEVI M.Tech., Department of ECE sbsridevi89@gmail.com 287 ABSTRACT Fingerprint identification is the most prominent method of biometric
AN EFFICIENT BINARIZATION TECHNIQUE FOR FINGERPRINT IMAGES S. B. SRIDEVI M.Tech., Department of ECE sbsridevi89@gmail.com 287 ABSTRACT Fingerprint identification is the most prominent method of biometric
MR IMAGE SEGMENTATION
 MR IMAGE SEGMENTATION Prepared by : Monil Shah What is Segmentation? Partitioning a region or regions of interest in images such that each region corresponds to one or more anatomic structures Classification
MR IMAGE SEGMENTATION Prepared by : Monil Shah What is Segmentation? Partitioning a region or regions of interest in images such that each region corresponds to one or more anatomic structures Classification
RESTORATION OF DEGRADED DOCUMENTS USING IMAGE BINARIZATION TECHNIQUE
 RESTORATION OF DEGRADED DOCUMENTS USING IMAGE BINARIZATION TECHNIQUE K. Kaviya Selvi 1 and R. S. Sabeenian 2 1 Department of Electronics and Communication Engineering, Communication Systems, Sona College
RESTORATION OF DEGRADED DOCUMENTS USING IMAGE BINARIZATION TECHNIQUE K. Kaviya Selvi 1 and R. S. Sabeenian 2 1 Department of Electronics and Communication Engineering, Communication Systems, Sona College
Introduction to Medical Imaging (5XSA0) Module 5
 Introduction to Medical Imaging (5XSA0) Module 5 Segmentation Jungong Han, Dirk Farin, Sveta Zinger ( s.zinger@tue.nl ) 1 Outline Introduction Color Segmentation region-growing region-merging watershed
Introduction to Medical Imaging (5XSA0) Module 5 Segmentation Jungong Han, Dirk Farin, Sveta Zinger ( s.zinger@tue.nl ) 1 Outline Introduction Color Segmentation region-growing region-merging watershed
Integrating Intensity and Texture in Markov Random Fields Segmentation. Amer Dawoud and Anton Netchaev. {amer.dawoud*,
 Integrating Intensity and Texture in Markov Random Fields Segmentation Amer Dawoud and Anton Netchaev {amer.dawoud*, anton.netchaev}@usm.edu School of Computing, University of Southern Mississippi 118
Integrating Intensity and Texture in Markov Random Fields Segmentation Amer Dawoud and Anton Netchaev {amer.dawoud*, anton.netchaev}@usm.edu School of Computing, University of Southern Mississippi 118
CHAPTER 6 MODIFIED FUZZY TECHNIQUES BASED IMAGE SEGMENTATION
 CHAPTER 6 MODIFIED FUZZY TECHNIQUES BASED IMAGE SEGMENTATION 6.1 INTRODUCTION Fuzzy logic based computational techniques are becoming increasingly important in the medical image analysis arena. The significant
CHAPTER 6 MODIFIED FUZZY TECHNIQUES BASED IMAGE SEGMENTATION 6.1 INTRODUCTION Fuzzy logic based computational techniques are becoming increasingly important in the medical image analysis arena. The significant
Fundamentals of Digital Image Processing
 \L\.6 Gw.i Fundamentals of Digital Image Processing A Practical Approach with Examples in Matlab Chris Solomon School of Physical Sciences, University of Kent, Canterbury, UK Toby Breckon School of Engineering,
\L\.6 Gw.i Fundamentals of Digital Image Processing A Practical Approach with Examples in Matlab Chris Solomon School of Physical Sciences, University of Kent, Canterbury, UK Toby Breckon School of Engineering,
Texture Image Segmentation using FCM
 Proceedings of 2012 4th International Conference on Machine Learning and Computing IPCSIT vol. 25 (2012) (2012) IACSIT Press, Singapore Texture Image Segmentation using FCM Kanchan S. Deshmukh + M.G.M
Proceedings of 2012 4th International Conference on Machine Learning and Computing IPCSIT vol. 25 (2012) (2012) IACSIT Press, Singapore Texture Image Segmentation using FCM Kanchan S. Deshmukh + M.G.M
Medical images, segmentation and analysis
 Medical images, segmentation and analysis ImageLab group http://imagelab.ing.unimo.it Università degli Studi di Modena e Reggio Emilia Medical Images Macroscopic Dermoscopic ELM enhance the features of
Medical images, segmentation and analysis ImageLab group http://imagelab.ing.unimo.it Università degli Studi di Modena e Reggio Emilia Medical Images Macroscopic Dermoscopic ELM enhance the features of
Mine Boundary Detection Using Markov Random Field Models
 Mine Boundary Detection Using Markov Random Field Models Xia Hua and Jennifer Davidson Department of Electrical & Computer Engineering Iowa State University Noel Cressie Department of Statistics Iowa State
Mine Boundary Detection Using Markov Random Field Models Xia Hua and Jennifer Davidson Department of Electrical & Computer Engineering Iowa State University Noel Cressie Department of Statistics Iowa State
IDENTIFYING GEOMETRICAL OBJECTS USING IMAGE ANALYSIS
 IDENTIFYING GEOMETRICAL OBJECTS USING IMAGE ANALYSIS Fathi M. O. Hamed and Salma F. Elkofhaifee Department of Statistics Faculty of Science University of Benghazi Benghazi Libya felramly@gmail.com and
IDENTIFYING GEOMETRICAL OBJECTS USING IMAGE ANALYSIS Fathi M. O. Hamed and Salma F. Elkofhaifee Department of Statistics Faculty of Science University of Benghazi Benghazi Libya felramly@gmail.com and
Texture Segmentation by Windowed Projection
 Texture Segmentation by Windowed Projection 1, 2 Fan-Chen Tseng, 2 Ching-Chi Hsu, 2 Chiou-Shann Fuh 1 Department of Electronic Engineering National I-Lan Institute of Technology e-mail : fctseng@ccmail.ilantech.edu.tw
Texture Segmentation by Windowed Projection 1, 2 Fan-Chen Tseng, 2 Ching-Chi Hsu, 2 Chiou-Shann Fuh 1 Department of Electronic Engineering National I-Lan Institute of Technology e-mail : fctseng@ccmail.ilantech.edu.tw
Adaptive Wavelet Image Denoising Based on the Entropy of Homogenus Regions
 International Journal of Electrical and Electronic Science 206; 3(4): 9-25 http://www.aascit.org/journal/ijees ISSN: 2375-2998 Adaptive Wavelet Image Denoising Based on the Entropy of Homogenus Regions
International Journal of Electrical and Electronic Science 206; 3(4): 9-25 http://www.aascit.org/journal/ijees ISSN: 2375-2998 Adaptive Wavelet Image Denoising Based on the Entropy of Homogenus Regions
A Quantitative Approach for Textural Image Segmentation with Median Filter
 International Journal of Advancements in Research & Technology, Volume 2, Issue 4, April-2013 1 179 A Quantitative Approach for Textural Image Segmentation with Median Filter Dr. D. Pugazhenthi 1, Priya
International Journal of Advancements in Research & Technology, Volume 2, Issue 4, April-2013 1 179 A Quantitative Approach for Textural Image Segmentation with Median Filter Dr. D. Pugazhenthi 1, Priya
doi: /
 Yiting Xie ; Anthony P. Reeves; Single 3D cell segmentation from optical CT microscope images. Proc. SPIE 934, Medical Imaging 214: Image Processing, 9343B (March 21, 214); doi:1.1117/12.243852. (214)
Yiting Xie ; Anthony P. Reeves; Single 3D cell segmentation from optical CT microscope images. Proc. SPIE 934, Medical Imaging 214: Image Processing, 9343B (March 21, 214); doi:1.1117/12.243852. (214)
Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging
 1 CS 9 Final Project Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging Feiyu Chen Department of Electrical Engineering ABSTRACT Subject motion is a significant
1 CS 9 Final Project Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging Feiyu Chen Department of Electrical Engineering ABSTRACT Subject motion is a significant
Efficient Image Compression of Medical Images Using the Wavelet Transform and Fuzzy c-means Clustering on Regions of Interest.
 Efficient Image Compression of Medical Images Using the Wavelet Transform and Fuzzy c-means Clustering on Regions of Interest. D.A. Karras, S.A. Karkanis and D. E. Maroulis University of Piraeus, Dept.
Efficient Image Compression of Medical Images Using the Wavelet Transform and Fuzzy c-means Clustering on Regions of Interest. D.A. Karras, S.A. Karkanis and D. E. Maroulis University of Piraeus, Dept.
Image Segmentation Based on Watershed and Edge Detection Techniques
 0 The International Arab Journal of Information Technology, Vol., No., April 00 Image Segmentation Based on Watershed and Edge Detection Techniques Nassir Salman Computer Science Department, Zarqa Private
0 The International Arab Journal of Information Technology, Vol., No., April 00 Image Segmentation Based on Watershed and Edge Detection Techniques Nassir Salman Computer Science Department, Zarqa Private
Evaluation of texture features for image segmentation
 RIT Scholar Works Articles 9-14-2001 Evaluation of texture features for image segmentation Navid Serrano Jiebo Luo Andreas Savakis Follow this and additional works at: http://scholarworks.rit.edu/article
RIT Scholar Works Articles 9-14-2001 Evaluation of texture features for image segmentation Navid Serrano Jiebo Luo Andreas Savakis Follow this and additional works at: http://scholarworks.rit.edu/article
Segmentation of Images
 Segmentation of Images SEGMENTATION If an image has been preprocessed appropriately to remove noise and artifacts, segmentation is often the key step in interpreting the image. Image segmentation is a
Segmentation of Images SEGMENTATION If an image has been preprocessed appropriately to remove noise and artifacts, segmentation is often the key step in interpreting the image. Image segmentation is a
Keywords: Thresholding, Morphological operations, Image filtering, Adaptive histogram equalization, Ceramic tile.
 Volume 3, Issue 7, July 2013 ISSN: 2277 128X International Journal of Advanced Research in Computer Science and Software Engineering Research Paper Available online at: www.ijarcsse.com Blobs and Cracks
Volume 3, Issue 7, July 2013 ISSN: 2277 128X International Journal of Advanced Research in Computer Science and Software Engineering Research Paper Available online at: www.ijarcsse.com Blobs and Cracks
Model-based segmentation and recognition from range data
 Model-based segmentation and recognition from range data Jan Boehm Institute for Photogrammetry Universität Stuttgart Germany Keywords: range image, segmentation, object recognition, CAD ABSTRACT This
Model-based segmentation and recognition from range data Jan Boehm Institute for Photogrammetry Universität Stuttgart Germany Keywords: range image, segmentation, object recognition, CAD ABSTRACT This
CHAPTER 6 DETECTION OF MASS USING NOVEL SEGMENTATION, GLCM AND NEURAL NETWORKS
 130 CHAPTER 6 DETECTION OF MASS USING NOVEL SEGMENTATION, GLCM AND NEURAL NETWORKS A mass is defined as a space-occupying lesion seen in more than one projection and it is described by its shapes and margin
130 CHAPTER 6 DETECTION OF MASS USING NOVEL SEGMENTATION, GLCM AND NEURAL NETWORKS A mass is defined as a space-occupying lesion seen in more than one projection and it is described by its shapes and margin
Data: a collection of numbers or facts that require further processing before they are meaningful
 Digital Image Classification Data vs. Information Data: a collection of numbers or facts that require further processing before they are meaningful Information: Derived knowledge from raw data. Something
Digital Image Classification Data vs. Information Data: a collection of numbers or facts that require further processing before they are meaningful Information: Derived knowledge from raw data. Something
Fractional Discrimination for Texture Image Segmentation
 Fractional Discrimination for Texture Image Segmentation Author You, Jia, Sattar, Abdul Published 1997 Conference Title IEEE 1997 International Conference on Image Processing, Proceedings of the DOI https://doi.org/10.1109/icip.1997.647743
Fractional Discrimination for Texture Image Segmentation Author You, Jia, Sattar, Abdul Published 1997 Conference Title IEEE 1997 International Conference on Image Processing, Proceedings of the DOI https://doi.org/10.1109/icip.1997.647743
Method of Background Subtraction for Medical Image Segmentation
 Method of Background Subtraction for Medical Image Segmentation Seongjai Kim Department of Mathematics and Statistics, Mississippi State University Mississippi State, MS 39762, USA and Hyeona Lim Department
Method of Background Subtraction for Medical Image Segmentation Seongjai Kim Department of Mathematics and Statistics, Mississippi State University Mississippi State, MS 39762, USA and Hyeona Lim Department
Experiments with Edge Detection using One-dimensional Surface Fitting
 Experiments with Edge Detection using One-dimensional Surface Fitting Gabor Terei, Jorge Luis Nunes e Silva Brito The Ohio State University, Department of Geodetic Science and Surveying 1958 Neil Avenue,
Experiments with Edge Detection using One-dimensional Surface Fitting Gabor Terei, Jorge Luis Nunes e Silva Brito The Ohio State University, Department of Geodetic Science and Surveying 1958 Neil Avenue,
Texture Based Image Segmentation and analysis of medical image
 Texture Based Image Segmentation and analysis of medical image 1. The Image Segmentation Problem Dealing with information extracted from a natural image, a medical scan, satellite data or a frame in a
Texture Based Image Segmentation and analysis of medical image 1. The Image Segmentation Problem Dealing with information extracted from a natural image, a medical scan, satellite data or a frame in a
Color Image Segmentation
 Color Image Segmentation Yining Deng, B. S. Manjunath and Hyundoo Shin* Department of Electrical and Computer Engineering University of California, Santa Barbara, CA 93106-9560 *Samsung Electronics Inc.
Color Image Segmentation Yining Deng, B. S. Manjunath and Hyundoo Shin* Department of Electrical and Computer Engineering University of California, Santa Barbara, CA 93106-9560 *Samsung Electronics Inc.
Detection & Classification of Lung Nodules Using multi resolution MTANN in Chest Radiography Images
 The International Journal Of Engineering And Science (IJES) ISSN (e): 2319 1813 ISSN (p): 2319 1805 Pages 98-104 March - 2015 Detection & Classification of Lung Nodules Using multi resolution MTANN in
The International Journal Of Engineering And Science (IJES) ISSN (e): 2319 1813 ISSN (p): 2319 1805 Pages 98-104 March - 2015 Detection & Classification of Lung Nodules Using multi resolution MTANN in
COMPUTER AND ROBOT VISION
 VOLUME COMPUTER AND ROBOT VISION Robert M. Haralick University of Washington Linda G. Shapiro University of Washington A^ ADDISON-WESLEY PUBLISHING COMPANY Reading, Massachusetts Menlo Park, California
VOLUME COMPUTER AND ROBOT VISION Robert M. Haralick University of Washington Linda G. Shapiro University of Washington A^ ADDISON-WESLEY PUBLISHING COMPANY Reading, Massachusetts Menlo Park, California
Perceptual Quality Improvement of Stereoscopic Images
 Perceptual Quality Improvement of Stereoscopic Images Jong In Gil and Manbae Kim Dept. of Computer and Communications Engineering Kangwon National University Chunchon, Republic of Korea, 200-701 E-mail:
Perceptual Quality Improvement of Stereoscopic Images Jong In Gil and Manbae Kim Dept. of Computer and Communications Engineering Kangwon National University Chunchon, Republic of Korea, 200-701 E-mail:
An ICA based Approach for Complex Color Scene Text Binarization
 An ICA based Approach for Complex Color Scene Text Binarization Siddharth Kherada IIIT-Hyderabad, India siddharth.kherada@research.iiit.ac.in Anoop M. Namboodiri IIIT-Hyderabad, India anoop@iiit.ac.in
An ICA based Approach for Complex Color Scene Text Binarization Siddharth Kherada IIIT-Hyderabad, India siddharth.kherada@research.iiit.ac.in Anoop M. Namboodiri IIIT-Hyderabad, India anoop@iiit.ac.in
Image Segmentation Techniques for Object-Based Coding
 Image Techniques for Object-Based Coding Junaid Ahmed, Joseph Bosworth, and Scott T. Acton The Oklahoma Imaging Laboratory School of Electrical and Computer Engineering Oklahoma State University {ajunaid,bosworj,sacton}@okstate.edu
Image Techniques for Object-Based Coding Junaid Ahmed, Joseph Bosworth, and Scott T. Acton The Oklahoma Imaging Laboratory School of Electrical and Computer Engineering Oklahoma State University {ajunaid,bosworj,sacton}@okstate.edu
Learning and Inferring Depth from Monocular Images. Jiyan Pan April 1, 2009
 Learning and Inferring Depth from Monocular Images Jiyan Pan April 1, 2009 Traditional ways of inferring depth Binocular disparity Structure from motion Defocus Given a single monocular image, how to infer
Learning and Inferring Depth from Monocular Images Jiyan Pan April 1, 2009 Traditional ways of inferring depth Binocular disparity Structure from motion Defocus Given a single monocular image, how to infer
Motivation. Intensity Levels
 Motivation Image Intensity and Point Operations Dr. Edmund Lam Department of Electrical and Electronic Engineering The University of Hong ong A digital image is a matrix of numbers, each corresponding
Motivation Image Intensity and Point Operations Dr. Edmund Lam Department of Electrical and Electronic Engineering The University of Hong ong A digital image is a matrix of numbers, each corresponding
Digital Image Processing. Prof. P.K. Biswas. Department of Electronics & Electrical Communication Engineering
 Digital Image Processing Prof. P.K. Biswas Department of Electronics & Electrical Communication Engineering Indian Institute of Technology, Kharagpur Image Segmentation - III Lecture - 31 Hello, welcome
Digital Image Processing Prof. P.K. Biswas Department of Electronics & Electrical Communication Engineering Indian Institute of Technology, Kharagpur Image Segmentation - III Lecture - 31 Hello, welcome
GENERAL AUTOMATED FLAW DETECTION SCHEME FOR NDE X-RAY IMAGES
 GENERAL AUTOMATED FLAW DETECTION SCHEME FOR NDE X-RAY IMAGES Karl W. Ulmer and John P. Basart Center for Nondestructive Evaluation Department of Electrical and Computer Engineering Iowa State University
GENERAL AUTOMATED FLAW DETECTION SCHEME FOR NDE X-RAY IMAGES Karl W. Ulmer and John P. Basart Center for Nondestructive Evaluation Department of Electrical and Computer Engineering Iowa State University
Lab 9. Julia Janicki. Introduction
 Lab 9 Julia Janicki Introduction My goal for this project is to map a general land cover in the area of Alexandria in Egypt using supervised classification, specifically the Maximum Likelihood and Support
Lab 9 Julia Janicki Introduction My goal for this project is to map a general land cover in the area of Alexandria in Egypt using supervised classification, specifically the Maximum Likelihood and Support
Artifacts and Textured Region Detection
 Artifacts and Textured Region Detection 1 Vishal Bangard ECE 738 - Spring 2003 I. INTRODUCTION A lot of transformations, when applied to images, lead to the development of various artifacts in them. In
Artifacts and Textured Region Detection 1 Vishal Bangard ECE 738 - Spring 2003 I. INTRODUCTION A lot of transformations, when applied to images, lead to the development of various artifacts in them. In
Image Enhancement Techniques for Fingerprint Identification
 March 2013 1 Image Enhancement Techniques for Fingerprint Identification Pankaj Deshmukh, Siraj Pathan, Riyaz Pathan Abstract The aim of this paper is to propose a new method in fingerprint enhancement
March 2013 1 Image Enhancement Techniques for Fingerprint Identification Pankaj Deshmukh, Siraj Pathan, Riyaz Pathan Abstract The aim of this paper is to propose a new method in fingerprint enhancement
Hybrid Approach for MRI Human Head Scans Classification using HTT based SFTA Texture Feature Extraction Technique
 Volume 118 No. 17 2018, 691-701 ISSN: 1311-8080 (printed version); ISSN: 1314-3395 (on-line version) url: http://www.ijpam.eu ijpam.eu Hybrid Approach for MRI Human Head Scans Classification using HTT
Volume 118 No. 17 2018, 691-701 ISSN: 1311-8080 (printed version); ISSN: 1314-3395 (on-line version) url: http://www.ijpam.eu ijpam.eu Hybrid Approach for MRI Human Head Scans Classification using HTT
An Approach for Reduction of Rain Streaks from a Single Image
 An Approach for Reduction of Rain Streaks from a Single Image Vijayakumar Majjagi 1, Netravati U M 2 1 4 th Semester, M. Tech, Digital Electronics, Department of Electronics and Communication G M Institute
An Approach for Reduction of Rain Streaks from a Single Image Vijayakumar Majjagi 1, Netravati U M 2 1 4 th Semester, M. Tech, Digital Electronics, Department of Electronics and Communication G M Institute
MULTI ORIENTATION PERFORMANCE OF FEATURE EXTRACTION FOR HUMAN HEAD RECOGNITION
 MULTI ORIENTATION PERFORMANCE OF FEATURE EXTRACTION FOR HUMAN HEAD RECOGNITION Panca Mudjirahardjo, Rahmadwati, Nanang Sulistiyanto and R. Arief Setyawan Department of Electrical Engineering, Faculty of
MULTI ORIENTATION PERFORMANCE OF FEATURE EXTRACTION FOR HUMAN HEAD RECOGNITION Panca Mudjirahardjo, Rahmadwati, Nanang Sulistiyanto and R. Arief Setyawan Department of Electrical Engineering, Faculty of
6. Object Identification L AK S H M O U. E D U
 6. Object Identification L AK S H M AN @ O U. E D U Objects Information extracted from spatial grids often need to be associated with objects not just an individual pixel Group of pixels that form a real-world
6. Object Identification L AK S H M AN @ O U. E D U Objects Information extracted from spatial grids often need to be associated with objects not just an individual pixel Group of pixels that form a real-world
Image Segmentation Using Iterated Graph Cuts Based on Multi-scale Smoothing
 Image Segmentation Using Iterated Graph Cuts Based on Multi-scale Smoothing Tomoyuki Nagahashi 1, Hironobu Fujiyoshi 1, and Takeo Kanade 2 1 Dept. of Computer Science, Chubu University. Matsumoto 1200,
Image Segmentation Using Iterated Graph Cuts Based on Multi-scale Smoothing Tomoyuki Nagahashi 1, Hironobu Fujiyoshi 1, and Takeo Kanade 2 1 Dept. of Computer Science, Chubu University. Matsumoto 1200,
Statistical Analysis of Metabolomics Data. Xiuxia Du Department of Bioinformatics & Genomics University of North Carolina at Charlotte
 Statistical Analysis of Metabolomics Data Xiuxia Du Department of Bioinformatics & Genomics University of North Carolina at Charlotte Outline Introduction Data pre-treatment 1. Normalization 2. Centering,
Statistical Analysis of Metabolomics Data Xiuxia Du Department of Bioinformatics & Genomics University of North Carolina at Charlotte Outline Introduction Data pre-treatment 1. Normalization 2. Centering,
Cellular Learning Automata-Based Color Image Segmentation using Adaptive Chains
 Cellular Learning Automata-Based Color Image Segmentation using Adaptive Chains Ahmad Ali Abin, Mehran Fotouhi, Shohreh Kasaei, Senior Member, IEEE Sharif University of Technology, Tehran, Iran abin@ce.sharif.edu,
Cellular Learning Automata-Based Color Image Segmentation using Adaptive Chains Ahmad Ali Abin, Mehran Fotouhi, Shohreh Kasaei, Senior Member, IEEE Sharif University of Technology, Tehran, Iran abin@ce.sharif.edu,
Motivation. Gray Levels
 Motivation Image Intensity and Point Operations Dr. Edmund Lam Department of Electrical and Electronic Engineering The University of Hong ong A digital image is a matrix of numbers, each corresponding
Motivation Image Intensity and Point Operations Dr. Edmund Lam Department of Electrical and Electronic Engineering The University of Hong ong A digital image is a matrix of numbers, each corresponding
Image Processing, Analysis and Machine Vision
 Image Processing, Analysis and Machine Vision Milan Sonka PhD University of Iowa Iowa City, USA Vaclav Hlavac PhD Czech Technical University Prague, Czech Republic and Roger Boyle DPhil, MBCS, CEng University
Image Processing, Analysis and Machine Vision Milan Sonka PhD University of Iowa Iowa City, USA Vaclav Hlavac PhD Czech Technical University Prague, Czech Republic and Roger Boyle DPhil, MBCS, CEng University
Denoising and Edge Detection Using Sobelmethod
 International OPEN ACCESS Journal Of Modern Engineering Research (IJMER) Denoising and Edge Detection Using Sobelmethod P. Sravya 1, T. Rupa devi 2, M. Janardhana Rao 3, K. Sai Jagadeesh 4, K. Prasanna
International OPEN ACCESS Journal Of Modern Engineering Research (IJMER) Denoising and Edge Detection Using Sobelmethod P. Sravya 1, T. Rupa devi 2, M. Janardhana Rao 3, K. Sai Jagadeesh 4, K. Prasanna
Adaptive Fingerprint Image Enhancement Techniques and Performance Evaluations
 Adaptive Fingerprint Image Enhancement Techniques and Performance Evaluations Kanpariya Nilam [1], Rahul Joshi [2] [1] PG Student, PIET, WAGHODIYA [2] Assistant Professor, PIET WAGHODIYA ABSTRACT: Image
Adaptive Fingerprint Image Enhancement Techniques and Performance Evaluations Kanpariya Nilam [1], Rahul Joshi [2] [1] PG Student, PIET, WAGHODIYA [2] Assistant Professor, PIET WAGHODIYA ABSTRACT: Image
Mobile Human Detection Systems based on Sliding Windows Approach-A Review
 Mobile Human Detection Systems based on Sliding Windows Approach-A Review Seminar: Mobile Human detection systems Njieutcheu Tassi cedrique Rovile Department of Computer Engineering University of Heidelberg
Mobile Human Detection Systems based on Sliding Windows Approach-A Review Seminar: Mobile Human detection systems Njieutcheu Tassi cedrique Rovile Department of Computer Engineering University of Heidelberg
Chapter 3 Set Redundancy in Magnetic Resonance Brain Images
 16 Chapter 3 Set Redundancy in Magnetic Resonance Brain Images 3.1 MRI (magnetic resonance imaging) MRI is a technique of measuring physical structure within the human anatomy. Our proposed research focuses
16 Chapter 3 Set Redundancy in Magnetic Resonance Brain Images 3.1 MRI (magnetic resonance imaging) MRI is a technique of measuring physical structure within the human anatomy. Our proposed research focuses
Semi-Automatic Detection of Cervical Vertebrae in X-ray Images Using Generalized Hough Transform
 Semi-Automatic Detection of Cervical Vertebrae in X-ray Images Using Generalized Hough Transform Mohamed Amine LARHMAM, Saïd MAHMOUDI and Mohammed BENJELLOUN Faculty of Engineering, University of Mons,
Semi-Automatic Detection of Cervical Vertebrae in X-ray Images Using Generalized Hough Transform Mohamed Amine LARHMAM, Saïd MAHMOUDI and Mohammed BENJELLOUN Faculty of Engineering, University of Mons,
2D image segmentation based on spatial coherence
 2D image segmentation based on spatial coherence Václav Hlaváč Czech Technical University in Prague Center for Machine Perception (bridging groups of the) Czech Institute of Informatics, Robotics and Cybernetics
2D image segmentation based on spatial coherence Václav Hlaváč Czech Technical University in Prague Center for Machine Perception (bridging groups of the) Czech Institute of Informatics, Robotics and Cybernetics
Texture Modeling using MRF and Parameters Estimation
 Texture Modeling using MRF and Parameters Estimation Ms. H. P. Lone 1, Prof. G. R. Gidveer 2 1 Postgraduate Student E & TC Department MGM J.N.E.C,Aurangabad 2 Professor E & TC Department MGM J.N.E.C,Aurangabad
Texture Modeling using MRF and Parameters Estimation Ms. H. P. Lone 1, Prof. G. R. Gidveer 2 1 Postgraduate Student E & TC Department MGM J.N.E.C,Aurangabad 2 Professor E & TC Department MGM J.N.E.C,Aurangabad
Adaptive Algorithm in Image Denoising Based on Data Mining
 Adaptive Algorithm in Image Denoising Based on Data Mining Yan-hua Ma 1 Chuan-jun Liu 2 1 Qingdao University of Science and Technology 2 Hisense Mobil Communication Technology Corporation Abstract An adaptive
Adaptive Algorithm in Image Denoising Based on Data Mining Yan-hua Ma 1 Chuan-jun Liu 2 1 Qingdao University of Science and Technology 2 Hisense Mobil Communication Technology Corporation Abstract An adaptive
Histogram and watershed based segmentation of color images
 Histogram and watershed based segmentation of color images O. Lezoray H. Cardot LUSAC EA 2607 IUT Saint-Lô, 120 rue de l'exode, 50000 Saint-Lô, FRANCE Abstract A novel method for color image segmentation
Histogram and watershed based segmentation of color images O. Lezoray H. Cardot LUSAC EA 2607 IUT Saint-Lô, 120 rue de l'exode, 50000 Saint-Lô, FRANCE Abstract A novel method for color image segmentation
IMAGE SEGMENTATION. Václav Hlaváč
 IMAGE SEGMENTATION Václav Hlaváč Czech Technical University in Prague Faculty of Electrical Engineering, Department of Cybernetics Center for Machine Perception http://cmp.felk.cvut.cz/ hlavac, hlavac@fel.cvut.cz
IMAGE SEGMENTATION Václav Hlaváč Czech Technical University in Prague Faculty of Electrical Engineering, Department of Cybernetics Center for Machine Perception http://cmp.felk.cvut.cz/ hlavac, hlavac@fel.cvut.cz
Improved Non-Local Means Algorithm Based on Dimensionality Reduction
 Improved Non-Local Means Algorithm Based on Dimensionality Reduction Golam M. Maruf and Mahmoud R. El-Sakka (&) Department of Computer Science, University of Western Ontario, London, Ontario, Canada {gmaruf,melsakka}@uwo.ca
Improved Non-Local Means Algorithm Based on Dimensionality Reduction Golam M. Maruf and Mahmoud R. El-Sakka (&) Department of Computer Science, University of Western Ontario, London, Ontario, Canada {gmaruf,melsakka}@uwo.ca
WEINER FILTER AND SUB-BLOCK DECOMPOSITION BASED IMAGE RESTORATION FOR MEDICAL APPLICATIONS
 WEINER FILTER AND SUB-BLOCK DECOMPOSITION BASED IMAGE RESTORATION FOR MEDICAL APPLICATIONS ARIFA SULTANA 1 & KANDARPA KUMAR SARMA 2 1,2 Department of Electronics and Communication Engineering, Gauhati
WEINER FILTER AND SUB-BLOCK DECOMPOSITION BASED IMAGE RESTORATION FOR MEDICAL APPLICATIONS ARIFA SULTANA 1 & KANDARPA KUMAR SARMA 2 1,2 Department of Electronics and Communication Engineering, Gauhati
A MORPHOLOGY-BASED FILTER STRUCTURE FOR EDGE-ENHANCING SMOOTHING
 Proceedings of the 1994 IEEE International Conference on Image Processing (ICIP-94), pp. 530-534. (Austin, Texas, 13-16 November 1994.) A MORPHOLOGY-BASED FILTER STRUCTURE FOR EDGE-ENHANCING SMOOTHING
Proceedings of the 1994 IEEE International Conference on Image Processing (ICIP-94), pp. 530-534. (Austin, Texas, 13-16 November 1994.) A MORPHOLOGY-BASED FILTER STRUCTURE FOR EDGE-ENHANCING SMOOTHING
Ulrik Söderström 16 Feb Image Processing. Segmentation
 Ulrik Söderström ulrik.soderstrom@tfe.umu.se 16 Feb 2011 Image Processing Segmentation What is Image Segmentation? To be able to extract information from an image it is common to subdivide it into background
Ulrik Söderström ulrik.soderstrom@tfe.umu.se 16 Feb 2011 Image Processing Segmentation What is Image Segmentation? To be able to extract information from an image it is common to subdivide it into background
Color Image Segmentation Editor Based on the Integration of Edge-Linking, Region Labeling and Deformable Model
 This paper appears in: IEEE International Conference on Systems, Man and Cybernetics, 1999 Color Image Segmentation Editor Based on the Integration of Edge-Linking, Region Labeling and Deformable Model
This paper appears in: IEEE International Conference on Systems, Man and Cybernetics, 1999 Color Image Segmentation Editor Based on the Integration of Edge-Linking, Region Labeling and Deformable Model
Image segmentation. Stefano Ferrari. Università degli Studi di Milano Methods for Image Processing. academic year
 Image segmentation Stefano Ferrari Università degli Studi di Milano stefano.ferrari@unimi.it Methods for Image Processing academic year 2017 2018 Segmentation by thresholding Thresholding is the simplest
Image segmentation Stefano Ferrari Università degli Studi di Milano stefano.ferrari@unimi.it Methods for Image Processing academic year 2017 2018 Segmentation by thresholding Thresholding is the simplest
Moving Object Segmentation Method Based on Motion Information Classification by X-means and Spatial Region Segmentation
 IJCSNS International Journal of Computer Science and Network Security, VOL.13 No.11, November 2013 1 Moving Object Segmentation Method Based on Motion Information Classification by X-means and Spatial
IJCSNS International Journal of Computer Science and Network Security, VOL.13 No.11, November 2013 1 Moving Object Segmentation Method Based on Motion Information Classification by X-means and Spatial
ECG782: Multidimensional Digital Signal Processing
 Professor Brendan Morris, SEB 3216, brendan.morris@unlv.edu ECG782: Multidimensional Digital Signal Processing Spring 2014 TTh 14:30-15:45 CBC C313 Lecture 10 Segmentation 14/02/27 http://www.ee.unlv.edu/~b1morris/ecg782/
Professor Brendan Morris, SEB 3216, brendan.morris@unlv.edu ECG782: Multidimensional Digital Signal Processing Spring 2014 TTh 14:30-15:45 CBC C313 Lecture 10 Segmentation 14/02/27 http://www.ee.unlv.edu/~b1morris/ecg782/
Automated Human Embryonic Stem Cell Detection
 212 IEEE Second Conference on Healthcare Informatics, Imaging and Systems Biology Automated Human Embryonic Stem Cell Detection Benjamin X. Guan, Bir Bhanu Center for Research in Intelligent Systems University
212 IEEE Second Conference on Healthcare Informatics, Imaging and Systems Biology Automated Human Embryonic Stem Cell Detection Benjamin X. Guan, Bir Bhanu Center for Research in Intelligent Systems University
Morphological Change Detection Algorithms for Surveillance Applications
 Morphological Change Detection Algorithms for Surveillance Applications Elena Stringa Joint Research Centre Institute for Systems, Informatics and Safety TP 270, Ispra (VA), Italy elena.stringa@jrc.it
Morphological Change Detection Algorithms for Surveillance Applications Elena Stringa Joint Research Centre Institute for Systems, Informatics and Safety TP 270, Ispra (VA), Italy elena.stringa@jrc.it
Region-based Segmentation
 Region-based Segmentation Image Segmentation Group similar components (such as, pixels in an image, image frames in a video) to obtain a compact representation. Applications: Finding tumors, veins, etc.
Region-based Segmentation Image Segmentation Group similar components (such as, pixels in an image, image frames in a video) to obtain a compact representation. Applications: Finding tumors, veins, etc.
Scene Text Detection Using Machine Learning Classifiers
 601 Scene Text Detection Using Machine Learning Classifiers Nafla C.N. 1, Sneha K. 2, Divya K.P. 3 1 (Department of CSE, RCET, Akkikkvu, Thrissur) 2 (Department of CSE, RCET, Akkikkvu, Thrissur) 3 (Department
601 Scene Text Detection Using Machine Learning Classifiers Nafla C.N. 1, Sneha K. 2, Divya K.P. 3 1 (Department of CSE, RCET, Akkikkvu, Thrissur) 2 (Department of CSE, RCET, Akkikkvu, Thrissur) 3 (Department
Image Restoration using Markov Random Fields
 Image Restoration using Markov Random Fields Based on the paper Stochastic Relaxation, Gibbs Distributions and Bayesian Restoration of Images, PAMI, 1984, Geman and Geman. and the book Markov Random Field
Image Restoration using Markov Random Fields Based on the paper Stochastic Relaxation, Gibbs Distributions and Bayesian Restoration of Images, PAMI, 1984, Geman and Geman. and the book Markov Random Field
[Programming Assignment] (1)
![[Programming Assignment] (1) [Programming Assignment] (1)](/thumbs/90/104460171.jpg) http://crcv.ucf.edu/people/faculty/bagci/ [Programming Assignment] (1) Computer Vision Dr. Ulas Bagci (Fall) 2015 University of Central Florida (UCF) Coding Standard and General Requirements Code for all
http://crcv.ucf.edu/people/faculty/bagci/ [Programming Assignment] (1) Computer Vision Dr. Ulas Bagci (Fall) 2015 University of Central Florida (UCF) Coding Standard and General Requirements Code for all
Character Recognition
 Character Recognition 5.1 INTRODUCTION Recognition is one of the important steps in image processing. There are different methods such as Histogram method, Hough transformation, Neural computing approaches
Character Recognition 5.1 INTRODUCTION Recognition is one of the important steps in image processing. There are different methods such as Histogram method, Hough transformation, Neural computing approaches
Processing Missing Values with Self-Organized Maps
 Processing Missing Values with Self-Organized Maps David Sommer, Tobias Grimm, Martin Golz University of Applied Sciences Schmalkalden Department of Computer Science D-98574 Schmalkalden, Germany Phone:
Processing Missing Values with Self-Organized Maps David Sommer, Tobias Grimm, Martin Golz University of Applied Sciences Schmalkalden Department of Computer Science D-98574 Schmalkalden, Germany Phone:
Enhanced Hybrid Compound Image Compression Algorithm Combining Block and Layer-based Segmentation
 Enhanced Hybrid Compound Image Compression Algorithm Combining Block and Layer-based Segmentation D. Maheswari 1, Dr. V.Radha 2 1 Department of Computer Science, Avinashilingam Deemed University for Women,
Enhanced Hybrid Compound Image Compression Algorithm Combining Block and Layer-based Segmentation D. Maheswari 1, Dr. V.Radha 2 1 Department of Computer Science, Avinashilingam Deemed University for Women,
Shadow detection and removal from a single image
 Shadow detection and removal from a single image Corina BLAJOVICI, Babes-Bolyai University, Romania Peter Jozsef KISS, University of Pannonia, Hungary Zoltan BONUS, Obuda University, Hungary Laszlo VARGA,
Shadow detection and removal from a single image Corina BLAJOVICI, Babes-Bolyai University, Romania Peter Jozsef KISS, University of Pannonia, Hungary Zoltan BONUS, Obuda University, Hungary Laszlo VARGA,
CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS
 CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS CHAPTER 4 CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS 4.1 Introduction Optical character recognition is one of
CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS CHAPTER 4 CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS 4.1 Introduction Optical character recognition is one of
Announcements. Recognition. Recognition. Recognition. Recognition. Homework 3 is due May 18, 11:59 PM Reading: Computer Vision I CSE 152 Lecture 14
 Announcements Computer Vision I CSE 152 Lecture 14 Homework 3 is due May 18, 11:59 PM Reading: Chapter 15: Learning to Classify Chapter 16: Classifying Images Chapter 17: Detecting Objects in Images Given
Announcements Computer Vision I CSE 152 Lecture 14 Homework 3 is due May 18, 11:59 PM Reading: Chapter 15: Learning to Classify Chapter 16: Classifying Images Chapter 17: Detecting Objects in Images Given
COMPUTER VISION. Dr. Sukhendu Das Deptt. of Computer Science and Engg., IIT Madras, Chennai
 COMPUTER VISION Dr. Sukhendu Das Deptt. of Computer Science and Engg., IIT Madras, Chennai 600036. Email: sdas@iitm.ac.in URL: //www.cs.iitm.ernet.in/~sdas 1 INTRODUCTION 2 Human Vision System (HVS) Vs.
COMPUTER VISION Dr. Sukhendu Das Deptt. of Computer Science and Engg., IIT Madras, Chennai 600036. Email: sdas@iitm.ac.in URL: //www.cs.iitm.ernet.in/~sdas 1 INTRODUCTION 2 Human Vision System (HVS) Vs.
Face Hallucination Based on Eigentransformation Learning
 Advanced Science and Technology etters, pp.32-37 http://dx.doi.org/10.14257/astl.2016. Face allucination Based on Eigentransformation earning Guohua Zou School of software, East China University of Technology,
Advanced Science and Technology etters, pp.32-37 http://dx.doi.org/10.14257/astl.2016. Face allucination Based on Eigentransformation earning Guohua Zou School of software, East China University of Technology,
Content Based Image Retrieval Using Color Quantizes, EDBTC and LBP Features
 Content Based Image Retrieval Using Color Quantizes, EDBTC and LBP Features 1 Kum Sharanamma, 2 Krishnapriya Sharma 1,2 SIR MVIT Abstract- To describe the image features the Local binary pattern (LBP)
Content Based Image Retrieval Using Color Quantizes, EDBTC and LBP Features 1 Kum Sharanamma, 2 Krishnapriya Sharma 1,2 SIR MVIT Abstract- To describe the image features the Local binary pattern (LBP)
Digital Image Analysis and Processing
 Digital Image Analysis and Processing CPE 0907544 Image Segmentation Part II Chapter 10 Sections : 10.3 10.4 Dr. Iyad Jafar Outline Introduction Thresholdingh Fundamentals Basic Global Thresholding Optimal
Digital Image Analysis and Processing CPE 0907544 Image Segmentation Part II Chapter 10 Sections : 10.3 10.4 Dr. Iyad Jafar Outline Introduction Thresholdingh Fundamentals Basic Global Thresholding Optimal
CHAPTER 1 INTRODUCTION
 CHAPTER 1 INTRODUCTION 1.1 Introduction Pattern recognition is a set of mathematical, statistical and heuristic techniques used in executing `man-like' tasks on computers. Pattern recognition plays an
CHAPTER 1 INTRODUCTION 1.1 Introduction Pattern recognition is a set of mathematical, statistical and heuristic techniques used in executing `man-like' tasks on computers. Pattern recognition plays an
Digital Image Processing COSC 6380/4393
 Digital Image Processing COSC 6380/4393 Lecture 21 Nov 16 th, 2017 Pranav Mantini Ack: Shah. M Image Processing Geometric Transformation Point Operations Filtering (spatial, Frequency) Input Restoration/
Digital Image Processing COSC 6380/4393 Lecture 21 Nov 16 th, 2017 Pranav Mantini Ack: Shah. M Image Processing Geometric Transformation Point Operations Filtering (spatial, Frequency) Input Restoration/
Image Segmentation and Registration
 Image Segmentation and Registration Dr. Christine Tanner (tanner@vision.ee.ethz.ch) Computer Vision Laboratory, ETH Zürich Dr. Verena Kaynig, Machine Learning Laboratory, ETH Zürich Outline Segmentation
Image Segmentation and Registration Dr. Christine Tanner (tanner@vision.ee.ethz.ch) Computer Vision Laboratory, ETH Zürich Dr. Verena Kaynig, Machine Learning Laboratory, ETH Zürich Outline Segmentation
Effects Of Shadow On Canny Edge Detection through a camera
 1523 Effects Of Shadow On Canny Edge Detection through a camera Srajit Mehrotra Shadow causes errors in computer vision as it is difficult to detect objects that are under the influence of shadows. Shadow
1523 Effects Of Shadow On Canny Edge Detection through a camera Srajit Mehrotra Shadow causes errors in computer vision as it is difficult to detect objects that are under the influence of shadows. Shadow
Modern Medical Image Analysis 8DC00 Exam
 Parts of answers are inside square brackets [... ]. These parts are optional. Answers can be written in Dutch or in English, as you prefer. You can use drawings and diagrams to support your textual answers.
Parts of answers are inside square brackets [... ]. These parts are optional. Answers can be written in Dutch or in English, as you prefer. You can use drawings and diagrams to support your textual answers.
MEDICAL IMAGE NOISE REDUCTION AND REGION CONTRAST ENHANCEMENT USING PARTIAL DIFFERENTIAL EQUATIONS
 MEDICAL IMAGE NOISE REDUCTION AND REGION CONTRAST ENHANCEMENT USING PARTIAL DIFFERENTIAL EQUATIONS Miguel Alemán-Flores, Luis Álvarez-León Departamento de Informática y Sistemas, Universidad de Las Palmas
MEDICAL IMAGE NOISE REDUCTION AND REGION CONTRAST ENHANCEMENT USING PARTIAL DIFFERENTIAL EQUATIONS Miguel Alemán-Flores, Luis Álvarez-León Departamento de Informática y Sistemas, Universidad de Las Palmas
Norbert Schuff VA Medical Center and UCSF
 Norbert Schuff Medical Center and UCSF Norbert.schuff@ucsf.edu Medical Imaging Informatics N.Schuff Course # 170.03 Slide 1/67 Objective Learn the principle segmentation techniques Understand the role
Norbert Schuff Medical Center and UCSF Norbert.schuff@ucsf.edu Medical Imaging Informatics N.Schuff Course # 170.03 Slide 1/67 Objective Learn the principle segmentation techniques Understand the role
Part-Based Skew Estimation for Mathematical Expressions
 Soma Shiraishi, Yaokai Feng, and Seiichi Uchida shiraishi@human.ait.kyushu-u.ac.jp {fengyk,uchida}@ait.kyushu-u.ac.jp Abstract We propose a novel method for the skew estimation on text images containing
Soma Shiraishi, Yaokai Feng, and Seiichi Uchida shiraishi@human.ait.kyushu-u.ac.jp {fengyk,uchida}@ait.kyushu-u.ac.jp Abstract We propose a novel method for the skew estimation on text images containing
Local Image Registration: An Adaptive Filtering Framework
 Local Image Registration: An Adaptive Filtering Framework Gulcin Caner a,a.murattekalp a,b, Gaurav Sharma a and Wendi Heinzelman a a Electrical and Computer Engineering Dept.,University of Rochester, Rochester,
Local Image Registration: An Adaptive Filtering Framework Gulcin Caner a,a.murattekalp a,b, Gaurav Sharma a and Wendi Heinzelman a a Electrical and Computer Engineering Dept.,University of Rochester, Rochester,
Improving Latent Fingerprint Matching Performance by Orientation Field Estimation using Localized Dictionaries
 Available Online at www.ijcsmc.com International Journal of Computer Science and Mobile Computing A Monthly Journal of Computer Science and Information Technology IJCSMC, Vol. 3, Issue. 11, November 2014,
Available Online at www.ijcsmc.com International Journal of Computer Science and Mobile Computing A Monthly Journal of Computer Science and Information Technology IJCSMC, Vol. 3, Issue. 11, November 2014,
Towards full-body X-ray images
 Towards full-body X-ray images Christoph Luckner 1,2, Thomas Mertelmeier 2, Andreas Maier 1, Ludwig Ritschl 2 1 Pattern Recognition Lab, FAU Erlangen-Nuernberg 2 Siemens Healthcare GmbH, Forchheim christoph.luckner@fau.de
Towards full-body X-ray images Christoph Luckner 1,2, Thomas Mertelmeier 2, Andreas Maier 1, Ludwig Ritschl 2 1 Pattern Recognition Lab, FAU Erlangen-Nuernberg 2 Siemens Healthcare GmbH, Forchheim christoph.luckner@fau.de
Part 3: Image Processing
 Part 3: Image Processing Image Filtering and Segmentation Georgy Gimel farb COMPSCI 373 Computer Graphics and Image Processing 1 / 60 1 Image filtering 2 Median filtering 3 Mean filtering 4 Image segmentation
Part 3: Image Processing Image Filtering and Segmentation Georgy Gimel farb COMPSCI 373 Computer Graphics and Image Processing 1 / 60 1 Image filtering 2 Median filtering 3 Mean filtering 4 Image segmentation
Automatic Colorization of Grayscale Images
 Automatic Colorization of Grayscale Images Austin Sousa Rasoul Kabirzadeh Patrick Blaes Department of Electrical Engineering, Stanford University 1 Introduction ere exists a wealth of photographic images,
Automatic Colorization of Grayscale Images Austin Sousa Rasoul Kabirzadeh Patrick Blaes Department of Electrical Engineering, Stanford University 1 Introduction ere exists a wealth of photographic images,
Analysis of Image and Video Using Color, Texture and Shape Features for Object Identification
 IOSR Journal of Computer Engineering (IOSR-JCE) e-issn: 2278-0661,p-ISSN: 2278-8727, Volume 16, Issue 6, Ver. VI (Nov Dec. 2014), PP 29-33 Analysis of Image and Video Using Color, Texture and Shape Features
IOSR Journal of Computer Engineering (IOSR-JCE) e-issn: 2278-0661,p-ISSN: 2278-8727, Volume 16, Issue 6, Ver. VI (Nov Dec. 2014), PP 29-33 Analysis of Image and Video Using Color, Texture and Shape Features
SURVEY ON IMAGE PROCESSING IN THE FIELD OF DE-NOISING TECHNIQUES AND EDGE DETECTION TECHNIQUES ON RADIOGRAPHIC IMAGES
 SURVEY ON IMAGE PROCESSING IN THE FIELD OF DE-NOISING TECHNIQUES AND EDGE DETECTION TECHNIQUES ON RADIOGRAPHIC IMAGES 1 B.THAMOTHARAN, 2 M.MENAKA, 3 SANDHYA VAIDYANATHAN, 3 SOWMYA RAVIKUMAR 1 Asst. Prof.,
SURVEY ON IMAGE PROCESSING IN THE FIELD OF DE-NOISING TECHNIQUES AND EDGE DETECTION TECHNIQUES ON RADIOGRAPHIC IMAGES 1 B.THAMOTHARAN, 2 M.MENAKA, 3 SANDHYA VAIDYANATHAN, 3 SOWMYA RAVIKUMAR 1 Asst. Prof.,
CORRELATION BASED CAR NUMBER PLATE EXTRACTION SYSTEM
 CORRELATION BASED CAR NUMBER PLATE EXTRACTION SYSTEM 1 PHYO THET KHIN, 2 LAI LAI WIN KYI 1,2 Department of Information Technology, Mandalay Technological University The Republic of the Union of Myanmar
CORRELATION BASED CAR NUMBER PLATE EXTRACTION SYSTEM 1 PHYO THET KHIN, 2 LAI LAI WIN KYI 1,2 Department of Information Technology, Mandalay Technological University The Republic of the Union of Myanmar
