Run-Length and Markov Compression Revisited in DNA Sequences
|
|
|
- Corey Lloyd
- 5 years ago
- Views:
Transcription
1 Run-Length and Markov Compression Revisited in DNA Sequences Saurabh Kadekodi M.S. Computer Science Efficient and economical storage of DNA sequences has always been an interesting problem to solve. There is no particular solution termed as the "best" solution for this problem although people have tried to attack the problem from various angles. Today, the digital size of an uncompressed human genome can be as large as 285 GB[1]. With this explosion in DNA data, especially since the usage of next generation sequencing methods[2], its compression has become imperative. Below is a graph that summarizes the history of genome data and the evident need for DNA compression. [Fig 1] - Comparison of disk costs in MB per US dollar to DNA costs in base pair per US dollar. The blue squares describe the historic cost of disk prices in megabytes per US dollar. The long-term trend (blue line, which is a straight line here because the plot is logarithmic) shows exponential growth in storage per dollar with a doubling time of roughly 1.5 years. The cost of DNA sequencing, expressed in base pairs per dollar, is shown by the red triangles. It follows an exponential curve (yellow line) with a doubling time slightly slower than disk storage until 2004, when next generation sequencing (NGS) causes an inflection in the curve to a doubling time of less than 6 months (red line). These curves are not corrected for inflation or for the 'fully loaded' cost of sequencing and disk storage, which would include personnel costs, depreciation and overhead[2]. Classes of Compression: Compression algorithms can broadly be classified into 3 categories, compression using run-length encoding techniques, dictionary based compression and Markov compression.
2 Random data is any compression algorithm's worst enemy. DNA sequences typically consist of 4 characters, viz. A, C, G and T, and their existence in a DNA sequence is close to random. There is very little that any compression algorithm can do in such a situation. Most algorithms today are based on the dictionary based approach, and are interesting extensions to the universal Lempel-Ziv compression algorithm[3]. We believe that it is due to the randomness and unstructured nature of genome sequences, that, till date, the dictionary based models have proven to be the best from the point of view of performance and size. Splitting the Genome: An interesting preprocessing technique shows that we can possibly have some order in this chaos making us revisit the run-length encoding schemes and Markov compression as possibly better options than the current compression techniques. Splitting involves duplicating the DNA sequence into as many copies as the unique characters that are there in DNA, viz. 4. Let's call each copy by the character for which it is intended. For A's copy, we represent all occurrences of the rest of the three characters by a hyphen. In A's copy we now have only 2 characters, A and hyphen. We perform the similar operation on C's copy, G's copy and T's copy. We can now observe that if we superimpose the 4 copies on one another, we get the original sequence back. Even though this preprocessing technique seems trivial and intuitively seems to make an already unwieldy genome sequence even larger in size (contradicting our original objective), it has some very interesting characteristics. Run-Length Encoding: Let us now look at how we can benefit from splitting of the genome from the run-length encoding perspective. To summarize, run-length encoding involves replacing the number of continuous occurrences of a certain entity by a single instance of that entity followed by a number denoting how many times it occurred. Before mentioning how we can exploit splitting, let us first introduce 2 more concepts, Genome Encoding and Variable Integers (VINT). Genome Encoding: As mentioned previously, DNA sequences consist of A, C, G and T. Since there are only 4 characters, we can reduce the 1-byte representation of the characters by only a 2-bit representation, according to the table below. This is one of the most primitive compression methods. A 00 C 01 G 10 T 11 [Table 1] - Genome Encoding of DNA characters
3 Variable Integers: The usual size of an integer is 4 bytes. An integer (of 4 bytes) is thus capable of storing numbers up to in unsigned representation and to in signed representation. Many times, we do not need these many numbers and we end up wasting a lot of space, just to store 0's. To use integers more efficiently, we have the concept of Variable Integers. In VINT, we use only as many bytes to represent the number as required. 1 bit from each byte holding the number is reserved as a flag bit to indicate whether the number ends in the current byte or has overflowed into the next byte. An example of a single byte integer: Integer 6 in the usual representation: Integer 6 in VINT: An example of a multiple byte integer: Integer 1722 in the usual representation: Integer 1722 in VINT: VINT reduces the effective length of each byte by 1 bit, but gives us the flexibility to use only as many bytes as needed, especially effective for small number representations. Proposed Technique: We propose the following technique for run-length based DNA compression: Splitting + Genome Encoding + Run-Length Encoding + VINT For a better explanation for the technique as a whole, let us take an example. In the example, let us do a comparative study of naive run-length encoding with the above mentioned technique. Following is a sample DNA sequence to be compressed: G G G G T C C C A A T T C A G T Naive run-length compression consists of initially genome encoding the sequence before we runlength encode it using normal integers. On genome encoding the above sequence, we get the following bit representation: On performing run-length encoding of the above bit sequence, we get the following:
4 We can observe that 1's and 0's (in black) are alternating. So, we can simply omit them (i.e. just make a note of whether to start with 1 or 0, but for the purpose of this discussion, omit them altogether) and just list their run-lengths to get: Considering 4 byte integers, the above sequence would be 92 bytes long. Let us now work on the same sequence using the technique we introduced above. The first step is to split the original sequence to obtain the following copies: A's copy: A A A - - C's copy: C C C C G's copy: G G G G G - T's copy: T T T T To genome encode the sequence, we can now take advantage of the fact that there are only 2 characters per split sequence. We can thus represent the hyphen by 0 and the corresponding character in each sequence by the bit 1. On genome encoding the split sequence according to the modified scheme, we get: A's copy: C's copy: G's copy: T's copy: If we run-length encode each of the above sequences, we get: A's copy: C's copy: G's copy: T's copy: Again, since there are alternating 0's and 1's, we can simply omit them to get: A's copy: C's copy: G's copy: T's copy: Considering 4 byte integers, to compare with the above results (despite having 4 copies), the total is just 80 bytes. This simple algorithm perform better than naive run-length compression only because of splitting the sequence. VINT is our trump card in distinctly outperforming naive run-length. Before delving into the details of how we use VINT, let us first understand why is it that we chose VINT.
5 Motivation for VINT (Variable Integer): To be able to efficiently encode the run-lengths of hyphens and characters, we need to know what the frequency of their various runs is. Let's plot the histogram of the occurrences of characters and hyphens within a split sequence. The split sequences used in the plot below are of chromosome 22 of the human genome[4]: Histogram"of"characters" Histogram"of"hyphens" [Fig 2] - Histogram of Split Sequence for A The above graph contains two histograms, one (red) for the occurrences of the hyphens within the split sequence and the other (blue) for the occurrences of the characters within the split sequence. The X axis denotes the number of continuous occurrences of hyphen / character and the Y axis shows the frequency of a articular run-length. The X axis is truncated to only runs of length 100 to emphasize on the interesting portion of the graph at the start. The tail of the graph tapers towards zero.
6 Histogram"of"Characters" Histogram"of"hyphens" [Fig 3] - Histogram of Split Sequence for C Histogram"of"Characters" Histogram"of"hyphens" [Fig 4] - Histogram of Split Sequence for G
7 Histogram"of"Characters" Histogram"of"hyphens" [Fig 5] - Histogram of Split Sequence for T Since the histograms of all the copies are similar, we will consider A's copy for the rest of the discussion. As soon as we see a histogram of this form, we are curious to find out what its distribution is like, viz. exponential, power law, etc. One of the main characteristics of any power law function is that its graphical plot on a log-log scale is linear. Following is the distribution of the above plot for character A: Histogram"of"characters" Histogram"of"hyphens" " 1"
8 [Fig 6] - Histogram of Split Sequence for A The above graph is a log-log plot of fig. 2. The linearity of the histograms, when plotted on a log-log scale reveal that in most likelyhood, they have a power-law distribution. The interesting part is the extremely linear plot of the occurrences of the hyphens within a split sequence. Whenever there is a power law distribution that can be associated with an entity, the problem of compression of that entity is reduced to finding the variable-length code that corresponds to the distribution[7]. Thinking on the same lines, we are in the process of finding the function that fits this distribution to find the optimal variable-length code. VINT has a 1-byte granularity. For the purpose of effective compression, we need to make this granularity finer and think in terms of bits. As a result multiple codes can fit within a byte, possibly unveiling a much more efficient compression of genome sequences from the view of run-length compression algorithms. Markov Compression: Another avenue that the power law characteristic opens up, is the possible usage of Markov models to aid compression of these sequences. In this technique, we try to find the best order Markov model that fits our sequence and try compressing the probability state transition matrix that is built as a part of the model analysis. Currently, on analyzing the Markov models of order 1 through 16 on the original DNA sequence and the split DNA sequence for character A, we get the following results. Split"Sequence" Original"Sequence" " 2" 3" 4" 5" 6" 7" 8" 9" 1 11" 12" 13" 14" 15" 16" [Fig 6] - Markov Model Predictability on Original and Split Sequences The X axis on the above graph is the order of the Markov model that was run on the original and split sequences of chr22. The Y axis denotes the percentage of correct predictions that could be done for each of order the Markov models from the X axis (1 through 16). The interesting aspect is the stability (and gradual increase) of predictability in the split sequence against the poor predictability of the original DNA sequence.
9 As seen above, the original sequence's predictability drops drastically after model 12, while the split sequence's predictability remains stable even till 16. The higher the model (assuming it is stable), the more contiguous set of characters are predictable. While better prediction is not directly related to better compression, we are in the process of finding out a way to harness the higher predictability, post splitting, to achieve higher compression. On searching more about Markov compression techniques, we came across a relatively newer algorithm called LZMA[5], which essentially is the universal LZ (dictionary based) technique, but whose dictionary size is allowed to be greater than in the standard technique, and, also whose dictionary is built based on the Markov modeling. As shown above, there is a certain predictability in DNA sequences (less in unsplit ones, but there nevertheless). Intuitively, this algorithm should outperform completely dictionary based techniques viz. gzip and bz2. On quickly running a few tests on split and original DNA sequences, we can see that in fact, it does perform better than both gzip and bzip2 (see table below). Also, like our graphs suggest, the compression is better in the split form than in the whole form. A gzipped split sequence performs slightly better than 100% when compared with the gzipped archive of the original sequence. That ratio is slightly reduced, but still in the high 90's when comparing the sequences using bzip2. The ratio is almost equally maintained by LZMA as well. When comparing amongst the split sequence archives in themselves, LZMA proves to be the algorithm providing maximum compression. Despite these results the overhead of 4 copies outweighs the advantage gained by better predictability. On this front, we are on the search of a more pure form of Markov compression for additional benefit. Chr 22 (Original) Chr 22 (Split - A) Uncompressed GZIP BZIP2 LZMA (~33 MB) (~33 MB) (~9.2 MB) (~4.8 MB) (~8.4 MB) (~4 MB) (~7.4 MB) (~3.4 MB) [Table 2] - Comparison of popular compression algorithms on original and split sequences with LZMA Parallelism in Markov: In many situations, Markov models have much higher compression ratio than LZ techniques, but their runtimes are too slow to be used in practice. Due to splitting, we now have higher predictability and lesser states (as there are only 2 characters within a sequence). This will result in faster compression using Markov techniques. We can parallelize the operation(s) by simultaneously compressing the 4 sequences, since they are completely unrelated with each other for the purpose of compression. DNA Generation - A side effect: When predicting a sequence, Markov models construct a probability state transition matrix. Suppose we are running our sequence through a Markov model of order 12, and our sequence can consist of 2 characters, then the number of states in the state transition matrix are 2^12, i.e., on a general note, the number of unique characters raised to the number of the order that we are
10 considering. This produces all possible arrangements of the characters having length equal to the order we have decided. Using this matrix, we can also construct sequences of arbitrary length having similar characteristics as that of the original. This can prove useful in situations where we want to create a large number of sufficiently random sequences that are based on the characteristics of one (or an average of multiple) sequences as a testbed. While generation of such data can prove helpful in the testing of a computational concept, its biological implications may be interesting (and worth exploring). Future Work: We are interestingly poised on two fronts regarding splitting the genome. On one hand, we are trying to find a function that fits the power-law distribution to construct the optimal variable-length encoding scheme to try and revive run-length compression as one of the best for DNA sequences. On the other hand, we are viewing Markov predictability (combined with parallelism) to be our possible answer for the best DNA compression algorithm. Conclusion: Splitting DNA sequences has given us an interesting view of genomic data. Preliminary tests and analysis are pointing at promising benefits in the area of DNA compression. With further study of this technique, we aim to unlock the answer for the smallest DNA footprint. Acknowledgements: The author thanks Prof. Ming-Yang Kao for his timely guidance and generous help during the course of this study. He also thanks Prof. Peter Dinda for his valuable insight into predictability and Markov techniques. Last, but not the least the author is grateful for the support and encouragement of Dr. Ankit Agarwal. References: [1] C. Kozanitis, C. Saunders, S. Kruglyak, V. Bafna, G. Varghese. Compressing Genomic Sequence Fragments Using SlimGene. Journal of Computation Biology. (2011) 18: [2] L. Stein. The case for cloud computing in genome informatics. Biomed Central. (2010). [3] J. Ziv, A. Lempel. Compression of individual sequences via variable-rate encoding. Information Theory, IEEE Transactions. (1978) 24: [4] Homo sapiens chromosome 22, alternate assembly CHM1_1.0, whole genome shotgun sequence. - GenBank [ [5] I. Pavlov. 7-Zip. Online - [ [6] D. Salomon. Variable-length Codes for Data Compression. Springer. (2007) 2:92-93
Simple variant of coding with a variable number of symbols and fixlength codewords.
 Dictionary coding Simple variant of coding with a variable number of symbols and fixlength codewords. Create a dictionary containing 2 b different symbol sequences and code them with codewords of length
Dictionary coding Simple variant of coding with a variable number of symbols and fixlength codewords. Create a dictionary containing 2 b different symbol sequences and code them with codewords of length
JULIA ENABLED COMPUTATION OF MOLECULAR LIBRARY COMPLEXITY IN DNA SEQUENCING
 JULIA ENABLED COMPUTATION OF MOLECULAR LIBRARY COMPLEXITY IN DNA SEQUENCING Larson Hogstrom, Mukarram Tahir, Andres Hasfura Massachusetts Institute of Technology, Cambridge, Massachusetts, USA 18.337/6.338
JULIA ENABLED COMPUTATION OF MOLECULAR LIBRARY COMPLEXITY IN DNA SEQUENCING Larson Hogstrom, Mukarram Tahir, Andres Hasfura Massachusetts Institute of Technology, Cambridge, Massachusetts, USA 18.337/6.338
Lossless Compression Algorithms
 Multimedia Data Compression Part I Chapter 7 Lossless Compression Algorithms 1 Chapter 7 Lossless Compression Algorithms 1. Introduction 2. Basics of Information Theory 3. Lossless Compression Algorithms
Multimedia Data Compression Part I Chapter 7 Lossless Compression Algorithms 1 Chapter 7 Lossless Compression Algorithms 1. Introduction 2. Basics of Information Theory 3. Lossless Compression Algorithms
Evolutionary Lossless Compression with GP-ZIP
 Evolutionary Lossless Compression with GP-ZIP Ahmad Kattan and Riccardo Poli Abstract In this paper we propose a new approach for applying Genetic Programming to lossless data compression based on combining
Evolutionary Lossless Compression with GP-ZIP Ahmad Kattan and Riccardo Poli Abstract In this paper we propose a new approach for applying Genetic Programming to lossless data compression based on combining
Basic Compression Library
 Basic Compression Library Manual API version 1.2 July 22, 2006 c 2003-2006 Marcus Geelnard Summary This document describes the algorithms used in the Basic Compression Library, and how to use the library
Basic Compression Library Manual API version 1.2 July 22, 2006 c 2003-2006 Marcus Geelnard Summary This document describes the algorithms used in the Basic Compression Library, and how to use the library
Wednesday, May 3, Several RAID "levels" have been defined. Some are more commercially viable than others.
 Wednesday, May 3, 2017 Topics for today RAID: Level 0 Level 1 Level 3 Level 4 Level 5 Beyond RAID 5 File systems RAID revisited Several RAID "levels" have been defined. Some are more commercially viable
Wednesday, May 3, 2017 Topics for today RAID: Level 0 Level 1 Level 3 Level 4 Level 5 Beyond RAID 5 File systems RAID revisited Several RAID "levels" have been defined. Some are more commercially viable
Data Compression for Bitmap Indexes. Y. Chen
 Data Compression for Bitmap Indexes Y. Chen Abstract Compression Ratio (CR) and Logical Operation Time (LOT) are two major measures of the efficiency of bitmap indexing. Previous works by [5, 9, 10, 11]
Data Compression for Bitmap Indexes Y. Chen Abstract Compression Ratio (CR) and Logical Operation Time (LOT) are two major measures of the efficiency of bitmap indexing. Previous works by [5, 9, 10, 11]
Demand fetching is commonly employed to bring the data
 Proceedings of 2nd Annual Conference on Theoretical and Applied Computer Science, November 2010, Stillwater, OK 14 Markov Prediction Scheme for Cache Prefetching Pranav Pathak, Mehedi Sarwar, Sohum Sohoni
Proceedings of 2nd Annual Conference on Theoretical and Applied Computer Science, November 2010, Stillwater, OK 14 Markov Prediction Scheme for Cache Prefetching Pranav Pathak, Mehedi Sarwar, Sohum Sohoni
6.2 DATA DISTRIBUTION AND EXPERIMENT DETAILS
 Chapter 6 Indexing Results 6. INTRODUCTION The generation of inverted indexes for text databases is a computationally intensive process that requires the exclusive use of processing resources for long
Chapter 6 Indexing Results 6. INTRODUCTION The generation of inverted indexes for text databases is a computationally intensive process that requires the exclusive use of processing resources for long
From Smith-Waterman to BLAST
 From Smith-Waterman to BLAST Jeremy Buhler July 23, 2015 Smith-Waterman is the fundamental tool that we use to decide how similar two sequences are. Isn t that all that BLAST does? In principle, it is
From Smith-Waterman to BLAST Jeremy Buhler July 23, 2015 Smith-Waterman is the fundamental tool that we use to decide how similar two sequences are. Isn t that all that BLAST does? In principle, it is
Multimedia Networking ECE 599
 Multimedia Networking ECE 599 Prof. Thinh Nguyen School of Electrical Engineering and Computer Science Based on B. Lee s lecture notes. 1 Outline Compression basics Entropy and information theory basics
Multimedia Networking ECE 599 Prof. Thinh Nguyen School of Electrical Engineering and Computer Science Based on B. Lee s lecture notes. 1 Outline Compression basics Entropy and information theory basics
Inverted Indexes. Indexing and Searching, Modern Information Retrieval, Addison Wesley, 2010 p. 5
 Inverted Indexes Indexing and Searching, Modern Information Retrieval, Addison Wesley, 2010 p. 5 Basic Concepts Inverted index: a word-oriented mechanism for indexing a text collection to speed up the
Inverted Indexes Indexing and Searching, Modern Information Retrieval, Addison Wesley, 2010 p. 5 Basic Concepts Inverted index: a word-oriented mechanism for indexing a text collection to speed up the
Optimized Compression and Decompression Software
 2015 IJSRSET Volume 1 Issue 3 Print ISSN : 2395-1990 Online ISSN : 2394-4099 Themed Section: Engineering and Technology Optimized Compression and Decompression Software Mohd Shafaat Hussain, Manoj Yadav
2015 IJSRSET Volume 1 Issue 3 Print ISSN : 2395-1990 Online ISSN : 2394-4099 Themed Section: Engineering and Technology Optimized Compression and Decompression Software Mohd Shafaat Hussain, Manoj Yadav
Efficiency of Data Distribution in BitTorrent-Like Systems
 Efficiency of Data Distribution in BitTorrent-Like Systems Ho-Leung Chan 1,, Tak-Wah Lam 2, and Prudence W.H. Wong 3 1 Department of Computer Science, University of Pittsburgh hlchan@cs.pitt.edu 2 Department
Efficiency of Data Distribution in BitTorrent-Like Systems Ho-Leung Chan 1,, Tak-Wah Lam 2, and Prudence W.H. Wong 3 1 Department of Computer Science, University of Pittsburgh hlchan@cs.pitt.edu 2 Department
Data Structures and Algorithms Dr. Naveen Garg Department of Computer Science and Engineering Indian Institute of Technology, Delhi.
 Data Structures and Algorithms Dr. Naveen Garg Department of Computer Science and Engineering Indian Institute of Technology, Delhi Lecture 18 Tries Today we are going to be talking about another data
Data Structures and Algorithms Dr. Naveen Garg Department of Computer Science and Engineering Indian Institute of Technology, Delhi Lecture 18 Tries Today we are going to be talking about another data
Parallel Computing Concepts. CSInParallel Project
 Parallel Computing Concepts CSInParallel Project July 26, 2012 CONTENTS 1 Introduction 1 1.1 Motivation................................................ 1 1.2 Some pairs of terms...........................................
Parallel Computing Concepts CSInParallel Project July 26, 2012 CONTENTS 1 Introduction 1 1.1 Motivation................................................ 1 1.2 Some pairs of terms...........................................
Ch. 2: Compression Basics Multimedia Systems
 Ch. 2: Compression Basics Multimedia Systems Prof. Ben Lee School of Electrical Engineering and Computer Science Oregon State University Outline Why compression? Classification Entropy and Information
Ch. 2: Compression Basics Multimedia Systems Prof. Ben Lee School of Electrical Engineering and Computer Science Oregon State University Outline Why compression? Classification Entropy and Information
Administrative. Distributed indexing. Index Compression! What I did last summer lunch talks today. Master. Tasks
 Administrative Index Compression! n Assignment 1? n Homework 2 out n What I did last summer lunch talks today David Kauchak cs458 Fall 2012 adapted from: http://www.stanford.edu/class/cs276/handouts/lecture5-indexcompression.ppt
Administrative Index Compression! n Assignment 1? n Homework 2 out n What I did last summer lunch talks today David Kauchak cs458 Fall 2012 adapted from: http://www.stanford.edu/class/cs276/handouts/lecture5-indexcompression.ppt
Introduction to carving File fragmentation Object validation Carving methods Conclusion
 Simson L. Garfinkel Presented by Jevin Sweval Introduction to carving File fragmentation Object validation Carving methods Conclusion 1 Carving is the recovery of files from a raw dump of a storage device
Simson L. Garfinkel Presented by Jevin Sweval Introduction to carving File fragmentation Object validation Carving methods Conclusion 1 Carving is the recovery of files from a raw dump of a storage device
MIGRATORY COMPRESSION Coarse-grained Data Reordering to Improve Compressibility
 MIGRATORY COMPRESSION Coarse-grained Data Reordering to Improve Compressibility Xing Lin *, Guanlin Lu, Fred Douglis, Philip Shilane, Grant Wallace * University of Utah EMC Corporation Data Protection
MIGRATORY COMPRESSION Coarse-grained Data Reordering to Improve Compressibility Xing Lin *, Guanlin Lu, Fred Douglis, Philip Shilane, Grant Wallace * University of Utah EMC Corporation Data Protection
To Zip or not to Zip. Effective Resource Usage for Real-Time Compression
 To Zip or not to Zip Effective Resource Usage for Real-Time Compression Danny Harnik, Oded Margalit, Ronen Kat, Dmitry Sotnikov, Avishay Traeger IBM Research - Haifa Our scope Real-Time Compression Compression
To Zip or not to Zip Effective Resource Usage for Real-Time Compression Danny Harnik, Oded Margalit, Ronen Kat, Dmitry Sotnikov, Avishay Traeger IBM Research - Haifa Our scope Real-Time Compression Compression
Index Compression. David Kauchak cs160 Fall 2009 adapted from:
 Index Compression David Kauchak cs160 Fall 2009 adapted from: http://www.stanford.edu/class/cs276/handouts/lecture5-indexcompression.ppt Administrative Homework 2 Assignment 1 Assignment 2 Pair programming?
Index Compression David Kauchak cs160 Fall 2009 adapted from: http://www.stanford.edu/class/cs276/handouts/lecture5-indexcompression.ppt Administrative Homework 2 Assignment 1 Assignment 2 Pair programming?
Scalable Trigram Backoff Language Models
 Scalable Trigram Backoff Language Models Kristie Seymore Ronald Rosenfeld May 1996 CMU-CS-96-139 School of Computer Science Carnegie Mellon University Pittsburgh, PA 15213 This material is based upon work
Scalable Trigram Backoff Language Models Kristie Seymore Ronald Rosenfeld May 1996 CMU-CS-96-139 School of Computer Science Carnegie Mellon University Pittsburgh, PA 15213 This material is based upon work
DESIGN AND ANALYSIS OF ALGORITHMS. Unit 1 Chapter 4 ITERATIVE ALGORITHM DESIGN ISSUES
 DESIGN AND ANALYSIS OF ALGORITHMS Unit 1 Chapter 4 ITERATIVE ALGORITHM DESIGN ISSUES http://milanvachhani.blogspot.in USE OF LOOPS As we break down algorithm into sub-algorithms, sooner or later we shall
DESIGN AND ANALYSIS OF ALGORITHMS Unit 1 Chapter 4 ITERATIVE ALGORITHM DESIGN ISSUES http://milanvachhani.blogspot.in USE OF LOOPS As we break down algorithm into sub-algorithms, sooner or later we shall
Comparing Implementations of Optimal Binary Search Trees
 Introduction Comparing Implementations of Optimal Binary Search Trees Corianna Jacoby and Alex King Tufts University May 2017 In this paper we sought to put together a practical comparison of the optimality
Introduction Comparing Implementations of Optimal Binary Search Trees Corianna Jacoby and Alex King Tufts University May 2017 In this paper we sought to put together a practical comparison of the optimality
Connections in Networks: A Hybrid Approach
 Connections in Networks: A Hybrid Approach Carla P. Gomes 1 Willem-Jan van Hoeve 2 Ashish Sabharwal 1 1 Department of Computer Science, Cornell University, Ithaca NY 14853, U.S.A. {gomes,sabhar}@cs.cornell.edu
Connections in Networks: A Hybrid Approach Carla P. Gomes 1 Willem-Jan van Hoeve 2 Ashish Sabharwal 1 1 Department of Computer Science, Cornell University, Ithaca NY 14853, U.S.A. {gomes,sabhar}@cs.cornell.edu
Determining the Number of CPUs for Query Processing
 Determining the Number of CPUs for Query Processing Fatemah Panahi Elizabeth Soechting CS747 Advanced Computer Systems Analysis Techniques The University of Wisconsin-Madison fatemeh@cs.wisc.edu, eas@cs.wisc.edu
Determining the Number of CPUs for Query Processing Fatemah Panahi Elizabeth Soechting CS747 Advanced Computer Systems Analysis Techniques The University of Wisconsin-Madison fatemeh@cs.wisc.edu, eas@cs.wisc.edu
Compression in Open Source Databases. Peter Zaitsev April 20, 2016
 Compression in Open Source Databases Peter Zaitsev April 20, 2016 About the Talk 2 A bit of the History Approaches to Data Compression What some of the popular systems implement 2 Lets Define The Term
Compression in Open Source Databases Peter Zaitsev April 20, 2016 About the Talk 2 A bit of the History Approaches to Data Compression What some of the popular systems implement 2 Lets Define The Term
Greedy Algorithms CHAPTER 16
 CHAPTER 16 Greedy Algorithms In dynamic programming, the optimal solution is described in a recursive manner, and then is computed ``bottom up''. Dynamic programming is a powerful technique, but it often
CHAPTER 16 Greedy Algorithms In dynamic programming, the optimal solution is described in a recursive manner, and then is computed ``bottom up''. Dynamic programming is a powerful technique, but it often
INSPIRE and SPIRES Log File Analysis
 INSPIRE and SPIRES Log File Analysis Cole Adams Science Undergraduate Laboratory Internship Program Wheaton College SLAC National Accelerator Laboratory August 5, 2011 Prepared in partial fulfillment of
INSPIRE and SPIRES Log File Analysis Cole Adams Science Undergraduate Laboratory Internship Program Wheaton College SLAC National Accelerator Laboratory August 5, 2011 Prepared in partial fulfillment of
Unit 1 Chapter 4 ITERATIVE ALGORITHM DESIGN ISSUES
 DESIGN AND ANALYSIS OF ALGORITHMS Unit 1 Chapter 4 ITERATIVE ALGORITHM DESIGN ISSUES http://milanvachhani.blogspot.in USE OF LOOPS As we break down algorithm into sub-algorithms, sooner or later we shall
DESIGN AND ANALYSIS OF ALGORITHMS Unit 1 Chapter 4 ITERATIVE ALGORITHM DESIGN ISSUES http://milanvachhani.blogspot.in USE OF LOOPS As we break down algorithm into sub-algorithms, sooner or later we shall
Technical lossless / near lossless data compression
 Technical lossless / near lossless data compression Nigel Atkinson (Met Office, UK) ECMWF/EUMETSAT NWP SAF Workshop 5-7 Nov 2013 Contents Survey of file compression tools Studies for AVIRIS imager Study
Technical lossless / near lossless data compression Nigel Atkinson (Met Office, UK) ECMWF/EUMETSAT NWP SAF Workshop 5-7 Nov 2013 Contents Survey of file compression tools Studies for AVIRIS imager Study
GenCodex - A Novel Algorithm for Compressing DNA sequences on Multi-cores and GPUs
 GenCodex - A Novel Algorithm for ing DNA sequences on Multi-cores and GPUs D Satyanvesh, Kaliuday Balleda, Ajith Padyana, P K Baruah Sri Sathya Sai Institute of Higher Learning, Prasanthi Nilayam, India
GenCodex - A Novel Algorithm for ing DNA sequences on Multi-cores and GPUs D Satyanvesh, Kaliuday Balleda, Ajith Padyana, P K Baruah Sri Sathya Sai Institute of Higher Learning, Prasanthi Nilayam, India
Memory Management. Kevin Webb Swarthmore College February 27, 2018
 Memory Management Kevin Webb Swarthmore College February 27, 2018 Today s Goals Shifting topics: different process resource memory Motivate virtual memory, including what it might look like without it
Memory Management Kevin Webb Swarthmore College February 27, 2018 Today s Goals Shifting topics: different process resource memory Motivate virtual memory, including what it might look like without it
Modeling Delta Encoding of Compressed Files
 Modeling Delta Encoding of Compressed Files EXTENDED ABSTRACT S.T. Klein, T.C. Serebro, and D. Shapira 1 Dept of CS Bar Ilan University Ramat Gan, Israel tomi@cs.biu.ac.il 2 Dept of CS Bar Ilan University
Modeling Delta Encoding of Compressed Files EXTENDED ABSTRACT S.T. Klein, T.C. Serebro, and D. Shapira 1 Dept of CS Bar Ilan University Ramat Gan, Israel tomi@cs.biu.ac.il 2 Dept of CS Bar Ilan University
CSE332: Data Abstractions Lecture 7: B Trees. James Fogarty Winter 2012
 CSE2: Data Abstractions Lecture 7: B Trees James Fogarty Winter 20 The Dictionary (a.k.a. Map) ADT Data: Set of (key, value) pairs keys must be comparable insert(jfogarty,.) Operations: insert(key,value)
CSE2: Data Abstractions Lecture 7: B Trees James Fogarty Winter 20 The Dictionary (a.k.a. Map) ADT Data: Set of (key, value) pairs keys must be comparable insert(jfogarty,.) Operations: insert(key,value)
Binary Encoded Attribute-Pairing Technique for Database Compression
 Binary Encoded Attribute-Pairing Technique for Database Compression Akanksha Baid and Swetha Krishnan Computer Sciences Department University of Wisconsin, Madison baid,swetha@cs.wisc.edu Abstract Data
Binary Encoded Attribute-Pairing Technique for Database Compression Akanksha Baid and Swetha Krishnan Computer Sciences Department University of Wisconsin, Madison baid,swetha@cs.wisc.edu Abstract Data
Study of LZ77 and LZ78 Data Compression Techniques
 Study of LZ77 and LZ78 Data Compression Techniques Suman M. Choudhary, Anjali S. Patel, Sonal J. Parmar Abstract Data Compression is defined as the science and art of the representation of information
Study of LZ77 and LZ78 Data Compression Techniques Suman M. Choudhary, Anjali S. Patel, Sonal J. Parmar Abstract Data Compression is defined as the science and art of the representation of information
Compressing Data. Konstantin Tretyakov
 Compressing Data Konstantin Tretyakov (kt@ut.ee) MTAT.03.238 Advanced April 26, 2012 Claude Elwood Shannon (1916-2001) C. E. Shannon. A mathematical theory of communication. 1948 C. E. Shannon. The mathematical
Compressing Data Konstantin Tretyakov (kt@ut.ee) MTAT.03.238 Advanced April 26, 2012 Claude Elwood Shannon (1916-2001) C. E. Shannon. A mathematical theory of communication. 1948 C. E. Shannon. The mathematical
Project Proposal. ECE 526 Spring Modified Data Structure of Aho-Corasick. Benfano Soewito, Ed Flanigan and John Pangrazio
 Project Proposal ECE 526 Spring 2006 Modified Data Structure of Aho-Corasick Benfano Soewito, Ed Flanigan and John Pangrazio 1. Introduction The internet becomes the most important tool in this decade
Project Proposal ECE 526 Spring 2006 Modified Data Structure of Aho-Corasick Benfano Soewito, Ed Flanigan and John Pangrazio 1. Introduction The internet becomes the most important tool in this decade
Information Retrieval
 Information Retrieval Suan Lee - Information Retrieval - 05 Index Compression 1 05 Index Compression - Information Retrieval - 05 Index Compression 2 Last lecture index construction Sort-based indexing
Information Retrieval Suan Lee - Information Retrieval - 05 Index Compression 1 05 Index Compression - Information Retrieval - 05 Index Compression 2 Last lecture index construction Sort-based indexing
Introduction to Operating Systems
 Introduction to Operating Systems Lecture 6: Memory Management MING GAO SE@ecnu (for course related communications) mgao@sei.ecnu.edu.cn Apr. 22, 2015 Outline 1 Issues of main memory 2 Main memory management
Introduction to Operating Systems Lecture 6: Memory Management MING GAO SE@ecnu (for course related communications) mgao@sei.ecnu.edu.cn Apr. 22, 2015 Outline 1 Issues of main memory 2 Main memory management
12 Algorithms in Bioinformatics I, WS 06, ZBIT, D. Huson, October 18, 2006
 12 Algorithms in Bioinformatics I, WS 06, ZBIT, D. Huson, October 18, 2006 3 Sequence comparison by compression This chapter is based on the following articles, which are all recommended reading: X. Chen,
12 Algorithms in Bioinformatics I, WS 06, ZBIT, D. Huson, October 18, 2006 3 Sequence comparison by compression This chapter is based on the following articles, which are all recommended reading: X. Chen,
Analyzing Data Properties using Statistical Sampling Techniques
 Analyzing Data Properties using Statistical Sampling Techniques Illustrated on Scientific File Formats and Compression Features Julian M. Kunkel kunkel@dkrz.de June 2, 2016 Outline 1 Introduction 2 Exploring
Analyzing Data Properties using Statistical Sampling Techniques Illustrated on Scientific File Formats and Compression Features Julian M. Kunkel kunkel@dkrz.de June 2, 2016 Outline 1 Introduction 2 Exploring
CS 445 HW#6 Solutions
 CS 445 HW#6 Solutions Text problem 6.1 From the figure, x = 0.43 and y = 0.4. Since x + y + z = 1, it follows that z = 0.17. These are the trichromatic coefficients. We are interested in tristimulus values
CS 445 HW#6 Solutions Text problem 6.1 From the figure, x = 0.43 and y = 0.4. Since x + y + z = 1, it follows that z = 0.17. These are the trichromatic coefficients. We are interested in tristimulus values
An Asymmetric, Semi-adaptive Text Compression Algorithm
 An Asymmetric, Semi-adaptive Text Compression Algorithm Harry Plantinga Department of Computer Science University of Pittsburgh Pittsburgh, PA 15260 planting@cs.pitt.edu Abstract A new heuristic for text
An Asymmetric, Semi-adaptive Text Compression Algorithm Harry Plantinga Department of Computer Science University of Pittsburgh Pittsburgh, PA 15260 planting@cs.pitt.edu Abstract A new heuristic for text
Brief review from last class
 Sequence Alignment Brief review from last class DNA is has direction, we will use only one (5 -> 3 ) and generate the opposite strand as needed. DNA is a 3D object (see lecture 1) but we will model it
Sequence Alignment Brief review from last class DNA is has direction, we will use only one (5 -> 3 ) and generate the opposite strand as needed. DNA is a 3D object (see lecture 1) but we will model it
Hardware Modeling using Verilog Prof. Indranil Sengupta Department of Computer Science and Engineering Indian Institute of Technology, Kharagpur
 Hardware Modeling using Verilog Prof. Indranil Sengupta Department of Computer Science and Engineering Indian Institute of Technology, Kharagpur Lecture 01 Introduction Welcome to the course on Hardware
Hardware Modeling using Verilog Prof. Indranil Sengupta Department of Computer Science and Engineering Indian Institute of Technology, Kharagpur Lecture 01 Introduction Welcome to the course on Hardware
Analysis of Parallelization Effects on Textual Data Compression
 Analysis of Parallelization Effects on Textual Data GORAN MARTINOVIC, CASLAV LIVADA, DRAGO ZAGAR Faculty of Electrical Engineering Josip Juraj Strossmayer University of Osijek Kneza Trpimira 2b, 31000
Analysis of Parallelization Effects on Textual Data GORAN MARTINOVIC, CASLAV LIVADA, DRAGO ZAGAR Faculty of Electrical Engineering Josip Juraj Strossmayer University of Osijek Kneza Trpimira 2b, 31000
Suffix Vector: A Space-Efficient Suffix Tree Representation
 Lecture Notes in Computer Science 1 Suffix Vector: A Space-Efficient Suffix Tree Representation Krisztián Monostori 1, Arkady Zaslavsky 1, and István Vajk 2 1 School of Computer Science and Software Engineering,
Lecture Notes in Computer Science 1 Suffix Vector: A Space-Efficient Suffix Tree Representation Krisztián Monostori 1, Arkady Zaslavsky 1, and István Vajk 2 1 School of Computer Science and Software Engineering,
A STUDY OF THE PERFORMANCE TRADEOFFS OF A TRADE ARCHIVE
 A STUDY OF THE PERFORMANCE TRADEOFFS OF A TRADE ARCHIVE CS737 PROJECT REPORT Anurag Gupta David Goldman Han-Yin Chen {anurag, goldman, han-yin}@cs.wisc.edu Computer Sciences Department University of Wisconsin,
A STUDY OF THE PERFORMANCE TRADEOFFS OF A TRADE ARCHIVE CS737 PROJECT REPORT Anurag Gupta David Goldman Han-Yin Chen {anurag, goldman, han-yin}@cs.wisc.edu Computer Sciences Department University of Wisconsin,
Information Retrieval
 Introduction to Information Retrieval CS3245 Information Retrieval Lecture 6: Index Compression 6 Last Time: index construction Sort- based indexing Blocked Sort- Based Indexing Merge sort is effective
Introduction to Information Retrieval CS3245 Information Retrieval Lecture 6: Index Compression 6 Last Time: index construction Sort- based indexing Blocked Sort- Based Indexing Merge sort is effective
So, what is data compression, and why do we need it?
 In the last decade we have been witnessing a revolution in the way we communicate 2 The major contributors in this revolution are: Internet; The explosive development of mobile communications; and The
In the last decade we have been witnessing a revolution in the way we communicate 2 The major contributors in this revolution are: Internet; The explosive development of mobile communications; and The
You can say that again! Text compression
 Activity 3 You can say that again! Text compression Age group Early elementary and up. Abilities assumed Copying written text. Time 10 minutes or more. Size of group From individuals to the whole class.
Activity 3 You can say that again! Text compression Age group Early elementary and up. Abilities assumed Copying written text. Time 10 minutes or more. Size of group From individuals to the whole class.
Genetic Programming. Charles Chilaka. Department of Computational Science Memorial University of Newfoundland
 Genetic Programming Charles Chilaka Department of Computational Science Memorial University of Newfoundland Class Project for Bio 4241 March 27, 2014 Charles Chilaka (MUN) Genetic algorithms and programming
Genetic Programming Charles Chilaka Department of Computational Science Memorial University of Newfoundland Class Project for Bio 4241 March 27, 2014 Charles Chilaka (MUN) Genetic algorithms and programming
Computational biology course IST 2015/2016
 Computational biology course IST 2015/2016 Introduc)on to Algorithms! Algorithms: problem- solving methods suitable for implementation as a computer program! Data structures: objects created to organize
Computational biology course IST 2015/2016 Introduc)on to Algorithms! Algorithms: problem- solving methods suitable for implementation as a computer program! Data structures: objects created to organize
(Refer Slide Time: 01.26)
 Data Structures and Algorithms Dr. Naveen Garg Department of Computer Science and Engineering Indian Institute of Technology, Delhi Lecture # 22 Why Sorting? Today we are going to be looking at sorting.
Data Structures and Algorithms Dr. Naveen Garg Department of Computer Science and Engineering Indian Institute of Technology, Delhi Lecture # 22 Why Sorting? Today we are going to be looking at sorting.
Indexing and Searching
 Indexing and Searching Introduction How to retrieval information? A simple alternative is to search the whole text sequentially Another option is to build data structures over the text (called indices)
Indexing and Searching Introduction How to retrieval information? A simple alternative is to search the whole text sequentially Another option is to build data structures over the text (called indices)
Digital Communication Prof. Bikash Kumar Dey Department of Electrical Engineering Indian Institute of Technology, Bombay
 Digital Communication Prof. Bikash Kumar Dey Department of Electrical Engineering Indian Institute of Technology, Bombay Lecture - 29 Source Coding (Part-4) We have already had 3 classes on source coding
Digital Communication Prof. Bikash Kumar Dey Department of Electrical Engineering Indian Institute of Technology, Bombay Lecture - 29 Source Coding (Part-4) We have already had 3 classes on source coding
PARALLEL MULTI-DELAY SIMULATION
 PARALLEL MULTI-DELAY SIMULATION Yun Sik Lee Peter M. Maurer Department of Computer Science and Engineering University of South Florida Tampa, FL 33620 CATEGORY: 7 - Discrete Simulation PARALLEL MULTI-DELAY
PARALLEL MULTI-DELAY SIMULATION Yun Sik Lee Peter M. Maurer Department of Computer Science and Engineering University of South Florida Tampa, FL 33620 CATEGORY: 7 - Discrete Simulation PARALLEL MULTI-DELAY
MODELING DELTA ENCODING OF COMPRESSED FILES. and. and
 International Journal of Foundations of Computer Science c World Scientific Publishing Company MODELING DELTA ENCODING OF COMPRESSED FILES SHMUEL T. KLEIN Department of Computer Science, Bar-Ilan University
International Journal of Foundations of Computer Science c World Scientific Publishing Company MODELING DELTA ENCODING OF COMPRESSED FILES SHMUEL T. KLEIN Department of Computer Science, Bar-Ilan University
DATABASE SCALABILITY AND CLUSTERING
 WHITE PAPER DATABASE SCALABILITY AND CLUSTERING As application architectures become increasingly dependent on distributed communication and processing, it is extremely important to understand where the
WHITE PAPER DATABASE SCALABILITY AND CLUSTERING As application architectures become increasingly dependent on distributed communication and processing, it is extremely important to understand where the
A QUAD-TREE DECOMPOSITION APPROACH TO CARTOON IMAGE COMPRESSION. Yi-Chen Tsai, Ming-Sui Lee, Meiyin Shen and C.-C. Jay Kuo
 A QUAD-TREE DECOMPOSITION APPROACH TO CARTOON IMAGE COMPRESSION Yi-Chen Tsai, Ming-Sui Lee, Meiyin Shen and C.-C. Jay Kuo Integrated Media Systems Center and Department of Electrical Engineering University
A QUAD-TREE DECOMPOSITION APPROACH TO CARTOON IMAGE COMPRESSION Yi-Chen Tsai, Ming-Sui Lee, Meiyin Shen and C.-C. Jay Kuo Integrated Media Systems Center and Department of Electrical Engineering University
AM 221: Advanced Optimization Spring 2016
 AM 221: Advanced Optimization Spring 2016 Prof Yaron Singer Lecture 3 February 1st 1 Overview In our previous lecture we presented fundamental results from convex analysis and in particular the separating
AM 221: Advanced Optimization Spring 2016 Prof Yaron Singer Lecture 3 February 1st 1 Overview In our previous lecture we presented fundamental results from convex analysis and in particular the separating
S U N G - E U I YO O N, K A I S T R E N D E R I N G F R E E LY A VA I L A B L E O N T H E I N T E R N E T
 S U N G - E U I YO O N, K A I S T R E N D E R I N G F R E E LY A VA I L A B L E O N T H E I N T E R N E T Copyright 2018 Sung-eui Yoon, KAIST freely available on the internet http://sglab.kaist.ac.kr/~sungeui/render
S U N G - E U I YO O N, K A I S T R E N D E R I N G F R E E LY A VA I L A B L E O N T H E I N T E R N E T Copyright 2018 Sung-eui Yoon, KAIST freely available on the internet http://sglab.kaist.ac.kr/~sungeui/render
Comparison of Existing Dictionary Based Data Compression Methods for English and Gujarati Text # MCA,SRIMCA
 Comparison of Existing Dictionary Based Data Compression Methods for English and Gujarati Sandip Maniya #1, Jikitsha Sheth *2, Dr. Kalpesh Lad *3 # MCA,SRIMCA Uka Tarsadia University, Maliba Campus, Dist.
Comparison of Existing Dictionary Based Data Compression Methods for English and Gujarati Sandip Maniya #1, Jikitsha Sheth *2, Dr. Kalpesh Lad *3 # MCA,SRIMCA Uka Tarsadia University, Maliba Campus, Dist.
Text Compression. Jayadev Misra The University of Texas at Austin July 1, A Very Incomplete Introduction To Information Theory 2
 Text Compression Jayadev Misra The University of Texas at Austin July 1, 2003 Contents 1 Introduction 1 2 A Very Incomplete Introduction To Information Theory 2 3 Huffman Coding 5 3.1 Uniquely Decodable
Text Compression Jayadev Misra The University of Texas at Austin July 1, 2003 Contents 1 Introduction 1 2 A Very Incomplete Introduction To Information Theory 2 3 Huffman Coding 5 3.1 Uniquely Decodable
Introduction to Information Retrieval
 Introduction to Information Retrieval http://informationretrieval.org IIR 5: Index Compression Hinrich Schütze Center for Information and Language Processing, University of Munich 2014-04-17 1/59 Overview
Introduction to Information Retrieval http://informationretrieval.org IIR 5: Index Compression Hinrich Schütze Center for Information and Language Processing, University of Munich 2014-04-17 1/59 Overview
Formal Model. Figure 1: The target concept T is a subset of the concept S = [0, 1]. The search agent needs to search S for a point in T.
![Formal Model. Figure 1: The target concept T is a subset of the concept S = [0, 1]. The search agent needs to search S for a point in T. Formal Model. Figure 1: The target concept T is a subset of the concept S = [0, 1]. The search agent needs to search S for a point in T.](/thumbs/72/66819767.jpg) Although this paper analyzes shaping with respect to its benefits on search problems, the reader should recognize that shaping is often intimately related to reinforcement learning. The objective in reinforcement
Although this paper analyzes shaping with respect to its benefits on search problems, the reader should recognize that shaping is often intimately related to reinforcement learning. The objective in reinforcement
Data Compression Techniques
 Data Compression Techniques Part 2: Text Compression Lecture 6: Dictionary Compression Juha Kärkkäinen 15.11.2017 1 / 17 Dictionary Compression The compression techniques we have seen so far replace individual
Data Compression Techniques Part 2: Text Compression Lecture 6: Dictionary Compression Juha Kärkkäinen 15.11.2017 1 / 17 Dictionary Compression The compression techniques we have seen so far replace individual
Untyped Memory in the Java Virtual Machine
 Untyped Memory in the Java Virtual Machine Andreas Gal and Michael Franz University of California, Irvine {gal,franz}@uci.edu Christian W. Probst Technical University of Denmark probst@imm.dtu.dk July
Untyped Memory in the Java Virtual Machine Andreas Gal and Michael Franz University of California, Irvine {gal,franz}@uci.edu Christian W. Probst Technical University of Denmark probst@imm.dtu.dk July
Database Management System Prof. D. Janakiram Department of Computer Science & Engineering Indian Institute of Technology, Madras Lecture No.
 Database Management System Prof. D. Janakiram Department of Computer Science & Engineering Indian Institute of Technology, Madras Lecture No. # 20 Concurrency Control Part -1 Foundations for concurrency
Database Management System Prof. D. Janakiram Department of Computer Science & Engineering Indian Institute of Technology, Madras Lecture No. # 20 Concurrency Control Part -1 Foundations for concurrency
EFFICIENT ATTACKS ON HOMOPHONIC SUBSTITUTION CIPHERS
 EFFICIENT ATTACKS ON HOMOPHONIC SUBSTITUTION CIPHERS A Project Report Presented to The faculty of the Department of Computer Science San Jose State University In Partial Fulfillment of the Requirements
EFFICIENT ATTACKS ON HOMOPHONIC SUBSTITUTION CIPHERS A Project Report Presented to The faculty of the Department of Computer Science San Jose State University In Partial Fulfillment of the Requirements
Figure-2.1. Information system with encoder/decoders.
 2. Entropy Coding In the section on Information Theory, information system is modeled as the generationtransmission-user triplet, as depicted in fig-1.1, to emphasize the information aspect of the system.
2. Entropy Coding In the section on Information Theory, information system is modeled as the generationtransmission-user triplet, as depicted in fig-1.1, to emphasize the information aspect of the system.
A Comparison of Alternative Encoding Mechanisms for Web Services 1
 A Comparison of Alternative Encoding Mechanisms for Web Services 1 Min Cai, Shahram Ghandeharizadeh, Rolfe Schmidt, Saihong Song Computer Science Department University of Southern California Los Angeles,
A Comparison of Alternative Encoding Mechanisms for Web Services 1 Min Cai, Shahram Ghandeharizadeh, Rolfe Schmidt, Saihong Song Computer Science Department University of Southern California Los Angeles,
Algorithm Analysis. Big Oh
 Algorithm Analysis with Big Oh Data Structures and Design with Java and JUnit Chapter 12 Rick Mercer Algorithm Analysis w Objectives Analyze the efficiency of algorithms Analyze a few classic algorithms
Algorithm Analysis with Big Oh Data Structures and Design with Java and JUnit Chapter 12 Rick Mercer Algorithm Analysis w Objectives Analyze the efficiency of algorithms Analyze a few classic algorithms
Improving Logic Obfuscation via Logic Cone Analysis
 Improving Logic Obfuscation via Logic Cone Analysis Yu-Wei Lee and Nur A. Touba Computer Engineering Research Center University of Texas, Austin, TX 78712 ywlee@utexas.edu, touba@utexas.edu Abstract -
Improving Logic Obfuscation via Logic Cone Analysis Yu-Wei Lee and Nur A. Touba Computer Engineering Research Center University of Texas, Austin, TX 78712 ywlee@utexas.edu, touba@utexas.edu Abstract -
Compact Encoding of the Web Graph Exploiting Various Power Laws
 Compact Encoding of the Web Graph Exploiting Various Power Laws Statistical Reason Behind Link Database Yasuhito Asano, Tsuyoshi Ito 2, Hiroshi Imai 2, Masashi Toyoda 3, and Masaru Kitsuregawa 3 Department
Compact Encoding of the Web Graph Exploiting Various Power Laws Statistical Reason Behind Link Database Yasuhito Asano, Tsuyoshi Ito 2, Hiroshi Imai 2, Masashi Toyoda 3, and Masaru Kitsuregawa 3 Department
MergeSort, Recurrences, Asymptotic Analysis Scribe: Michael P. Kim Date: April 1, 2015
 CS161, Lecture 2 MergeSort, Recurrences, Asymptotic Analysis Scribe: Michael P. Kim Date: April 1, 2015 1 Introduction Today, we will introduce a fundamental algorithm design paradigm, Divide-And-Conquer,
CS161, Lecture 2 MergeSort, Recurrences, Asymptotic Analysis Scribe: Michael P. Kim Date: April 1, 2015 1 Introduction Today, we will introduce a fundamental algorithm design paradigm, Divide-And-Conquer,
Chapter 4 File Systems. Tanenbaum, Modern Operating Systems 3 e, (c) 2008 Prentice-Hall, Inc. All rights reserved
 Chapter 4 File Systems File Systems The best way to store information: Store all information in virtual memory address space Use ordinary memory read/write to access information Not feasible: no enough
Chapter 4 File Systems File Systems The best way to store information: Store all information in virtual memory address space Use ordinary memory read/write to access information Not feasible: no enough
Introduction to Machine Learning Prof. Anirban Santara Department of Computer Science and Engineering Indian Institute of Technology, Kharagpur
 Introduction to Machine Learning Prof. Anirban Santara Department of Computer Science and Engineering Indian Institute of Technology, Kharagpur Lecture 14 Python Exercise on knn and PCA Hello everyone,
Introduction to Machine Learning Prof. Anirban Santara Department of Computer Science and Engineering Indian Institute of Technology, Kharagpur Lecture 14 Python Exercise on knn and PCA Hello everyone,
Statistical Modeling of Huffman Tables Coding
 Statistical Modeling of Huffman Tables Coding S. Battiato 1, C. Bosco 1, A. Bruna 2, G. Di Blasi 1, and G.Gallo 1 1 D.M.I. University of Catania - Viale A. Doria 6, 95125, Catania, Italy {battiato, bosco,
Statistical Modeling of Huffman Tables Coding S. Battiato 1, C. Bosco 1, A. Bruna 2, G. Di Blasi 1, and G.Gallo 1 1 D.M.I. University of Catania - Viale A. Doria 6, 95125, Catania, Italy {battiato, bosco,
Tutorial: De Novo Assembly of Paired Data
 : De Novo Assembly of Paired Data September 20, 2013 CLC bio Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 Fax: +45 86 20 12 22 www.clcbio.com support@clcbio.com : De Novo Assembly
: De Novo Assembly of Paired Data September 20, 2013 CLC bio Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 Fax: +45 86 20 12 22 www.clcbio.com support@clcbio.com : De Novo Assembly
Data Preprocessing. Next Generation Sequencing analysis DTU Bioinformatics Next Generation Sequencing Analysis
 Data Preprocessing Next Generation Sequencing analysis DTU Bioinformatics Generalized NGS analysis Data size Application Assembly: Compare Raw Pre- specific: Question Alignment / samples / Answer? reads
Data Preprocessing Next Generation Sequencing analysis DTU Bioinformatics Generalized NGS analysis Data size Application Assembly: Compare Raw Pre- specific: Question Alignment / samples / Answer? reads
Column Stores vs. Row Stores How Different Are They Really?
 Column Stores vs. Row Stores How Different Are They Really? Daniel J. Abadi (Yale) Samuel R. Madden (MIT) Nabil Hachem (AvantGarde) Presented By : Kanika Nagpal OUTLINE Introduction Motivation Background
Column Stores vs. Row Stores How Different Are They Really? Daniel J. Abadi (Yale) Samuel R. Madden (MIT) Nabil Hachem (AvantGarde) Presented By : Kanika Nagpal OUTLINE Introduction Motivation Background
1 Motivation for Improving Matrix Multiplication
 CS170 Spring 2007 Lecture 7 Feb 6 1 Motivation for Improving Matrix Multiplication Now we will just consider the best way to implement the usual algorithm for matrix multiplication, the one that take 2n
CS170 Spring 2007 Lecture 7 Feb 6 1 Motivation for Improving Matrix Multiplication Now we will just consider the best way to implement the usual algorithm for matrix multiplication, the one that take 2n
THE RELATIVE EFFICIENCY OF DATA COMPRESSION BY LZW AND LZSS
 THE RELATIVE EFFICIENCY OF DATA COMPRESSION BY LZW AND LZSS Yair Wiseman 1* * 1 Computer Science Department, Bar-Ilan University, Ramat-Gan 52900, Israel Email: wiseman@cs.huji.ac.il, http://www.cs.biu.ac.il/~wiseman
THE RELATIVE EFFICIENCY OF DATA COMPRESSION BY LZW AND LZSS Yair Wiseman 1* * 1 Computer Science Department, Bar-Ilan University, Ramat-Gan 52900, Israel Email: wiseman@cs.huji.ac.il, http://www.cs.biu.ac.il/~wiseman
Data Preprocessing : Next Generation Sequencing analysis CBS - DTU Next Generation Sequencing Analysis
 Data Preprocessing 27626: Next Generation Sequencing analysis CBS - DTU Generalized NGS analysis Data size Application Assembly: Compare Raw Pre- specific: Question Alignment / samples / Answer? reads
Data Preprocessing 27626: Next Generation Sequencing analysis CBS - DTU Generalized NGS analysis Data size Application Assembly: Compare Raw Pre- specific: Question Alignment / samples / Answer? reads
CS 112 Introduction to Programming
 Running Time CS 112 Introduction to Programming As soon as an Analytic Engine exists, it will necessarily guide the future course of the science. Whenever any result is sought by its aid, the question
Running Time CS 112 Introduction to Programming As soon as an Analytic Engine exists, it will necessarily guide the future course of the science. Whenever any result is sought by its aid, the question
High Dimensional Indexing by Clustering
 Yufei Tao ITEE University of Queensland Recall that, our discussion so far has assumed that the dimensionality d is moderately high, such that it can be regarded as a constant. This means that d should
Yufei Tao ITEE University of Queensland Recall that, our discussion so far has assumed that the dimensionality d is moderately high, such that it can be regarded as a constant. This means that d should
Joint Entity Resolution
 Joint Entity Resolution Steven Euijong Whang, Hector Garcia-Molina Computer Science Department, Stanford University 353 Serra Mall, Stanford, CA 94305, USA {swhang, hector}@cs.stanford.edu No Institute
Joint Entity Resolution Steven Euijong Whang, Hector Garcia-Molina Computer Science Department, Stanford University 353 Serra Mall, Stanford, CA 94305, USA {swhang, hector}@cs.stanford.edu No Institute
Implementing a Statically Adaptive Software RAID System
 Implementing a Statically Adaptive Software RAID System Matt McCormick mattmcc@cs.wisc.edu Master s Project Report Computer Sciences Department University of Wisconsin Madison Abstract Current RAID systems
Implementing a Statically Adaptive Software RAID System Matt McCormick mattmcc@cs.wisc.edu Master s Project Report Computer Sciences Department University of Wisconsin Madison Abstract Current RAID systems
White Paper. How the Meltdown and Spectre bugs work and what you can do to prevent a performance plummet. Contents
 White Paper How the Meltdown and Spectre bugs work and what you can do to prevent a performance plummet Programs that do a lot of I/O are likely to be the worst hit by the patches designed to fix the Meltdown
White Paper How the Meltdown and Spectre bugs work and what you can do to prevent a performance plummet Programs that do a lot of I/O are likely to be the worst hit by the patches designed to fix the Meltdown
Big Data Analytics: What is Big Data? Stony Brook University CSE545, Fall 2016 the inaugural edition
 Big Data Analytics: What is Big Data? Stony Brook University CSE545, Fall 2016 the inaugural edition What s the BIG deal?! 2011 2011 2008 2010 2012 What s the BIG deal?! (Gartner Hype Cycle) What s the
Big Data Analytics: What is Big Data? Stony Brook University CSE545, Fall 2016 the inaugural edition What s the BIG deal?! 2011 2011 2008 2010 2012 What s the BIG deal?! (Gartner Hype Cycle) What s the
Two-dimensional Totalistic Code 52
 Two-dimensional Totalistic Code 52 Todd Rowland Senior Research Associate, Wolfram Research, Inc. 100 Trade Center Drive, Champaign, IL The totalistic two-dimensional cellular automaton code 52 is capable
Two-dimensional Totalistic Code 52 Todd Rowland Senior Research Associate, Wolfram Research, Inc. 100 Trade Center Drive, Champaign, IL The totalistic two-dimensional cellular automaton code 52 is capable
Core Membership Computation for Succinct Representations of Coalitional Games
 Core Membership Computation for Succinct Representations of Coalitional Games Xi Alice Gao May 11, 2009 Abstract In this paper, I compare and contrast two formal results on the computational complexity
Core Membership Computation for Succinct Representations of Coalitional Games Xi Alice Gao May 11, 2009 Abstract In this paper, I compare and contrast two formal results on the computational complexity
File system internals Tanenbaum, Chapter 4. COMP3231 Operating Systems
 File system internals Tanenbaum, Chapter 4 COMP3231 Operating Systems Architecture of the OS storage stack Application File system: Hides physical location of data on the disk Exposes: directory hierarchy,
File system internals Tanenbaum, Chapter 4 COMP3231 Operating Systems Architecture of the OS storage stack Application File system: Hides physical location of data on the disk Exposes: directory hierarchy,
1. Performance Comparison of Interdependent and Isolated Systems
 Supplementary Information for: Fu, G., Dawson, R., Khoury, M., & Bullock, S. (2014) Interdependent networks: Vulnerability analysis and strategies to limit cascading failure, European Physical Journal
Supplementary Information for: Fu, G., Dawson, R., Khoury, M., & Bullock, S. (2014) Interdependent networks: Vulnerability analysis and strategies to limit cascading failure, European Physical Journal
Parallel LZ77 Decoding with a GPU. Emmanuel Morfiadakis Supervisor: Dr Eric McCreath College of Engineering and Computer Science, ANU
 Parallel LZ77 Decoding with a GPU Emmanuel Morfiadakis Supervisor: Dr Eric McCreath College of Engineering and Computer Science, ANU Outline Background (What?) Problem definition and motivation (Why?)
Parallel LZ77 Decoding with a GPU Emmanuel Morfiadakis Supervisor: Dr Eric McCreath College of Engineering and Computer Science, ANU Outline Background (What?) Problem definition and motivation (Why?)
The levels of a memory hierarchy. Main. Memory. 500 By 1MB 4GB 500GB 0.25 ns 1ns 20ns 5ms
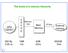 The levels of a memory hierarchy CPU registers C A C H E Memory bus Main Memory I/O bus External memory 500 By 1MB 4GB 500GB 0.25 ns 1ns 20ns 5ms 1 1 Some useful definitions When the CPU finds a requested
The levels of a memory hierarchy CPU registers C A C H E Memory bus Main Memory I/O bus External memory 500 By 1MB 4GB 500GB 0.25 ns 1ns 20ns 5ms 1 1 Some useful definitions When the CPU finds a requested
