A Multi-Relational Decision Tree Learning Algorithm - Implementation and Experiments
|
|
|
- Ophelia Leonard
- 6 years ago
- Views:
Transcription
1 A Multi-Relational Decision Tree Learning Algorithm - Implementation and Experiments Anna Atramentov, Hector Leiva and Vasant Honavar Artificial Intelligence Research Laboratory, Computer Science Department 226 Atanasoff Hall, Iowa State University Ames, IA , USA, {anjuta, aleiva, honavar}@cs.iastate.edu Abstract. We describe an efficient implementation (MRDTL-2) of the Multirelational decision tree learning (MRDTL) algorithm [23] which in turn was based on a proposal by Knobbe et al. [19] We describe some simple techniques for speeding up the calculation of sufficient statistics for decision trees and related hypothesis classes from multi-relational data. Because missing values are fairly common in many real-world applications of data mining, our implementation also includes some simple techniques for dealing with missing values. We describe results of experiments with several real-world data sets from the KDD Cup 2001 data mining competition and PKDD 2001 discovery challenge. Results of our experiments indicate that MRDTL is competitive with the state-of-the-art algorithms for learning classifiers from relational databases. 1 Introduction Recent advances in high throughput data acquisition, digital storage, and communications technologies have made it possible to gather very large amounts of data in many scientific and commercial domains. Much of this data resides in relational databases. Even when the data repository is not a relational database, it is often convenient to view heterogeneous data sources as if they were a collection of relations [28] for the purpose of extracting and organizing information from multiple sources. Thus, the task of learning from relational data has begun to receive significant attention in the literature [1, 19, 11, 21, 22, 12, 18, 26, 9, 8, 14, 16]. Knobbe et al. [19] outlined a general framework for multi-relational data mining which exploits structured query language (SQL) to gather the information needed for constructing classifiers (e.g., decision trees) from multi-relational data. Based on this framework, Leiva [23] developed a multi-relational decision tree learning algorithm (MRDTL). Experiments reported by Leiva [23] have shown that decision trees constructed using MRDTL have accuracies that are comparable to that obtained using other algorithms on several multi-relational data sets. However, MRDTL has two significant limitations from the standpoint of multi-relational data mining from large, real-world data sets: (a) Slow running time: MRDTL (like other algorithms based on the multi-relational data mining framework proposed by Knobbe et al. [19]) uses selection graphs to
2 query the relevant databases to obtain the statistics needed for constructing the classifier. Our experiments with MRDTL on data from KDD Cup 2001 [5] showed that the execution of queries encoded by such selection graphs is a major bottleneck in terms of the running time of the algorithm. (b) Inability to handle missing attribute values: In multi-relational databases encountered in many real-world applications of data mining, a significant fraction of the data have one or more missing attribute values. For example, in the case of gene localization task from KDD Cup 2001 [5], 70% of CLASS, 50% of COMPLEX and 50% of MOTIF attribute values are missing. Leiva s implementation of MRDTL [23] treats each missing value as a special value ( missing ) and does not include any statistically well-founded techniques for dealing with missing values. Consequently, the accuracy of decision trees constructed using MRDTL is far from satisfactory on classification tasks in which missing attribute values are commonplace. For example, the accuracy of MRDTL on the gene localization task was approximately 50%. Against this background, this paper describes MRDTL-2 which attempts to overcome these limitations of Leiva s implementation of MRDTL: (a) MRDTL-2 includes techniques for significantly speeding up some of the most time consuming components of multi-relational data mining algorithms like MRDTL that rely on the use of selection graphs. (b) MRDTL-2 includes a simple and computationally efficient technique which uses Naive Bayes classifiers for filling in missing attribute values. These enhancements enable us to apply multi-relational decision tree learning algorithms to significantly more complex classification tasks involving larger data sets and larger percentage of missing attribute values than was feasible in the case of MRDTL. Our experiments with several classification tasks drawn from KDD Cup 2001 [5], PKDD 2001 [7] and the widely studied Mutagenesis data set [25] show that MRDTL-2 (a) significantly outperforms MRDTL in terms of running time (b) yields results that are comparable to the best reported results obtained using multirelational data mining algorithms (c) compares favorably with feature-based learners that are based on clever propositionalization methods [22] The rest of the paper is organized as follows: Section 2 reviews the multi-relational data-mining framework; Section 3 describes our implementation of MRDTL-2, a multirelational decision tree learning algorithm; Section 4 describes the results of our experiments with MRDTL-2 on several representative multi-relational data mining tasks and compares them with the results of other approaches available in the literature; Section 5 concludes with a brief summary and discussion of the main results and an outline of some directions for further research.
3 2 Multi-Relational Data Mining 2.1 Relational Databases A relational database consists of a set of tables D = {X 1, X 2,...X n }, and a set of associations between pairs of tables. In each table a row represents description of one record. A column represents values of some attribute for the records in the table. An attribute A from table X is denoted by X.A. Definition 1. The domain of the attribute X.A is denoted as DOM(X.A) and is defined as the set of all different values that the records from table X have in the column of attribute A. Associations between tables are defined through primary and foreign key attributes. Definition 2. A primary key attribute of table X, denoted as X.ID, has a unique value for each row in this table. Definition 3. A foreign key attribute in table Y referencing table X, denoted as Y.X ID, takes values from DOM(X.ID). An example of a relational database is shown in Figure 1. There are three tables and three associations between tables. The primary keys of the tables, COM- POSITION, and are: ID, C ID, and I ID, respectively. Each COMPOSITION record references some record through the foreign key COM- POSITION. ID, and each record references two records through the foreign keys. ID1 and. ID2. In this setting the attribute of interest (e.g., class label) is called target attribute, and the _ID1 _ID2 TYPE EXPRESSION_CORR I_ID _ID ESSENTIAL CHROMOSOME LOCALIZATION COMPOSITION _ID CLASS COMPLEX PHENOTYPE MOTIF C_ID Fig. 1. Example database table in which this attribute is stored is called target table and is denoted by T 0. Each record in T 0 corresponds to a single object. Additional information about an object is stored in other tables of the database that can be looked up by following the associations between tables.
4 2.2 Multi-Relational Data Mining Framework Multi-relational data mining framework is based on the search for interesting patterns in the relational database, where multi-relational patterns can be viewed as pieces of substructure encountered in the structure of the objects of interest [19]. Definition 4. ([19]) A multi-relational object is covered by a multi-relational pattern iff the substructure described by the multi-relational pattern, in terms of both attributevalue conditions and structural conditions, occurs at least once in the multi-relational object. Multi-relational patterns also can be viewed as subsets of the objects from the database having some property. The most interesting subsets are chosen according to some measure (i.e. information gain for classification task), which guides the search in the space of all patterns. The search for interesting patterns usually proceeds by a top-down induction. For each interesting pattern, subpatterns are obtained with the help of refinement operator, which can be seen as further division of the set of objects covered by initial pattern. Top-down induction of interesting pattern proceeds recursively applying such refinement operators to the best patterns. Multi-relational pattern language is defined in terms of selection graphs and refinements which are described in the following sections. 2.3 Selection Graphs Multi-relational patterns are expressed in a graphical language of selection graphs [20]. Definition 5. ([19]) A selection graph S is a directed graph S = (N, E). N represents the set of nodes in S in the form of tuples (X, C, s, f), where X is a table from D, C is the set of conditions on attributes in X (for example, X.color = red or X.salary > 5,000), s is a flag with possible values open and closed, and f is a flag with possible values front and back. E represents edges in S in the form of tuples (p, q, a, e), where p and q are nodes and a is a relation between p and q in the data model (for example, X.ID = Y.X ID), and e is a flag with possible values present and absent. The selection graph should contain at least one node n 0 that corresponds to the target table T 0. An example of the selection graph for the data model from Figure 1 is shown in Figure 2. This selection graph corresponds to those (s) that belong to chromosome number 5, that have at least one record of type Genetic with a corresponding on chromosome number 11, but for which none of the INTERAC- TION records have type value Genetic and expression corr value 0.2. In this example the target table is, and within the target attribute is LOCALIZATION. In graphical representation of a selection graph, the value of s is represented by the absence or presence of a cross in the node, representing values open and closed, respectively. The value for e, in turn, is indicated by the presence (absent value) or absence (present value) of a cross on the corresponding arrow representing the edge. An edge between nodes p and q chooses the records in the database that match the join condition, a, between the tables which is defined by the relation between the primary key in p and a foreign key in q, or the other way around. For example, the join condition, a, between
5 .gene_id =.gene_id2 Type = Genetic.gene_id1 =.gene_id Chromosome=11.gene_id =.gene_id2 Chromosome=5 Type = Genetic and Expression_corr = 0.2 Fig. 2. Selection graph corresponding to those s which belong to chromosome number 5, that have at least one interaction of type Genetic with a corresponding gene on chromosome number 11 but for which none of the interaction records are of the type Genetic and expression corr value 0.2. table and in selection graph from Figure 2 is. ID =. ID2. A present edge between tables p and q combined with a list of conditions, q.c and p.c, selects those objects that match the list of conditions, q.c and p.c, and belong to the join between p and q, specified by join condition, e.a. On the other hand, an absent edge between tables p and q combined with a list of conditions, q.c and p.c, selects those objects that match condition p.c but do not satisfy the following: match q.c and belong to the join between tables at the same time. Flag f is set to front for those nodes that on their path to n 0 have no closed edges. For all the other nodes flag f is set to back. Knobbe et al. [20] introduce the algorithm (Figure 3) for translating a selection graph into SQL query. This algorithm returns the records in the target table covered by this selection graph. The subgraph(s, j.q) procedure returns the subgraph of the selection graph S starting with the node q as the target node, which label s is reset to open, removing the part of the graph that was connected to this node with the edge j and resetting all the values of flag f at the resulting selection graph by definition of f. Notation j.q.key means the name of the attribute (primary or foreign key) in the table q that is associated with the table p in relation j.a. Using this procedure the graph in Figure 2 translates to the SQL statement shown in Figure Refinements of Selection Graphs Multi-relational data mining algorithms search for and successively refine interesting patterns and select promising ones based on some impurity measure (e.g. information gain). The set of refinements introduced by [20] are given below. We will illustrate all the refinements on the selection graph from Figure 2. Labels s and f help identify the nodes that needs to be refined. Note that a refinement can only be applied to the open and front nodes in the selection graph S.
6 TRANSLATE(S, key) Input Selection graph S, key (primary or foreign) in the target node of the selection graph S Output SQL query for objects covered by selection graph S 1 table list := 2 condition list := 3 join list := 4 for each node i in S do 5 if (i.s = open and i.f = front ) 6 table list.add(i.table name + T + i) 7 for each condition c in i do 8 condition list.add(c) 9 for each edge j in S do 10 if (j.e = present ) 11 if (j.q.s = open and j.q.f = front ) 12 join list.add(j.a) 13 else join list.add(j.p +. + j.p.primary key + not in + TRANSLATE( subgraph(s, j.q), j.q.key) 15 return select distinct + T 0. + key + from + table list + where + join list + and + condition list Fig. 3. Translation of selection graph into SQL query select distinct T 0.gene id from T 0, T 1, T 2 where T 0.gene id = T 1.gene id2 and T 1.gene id2 = T 2.gene id and T 0.chromosome = 5 and T 1.type = Genetic and T 2.chromosome = 11 and T 0.gene id not in ( select T 0.gene id2 from T 0 where T 0.type = Genetic and T 0.expression corr = 0.2) Fig. 4. SQL query corresponding to the selection graph in Figure 2 (a) Add positive condition. This refinement simply adds a condition c to the set of conditions C in the node that is being refined in the selection graph S without actually changing the structure of S. For the selection graph from Figure 2 positive condition expression corr=0.5 applied to the node results in the selection graph shown on Figure 5 a. (b) Add negative condition (Figure 5 b). If the node which is refined is not n 0, the corresponding refinement will introduce a new absent edge from the parent of the selection node in question. The condition list of the selection node is copied to the new closed node and extended by the new condition. This node gets the copies of the children of the selection graph in question and open edges to those children are added. If the node which is refined represents the target table, the condition is simply negated and added to the current list of conditions for this node. This refinement is the complement of the add positive condition refinement, in the
7 Type = Genetic Expression_corr = 0.5 Chromosome=11 Type = Genetic Expression_corr = 0.5 Chromosome=11 Type = Genetic Chromosome=11 Chromosome=5 Chromosome=5 Type = Genetic and Expression_corr = 0.2 Type = Genetic and Expression_corr = 0.2 a) b) Fig. 5. Complement refinements for adding condition expression corr=0.5 to the node INTER- ACTION in the selection graph from Figure 2: a) positive condition, b) negative condition sense that it covers those objects from the original selection graph which were not covered by corresponding add positive condition refinement. (c) Add present edge and open node. This refinement introduces a present edge together with its corresponding table to the selection graph S. For the selection graph from Figure 2 adding edge from node to COMPOSITION node results in the selection graph shown in (Figure 6 a). COMPOSITION COMPOSITION Type = Genetic Chromosome=11 Type = Genetic Chromosome=11 Chromosome=5 Chromosome=5 Type = Genetic and Expression_corr = 0.2 Type = Genetic and Expression_corr = 0.2 a) b) Fig. 6. Complement refinements for adding edge from node to COMPOSITION node to selection graph from Figure 2: a) adding present edge and open node, b) adding absent edge and closed node (d) Add absent edge and closed node (Figure 6 b). This refinement introduces an absent edge together with its corresponding table to the selection graph S. This refinement is complement to the add present edge and open node, in the sense that it covers
8 those objects from the original selection graph which were not covered by add present edge and open node refinement. It is important to note that only through the add edge refinements the exploration of all the tables in the database is carried out. We can consider add condition refinement on some attribute from some table only after the edge to that table has been added to the selection graph. This raises the question as to what happens if the values of the attributes in some table are important for the task but the edge to this table can never be added, i.e. adding edge doesn t result in further split of the data covered by the refined selection graph. Look ahead refinements, which are a sequence of several refinements, are used for dealing with this situation. In the case when some refinement does not split the data covered by the selection graph, the next set of refinements is also considered as refinements of the original selection graph. 3 MDRTL-2: An Efficient Multi-Relational Decision Tree Learning Algorithm 3.1 Decision Tree Construction Multi-relational decision tree learning algorithm constructs a decision tree whose nodes are multi-relational patterns i.e., selection graphs. MRDTL-2 that we describe below is based on MRDTL proposed by Leiva [23] which in turn is based on the algorithm described by [20] and the logical decision tree induction algorithm called TILDE proposed by [1]. TILDE uses first order logic clauses to represent decisions (nodes) in the tree, when data are represented in first order logic rather than a collection of records in a relational database. MRDTL deals with records in relational databases, similarly to the TILDE s approach. Essentially, MRDTL adds selection graphs as the nodes to the tree through a process of successive refinement until some termination criterion is met (e.g., correct classification of instances in the training set). The choice of refinement to be added at each step is guided by a suitable impurity measure (e.g., information gain). MRDTL starts with the selection graph containing a single node at the root of the tree, which represents the set of all objects of interest in the relational database. This node corresponds to the target table T 0. The pseudocode for MRDTL is shown in Figure 7. The function optimal refinement(s) considers every possible refinement that can be made to the current selection graph S and selects the (locally) optimal refinement (i.e., one that maximizes information gain). Here, R(S) denotes the selection graph resulting from applying the refinement R to the selection graph S. R(S) denotes the application of the complement of R to the selection graph S. Our implementation of MRDTL considers the refinements described in the Section 2 as well as the look ahead refinements. The program automatically determines from the relational schema of the current database when look ahead might be needed. When adding an edge does not result in further split of the data, two-step refinements of the original selection graph are considered. Each candidate refinement is evaluated in terms of the split of the data induced by the refinement with respect to the target attribute, as in the case of the propositional version of the decision tree learning algorithm [30]. Splits based on numerical attributes
9 TREE INDUCTION(D, S) Input Database D, selection graph S Output The root of the tree, T 1 R := optimal refinement(s) 2 if stopping criteria(s) 3 return leaf 4 else 6 T left := TREE INDUCTION(D, R(S)) 8 T right := TREE INDUCTION(D, R(S)) 9 return node(t left, T right, R) Fig. 7. MRDTL algorithm are handled using a technique similar to that of C4.5 algorithm [30] with modifications proposed in [10, 29]. Our implementation of MRDTL uses SQL operations to obtain the counts needed for calculating information gain associated with the refinements. First we show the calculation of the information gain associated with add condition refinements. Let X be the table associated with one of the nodes in the current selection graph S and X.A be the attribute to be refined, and R vj (S) and R vj (S) be the add condition X.A = v j refinement and the complement of it respectively. The goal is to calculate entropies associated with the split based on these two refinements. This requires the following counts: count(c i, R vj (S)) and count(c i, R vj (S)), where count(c i, S) is the number of objects covered with selection graph S which have classification attribute T 0.target attribute = c i DOM(T 0.target attribute). The result of the SQL query shown in Figure 8 returns a list of the necessary counts: count(c i, R vj (S)) for each possible values c i DOM(T 0.target attribute) and v j DOM(X.A). select T 0.target attribute, X.A, count(distinct T 0.id) from TRANSLATE(S).get table list where TRANSLATE(S).get join list and TRANSLATE(S).get condition list Fig. 8. SQL query that returns counts of the type count(c i, R vj (S)) for each of the possible values c i DOM(T 0.target attribute) and v j DOM(X.A). The rest of the counts needed for the computation of the information gain can be obtained from the formula: count(c i, R vj (S)) = count(c i, S) count(c i, R vj (S)) Consider the SQL queries needed for calculating information gain associated with add edge refinements. Let X be the table associated with one of the nodes in the current selection graph S and e be the edge to be added from table X to table Y, and R e (S) and R e (S) be the add edge e refinement and its complement respectively. In order to
10 calculate the entropies associated with the split based on these refinements we need to gather the following counts: count(c i, R e (S)) and count(c i, R e (S)). The result of the SQL query shown in Figure 9 returns a list of the desired counts: count(c i, R e (S)) for each possible value c i DOM(T 0.target attribute). select T 0.target attribute, count(distinct T 0.id) from TRANSLATE(S).get table list, Y where TRANSLATE(S).get join list and TRANSLATE(S).get command list and e.a Fig. 9. SQL query returning counts of the type count(c i, R e(s)) for each possible value c i DOM(T 0.target attribute). The rest of the counts needed for the computation of the information gain can be obtained from the formula: count(c i, R e (S)) = count(c i, S) count(c i, R e (S)) The straightforward implementation of the algorithm based on the description given so far suffers from an efficiency problem which makes its application to complex realworld data sets infeasible in practice. As one gets further down in the decision tree the selection graph at the corresponding node grows. Thus, as more and more nodes are added to the decision tree the longer it takes to execute the corresponding SQL queries (Figures 8, 9) needed to examine the candidate refinements of the corresponding selection graph. Consequently, the straightforward implementation of MRDTL as described in [23] is too slow to be useful in practice. MRDTL-2 is a more efficient version of MRDTL. It exploits the fact that some of the results of computations that were carried out in the course of adding nodes at higher levels in the decision tree can be reused at lower levels in the course of refining the tree. Note that the queries from Figures 8 and 9 unnecessarily repeats work done earlier by retrieving instances covered by the selection graph whenever refining an existing selection graph. This query can be significantly simplified by storing the instances covered by the selection graph from previous iteration in a table to avoid retrieving them from the database. Thus, refining an existing selection graph reduces to finding a (locally) optimal split of the relevant set of instances. Now we proceed to show how the calculation of the SQL queries in Figures 8 and 9 is carried out in MRDTL-2. In each iteration of the algorithm, we store the primary keys from all open, front nodes of the selection graph for every object covered by it together with its classification value. This can be viewed as storing the skeletons of the objects covered by the selection graph, because it stores no other attribute information about records except for their primary keys. The SQL query for generating such a table for the selection graph S is shown in Figure 10. We call the resulting table of primary keys the sufficient table for S and denote it by I S. Given a sufficient table I S, we can obtain the counts needed for the calculation of the entropy for the add condition refinements as shown in Figure 11, and for the add edge refinements as shown in Figure 12.
11 SUF TABLE(S) Input Selection graph S Output SQL query for creating sufficient table I S 1 table list, condition list, join list := extract from(translate(s)) 2 primary key list := T 0.target attribute 3 for each node i in S do 4 if (i.s = open and i.f = front ) 5 primary key list.add(i.id) 6 return create table I S as ( select + primary key list + from + table list + where + join list + and + condition list + ) Fig. 10. Algorithm for generating SQL query corresponding to the sufficient table I S of the selection graph S select I S.T 0 target attribute, X.A, count(distinct I S.T 0 id) from I S, X where I S.X ID = X.ID Fig. 11. SQL query which returns the counts needed for calculating entropy for the splits based on add condition refinements to the node X for the attribute X.A using sufficient table I S It is easy to see that now the number of tables that need to be joined is not more than 3, whereas the number of tables needed to be joined in Figures 8 and 9 grows with the size of the selection graph. It is this growth that was responsible for the significant performance deterioration of MRDTL as nodes get added to the decision tree. It is important to note that it is inefficient to use the algorithm from Figure 10 in each iteration, since again the size of the query would increase with the growth of the selection graph. It is possible to create the sufficient table for the refined selection graph using only the information about refinement and sufficient table of the original selection graph as shown in Figure 13. The simple modifications described above make MRDTL-2 significantly faster to execute compared to MRDTL. This is confirmed by experimental results presented in section Handling missing values The current implementation of MRDTL-2 incorporates a simple approach to dealing with missing attribute values in the data. A Naive Bayes model for each attribute in a table is built based on the other attributes (excluding the class attribute). Missing attribute values are filled in with the most likely value predicted by the Naive Bayes predictor for the corresponding attribute. Thus, for each record, r, from table X we replace its missing value for the attribute X.A with the following value:
12 select I S.T 0 target attribute, count(distinct I S.T 0 id) from I S, X, Y where I S.X ID = X.ID and e.a Fig. 12. SQL query which returns counts needed for calculating entropy for the splits based on add edge e refinements from the node X to the node Y using sufficient table I S v NB = argmax v j DOM(X.A) P (v j ) X l D X l.a i,a i A, A i T 0.target attribute rn X l,rn associatedwithr P (X l.a i v j ) Here the first product is taken over the tables in the training database; The second product is taken over all the attributes in that table, except for the target attribute, T 0.target attribute, and the attribute which is being predicted, namely, X.A; The third product is taken over all the records in the table X l which are connected to the record r through the associations between the tables X and X l. In the case of one-to-many relation between the tables X and X l, one record from table X may have several corresponding records in the table X l. The value P (X l.a i v j ) is defined as the probability that some random element from table X has at least one corresponding record from table X l. Once the tables in the database are preprocessed in this manner, MRDTL-2 proceeds to build a decision tree from the resulting tables that contain no missing attribute values. In the future it would be interesting to investigate more sophisticated techniques for dealing with missing values. 3.3 Using the Decision Tree for Classification Before classifying an instance, any missing attribute values are filled in by preprocessing the tables using the method described above on the database of instances to be classified. The decision tree produced by MRDTL-2, as in the case of MRDTL, can be viewed as a set of SQL queries associated with the selection graphs that correspond to the leaves of the decision tree. Each selection graph (query) has a class label associated with it. If the corresponding node is not a pure node, (i.e., it does not unambiguously classify the training instances that match the query), the label associated with the node is based on the classification of the majority of training instances that match the corresponding selection graph in our implementation. (Alternatively, we could use probabilistic assignment of labels based on the distribution of class labels among the training instances that match the corresponding selection graph). The complementary nature of the different branches of a decision tree ensures that a given instance will not be assigned conflicting labels. It is also worth noting that it is not necessary to traverse the entire tree in order to classify a new instance; all the constraints on a certain path are stored in the selection graph associated with the corresponding leaf node.
13 REFINEMENT SUF TABLE(I S, R) Input Sufficient table I S for selection graph S, refinement R Output SQL query for sufficient table for R(S) 1 table list := I S 2 condition list := 3 join list := 4 primary key list := primary keys(i S) 5 if R == add positive condition, c, in table T i 6 table list += T i 7 condition list += T i.c 8 join list += T i.id+ = +I S.T i ID 9 else if R == add negative condition, c, in table T i 10 condition list += T 0.ID + is not in ( select distinct + I S.T 0 ID + from + I S, T i + where + T i.c + and + T i.id+ = +I S.T i ID+ ) 12 else if R = add present edge, e, from T i to T j 13 table list += T i+, +T j 14 join list += T i.id+ = +I S.T i ID+ and + e.a 15 primary key list += T j.id 16 else if R == add absent edge, e from T i to T j 17 condition list += T 0.ID + is not in ( select distinct + I S.T 0 ID + from + I S+, +T i+, +T j + where + T i.id+ = +I S.T i ID+ and + e.a+ ) 19 return create table I R as ( select + primary key list + from + table list + where + join list + and + condition list + ) Fig. 13. Algorithm for generating SQL query corresponding to sufficient table I R(S) 4 Experimental Results Our experiments focused on three data sets - the mutagenesis database which has been widely used in Inductive Logic Programming (ILP) research [25], the data for prediction of protein/gene localization and function from KDD Cup 2001 [17] and the data for predicting thrombosis from PKDD 2001 Discovery Challenge [27]. We compared the results we obtained using MRDTL-2 algorithm with those reported in the literature for the same datasets. 4.1 Mutagenesis Data Set The entity-relation diagram for the part of the Mutagenesis database [25] we used in our experiments is shown in Figure 14. The data set consists of 230 molecules divided into two subsets: 188 molecules for which linear regression yields good results and 42 molecules that are regression-unfriendly. This database contains descriptions of molecules and the characteristic to be predicted is their mutagenic activity (ability to cause DNA to mutate) represented by attribute label in molecule table.
14 MOLECULE ATOM ATOME_ID MOLECULE_ID ELEMNT TYPE CHARGE BOND MOLECULE_ID ATOM_ID1 ATOM_ID2 TYPE MOLECULE_ID LOG_MUT LOGP LUGMO IND1 INDA LABEL Fig. 14. Schema of the mutagenesis database accuracy time with MRDTL-2 time with MRDTL mutagenesis 87.5 % secs secs Table 1. Experimental results for mutagenesis data This dataset comes with different levels of background knowledge B 0, B 1, B 2, and B 3. In our experiments we chose to use the background knowledge B 2 and regression friendly subset of the dataset in order to compare the performance of MRDTL-2 with other methods for which experimental results are available in the literature. The results averaged with ten-fold crossvalidation are shown in the Table KDD Cup 2001 Data Set We also considered the two tasks from the KDD Cup 2001 data mining competition [5]: prediction of gene/protein function and localization. We normalized the data given in the task which resulted in the schema shown in Table 1. The resulting database consists of 3 tables,, and COM- POSITION, where table contains 862 entries in the training set and 381 in the testing set. Gene localization and function tasks present significant challenges because many attribute values in the data set are missing. We have conducted experiments both using technique for handling missing values, denoted mvh in the tables 2 and 3, and not using it (considering each missing values as a special value missing ). Table 2 summarizes the results we obtained for predicting.localization value on this set. In the case gene/protein function prediction, instances often have several class labels, since a protein may have several functions. MRDTL-2, like its propositional counterpart C4.5, assumes that each instance can be assigned to only one of several nonlocalization accuracy time with MRDTL-2 time with MRDTL accuracy with mvh % secs secs accuracy without mvh % secs secs Table 2. Experimental results for gene/protein localization prediction task
15 function accuracy time with MRDTL-2 time with MRDTL accuracy with mvh % secs secs accuracy without mvh % secs secs Table 3. Experimental results for gene/protein function prediction task accuracy time with MRDTL-2 time with MRDTL thrombosis 98.1 % secs secs Table 4. Experimental results for Thrombosis data set overlapping classes. To deal with multivalued class attributes, we transformed the problem into one of separately predicting membership in each possible class. i.e. for each possible function label we predicted whether the protein has this function or not. The overall accuracy was obtained from the formula: (true positive + true negative) (true positive + true negative + false positive + false negative) for all binary predictions. Table 3 summarizes the results we got for predicting the function of the protein. 4.3 PKDD 2001 Discovery Challenge Data Set The Thrombosis Data from the PKDD 2001 Discovery Challenge Data Set [7] consists of seven tables. PATIENT INFO contains 1239 records about patients. For our experiments we used 4 other tables (DIAGNOSIS, ANTIBODY EXAM, ANA PATTERN and THROMBOSIS) which all have a foreign key to the PATIENT INFO table. There are no other relations between the tables in this dataset. The task is to predict the degree of thrombosis attribute from ANTIBODY EXAM table. The results we obtained with 5:2 crossvalidation are shown in Table 4. The crossvalidation was done by partitioning the set of all records in the ANTIBODY EXAM table and their corresponding records from other tables into training and test sets. 4.4 Comparative evaluation The results of the comparison of MRDTL-2 performance with the best-known reported results for the datasets we described above are shown in the Table 5. 5 Summary and Discussion Advances in data acquisition, digital communication, and storage technologies have made it possible to gather and store large volumes data in digital form. A large fraction of this data resides in relational databases. Even when the data repository is not a relational database, it is possible to extract information from heterogeneous, autonomous,
16 dataset MRDTL accuracy best reported accuracy reference mutagenesis 87.5 % 86 % [31] localization % 72.1 % [5] function % 93.6 % [5] thrombosis 98.1 % % [7] Table 5. Comparison of MRDTL-2 performance with the best-known reported results distributed data sources using domain specific ontologies [28]. The result of such data integration is in the form of relational tables. Effective use of such data in data-driven scientific discovery and decision-making calls for sophisticated algorithms for knowledge acquisition from relational databases or multi-relational data mining algorithms is an important problem in multi-relational data mining which has begun to receive significant attention in the literature [1, 19, 11, 21, 22, 12, 18, 26, 9, 8, 14, 16, 2, 6, 13, 24]. Learning classifiers from relational databases has also been a focus of KDD Cup 2001 Competition and the PKDD 2001 Discovery Challenge. Against this background, this paper describes the design and implementation of MRDTL-2 - an algorithm for learning decision tree classifiers from relational databases, which is based on the framework for multi-relational data mining originally proposed by Knobbe et al. [19]. MRDTL- 2 extends an MRDTL, an algorithm for learning decision tree classifiers described by Leiva [23]. MRDTL-2 includes enhancements that overcome two significant limitations of MRDTL: (a) Slow running time: MRDTL-2 incorporates methods for speeding up MRDTL. Experiments using several data sets from KDD Cup 2001 and PKDD 2001 Discovery Challenge show that the proposed methods can significantly reduce the running time of the algorithm, thereby making it possible to apply multi-relational decision tree learning algorithms on far more complex data mining tasks. The proposed methods are potentially applicable to a broad class of multi-relational data mining algorithms based on the framework proposed by Knobbe et al. [19]. (b) Inability to handle missing attribute values: MRDTL-2 includes a simple and computationally efficient technique using Naive Bayes classifiers for filling in missing attribute values, which significantly enhances the applicability of multi-relational decision tree learning algorithms to the real-world classification tasks. Our experiments with several classification tasks drawn from KDD Cup 2001 [5] and PKDD 2001 Discovery Challenge [7] and the widely studied Mutagenesis data set show that MRDTL-2 (a) significantly outperforms MRDTL in terms of running time (b) yields results that are comparable to the best reported results obtained using multirelational data mining algorithms (Table 5) (c) compares favorably with feature-based learners that are based on clever propositionalization methods [22] Work in progress is aimed at:
17 (a) Incorporation of more sophisticated methods for handling missing attribute values into MRDTL-2 (b) Incorporation of sophisticated pruning methods or complexity regularization techniques into MRDTL-2 to minimize overfitting and improve generalization (c) Development of ontology-guided multi-relational decision tree learning algorithms to generate classifiers at multiple levels of abstraction [33] (d) Development of variants of MRDTL that can learn from heterogeneous, distributed, autonomous data [4, 3, 28] (e) More extensive experimental evaluation of MRDTL-2 on real-world data sets. (f) Incorporation of more sophisticated methods for evaluation of MRDTL-2 [15]. Acknowledgements This work is supported in part by an Information Technology Research (ITR) grant ( ) from the National Science Foundation, and a Biological Information Science and Technology Initiative (BISTI) award (GM066387) from the National Institutes of Health. The paper has benefited from discussions with Doina Caragea and Adrian Silvescu of the Iowa State University Artificial Intelligence Research Laboratory. References 1. Hendrik Blockeel: Top-down induction of first order logical decision trees. Department of Computer Science, Katholieke Universiteit Leuven (1998) 2. Blockeel, H. and De Raedt, L.: Relational Knowledge Discovery in Databases. In: Proceedings of the sixth internal workshop of Inductive Logic Programming, volume 1312 of Lecture Notes in Artificial Intelligence, , Springer-Verlag (1996) 3. Caragea, D., Silvescu, A., and Honavar, V. (ISDA 2003)Decision Tree Induction from Distributed, Heterogeneous, Autonomous Data Sources. In: Proceedings of the Conference on Intelligent Systems Design and Applications. In press. 4. Caragea, D. and Silvescu, A. and Honavar, V.: Invited Chapter. Toward a Theoretical Framework for Analysis and Synthesis of Agents That Learn from Distributed Dynamic Data SourcesTechnical Report. In: Emerging Neural Architectures Based on Neuroscience, Berlin: Springer-Verlag (2001) 5. Cheng, J. and Krogel, M. and Sese, J. and Hatzis C. and Morishita, S. and Hayashi, H. and Page, D.: KDD Cup 2001 Report. In: ACM Special Interest Group on Knowledge Discovery and Data Mining (SIGKDD) Explorations, vol. 3, issue 2 (2002) 6. Santos Costa et al.: Query Transformations for Improving the Efficiency of ILP Systems. In: Journal of Machine Learning Research (2002) 7. Coursac,I. - Duteil,N. - Lucas,N.: pkdd 2001 Discovery Challenge - Medical Domain. In: PKDD Discovery Challenge 2001, vol. 3, issue 2 (2002) 8. L. Dehaspe and L. De Raedt: Mining Association Rules in Multiple Relations. In: Proceedings of the 7th International Workshop on Inductive Logic Programming, vol. 1297, p (1997) 9. Dzeroski, S. and Lavrac, N: Relational Data Mining. Springer-Verlag (2001) 10. Fayyad, U. M. and Irani, K. B: On the handling of continuous-valued attributes in decision tree generation. In: Machine Learning, vol.8 (1992) 11. Friedman, N. and Getoor, L. and Koller, D. and and Pfeffer: Learning probabilistic relational models. In: Proceedings of the 6th International Joint Conference on Artificial Intelligence (1999)
18 12. Getoor, L.: Multi-relational data mining using probabilistic relational models: research summary. In: Proceedings of the First Workshop in Multi-relational Data Mining (2001) 13. Masaki Ito and Hayato Ohwada: Efficient Database Access for Implementing a Scalable ILP engine. In: Work-In-Progress Report of the Eleventh International Conference on Inductive Logic Programming (2001) 14. Jaeger, M: Relational Bayesian networks. In: Proceedings of the 13th Annual Conference on Uncertainty in Artificial Intelligence (UAI-1997) 15. Jensen, D. and J. Neville: Autocorrelation and Linkage Cause Bias in Evaluation of Relational Learners. In: Proceedings of the 12th International Conference on Inductive Logic Programming (ILP 2002) 16. Karalic and Bratko: First order regression. In: Machine Learning 26, vol The KDD Cup 2001 dataset: dpage/kddcup2001/ 18. Kersting, K. and De Raedt, L.: Bayesian Logic Programs. In: Proceedings of the Work-in- Progress Track at the 10th International Conference on Inductive Logic Programming (2000) 19. Knobbe, J. and Blockeel, H. and Siebes, A. and Van der Wallen D.: Multi-relational Data Mining. In: Proceedings of Benelearn (1999) 20. Knobbe, J. and Blockeel, H. and Siebes, A. and Van der Wallen D.: Multi-relational decision tree induction. In: Proceedings of the 3rdEuropean Conference on Principles and Practice of Knowledge Discovery in Databases, PKDD-99 (1999) 21. Koller, D: Probabilistic Relational Models. In: Proceedings of 9th International Workshop on Inductive Logic Programming (ILP-99) 22. Krogel, M. and Wrobel, S.: Transformation-Based Learning Using Multirelational Aggregation. In: Proceedings of the 11th International Conference on Inductive Logic Programming, vol (2001) 23. Hector Ariel Leiva: A multi-relational decision tree learning algorithm. M.S. thesis. Deparment of Computer Science. Iowa State University (2002) 24. Morik, K. and Brockhausen, P.: A multistrategy approach to relational discovery in databases. In: Machine Learning, 27(3), (1997) 25. The mutagenesis dataset: Pfeffer, A.: A Bayesian Language for Cumulative Learning. In: Proceedings of AAAI 2000 Workshop on Learning Statistical Models from Relational Data, AAAI Press (2000) 27. The PKDD 2001 Discovery Challenge dataset: Jaime Reinoso-Castillo: Ontology-driven information extraction and integration from Heterogeneous Distributed Autonomous Data Sources. M.S. Thesis. Department of Computer Science. Iowa State University (2002) 29. Quinlan, R.: Improved Use of Continuous Attributes in C4.5. In: Journal of Artificial Intelligence Research, vol.4 (1996) 30. Quinlan, R.: C4.5: Programs for Machine Learning. In: San Mateo: Morgan Kaufmann (1993) 31. Srinivasan, A., King, R. D., and Muggleton, S.: The role of background knowledge: using a problem from chemistry to examine the performance of an ILP program. Technical Report PRG-TR-08-99, Oxford University Computing Laboratory, Oxford, Wang, X., Schroeder, D., Dobbs, D., and Honavar, V. (2003). Data-Driven Discovery of Rules for Protein Function Classification Based on Sequence Motifs. Information Sciences. In press. 33. Zhang, J. and Honavar, V. (2003). Learning Decision Tree Classifiers from Attribute-Value Taxonomies and Partially Specified Data. In: Proceedings of the International Conference on Machine Learning. Washington, DC. In press.
Experiments with MRDTL -- A Multi-relational Decision Tree Learning Algorithm
 Experiments with MRDTL -- A Multi-relational Decision Tree Learning Algorithm Hector Leiva, Anna Atramentov and Vasant Honavar 1 Artificial Intelligence Laboratory Department of Computer Science and Graduate
Experiments with MRDTL -- A Multi-relational Decision Tree Learning Algorithm Hector Leiva, Anna Atramentov and Vasant Honavar 1 Artificial Intelligence Laboratory Department of Computer Science and Graduate
Multi-relational decision tree algorithm - implementation and experiments. Anna Atramentov. A thesis submitted to the graduate faculty
 Multi-relational decision tree algorithm - implementation and experiments by Anna Atramentov A thesis submitted to the graduate faculty in partial fulfillment of the requirements for the degree of MASTER
Multi-relational decision tree algorithm - implementation and experiments by Anna Atramentov A thesis submitted to the graduate faculty in partial fulfillment of the requirements for the degree of MASTER
Multi-relational Decision Tree Induction
 Multi-relational Decision Tree Induction Arno J. Knobbe 1,2, Arno Siebes 2, Daniël van der Wallen 1 1 Syllogic B.V., Hoefseweg 1, 3821 AE, Amersfoort, The Netherlands, {a.knobbe, d.van.der.wallen}@syllogic.com
Multi-relational Decision Tree Induction Arno J. Knobbe 1,2, Arno Siebes 2, Daniël van der Wallen 1 1 Syllogic B.V., Hoefseweg 1, 3821 AE, Amersfoort, The Netherlands, {a.knobbe, d.van.der.wallen}@syllogic.com
SRL2003 IJCAI 2003 Workshop on Learning Statistical Models from Relational Data
 University of Massachusetts Amherst ScholarWorks@UMass Amherst Computer Science Department Faculty Publication Series Computer Science 2003 SRL2003 IJCAI 2003 Workshop on Learning Statistical Models from
University of Massachusetts Amherst ScholarWorks@UMass Amherst Computer Science Department Faculty Publication Series Computer Science 2003 SRL2003 IJCAI 2003 Workshop on Learning Statistical Models from
Stochastic propositionalization of relational data using aggregates
 Stochastic propositionalization of relational data using aggregates Valentin Gjorgjioski and Sašo Dzeroski Jožef Stefan Institute Abstract. The fact that data is already stored in relational databases
Stochastic propositionalization of relational data using aggregates Valentin Gjorgjioski and Sašo Dzeroski Jožef Stefan Institute Abstract. The fact that data is already stored in relational databases
DRILA: A Distributed Relational Inductive Learning Algorithm
 DRILA: A Distributed Relational Inductive Learning Algorithm SALEH M. ABU-SOUD Computer Science Department New York Institute of Technology Amman Campus P.O. Box (1202), Amman, 11941 JORDAN sabusoud@nyit.edu
DRILA: A Distributed Relational Inductive Learning Algorithm SALEH M. ABU-SOUD Computer Science Department New York Institute of Technology Amman Campus P.O. Box (1202), Amman, 11941 JORDAN sabusoud@nyit.edu
Decision Tree Induction from Distributed Heterogeneous Autonomous Data Sources
 Decision Tree Induction from Distributed Heterogeneous Autonomous Data Sources Doina Caragea, Adrian Silvescu, and Vasant Honavar Artificial Intelligence Research Laboratory, Computer Science Department,
Decision Tree Induction from Distributed Heterogeneous Autonomous Data Sources Doina Caragea, Adrian Silvescu, and Vasant Honavar Artificial Intelligence Research Laboratory, Computer Science Department,
A Framework for Learning from Distributed Data Using Sufficient Statistics and its Application to Learning Decision Trees
 A Framework for Learning from Distributed Data Using Sufficient Statistics and its Application to Learning Decision Trees Doina Caragea, Adrian Silvescu and Vasant Honavar Artificial Intelligence Research
A Framework for Learning from Distributed Data Using Sufficient Statistics and its Application to Learning Decision Trees Doina Caragea, Adrian Silvescu and Vasant Honavar Artificial Intelligence Research
Efficient Multi-relational Classification by Tuple ID Propagation
 Efficient Multi-relational Classification by Tuple ID Propagation Xiaoxin Yin, Jiawei Han and Jiong Yang Department of Computer Science University of Illinois at Urbana-Champaign {xyin1, hanj, jioyang}@uiuc.edu
Efficient Multi-relational Classification by Tuple ID Propagation Xiaoxin Yin, Jiawei Han and Jiong Yang Department of Computer Science University of Illinois at Urbana-Champaign {xyin1, hanj, jioyang}@uiuc.edu
Mr-SBC: A Multi-relational Naïve Bayes Classifier
 Mr-SBC: A Multi-relational Naïve Bayes Classifier Michelangelo Ceci, Annalisa Appice, and Donato Malerba Dipartimento di Informatica, Università degli Studi via Orabona, 4-70126 Bari - Italy _GIGMETTMGIQEPIVFEa$HMYRMFEMX
Mr-SBC: A Multi-relational Naïve Bayes Classifier Michelangelo Ceci, Annalisa Appice, and Donato Malerba Dipartimento di Informatica, Università degli Studi via Orabona, 4-70126 Bari - Italy _GIGMETTMGIQEPIVFEa$HMYRMFEMX
MRDTL: A multi-relational decision tree learning algorithm. Héctor Ariel Leiva. A thesis submitted to the graduate faculty
 MRDTL: A multi-relational decision tree learning algorithm by Héctor Ariel Leiva A thesis submitted to the graduate faculty in partial fulfillment of the requirements for the degree of MASTER OF SCIENCE
MRDTL: A multi-relational decision tree learning algorithm by Héctor Ariel Leiva A thesis submitted to the graduate faculty in partial fulfillment of the requirements for the degree of MASTER OF SCIENCE
Learning Directed Probabilistic Logical Models using Ordering-search
 Learning Directed Probabilistic Logical Models using Ordering-search Daan Fierens, Jan Ramon, Maurice Bruynooghe, and Hendrik Blockeel K.U.Leuven, Dept. of Computer Science, Celestijnenlaan 200A, 3001
Learning Directed Probabilistic Logical Models using Ordering-search Daan Fierens, Jan Ramon, Maurice Bruynooghe, and Hendrik Blockeel K.U.Leuven, Dept. of Computer Science, Celestijnenlaan 200A, 3001
Aggregation and Selection in Relational Data Mining
 in Relational Data Mining Celine Vens Anneleen Van Assche Hendrik Blockeel Sašo Džeroski Department of Computer Science - K.U.Leuven Department of Knowledge Technologies - Jozef Stefan Institute, Slovenia
in Relational Data Mining Celine Vens Anneleen Van Assche Hendrik Blockeel Sašo Džeroski Department of Computer Science - K.U.Leuven Department of Knowledge Technologies - Jozef Stefan Institute, Slovenia
Relational Learning. Jan Struyf and Hendrik Blockeel
 Relational Learning Jan Struyf and Hendrik Blockeel Dept. of Computer Science, Katholieke Universiteit Leuven Celestijnenlaan 200A, 3001 Leuven, Belgium 1 Problem definition Relational learning refers
Relational Learning Jan Struyf and Hendrik Blockeel Dept. of Computer Science, Katholieke Universiteit Leuven Celestijnenlaan 200A, 3001 Leuven, Belgium 1 Problem definition Relational learning refers
ReMauve: A Relational Model Tree Learner
 ReMauve: A Relational Model Tree Learner Celine Vens, Jan Ramon, and Hendrik Blockeel Katholieke Universiteit Leuven - Department of Computer Science, Celestijnenlaan 200 A, 3001 Leuven, Belgium {celine.vens,
ReMauve: A Relational Model Tree Learner Celine Vens, Jan Ramon, and Hendrik Blockeel Katholieke Universiteit Leuven - Department of Computer Science, Celestijnenlaan 200 A, 3001 Leuven, Belgium {celine.vens,
Bias-free Hypothesis Evaluation in Multirelational Domains
 Bias-free Hypothesis Evaluation in Multirelational Domains Christine Körner Fraunhofer Institut AIS, Germany christine.koerner@ais.fraunhofer.de Stefan Wrobel Fraunhofer Institut AIS and Dept. of Computer
Bias-free Hypothesis Evaluation in Multirelational Domains Christine Körner Fraunhofer Institut AIS, Germany christine.koerner@ais.fraunhofer.de Stefan Wrobel Fraunhofer Institut AIS and Dept. of Computer
Enhancing Forecasting Performance of Naïve-Bayes Classifiers with Discretization Techniques
 24 Enhancing Forecasting Performance of Naïve-Bayes Classifiers with Discretization Techniques Enhancing Forecasting Performance of Naïve-Bayes Classifiers with Discretization Techniques Ruxandra PETRE
24 Enhancing Forecasting Performance of Naïve-Bayes Classifiers with Discretization Techniques Enhancing Forecasting Performance of Naïve-Bayes Classifiers with Discretization Techniques Ruxandra PETRE
Hybrid Feature Selection for Modeling Intrusion Detection Systems
 Hybrid Feature Selection for Modeling Intrusion Detection Systems Srilatha Chebrolu, Ajith Abraham and Johnson P Thomas Department of Computer Science, Oklahoma State University, USA ajith.abraham@ieee.org,
Hybrid Feature Selection for Modeling Intrusion Detection Systems Srilatha Chebrolu, Ajith Abraham and Johnson P Thomas Department of Computer Science, Oklahoma State University, USA ajith.abraham@ieee.org,
An Information-Theoretic Approach to the Prepruning of Classification Rules
 An Information-Theoretic Approach to the Prepruning of Classification Rules Max Bramer University of Portsmouth, Portsmouth, UK Abstract: Keywords: The automatic induction of classification rules from
An Information-Theoretic Approach to the Prepruning of Classification Rules Max Bramer University of Portsmouth, Portsmouth, UK Abstract: Keywords: The automatic induction of classification rules from
Centrum voor Wiskunde en Informatica. Information Systems (INS)
 Centrum voor Wiskunde en Informatica! "# $ % &('( %!*) +,-.(!/0**1243 56 78 % Information Systems (INS) INS-R9908 May 31, 1999 Report INS-R9908 ISSN 1386-3681 CWI P.O. Box 94079 1090 GB Amsterdam The Netherlands!
Centrum voor Wiskunde en Informatica! "# $ % &('( %!*) +,-.(!/0**1243 56 78 % Information Systems (INS) INS-R9908 May 31, 1999 Report INS-R9908 ISSN 1386-3681 CWI P.O. Box 94079 1090 GB Amsterdam The Netherlands!
An Efficient Approximation to Lookahead in Relational Learners
 An Efficient Approximation to Lookahead in Relational Learners Jan Struyf 1, Jesse Davis 2, and David Page 2 1 Katholieke Universiteit Leuven, Dept. of Computer Science Celestijnenlaan 200A, 3001 Leuven,
An Efficient Approximation to Lookahead in Relational Learners Jan Struyf 1, Jesse Davis 2, and David Page 2 1 Katholieke Universiteit Leuven, Dept. of Computer Science Celestijnenlaan 200A, 3001 Leuven,
Estimating Missing Attribute Values Using Dynamically-Ordered Attribute Trees
 Estimating Missing Attribute Values Using Dynamically-Ordered Attribute Trees Jing Wang Computer Science Department, The University of Iowa jing-wang-1@uiowa.edu W. Nick Street Management Sciences Department,
Estimating Missing Attribute Values Using Dynamically-Ordered Attribute Trees Jing Wang Computer Science Department, The University of Iowa jing-wang-1@uiowa.edu W. Nick Street Management Sciences Department,
Learning Relational Probability Trees Jennifer Neville David Jensen Lisa Friedland Michael Hay
 Learning Relational Probability Trees Jennifer Neville David Jensen Lisa Friedland Michael Hay Computer Science Department University of Massachusetts Amherst, MA 01003 USA [jneville jensen lfriedl mhay]@cs.umass.edu
Learning Relational Probability Trees Jennifer Neville David Jensen Lisa Friedland Michael Hay Computer Science Department University of Massachusetts Amherst, MA 01003 USA [jneville jensen lfriedl mhay]@cs.umass.edu
A Clustering Approach to Generalized Pattern Identification Based on Multi-instanced Objects with DARA
 A Clustering Approach to Generalized Pattern Identification Based on Multi-instanced Objects with DARA Rayner Alfred 1,2, Dimitar Kazakov 1 1 University of York, Computer Science Department, Heslington,
A Clustering Approach to Generalized Pattern Identification Based on Multi-instanced Objects with DARA Rayner Alfred 1,2, Dimitar Kazakov 1 1 University of York, Computer Science Department, Heslington,
7. Decision or classification trees
 7. Decision or classification trees Next we are going to consider a rather different approach from those presented so far to machine learning that use one of the most common and important data structure,
7. Decision or classification trees Next we are going to consider a rather different approach from those presented so far to machine learning that use one of the most common and important data structure,
Efficient SQL-Querying Method for Data Mining in Large Data Bases
 Efficient SQL-Querying Method for Data Mining in Large Data Bases Nguyen Hung Son Institute of Mathematics Warsaw University Banacha 2, 02095, Warsaw, Poland Abstract Data mining can be understood as a
Efficient SQL-Querying Method for Data Mining in Large Data Bases Nguyen Hung Son Institute of Mathematics Warsaw University Banacha 2, 02095, Warsaw, Poland Abstract Data mining can be understood as a
Multi-relational data mining in Microsoft SQL Server 2005
 Data Mining VII: Data, Text and Web Mining and their Business Applications 151 Multi-relational data mining in Microsoft SQL Server 005 C. L. Curotto 1 & N. F. F. Ebecken & H. Blockeel 1 PPGCC/UFPR, Universidade
Data Mining VII: Data, Text and Web Mining and their Business Applications 151 Multi-relational data mining in Microsoft SQL Server 005 C. L. Curotto 1 & N. F. F. Ebecken & H. Blockeel 1 PPGCC/UFPR, Universidade
Learning from Semantically Heterogeneous Data
 Learning from Semantically Heterogeneous Data Doina Caragea* Department of Computing and Information Sciences Kansas State University 234 Nichols Hall Manhattan, KS 66506 USA voice: +1 785-532-7908 fax:
Learning from Semantically Heterogeneous Data Doina Caragea* Department of Computing and Information Sciences Kansas State University 234 Nichols Hall Manhattan, KS 66506 USA voice: +1 785-532-7908 fax:
Fuzzy Partitioning with FID3.1
 Fuzzy Partitioning with FID3.1 Cezary Z. Janikow Dept. of Mathematics and Computer Science University of Missouri St. Louis St. Louis, Missouri 63121 janikow@umsl.edu Maciej Fajfer Institute of Computing
Fuzzy Partitioning with FID3.1 Cezary Z. Janikow Dept. of Mathematics and Computer Science University of Missouri St. Louis St. Louis, Missouri 63121 janikow@umsl.edu Maciej Fajfer Institute of Computing
Decision Trees Dr. G. Bharadwaja Kumar VIT Chennai
 Decision Trees Decision Tree Decision Trees (DTs) are a nonparametric supervised learning method used for classification and regression. The goal is to create a model that predicts the value of a target
Decision Trees Decision Tree Decision Trees (DTs) are a nonparametric supervised learning method used for classification and regression. The goal is to create a model that predicts the value of a target
A Parallel Evolutionary Algorithm for Discovery of Decision Rules
 A Parallel Evolutionary Algorithm for Discovery of Decision Rules Wojciech Kwedlo Faculty of Computer Science Technical University of Bia lystok Wiejska 45a, 15-351 Bia lystok, Poland wkwedlo@ii.pb.bialystok.pl
A Parallel Evolutionary Algorithm for Discovery of Decision Rules Wojciech Kwedlo Faculty of Computer Science Technical University of Bia lystok Wiejska 45a, 15-351 Bia lystok, Poland wkwedlo@ii.pb.bialystok.pl
Graph Classification in Heterogeneous
 Title: Graph Classification in Heterogeneous Networks Name: Xiangnan Kong 1, Philip S. Yu 1 Affil./Addr.: Department of Computer Science University of Illinois at Chicago Chicago, IL, USA E-mail: {xkong4,
Title: Graph Classification in Heterogeneous Networks Name: Xiangnan Kong 1, Philip S. Yu 1 Affil./Addr.: Department of Computer Science University of Illinois at Chicago Chicago, IL, USA E-mail: {xkong4,
Improving Quality of Products in Hard Drive Manufacturing by Decision Tree Technique
 Improving Quality of Products in Hard Drive Manufacturing by Decision Tree Technique Anotai Siltepavet 1, Sukree Sinthupinyo 2 and Prabhas Chongstitvatana 3 1 Computer Engineering, Chulalongkorn University,
Improving Quality of Products in Hard Drive Manufacturing by Decision Tree Technique Anotai Siltepavet 1, Sukree Sinthupinyo 2 and Prabhas Chongstitvatana 3 1 Computer Engineering, Chulalongkorn University,
Constraint Based Induction of Multi-Objective Regression Trees
 Constraint Based Induction of Multi-Objective Regression Trees Jan Struyf 1 and Sašo Džeroski 2 1 Katholieke Universiteit Leuven, Dept. of Computer Science Celestijnenlaan 200A, B-3001 Leuven, Belgium
Constraint Based Induction of Multi-Objective Regression Trees Jan Struyf 1 and Sašo Džeroski 2 1 Katholieke Universiteit Leuven, Dept. of Computer Science Celestijnenlaan 200A, B-3001 Leuven, Belgium
Learning Link-Based Naïve Bayes Classifiers from Ontology-Extended Distributed Data
 Learning Link-Based Naïve Bayes Classifiers from Ontology-Extended Distributed Data Cornelia Caragea 1, Doina Caragea 2, and Vasant Honavar 1 1 Computer Science Department, Iowa State University 2 Computer
Learning Link-Based Naïve Bayes Classifiers from Ontology-Extended Distributed Data Cornelia Caragea 1, Doina Caragea 2, and Vasant Honavar 1 1 Computer Science Department, Iowa State University 2 Computer
SSV Criterion Based Discretization for Naive Bayes Classifiers
 SSV Criterion Based Discretization for Naive Bayes Classifiers Krzysztof Grąbczewski kgrabcze@phys.uni.torun.pl Department of Informatics, Nicolaus Copernicus University, ul. Grudziądzka 5, 87-100 Toruń,
SSV Criterion Based Discretization for Naive Bayes Classifiers Krzysztof Grąbczewski kgrabcze@phys.uni.torun.pl Department of Informatics, Nicolaus Copernicus University, ul. Grudziądzka 5, 87-100 Toruń,
Classifying Relational Data with Neural Networks
 Classifying Relational Data with Neural Networks Werner Uwents and Hendrik Blockeel Katholieke Universiteit Leuven, Department of Computer Science, Celestijnenlaan 200A, B-3001 Leuven {werner.uwents, hendrik.blockeel}@cs.kuleuven.be
Classifying Relational Data with Neural Networks Werner Uwents and Hendrik Blockeel Katholieke Universiteit Leuven, Department of Computer Science, Celestijnenlaan 200A, B-3001 Leuven {werner.uwents, hendrik.blockeel}@cs.kuleuven.be
Logistic Model Tree With Modified AIC
 Logistic Model Tree With Modified AIC Mitesh J. Thakkar Neha J. Thakkar Dr. J.S.Shah Student of M.E.I.T. Asst.Prof.Computer Dept. Prof.&Head Computer Dept. S.S.Engineering College, Indus Engineering College
Logistic Model Tree With Modified AIC Mitesh J. Thakkar Neha J. Thakkar Dr. J.S.Shah Student of M.E.I.T. Asst.Prof.Computer Dept. Prof.&Head Computer Dept. S.S.Engineering College, Indus Engineering College
Performance Analysis of Data Mining Classification Techniques
 Performance Analysis of Data Mining Classification Techniques Tejas Mehta 1, Dr. Dhaval Kathiriya 2 Ph.D. Student, School of Computer Science, Dr. Babasaheb Ambedkar Open University, Gujarat, India 1 Principal
Performance Analysis of Data Mining Classification Techniques Tejas Mehta 1, Dr. Dhaval Kathiriya 2 Ph.D. Student, School of Computer Science, Dr. Babasaheb Ambedkar Open University, Gujarat, India 1 Principal
A MODULAR APPROACH TO RELATIONAL DATA MINING
 Association for Information Systems AIS Electronic Library (AISeL) AMCIS 2002 Proceedings Americas Conference on Information Systems (AMCIS) December 2002 A MODULAR APPROACH TO RELATIONAL DATA MINING Claudia
Association for Information Systems AIS Electronic Library (AISeL) AMCIS 2002 Proceedings Americas Conference on Information Systems (AMCIS) December 2002 A MODULAR APPROACH TO RELATIONAL DATA MINING Claudia
Improving Quality of Products in Hard Drive Manufacturing by Decision Tree Technique
 www.ijcsi.org 29 Improving Quality of Products in Hard Drive Manufacturing by Decision Tree Technique Anotai Siltepavet 1, Sukree Sinthupinyo 2 and Prabhas Chongstitvatana 3 1 Computer Engineering, Chulalongkorn
www.ijcsi.org 29 Improving Quality of Products in Hard Drive Manufacturing by Decision Tree Technique Anotai Siltepavet 1, Sukree Sinthupinyo 2 and Prabhas Chongstitvatana 3 1 Computer Engineering, Chulalongkorn
Multi-relational Data Mining, Using UML for ILP
 Multi-relational Data Mining, Using UML for ILP Arno J. Knobbe 1,2, Arno Siebes 2, Hendrik Blockeel 3, and Daniël Van Der Wallen 4 1 Perot Systems Nederland B.V., Hoefseweg 1, 3821 AE Amersfoort, The Netherlands,
Multi-relational Data Mining, Using UML for ILP Arno J. Knobbe 1,2, Arno Siebes 2, Hendrik Blockeel 3, and Daniël Van Der Wallen 4 1 Perot Systems Nederland B.V., Hoefseweg 1, 3821 AE Amersfoort, The Netherlands,
1 DATA MINING IN DATA WAREHOUSE
 Sborník vědeckých prací Vysoké školy báňské - Technické univerzity Ostrava číslo 2, rok 2005, ročník LI, řada strojní článek č. 1484 Abstract Tibor SZAPPANOS *, Iveta ZOLOTOVÁ *, Lenka LANDRYOVÁ ** DISTIRIBUTED
Sborník vědeckých prací Vysoké školy báňské - Technické univerzity Ostrava číslo 2, rok 2005, ročník LI, řada strojní článek č. 1484 Abstract Tibor SZAPPANOS *, Iveta ZOLOTOVÁ *, Lenka LANDRYOVÁ ** DISTIRIBUTED
Mining High Order Decision Rules
 Mining High Order Decision Rules Y.Y. Yao Department of Computer Science, University of Regina Regina, Saskatchewan, Canada S4S 0A2 e-mail: yyao@cs.uregina.ca Abstract. We introduce the notion of high
Mining High Order Decision Rules Y.Y. Yao Department of Computer Science, University of Regina Regina, Saskatchewan, Canada S4S 0A2 e-mail: yyao@cs.uregina.ca Abstract. We introduce the notion of high
BITS F464: MACHINE LEARNING
 BITS F464: MACHINE LEARNING Lecture-16: Decision Tree (contd.) + Random Forest Dr. Kamlesh Tiwari Assistant Professor Department of Computer Science and Information Systems Engineering, BITS Pilani, Rajasthan-333031
BITS F464: MACHINE LEARNING Lecture-16: Decision Tree (contd.) + Random Forest Dr. Kamlesh Tiwari Assistant Professor Department of Computer Science and Information Systems Engineering, BITS Pilani, Rajasthan-333031
Discretizing Continuous Attributes Using Information Theory
 Discretizing Continuous Attributes Using Information Theory Chang-Hwan Lee Department of Information and Communications, DongGuk University, Seoul, Korea 100-715 chlee@dgu.ac.kr Abstract. Many classification
Discretizing Continuous Attributes Using Information Theory Chang-Hwan Lee Department of Information and Communications, DongGuk University, Seoul, Korea 100-715 chlee@dgu.ac.kr Abstract. Many classification
Supporting Relational Knowledge Discovery: Lessons in Architecture and Algorithm Design
 J. eville and D. Jensen (2002). Supporting relational knowledge discovery: Lessons in architecture and algorithm design. Papers of the ICML 2002 Workshop on Data Mining Lessons Learned. Supporting Relational
J. eville and D. Jensen (2002). Supporting relational knowledge discovery: Lessons in architecture and algorithm design. Papers of the ICML 2002 Workshop on Data Mining Lessons Learned. Supporting Relational
Combining Gradient Boosting Machines with Collective Inference to Predict Continuous Values
 Combining Gradient Boosting Machines with Collective Inference to Predict Continuous Values Iman Alodah Computer Science Department Purdue University West Lafayette, Indiana 47906 Email: ialodah@purdue.edu
Combining Gradient Boosting Machines with Collective Inference to Predict Continuous Values Iman Alodah Computer Science Department Purdue University West Lafayette, Indiana 47906 Email: ialodah@purdue.edu
Discretization Numerical Data for Relational Data with One-to-Many Relations
 Journal of Computer Science 5 (7): 519-528, 2009 ISSN 1549-3636 2009 Science Publications Discretization Numerical Data for Relational Data with One-to-Many Relations Rayner Alfred School of Engineering
Journal of Computer Science 5 (7): 519-528, 2009 ISSN 1549-3636 2009 Science Publications Discretization Numerical Data for Relational Data with One-to-Many Relations Rayner Alfred School of Engineering
International Journal of Scientific Research & Engineering Trends Volume 4, Issue 6, Nov-Dec-2018, ISSN (Online): X
 Analysis about Classification Techniques on Categorical Data in Data Mining Assistant Professor P. Meena Department of Computer Science Adhiyaman Arts and Science College for Women Uthangarai, Krishnagiri,
Analysis about Classification Techniques on Categorical Data in Data Mining Assistant Professor P. Meena Department of Computer Science Adhiyaman Arts and Science College for Women Uthangarai, Krishnagiri,
Improving Tree-Based Classification Rules Using a Particle Swarm Optimization
 Improving Tree-Based Classification Rules Using a Particle Swarm Optimization Chi-Hyuck Jun *, Yun-Ju Cho, and Hyeseon Lee Department of Industrial and Management Engineering Pohang University of Science
Improving Tree-Based Classification Rules Using a Particle Swarm Optimization Chi-Hyuck Jun *, Yun-Ju Cho, and Hyeseon Lee Department of Industrial and Management Engineering Pohang University of Science
A Survey on Postive and Unlabelled Learning
 A Survey on Postive and Unlabelled Learning Gang Li Computer & Information Sciences University of Delaware ligang@udel.edu Abstract In this paper we survey the main algorithms used in positive and unlabeled
A Survey on Postive and Unlabelled Learning Gang Li Computer & Information Sciences University of Delaware ligang@udel.edu Abstract In this paper we survey the main algorithms used in positive and unlabeled
CloNI: clustering of JN -interval discretization
 CloNI: clustering of JN -interval discretization C. Ratanamahatana Department of Computer Science, University of California, Riverside, USA Abstract It is known that the naive Bayesian classifier typically
CloNI: clustering of JN -interval discretization C. Ratanamahatana Department of Computer Science, University of California, Riverside, USA Abstract It is known that the naive Bayesian classifier typically
Forward Feature Selection Using Residual Mutual Information
 Forward Feature Selection Using Residual Mutual Information Erik Schaffernicht, Christoph Möller, Klaus Debes and Horst-Michael Gross Ilmenau University of Technology - Neuroinformatics and Cognitive Robotics
Forward Feature Selection Using Residual Mutual Information Erik Schaffernicht, Christoph Möller, Klaus Debes and Horst-Michael Gross Ilmenau University of Technology - Neuroinformatics and Cognitive Robotics
Lars Schmidt-Thieme, Information Systems and Machine Learning Lab (ISMLL), University of Hildesheim, Germany
 Syllabus Fri. 27.10. (1) 0. Introduction A. Supervised Learning: Linear Models & Fundamentals Fri. 3.11. (2) A.1 Linear Regression Fri. 10.11. (3) A.2 Linear Classification Fri. 17.11. (4) A.3 Regularization
Syllabus Fri. 27.10. (1) 0. Introduction A. Supervised Learning: Linear Models & Fundamentals Fri. 3.11. (2) A.1 Linear Regression Fri. 10.11. (3) A.2 Linear Classification Fri. 17.11. (4) A.3 Regularization
Business Club. Decision Trees
 Business Club Decision Trees Business Club Analytics Team December 2017 Index 1. Motivation- A Case Study 2. The Trees a. What is a decision tree b. Representation 3. Regression v/s Classification 4. Building
Business Club Decision Trees Business Club Analytics Team December 2017 Index 1. Motivation- A Case Study 2. The Trees a. What is a decision tree b. Representation 3. Regression v/s Classification 4. Building
Probabilistic Abstraction Lattices: A Computationally Efficient Model for Conditional Probability Estimation
 Probabilistic Abstraction Lattices: A Computationally Efficient Model for Conditional Probability Estimation Daniel Lowd January 14, 2004 1 Introduction Probabilistic models have shown increasing popularity
Probabilistic Abstraction Lattices: A Computationally Efficient Model for Conditional Probability Estimation Daniel Lowd January 14, 2004 1 Introduction Probabilistic models have shown increasing popularity
Algorithms: Decision Trees
 Algorithms: Decision Trees A small dataset: Miles Per Gallon Suppose we want to predict MPG From the UCI repository A Decision Stump Recursion Step Records in which cylinders = 4 Records in which cylinders
Algorithms: Decision Trees A small dataset: Miles Per Gallon Suppose we want to predict MPG From the UCI repository A Decision Stump Recursion Step Records in which cylinders = 4 Records in which cylinders
CSE4334/5334 DATA MINING
 CSE4334/5334 DATA MINING Lecture 4: Classification (1) CSE4334/5334 Data Mining, Fall 2014 Department of Computer Science and Engineering, University of Texas at Arlington Chengkai Li (Slides courtesy
CSE4334/5334 DATA MINING Lecture 4: Classification (1) CSE4334/5334 Data Mining, Fall 2014 Department of Computer Science and Engineering, University of Texas at Arlington Chengkai Li (Slides courtesy
International Journal of Software and Web Sciences (IJSWS)
 International Association of Scientific Innovation and Research (IASIR) (An Association Unifying the Sciences, Engineering, and Applied Research) ISSN (Print): 2279-0063 ISSN (Online): 2279-0071 International
International Association of Scientific Innovation and Research (IASIR) (An Association Unifying the Sciences, Engineering, and Applied Research) ISSN (Print): 2279-0063 ISSN (Online): 2279-0071 International
Comparing Univariate and Multivariate Decision Trees *
 Comparing Univariate and Multivariate Decision Trees * Olcay Taner Yıldız, Ethem Alpaydın Department of Computer Engineering Boğaziçi University, 80815 İstanbul Turkey yildizol@cmpe.boun.edu.tr, alpaydin@boun.edu.tr
Comparing Univariate and Multivariate Decision Trees * Olcay Taner Yıldız, Ethem Alpaydın Department of Computer Engineering Boğaziçi University, 80815 İstanbul Turkey yildizol@cmpe.boun.edu.tr, alpaydin@boun.edu.tr
Join Bayes Nets: A New Type of Bayes net for Relational Data
 Join Bayes Nets: A New Type of Bayes net for Relational Data Oliver Schulte oschulte@cs.sfu.ca Hassan Khosravi hkhosrav@cs.sfu.ca Bahareh Bina bba18@cs.sfu.ca Flavia Moser fmoser@cs.sfu.ca Abstract Many
Join Bayes Nets: A New Type of Bayes net for Relational Data Oliver Schulte oschulte@cs.sfu.ca Hassan Khosravi hkhosrav@cs.sfu.ca Bahareh Bina bba18@cs.sfu.ca Flavia Moser fmoser@cs.sfu.ca Abstract Many
.. Cal Poly CSC 466: Knowledge Discovery from Data Alexander Dekhtyar.. for each element of the dataset we are given its class label.
 .. Cal Poly CSC 466: Knowledge Discovery from Data Alexander Dekhtyar.. Data Mining: Classification/Supervised Learning Definitions Data. Consider a set A = {A 1,...,A n } of attributes, and an additional
.. Cal Poly CSC 466: Knowledge Discovery from Data Alexander Dekhtyar.. Data Mining: Classification/Supervised Learning Definitions Data. Consider a set A = {A 1,...,A n } of attributes, and an additional
A Two-level Learning Method for Generalized Multi-instance Problems
 A wo-level Learning Method for Generalized Multi-instance Problems Nils Weidmann 1,2, Eibe Frank 2, and Bernhard Pfahringer 2 1 Department of Computer Science University of Freiburg Freiburg, Germany weidmann@informatik.uni-freiburg.de
A wo-level Learning Method for Generalized Multi-instance Problems Nils Weidmann 1,2, Eibe Frank 2, and Bernhard Pfahringer 2 1 Department of Computer Science University of Freiburg Freiburg, Germany weidmann@informatik.uni-freiburg.de
Data Mining. 3.2 Decision Tree Classifier. Fall Instructor: Dr. Masoud Yaghini. Chapter 5: Decision Tree Classifier
 Data Mining 3.2 Decision Tree Classifier Fall 2008 Instructor: Dr. Masoud Yaghini Outline Introduction Basic Algorithm for Decision Tree Induction Attribute Selection Measures Information Gain Gain Ratio
Data Mining 3.2 Decision Tree Classifier Fall 2008 Instructor: Dr. Masoud Yaghini Outline Introduction Basic Algorithm for Decision Tree Induction Attribute Selection Measures Information Gain Gain Ratio
AMOL MUKUND LONDHE, DR.CHELPA LINGAM
 International Journal of Advances in Applied Science and Engineering (IJAEAS) ISSN (P): 2348-1811; ISSN (E): 2348-182X Vol. 2, Issue 4, Dec 2015, 53-58 IIST COMPARATIVE ANALYSIS OF ANN WITH TRADITIONAL
International Journal of Advances in Applied Science and Engineering (IJAEAS) ISSN (P): 2348-1811; ISSN (E): 2348-182X Vol. 2, Issue 4, Dec 2015, 53-58 IIST COMPARATIVE ANALYSIS OF ANN WITH TRADITIONAL
Credit card Fraud Detection using Predictive Modeling: a Review
 February 207 IJIRT Volume 3 Issue 9 ISSN: 2396002 Credit card Fraud Detection using Predictive Modeling: a Review Varre.Perantalu, K. BhargavKiran 2 PG Scholar, CSE, Vishnu Institute of Technology, Bhimavaram,
February 207 IJIRT Volume 3 Issue 9 ISSN: 2396002 Credit card Fraud Detection using Predictive Modeling: a Review Varre.Perantalu, K. BhargavKiran 2 PG Scholar, CSE, Vishnu Institute of Technology, Bhimavaram,
The role of feature construction in inductive rule learning
 The role of feature construction in inductive rule learning Peter A. Flach 1 and Nada Lavrač 2 1 Department of Computer Science University of Bristol, United Kingdom 2 Department of Intelligent Systems
The role of feature construction in inductive rule learning Peter A. Flach 1 and Nada Lavrač 2 1 Department of Computer Science University of Bristol, United Kingdom 2 Department of Intelligent Systems
Study on Classifiers using Genetic Algorithm and Class based Rules Generation
 2012 International Conference on Software and Computer Applications (ICSCA 2012) IPCSIT vol. 41 (2012) (2012) IACSIT Press, Singapore Study on Classifiers using Genetic Algorithm and Class based Rules
2012 International Conference on Software and Computer Applications (ICSCA 2012) IPCSIT vol. 41 (2012) (2012) IACSIT Press, Singapore Study on Classifiers using Genetic Algorithm and Class based Rules
A Fast Decision Tree Learning Algorithm
 A Fast Decision Tree Learning Algorithm Jiang Su and Harry Zhang Faculty of Computer Science University of New Brunswick, NB, Canada, E3B 5A3 {jiang.su, hzhang}@unb.ca Abstract There is growing interest
A Fast Decision Tree Learning Algorithm Jiang Su and Harry Zhang Faculty of Computer Science University of New Brunswick, NB, Canada, E3B 5A3 {jiang.su, hzhang}@unb.ca Abstract There is growing interest
Network. Department of Statistics. University of California, Berkeley. January, Abstract
 Parallelizing CART Using a Workstation Network Phil Spector Leo Breiman Department of Statistics University of California, Berkeley January, 1995 Abstract The CART (Classication and Regression Trees) program,
Parallelizing CART Using a Workstation Network Phil Spector Leo Breiman Department of Statistics University of California, Berkeley January, 1995 Abstract The CART (Classication and Regression Trees) program,
Rule-Based Method for Entity Resolution Using Optimized Root Discovery (ORD)
 American-Eurasian Journal of Scientific Research 12 (5): 255-259, 2017 ISSN 1818-6785 IDOSI Publications, 2017 DOI: 10.5829/idosi.aejsr.2017.255.259 Rule-Based Method for Entity Resolution Using Optimized
American-Eurasian Journal of Scientific Research 12 (5): 255-259, 2017 ISSN 1818-6785 IDOSI Publications, 2017 DOI: 10.5829/idosi.aejsr.2017.255.259 Rule-Based Method for Entity Resolution Using Optimized
On Demand Phenotype Ranking through Subspace Clustering
 On Demand Phenotype Ranking through Subspace Clustering Xiang Zhang, Wei Wang Department of Computer Science University of North Carolina at Chapel Hill Chapel Hill, NC 27599, USA {xiang, weiwang}@cs.unc.edu
On Demand Phenotype Ranking through Subspace Clustering Xiang Zhang, Wei Wang Department of Computer Science University of North Carolina at Chapel Hill Chapel Hill, NC 27599, USA {xiang, weiwang}@cs.unc.edu
Extra readings beyond the lecture slides are important:
 1 Notes To preview next lecture: Check the lecture notes, if slides are not available: http://web.cse.ohio-state.edu/~sun.397/courses/au2017/cse5243-new.html Check UIUC course on the same topic. All their
1 Notes To preview next lecture: Check the lecture notes, if slides are not available: http://web.cse.ohio-state.edu/~sun.397/courses/au2017/cse5243-new.html Check UIUC course on the same topic. All their
Machine Learning in Real World: C4.5
 Machine Learning in Real World: C4.5 Industrial-strength algorithms For an algorithm to be useful in a wide range of realworld applications it must: Permit numeric attributes with adaptive discretization
Machine Learning in Real World: C4.5 Industrial-strength algorithms For an algorithm to be useful in a wide range of realworld applications it must: Permit numeric attributes with adaptive discretization
Generation of Near-Optimal Solutions Using ILP-Guided Sampling
 Generation of Near-Optimal Solutions Using ILP-Guided Sampling Ashwin Srinivasan 1, Gautam Shroff 2, Lovekesh Vig 2, and Sarmimala Saikia 2 1 Department of Computer Science & Information Systems BITS Pilani,
Generation of Near-Optimal Solutions Using ILP-Guided Sampling Ashwin Srinivasan 1, Gautam Shroff 2, Lovekesh Vig 2, and Sarmimala Saikia 2 1 Department of Computer Science & Information Systems BITS Pilani,
The Role of Biomedical Dataset in Classification
 The Role of Biomedical Dataset in Classification Ajay Kumar Tanwani and Muddassar Farooq Next Generation Intelligent Networks Research Center (nexgin RC) National University of Computer & Emerging Sciences
The Role of Biomedical Dataset in Classification Ajay Kumar Tanwani and Muddassar Farooq Next Generation Intelligent Networks Research Center (nexgin RC) National University of Computer & Emerging Sciences
has to choose. Important questions are: which relations should be dened intensionally,
 Automated Design of Deductive Databases (Extended abstract) Hendrik Blockeel and Luc De Raedt Department of Computer Science, Katholieke Universiteit Leuven Celestijnenlaan 200A B-3001 Heverlee, Belgium
Automated Design of Deductive Databases (Extended abstract) Hendrik Blockeel and Luc De Raedt Department of Computer Science, Katholieke Universiteit Leuven Celestijnenlaan 200A B-3001 Heverlee, Belgium
CONCEPT FORMATION AND DECISION TREE INDUCTION USING THE GENETIC PROGRAMMING PARADIGM
 1 CONCEPT FORMATION AND DECISION TREE INDUCTION USING THE GENETIC PROGRAMMING PARADIGM John R. Koza Computer Science Department Stanford University Stanford, California 94305 USA E-MAIL: Koza@Sunburn.Stanford.Edu
1 CONCEPT FORMATION AND DECISION TREE INDUCTION USING THE GENETIC PROGRAMMING PARADIGM John R. Koza Computer Science Department Stanford University Stanford, California 94305 USA E-MAIL: Koza@Sunburn.Stanford.Edu
Part I. Instructor: Wei Ding
 Classification Part I Instructor: Wei Ding Tan,Steinbach, Kumar Introduction to Data Mining 4/18/2004 1 Classification: Definition Given a collection of records (training set ) Each record contains a set
Classification Part I Instructor: Wei Ding Tan,Steinbach, Kumar Introduction to Data Mining 4/18/2004 1 Classification: Definition Given a collection of records (training set ) Each record contains a set
ISSN: (Online) Volume 3, Issue 9, September 2015 International Journal of Advance Research in Computer Science and Management Studies
 ISSN: 2321-7782 (Online) Volume 3, Issue 9, September 2015 International Journal of Advance Research in Computer Science and Management Studies Research Article / Survey Paper / Case Study Available online
ISSN: 2321-7782 (Online) Volume 3, Issue 9, September 2015 International Journal of Advance Research in Computer Science and Management Studies Research Article / Survey Paper / Case Study Available online
Perceptron-Based Oblique Tree (P-BOT)
 Perceptron-Based Oblique Tree (P-BOT) Ben Axelrod Stephen Campos John Envarli G.I.T. G.I.T. G.I.T. baxelrod@cc.gatech sjcampos@cc.gatech envarli@cc.gatech Abstract Decision trees are simple and fast data
Perceptron-Based Oblique Tree (P-BOT) Ben Axelrod Stephen Campos John Envarli G.I.T. G.I.T. G.I.T. baxelrod@cc.gatech sjcampos@cc.gatech envarli@cc.gatech Abstract Decision trees are simple and fast data
Comparison of Online Record Linkage Techniques
 International Research Journal of Engineering and Technology (IRJET) e-issn: 2395-0056 Volume: 02 Issue: 09 Dec-2015 p-issn: 2395-0072 www.irjet.net Comparison of Online Record Linkage Techniques Ms. SRUTHI.
International Research Journal of Engineering and Technology (IRJET) e-issn: 2395-0056 Volume: 02 Issue: 09 Dec-2015 p-issn: 2395-0072 www.irjet.net Comparison of Online Record Linkage Techniques Ms. SRUTHI.
KDD, SEMMA AND CRISP-DM: A PARALLEL OVERVIEW. Ana Azevedo and M.F. Santos
 KDD, SEMMA AND CRISP-DM: A PARALLEL OVERVIEW Ana Azevedo and M.F. Santos ABSTRACT In the last years there has been a huge growth and consolidation of the Data Mining field. Some efforts are being done
KDD, SEMMA AND CRISP-DM: A PARALLEL OVERVIEW Ana Azevedo and M.F. Santos ABSTRACT In the last years there has been a huge growth and consolidation of the Data Mining field. Some efforts are being done
A New Approach To Graph Based Object Classification On Images
 A New Approach To Graph Based Object Classification On Images Sandhya S Krishnan,Kavitha V K P.G Scholar, Dept of CSE, BMCE, Kollam, Kerala, India Sandhya4parvathy@gmail.com Abstract: The main idea of
A New Approach To Graph Based Object Classification On Images Sandhya S Krishnan,Kavitha V K P.G Scholar, Dept of CSE, BMCE, Kollam, Kerala, India Sandhya4parvathy@gmail.com Abstract: The main idea of
Classification: Basic Concepts, Decision Trees, and Model Evaluation
 Classification: Basic Concepts, Decision Trees, and Model Evaluation Data Warehousing and Mining Lecture 4 by Hossen Asiful Mustafa Classification: Definition Given a collection of records (training set
Classification: Basic Concepts, Decision Trees, and Model Evaluation Data Warehousing and Mining Lecture 4 by Hossen Asiful Mustafa Classification: Definition Given a collection of records (training set
Ordering attributes for missing values prediction and data classification
 Ordering attributes for missing values prediction and data classification E. R. Hruschka Jr., N. F. F. Ebecken COPPE /Federal University of Rio de Janeiro, Brazil. Abstract This work shows the application
Ordering attributes for missing values prediction and data classification E. R. Hruschka Jr., N. F. F. Ebecken COPPE /Federal University of Rio de Janeiro, Brazil. Abstract This work shows the application
What is Learning? CS 343: Artificial Intelligence Machine Learning. Raymond J. Mooney. Problem Solving / Planning / Control.
 What is Learning? CS 343: Artificial Intelligence Machine Learning Herbert Simon: Learning is any process by which a system improves performance from experience. What is the task? Classification Problem
What is Learning? CS 343: Artificial Intelligence Machine Learning Herbert Simon: Learning is any process by which a system improves performance from experience. What is the task? Classification Problem
Rank Measures for Ordering
 Rank Measures for Ordering Jin Huang and Charles X. Ling Department of Computer Science The University of Western Ontario London, Ontario, Canada N6A 5B7 email: fjhuang33, clingg@csd.uwo.ca Abstract. Many
Rank Measures for Ordering Jin Huang and Charles X. Ling Department of Computer Science The University of Western Ontario London, Ontario, Canada N6A 5B7 email: fjhuang33, clingg@csd.uwo.ca Abstract. Many
Preprocessing of Stream Data using Attribute Selection based on Survival of the Fittest
 Preprocessing of Stream Data using Attribute Selection based on Survival of the Fittest Bhakti V. Gavali 1, Prof. Vivekanand Reddy 2 1 Department of Computer Science and Engineering, Visvesvaraya Technological
Preprocessing of Stream Data using Attribute Selection based on Survival of the Fittest Bhakti V. Gavali 1, Prof. Vivekanand Reddy 2 1 Department of Computer Science and Engineering, Visvesvaraya Technological
Dynamic Load Balancing of Unstructured Computations in Decision Tree Classifiers
 Dynamic Load Balancing of Unstructured Computations in Decision Tree Classifiers A. Srivastava E. Han V. Kumar V. Singh Information Technology Lab Dept. of Computer Science Information Technology Lab Hitachi
Dynamic Load Balancing of Unstructured Computations in Decision Tree Classifiers A. Srivastava E. Han V. Kumar V. Singh Information Technology Lab Dept. of Computer Science Information Technology Lab Hitachi
Development of Prediction Model for Linked Data based on the Decision Tree for Track A, Task A1
 Development of Prediction Model for Linked Data based on the Decision Tree for Track A, Task A1 Dongkyu Jeon and Wooju Kim Dept. of Information and Industrial Engineering, Yonsei University, Seoul, Korea
Development of Prediction Model for Linked Data based on the Decision Tree for Track A, Task A1 Dongkyu Jeon and Wooju Kim Dept. of Information and Industrial Engineering, Yonsei University, Seoul, Korea
Using Association Rules for Better Treatment of Missing Values
 Using Association Rules for Better Treatment of Missing Values SHARIQ BASHIR, SAAD RAZZAQ, UMER MAQBOOL, SONYA TAHIR, A. RAUF BAIG Department of Computer Science (Machine Intelligence Group) National University
Using Association Rules for Better Treatment of Missing Values SHARIQ BASHIR, SAAD RAZZAQ, UMER MAQBOOL, SONYA TAHIR, A. RAUF BAIG Department of Computer Science (Machine Intelligence Group) National University
Data Mining. Introduction. Hamid Beigy. Sharif University of Technology. Fall 1395
 Data Mining Introduction Hamid Beigy Sharif University of Technology Fall 1395 Hamid Beigy (Sharif University of Technology) Data Mining Fall 1395 1 / 21 Table of contents 1 Introduction 2 Data mining
Data Mining Introduction Hamid Beigy Sharif University of Technology Fall 1395 Hamid Beigy (Sharif University of Technology) Data Mining Fall 1395 1 / 21 Table of contents 1 Introduction 2 Data mining
RECORD DEDUPLICATION USING GENETIC PROGRAMMING APPROACH
 Int. J. Engg. Res. & Sci. & Tech. 2013 V Karthika et al., 2013 Research Paper ISSN 2319-5991 www.ijerst.com Vol. 2, No. 2, May 2013 2013 IJERST. All Rights Reserved RECORD DEDUPLICATION USING GENETIC PROGRAMMING
Int. J. Engg. Res. & Sci. & Tech. 2013 V Karthika et al., 2013 Research Paper ISSN 2319-5991 www.ijerst.com Vol. 2, No. 2, May 2013 2013 IJERST. All Rights Reserved RECORD DEDUPLICATION USING GENETIC PROGRAMMING
CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS
 CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS CHAPTER 4 CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS 4.1 Introduction Optical character recognition is one of
CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS CHAPTER 4 CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS 4.1 Introduction Optical character recognition is one of
Structural Logistic Regression for Link Analysis
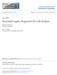 University of Pennsylvania ScholarlyCommons Departmental Papers (CIS) Department of Computer & Information Science August 2003 Structural Logistic Regression for Link Analysis Alexandrin Popescul University
University of Pennsylvania ScholarlyCommons Departmental Papers (CIS) Department of Computer & Information Science August 2003 Structural Logistic Regression for Link Analysis Alexandrin Popescul University
First order random forests: Learning relational classifiers with complex aggregates
 Mach Learn (2006) 64:149 182 DOI 10.1007/s10994-006-8713-9 First order random forests: Learning relational classifiers with complex aggregates Anneleen Van Assche Celine Vens Hendrik Blockeel Sašo Džeroski
Mach Learn (2006) 64:149 182 DOI 10.1007/s10994-006-8713-9 First order random forests: Learning relational classifiers with complex aggregates Anneleen Van Assche Celine Vens Hendrik Blockeel Sašo Džeroski
Evolving SQL Queries for Data Mining
 Evolving SQL Queries for Data Mining Majid Salim and Xin Yao School of Computer Science, The University of Birmingham Edgbaston, Birmingham B15 2TT, UK {msc30mms,x.yao}@cs.bham.ac.uk Abstract. This paper
Evolving SQL Queries for Data Mining Majid Salim and Xin Yao School of Computer Science, The University of Birmingham Edgbaston, Birmingham B15 2TT, UK {msc30mms,x.yao}@cs.bham.ac.uk Abstract. This paper
Building a Concept Hierarchy from a Distance Matrix
 Building a Concept Hierarchy from a Distance Matrix Huang-Cheng Kuo 1 and Jen-Peng Huang 2 1 Department of Computer Science and Information Engineering National Chiayi University, Taiwan 600 hckuo@mail.ncyu.edu.tw
Building a Concept Hierarchy from a Distance Matrix Huang-Cheng Kuo 1 and Jen-Peng Huang 2 1 Department of Computer Science and Information Engineering National Chiayi University, Taiwan 600 hckuo@mail.ncyu.edu.tw
