Package randomglm. R topics documented: February 20, 2015
|
|
|
- Olivia Johnson
- 6 years ago
- Views:
Transcription
1 Package randomglm February 20, 2015 Version Date Title Random General Linear Model Prediction Author Lin Song, Peter Langfelder Maintainer Peter Langfelder Depends R (>= ), MASS, foreach, doparallel ZipData no License GPL (>= 2) Description The package implements a bagging predictor based on general linear models URL Repository CRAN Date/Publication :37:33 NeedsCompilation no R topics documented: braincancer mini predict.randomglm randomglm thinrandomglm Index 16 1
2 2 braincancer braincancer The brain cancer data set Description 2 sets containing the gene expression profiles of 55 and 65 brain cancer patients respectively. Usage data(braincancer) Format braincancer is a list of 2 components: train and test. "train" is a numeric matrix with 55 samples (rows) across 5000 genes (columns). "test" is a numeric matrix with 65 samples (rows) across the same 5000 genes (columns). Author(s) Lin Song, Steve Horvath Source Horvath S, Zhang B, Carlson M, Lu K, Zhu S, Felciano R, Laurance M, Zhao W, Shu Q, Lee Y, Scheck A, Liau L, Wu H, Geschwind D, Febbo P, Kornblum H, TF C, Nelson S, Mischel P: Analysis of Oncogenic Signaling Networks in Glioblastoma Identifies ASPM as a Novel Molecular Target. Proc Natl Acad Sci U S A 2006, 103(46): References Lin Song, Peter Langfelder, Steve Horvath: Random generalized linear model: a highly accurate and interpretable ensemble predictor. BMC Bioinformatics (2013) Examples data(braincancer)
3 mini 3 mini An example data set derived from the brain cancer data set Description This example contains one training set, one test set, a corresponding binary outcome and a corresponding continuous outcome. Outcomes are gene traits derived from the brain cancer data set. Usage data(mini) Format mini is a list of 6 components: x, xtest, yb, ybtest, yc and yctest. "x" is a numeric matrix with 55 samples (rows) across 4999 genes (columns). "xtest" is a numeric matrix with 65 samples (rows) across the same 4999 genes (columns). They are subsets of the original braincancer data. One gene is left out. "yc" and "yctest" are continuous outcomes equal to the expression values of the left out gene in the training and test set respectively. "yb" and "ybtest" are binary outcomes dichotomized at the median based on "yc" and "yctest". Author(s) Lin Song, Steve Horvath Source Horvath S, Zhang B, Carlson M, Lu K, Zhu S, Felciano R, Laurance M, Zhao W, Shu Q, Lee Y, Scheck A, Liau L, Wu H, Geschwind D, Febbo P, Kornblum H, TF C, Nelson S, Mischel P: Analysis of Oncogenic Signaling Networks in Glioblastoma Identifies ASPM as a Novel Molecular Target. Proc Natl Acad Sci U S A 2006, 103(46): References Lin Song, Peter Langfelder, Steve Horvath: Random generalized linear model: a highly accurate and interpretable ensemble predictor. BMC Bioinformatics (2013) Examples data(mini)
4 4 predict.randomglm predict.randomglm Prediction from a random generalized linear model predictor Description Implements a predict method on a previously-constructed random generalized linear model predictor and new data. Usage ## S3 method for class randomglm predict(object, newdata, type=c("response", "class"), thresholdclassprob = object$thresholdclassprob,...) Arguments object newdata a randomglm object such as one returned by randomglm. specification of test data for which to calculate the prediction. type type of prediction required. Type "response" gives the fitted probabilities for classification, the fitted values for regression. Type "class" applies only to classification, and produces the predicted class labels. thresholdclassprob the threshold of predictive probabilities to arrive at classification. Takes values between 0 and 1. Only used for binary outcomes.... other arguments that may be passed to and from methods. Currently unused. Details The function calculates prediction on new test data. It only works if object contains the regression models that were used to construct the predictor (see argument keepmodels of the function randomglm). If the predictor was trained on a multi-class response, the prediction is applied to each of the representing binary variables (see randomglm for details). Value For continuous prediction, the predicted values. For classification of binary response, predicted class when type="class"; or a two-column matrix giving the class probabilities if type="response". If the predictor was trained on a multi-class response, the returned value is a matrix of "cbind"-ed results for the representing individual binary variables (see randomglm for details). Author(s) Lin Song, Steve Horvath and Peter Langfelder.
5 randomglm 5 References Lin Song, Peter Langfelder, Steve Horvath: Random generalized linear model: a highly accurate and interpretable ensemble predictor. BMC Bioinformatics (2013) Examples ## binary outcome prediction # data generation data(iris) # Restrict data to first 100 observations iris=iris[1:100,] # Turn Species into a factor iris$species = as.factor(as.character(iris$species)) # Select a training and a test subset of the 100 observations set.seed(1) indx = sample(100, 67, replace=false) xytrain = iris[indx,] xytest = iris[-indx,] xtrain = xytrain[, -5] ytrain = xytrain[, 5] xtest = xytest[, -5] ytest = xytest[, 5] # predict with a small number of bags - normally nbags should be at least 100. RGLM = randomglm(xtrain, ytrain, ncandidatecovariates=ncol(xtrain), nbags=30, keepmodels = TRUE, nthreads = 1) predicted = predict(rglm, newdata = xtest, type="class") table(predicted, ytest) ## continuous outcome prediction x=matrix(rnorm(100*20),100,20) y=rnorm(100) xtrain = x[1:50,] ytrain = y[1:50] xtest = x[51:100,] ytest = y[51:100] RGLM = randomglm(xtrain, ytrain, classify=false, ncandidatecovariates=ncol(xtrain), nbags=10, keepmodels = TRUE, nthreads = 1) predicted = predict(rglm, newdata = xtest) randomglm Random generalized linear model predictor Description Ensemble predictor comprised of individual generalized linear model predictors.
6 6 randomglm Usage randomglm( # Input data x, y, xtest = NULL, # Include interactions? maxinteractionorder = 1, # Prediction type classify = is.factor(y) length(unique(y)) < 4, # Multi-level classification options - only apply to classification with multi-level response multiclass.global = TRUE, multiclass.pairwise = FALSE, multiclass.minobs = 1, multiclass.ignorelevels = NULL, # Sampling options nbags = 100, replace = TRUE, sampleweight=null, nobsinbag = if (replace) nrow(x) else as.integer(0.632 * nrow(x)), nfeaturesinbag = ceiling(ifelse(ncol(x)<=10, ncol(x), ifelse(ncol(x)<=300, ( *ncol(x))*ncol(x), ncol(x)/5))), mininbagobs = min( max( nrow(x)/2, 5), 2*nrow(x)/3), # Individual ensemble member predictor options ncandidatecovariates=50, corfncforcandidatecovariates= cor, coroptionsforcandidatecovariates = list(method = "pearson", use="p"), mandatorycovariates = NULL, interactionsmandatory = FALSE, keepmodels = is.null(xtest), # Miscellaneous options thresholdclassprob = 0.5, interactionseparatorforcoefnames = ".times.", randomseed = 12345, nthreads = NULL, verbose =0 ) Arguments x y a matrix whose rows correspond to observations (samples) and whose columns correspond to features (also known as covariates or variables). Thus, x contains the training data sets. outcome variable corresponding to the rows of x: at this point, one can either
7 randomglm 7 use a binary class outcome (factor variable) or a quantitative outcome (numeric variable). xtest an optional matrix of a second data set (referred to as test data set while the data in x are interpreted as training data). The number of rows can (and typically will) be different from the number of rows in x. maxinteractionorder integer specifying the maximum interaction level. The default is to have no interactions; numbers higher than 1 specify interactions up to that order. For example, 3 means quadratic and cubic interactions will be included. Warning: higher order interactions greatly increase the computation time. We see no benefit of using maxinteractionorder>2. classify logical: If TRUE the response y will be interpreted as a binary variable and logistic regression will be used. If FALSE the response y will be interpreted as a quantitative numeric variable and a least squares regression model will be used to arrive at base learners. Multi-level classification is split into a series of binary classification problems according to the multiclass... arguments described below. multiclass.global for multi-level classification, this logical argument controls whether binary variables of the type "level vs. all others" are included in the series of binary variables to which classification is applied. multiclass.pairwise for multi-level classification, this logical argument controls whether binary variables of the type "level A vs. level B" are included in the series of binary variables to which classification is applied. multiclass.minobs an integer specifying the minimum number of observations for each level for the level to be considered when creating "level vs. all" and "level vs. level" binary variables. multiclass.ignorelevels optional specifications of the values (levels) of the input response y that are to be ignored when constructing level vs. all and level vs. level binary responses. Note that observation with these values will be included in the "all" but will not have their own "level vs. all" variables. nbags replace sampleweight nobsinbag number of bags (bootstrap samples) for defining the ensemble predictor, i.e. this also corresponds to the number of individual GLMs. logical. If TRUE then each bootstrap sample (bag) is defined by sampling with replacement. Otherwise, sampling is carried out without replacement. We recommend to choose TRUE. weight assigned to each observations (sample) during bootstrap sampling. Default NULL corresponds to equal weights. number of observations selected for each bag. Typically, a bootstrap sample (bag) has the same number of observations as in the original data set (i.e. the rows of x). nfeaturesinbag number of features randomly selected for each bag. Features are randomly selected without replacement. If there are no interaction terms, then this number should be smaller than or equal to the number of rows of x.
8 8 randomglm mininbagobs minimum number of unique observations that constitute a valid bag. If the sampling produces a bag with fewer than this number of unique observations, the bag is discarded and re-sampled again until the number of unique observations is at least mininbagobs. This helps prevent too few unique observations in a bag which would lead to problems with model selection. ncandidatecovariates Positive integer. The number of features that are being considered for forward selection in each GLM (and in each bag). For each bag, the covariates are being chosen according their highest absolute correlation with the outcome. In case of a binary outcome, it is first turned into a binary numeric variable. corfncforcandidatecovariates the correlation function used to select candidate covariates. Choices include cor or biweight midcorrelation, bicor, implemented in the package WGCNA. The biweight mid-correlation is a robust alternative to the Pearson correlation. coroptionsforcandidatecovariates list of arguments to the correlation function. Note that robust correlations are sometimes problematic for binary class outcomes. When using the robust correlation bicor, use the argument "robusty=false". mandatorycovariates indices of features that are included as mandatory covariates in each GLM model. The default is no mandatory features. This allows the user to "force" variables into each GLM. interactionsmandatory logical: should interactions of mandatory covariates be mandatory as well? Interactions are only included up to the level specified in maxinteractionorder. keepmodels logical: should the regression models for each bag be kept? The models are necessary for future predictions using the predict function, predict() generic. thresholdclassprob number in the interval [0,1]. Recommended value 0.5. This parameter is only relevant in case of a binary outcome, i.e. for a logistic regression model. Then this threshold will be applied to the predictive class probabilities to arrive at binary outcome (class outcome). interactionseparatorforcoefnames a character string that will be used to separate feature names when forming names of interaction terms. This is only used when interactions are actually taken into account (see maxinteractionlevel above) and only affects coefficient names in models and columns names in returned featuresinforwardregression (see output value below). We recommend setting it so the interaction separator does not conflict with any feature name since this may improve interpretability of the results. randomseed nthreads NULL or integer. The seed for the random number generator. If NULL, the seed will not be set. If non-null and the random generator has been initialized prior to the function call, the latter s state is saved and restored upon exit. number of threads (worker processes) to perform the calculation. If not given, will be determined automatically as the number of available cores if the latter is 3 or less, and number of cores minus 1 if the number of available cores is 4 or
9 randomglm 9 verbose more. Invalid entries (missing value, zero or negative values etc.) are changed to 1, with a warning. value 0 or 1 which determines the level of verbosity. Zero means silent, 1 reports the bag number the function is working on. At this point verbose output only works if nthreads=1 Details At this point, the function randomglm can be used to predict a binary outcome or a quantitative numeric outcome. This ensemble predictor proceeds along the following steps. Step 1 (bagging): nbags bootstrapped data sets are being generated based on random sampling from the original training data set (x,y). If a bag contains less than mininbagobs unique observations or it contains all observations, it is discarded and re-sampled again. Step 2 (random subspace): For each bag, nfeaturesinbag features are randomly selected (without replacement) from the columns of x. Optionally, interaction terms between the selected features can be formed (see the argument maxinteractionorder). Step 3 (feature ranking): In each bag, features are ranked according to their correlation with the outcome measure. Next the top ncandidatecovariates are being considered for forward selection in each GLM (and in each bag). Step 4 (forward selection): Forward variable selection is employed to define a multivariate GLM model of the outcome in each bag. Step 5 (aggregating the predictions): Prediction from each bag are aggregated. In case, of a quantitative outcome, the predictions are simply averaged across the bags. Generally, ncandidatecovariates>100 is not recommended, because the forward selection process is time-consuming. If arguments "nbags=1, replace=false, nobsinbag=nrow(x)" are used, the function becomes a forward selection GLM predictor without bagging. Classification of multi-level categorical responses is performed indirectly by turning the single multi-class response into a set of binary variables. The set can include two types of binary variables: Level vs. all others (this binary variable is 1 when the original response equals the level and zero otherwise), and level A vs. level B (this binary variable is 0 when the response equals level A, 1 when the response equals level B, and NA otherwise). For example, if the input response y contains observations with values (levels) "A", "B", "C", the binary variables will have names "all.vs.a" (1 means "A", 0 means all others), "all.vs.b", "all.vs.c", and optionally also "A.vs.B" (0 means "A", 1 means "B", NA means neither "A" nor "B"), "A.vs.C", and "B.vs.C". Note that using pairwise level vs. level binary variables be very time-consuming since the number of such binary variables grows quadratically with the number of levels in the response. The user has the option to limit which levels of the original response will have their "own" binary variables, by setting the minimum observations a level must have to qualify for its own binary variable, and by explicitly enumerating levels that should not have their own binary variables. Note that such "ignored" levels are still included on the "all" side of "level vs. all" binary variables. At this time the predictor does not attempt to summarize the binary variable classifications into a single multi-level classification. Training this predictor on data with fewer than 8 observations is not recommended (and the function will warn about it). Due to the bagging step, the number of unique observations in each bag is less than the number of observations in the input data; the low number of unique observations can
10 10 randomglm Value (and often will) lead to an essentially perfect fit which makes it impossible to perfrom meaningful stepwise model selection. Feature names: In general, the column names of input x are assumed to be the feature names. If x has no column names (i.e., colnames(x) is NULL), stadard column names of the form "F01", "F02",... are used. If x has non-null column names, they are turned into valid and unique names using the function make.names. If the function make.names returns names that are not the same as the column names of x, the component featurenameschanged will be TRUE and the component nametranslationtable contains the information about input and actual used feature names. The feature names are used as predictor names in the individual models in each bag. The function returns an object of class randomglm. For continuous prediction or two-level classification, this is a list with the following components: predictedoob the continuous prediction (if classify is FALSE) or predicted classification (if classify is TRUE) of the input data based on out-of-bag samples. predictedoob.response In case of a binary outcome, this is the predicted probability of each outcome specified by y based on out-of-bag samples. In case of a continuous outcome, this is the predicted value based on out-of-bag samples (i.e., a copy of predictedoob). predictedtest.cont if test set is given, the predicted probability of each outcome specified by y for test data for binary outcomes. In case of a continuous outcome, this is the test set predicted value. predictedtest if test set is given, the predicted classification for test data. Only for binary outcomes. candidatefeatures candidate features in each bag. A list with one component per bag. Each component is a matrix with maxinteractionorder rows and ncandidatecovariates columns. Each column represents one interaction obtained by multiplying the features indicated by the entries in each column (0 means no feature, i.e. a lower order interaction). featuresinforwardregression features selected by forward selection in each bag. A list with one component per bag. Each component is a matrix with maxinteractionorder rows. Each column represents one interaction obtained by multiplying the features indicated by the entries in each column (0 means no feature, i.e. a lower order interaction). The column names contain human-readable names for the terms. coefofforwardregression coefficients of forward regression. A list with one component per bag. Each component is a vector giving the coefficients of the model determined by forward selection in the corresponding bag. The order of the coefficients is the same as the order of the terms in the corresponding component of featuresinforwardregression. interceptofforwardregression a vector with one component per bag giving the intercept of the regression model in each bag.
11 randomglm 11 bagobsindx a matrix with nobsinbag rows and nbags columns, giving the indices of observations selected for each bag. timesselectedbyforwardregression a matrix of maxinteractionorder rows and number of features columns. Each entry gives the number of times the corresponding feature appeared in a predictor model at the corresponding order of interactions. Interactions where a single feature enters more than once (e.g., a quadratic interaction of the feature with itself) are counted once. models the regression models for each bag. Predictor features in each bag model are named using their featurenameschanged logical indicating whether feature names were copied verbatim from column names of x (FALSE) or whether they had to be changed to make them valid and unique names (TRUE). nametranslationtable only present if above featurenameschanged is TRUE. A data frame with three columns and one row per input feature (column of input x) giving the feature number, original feature name, and modified feature name that is used for model fitting. In addition, the output value contains a copy of several input arguments. These are included to facilitate prediction using the predict method. These returned values should be considered undocumented and may change in the future. In the multi-level classification classification case, the returned list (still considered a valid randomglm object) contains the following components: binarypredictors a list with one component per binary variable, containing the randomglm predictor trained on that binary variable as the response. The list is named by the corresponding binary variable. For example, if the input response y contains observations with values (levels) "A", "B", "C", the binary variables (and components of this list) will have names "all.vs.a" (1 means "A", 0 means all others), "all.vs.b", "all.vs.c", and optionally also "A.vs.B" (0 means "A", 1 means "B", NA means neither "A" nor "B"), "A.vs.C", and "B.vs.C". predictedoob a matrix in which columns correspond to the binary variables and rows to samples, containing the predicted binary classification for each binary variable. Columns names and meaning of 0 and 1 are described above. predictedoob.response a matrix with two columns per binary variable, giving the class probabilities for each of the two classes in each binary variables. Column names contain the variable and class names. levelmatrix a character matrix with two rows and one column per binary variable, giving the level corresponding to value 0 (row 1) and level corresponding to value 1 (row 2). This encodes the same information as the names of the binarypredictors list but in a more programmer-friendly way. If input xtest is non-null, the components predictedtest and predictedtest.response contain test set predictions analogous to predictedoob and predictedoob.response.
12 12 randomglm Author(s) Lin Song, Steve Horvath, Peter Langfelder. The function makes use of the glm function and other standard R functions. References Lin Song, Peter Langfelder, Steve Horvath: Random generalized linear model: a highly accurate and interpretable ensemble predictor. BMC Bioinformatics (2013) Examples ## binary outcome prediction # data generation data(iris) # Restrict data to first 100 observations iris=iris[1:100,] # Turn Species into a factor iris$species = as.factor(as.character(iris$species)) # Select a training and a test subset of the 100 observations set.seed(1) indx = sample(100, 67, replace=false) xytrain = iris[indx,] xytest = iris[-indx,] xtrain = xytrain[, -5] ytrain = xytrain[, 5] xtest = xytest[, -5] ytest = xytest[, 5] # predict with a small number of bags - normally nbags should be at least 100. RGLM = randomglm(xtrain, ytrain, xtest, ncandidatecovariates=ncol(xtrain), nbags=30, nthreads = 1) ypredicted = RGLM$predictedTest table(ypredicted, ytest) ## continuous outcome prediction x=matrix(rnorm(100*20),100,20) y=rnorm(100) xtrain = x[1:50,] ytrain = y[1:50] xtest = x[51:100,] ytest = y[51:100] RGLM = randomglm(xtrain, ytrain, xtest, classify=false, ncandidatecovariates=ncol(xtrain), nbags=10, keepmodels = TRUE, nthreads = 1)
13 thinrandomglm 13 thinrandomglm Random generalized linear model predictor thinning Description Usage This function allows the user to define a "thinned" version of a random generalized linear model predictor by focusing on those features that occur relatively frequently. thinrandomglm(rglm, threshold) Arguments rglm threshold a randomglm object such as one returned by randomglm. integer specifying the minimum of times a feature was selected across the bags in rglm for the feature to be kept. Note that only features selected threshold +1 times and more are retained. For the purposes of this count, appearances in interactions are not counted. Features that appear threshold times or fewer are removed from the underlying regression models when the models are re-fit. Details Value The function "thins out" (reduces) a previously-constructed random generalized linear model predictor by removing rarely selected features and refitting each (generalized) linear model (GLM). Each GLM (per bag) is refit using only those features that occur more than threshold times across the nbags number of bags. The occurrence count excludes interactions (in other words, the threshold will be applied to the first row of timesselectedbyforwardregression). The function returns a valid randomglm object (see randomglm for details) that can be used as input to the predict() method (see predict.randomglm). The returned object contains a copy of the input rglm in which the following components were modified: predictedoob the updated continuous prediction (if classify is FALSE) or predicted classification (if classify is TRUE) of the input data based on out-of-bag samples. predictedoob.response In case of a binary outcome, the updated predicted probability of each outcome specified by y based on out-of-bag samples. In case of a continuous outcome, this is the predicted value based on out-of-bag samples (i.e., a copy of predictedoob). featuresinforwardregression features selected by forward selection in each bag. A list with one component per bag. Each component is a matrix with maxinteractionorder rows. Each column represents one interaction obtained by multiplying the features indicated by the entries in each column (0 means no feature, i.e. a lower order interaction).
14 14 thinrandomglm coefofforwardregression coefficients of forward regression. A list with one component per bag. Each component is a vector giving the coefficients of the model determined by forward selection in the corresponding bag. The order of the coefficients is the same as the order of the terms in the corresponding component of featuresinforwardregression. interceptofforwardregression a vector with one component per bag giving the intercept of the regression model in each bag. timesselectedbyforwardregression a matrix of maxinteractionorder rows and number of features columns. Each entry gives the number of times the corresponding feature appeared in a predictor model at the corresponding order of interactions. Interactions where a single feature enters more than once (e.g., a quadratic interaction of the feature with itself) are counted once. models Author(s) the "thinned" regression models for each bag. Lin Song, Steve Horvath, Peter Langfelder References Lin Song, Peter Langfelder, Steve Horvath: Random generalized linear model: a highly accurate and interpretable ensemble predictor. BMC Bioinformatics (2013) Examples ## binary outcome prediction # data generation data(iris) # Restrict data to first 100 observations iris=iris[1:100,] # Turn Species into a factor iris$species = as.factor(as.character(iris$species)) # Select a training and a test subset of the 100 observations set.seed(1) indx = sample(100, 67, replace=false) xytrain = iris[indx,] xytest = iris[-indx,] xtrain = xytrain[, -5] ytrain = xytrain[, 5] xtest = xytest[, -5] ytest = xytest[, 5] # predict with a small number of bags - normally nbags should be at least 100. RGLM = randomglm(xtrain, ytrain, ncandidatecovariates=ncol(xtrain), nbags=30, keepmodels = TRUE, nthreads = 1) table(rglm$timesselectedbyforwardregression[1, ]) # # 2 1 1
15 thinrandomglm 15 thinnedrglm = thinrandomglm(rglm, threshold=7) predicted = predict(thinnedrglm, newdata = xtest, type="class") predicted = predict(rglm, newdata = xtest, type="class")
16 Index Topic misc predict.randomglm, 4 randomglm, 5 thinrandomglm, 13 bicor, 8 braincancer, 2 cor, 8 make.names, 10 mini, 3 predict.randomglm, 4, 13 randomglm, 4, 5, 13 thinrandomglm, 13 16
Package obliquerf. February 20, Index 8. Extract variable importance measure
 Package obliquerf February 20, 2015 Title Oblique Random Forests from Recursive Linear Model Splits Version 0.3 Date 2012-08-10 Depends R (>= 2.0.0), stats, ROCR, pls, mda, e1071 Author Bjoern Menze and
Package obliquerf February 20, 2015 Title Oblique Random Forests from Recursive Linear Model Splits Version 0.3 Date 2012-08-10 Depends R (>= 2.0.0), stats, ROCR, pls, mda, e1071 Author Bjoern Menze and
Package randomforest.ddr
 Package randomforest.ddr March 10, 2017 Type Package Title Distributed 'randomforest' for Big Data using 'ddr' API Version 0.1.2 Date 2017-03-09 Author Vishrut Gupta, Arash Fard, Winston Li, Matthew Saltz
Package randomforest.ddr March 10, 2017 Type Package Title Distributed 'randomforest' for Big Data using 'ddr' API Version 0.1.2 Date 2017-03-09 Author Vishrut Gupta, Arash Fard, Winston Li, Matthew Saltz
Package rferns. November 15, 2017
 Package rferns November 15, 2017 Type Package Title Random Ferns Classifier Version 2.0.3 Author Miron B. Kursa Maintainer Miron B. Kursa Description An R implementation of the random
Package rferns November 15, 2017 Type Package Title Random Ferns Classifier Version 2.0.3 Author Miron B. Kursa Maintainer Miron B. Kursa Description An R implementation of the random
Package gbts. February 27, 2017
 Type Package Package gbts February 27, 2017 Title Hyperparameter Search for Gradient Boosted Trees Version 1.2.0 Date 2017-02-26 Author Waley W. J. Liang Maintainer Waley W. J. Liang
Type Package Package gbts February 27, 2017 Title Hyperparameter Search for Gradient Boosted Trees Version 1.2.0 Date 2017-02-26 Author Waley W. J. Liang Maintainer Waley W. J. Liang
Package PropClust. September 15, 2018
 Type Package Title Propensity Clustering and Decomposition Version 1.4-6 Date 2018-09-12 Package PropClust September 15, 2018 Author John Michael O Ranola, Kenneth Lange, Steve Horvath, Peter Langfelder
Type Package Title Propensity Clustering and Decomposition Version 1.4-6 Date 2018-09-12 Package PropClust September 15, 2018 Author John Michael O Ranola, Kenneth Lange, Steve Horvath, Peter Langfelder
Package EFS. R topics documented:
 Package EFS July 24, 2017 Title Tool for Ensemble Feature Selection Description Provides a function to check the importance of a feature based on a dependent classification variable. An ensemble of feature
Package EFS July 24, 2017 Title Tool for Ensemble Feature Selection Description Provides a function to check the importance of a feature based on a dependent classification variable. An ensemble of feature
The supclust Package
 The supclust Package May 18, 2005 Title Supervised Clustering of Genes Version 1.0-5 Date 2005-05-18 Methodology for Supervised Grouping of Predictor Variables Author Marcel Dettling and Martin Maechler
The supclust Package May 18, 2005 Title Supervised Clustering of Genes Version 1.0-5 Date 2005-05-18 Methodology for Supervised Grouping of Predictor Variables Author Marcel Dettling and Martin Maechler
Package RAMP. May 25, 2017
 Type Package Package RAMP May 25, 2017 Title Regularized Generalized Linear Models with Interaction Effects Version 2.0.1 Date 2017-05-24 Author Yang Feng, Ning Hao and Hao Helen Zhang Maintainer Yang
Type Package Package RAMP May 25, 2017 Title Regularized Generalized Linear Models with Interaction Effects Version 2.0.1 Date 2017-05-24 Author Yang Feng, Ning Hao and Hao Helen Zhang Maintainer Yang
Package rocc. February 20, 2015
 Package rocc February 20, 2015 Title ROC based classification Version 1.2 Depends ROCR Author Martin Lauss Description Functions for a classification method based on receiver operating characteristics
Package rocc February 20, 2015 Title ROC based classification Version 1.2 Depends ROCR Author Martin Lauss Description Functions for a classification method based on receiver operating characteristics
Package glmnetutils. August 1, 2017
 Type Package Version 1.1 Title Utilities for 'Glmnet' Package glmnetutils August 1, 2017 Description Provides a formula interface for the 'glmnet' package for elasticnet regression, a method for cross-validating
Type Package Version 1.1 Title Utilities for 'Glmnet' Package glmnetutils August 1, 2017 Description Provides a formula interface for the 'glmnet' package for elasticnet regression, a method for cross-validating
Package bayescl. April 14, 2017
 Package bayescl April 14, 2017 Version 0.0.1 Date 2017-04-10 Title Bayesian Inference on a GPU using OpenCL Author Rok Cesnovar, Erik Strumbelj Maintainer Rok Cesnovar Description
Package bayescl April 14, 2017 Version 0.0.1 Date 2017-04-10 Title Bayesian Inference on a GPU using OpenCL Author Rok Cesnovar, Erik Strumbelj Maintainer Rok Cesnovar Description
Package missforest. February 15, 2013
 Type Package Package missforest February 15, 2013 Title Nonparametric Missing Value Imputation using Random Forest Version 1.3 Date 2012-06-26 Author Maintainer Depends randomforest The function missforest
Type Package Package missforest February 15, 2013 Title Nonparametric Missing Value Imputation using Random Forest Version 1.3 Date 2012-06-26 Author Maintainer Depends randomforest The function missforest
Package sprinter. February 20, 2015
 Type Package Package sprinter February 20, 2015 Title Framework for Screening Prognostic Interactions Version 1.1.0 Date 2014-04-11 Author Isabell Hoffmann Maintainer Isabell Hoffmann
Type Package Package sprinter February 20, 2015 Title Framework for Screening Prognostic Interactions Version 1.1.0 Date 2014-04-11 Author Isabell Hoffmann Maintainer Isabell Hoffmann
Package misclassglm. September 3, 2016
 Type Package Package misclassglm September 3, 2016 Title Computation of Generalized Linear Models with Misclassified Covariates Using Side Information Version 0.2.0 Date 2016-09-02 Author Stephan Dlugosz
Type Package Package misclassglm September 3, 2016 Title Computation of Generalized Linear Models with Misclassified Covariates Using Side Information Version 0.2.0 Date 2016-09-02 Author Stephan Dlugosz
Package logicfs. R topics documented:
 Package logicfs November 21, 2017 Title Identification of SNP Interactions Version 1.48.0 Date 2013-09-12 Author Holger Schwender Maintainer Holger Schwender Depends LogicReg, mcbiopi
Package logicfs November 21, 2017 Title Identification of SNP Interactions Version 1.48.0 Date 2013-09-12 Author Holger Schwender Maintainer Holger Schwender Depends LogicReg, mcbiopi
Package PrivateLR. March 20, 2018
 Type Package Package PrivateLR March 20, 2018 Title Differentially Private Regularized Logistic Regression Version 1.2-22 Date 2018-03-19 Author Staal A. Vinterbo Maintainer Staal
Type Package Package PrivateLR March 20, 2018 Title Differentially Private Regularized Logistic Regression Version 1.2-22 Date 2018-03-19 Author Staal A. Vinterbo Maintainer Staal
Package maboost. R topics documented: February 20, 2015
 Version 1.0-0 Date 2014-11-01 Title Binary and Multiclass Boosting Algorithms Author Tofigh Naghibi Depends R(>= 2.10),rpart,C50 Package maboost February 20, 2015 Performs binary and multiclass boosting
Version 1.0-0 Date 2014-11-01 Title Binary and Multiclass Boosting Algorithms Author Tofigh Naghibi Depends R(>= 2.10),rpart,C50 Package maboost February 20, 2015 Performs binary and multiclass boosting
Package RPEnsemble. October 7, 2017
 Version 0.4 Date 2017-10-06 Title Random Projection Ensemble Classification Package RPEnsemble October 7, 2017 Author Maintainer Timothy I. Cannings Implements the methodology
Version 0.4 Date 2017-10-06 Title Random Projection Ensemble Classification Package RPEnsemble October 7, 2017 Author Maintainer Timothy I. Cannings Implements the methodology
Package ridge. R topics documented: February 15, Title Ridge Regression with automatic selection of the penalty parameter. Version 2.
 Package ridge February 15, 2013 Title Ridge Regression with automatic selection of the penalty parameter Version 2.1-2 Date 2012-25-09 Author Erika Cule Linear and logistic ridge regression for small data
Package ridge February 15, 2013 Title Ridge Regression with automatic selection of the penalty parameter Version 2.1-2 Date 2012-25-09 Author Erika Cule Linear and logistic ridge regression for small data
In stochastic gradient descent implementations, the fixed learning rate η is often replaced by an adaptive learning rate that decreases over time,
 Chapter 2 Although stochastic gradient descent can be considered as an approximation of gradient descent, it typically reaches convergence much faster because of the more frequent weight updates. Since
Chapter 2 Although stochastic gradient descent can be considered as an approximation of gradient descent, it typically reaches convergence much faster because of the more frequent weight updates. Since
Package Modeler. May 18, 2018
 Version 3.4.3 Date 2018-05-17 Package Modeler May 18, 2018 Title Classes and Methods for Training and Using Binary Prediction Models Author Kevin R. Coombes Maintainer Kevin R. Coombes
Version 3.4.3 Date 2018-05-17 Package Modeler May 18, 2018 Title Classes and Methods for Training and Using Binary Prediction Models Author Kevin R. Coombes Maintainer Kevin R. Coombes
Package rpst. June 6, 2017
 Type Package Title Recursive Partitioning Survival Trees Version 1.0.0 Date 2017-6-6 Author Maintainer Package rpst June 6, 2017 An implementation of Recursive Partitioning Survival Trees
Type Package Title Recursive Partitioning Survival Trees Version 1.0.0 Date 2017-6-6 Author Maintainer Package rpst June 6, 2017 An implementation of Recursive Partitioning Survival Trees
Package binomlogit. February 19, 2015
 Type Package Title Efficient MCMC for Binomial Logit Models Version 1.2 Date 2014-03-12 Author Agnes Fussl Maintainer Agnes Fussl Package binomlogit February 19, 2015 Description The R package
Type Package Title Efficient MCMC for Binomial Logit Models Version 1.2 Date 2014-03-12 Author Agnes Fussl Maintainer Agnes Fussl Package binomlogit February 19, 2015 Description The R package
Supplementary text S6 Comparison studies on simulated data
 Supplementary text S Comparison studies on simulated data Peter Langfelder, Rui Luo, Michael C. Oldham, and Steve Horvath Corresponding author: shorvath@mednet.ucla.edu Overview In this document we illustrate
Supplementary text S Comparison studies on simulated data Peter Langfelder, Rui Luo, Michael C. Oldham, and Steve Horvath Corresponding author: shorvath@mednet.ucla.edu Overview In this document we illustrate
Package mvprobit. November 2, 2015
 Version 0.1-8 Date 2015-11-02 Title Multivariate Probit Models Package mvprobit November 2, 2015 Author Arne Henningsen Maintainer Arne Henningsen
Version 0.1-8 Date 2015-11-02 Title Multivariate Probit Models Package mvprobit November 2, 2015 Author Arne Henningsen Maintainer Arne Henningsen
Package nodeharvest. June 12, 2015
 Type Package Package nodeharvest June 12, 2015 Title Node Harvest for Regression and Classification Version 0.7-3 Date 2015-06-10 Author Nicolai Meinshausen Maintainer Nicolai Meinshausen
Type Package Package nodeharvest June 12, 2015 Title Node Harvest for Regression and Classification Version 0.7-3 Date 2015-06-10 Author Nicolai Meinshausen Maintainer Nicolai Meinshausen
Package gensemble. R topics documented: February 19, 2015
 Type Package Title generalized ensemble methods Version 1.0 Date 2013-01-25 Author Peter Werner, Eugene Dubossarsky Package gensemble February 19, 2015 Maintainer Peter Werner Generalized
Type Package Title generalized ensemble methods Version 1.0 Date 2013-01-25 Author Peter Werner, Eugene Dubossarsky Package gensemble February 19, 2015 Maintainer Peter Werner Generalized
Package TANDEM. R topics documented: June 15, Type Package
 Type Package Package TANDEM June 15, 2017 Title A Two-Stage Approach to Maximize Interpretability of Drug Response Models Based on Multiple Molecular Data Types Version 1.0.2 Date 2017-04-07 Author Nanne
Type Package Package TANDEM June 15, 2017 Title A Two-Stage Approach to Maximize Interpretability of Drug Response Models Based on Multiple Molecular Data Types Version 1.0.2 Date 2017-04-07 Author Nanne
Package kerasformula
 Package kerasformula August 23, 2018 Type Package Title A High-Level R Interface for Neural Nets Version 1.5.1 Author Pete Mohanty [aut, cre] Maintainer Pete Mohanty Description
Package kerasformula August 23, 2018 Type Package Title A High-Level R Interface for Neural Nets Version 1.5.1 Author Pete Mohanty [aut, cre] Maintainer Pete Mohanty Description
Package Cubist. December 2, 2017
 Type Package Package Cubist December 2, 2017 Title Rule- And Instance-Based Regression Modeling Version 0.2.1 Date 2017-12-01 Maintainer Max Kuhn Description Regression modeling using
Type Package Package Cubist December 2, 2017 Title Rule- And Instance-Based Regression Modeling Version 0.2.1 Date 2017-12-01 Maintainer Max Kuhn Description Regression modeling using
Package GWRM. R topics documented: July 31, Type Package
 Type Package Package GWRM July 31, 2017 Title Generalized Waring Regression Model for Count Data Version 2.1.0.3 Date 2017-07-18 Maintainer Antonio Jose Saez-Castillo Depends R (>= 3.0.0)
Type Package Package GWRM July 31, 2017 Title Generalized Waring Regression Model for Count Data Version 2.1.0.3 Date 2017-07-18 Maintainer Antonio Jose Saez-Castillo Depends R (>= 3.0.0)
Package naivebayes. R topics documented: January 3, Type Package. Title High Performance Implementation of the Naive Bayes Algorithm
 Package naivebayes January 3, 2018 Type Package Title High Performance Implementation of the Naive Bayes Algorithm Version 0.9.2 Author Michal Majka Maintainer Michal Majka Description
Package naivebayes January 3, 2018 Type Package Title High Performance Implementation of the Naive Bayes Algorithm Version 0.9.2 Author Michal Majka Maintainer Michal Majka Description
Package enpls. May 14, 2018
 Package enpls May 14, 2018 Type Package Title Ensemble Partial Least Squares Regression Version 6.0 Maintainer Nan Xiao An algorithmic framework for measuring feature importance, outlier detection,
Package enpls May 14, 2018 Type Package Title Ensemble Partial Least Squares Regression Version 6.0 Maintainer Nan Xiao An algorithmic framework for measuring feature importance, outlier detection,
Package hiernet. March 18, 2018
 Title A Lasso for Hierarchical Interactions Version 1.7 Author Jacob Bien and Rob Tibshirani Package hiernet March 18, 2018 Fits sparse interaction models for continuous and binary responses subject to
Title A Lasso for Hierarchical Interactions Version 1.7 Author Jacob Bien and Rob Tibshirani Package hiernet March 18, 2018 Fits sparse interaction models for continuous and binary responses subject to
Package parcor. February 20, 2015
 Type Package Package parcor February 20, 2015 Title Regularized estimation of partial correlation matrices Version 0.2-6 Date 2014-09-04 Depends MASS, glmnet, ppls, Epi, GeneNet Author, Juliane Schaefer
Type Package Package parcor February 20, 2015 Title Regularized estimation of partial correlation matrices Version 0.2-6 Date 2014-09-04 Depends MASS, glmnet, ppls, Epi, GeneNet Author, Juliane Schaefer
Package cgh. R topics documented: February 19, 2015
 Package cgh February 19, 2015 Version 1.0-7.1 Date 2009-11-20 Title Microarray CGH analysis using the Smith-Waterman algorithm Author Tom Price Maintainer Tom Price
Package cgh February 19, 2015 Version 1.0-7.1 Date 2009-11-20 Title Microarray CGH analysis using the Smith-Waterman algorithm Author Tom Price Maintainer Tom Price
Final Exam. Advanced Methods for Data Analysis (36-402/36-608) Due Thursday May 8, 2014 at 11:59pm
 Final Exam Advanced Methods for Data Analysis (36-402/36-608) Due Thursday May 8, 2014 at 11:59pm Instructions: you will submit this take-home final exam in three parts. 1. Writeup. This will be a complete
Final Exam Advanced Methods for Data Analysis (36-402/36-608) Due Thursday May 8, 2014 at 11:59pm Instructions: you will submit this take-home final exam in three parts. 1. Writeup. This will be a complete
Package tspair. July 18, 2013
 Package tspair July 18, 2013 Title Top Scoring Pairs for Microarray Classification Version 1.18.0 Author These functions calculate the pair of genes that show the maximum difference in ranking between
Package tspair July 18, 2013 Title Top Scoring Pairs for Microarray Classification Version 1.18.0 Author These functions calculate the pair of genes that show the maximum difference in ranking between
Applying Supervised Learning
 Applying Supervised Learning When to Consider Supervised Learning A supervised learning algorithm takes a known set of input data (the training set) and known responses to the data (output), and trains
Applying Supervised Learning When to Consider Supervised Learning A supervised learning algorithm takes a known set of input data (the training set) and known responses to the data (output), and trains
Package EBglmnet. January 30, 2016
 Type Package Package EBglmnet January 30, 2016 Title Empirical Bayesian Lasso and Elastic Net Methods for Generalized Linear Models Version 4.1 Date 2016-01-15 Author Anhui Huang, Dianting Liu Maintainer
Type Package Package EBglmnet January 30, 2016 Title Empirical Bayesian Lasso and Elastic Net Methods for Generalized Linear Models Version 4.1 Date 2016-01-15 Author Anhui Huang, Dianting Liu Maintainer
Package ranger. November 10, 2015
 Type Package Title A Fast Implementation of Random Forests Version 0.3.0 Date 2015-11-10 Author Package November 10, 2015 Maintainer A fast implementation of Random Forests,
Type Package Title A Fast Implementation of Random Forests Version 0.3.0 Date 2015-11-10 Author Package November 10, 2015 Maintainer A fast implementation of Random Forests,
Package reghelper. April 8, 2017
 Type Package Title Helper Functions for Regression Analysis Version 0.3.3 Date 2017-04-07 Package reghelper April 8, 2017 A set of functions used to automate commonly used methods in regression analysis.
Type Package Title Helper Functions for Regression Analysis Version 0.3.3 Date 2017-04-07 Package reghelper April 8, 2017 A set of functions used to automate commonly used methods in regression analysis.
Package kernelfactory
 Package kernelfactory September 29, 2015 Type Package Title Kernel Factory: An Ensemble of Kernel Machines Version 0.3.0 Date 2015-09-29 Imports randomforest, AUC, genalg, kernlab, stats Author Michel
Package kernelfactory September 29, 2015 Type Package Title Kernel Factory: An Ensemble of Kernel Machines Version 0.3.0 Date 2015-09-29 Imports randomforest, AUC, genalg, kernlab, stats Author Michel
Linear Methods for Regression and Shrinkage Methods
 Linear Methods for Regression and Shrinkage Methods Reference: The Elements of Statistical Learning, by T. Hastie, R. Tibshirani, J. Friedman, Springer 1 Linear Regression Models Least Squares Input vectors
Linear Methods for Regression and Shrinkage Methods Reference: The Elements of Statistical Learning, by T. Hastie, R. Tibshirani, J. Friedman, Springer 1 Linear Regression Models Least Squares Input vectors
2 cubist.default. Fit a Cubist model
 Package Cubist August 29, 2013 Type Package Title Rule- and Instance-Based Regression Modeling Version 0.0.13 Date 2013-01-28 Author Max Kuhn, Steve Weston, Chris Keefer, Nathan Coulter. C code for Cubist
Package Cubist August 29, 2013 Type Package Title Rule- and Instance-Based Regression Modeling Version 0.0.13 Date 2013-01-28 Author Max Kuhn, Steve Weston, Chris Keefer, Nathan Coulter. C code for Cubist
Package glinternet. June 15, 2018
 Type Package Package glinternet June 15, 2018 Title Learning Interactions via Hierarchical Group-Lasso Regularization Version 1.0.8 Date 2018-06-20 Author Michael Lim, Trevor Hastie Maintainer Michael
Type Package Package glinternet June 15, 2018 Title Learning Interactions via Hierarchical Group-Lasso Regularization Version 1.0.8 Date 2018-06-20 Author Michael Lim, Trevor Hastie Maintainer Michael
Package DTRreg. August 30, 2017
 Package DTRreg August 30, 2017 Type Package Title DTR Estimation and Inference via G-Estimation, Dynamic WOLS, and Q-Learning Version 1.3 Date 2017-08-30 Author Michael Wallace, Erica E M Moodie, David
Package DTRreg August 30, 2017 Type Package Title DTR Estimation and Inference via G-Estimation, Dynamic WOLS, and Q-Learning Version 1.3 Date 2017-08-30 Author Michael Wallace, Erica E M Moodie, David
Package C50. December 1, 2017
 Type Package Title C5.0 Decision Trees and Rule-Based Models Version 0.1.1 Date 2017-11-20 Maintainer Max Kuhn Package C50 December 1, 2017 C5.0 decision trees and rulebased models for
Type Package Title C5.0 Decision Trees and Rule-Based Models Version 0.1.1 Date 2017-11-20 Maintainer Max Kuhn Package C50 December 1, 2017 C5.0 decision trees and rulebased models for
stepwisecm: Stepwise Classification of Cancer Samples using High-dimensional Data Sets
 stepwisecm: Stepwise Classification of Cancer Samples using High-dimensional Data Sets Askar Obulkasim Department of Epidemiology and Biostatistics, VU University Medical Center P.O. Box 7075, 1007 MB
stepwisecm: Stepwise Classification of Cancer Samples using High-dimensional Data Sets Askar Obulkasim Department of Epidemiology and Biostatistics, VU University Medical Center P.O. Box 7075, 1007 MB
K-fold cross validation in the Tidyverse Stephanie J. Spielman 11/7/2017
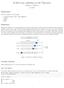 K-fold cross validation in the Tidyverse Stephanie J. Spielman 11/7/2017 Requirements This demo requires several packages: tidyverse (dplyr, tidyr, tibble, ggplot2) modelr broom proc Background K-fold
K-fold cross validation in the Tidyverse Stephanie J. Spielman 11/7/2017 Requirements This demo requires several packages: tidyverse (dplyr, tidyr, tibble, ggplot2) modelr broom proc Background K-fold
Package arulescba. April 24, 2018
 Version 1.1.3-1 Date 2018-04-23 Title Classification Based on Association Rules Package arulescba April 24, 2018 Provides a function to build an association rulebased classifier for data frames, and to
Version 1.1.3-1 Date 2018-04-23 Title Classification Based on Association Rules Package arulescba April 24, 2018 Provides a function to build an association rulebased classifier for data frames, and to
Package spikeslab. February 20, 2015
 Version 1.1.5 Date 2013-04-18 Package spikeslab February 20, 2015 Title Prediction and variable selection using spike and slab regression Author Hemant Ishwaran Maintainer Udaya
Version 1.1.5 Date 2013-04-18 Package spikeslab February 20, 2015 Title Prediction and variable selection using spike and slab regression Author Hemant Ishwaran Maintainer Udaya
Package subsemble. February 20, 2015
 Type Package Package subsemble February 20, 2015 Title An Ensemble Method for Combining Subset-Specific Algorithm Fits Version 0.0.9 Date 2014-07-01 Author Erin LeDell, Stephanie Sapp, Mark van der Laan
Type Package Package subsemble February 20, 2015 Title An Ensemble Method for Combining Subset-Specific Algorithm Fits Version 0.0.9 Date 2014-07-01 Author Erin LeDell, Stephanie Sapp, Mark van der Laan
Package SAENET. June 4, 2015
 Type Package Package SAENET June 4, 2015 Title A Stacked Autoencoder Implementation with Interface to 'neuralnet' Version 1.1 Date 2015-06-04 An implementation of a stacked sparse autoencoder for dimension
Type Package Package SAENET June 4, 2015 Title A Stacked Autoencoder Implementation with Interface to 'neuralnet' Version 1.1 Date 2015-06-04 An implementation of a stacked sparse autoencoder for dimension
Package ordinalforest
 Type Package Package ordinalforest July 16, 2018 Title Ordinal Forests: Prediction and Variable Ranking with Ordinal Target Variables Version 2.2 Date 2018-07-16 Author Roman Hornung Maintainer Roman Hornung
Type Package Package ordinalforest July 16, 2018 Title Ordinal Forests: Prediction and Variable Ranking with Ordinal Target Variables Version 2.2 Date 2018-07-16 Author Roman Hornung Maintainer Roman Hornung
Package nscancor. February 15, 2018
 Version 0.6.1-25 Title Non-Negative and Sparse CCA Package nscancor February 15, 2018 Description Two implementations of canonical correlation analysis (CCA) that are based on iterated regression. By choosing
Version 0.6.1-25 Title Non-Negative and Sparse CCA Package nscancor February 15, 2018 Description Two implementations of canonical correlation analysis (CCA) that are based on iterated regression. By choosing
Package knnflex. April 17, 2009
 Package knnflex April 17, 2009 Type Package Title A more flexible KNN Version 1.1.1 Depends R (>= 2.4.0) Date 2007-02-01 Author Atina Dunlap Brooks Maintainer ORPHANED Description A KNN implementaion which
Package knnflex April 17, 2009 Type Package Title A more flexible KNN Version 1.1.1 Depends R (>= 2.4.0) Date 2007-02-01 Author Atina Dunlap Brooks Maintainer ORPHANED Description A KNN implementaion which
Package PTE. October 10, 2017
 Type Package Title Personalized Treatment Evaluator Version 1.6 Date 2017-10-9 Package PTE October 10, 2017 Author Adam Kapelner, Alina Levine & Justin Bleich Maintainer Adam Kapelner
Type Package Title Personalized Treatment Evaluator Version 1.6 Date 2017-10-9 Package PTE October 10, 2017 Author Adam Kapelner, Alina Levine & Justin Bleich Maintainer Adam Kapelner
Package texteffect. November 27, 2017
 Version 0.1 Date 2017-11-27 Package texteffect November 27, 2017 Title Discovering Latent Treatments in Text Corpora and Estimating Their Causal Effects Author Christian Fong
Version 0.1 Date 2017-11-27 Package texteffect November 27, 2017 Title Discovering Latent Treatments in Text Corpora and Estimating Their Causal Effects Author Christian Fong
Package caretensemble
 Package caretensemble Type Package Title Ensembles of Caret Models Version 2.0.0 Date 2016-02-06 August 29, 2016 URL https://github.com/zachmayer/caretensemble BugReports https://github.com/zachmayer/caretensemble/issues
Package caretensemble Type Package Title Ensembles of Caret Models Version 2.0.0 Date 2016-02-06 August 29, 2016 URL https://github.com/zachmayer/caretensemble BugReports https://github.com/zachmayer/caretensemble/issues
Package blockforest. December 30, 2018
 Type Package Package blockforest December 30, 2018 Title Block Forests: Random Forests for Blocks of Clinical and Omics Covariate Data Version 0.1.7 Date 2018-12-18 Author Roman Hornung, Marvin N. Wright
Type Package Package blockforest December 30, 2018 Title Block Forests: Random Forests for Blocks of Clinical and Omics Covariate Data Version 0.1.7 Date 2018-12-18 Author Roman Hornung, Marvin N. Wright
Package glmmml. R topics documented: March 25, Encoding UTF-8 Version Date Title Generalized Linear Models with Clustering
 Encoding UTF-8 Version 1.0.3 Date 2018-03-25 Title Generalized Linear Models with Clustering Package glmmml March 25, 2018 Binomial and Poisson regression for clustered data, fixed and random effects with
Encoding UTF-8 Version 1.0.3 Date 2018-03-25 Title Generalized Linear Models with Clustering Package glmmml March 25, 2018 Binomial and Poisson regression for clustered data, fixed and random effects with
Resampling Methods. Levi Waldron, CUNY School of Public Health. July 13, 2016
 Resampling Methods Levi Waldron, CUNY School of Public Health July 13, 2016 Outline and introduction Objectives: prediction or inference? Cross-validation Bootstrap Permutation Test Monte Carlo Simulation
Resampling Methods Levi Waldron, CUNY School of Public Health July 13, 2016 Outline and introduction Objectives: prediction or inference? Cross-validation Bootstrap Permutation Test Monte Carlo Simulation
Package intccr. September 12, 2017
 Type Package Package intccr September 12, 2017 Title Semiparametric Competing Risks Regression under Interval Censoring Version 0.2.0 Author Giorgos Bakoyannis , Jun Park
Type Package Package intccr September 12, 2017 Title Semiparametric Competing Risks Regression under Interval Censoring Version 0.2.0 Author Giorgos Bakoyannis , Jun Park
Machine Learning: An Applied Econometric Approach Online Appendix
 Machine Learning: An Applied Econometric Approach Online Appendix Sendhil Mullainathan mullain@fas.harvard.edu Jann Spiess jspiess@fas.harvard.edu April 2017 A How We Predict In this section, we detail
Machine Learning: An Applied Econometric Approach Online Appendix Sendhil Mullainathan mullain@fas.harvard.edu Jann Spiess jspiess@fas.harvard.edu April 2017 A How We Predict In this section, we detail
Linear discriminant analysis and logistic
 Practical 6: classifiers Linear discriminant analysis and logistic This practical looks at two different methods of fitting linear classifiers. The linear discriminant analysis is implemented in the MASS
Practical 6: classifiers Linear discriminant analysis and logistic This practical looks at two different methods of fitting linear classifiers. The linear discriminant analysis is implemented in the MASS
Package TVsMiss. April 5, 2018
 Type Package Title Variable Selection for Missing Data Version 0.1.1 Date 2018-04-05 Author Jiwei Zhao, Yang Yang, and Ning Yang Maintainer Yang Yang Package TVsMiss April 5, 2018
Type Package Title Variable Selection for Missing Data Version 0.1.1 Date 2018-04-05 Author Jiwei Zhao, Yang Yang, and Ning Yang Maintainer Yang Yang Package TVsMiss April 5, 2018
Package MTLR. March 9, 2019
 Type Package Package MTLR March 9, 2019 Title Survival Prediction with Multi-Task Logistic Regression Version 0.2.0 Author Humza Haider Maintainer Humza Haider URL https://github.com/haiderstats/mtlr
Type Package Package MTLR March 9, 2019 Title Survival Prediction with Multi-Task Logistic Regression Version 0.2.0 Author Humza Haider Maintainer Humza Haider URL https://github.com/haiderstats/mtlr
Hierarchical Clustering
 What is clustering Partitioning of a data set into subsets. A cluster is a group of relatively homogeneous cases or observations Hierarchical Clustering Mikhail Dozmorov Fall 2016 2/61 What is clustering
What is clustering Partitioning of a data set into subsets. A cluster is a group of relatively homogeneous cases or observations Hierarchical Clustering Mikhail Dozmorov Fall 2016 2/61 What is clustering
Package clustvarsel. April 9, 2018
 Version 2.3.2 Date 2018-04-09 Package clustvarsel April 9, 2018 Title Variable Selection for Gaussian Model-Based Clustering Description Variable selection for Gaussian model-based clustering as implemented
Version 2.3.2 Date 2018-04-09 Package clustvarsel April 9, 2018 Title Variable Selection for Gaussian Model-Based Clustering Description Variable selection for Gaussian model-based clustering as implemented
Package gee. June 29, 2015
 Title Generalized Estimation Equation Solver Version 4.13-19 Depends stats Suggests MASS Date 2015-06-29 DateNote Gee version 1998-01-27 Package gee June 29, 2015 Author Vincent J Carey. Ported to R by
Title Generalized Estimation Equation Solver Version 4.13-19 Depends stats Suggests MASS Date 2015-06-29 DateNote Gee version 1998-01-27 Package gee June 29, 2015 Author Vincent J Carey. Ported to R by
Big Data Methods. Chapter 5: Machine learning. Big Data Methods, Chapter 5, Slide 1
 Big Data Methods Chapter 5: Machine learning Big Data Methods, Chapter 5, Slide 1 5.1 Introduction to machine learning What is machine learning? Concerned with the study and development of algorithms that
Big Data Methods Chapter 5: Machine learning Big Data Methods, Chapter 5, Slide 1 5.1 Introduction to machine learning What is machine learning? Concerned with the study and development of algorithms that
Package automl. September 13, 2018
 Type Package Title Deep Learning with Metaheuristic Version 1.0.5 Author Alex Boulangé Package automl September 13, 2018 Maintainer Alex Boulangé Fits from
Type Package Title Deep Learning with Metaheuristic Version 1.0.5 Author Alex Boulangé Package automl September 13, 2018 Maintainer Alex Boulangé Fits from
Package StVAR. February 11, 2017
 Type Package Title Student's t Vector Autoregression (StVAR) Version 1.1 Date 2017-02-10 Author Niraj Poudyal Maintainer Niraj Poudyal Package StVAR February 11, 2017 Description Estimation
Type Package Title Student's t Vector Autoregression (StVAR) Version 1.1 Date 2017-02-10 Author Niraj Poudyal Maintainer Niraj Poudyal Package StVAR February 11, 2017 Description Estimation
Package BigTSP. February 19, 2015
 Type Package Package BigTSP February 19, 2015 Title Top Scoring Pair based methods for classification Version 1.0 Date 2012-08-20 Author Xiaolin Yang,Han Liu Maintainer Xiaolin Yang
Type Package Package BigTSP February 19, 2015 Title Top Scoring Pair based methods for classification Version 1.0 Date 2012-08-20 Author Xiaolin Yang,Han Liu Maintainer Xiaolin Yang
Package svmpath. R topics documented: August 30, Title The SVM Path Algorithm Date Version Author Trevor Hastie
 Title The SVM Path Algorithm Date 2016-08-29 Version 0.955 Author Package svmpath August 30, 2016 Computes the entire regularization path for the two-class svm classifier with essentially the same cost
Title The SVM Path Algorithm Date 2016-08-29 Version 0.955 Author Package svmpath August 30, 2016 Computes the entire regularization path for the two-class svm classifier with essentially the same cost
Package fasttextr. May 12, 2017
 Type Package Title An Interface to the 'fasttext' Library Version 1.0 Date 2016-09-22 Author Florian Schwendinger [aut, cre] Package fasttextr May 12, 2017 Maintainer Florian Schwendinger
Type Package Title An Interface to the 'fasttext' Library Version 1.0 Date 2016-09-22 Author Florian Schwendinger [aut, cre] Package fasttextr May 12, 2017 Maintainer Florian Schwendinger
Package apricom. R topics documented: November 24, 2015
 Package apricom November 24, 2015 Title Tools for the a Priori Comparison of Regression Modelling Strategies Version 1.0.0 Date 2015-11-11 Maintainer Romin Pajouheshnia Tools
Package apricom November 24, 2015 Title Tools for the a Priori Comparison of Regression Modelling Strategies Version 1.0.0 Date 2015-11-11 Maintainer Romin Pajouheshnia Tools
Package RSNPset. December 14, 2017
 Type Package Package RSNPset December 14, 2017 Title Efficient Score Statistics for Genome-Wide SNP Set Analysis Version 0.5.3 Date 2017-12-11 Author Chanhee Yi, Alexander Sibley, and Kouros Owzar Maintainer
Type Package Package RSNPset December 14, 2017 Title Efficient Score Statistics for Genome-Wide SNP Set Analysis Version 0.5.3 Date 2017-12-11 Author Chanhee Yi, Alexander Sibley, and Kouros Owzar Maintainer
Package RaPKod. February 5, 2018
 Package RaPKod February 5, 2018 Type Package Title Random Projection Kernel Outlier Detector Version 0.9 Date 2018-01-30 Author Jeremie Kellner Maintainer Jeremie Kellner
Package RaPKod February 5, 2018 Type Package Title Random Projection Kernel Outlier Detector Version 0.9 Date 2018-01-30 Author Jeremie Kellner Maintainer Jeremie Kellner
Package mgc. April 13, 2018
 Type Package Title Multiscale Graph Correlation Version 1.0.1 Date 2018-04-12 Package mgc April 13, 2018 Maintainer Multiscale Graph Correlation (MGC) is a framework developed by Shen
Type Package Title Multiscale Graph Correlation Version 1.0.1 Date 2018-04-12 Package mgc April 13, 2018 Maintainer Multiscale Graph Correlation (MGC) is a framework developed by Shen
Package milr. June 8, 2017
 Type Package Package milr June 8, 2017 Title Multiple-Instance Logistic Regression with LASSO Penalty Version 0.3.0 Date 2017-06-05 The multiple instance data set consists of many independent subjects
Type Package Package milr June 8, 2017 Title Multiple-Instance Logistic Regression with LASSO Penalty Version 0.3.0 Date 2017-06-05 The multiple instance data set consists of many independent subjects
k-nn classification with R QMMA
 k-nn classification with R QMMA Emanuele Taufer file:///c:/users/emanuele.taufer/google%20drive/2%20corsi/5%20qmma%20-%20mim/0%20labs/l1-knn-eng.html#(1) 1/16 HW (Height and weight) of adults Statistics
k-nn classification with R QMMA Emanuele Taufer file:///c:/users/emanuele.taufer/google%20drive/2%20corsi/5%20qmma%20-%20mim/0%20labs/l1-knn-eng.html#(1) 1/16 HW (Height and weight) of adults Statistics
Package sparsereg. R topics documented: March 10, Type Package
 Type Package Package sparsereg March 10, 2016 Title Sparse Bayesian Models for Regression, Subgroup Analysis, and Panel Data Version 1.2 Date 2016-03-01 Author Marc Ratkovic and Dustin Tingley Maintainer
Type Package Package sparsereg March 10, 2016 Title Sparse Bayesian Models for Regression, Subgroup Analysis, and Panel Data Version 1.2 Date 2016-03-01 Author Marc Ratkovic and Dustin Tingley Maintainer
Package JOUSBoost. July 12, 2017
 Type Package Package JOUSBoost July 12, 2017 Title Implements Under/Oversampling for Probability Estimation Version 2.1.0 Implements under/oversampling for probability estimation. To be used with machine
Type Package Package JOUSBoost July 12, 2017 Title Implements Under/Oversampling for Probability Estimation Version 2.1.0 Implements under/oversampling for probability estimation. To be used with machine
Supervised vs unsupervised clustering
 Classification Supervised vs unsupervised clustering Cluster analysis: Classes are not known a- priori. Classification: Classes are defined a-priori Sometimes called supervised clustering Extract useful
Classification Supervised vs unsupervised clustering Cluster analysis: Classes are not known a- priori. Classification: Classes are defined a-priori Sometimes called supervised clustering Extract useful
The Generalized Topological Overlap Matrix in Biological Network Analysis
 The Generalized Topological Overlap Matrix in Biological Network Analysis Andy Yip, Steve Horvath Email: shorvath@mednet.ucla.edu Depts Human Genetics and Biostatistics, University of California, Los Angeles
The Generalized Topological Overlap Matrix in Biological Network Analysis Andy Yip, Steve Horvath Email: shorvath@mednet.ucla.edu Depts Human Genetics and Biostatistics, University of California, Los Angeles
Package hsiccca. February 20, 2015
 Package hsiccca February 20, 2015 Type Package Title Canonical Correlation Analysis based on Kernel Independence Measures Version 1.0 Date 2013-03-13 Author Billy Chang Maintainer Billy Chang
Package hsiccca February 20, 2015 Type Package Title Canonical Correlation Analysis based on Kernel Independence Measures Version 1.0 Date 2013-03-13 Author Billy Chang Maintainer Billy Chang
Fast or furious? - User analysis of SF Express Inc
 CS 229 PROJECT, DEC. 2017 1 Fast or furious? - User analysis of SF Express Inc Gege Wen@gegewen, Yiyuan Zhang@yiyuan12, Kezhen Zhao@zkz I. MOTIVATION The motivation of this project is to predict the likelihood
CS 229 PROJECT, DEC. 2017 1 Fast or furious? - User analysis of SF Express Inc Gege Wen@gegewen, Yiyuan Zhang@yiyuan12, Kezhen Zhao@zkz I. MOTIVATION The motivation of this project is to predict the likelihood
Package ESKNN. September 13, 2015
 Type Package Package ESKNN September 13, 2015 Title Ensemble of Subset of K-Nearest Neighbours Classifiers for Classification and Class Membership Probability Estimation Version 1.0 Date 2015-09-13 Author
Type Package Package ESKNN September 13, 2015 Title Ensemble of Subset of K-Nearest Neighbours Classifiers for Classification and Class Membership Probability Estimation Version 1.0 Date 2015-09-13 Author
Machine Learning with MATLAB --classification
 Machine Learning with MATLAB --classification Stanley Liang, PhD York University Classification the definition In machine learning and statistics, classification is the problem of identifying to which
Machine Learning with MATLAB --classification Stanley Liang, PhD York University Classification the definition In machine learning and statistics, classification is the problem of identifying to which
The grplasso Package
 The grplasso Package June 27, 2007 Type Package Title Fitting user specified models with Group Lasso penalty Version 0.2-1 Date 2007-06-27 Author Lukas Meier Maintainer Lukas Meier
The grplasso Package June 27, 2007 Type Package Title Fitting user specified models with Group Lasso penalty Version 0.2-1 Date 2007-06-27 Author Lukas Meier Maintainer Lukas Meier
Package RCA. R topics documented: February 29, 2016
 Type Package Title Relational Class Analysis Version 2.0 Date 2016-02-25 Author Amir Goldberg, Sarah K. Stein Package RCA February 29, 2016 Maintainer Amir Goldberg Depends igraph,
Type Package Title Relational Class Analysis Version 2.0 Date 2016-02-25 Author Amir Goldberg, Sarah K. Stein Package RCA February 29, 2016 Maintainer Amir Goldberg Depends igraph,
Package sure. September 19, 2017
 Type Package Package sure September 19, 2017 Title Surrogate Residuals for Ordinal and General Regression Models An implementation of the surrogate approach to residuals and diagnostics for ordinal and
Type Package Package sure September 19, 2017 Title Surrogate Residuals for Ordinal and General Regression Models An implementation of the surrogate approach to residuals and diagnostics for ordinal and
Package TPD. June 14, 2018
 Type Package Package TPD June 14, 2018 Title Methods for Measuring Functional Diversity Based on Trait Probability Density Version 1.0.0 Date 2018-06-13 Author Carlos P. Carmona
Type Package Package TPD June 14, 2018 Title Methods for Measuring Functional Diversity Based on Trait Probability Density Version 1.0.0 Date 2018-06-13 Author Carlos P. Carmona
RESAMPLING METHODS. Chapter 05
 1 RESAMPLING METHODS Chapter 05 2 Outline Cross Validation The Validation Set Approach Leave-One-Out Cross Validation K-fold Cross Validation Bias-Variance Trade-off for k-fold Cross Validation Cross Validation
1 RESAMPLING METHODS Chapter 05 2 Outline Cross Validation The Validation Set Approach Leave-One-Out Cross Validation K-fold Cross Validation Bias-Variance Trade-off for k-fold Cross Validation Cross Validation
Stat 342 Exam 3 Fall 2014
 Stat 34 Exam 3 Fall 04 I have neither given nor received unauthorized assistance on this exam. Name Signed Date Name Printed There are questions on the following 6 pages. Do as many of them as you can
Stat 34 Exam 3 Fall 04 I have neither given nor received unauthorized assistance on this exam. Name Signed Date Name Printed There are questions on the following 6 pages. Do as many of them as you can
Package rgcvpack. February 20, Index 6. Fitting Thin Plate Smoothing Spline. Fit thin plate splines of any order with user specified knots
 Version 0.1-4 Date 2013/10/25 Title R Interface for GCVPACK Fortran Package Author Xianhong Xie Package rgcvpack February 20, 2015 Maintainer Xianhong Xie
Version 0.1-4 Date 2013/10/25 Title R Interface for GCVPACK Fortran Package Author Xianhong Xie Package rgcvpack February 20, 2015 Maintainer Xianhong Xie
Package nricens. R topics documented: May 30, Type Package
 Type Package Package nricens May 30, 2018 Title NRI for Risk Prediction Models with Time to Event and Binary Response Data Version 1.6 Date 2018-5-30 Author Eisuke Inoue Depends survival Maintainer Eisuke
Type Package Package nricens May 30, 2018 Title NRI for Risk Prediction Models with Time to Event and Binary Response Data Version 1.6 Date 2018-5-30 Author Eisuke Inoue Depends survival Maintainer Eisuke
Package subtype. February 20, 2015
 Type Package Package subtype February 20, 2015 Title Cluster analysis to find molecular subtypes and their assessment Version 1.0 Date 2013-01-14 Author Andrey Alexeyenko, Woojoo Lee and Yudi Pawitan Maintainer
Type Package Package subtype February 20, 2015 Title Cluster analysis to find molecular subtypes and their assessment Version 1.0 Date 2013-01-14 Author Andrey Alexeyenko, Woojoo Lee and Yudi Pawitan Maintainer
