NacoStringQCPro. Dorothee Nickles, Thomas Sandmann, Robert Ziman, Richard Bourgon. Modified: April 10, Compiled: April 24, 2017.
|
|
|
- Amber Pope
- 5 years ago
- Views:
Transcription
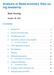
1 NacoStringQCPro Dorothee Nickles, Thomas Sandmann, Robert Ziman, Richard Bourgon Modified: April 10, Compiled: April 24, 2017 Contents 1 Scope 1 2 Quick start 1 3 RccSet loaded with NanoString data Sample annotation Codeset annotation Preprocessing Positive control normalization Background correction RNA content normalization QC report and quality control of NanoString expression data 9 6 Example files 9 7 Session info 9 1 Scope The NanoStringQCPro package facilitates preprocessing and quality control of NanoString gene expression data. It provides functions for creating an RccSet (an extension of the standard ExpressionSet class) from the files produced by an ncounter mrna Gene Expression assay, performing NanoString -recommended background correction and normalization, and generating a QC report with metrics that can be used to identify outlier samples and poorly performing probes (i.e. probes with signals within noise range). The overall workflow is outlined in Figure 1 (see next page). 2 Quick start The package s newrccset() function is the main wrapper function for reading in ncounter mrna Gene Expression count data and annotations: 1
2 NacoStringQCPro 2 > library(nanostringqcpro) > exampledatadir <- system.file("extdata", package="nanostringqcpro") > rccdir <- file.path(exampledatadir, "RCC") > example_rccset <- newrccset( + rccfiles = dir(rccdir, full.names=true) + #,rcccollectortoolexport = file.path(exampledatadir, "nsolver", "RCC_collector_tool_export.csv") +,rlf = file.path(exampledatadir, "RLF", "NQCP_example.rlf") +,cdrdesigndata = file.path(exampledatadir, "CDR", "CDR-DesignData.csv") +,extrapdata = file.path(exampledatadir, "extrapdata", "SampleType.txt") +,blanklabel = "blank" +,experimentdata.name = "Dorothee Nickles" +,experimentdata.lab = "Richard Bourgon" +,experimentdata.contact = "nickles.dorothee@gene.com" +,experimentdata.title = "NanoStringQCPro example dataset" +,experimentdata.abstract= "Example data for the NanoStringQCPro package" + ) > > # Reading RCC files... > # checkrccset() messages: > # The following panel housekeeping genes were found: RBCK1, USP19 Raw count data is stored in a series of Reporter Code Count (.RCC) files where each file holds the data for one sample. The rccfiles argument should be assigned a vector of paths to these files. Alternatively, the.csv file produced using the RCC Collector Tool Format Export feature of NanoString nsolver Analysis Software can be imported instead via the rcccollectortoolexport argument (commented out in the example above), but we recommend importing the.rcc files directly if they are available. Details for each probe in the codeset used RCC file RCC file RCC file...or... nsolver RCC Collector Tool Export newrccset() Annotated RccSet preprocrccset() positive control normalization background correction content normalization presence/absence call CDR file optional RLF file optional Processed RccSet Misc info from experimentalist makeqcreport() Extra sample annotation QC report optional Figure 1: Overview of the NanoStringQCPro work flow.
3 NacoStringQCPro 3 in the experiment are specified in a Reporter Library File (.RLF): a path to this file must always be specified via the rlf argument. Optional additional details for each probe are usually provided by NanoString in the Design Data tab of the Codeset Design Report (CDR) an Excel spreadsheet that accompanies each codeset order. To import these details, an extract of the Design Data tab should be saved as a.csv file (see extdata/cdr for an example) and then its path should be specified in the cdrdesigndata argument. The function has a few optional arguments that are described in more detail in its man page and in Sample annotation and Codeset annotation in the following section. For examples of the various input files, see the extdata directory included with the package and the Example files section at the end of this vignette. 3 RccSet loaded with NanoString data The object returned by newrccset() is an RccSet. This class is a light extension of the standard ExpressionSet class, and it enables NanoStringQCPro to more rigorously make use of the base ExpressionSet data structure in a manner suitable for the preprocessing and quality control of NanoString data. > str(max.level=2, example_rccset) Formal class 'RccSet' [package "NanoStringQCPro"] with 7 slots..@ experimentdata :Formal class 'MIAME' [package "Biobase"] with 13 slots..@ assaydata :<environment: 0xb6679e0>..@ phenodata :Formal class 'AnnotatedDataFrame' [package "Biobase"] with 4 slots..@ featuredata :Formal class 'AnnotatedDataFrame' [package "Biobase"] with 4 slots..@ annotation : chr "NQCP_example"..@ protocoldata :Formal class 'AnnotatedDataFrame' [package "Biobase"] with 4 slots..@. classversion :Formal class 'Versions' [package "Biobase"] with 1 slot The experimentdata slot contains information about the experiment: its title and an abstract, and the name, lab, and contact information of the people who generated the data. assaydata is a pointer to an environment that contains a matrix called exprs which initially contains the raw expression data imported from the.rcc files. (Note: a pseudo-count of 1 is always added to all measurements to enable subsequent log transformation of the data in cases where zero-counts are present.) After preprocessing, this matrix will contain background-corrected and normalized data and there will be additional matrices in assaydata for each preprocessing step see sections below. phenodata is an AnnotatedDataFrame with information about the samples, and featuredata is a similar AnnotatedDataFrame with information about the probes. The elements of phenodata correspond to the columns of the exprs matrix while those of featuredata correspond to its rows. The annotation slot shows the name of the codeset. The protocoldata slot is currently unused. Note that Biobase provides accessor functions for most key content: experimentdata(), assaydata(), exprs(), phenodata() (or pdata() for the same content in a standard data frame), featuredata() (or fdata()), and annotation(). For more information about any of these functions or the ExpressionSet class overall, see the Biobase documentation. For more information on the additional methods provided by the RccSet class, see the package s man pages. 3.1 Sample annotation Any descriptions or details about the samples that are already present in the.rcc files will be extracted by newrccset() and stored in the phenodata slot. Such details are often limited and insufficient for downstream work, so additional sample annotation can be added via.csv files specified in a vector of paths passed to the function s extrapdata argument. Each of these annotation files should be comma-separated and should
4 NacoStringQCPro 4 contain a column labeled FileName whose values are the exact.rcc file names. The FileName column is used as the primary key for merging the annotation columns into phenodata, and the function will return an error if it contains any missing or additional.rcc filenames. One particularly important bit of sample information that can be added via one of these annotation files is the SampleType column used primarily by the preprocessing and QC report functions to identify blank samples (i.e. water runs). If this column isn t specified as such, it will be assigned with default value NA for all samples. (Note also that the blanklabel argument to newrccset() should be the same as the value used in the annotation file.) 3.2 Codeset annotation Three feature annotation columns (CodeClass, GeneName, Accession) are extracted from the.rcc files and stored in the featuredata slot. Their concatenation ( <CodeClass>_<GeneName>_<Accession> ) is used for the featuredata row names and thus serves as a primary key for features; probe details in the.rlf file are merged into featuredata using this key. An additional column named SpikeInInput is added to record the RNA spike in input concentrations for the control probes in each codeset. These probes are identified by Positive and Negative values in the CodeClass column, and their concentrations are parsed from the parenthesized label in the GeneName (128, 32, 8, 2, 0.5, and fm for positive control probes and 0 for all negative control probes in the mrna Gene Expression assay). If addegannotions == TRUE, then featuredata is further annotated with EntrezGene identifiers and HGNC symbols by doing look ups in the org.hs.eg.db package using the RefSeq accessions indicated in the.rlf. Note that NA values will be assigned as the annotations for any accessions not found in org.hs.eg.db, and the values for other accessions will depend on the specific version of org.hs.eg.db that you have installed. The CDR for a given codeset is provided as an Excel file with multiple tabs containing information about the probe design history and the overall order. For the annotation of a codeset, only the Design Data tab is relevant: it has information such as probe identifiers and target sequences for all probes in the codeset. If the CDR is available, the Design Data tab can be saved as a comma delimited.csv file, and the cdrdesigndata argument of newrccset() can be pointed at this file. The function will then add the relevant columns from the file to featuredata. See the buildcodesetannotation() man page for more details. 4 Preprocessing NanoString proposes several data preprocessing steps for NanoString gene expression data: ˆ Positive control normalization. Positive control normalization can be used to adjust for all platformassociated sources of variation (e.g., automated purification, hybridization conditions, etc.). This type of normalization will not, however, account for differences in sample input which is also an important source of systematic variation. ˆ Background correction. Background signal estimates ideally, probe-specific estimates are the basis for determining whether or not a specific transcript was detected in a given sample. In addition, background subtraction improves accuracy of fold change estimation across samples, especially for targets whose signal is above but still close to the background level. Global background signal estimates can be computed using negative controls. Probe-specific background signal estimates require running a small number of blanks in addition to the primary samples of interest (three or more blanks is recommended). ˆ RNA content normalization. To account for inevitable differences in total RNA input per sample, an additional step beyond positive control normalization is required. One such approach uses scaling to
5 NacoStringQCPro 5 make the average signal for so-called housekeeping genes the same across all samples. Other non-linear approaches exist as well, but those in common use for microarray or whole-transcriptome RNA-seq data typically assume that most targets show no differential expression between conditions. For NanoString panels that include target sets that are highly focused on particular pathways or processes, this is often not a reasonable assumption, and as a consequence such approaches may not be appropriate. For most data sets, we typically use the following approach: (i) Positive control normalization, to first reduce technical variability. (ii) Subtraction of background estimates. Our preferred method (described in more detail below) is identical to that of NanoString, and it makes use of both blank samples and negative control probes. The method smoothly combines blanks and negative controls relying wholly on the latter if no blanks are available, but largely ignoring negative controls as the number of blanks increases. We have observed strong and systematic differences in probe-specific background, and we therefore recommend including enough blanks to make the negative control impact on background estimation minor. (iii) RNA content normalization via housekeeping or global median scaling. Note that if background subtraction is omitted, there is no need to do both positive control and RNA content normalization: the latter will override the former. The preprocrccset() function can be used to perform all recommended preprocessing steps (or any combination of them) in one go. It takes an RccSet and the preprocessing parameters as input, and it returns a new RccSet with exprs containing the final result of the preprocessing. Results for intermediate preprocessing steps are included in additional matrices in assaydata, and the settings for each preprocessing step are recorded in correspondingly named elements in the preprocessing slot of experimentdata (accessible via the preproc() function provided by Biobase). Note: Most downstream analysis is best performed on a logarithmic scale, so in the output s assaydata, the normdata matrix is on the log2 scale; all other matrices in assaydata are on the natural (original) scale. > norm_example_rccset <- preprocrccset(rccset = example_rccset, normmethod = "housekeeping") > # Warning message: > # In.local(rccSet,...) : Less than three housekeeping features are defined > > ls(assaydata(norm_example_rccset)) [1] "bgcorrdata" "bgestimates" "exprs" "normdata" "padata" [6] "posctrldata" > preproc(norm_example_rccset) $org.hs.eg.db_version $posctrldata_summaryfunction [1] "sum" $bgestimates_bgreference [1] "both" $bgestimates_summaryfunction [1] "median" $bgestimates_stringency [1] 1 $bgestimates_nsolverbackground.w1 [1] 2.18
6 NacoStringQCPro 6 $bgestimates_nsolverbackground.shrink [1] TRUE $bgestimates_inputmatrix [1] "posctrldata" $padata_stringency [1] 2 $padata_inputmatrix [1] "posctrldata" $normdata_method [1] "housekeeping" $normdata_summaryfunction [1] "median" $normdata_hkgenes [1] "RBCK1" "USP19" $normdata_hkfeatures [1] "Housekeeping_RBCK1_NM_006462" "Housekeeping_USP19_XP_ " $normdata_inputmatrix [1] "bgcorrdata" The following subsections further describe each individual preprocessing step. 4.1 Positive control normalization The positive spike-in RNA hybridization controls for each lane may be used to estimate the overall efficiency of hybridization and recovery for each lane. To do so, a lane-specific summary (e.g., sum or, equivalently, average) of positive control counts is first calculated. The average of these per-lane values is then used as the target, and all counts for each lane are then scaled so that all lane-specific positive control summaries match the target. The posctrlnorm() function performs positive control normalization and stores the results in a matrix called posctrldata in the assaydata slot. (Note: this differs from the behavior of other packages dealing with ExpressionSet objects that modify the values in exprs. The implementation was chosen in order to handle certain cases where the QC report needs to perform its own preprocessing on a given RccSet.) You can choose mean, median, or sum (the default) as the summary of positive control counts. NanoString advises caution with interpreting results of samples with positive control scaling factors outside the range of 0.3 to 3 since values outside this range may indicate significant under-performance of the respective lanes. A plot highlighting any outliers is provided in the QC report, and posctrlnorm() records the positive control scaling factors in a column named PosFactor in the output s phenodata for additional inspection. > adj_example_rccset <- posctrlnorm(example_rccset, summaryfunction="sum") > ls(assaydata(adj_example_rccset)) [1] "exprs" "posctrldata" > preproc(adj_example_rccset)
7 NacoStringQCPro 7 $org.hs.eg.db_version $posctrldata_summaryfunction [1] "sum" > head(pdata(adj_example_rccset)$posfactor) [1] Background correction As mentioned above, NanoString includes several probes in each codeset for which no target is expected to be present. These negative controls can be used to produce global estimates for each lane. Importantly, such global estimates will not capture probe-specific differences in background which we have seen to often be quite large. The NanoStringQCPro package offers three ways to estimate the non-specific noise in a NanoString gene expression experiment, two of which are probe specific: ˆ Lane-sepecific background, based on signals obtained for negative controls in each lane. ˆ Probe-specific background, based on signals obtained for each probe in blank measurements without RNA present. ˆ Combined probe- and lane-specific background. A few examples are shown below using the getbackground() function. The algorithm that mimics the calculation used in NanoString nsolver Analysis Software takes effect when the option to do combined probe- and lane-specific background is selected, and a full description of the algorithm and its parameters can be found in the man page for the nsolverbackground() function in this package. Note: for datasets that do not contain blanks, use bgreference="negatives" or preprorccset() will throw an error. > # Get background based on median signal for each probe in blank measurements: > bg1 <- getbackground(adj_example_rccset, bgreference="blanks", summaryfunction="median") > #...based on mean of negative controls: > bg2 <- getbackground(adj_example_rccset, bgreference="negatives", summaryfunction="mean") > #...using an implementation mimicking that of the nsolver software: > bg3 <- getbackground(adj_example_rccset, bgreference="both", stringency=1) Background signal can then be subtracted from the data by calling subtractbackground() on the background estimates matrix as obtained above. > bgcorr_example_rccset <- subtractbackground(adj_example_rccset, bgestimates=bg1) > ls(assaydata(bgcorr_example_rccset)) [1] "bgcorrdata" "bgestimates" "exprs" "posctrldata" > preproc(bgcorr_example_rccset) $org.hs.eg.db_version $posctrldata_summaryfunction [1] "sum" $bgestimates_bgreference
8 NacoStringQCPro 8 $bgestimates_summaryfunction $bgestimates_stringency $bgestimates_nsolverbackground.w1 $bgestimates_nsolverbackground.shrink $bgestimates_inputmatrix 4.3 RNA content normalization NanoStringQCPro offers two ways to normalize the NanoString count data to account for differences in input RNA content: ˆ Median normalization. Median expression values across all probes are equated (typically after positive control normalization and background correction have already been applied). ˆ Housekeeping normalization. Expression values for each probe are scaled relative the average signal from a small number of pre-specified genes housekeeping genes. Median normalization is only recommended if a large number of genes are assayed in the experiment (NanoString recommends >300 probes) and most of them are not expected to change dramatically across conditions. Note that in this setting, other non-linear normalization techniques traditionally applied to microarray or RNA-seq data e.g., variance stabilizing transforms or quantile normalization may also be appropriate. (These are not currently implemented in NanoStringQCPro, but relevant code in other Bioconductor packages can be applied directly to the matrices in assaydata.) Housekeeping normalization is recommended if a reasonable number of low-variance features can be identified for the experiment. As in PCR experiments, the housekeeping genes should be carefully evaluated as part of the codeset design process. NanoString recommends using at least 3 housekeeping features, and ideally even more. If housekeeping normalization is selected, the normalization function will by default use the panelspecified housekeeping features (i.e., those whose CodeClass is recorded as Housekeeping in the.rcc and.rlf files), but these can be overridden by passing in a different list of feature identifiers or gene symbols. The RccSet output by the normalization function will have a featuredata column named Housekeeping that indicates which features were used for the normalization. Note: Final normalized data are on the log2 scale, unlike all other matrices in assaydata. > gmnorm_example_rccset <- contentnorm(rccset=bgcorr_example_rccset, method="median") > hknorm_example_rccset <- contentnorm(rccset=bgcorr_example_rccset, method="housekeeping") > > # Warning message: > # In contentnorm(srccset, method = "housekeeping", hk = hk) : > # Less than three housekeeping features are defined
9 NacoStringQCPro 9 5 QC report and quality control of NanoString expression data For quality control of NanoString expression data, NanoStringQCPro provides a single function, makeqcreport(), that generates a QC report in HTML format: > qc_example_rccset <- makeqcreport(norm_example_rccset, "example_qc_report") > > # Generating QC report... > # Report file has been generated: /gne/home/richarbo/tmp/example_qc_report.html The report includes detailed descriptions of the different quality control steps it performs and metrics it uses. Please see the example report generated above for full detail (a copy of the report can be found in extdata/example_output). In addition to generating the report, makeqcreport() also generates additional QC files: ˆ SampleFlags.txt: a tab-delimited file listing all samples along with their sample identifiers and sample types. Three TRUE/FALSE columns specify QC flags for each sample: one flag based on technical assay run information (TechnicalFlags), one based on QC steps using signals obtained for positive and negative controls (ControlFlags), and one based on QC steps using signals obtained for endogenous genes (CountFlags). ˆ HousekeepingGeneStats.txt: a tab-delimited file providing some metrics on the housekeeping genes. For each gene, values are provided for the variance, inter-quartile range (IQR), median expression, and correlation (cor) with all other housekeeping genes across all samples. (Note that this file will only be generated if housekeeping genes are indicated in the input via CodeClass Housekeeping or if housekeeping normalization has been evinced by the featuredata Housekeeping column.) 6 Example files The package comes with the following example files corresponding to the example discussed throughout the vignette: ˆ extdata/rcc contains.rcc files as they would be produced by the ncounter instrument. ˆ extdata/nsolver/rcc_collector_tool_export.csv contains a.csv file as would be generated by the RCC Collector Tool Format Export feature of the NanoString nsolver Analysis Software. (This file contains the same data as the.rcc files.) ˆ extdata/rlf has the.rlf annotation corresponding to the codeset used in the example. ˆ extdata/cdr has the CDR Design Data annotation corresponding to the codeset used in the example. ˆ extdata/extrapdata has additional sample annotation passed to the extrapdata argument of newrccset(). ˆ examples contains.r scripts illustrating the overall workflow. 7 Session info > sessioninfo() R version ( ) Platform: x86_64-pc-linux-gnu (64-bit) Running under: Ubuntu LTS
10 NacoStringQCPro 10 Matrix products: default BLAS: /home/biocbuild/bbs-3.5-bioc/r/lib/librblas.so LAPACK: /home/biocbuild/bbs-3.5-bioc/r/lib/librlapack.so locale: [1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C [3] LC_TIME=en_US.UTF-8 LC_COLLATE=C [5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8 [7] LC_PAPER=en_US.UTF-8 LC_NAME=C [9] LC_ADDRESS=C LC_TELEPHONE=C [11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C attached base packages: [1] parallel stats graphics grdevices utils datasets methods [8] base other attached packages: [1] NanoStringQCPro_1.8.0 bigmemory_ bigmemory.sri_0.1.3 [4] Biobase_ BiocGenerics_ loaded via a namespace (and not attached): [1] Rcpp_ highr_0.6 plyr_1.8.4 [4] compiler_3.4.0 RColorBrewer_1.1-2 iterators_1.0.8 [7] tools_3.4.0 rngtools_1.2.4 digest_ [10] tibble_1.3.0 RSQLite_1.1-2 evaluate_0.10 [13] memoise_1.1.0 gtable_0.2.0 gridbase_0.4-7 [16] NMF_ png_0.1-7 foreach_1.4.3 [19] DBI_0.6-1 registry_0.3 yaml_ [22] stringr_1.2.0 pkgmaker_0.22 knitr_ [25] cluster_2.0.6 S4Vectors_ IRanges_ [28] stats4_3.4.0 rprojroot_1.2 grid_3.4.0 [31] AnnotationDbi_ rmarkdown_1.4 reshape2_1.4.2 [34] ggplot2_2.2.1 org.hs.eg.db_3.4.1 magrittr_1.5 [37] backports_1.0.5 scales_0.4.1 codetools_ [40] htmltools_0.3.5 BiocStyle_2.4.0 xtable_1.8-2 [43] colorspace_1.3-2 stringi_1.1.5 lazyeval_0.2.0 [46] munsell_0.4.3 doparallel_ markdown_0.8
Package NanoStringQCPro
 Package NanoStringQCPro October 12, 2016 Title Quality metrics and data processing methods for NanoString mrna gene expression data Version 1.4.0 Date 2016-03-03 Author , Thomas
Package NanoStringQCPro October 12, 2016 Title Quality metrics and data processing methods for NanoString mrna gene expression data Version 1.4.0 Date 2016-03-03 Author , Thomas
Package NanoStringQCPro
 Package NanoStringQCPro March 7, 2019 Title Quality metrics and data processing methods for NanoString mrna gene expression data Version 1.14.0 Date 2018-04-10 Author , Thomas
Package NanoStringQCPro March 7, 2019 Title Quality metrics and data processing methods for NanoString mrna gene expression data Version 1.14.0 Date 2018-04-10 Author , Thomas
Image Analysis with beadarray
 Mike Smith October 30, 2017 Introduction From version 2.0 beadarray provides more flexibility in the processing of array images and the extraction of bead intensities than its predecessor. In the past
Mike Smith October 30, 2017 Introduction From version 2.0 beadarray provides more flexibility in the processing of array images and the extraction of bead intensities than its predecessor. In the past
Introduction to the Codelink package
 Introduction to the Codelink package Diego Diez October 30, 2018 1 Introduction This package implements methods to facilitate the preprocessing and analysis of Codelink microarrays. Codelink is a microarray
Introduction to the Codelink package Diego Diez October 30, 2018 1 Introduction This package implements methods to facilitate the preprocessing and analysis of Codelink microarrays. Codelink is a microarray
Visualisation, transformations and arithmetic operations for grouped genomic intervals
 ## Warning: replacing previous import ggplot2::position by BiocGenerics::Position when loading soggi Visualisation, transformations and arithmetic operations for grouped genomic intervals Thomas Carroll
## Warning: replacing previous import ggplot2::position by BiocGenerics::Position when loading soggi Visualisation, transformations and arithmetic operations for grouped genomic intervals Thomas Carroll
An Introduction to the methylumi package
 An Introduction to the methylumi package Sean Davis and Sven Bilke October 13, 2014 1 Introduction Gene expression patterns are very important in understanding any biologic system. The regulation of gene
An Introduction to the methylumi package Sean Davis and Sven Bilke October 13, 2014 1 Introduction Gene expression patterns are very important in understanding any biologic system. The regulation of gene
The analysis of rtpcr data
 The analysis of rtpcr data Jitao David Zhang, Markus Ruschhaupt October 30, 2017 With the help of this document, an analysis of rtpcr data can be performed. For this, the user has to specify several parameters
The analysis of rtpcr data Jitao David Zhang, Markus Ruschhaupt October 30, 2017 With the help of this document, an analysis of rtpcr data can be performed. For this, the user has to specify several parameters
Introduction: microarray quality assessment with arrayqualitymetrics
 : microarray quality assessment with arrayqualitymetrics Audrey Kauffmann, Wolfgang Huber October 30, 2017 Contents 1 Basic use............................... 3 1.1 Affymetrix data - before preprocessing.............
: microarray quality assessment with arrayqualitymetrics Audrey Kauffmann, Wolfgang Huber October 30, 2017 Contents 1 Basic use............................... 3 1.1 Affymetrix data - before preprocessing.............
An Introduction to the methylumi package
 An Introduction to the methylumi package Sean Davis and Sven Bilke April 30, 2018 1 Introduction Gene expression patterns are very important in understanding any biologic system. The regulation of gene
An Introduction to the methylumi package Sean Davis and Sven Bilke April 30, 2018 1 Introduction Gene expression patterns are very important in understanding any biologic system. The regulation of gene
riboseqr Introduction Getting Data Workflow Example Thomas J. Hardcastle, Betty Y.W. Chung October 30, 2017
 riboseqr Thomas J. Hardcastle, Betty Y.W. Chung October 30, 2017 Introduction Ribosome profiling extracts those parts of a coding sequence currently bound by a ribosome (and thus, are likely to be undergoing
riboseqr Thomas J. Hardcastle, Betty Y.W. Chung October 30, 2017 Introduction Ribosome profiling extracts those parts of a coding sequence currently bound by a ribosome (and thus, are likely to be undergoing
Creating a New Annotation Package using SQLForge
 Creating a New Annotation Package using SQLForge Marc Carlson, HervÃľ PagÃĺs, Nianhua Li April 30, 2018 1 Introduction The AnnotationForge package provides a series of functions that can be used to build
Creating a New Annotation Package using SQLForge Marc Carlson, HervÃľ PagÃĺs, Nianhua Li April 30, 2018 1 Introduction The AnnotationForge package provides a series of functions that can be used to build
Cardinal design and development
 Kylie A. Bemis October 30, 2017 Contents 1 Introduction.............................. 2 2 Design overview........................... 2 3 iset: high-throughput imaging experiments............ 3 3.1 SImageSet:
Kylie A. Bemis October 30, 2017 Contents 1 Introduction.............................. 2 2 Design overview........................... 2 3 iset: high-throughput imaging experiments............ 3 3.1 SImageSet:
INSPEcT - INference of Synthesis, Processing and degradation rates in Time-course analysis
 INSPEcT - INference of Synthesis, Processing and degradation rates in Time-course analysis Stefano de Pretis October 30, 2017 Contents 1 Introduction.............................. 1 2 Quantification of
INSPEcT - INference of Synthesis, Processing and degradation rates in Time-course analysis Stefano de Pretis October 30, 2017 Contents 1 Introduction.............................. 1 2 Quantification of
MIGSA: Getting pbcmc datasets
 MIGSA: Getting pbcmc datasets Juan C Rodriguez Universidad Católica de Córdoba Universidad Nacional de Córdoba Cristóbal Fresno Instituto Nacional de Medicina Genómica Andrea S Llera Fundación Instituto
MIGSA: Getting pbcmc datasets Juan C Rodriguez Universidad Católica de Córdoba Universidad Nacional de Córdoba Cristóbal Fresno Instituto Nacional de Medicina Genómica Andrea S Llera Fundación Instituto
segmentseq: methods for detecting methylation loci and differential methylation
 segmentseq: methods for detecting methylation loci and differential methylation Thomas J. Hardcastle October 30, 2018 1 Introduction This vignette introduces analysis methods for data from high-throughput
segmentseq: methods for detecting methylation loci and differential methylation Thomas J. Hardcastle October 30, 2018 1 Introduction This vignette introduces analysis methods for data from high-throughput
Advanced analysis using bayseq; generic distribution definitions
 Advanced analysis using bayseq; generic distribution definitions Thomas J Hardcastle October 30, 2017 1 Generic Prior Distributions bayseq now offers complete user-specification of underlying distributions
Advanced analysis using bayseq; generic distribution definitions Thomas J Hardcastle October 30, 2017 1 Generic Prior Distributions bayseq now offers complete user-specification of underlying distributions
RMassBank for XCMS. Erik Müller. January 4, Introduction 2. 2 Input files LC/MS data Additional Workflow-Methods 2
 RMassBank for XCMS Erik Müller January 4, 2019 Contents 1 Introduction 2 2 Input files 2 2.1 LC/MS data........................... 2 3 Additional Workflow-Methods 2 3.1 Options..............................
RMassBank for XCMS Erik Müller January 4, 2019 Contents 1 Introduction 2 2 Input files 2 2.1 LC/MS data........................... 2 3 Additional Workflow-Methods 2 3.1 Options..............................
MMDiff2: Statistical Testing for ChIP-Seq Data Sets
 MMDiff2: Statistical Testing for ChIP-Seq Data Sets Gabriele Schwikert 1 and David Kuo 2 [1em] 1 The Informatics Forum, University of Edinburgh and The Wellcome
MMDiff2: Statistical Testing for ChIP-Seq Data Sets Gabriele Schwikert 1 and David Kuo 2 [1em] 1 The Informatics Forum, University of Edinburgh and The Wellcome
An Overview of the S4Vectors package
 Patrick Aboyoun, Michael Lawrence, Hervé Pagès Edited: February 2018; Compiled: June 7, 2018 Contents 1 Introduction.............................. 1 2 Vector-like and list-like objects...................
Patrick Aboyoun, Michael Lawrence, Hervé Pagès Edited: February 2018; Compiled: June 7, 2018 Contents 1 Introduction.............................. 1 2 Vector-like and list-like objects...................
How to Use pkgdeptools
 How to Use pkgdeptools Seth Falcon February 10, 2018 1 Introduction The pkgdeptools package provides tools for computing and analyzing dependency relationships among R packages. With it, you can build
How to Use pkgdeptools Seth Falcon February 10, 2018 1 Introduction The pkgdeptools package provides tools for computing and analyzing dependency relationships among R packages. With it, you can build
survsnp: Power and Sample Size Calculations for SNP Association Studies with Censored Time to Event Outcomes
 survsnp: Power and Sample Size Calculations for SNP Association Studies with Censored Time to Event Outcomes Kouros Owzar Zhiguo Li Nancy Cox Sin-Ho Jung Chanhee Yi June 29, 2016 1 Introduction This vignette
survsnp: Power and Sample Size Calculations for SNP Association Studies with Censored Time to Event Outcomes Kouros Owzar Zhiguo Li Nancy Cox Sin-Ho Jung Chanhee Yi June 29, 2016 1 Introduction This vignette
Introduction: microarray quality assessment with arrayqualitymetrics
 Introduction: microarray quality assessment with arrayqualitymetrics Audrey Kauffmann, Wolfgang Huber April 4, 2013 Contents 1 Basic use 3 1.1 Affymetrix data - before preprocessing.......................
Introduction: microarray quality assessment with arrayqualitymetrics Audrey Kauffmann, Wolfgang Huber April 4, 2013 Contents 1 Basic use 3 1.1 Affymetrix data - before preprocessing.......................
SimBindProfiles: Similar Binding Profiles, identifies common and unique regions in array genome tiling array data
 SimBindProfiles: Similar Binding Profiles, identifies common and unique regions in array genome tiling array data Bettina Fischer October 30, 2018 Contents 1 Introduction 1 2 Reading data and normalisation
SimBindProfiles: Similar Binding Profiles, identifies common and unique regions in array genome tiling array data Bettina Fischer October 30, 2018 Contents 1 Introduction 1 2 Reading data and normalisation
Working with aligned nucleotides (WORK- IN-PROGRESS!)
 Working with aligned nucleotides (WORK- IN-PROGRESS!) Hervé Pagès Last modified: January 2014; Compiled: November 17, 2017 Contents 1 Introduction.............................. 1 2 Load the aligned reads
Working with aligned nucleotides (WORK- IN-PROGRESS!) Hervé Pagès Last modified: January 2014; Compiled: November 17, 2017 Contents 1 Introduction.............................. 1 2 Load the aligned reads
PROPER: PROspective Power Evaluation for RNAseq
 PROPER: PROspective Power Evaluation for RNAseq Hao Wu [1em]Department of Biostatistics and Bioinformatics Emory University Atlanta, GA 303022 [1em] hao.wu@emory.edu October 30, 2017 Contents 1 Introduction..............................
PROPER: PROspective Power Evaluation for RNAseq Hao Wu [1em]Department of Biostatistics and Bioinformatics Emory University Atlanta, GA 303022 [1em] hao.wu@emory.edu October 30, 2017 Contents 1 Introduction..............................
Analysis of two-way cell-based assays
 Analysis of two-way cell-based assays Lígia Brás, Michael Boutros and Wolfgang Huber April 16, 2015 Contents 1 Introduction 1 2 Assembling the data 2 2.1 Reading the raw intensity files..................
Analysis of two-way cell-based assays Lígia Brás, Michael Boutros and Wolfgang Huber April 16, 2015 Contents 1 Introduction 1 2 Assembling the data 2 2.1 Reading the raw intensity files..................
INSPEcT - INference of Synthesis, Processing and degradation rates in Time-course analysis
 INSPEcT - INference of Synthesis, Processing and degradation rates in Time-course analysis Stefano de Pretis April 7, 20 Contents 1 Introduction 1 2 Quantification of Exon and Intron features 3 3 Time-course
INSPEcT - INference of Synthesis, Processing and degradation rates in Time-course analysis Stefano de Pretis April 7, 20 Contents 1 Introduction 1 2 Quantification of Exon and Intron features 3 3 Time-course
Biobase development and the new eset
 Biobase development and the new eset Martin T. Morgan * 7 August, 2006 Revised 4 September, 2006 featuredata slot. Revised 20 April 2007 minor wording changes; verbose and other arguments passed through
Biobase development and the new eset Martin T. Morgan * 7 August, 2006 Revised 4 September, 2006 featuredata slot. Revised 20 April 2007 minor wording changes; verbose and other arguments passed through
HowTo: Querying online Data
 HowTo: Querying online Data Jeff Gentry and Robert Gentleman November 12, 2017 1 Overview This article demonstrates how you can make use of the tools that have been provided for on-line querying of data
HowTo: Querying online Data Jeff Gentry and Robert Gentleman November 12, 2017 1 Overview This article demonstrates how you can make use of the tools that have been provided for on-line querying of data
Bioconductor: Annotation Package Overview
 Bioconductor: Annotation Package Overview April 30, 2018 1 Overview In its current state the basic purpose of annotate is to supply interface routines that support user actions that rely on the different
Bioconductor: Annotation Package Overview April 30, 2018 1 Overview In its current state the basic purpose of annotate is to supply interface routines that support user actions that rely on the different
All About PlexSet Technology Data Analysis in nsolver Software
 All About PlexSet Technology Data Analysis in nsolver Software PlexSet is a multiplexed gene expression technology which allows pooling of up to 8 samples per ncounter cartridge lane, enabling users to
All About PlexSet Technology Data Analysis in nsolver Software PlexSet is a multiplexed gene expression technology which allows pooling of up to 8 samples per ncounter cartridge lane, enabling users to
A complete analysis of peptide microarray binding data using the pepstat framework
 A complete analysis of peptide microarray binding data using the pepstat framework Greg Imholte, Renan Sauteraud, Mike Jiang and Raphael Gottardo April 30, 2018 This document present a full analysis, from
A complete analysis of peptide microarray binding data using the pepstat framework Greg Imholte, Renan Sauteraud, Mike Jiang and Raphael Gottardo April 30, 2018 This document present a full analysis, from
An Introduction to Bioconductor s ExpressionSet Class
 An Introduction to Bioconductor s ExpressionSet Class Seth Falcon, Martin Morgan, and Robert Gentleman 6 October, 2006; revised 9 February, 2007 1 Introduction Biobase is part of the Bioconductor project,
An Introduction to Bioconductor s ExpressionSet Class Seth Falcon, Martin Morgan, and Robert Gentleman 6 October, 2006; revised 9 February, 2007 1 Introduction Biobase is part of the Bioconductor project,
Creating IGV HTML reports with tracktables
 Thomas Carroll 1 [1em] 1 Bioinformatics Core, MRC Clinical Sciences Centre; thomas.carroll (at)imperial.ac.uk June 13, 2018 Contents 1 The tracktables package.................... 1 2 Creating IGV sessions
Thomas Carroll 1 [1em] 1 Bioinformatics Core, MRC Clinical Sciences Centre; thomas.carroll (at)imperial.ac.uk June 13, 2018 Contents 1 The tracktables package.................... 1 2 Creating IGV sessions
Analysis of Bead-level Data using beadarray
 Mark Dunning October 30, 2017 Introduction beadarray is a package for the pre-processing and analysis of Illumina BeadArray. The main advantage is being able to read raw data output by Illumina s scanning
Mark Dunning October 30, 2017 Introduction beadarray is a package for the pre-processing and analysis of Illumina BeadArray. The main advantage is being able to read raw data output by Illumina s scanning
Introduction to the TSSi package: Identification of Transcription Start Sites
 Introduction to the TSSi package: Identification of Transcription Start Sites Julian Gehring, Clemens Kreutz October 30, 2017 Along with the advances in high-throughput sequencing, the detection of transcription
Introduction to the TSSi package: Identification of Transcription Start Sites Julian Gehring, Clemens Kreutz October 30, 2017 Along with the advances in high-throughput sequencing, the detection of transcription
Analyse RT PCR data with ddct
 Analyse RT PCR data with ddct Jitao David Zhang, Rudolf Biczok and Markus Ruschhaupt October 30, 2017 Abstract Quantitative real time PCR (qrt PCR or RT PCR for short) is a laboratory technique based on
Analyse RT PCR data with ddct Jitao David Zhang, Rudolf Biczok and Markus Ruschhaupt October 30, 2017 Abstract Quantitative real time PCR (qrt PCR or RT PCR for short) is a laboratory technique based on
SCnorm: robust normalization of single-cell RNA-seq data
 SCnorm: robust normalization of single-cell RNA-seq data Rhonda Bacher and Christina Kendziorski February 18, 2019 Contents 1 Introduction.............................. 1 2 Run SCnorm.............................
SCnorm: robust normalization of single-cell RNA-seq data Rhonda Bacher and Christina Kendziorski February 18, 2019 Contents 1 Introduction.............................. 1 2 Run SCnorm.............................
Creating a New Annotation Package using SQLForge
 Creating a New Annotation Package using SQLForge Marc Carlson, Herve Pages, Nianhua Li November 19, 2013 1 Introduction The AnnotationForge package provides a series of functions that can be used to build
Creating a New Annotation Package using SQLForge Marc Carlson, Herve Pages, Nianhua Li November 19, 2013 1 Introduction The AnnotationForge package provides a series of functions that can be used to build
Triform: peak finding in ChIP-Seq enrichment profiles for transcription factors
 Triform: peak finding in ChIP-Seq enrichment profiles for transcription factors Karl Kornacker * and Tony Håndstad October 30, 2018 A guide for using the Triform algorithm to predict transcription factor
Triform: peak finding in ChIP-Seq enrichment profiles for transcription factors Karl Kornacker * and Tony Håndstad October 30, 2018 A guide for using the Triform algorithm to predict transcription factor
How To Use GOstats and Category to do Hypergeometric testing with unsupported model organisms
 How To Use GOstats and Category to do Hypergeometric testing with unsupported model organisms M. Carlson October 30, 2017 1 Introduction This vignette is meant as an extension of what already exists in
How To Use GOstats and Category to do Hypergeometric testing with unsupported model organisms M. Carlson October 30, 2017 1 Introduction This vignette is meant as an extension of what already exists in
How to use CNTools. Overview. Algorithms. Jianhua Zhang. April 14, 2011
 How to use CNTools Jianhua Zhang April 14, 2011 Overview Studies have shown that genomic alterations measured as DNA copy number variations invariably occur across chromosomal regions that span over several
How to use CNTools Jianhua Zhang April 14, 2011 Overview Studies have shown that genomic alterations measured as DNA copy number variations invariably occur across chromosomal regions that span over several
Analysis of screens with enhancer and suppressor controls
 Analysis of screens with enhancer and suppressor controls Lígia Brás, Michael Boutros and Wolfgang Huber April 16, 2015 Contents 1 Introduction 1 2 Assembling the data 2 2.1 Reading the raw intensity files..................
Analysis of screens with enhancer and suppressor controls Lígia Brás, Michael Boutros and Wolfgang Huber April 16, 2015 Contents 1 Introduction 1 2 Assembling the data 2 2.1 Reading the raw intensity files..................
The bumphunter user s guide
 The bumphunter user s guide Kasper Daniel Hansen khansen@jhsph.edu Martin Aryee aryee.martin@mgh.harvard.edu Rafael A. Irizarry rafa@jhu.edu Modified: November 23, 2012. Compiled: April 30, 2018 Introduction
The bumphunter user s guide Kasper Daniel Hansen khansen@jhsph.edu Martin Aryee aryee.martin@mgh.harvard.edu Rafael A. Irizarry rafa@jhu.edu Modified: November 23, 2012. Compiled: April 30, 2018 Introduction
CQN (Conditional Quantile Normalization)
 CQN (Conditional Quantile Normalization) Kasper Daniel Hansen khansen@jhsph.edu Zhijin Wu zhijin_wu@brown.edu Modified: August 8, 2012. Compiled: April 30, 2018 Introduction This package contains the CQN
CQN (Conditional Quantile Normalization) Kasper Daniel Hansen khansen@jhsph.edu Zhijin Wu zhijin_wu@brown.edu Modified: August 8, 2012. Compiled: April 30, 2018 Introduction This package contains the CQN
An Introduction to ShortRead
 Martin Morgan Modified: 21 October, 2013. Compiled: April 30, 2018 > library("shortread") The ShortRead package provides functionality for working with FASTQ files from high throughput sequence analysis.
Martin Morgan Modified: 21 October, 2013. Compiled: April 30, 2018 > library("shortread") The ShortRead package provides functionality for working with FASTQ files from high throughput sequence analysis.
How to use cghmcr. October 30, 2017
 How to use cghmcr Jianhua Zhang Bin Feng October 30, 2017 1 Overview Copy number data (arraycgh or SNP) can be used to identify genomic regions (Regions Of Interest or ROI) showing gains or losses that
How to use cghmcr Jianhua Zhang Bin Feng October 30, 2017 1 Overview Copy number data (arraycgh or SNP) can be used to identify genomic regions (Regions Of Interest or ROI) showing gains or losses that
segmentseq: methods for detecting methylation loci and differential methylation
 segmentseq: methods for detecting methylation loci and differential methylation Thomas J. Hardcastle October 13, 2015 1 Introduction This vignette introduces analysis methods for data from high-throughput
segmentseq: methods for detecting methylation loci and differential methylation Thomas J. Hardcastle October 13, 2015 1 Introduction This vignette introduces analysis methods for data from high-throughput
Count outlier detection using Cook s distance
 Count outlier detection using Cook s distance Michael Love August 9, 2014 1 Run DE analysis with and without outlier removal The following vignette produces the Supplemental Figure of the effect of replacing
Count outlier detection using Cook s distance Michael Love August 9, 2014 1 Run DE analysis with and without outlier removal The following vignette produces the Supplemental Figure of the effect of replacing
Introduction to BatchtoolsParam
 Nitesh Turaga 1, Martin Morgan 2 Edited: March 22, 2018; Compiled: January 4, 2019 1 Nitesh.Turaga@ RoswellPark.org 2 Martin.Morgan@ RoswellPark.org Contents 1 Introduction..............................
Nitesh Turaga 1, Martin Morgan 2 Edited: March 22, 2018; Compiled: January 4, 2019 1 Nitesh.Turaga@ RoswellPark.org 2 Martin.Morgan@ RoswellPark.org Contents 1 Introduction..............................
mocluster: Integrative clustering using multiple
 mocluster: Integrative clustering using multiple omics data Chen Meng Modified: June 03, 2015. Compiled: October 30, 2018. Contents 1 mocluster overview......................... 1 2 Run mocluster............................
mocluster: Integrative clustering using multiple omics data Chen Meng Modified: June 03, 2015. Compiled: October 30, 2018. Contents 1 mocluster overview......................... 1 2 Run mocluster............................
rhdf5 - HDF5 interface for R
 Bernd Fischer October 30, 2017 Contents 1 Introduction 1 2 Installation of the HDF5 package 2 3 High level R -HDF5 functions 2 31 Creating an HDF5 file and group hierarchy 2 32 Writing and reading objects
Bernd Fischer October 30, 2017 Contents 1 Introduction 1 2 Installation of the HDF5 package 2 3 High level R -HDF5 functions 2 31 Creating an HDF5 file and group hierarchy 2 32 Writing and reading objects
bayseq: Empirical Bayesian analysis of patterns of differential expression in count data
 bayseq: Empirical Bayesian analysis of patterns of differential expression in count data Thomas J. Hardcastle October 30, 2017 1 Introduction This vignette is intended to give a rapid introduction to the
bayseq: Empirical Bayesian analysis of patterns of differential expression in count data Thomas J. Hardcastle October 30, 2017 1 Introduction This vignette is intended to give a rapid introduction to the
SCnorm: robust normalization of single-cell RNA-seq data
 SCnorm: robust normalization of single-cell RNA-seq data Rhonda Bacher and Christina Kendziorski September 24, 207 Contents Introduction.............................. 2 Run SCnorm.............................
SCnorm: robust normalization of single-cell RNA-seq data Rhonda Bacher and Christina Kendziorski September 24, 207 Contents Introduction.............................. 2 Run SCnorm.............................
RNAseq Differential Expression Analysis Jana Schor and Jörg Hackermüller November, 2017
 RNAseq Differential Expression Analysis Jana Schor and Jörg Hackermüller November, 2017 RNA-Seq: Example dataset and download} Dataset taken from Somel et al. 2012: (check publication for details) 3 human
RNAseq Differential Expression Analysis Jana Schor and Jörg Hackermüller November, 2017 RNA-Seq: Example dataset and download} Dataset taken from Somel et al. 2012: (check publication for details) 3 human
MultipleAlignment Objects
 MultipleAlignment Objects Marc Carlson Bioconductor Core Team Fred Hutchinson Cancer Research Center Seattle, WA April 30, 2018 Contents 1 Introduction 1 2 Creation and masking 1 3 Analytic utilities 7
MultipleAlignment Objects Marc Carlson Bioconductor Core Team Fred Hutchinson Cancer Research Center Seattle, WA April 30, 2018 Contents 1 Introduction 1 2 Creation and masking 1 3 Analytic utilities 7
DupChecker: a bioconductor package for checking high-throughput genomic data redundancy in meta-analysis
 DupChecker: a bioconductor package for checking high-throughput genomic data redundancy in meta-analysis Quanhu Sheng, Yu Shyr, Xi Chen Center for Quantitative Sciences, Vanderbilt University, Nashville,
DupChecker: a bioconductor package for checking high-throughput genomic data redundancy in meta-analysis Quanhu Sheng, Yu Shyr, Xi Chen Center for Quantitative Sciences, Vanderbilt University, Nashville,
Comparing methods for differential expression analysis of RNAseq data with the compcoder
 Comparing methods for differential expression analysis of RNAseq data with the compcoder package Charlotte Soneson October 30, 2017 Contents 1 Introduction.............................. 2 2 The compdata
Comparing methods for differential expression analysis of RNAseq data with the compcoder package Charlotte Soneson October 30, 2017 Contents 1 Introduction.............................. 2 2 The compdata
RDAVIDWebService: a versatile R interface to DAVID
 RDAVIDWebService: a versatile R interface to DAVID Cristóbal Fresno 1,2 and Elmer A. Fernández 1,2 1 Bio-science Data Mining Group, Catholic University of Córdoba, Córdoba, Argentina. 2 CONICET, Argentina
RDAVIDWebService: a versatile R interface to DAVID Cristóbal Fresno 1,2 and Elmer A. Fernández 1,2 1 Bio-science Data Mining Group, Catholic University of Córdoba, Córdoba, Argentina. 2 CONICET, Argentina
Shrinkage of logarithmic fold changes
 Shrinkage of logarithmic fold changes Michael Love August 9, 2014 1 Comparing the posterior distribution for two genes First, we run a DE analysis on the Bottomly et al. dataset, once with shrunken LFCs
Shrinkage of logarithmic fold changes Michael Love August 9, 2014 1 Comparing the posterior distribution for two genes First, we run a DE analysis on the Bottomly et al. dataset, once with shrunken LFCs
ssviz: A small RNA-seq visualizer and analysis toolkit
 ssviz: A small RNA-seq visualizer and analysis toolkit Diana HP Low Institute of Molecular and Cell Biology Agency for Science, Technology and Research (A*STAR), Singapore dlow@imcb.a-star.edu.sg August
ssviz: A small RNA-seq visualizer and analysis toolkit Diana HP Low Institute of Molecular and Cell Biology Agency for Science, Technology and Research (A*STAR), Singapore dlow@imcb.a-star.edu.sg August
Overview of ensemblvep
 Overview of ensemblvep Lori Shepherd and Valerie Obenchain October 30, 2017 Contents 1 Introduction 1 2 Results as R objects 1 3 Write results to a file 4 4 Configuring runtime options 5 5 sessioninfo()
Overview of ensemblvep Lori Shepherd and Valerie Obenchain October 30, 2017 Contents 1 Introduction 1 2 Results as R objects 1 3 Write results to a file 4 4 Configuring runtime options 5 5 sessioninfo()
Getting started with flowstats
 Getting started with flowstats F. Hahne, N. Gopalakrishnan October, 7 Abstract flowstats is a collection of algorithms for the statistical analysis of flow cytometry data. So far, the focus is on automated
Getting started with flowstats F. Hahne, N. Gopalakrishnan October, 7 Abstract flowstats is a collection of algorithms for the statistical analysis of flow cytometry data. So far, the focus is on automated
Raman Spectra of Chondrocytes in Cartilage: hyperspec s chondro data set
 Raman Spectra of Chondrocytes in Cartilage: hyperspec s chondro data set Claudia Beleites CENMAT and DI3, University of Trieste Spectroscopy Imaging, IPHT Jena e.v. February 13,
Raman Spectra of Chondrocytes in Cartilage: hyperspec s chondro data set Claudia Beleites CENMAT and DI3, University of Trieste Spectroscopy Imaging, IPHT Jena e.v. February 13,
Fitting Baselines to Spectra
 Fitting Baselines to Spectra Claudia Beleites DIA Raman Spectroscopy Group, University of Trieste/Italy (2005 2008) Spectroscopy Imaging, IPHT, Jena/Germany (2008 2016) ÖPV, JKI,
Fitting Baselines to Spectra Claudia Beleites DIA Raman Spectroscopy Group, University of Trieste/Italy (2005 2008) Spectroscopy Imaging, IPHT, Jena/Germany (2008 2016) ÖPV, JKI,
SIBER User Manual. Pan Tong and Kevin R Coombes. May 27, Introduction 1
 SIBER User Manual Pan Tong and Kevin R Coombes May 27, 2015 Contents 1 Introduction 1 2 Using SIBER 1 2.1 A Quick Example........................................... 1 2.2 Dealing With RNAseq Normalization................................
SIBER User Manual Pan Tong and Kevin R Coombes May 27, 2015 Contents 1 Introduction 1 2 Using SIBER 1 2.1 A Quick Example........................................... 1 2.2 Dealing With RNAseq Normalization................................
qplexanalyzer Matthew Eldridge, Kamal Kishore, Ashley Sawle January 4, Overview 1 2 Import quantitative dataset 2 3 Quality control 2
 qplexanalyzer Matthew Eldridge, Kamal Kishore, Ashley Sawle January 4, 2019 Contents 1 Overview 1 2 Import quantitative dataset 2 3 Quality control 2 4 Data normalization 8 5 Aggregation of peptide intensities
qplexanalyzer Matthew Eldridge, Kamal Kishore, Ashley Sawle January 4, 2019 Contents 1 Overview 1 2 Import quantitative dataset 2 3 Quality control 2 4 Data normalization 8 5 Aggregation of peptide intensities
Introduction to QDNAseq
 Ilari Scheinin October 30, 2017 Contents 1 Running QDNAseq.......................... 1 1.1 Bin annotations.......................... 1 1.2 Processing bam files....................... 2 1.3 Downstream analyses......................
Ilari Scheinin October 30, 2017 Contents 1 Running QDNAseq.......................... 1 1.1 Bin annotations.......................... 1 1.2 Processing bam files....................... 2 1.3 Downstream analyses......................
BioConductor Overviewr
 BioConductor Overviewr 2016-09-28 Contents Installing Bioconductor 1 Bioconductor basics 1 ExressionSet 2 assaydata (gene expression)........................................ 2 phenodata (sample annotations).....................................
BioConductor Overviewr 2016-09-28 Contents Installing Bioconductor 1 Bioconductor basics 1 ExressionSet 2 assaydata (gene expression)........................................ 2 phenodata (sample annotations).....................................
MLSeq package: Machine Learning Interface to RNA-Seq Data
 MLSeq package: Machine Learning Interface to RNA-Seq Data Gokmen Zararsiz 1, Dincer Goksuluk 2, Selcuk Korkmaz 2, Vahap Eldem 3, Izzet Parug Duru 4, Turgay Unver 5, Ahmet Ozturk 5 1 Erciyes University,
MLSeq package: Machine Learning Interface to RNA-Seq Data Gokmen Zararsiz 1, Dincer Goksuluk 2, Selcuk Korkmaz 2, Vahap Eldem 3, Izzet Parug Duru 4, Turgay Unver 5, Ahmet Ozturk 5 1 Erciyes University,
Using branchpointer for annotation of intronic human splicing branchpoints
 Using branchpointer for annotation of intronic human splicing branchpoints Beth Signal October 30, 2018 Contents 1 Introduction.............................. 1 2 Preparation..............................
Using branchpointer for annotation of intronic human splicing branchpoints Beth Signal October 30, 2018 Contents 1 Introduction.............................. 1 2 Preparation..............................
The rtracklayer package
 The rtracklayer package Michael Lawrence January 22, 2018 Contents 1 Introduction 2 2 Gene expression and microrna target sites 2 2.1 Creating a target site track..................... 2 2.1.1 Constructing
The rtracklayer package Michael Lawrence January 22, 2018 Contents 1 Introduction 2 2 Gene expression and microrna target sites 2 2.1 Creating a target site track..................... 2 2.1.1 Constructing
Overview of ensemblvep
 Overview of ensemblvep Lori Shepherd and Valerie Obenchain October 30, 2018 Contents 1 Introduction 1 2 Results as R objects 1 3 Write results to a file 4 4 Configuring runtime options 5 5 sessioninfo()
Overview of ensemblvep Lori Shepherd and Valerie Obenchain October 30, 2018 Contents 1 Introduction 1 2 Results as R objects 1 3 Write results to a file 4 4 Configuring runtime options 5 5 sessioninfo()
RAPIDR. Kitty Lo. November 20, Intended use of RAPIDR 1. 2 Create binned counts file from BAMs Masking... 1
 RAPIDR Kitty Lo November 20, 2014 Contents 1 Intended use of RAPIDR 1 2 Create binned counts file from BAMs 1 2.1 Masking.................................................... 1 3 Build the reference 2 3.1
RAPIDR Kitty Lo November 20, 2014 Contents 1 Intended use of RAPIDR 1 2 Create binned counts file from BAMs 1 2.1 Masking.................................................... 1 3 Build the reference 2 3.1
Analyze Illumina Infinium methylation microarray data
 Analyze Illumina Infinium methylation microarray data Pan Du, Gang Feng, Spencer Huang, Warren A. Kibbe, Simon Lin May 3, 2016 Genentech Inc., South San Francisco, CA, 94080, USA Biomedical Informatics
Analyze Illumina Infinium methylation microarray data Pan Du, Gang Feng, Spencer Huang, Warren A. Kibbe, Simon Lin May 3, 2016 Genentech Inc., South San Francisco, CA, 94080, USA Biomedical Informatics
An Introduction to GenomeInfoDb
 Martin Morgan, Hervé Pagès, Marc Carlson, Sonali Arora Modified: 17 January, 2014. Compiled: January 21, 2018 Contents 1 Introduction.............................. 2 2 Functionality for all existing organisms...............
Martin Morgan, Hervé Pagès, Marc Carlson, Sonali Arora Modified: 17 January, 2014. Compiled: January 21, 2018 Contents 1 Introduction.............................. 2 2 Functionality for all existing organisms...............
Introduction to R (BaRC Hot Topics)
 Introduction to R (BaRC Hot Topics) George Bell September 30, 2011 This document accompanies the slides from BaRC s Introduction to R and shows the use of some simple commands. See the accompanying slides
Introduction to R (BaRC Hot Topics) George Bell September 30, 2011 This document accompanies the slides from BaRC s Introduction to R and shows the use of some simple commands. See the accompanying slides
Introduction to GenomicFiles
 Valerie Obenchain, Michael Love, Martin Morgan Last modified: October 2014; Compiled: October 30, 2018 Contents 1 Introduction.............................. 1 2 Quick Start..............................
Valerie Obenchain, Michael Love, Martin Morgan Last modified: October 2014; Compiled: October 30, 2018 Contents 1 Introduction.............................. 1 2 Quick Start..............................
Evaluation of VST algorithm in lumi package
 Evaluation of VST algorithm in lumi package Pan Du 1, Simon Lin 1, Wolfgang Huber 2, Warrren A. Kibbe 1 December 22, 2010 1 Robert H. Lurie Comprehensive Cancer Center Northwestern University, Chicago,
Evaluation of VST algorithm in lumi package Pan Du 1, Simon Lin 1, Wolfgang Huber 2, Warrren A. Kibbe 1 December 22, 2010 1 Robert H. Lurie Comprehensive Cancer Center Northwestern University, Chicago,
MethylAid: Visual and Interactive quality control of Illumina Human DNA Methylation array data
 MethylAid: Visual and Interactive quality control of Illumina Human DNA Methylation array Maarten van Iterson, Elmar Tobi, Roderick Slieker, Wouter den Hollander, Rene Luijk, Eline Slagboom and Bas Heijmans
MethylAid: Visual and Interactive quality control of Illumina Human DNA Methylation array Maarten van Iterson, Elmar Tobi, Roderick Slieker, Wouter den Hollander, Rene Luijk, Eline Slagboom and Bas Heijmans
highscreen: High-Throughput Screening for Plate Based Assays
 highscreen: High-Throughput Screening for Plate Based Assays Ivo D. Shterev Cliburn Chan Gregory D. Sempowski August 22, 2018 1 Introduction This vignette describes the use of the R extension package highscreen
highscreen: High-Throughput Screening for Plate Based Assays Ivo D. Shterev Cliburn Chan Gregory D. Sempowski August 22, 2018 1 Introduction This vignette describes the use of the R extension package highscreen
GenoGAM: Genome-wide generalized additive
 GenoGAM: Genome-wide generalized additive models Georg Stricker 1, Julien Gagneur 1 [1em] 1 Technische Universität München, Department of Informatics, Garching, Germany Abstract April 30, 2018 Many genomic
GenoGAM: Genome-wide generalized additive models Georg Stricker 1, Julien Gagneur 1 [1em] 1 Technische Universität München, Department of Informatics, Garching, Germany Abstract April 30, 2018 Many genomic
Starr: Simple Tiling ARRay analysis for Affymetrix ChIP-chip data
 Starr: Simple Tiling ARRay analysis for Affymetrix ChIP-chip data Benedikt Zacher, Achim Tresch October 30, 2017 1 Introduction Starr is an extension of the Ringo [6] package for the analysis of ChIP-chip
Starr: Simple Tiling ARRay analysis for Affymetrix ChIP-chip data Benedikt Zacher, Achim Tresch October 30, 2017 1 Introduction Starr is an extension of the Ringo [6] package for the analysis of ChIP-chip
Using metama for differential gene expression analysis from multiple studies
 Using metama for differential gene expression analysis from multiple studies Guillemette Marot and Rémi Bruyère Modified: January 28, 2015. Compiled: January 28, 2015 Abstract This vignette illustrates
Using metama for differential gene expression analysis from multiple studies Guillemette Marot and Rémi Bruyère Modified: January 28, 2015. Compiled: January 28, 2015 Abstract This vignette illustrates
A wrapper to query DGIdb using R
 A wrapper to query DGIdb using R Thomas Thurnherr 1, Franziska Singer, Daniel J. Stekhoven, Niko Beerenwinkel 1 thomas.thurnherr@gmail.com Abstract February 16, 2018 Annotation and interpretation of DNA
A wrapper to query DGIdb using R Thomas Thurnherr 1, Franziska Singer, Daniel J. Stekhoven, Niko Beerenwinkel 1 thomas.thurnherr@gmail.com Abstract February 16, 2018 Annotation and interpretation of DNA
Package frma. R topics documented: March 8, Version Date Title Frozen RMA and Barcode
 Version 1.34.0 Date 2017-11-08 Title Frozen RMA and Barcode Package frma March 8, 2019 Preprocessing and analysis for single microarrays and microarray batches. Author Matthew N. McCall ,
Version 1.34.0 Date 2017-11-08 Title Frozen RMA and Barcode Package frma March 8, 2019 Preprocessing and analysis for single microarrays and microarray batches. Author Matthew N. McCall ,
M 3 D: Statistical Testing for RRBS Data Sets
 Tom Mayo Modified: 5 September, 2016. Compiled: October 30, 2018 Contents 1 Introduction 1 2 Analysis Pipeline 2 2.1 Reading in Data. 2 2.2 Computing the MMD, and coverage-mmd. 5 2.3 Generating p-values..
Tom Mayo Modified: 5 September, 2016. Compiled: October 30, 2018 Contents 1 Introduction 1 2 Analysis Pipeline 2 2.1 Reading in Data. 2 2.2 Computing the MMD, and coverage-mmd. 5 2.3 Generating p-values..
Preprocessing and Genotyping Illumina Arrays for Copy Number Analysis
 Preprocessing and Genotyping Illumina Arrays for Copy Number Analysis Rob Scharpf September 18, 2012 Abstract This vignette illustrates the steps required prior to copy number analysis for Infinium platforms.
Preprocessing and Genotyping Illumina Arrays for Copy Number Analysis Rob Scharpf September 18, 2012 Abstract This vignette illustrates the steps required prior to copy number analysis for Infinium platforms.
Making and Utilizing TxDb Objects
 Marc Carlson, Patrick Aboyoun, HervÃľ PagÃĺs, Seth Falcon, Martin Morgan October 30, 2017 1 Introduction The GenomicFeatures package retrieves and manages transcript-related features from the UCSC Genome
Marc Carlson, Patrick Aboyoun, HervÃľ PagÃĺs, Seth Falcon, Martin Morgan October 30, 2017 1 Introduction The GenomicFeatures package retrieves and manages transcript-related features from the UCSC Genome
Bioconductor L A T E X Style 2.0
 Andrzej Oleś 1, Martin Morgan 2, and Wolfgang Huber 1 1 European Molecular Biology Laboratory, Heidelberg, Germany 2 Roswell Park Cancer Institute, Buffalo, NY Abstract Package November 29, 2017 This vignette
Andrzej Oleś 1, Martin Morgan 2, and Wolfgang Huber 1 1 European Molecular Biology Laboratory, Heidelberg, Germany 2 Roswell Park Cancer Institute, Buffalo, NY Abstract Package November 29, 2017 This vignette
An introduction to gaucho
 An introduction to gaucho Alex Murison Alexander.Murison@icr.ac.uk Christopher P Wardell Christopher.Wardell@icr.ac.uk October 30, 2017 This vignette serves as an introduction to the R package gaucho.
An introduction to gaucho Alex Murison Alexander.Murison@icr.ac.uk Christopher P Wardell Christopher.Wardell@icr.ac.uk October 30, 2017 This vignette serves as an introduction to the R package gaucho.
MiChip. Jonathon Blake. October 30, Introduction 1. 5 Plotting Functions 3. 6 Normalization 3. 7 Writing Output Files 3
 MiChip Jonathon Blake October 30, 2018 Contents 1 Introduction 1 2 Reading the Hybridization Files 1 3 Removing Unwanted Rows and Correcting for Flags 2 4 Summarizing Intensities 3 5 Plotting Functions
MiChip Jonathon Blake October 30, 2018 Contents 1 Introduction 1 2 Reading the Hybridization Files 1 3 Removing Unwanted Rows and Correcting for Flags 2 4 Summarizing Intensities 3 5 Plotting Functions
The REDseq user s guide
 The REDseq user s guide Lihua Julie Zhu * October 30, 2017 Contents 1 Introduction 1 2 Examples of using REDseq 2 2.1 Task 1: Build a RE map for a genome..................... 2 2.2 Task 2: Assign mapped
The REDseq user s guide Lihua Julie Zhu * October 30, 2017 Contents 1 Introduction 1 2 Examples of using REDseq 2 2.1 Task 1: Build a RE map for a genome..................... 2 2.2 Task 2: Assign mapped
Analysis of Pairwise Interaction Screens
 Analysis of Pairwise Interaction Screens Bernd Fischer April 11, 2014 Contents 1 Introduction 1 2 Installation of the RNAinteract package 1 3 Creating an RNAinteract object 2 4 Single Perturbation effects
Analysis of Pairwise Interaction Screens Bernd Fischer April 11, 2014 Contents 1 Introduction 1 2 Installation of the RNAinteract package 1 3 Creating an RNAinteract object 2 4 Single Perturbation effects
Djork-Arné Clevert and Andreas Mitterecker. Institute of Bioinformatics, Johannes Kepler University Linz
 Software Manual Institute of Bioinformatics, Johannes Kepler University Linz cn.farms: a latent variable model to detect copy number variations in microarray data with a low false discovery rate Manual
Software Manual Institute of Bioinformatics, Johannes Kepler University Linz cn.farms: a latent variable model to detect copy number variations in microarray data with a low false discovery rate Manual
Package BEARscc. May 2, 2018
 Type Package Package BEARscc May 2, 2018 Title BEARscc (Bayesian ERCC Assesstment of Robustness of Single Cell Clusters) Version 1.0.0 Author David T. Severson Maintainer
Type Package Package BEARscc May 2, 2018 Title BEARscc (Bayesian ERCC Assesstment of Robustness of Single Cell Clusters) Version 1.0.0 Author David T. Severson Maintainer
Analysis of Bead-summary Data using beadarray
 Analysis of Bead-summary Data using beadarray Mark Dunning October 30, 2017 Contents 1 Introduction............................. 2 2 feature and pheno data..................... 4 3 Subsetting the data.......................
Analysis of Bead-summary Data using beadarray Mark Dunning October 30, 2017 Contents 1 Introduction............................. 2 2 feature and pheno data..................... 4 3 Subsetting the data.......................
crlmm to downstream data analysis
 crlmm to downstream data analysis VJ Carey, B Carvalho March, 2012 1 Running CRLMM on a nontrivial set of CEL files To use the crlmm algorithm, the user must load the crlmm package, as described below:
crlmm to downstream data analysis VJ Carey, B Carvalho March, 2012 1 Running CRLMM on a nontrivial set of CEL files To use the crlmm algorithm, the user must load the crlmm package, as described below:
Djork-Arné Clevert and Andreas Mitterecker. Institute of Bioinformatics, Johannes Kepler University Linz
 Software Manual Institute of Bioinformatics, Johannes Kepler University Linz cn.farms: a latent variable model to detect copy number variations in microarray data with a low false discovery rate Manual
Software Manual Institute of Bioinformatics, Johannes Kepler University Linz cn.farms: a latent variable model to detect copy number variations in microarray data with a low false discovery rate Manual
Bioconductor L A T E X Style 2.0
 Andrzej Oleś 1, Martin Morgan 2, and Wolfgang Huber 1 1 European Molecular Biology Laboratory, Heidelberg, Germany 2 Roswell Park Cancer Institute, Buffalo, NY Abstract Package November 23, 2016 This vignette
Andrzej Oleś 1, Martin Morgan 2, and Wolfgang Huber 1 1 European Molecular Biology Laboratory, Heidelberg, Germany 2 Roswell Park Cancer Institute, Buffalo, NY Abstract Package November 23, 2016 This vignette
