Lab 07: Maximum Likelihood Model Selection and RAxML Using CIPRES
|
|
|
- Samuel Webb
- 6 years ago
- Views:
Transcription
1 Integrative Biology 200, Spring 2014 Principles of Phylogenetics: Systematics University of California, Berkeley Updated by Traci L. Grzymala Lab 07: Maximum Likelihood Model Selection and RAxML Using CIPRES Exercise 1: Maximum Likelihood (ML) Model Selection Maximum likelihood (ML) is a statistical method for reconstructing trees. Basically, ML operates by calculating the following conditional equation: What is the likelihood of observing a data set given a phylogeny and a model of DNA sequence evolution? The tree with the highest likelihood score is considered the best tree. When using maximum likelihood to build trees, we have to first select a model of DNA sequence evolution. Let s do this first and then learn a bit more about ML and model selection while we wait for it to run. jmodeltest 1. Open jmodeltest by clicking jmodeltest.jar. The main window and menu should now be open. 2. Select File à Load DNA Alignment and then select your Nexus file (Use your own data or use the primate-mtdna.nex file). Okay, jmodeltest should have read the file and tells you how many sequences (basically how many OTUs) and how many sites (basically how many nucleotides) the file has. 3. Now select Analysis à Compute likelihood scores 4. A new window should now pop up with several options to choose from. Feel free to examine these different parameters on your own. For now set the default settings and make sure under Base tree for likelihood calculations that ML Optimized is selected. Select Compute Likelihoods. 5. Okay, let that run. How fast it takes will depend on your computer and your data, but it will be at least a few minutes. The program is computing likelihood scores for 88 different nucleotide substitution models. Let s learn a bit more about these different models while we twiddle our thumbs and wait. Models of Nucleotide Change The Transition Matrix The transition matrix (not as in transition/transversion) is a matrix showing the instantaneous stochastic rate of change between any two nucleotides. It can be used to calculate the chance of one nucleotide changing into another on a branch with a given length. 1
2 The most unrestrained matrix would look like this: A C G T A α β γ α β γ C δ δ ε ζ ε ζ G η θ η θ ι ι T κ λ µ κ λ µ As you can see, the diagonals are all negative as each nucleotide will be changing away from itself at any instant, so that each row adds up to 0. Furthermore, the average rate of change of all the off diagonals is normalized to 1, so that you can eliminate another parameter for a total of 11 parameters. On the other hand the Kimura two parameter model would look like this: A C G T A α 2β β α β C β α 2β β α G α β α 2β β T β α β α 2β Here there are two parameters, transition and transversion rate, which can be reduced to just one by normalizing the matrix. Most programs (PAUP* included) can only calculate matrices with reversible models. This means that change has an equal probability of happening in either direction on a branch. Thus trees can be evaluated as unrooted networks, making the computationally-intensive likelihood calculations much easier. For a model to be reversible it must be true that: π X R X>Y =π Y R Y>X where R X>Y is the instantaneous rate of change from nucleotide X to nucleotide Y, and π X is the equilibrium frequency of nucleotide X. The equilibrium frequency is the frequency of that nucleotide if the substitution process is allowed to run forever, and can be considered another parameter. Thus any model in which R X>Y =π Y r XY, will be reversible. So the General Time Reversible (GTR) matrix looks like: A C G T A π C r AC π G r AG π T r AT C π A r AC π G r CG π T r CT G π A r AG π C r CG π T r GT T π A r AT π C r CT π G r GT with the diagonal filled in appropriately. The sum of the equilibrium frequencies for all four bases must equal one, so that there are three equilibrium frequency parameters. Furthermore, 2
3 one of the rate parameters can be eliminated by normalizing the matrix, leaving eight parameters total. General Time Reversible (GTR) represents a family of nested models that encompass 64 models with different combinations of parameters. Nested models are special cases of more general models. We briefly learned about some of these nested models already: JC : Jukes and Cantor (1969) - All nucleotide substitutions are equal and all base frequencies are equal. This is the most restricted (=specific) model of substitution because it assumes all changes are equal. F81 : Felsenstein (1981) - All nucleotide substitutions are equal, base frequencies allowed to vary. K2P : Kimura two-parameter model, Kimura (1980) - Two nucleotide substitutions types are allowed, those between transitions and transversions. Base frequencies are assumed equal. HKY85: Hasegawa-Kishino-Yano (1985) - Two nucleotide substitutions types are allowed, those between transitions and transversions. Base frequencies are allowed to vary. Question #1: Which of these models is the least complex? Which of these models is the most parameter rich? Proportion of Invariable Sites (I) This is a model that assumes some proportion of the sites, p i, cannot change. Thus it makes two calculations for each base pair. First it calculates the chance, λ i, that that base pair would have the observed distribution that it does if it could not change. This will be 1, if it is the same in all taxa, or 0, if there are any differences among the taxa. It then calculates the probability, λ v, that it would have the observed distribution if it could change, using the transition matrix and the tree. Then it calculates the overall likelihood for that base as: λ = p i λ i + (1-p i ) λ v Among-site rate variation (Γ, also abbreviated G by jmodeltest) Under the null hypothesis, all sites are assumed to have equal rates of substitution. One way of relaxing this assumption is to allow the rates at different sites to be drawn from a gamma distribution (with the mean value across all sites within a class, such as A-T, represented in the substitution matrix). The gamma distribution is used because the shape of the curve (α = shape parameter) changes dramatically depending on the parameter values of the distribution. This calculation is done essentially the same way as it is for invariable sites. The likelihood is calculated for each value of the gamma distribution for each base pair and added together. In practice this is only done for a few values of the gamma distribution, as there are an infinite number of possible values for the gamma distribution and each likelihood calculation is computationally burdensome. This serves as a good approximation of a true gamma distribution. jmodeltest continued: Once the program is finished computing the likelihood scores, we need a way to evaluate which one is best. Adding parameters to a model always increases the maximum likelihood of the data. However, if a model has too many parameters, then maximum likelihood becomes unreliable. Therefore to accept a new parameter into your model it must produce a significant increase in the 3
4 likelihood. How do you tell if a difference in likelihood is significant? We want the model that best explains our data without adding too many parameters. 6. Select Analysis. You ll notice that you can now select Do AIC Calculations, Do BIC Calculations, or Do DT Calculations 7. Select Do AIC Calculations. Another window will pop up. Select Use AICc correction, Calculate parameter importances, Do model averaging, and Write PAUP* block. Select Do AIC calculations. That was much quicker. What is AIC or AICc for that matter? What about BIC? The Akaike Information Criterion (AIC) can be thought of as the amount of information that is lost when we use a specific model to approximate the real process of molecular evolution. Basically AIC compares several candidate models simultaneously and is used to compare both nested and non-nested models. AICc is used to correct for small sample size. AICc will approach AIC with larger sample sizes. The Bayesian Information Criterion (BIC) can be alternately used. This criterion gives equal priors for all competing models and choosing the model with the smallest BIC is equivalent to selecting the model with the maximum posterior probability. [If you have never been introduced to Bayesian statistics, what I just said probably doesn t make any sense to you. Don t worry, this will become clearer after we introduce Bayesian Inference next week.] 8. Select Results -> Show results table. Click on AICc and then select the AICc column. The chosen model has the lowest AICc score and will be highlighted. Question #2: Which model was selected using AICc? What is the likelihood value for this? Use the BIC calculation (remember to select Write PAUP* block). Was the same model selected as with AICc? What is the likelihood value for this? What if these two criteria differ in their model selection? Currently AICc and BIC are generally accepted as the two best criteria. I would suggest running each in PAUP* to see if they affect your tree inference. Keep the jmodeltest window open. Exercise 2: Maximum Likelihood (ML) in PAUP* First let s use the parameter values chosen by jmodeltest. In the jmodeltest output file you will find a PAUP block that can be inserted directly into the Nexus file. Scroll up in the window to find this for the AICc. It starts BEGIN PAUP; and ends with END; This block changes the Likelihood Settings (Lset), by setting the base frequencies at equilibrium (Base), the number of substitution types (Nst), the rate matrix of instantaneous substitution rates 4
5 (Rmat), the among site rate variation (Rates), the shape of the gamma distribution (Shape), and the proportion of invariant sites (Pinvar). Copy the PAUP block from the text file. I also include the line above this [! Likelihood settings etc]. Edit your Nexus file and paste the PAUP block from jmodeltest directly into it. It can go after any END; statement. Execute the newly-edited sequence file in PAUP* again. Change your working directory to a folder for the day if you d like. paup> execute filename Set the optimality criteria to likelihood: paup> set criterion=likelihood Make sure you run a heuristic search. paup> hs You can use savetrees to write your tree to file and save information about the branch lengths: paup> savetrees file=aiccmltree.tre brlens A new tree file called aiccmltree.tre should appear in your specified folder. Question #3: Take a screen shot of your tree using the AICc PAUP block in FigTree and send it to me. Do the same for the BIC PAUP block. Exercise 3: Get Acquainted with CIPRES and RAxML The CIPRES Science Gateway is a computational server that hosts popular phylogenetic research tools. There are four necessary steps to use these tools: Create an account, upload your data, submit your tasks, and analyze your output. Go to the CIPRES Gateway website: Click on the icon: If you have never used CIPRES, you ll need to register. So go ahead and do that, then login. Under the tab My Workbench create a new folder for today. When you create the folder, you ll notice that two subfolders are created à Data and Tasks. Let s examine the Toolkit tab. Look at all the great resources for you to use! Today we re going to use RAxML-HPC2 on XSEDE, but MrBayes and BEAST are available here and can be very useful for your projects. We ll discuss both of these programs next week in lab. RAxML (Randomized Axelerated Maximum Likelihood) is a program for Maximum Likelihood inference of large phylogenetic trees. This program employs heuristics to reduce 5
6 likelihood search time including building an initial tree under parsimony and incorporating a cooling schedule that allows backward steps during the hill-climbing process (see Stamatakis et al for more details). Keep in mind that RAxML gets your likelihood tree quickly, but does not search through tree space as rigorously there is always a trade-off. You will first need to upload the file primates.phy. Select your Data subfolder and Upload data. RAxML requires files in Phylip format like this file. Question #4: Describe the Phylip file format for me. What do the numbers at the top of the file indicate? Select RAxML-HPC2 on XSEDE and a new window pops up. Add a description for what you wan to call today s exercise. Select Input Data and select the primates.phy file you just uploaded. Select 38 Parametes Set. There are many options here and I won t go into all of them. 1. You can select the maximum number of hours you want to run the analysis. This is a small data set so the default of 0.25 right now is fine. 2. Sequence Type is Nucleotide 3. Enter the number of taxa and enter the number of patterns (=nucleotides). For some reason I can only edit this after checking the box I have a data set that may require more than 125 GB of memory (which is not true in this case). Go ahead and click that so you can edit these fields and then un-click. 4. Set the outgroup as Lemur_catta 5. Leave these defaults (explore these when you run your own analyses though) 6. Advanced Parameters à Nucleic Acid Options 7. Use GTRGAMMA for the bootstrapping phase and GTRGAMMA for the final tree (takes longer) 8. Configure Bootstrapping à Select the box for Conduct Rapid Bootstrapping 9. If you want the best ML tree select Conduct a rapid Bootstrap analysis and search for the best-scoring ML tree in one single program run. (-f a) 10. Change the # of bootstrap iterations if you d like we can easily run 1000 on this data set. 11. Save Parameters à Save and Run Task You will be sent an when your task is finished (very quickly for this example). Go back to CIPRES, select view output. Download the stdout.txt file and examine this file to make sure your parameters were set correctly. You can check the likelihood value as well. Also download RAxML_bipartitions.result. This is your best ML tree with bootstrap values. Question #5: What was the likelihood value? How does this compare with your previous analyses from today? Load your tree into FigTree and make sure the bootstrap values are visible, take a screen shot, and send it to me. 6
Distance Methods. "PRINCIPLES OF PHYLOGENETICS" Spring 2006
 Integrative Biology 200A University of California, Berkeley "PRINCIPLES OF PHYLOGENETICS" Spring 2006 Distance Methods Due at the end of class: - Distance matrices and trees for two different distance
Integrative Biology 200A University of California, Berkeley "PRINCIPLES OF PHYLOGENETICS" Spring 2006 Distance Methods Due at the end of class: - Distance matrices and trees for two different distance
Lab 06: Support Measures Bootstrap, Jackknife, and Bremer
 Integrative Biology 200, Spring 2014 Principles of Phylogenetics: Systematics University of California, Berkeley Updated by Traci L. Grzymala Lab 06: Support Measures Bootstrap, Jackknife, and Bremer So
Integrative Biology 200, Spring 2014 Principles of Phylogenetics: Systematics University of California, Berkeley Updated by Traci L. Grzymala Lab 06: Support Measures Bootstrap, Jackknife, and Bremer So
"PRINCIPLES OF PHYLOGENETICS" Spring 2008
 Integrative Biology 200A University of California, Berkeley "PRINCIPLES OF PHYLOGENETICS" Spring 2008 Lab 7: Introduction to PAUP* Today we will be learning about some of the basic features of PAUP* (Phylogenetic
Integrative Biology 200A University of California, Berkeley "PRINCIPLES OF PHYLOGENETICS" Spring 2008 Lab 7: Introduction to PAUP* Today we will be learning about some of the basic features of PAUP* (Phylogenetic
PRINCIPLES OF PHYLOGENETICS Spring 2008 Updated by Nick Matzke. Lab 11: MrBayes Lab
 Integrative Biology 200A University of California, Berkeley PRINCIPLES OF PHYLOGENETICS Spring 2008 Updated by Nick Matzke Lab 11: MrBayes Lab Note: try downloading and installing MrBayes on your own laptop,
Integrative Biology 200A University of California, Berkeley PRINCIPLES OF PHYLOGENETICS Spring 2008 Updated by Nick Matzke Lab 11: MrBayes Lab Note: try downloading and installing MrBayes on your own laptop,
CS 581. Tandy Warnow
 CS 581 Tandy Warnow This week Maximum parsimony: solving it on small datasets Maximum Likelihood optimization problem Felsenstein s pruning algorithm Bayesian MCMC methods Research opportunities Maximum
CS 581 Tandy Warnow This week Maximum parsimony: solving it on small datasets Maximum Likelihood optimization problem Felsenstein s pruning algorithm Bayesian MCMC methods Research opportunities Maximum
Codon models. In reality we use codon model Amino acid substitution rates meet nucleotide models Codon(nucleotide triplet)
 Phylogeny Codon models Last lecture: poor man s way of calculating dn/ds (Ka/Ks) Tabulate synonymous/non- synonymous substitutions Normalize by the possibilities Transform to genetic distance K JC or K
Phylogeny Codon models Last lecture: poor man s way of calculating dn/ds (Ka/Ks) Tabulate synonymous/non- synonymous substitutions Normalize by the possibilities Transform to genetic distance K JC or K
Sistemática Teórica. Hernán Dopazo. Biomedical Genomics and Evolution Lab. Lesson 03 Statistical Model Selection
 Sistemática Teórica Hernán Dopazo Biomedical Genomics and Evolution Lab Lesson 03 Statistical Model Selection Facultad de Ciencias Exactas y Naturales Universidad de Buenos Aires Argentina 2013 Statistical
Sistemática Teórica Hernán Dopazo Biomedical Genomics and Evolution Lab Lesson 03 Statistical Model Selection Facultad de Ciencias Exactas y Naturales Universidad de Buenos Aires Argentina 2013 Statistical
Which is more useful?
 Which is more useful? Reality Detailed map Detailed public transporta:on Simplified metro Models don t need to reflect reality A model is an inten:onal simplifica:on of a complex situa:on designed to eliminate
Which is more useful? Reality Detailed map Detailed public transporta:on Simplified metro Models don t need to reflect reality A model is an inten:onal simplifica:on of a complex situa:on designed to eliminate
CLC Phylogeny Module User manual
 CLC Phylogeny Module User manual User manual for Phylogeny Module 1.0 Windows, Mac OS X and Linux September 13, 2013 This software is for research purposes only. CLC bio Silkeborgvej 2 Prismet DK-8000
CLC Phylogeny Module User manual User manual for Phylogeny Module 1.0 Windows, Mac OS X and Linux September 13, 2013 This software is for research purposes only. CLC bio Silkeborgvej 2 Prismet DK-8000
Lab 8: Molecular Evolution
 Integrative Biology 200B University of California, Berkeley, Spring 2011 "Ecology and Evolution" by NM Hallinan, updated by Nick Matzke Lab 8: Molecular Evolution There are many different features of genes
Integrative Biology 200B University of California, Berkeley, Spring 2011 "Ecology and Evolution" by NM Hallinan, updated by Nick Matzke Lab 8: Molecular Evolution There are many different features of genes
ML phylogenetic inference and GARLI. Derrick Zwickl. University of Arizona (and University of Kansas) Workshop on Molecular Evolution 2015
 ML phylogenetic inference and GARLI Derrick Zwickl University of Arizona (and University of Kansas) Workshop on Molecular Evolution 2015 Outline Heuristics and tree searches ML phylogeny inference and
ML phylogenetic inference and GARLI Derrick Zwickl University of Arizona (and University of Kansas) Workshop on Molecular Evolution 2015 Outline Heuristics and tree searches ML phylogeny inference and
in interleaved format. The same data set in sequential format:
 PHYML user's guide Introduction PHYML is a software implementing a new method for building phylogenies from sequences using maximum likelihood. The executables can be downloaded at: http://www.lirmm.fr/~guindon/phyml.html.
PHYML user's guide Introduction PHYML is a software implementing a new method for building phylogenies from sequences using maximum likelihood. The executables can be downloaded at: http://www.lirmm.fr/~guindon/phyml.html.
Heterotachy models in BayesPhylogenies
 Heterotachy models in is a general software package for inferring phylogenetic trees using Bayesian Markov Chain Monte Carlo (MCMC) methods. The program allows a range of models of gene sequence evolution,
Heterotachy models in is a general software package for inferring phylogenetic trees using Bayesian Markov Chain Monte Carlo (MCMC) methods. The program allows a range of models of gene sequence evolution,
( ylogenetics/bayesian_workshop/bayesian%20mini conference.htm#_toc )
 (http://www.nematodes.org/teaching/tutorials/ph ylogenetics/bayesian_workshop/bayesian%20mini conference.htm#_toc145477467) Model selection criteria Review Posada D & Buckley TR (2004) Model selection
(http://www.nematodes.org/teaching/tutorials/ph ylogenetics/bayesian_workshop/bayesian%20mini conference.htm#_toc145477467) Model selection criteria Review Posada D & Buckley TR (2004) Model selection
Bayesian phylogenetic inference MrBayes (Practice)
 Bayesian phylogenetic inference MrBayes (Practice) The aim of this tutorial is to give a very short introduction to MrBayes. There is a website with most information about MrBayes: www.mrbayes.net (which
Bayesian phylogenetic inference MrBayes (Practice) The aim of this tutorial is to give a very short introduction to MrBayes. There is a website with most information about MrBayes: www.mrbayes.net (which
Lab 8: Using POY from your desktop and through CIPRES
 Integrative Biology 200A University of California, Berkeley PRINCIPLES OF PHYLOGENETICS Spring 2012 Updated by Michael Landis Lab 8: Using POY from your desktop and through CIPRES In this lab we re going
Integrative Biology 200A University of California, Berkeley PRINCIPLES OF PHYLOGENETICS Spring 2012 Updated by Michael Landis Lab 8: Using POY from your desktop and through CIPRES In this lab we re going
Introduction to MrBayes
 Introduction to MrBayes Fred(rik) Ronquist Dept. Bioinformatics and Genetics Swedish Museum of Natural History, Stockholm, Sweden Installing MrBayes! Two options:! Go to mrbayes.net, click Download and
Introduction to MrBayes Fred(rik) Ronquist Dept. Bioinformatics and Genetics Swedish Museum of Natural History, Stockholm, Sweden Installing MrBayes! Two options:! Go to mrbayes.net, click Download and
Molecular Evolution & Phylogenetics Complexity of the search space, distance matrix methods, maximum parsimony
 Molecular Evolution & Phylogenetics Complexity of the search space, distance matrix methods, maximum parsimony Basic Bioinformatics Workshop, ILRI Addis Ababa, 12 December 2017 Learning Objectives understand
Molecular Evolution & Phylogenetics Complexity of the search space, distance matrix methods, maximum parsimony Basic Bioinformatics Workshop, ILRI Addis Ababa, 12 December 2017 Learning Objectives understand
Phylogenetics on CUDA (Parallel) Architectures Bradly Alicea
 Descent w/modification Descent w/modification Descent w/modification Descent w/modification CPU Descent w/modification Descent w/modification Phylogenetics on CUDA (Parallel) Architectures Bradly Alicea
Descent w/modification Descent w/modification Descent w/modification Descent w/modification CPU Descent w/modification Descent w/modification Phylogenetics on CUDA (Parallel) Architectures Bradly Alicea
Parsimony-Based Approaches to Inferring Phylogenetic Trees
 Parsimony-Based Approaches to Inferring Phylogenetic Trees BMI/CS 576 www.biostat.wisc.edu/bmi576.html Mark Craven craven@biostat.wisc.edu Fall 0 Phylogenetic tree approaches! three general types! distance:
Parsimony-Based Approaches to Inferring Phylogenetic Trees BMI/CS 576 www.biostat.wisc.edu/bmi576.html Mark Craven craven@biostat.wisc.edu Fall 0 Phylogenetic tree approaches! three general types! distance:
Supplementary Information Nature Methods
 Supplementary Information Nature Methods jmodeltest 2: more models, new heuristics and parallel computing Diego Darriba, Guillermo L. Taboada, Ramón Doallo and David Posada Supplementary Table 1. New features
Supplementary Information Nature Methods jmodeltest 2: more models, new heuristics and parallel computing Diego Darriba, Guillermo L. Taboada, Ramón Doallo and David Posada Supplementary Table 1. New features
Phylogenetics. Introduction to Bioinformatics Dortmund, Lectures: Sven Rahmann. Exercises: Udo Feldkamp, Michael Wurst
 Phylogenetics Introduction to Bioinformatics Dortmund, 16.-20.07.2007 Lectures: Sven Rahmann Exercises: Udo Feldkamp, Michael Wurst 1 Phylogenetics phylum = tree phylogenetics: reconstruction of evolutionary
Phylogenetics Introduction to Bioinformatics Dortmund, 16.-20.07.2007 Lectures: Sven Rahmann Exercises: Udo Feldkamp, Michael Wurst 1 Phylogenetics phylum = tree phylogenetics: reconstruction of evolutionary
Hybrid Parallelization of the MrBayes & RAxML Phylogenetics Codes
 Hybrid Parallelization of the MrBayes & RAxML Phylogenetics Codes Wayne Pfeiffer (SDSC/UCSD) & Alexandros Stamatakis (TUM) February 25, 2010 What was done? Why is it important? Who cares? Hybrid MPI/OpenMP
Hybrid Parallelization of the MrBayes & RAxML Phylogenetics Codes Wayne Pfeiffer (SDSC/UCSD) & Alexandros Stamatakis (TUM) February 25, 2010 What was done? Why is it important? Who cares? Hybrid MPI/OpenMP
Protein phylogenetics
 Protein phylogenetics Robert Hirt PAUP4.0* can be used for an impressive range of analytical methods involving DNA alignments. This, unfortunately is not the case for estimating protein phylogenies. Only
Protein phylogenetics Robert Hirt PAUP4.0* can be used for an impressive range of analytical methods involving DNA alignments. This, unfortunately is not the case for estimating protein phylogenies. Only
STATISTICS IN THE BILLERA-HOLMES- VOGTMANN TREESPACE
 University of Kentucky UKnowledge Theses and Dissertations--Statistics Statistics 2015 STATISTICS IN THE BILLERA-HOLMES- VOGTMANN TREESPACE Grady S. Weyenberg University of Kentucky, gradysw@gmail.com
University of Kentucky UKnowledge Theses and Dissertations--Statistics Statistics 2015 STATISTICS IN THE BILLERA-HOLMES- VOGTMANN TREESPACE Grady S. Weyenberg University of Kentucky, gradysw@gmail.com
Tutorial using BEAST v2.4.7 MASCOT Tutorial Nicola F. Müller
 Tutorial using BEAST v2.4.7 MASCOT Tutorial Nicola F. Müller Parameter and State inference using the approximate structured coalescent 1 Background Phylogeographic methods can help reveal the movement
Tutorial using BEAST v2.4.7 MASCOT Tutorial Nicola F. Müller Parameter and State inference using the approximate structured coalescent 1 Background Phylogeographic methods can help reveal the movement
HIDDEN MARKOV MODELS AND SEQUENCE ALIGNMENT
 HIDDEN MARKOV MODELS AND SEQUENCE ALIGNMENT - Swarbhanu Chatterjee. Hidden Markov models are a sophisticated and flexible statistical tool for the study of protein models. Using HMMs to analyze proteins
HIDDEN MARKOV MODELS AND SEQUENCE ALIGNMENT - Swarbhanu Chatterjee. Hidden Markov models are a sophisticated and flexible statistical tool for the study of protein models. Using HMMs to analyze proteins
Lab 13: The What the Hell Do I Do with All These Trees Lab
 Integrative Biology 200A University of California, Berkeley Principals of Phylogenetics Spring 2012 Updated by Michael Landis Lab 13: The What the Hell Do I Do with All These Trees Lab We ve generated
Integrative Biology 200A University of California, Berkeley Principals of Phylogenetics Spring 2012 Updated by Michael Landis Lab 13: The What the Hell Do I Do with All These Trees Lab We ve generated
CSE 549: Computational Biology
 CSE 549: Computational Biology Phylogenomics 1 slides marked with * by Carl Kingsford Tree of Life 2 * H5N1 Influenza Strains Salzberg, Kingsford, et al., 2007 3 * H5N1 Influenza Strains The 2007 outbreak
CSE 549: Computational Biology Phylogenomics 1 slides marked with * by Carl Kingsford Tree of Life 2 * H5N1 Influenza Strains Salzberg, Kingsford, et al., 2007 3 * H5N1 Influenza Strains The 2007 outbreak
Lab 10: Introduction to RevBayes: Phylogenetic Analysis Using Graphical Models and Markov chain Monte Carlo By Will Freyman, edited by Carrie Tribble
 IB200, Spring 2018 University of California, Berkeley Lab 10: Introduction to RevBayes: Phylogenetic Analysis Using Graphical Models and Markov chain Monte Carlo By Will Freyman, edited by Carrie Tribble
IB200, Spring 2018 University of California, Berkeley Lab 10: Introduction to RevBayes: Phylogenetic Analysis Using Graphical Models and Markov chain Monte Carlo By Will Freyman, edited by Carrie Tribble
Tutorial: Phylogenetic Analysis on BioHealthBase Written by: Catherine A. Macken Version 1: February 2009
 Tutorial: Phylogenetic Analysis on BioHealthBase Written by: Catherine A. Macken Version 1: February 2009 BioHealthBase provides multiple functions for inferring phylogenetic trees, through the Phylogenetic
Tutorial: Phylogenetic Analysis on BioHealthBase Written by: Catherine A. Macken Version 1: February 2009 BioHealthBase provides multiple functions for inferring phylogenetic trees, through the Phylogenetic
Massachusetts Institute of Technology Computational Evolutionary Biology, Fall, 2005 Laboratory 3: Detecting selection
 Massachusetts Institute of Technology 6.877 Computational Evolutionary Biology, Fall, 2005 Laboratory 3: Detecting selection Handed out: November 28 Due: December 14 Part 2. Detecting selection likelihood
Massachusetts Institute of Technology 6.877 Computational Evolutionary Biology, Fall, 2005 Laboratory 3: Detecting selection Handed out: November 28 Due: December 14 Part 2. Detecting selection likelihood
Tutorial using BEAST v2.4.1 Troubleshooting David A. Rasmussen
 Tutorial using BEAST v2.4.1 Troubleshooting David A. Rasmussen 1 Background The primary goal of most phylogenetic analyses in BEAST is to infer the posterior distribution of trees and associated model
Tutorial using BEAST v2.4.1 Troubleshooting David A. Rasmussen 1 Background The primary goal of most phylogenetic analyses in BEAST is to infer the posterior distribution of trees and associated model
B Y P A S S R D E G R Version 1.0.2
 B Y P A S S R D E G R Version 1.0.2 March, 2008 Contents Overview.................................. 2 Installation notes.............................. 2 Quick Start................................. 3 Examples
B Y P A S S R D E G R Version 1.0.2 March, 2008 Contents Overview.................................. 2 Installation notes.............................. 2 Quick Start................................. 3 Examples
One report (in pdf format) addressing each of following questions.
 MSCBIO 2070/02-710: Computational Genomics, Spring 2016 HW1: Sequence alignment and Evolution Due: 24:00 EST, Feb 15, 2016 by autolab Your goals in this assignment are to 1. Complete a genome assembler
MSCBIO 2070/02-710: Computational Genomics, Spring 2016 HW1: Sequence alignment and Evolution Due: 24:00 EST, Feb 15, 2016 by autolab Your goals in this assignment are to 1. Complete a genome assembler
PHYLIP. Joe Felsenstein. Depts. of Genome Sciences and of Biology, University of Washington. PHYLIP p.1/13
 PHYLIP p.1/13 PHYLIP Joe Felsenstein Depts. of Genome Sciences and of Biology, University of Washington PHYLIP p.2/13 Software for this lab This lab is intended to introduce the PHYLIP package and a number
PHYLIP p.1/13 PHYLIP Joe Felsenstein Depts. of Genome Sciences and of Biology, University of Washington PHYLIP p.2/13 Software for this lab This lab is intended to introduce the PHYLIP package and a number
Workshop Practical on concatenation and model testing
 Workshop Practical on concatenation and model testing Jacob L. Steenwyk & Antonis Rokas Programs that you will use: Bash, Python, Perl, Phyutility, PartitionFinder, awk To infer a putative species phylogeny
Workshop Practical on concatenation and model testing Jacob L. Steenwyk & Antonis Rokas Programs that you will use: Bash, Python, Perl, Phyutility, PartitionFinder, awk To infer a putative species phylogeny
Evolutionary tree reconstruction (Chapter 10)
 Evolutionary tree reconstruction (Chapter 10) Early Evolutionary Studies Anatomical features were the dominant criteria used to derive evolutionary relationships between species since Darwin till early
Evolutionary tree reconstruction (Chapter 10) Early Evolutionary Studies Anatomical features were the dominant criteria used to derive evolutionary relationships between species since Darwin till early
A STEP-BY-STEP TUTORIAL FOR DISCRETE STATE PHYLOGEOGRAPHY INFERENCE
 BEAST: Bayesian Evolutionary Analysis by Sampling Trees A STEP-BY-STEP TUTORIAL FOR DISCRETE STATE PHYLOGEOGRAPHY INFERENCE This step-by-step tutorial guides you through a discrete phylogeography analysis
BEAST: Bayesian Evolutionary Analysis by Sampling Trees A STEP-BY-STEP TUTORIAL FOR DISCRETE STATE PHYLOGEOGRAPHY INFERENCE This step-by-step tutorial guides you through a discrete phylogeography analysis
BIOL 848: Phylogenetic Methods MrBayes Tutorial Fall, 2010
 MrBayes Lab Note: This computer lab exercise was written by Paul O. Lewis. Paul has graciously allowed Mark Holder to use and modify the lab for the Summer Institute in Statistical Genetics and BIOL 848.
MrBayes Lab Note: This computer lab exercise was written by Paul O. Lewis. Paul has graciously allowed Mark Holder to use and modify the lab for the Summer Institute in Statistical Genetics and BIOL 848.
Parsimony methods. Chapter 1
 Chapter 1 Parsimony methods Parsimony methods are the easiest ones to explain, and were also among the first methods for inferring phylogenies. The issues that they raise also involve many of the phenomena
Chapter 1 Parsimony methods Parsimony methods are the easiest ones to explain, and were also among the first methods for inferring phylogenies. The issues that they raise also involve many of the phenomena
TreeTime User Manual
 TreeTime User Manual Lin Himmelmann www.linhi.de Dirk Metzler www.zi.biologie.uni-muenchen.de/evol/statgen.html March 2009 TreeTime is controlled via an input file in Nexus file format (view Maddison 1997).
TreeTime User Manual Lin Himmelmann www.linhi.de Dirk Metzler www.zi.biologie.uni-muenchen.de/evol/statgen.html March 2009 TreeTime is controlled via an input file in Nexus file format (view Maddison 1997).
HORIZONTAL GENE TRANSFER DETECTION
 HORIZONTAL GENE TRANSFER DETECTION Sequenzanalyse und Genomik (Modul 10-202-2207) Alejandro Nabor Lozada-Chávez Before start, the user must create a new folder or directory (WORKING DIRECTORY) for all
HORIZONTAL GENE TRANSFER DETECTION Sequenzanalyse und Genomik (Modul 10-202-2207) Alejandro Nabor Lozada-Chávez Before start, the user must create a new folder or directory (WORKING DIRECTORY) for all
Lab 12: The What the Hell Do I Do with All These Trees Lab
 Integrative Biology 200A University of California, Berkeley Principals of Phylogenetics Spring 2010 Updated by Nick Matzke Lab 12: The What the Hell Do I Do with All These Trees Lab We ve generated a lot
Integrative Biology 200A University of California, Berkeley Principals of Phylogenetics Spring 2010 Updated by Nick Matzke Lab 12: The What the Hell Do I Do with All These Trees Lab We ve generated a lot
Lab 8 Phylogenetics I: creating and analysing a data matrix
 G44 Geobiology Fall 23 Name Lab 8 Phylogenetics I: creating and analysing a data matrix For this lab and the next you will need to download and install the Mesquite and PHYLIP packages: http://mesquiteproject.org/mesquite/mesquite.html
G44 Geobiology Fall 23 Name Lab 8 Phylogenetics I: creating and analysing a data matrix For this lab and the next you will need to download and install the Mesquite and PHYLIP packages: http://mesquiteproject.org/mesquite/mesquite.html
Lesson 13 Molecular Evolution
 Sequence Analysis Spring 2000 Dr. Richard Friedman (212)305-6901 (76901) friedman@cuccfa.ccc.columbia.edu 130BB Lesson 13 Molecular Evolution In this class we learn how to draw molecular evolutionary trees
Sequence Analysis Spring 2000 Dr. Richard Friedman (212)305-6901 (76901) friedman@cuccfa.ccc.columbia.edu 130BB Lesson 13 Molecular Evolution In this class we learn how to draw molecular evolutionary trees
Become familiar with the NEXUS data file format used by PAUP* (as well as several other prominent phylogeny programs such as Mesquite and MrBayes)
 PAUP* Lab Note: This computer lab exercise was written by Paul O. Lewis. Paul has graciously allowed Mark Holder to use and modify the lab for the Summer Institute in Statistical Genetics. Thanks, Paul!
PAUP* Lab Note: This computer lab exercise was written by Paul O. Lewis. Paul has graciously allowed Mark Holder to use and modify the lab for the Summer Institute in Statistical Genetics. Thanks, Paul!
Quick Start with CASSY Lab. Bi-05-05
 Quick Start with CASSY Lab Bi-05-05 About this manual This manual helps you getting started with the CASSY system. The manual does provide you the information you need to start quickly a simple CASSY experiment
Quick Start with CASSY Lab Bi-05-05 About this manual This manual helps you getting started with the CASSY system. The manual does provide you the information you need to start quickly a simple CASSY experiment
Simulation of Molecular Evolution with Bioinformatics Analysis
 Simulation of Molecular Evolution with Bioinformatics Analysis Barbara N. Beck, Rochester Community and Technical College, Rochester, MN Project created by: Barbara N. Beck, Ph.D., Rochester Community
Simulation of Molecular Evolution with Bioinformatics Analysis Barbara N. Beck, Rochester Community and Technical College, Rochester, MN Project created by: Barbara N. Beck, Ph.D., Rochester Community
Main Reference. Marc A. Suchard: Stochastic Models for Horizontal Gene Transfer: Taking a Random Walk through Tree Space Genetics 2005
 Stochastic Models for Horizontal Gene Transfer Dajiang Liu Department of Statistics Main Reference Marc A. Suchard: Stochastic Models for Horizontal Gene Transfer: Taing a Random Wal through Tree Space
Stochastic Models for Horizontal Gene Transfer Dajiang Liu Department of Statistics Main Reference Marc A. Suchard: Stochastic Models for Horizontal Gene Transfer: Taing a Random Wal through Tree Space
Using Hidden Markov Models to analyse time series data
 Using Hidden Markov Models to analyse time series data September 9, 2011 Background Want to analyse time series data coming from accelerometer measurements. 19 different datasets corresponding to different
Using Hidden Markov Models to analyse time series data September 9, 2011 Background Want to analyse time series data coming from accelerometer measurements. 19 different datasets corresponding to different
Estimating rates and dates from time-stamped sequences A hands-on practical
 Estimating rates and dates from time-stamped sequences A hands-on practical This chapter provides a step-by-step tutorial for analyzing a set of virus sequences which have been isolated at different points
Estimating rates and dates from time-stamped sequences A hands-on practical This chapter provides a step-by-step tutorial for analyzing a set of virus sequences which have been isolated at different points
Depending on the computer you find yourself in front of, here s what you ll need to do to open SPSS.
 1 SPSS 11.5 for Windows Introductory Assignment Material covered: Opening an existing SPSS data file, creating new data files, generating frequency distributions and descriptive statistics, obtaining printouts
1 SPSS 11.5 for Windows Introductory Assignment Material covered: Opening an existing SPSS data file, creating new data files, generating frequency distributions and descriptive statistics, obtaining printouts
Basic Tree Building With PAUP
 Phylogenetic Tree Building Objectives 1. Understand the principles of phylogenetic thinking. 2. Be able to develop and test a phylogenetic hypothesis. 3. Be able to interpret a phylogenetic tree. Overview
Phylogenetic Tree Building Objectives 1. Understand the principles of phylogenetic thinking. 2. Be able to develop and test a phylogenetic hypothesis. 3. Be able to interpret a phylogenetic tree. Overview
Bootstrapping Methods
 Bootstrapping Methods example of a Monte Carlo method these are one Monte Carlo statistical method some Bayesian statistical methods are Monte Carlo we can also simulate models using Monte Carlo methods
Bootstrapping Methods example of a Monte Carlo method these are one Monte Carlo statistical method some Bayesian statistical methods are Monte Carlo we can also simulate models using Monte Carlo methods
Introduction to Trees
 Introduction to Trees Tandy Warnow December 28, 2016 Introduction to Trees Tandy Warnow Clades of a rooted tree Every node v in a leaf-labelled rooted tree defines a subset of the leafset that is below
Introduction to Trees Tandy Warnow December 28, 2016 Introduction to Trees Tandy Warnow Clades of a rooted tree Every node v in a leaf-labelled rooted tree defines a subset of the leafset that is below
The NEXUS file includes a variety of node-date calibrations, including an offsetexp(min=45, mean=50) calibration for the root node:
 Appendix 1: Issues with the MrBayes dating analysis of Slater (2015). In setting up variant MrBayes analyses (Supplemental Table S2), a number of issues became apparent with the NEXUS file of the original
Appendix 1: Issues with the MrBayes dating analysis of Slater (2015). In setting up variant MrBayes analyses (Supplemental Table S2), a number of issues became apparent with the NEXUS file of the original
PartitionFinder2. Manual Rob Lanfear, December 2016
 PartitionFinder2 = new methods for selecting partitioned models of evolution for phylogenetic analyses. 1 PartitionFinder2 Manual Rob Lanfear, December 2016 Icon Ainsley Seago. Thanks Ainsley! Questions,
PartitionFinder2 = new methods for selecting partitioned models of evolution for phylogenetic analyses. 1 PartitionFinder2 Manual Rob Lanfear, December 2016 Icon Ainsley Seago. Thanks Ainsley! Questions,
A Statistical Test for Clades in Phylogenies
 A STATISTICAL TEST FOR CLADES A Statistical Test for Clades in Phylogenies Thurston H. Y. Dang 1, and Elchanan Mossel 2 1 Department of Electrical Engineering and Computer Sciences, University of California,
A STATISTICAL TEST FOR CLADES A Statistical Test for Clades in Phylogenies Thurston H. Y. Dang 1, and Elchanan Mossel 2 1 Department of Electrical Engineering and Computer Sciences, University of California,
MrBayes Lab. Exercise 1: Basics
 MrBayes Lab Note: This computer lab exercise was written by Paul O. Lewis. Paul has graciously allowed us to use and modify the lab for the Summer Institute in Statistical Genetics. Thanks, Paul! The main
MrBayes Lab Note: This computer lab exercise was written by Paul O. Lewis. Paul has graciously allowed us to use and modify the lab for the Summer Institute in Statistical Genetics. Thanks, Paul! The main
New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0
 Zurich Open Repository and Archive University of Zurich Main Library Strickhofstrasse 39 CH-8057 Zurich www.zora.uzh.ch Year: 2010 New algorithms and methods to estimate maximum-likelihood phylogenies:
Zurich Open Repository and Archive University of Zurich Main Library Strickhofstrasse 39 CH-8057 Zurich www.zora.uzh.ch Year: 2010 New algorithms and methods to estimate maximum-likelihood phylogenies:
Scaling species tree estimation methods to large datasets using NJMerge
 Scaling species tree estimation methods to large datasets using NJMerge Erin Molloy and Tandy Warnow {emolloy2, warnow}@illinois.edu University of Illinois at Urbana Champaign 2018 Phylogenomics Software
Scaling species tree estimation methods to large datasets using NJMerge Erin Molloy and Tandy Warnow {emolloy2, warnow}@illinois.edu University of Illinois at Urbana Champaign 2018 Phylogenomics Software
Genetics/MBT 541 Spring, 2002 Lecture 1 Joe Felsenstein Department of Genome Sciences Phylogeny methods, part 1 (Parsimony and such)
 Genetics/MBT 541 Spring, 2002 Lecture 1 Joe Felsenstein Department of Genome Sciences joe@gs Phylogeny methods, part 1 (Parsimony and such) Methods of reconstructing phylogenies (evolutionary trees) Parsimony
Genetics/MBT 541 Spring, 2002 Lecture 1 Joe Felsenstein Department of Genome Sciences joe@gs Phylogeny methods, part 1 (Parsimony and such) Methods of reconstructing phylogenies (evolutionary trees) Parsimony
Filogeografía BIOL 4211, Universidad de los Andes 25 de enero a 01 de abril 2006
 Laboratory excercise written by Andrew J. Crawford with the support of CIES Fulbright Program and Fulbright Colombia. Enjoy! Filogeografía BIOL 4211, Universidad de los Andes 25 de enero
Laboratory excercise written by Andrew J. Crawford with the support of CIES Fulbright Program and Fulbright Colombia. Enjoy! Filogeografía BIOL 4211, Universidad de los Andes 25 de enero
A Simple, Fast, and Accurate Method to Estimate Large Phylogenies by Maximum Likelihood
 A Simple, Fast, and Accurate Method to Estimate Large Phylogenies by Maximum Likelihood Stéphane Guindon, Olivier Gascuel To cite this version: Stéphane Guindon, Olivier Gascuel. A Simple, Fast, and Accurate
A Simple, Fast, and Accurate Method to Estimate Large Phylogenies by Maximum Likelihood Stéphane Guindon, Olivier Gascuel To cite this version: Stéphane Guindon, Olivier Gascuel. A Simple, Fast, and Accurate
Tutorial using BEAST v2.4.2 Introduction to BEAST2 Jūlija Pečerska and Veronika Bošková
 Tutorial using BEAST v2.4.2 Introduction to BEAST2 Jūlija Pečerska and Veronika Bošková This is a simple introductory tutorial to help you get started with using BEAST2 and its accomplices. 1 Background
Tutorial using BEAST v2.4.2 Introduction to BEAST2 Jūlija Pečerska and Veronika Bošková This is a simple introductory tutorial to help you get started with using BEAST2 and its accomplices. 1 Background
GARLI manual (version 0.95)
 GARLI manual (version 0.95) Short serial algorithm description... 2 A quick guide to running the program... 3 Serial FAQ... 4 Descriptions of serial GARLI settings... 9 General settings... 9 Model specification
GARLI manual (version 0.95) Short serial algorithm description... 2 A quick guide to running the program... 3 Serial FAQ... 4 Descriptions of serial GARLI settings... 9 General settings... 9 Model specification
Computational Genomics and Molecular Biology, Fall
 Computational Genomics and Molecular Biology, Fall 2015 1 Sequence Alignment Dannie Durand Pairwise Sequence Alignment The goal of pairwise sequence alignment is to establish a correspondence between the
Computational Genomics and Molecular Biology, Fall 2015 1 Sequence Alignment Dannie Durand Pairwise Sequence Alignment The goal of pairwise sequence alignment is to establish a correspondence between the
Generation of distancebased phylogenetic trees
 primer for practical phylogenetic data gathering. Uconn EEB3899-007. Spring 2015 Session 12 Generation of distancebased phylogenetic trees Rafael Medina (rafael.medina.bry@gmail.com) Yang Liu (yang.liu@uconn.edu)
primer for practical phylogenetic data gathering. Uconn EEB3899-007. Spring 2015 Session 12 Generation of distancebased phylogenetic trees Rafael Medina (rafael.medina.bry@gmail.com) Yang Liu (yang.liu@uconn.edu)
Lab 1: Introduction to Mesquite
 Integrative Biology 200B University of California, Berkeley Principles of Phylogenetics: Systematics Spring 2011 Updated by Nick Matzke (before and after lab, 1/27/11) Lab 1: Introduction to Mesquite Today
Integrative Biology 200B University of California, Berkeley Principles of Phylogenetics: Systematics Spring 2011 Updated by Nick Matzke (before and after lab, 1/27/11) Lab 1: Introduction to Mesquite Today
Enabling Phylogenetic Research via the CIPRES Science Gateway!
 Enabling Phylogenetic Research via the CIPRES Science Gateway Wayne Pfeiffer SDSC/UCSD August 5, 2013 In collaboration with Mark A. Miller, Terri Schwartz, & Bryan Lunt SDSC/UCSD Supported by NSF Phylogenetics
Enabling Phylogenetic Research via the CIPRES Science Gateway Wayne Pfeiffer SDSC/UCSD August 5, 2013 In collaboration with Mark A. Miller, Terri Schwartz, & Bryan Lunt SDSC/UCSD Supported by NSF Phylogenetics
Shortest Path Algorithm
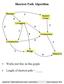 Shortest Path Algorithm C Works just fine on this graph. C Length of shortest path = Copyright 2005 DIMACS BioMath Connect Institute Robert Hochberg Dynamic Programming SP #1 Same Questions, Different
Shortest Path Algorithm C Works just fine on this graph. C Length of shortest path = Copyright 2005 DIMACS BioMath Connect Institute Robert Hochberg Dynamic Programming SP #1 Same Questions, Different
7.36/7.91/20.390/20.490/6.802/6.874 PROBLEM SET 3. Gibbs Sampler, RNA secondary structure, Protein Structure with PyRosetta, Connections (25 Points)
 7.36/7.91/20.390/20.490/6.802/6.874 PROBLEM SET 3. Gibbs Sampler, RNA secondary structure, Protein Structure with PyRosetta, Connections (25 Points) Due: Thursday, April 3 th at noon. Python Scripts All
7.36/7.91/20.390/20.490/6.802/6.874 PROBLEM SET 3. Gibbs Sampler, RNA secondary structure, Protein Structure with PyRosetta, Connections (25 Points) Due: Thursday, April 3 th at noon. Python Scripts All
Tips and Guidance for Analyzing Data. Executive Summary
 Tips and Guidance for Analyzing Data Executive Summary This document has information and suggestions about three things: 1) how to quickly do a preliminary analysis of time-series data; 2) key things to
Tips and Guidance for Analyzing Data Executive Summary This document has information and suggestions about three things: 1) how to quickly do a preliminary analysis of time-series data; 2) key things to
Stephen Scott.
 1 / 33 sscott@cse.unl.edu 2 / 33 Start with a set of sequences In each column, residues are homolgous Residues occupy similar positions in 3D structure Residues diverge from a common ancestral residue
1 / 33 sscott@cse.unl.edu 2 / 33 Start with a set of sequences In each column, residues are homolgous Residues occupy similar positions in 3D structure Residues diverge from a common ancestral residue
Recent Research Results. Evolutionary Trees Distance Methods
 Recent Research Results Evolutionary Trees Distance Methods Indo-European Languages After Tandy Warnow What is the purpose? Understand evolutionary history (relationship between species). Uderstand how
Recent Research Results Evolutionary Trees Distance Methods Indo-European Languages After Tandy Warnow What is the purpose? Understand evolutionary history (relationship between species). Uderstand how
SPSS 11.5 for Windows Assignment 2
 1 SPSS 11.5 for Windows Assignment 2 Material covered: Generating frequency distributions and descriptive statistics, converting raw scores to standard scores, creating variables using the Compute option,
1 SPSS 11.5 for Windows Assignment 2 Material covered: Generating frequency distributions and descriptive statistics, converting raw scores to standard scores, creating variables using the Compute option,
Barchard Introduction to SPSS Marks
 Barchard Introduction to SPSS 21.0 3 Marks Purpose The purpose of this assignment is to introduce you to SPSS, the most commonly used statistical package in the social sciences. You will create a new data
Barchard Introduction to SPSS 21.0 3 Marks Purpose The purpose of this assignment is to introduce you to SPSS, the most commonly used statistical package in the social sciences. You will create a new data
Tracy Heath Workshop on Molecular Evolution, Woods Hole USA
 INTRODUCTION TO BAYESIAN PHYLOGENETIC SOFTWARE Tracy Heath Integrative Biology, University of California, Berkeley Ecology & Evolutionary Biology, University of Kansas 2013 Workshop on Molecular Evolution,
INTRODUCTION TO BAYESIAN PHYLOGENETIC SOFTWARE Tracy Heath Integrative Biology, University of California, Berkeley Ecology & Evolutionary Biology, University of Kansas 2013 Workshop on Molecular Evolution,
Quartet Inference from SNP Data Under the Coalescent Model
 Quartet Inference from SNP Data Under the Coalescent Model Julia Chifman and Laura Kubatko By Shashank Yaduvanshi EsDmaDng Species Tree from Gene Sequences Input: Alignments from muldple genes Output:
Quartet Inference from SNP Data Under the Coalescent Model Julia Chifman and Laura Kubatko By Shashank Yaduvanshi EsDmaDng Species Tree from Gene Sequences Input: Alignments from muldple genes Output:
MetaPIGA v2.1. Maximum likelihood large phylogeny estimation using the metapopulation genetic algorithm (MetaGA) and other stochastic heuristics
 MetaPIGA v2.1 Maximum likelihood large phylogeny estimation using the metapopulation genetic algorithm (MetaGA) and other stochastic heuristics Manual version 2.1 (June 20, 2011) Michel C. Milinkovitch
MetaPIGA v2.1 Maximum likelihood large phylogeny estimation using the metapopulation genetic algorithm (MetaGA) and other stochastic heuristics Manual version 2.1 (June 20, 2011) Michel C. Milinkovitch
Barchard Introduction to SPSS Marks
 Barchard Introduction to SPSS 22.0 3 Marks Purpose The purpose of this assignment is to introduce you to SPSS, the most commonly used statistical package in the social sciences. You will create a new data
Barchard Introduction to SPSS 22.0 3 Marks Purpose The purpose of this assignment is to introduce you to SPSS, the most commonly used statistical package in the social sciences. You will create a new data
Species Trees with Relaxed Molecular Clocks Estimating per-species substitution rates using StarBEAST2
 Species Trees with Relaxed Molecular Clocks Estimating per-species substitution rates using StarBEAST2 Joseph Heled, Remco Bouckaert, Walter Xie, Alexei J. Drummond and Huw A. Ogilvie 1 Background In this
Species Trees with Relaxed Molecular Clocks Estimating per-species substitution rates using StarBEAST2 Joseph Heled, Remco Bouckaert, Walter Xie, Alexei J. Drummond and Huw A. Ogilvie 1 Background In this
PAUP tutorial - Macintosh
 1 PAUP tutorial - Macintosh The following tutorial is based on the tutorial provided with the PAUP package (but modified to emphasise morphological analysis, and updated to refer to the current command-line
1 PAUP tutorial - Macintosh The following tutorial is based on the tutorial provided with the PAUP package (but modified to emphasise morphological analysis, and updated to refer to the current command-line
Part I. Hierarchical clustering. Hierarchical Clustering. Hierarchical clustering. Produces a set of nested clusters organized as a
 Week 9 Based in part on slides from textbook, slides of Susan Holmes Part I December 2, 2012 Hierarchical Clustering 1 / 1 Produces a set of nested clusters organized as a Hierarchical hierarchical clustering
Week 9 Based in part on slides from textbook, slides of Susan Holmes Part I December 2, 2012 Hierarchical Clustering 1 / 1 Produces a set of nested clusters organized as a Hierarchical hierarchical clustering
Package markophylo. December 31, 2015
 Type Package Package markophylo December 31, 2015 Title Markov Chain Models for Phylogenetic Trees Version 1.0.4 Date 2015-12-31 Author Utkarsh J. Dang and G. Brian Golding Maintainer Utkarsh J. Dang
Type Package Package markophylo December 31, 2015 Title Markov Chain Models for Phylogenetic Trees Version 1.0.4 Date 2015-12-31 Author Utkarsh J. Dang and G. Brian Golding Maintainer Utkarsh J. Dang
10kTrees - Exercise #2. Viewing Trees Downloaded from 10kTrees: FigTree, R, and Mesquite
 10kTrees - Exercise #2 Viewing Trees Downloaded from 10kTrees: FigTree, R, and Mesquite The goal of this worked exercise is to view trees downloaded from 10kTrees, including tree blocks. You may wish to
10kTrees - Exercise #2 Viewing Trees Downloaded from 10kTrees: FigTree, R, and Mesquite The goal of this worked exercise is to view trees downloaded from 10kTrees, including tree blocks. You may wish to
Manual, ver. 03/01/2008, for Lever_1.1, and the associated programs PhylCRM_preprocess_1.1 and Lever_statistics_1.1 utilized in the paper:
 Manual, ver. 03/01/2008, for Lever_1.1, and the associated programs PhylCRM_preprocess_1.1 and Lever_statistics_1.1 utilized in the paper: Jason B. Warner 1,6, Anthony A. Philippakis 1,3,4,6, Savina A.
Manual, ver. 03/01/2008, for Lever_1.1, and the associated programs PhylCRM_preprocess_1.1 and Lever_statistics_1.1 utilized in the paper: Jason B. Warner 1,6, Anthony A. Philippakis 1,3,4,6, Savina A.
Stat 547 Assignment 3
 Stat 547 Assignment 3 Release Date: Saturday April 16, 2011 Due Date: Wednesday, April 27, 2011 at 4:30 PST Note that the deadline for this assignment is one day before the final project deadline, and
Stat 547 Assignment 3 Release Date: Saturday April 16, 2011 Due Date: Wednesday, April 27, 2011 at 4:30 PST Note that the deadline for this assignment is one day before the final project deadline, and
The CIPRES Science Gateway: Enabling High-Impact Science for Phylogenetics Researchers with Limited Resources
 The CIPRES Science Gateway: Enabling High-Impact Science for Phylogenetics Researchers with Limited Resources Mark Miller, Wayne Pfeiffer, and Terri Schwartz San Diego Supercomputer Center Phylogenetics
The CIPRES Science Gateway: Enabling High-Impact Science for Phylogenetics Researchers with Limited Resources Mark Miller, Wayne Pfeiffer, and Terri Schwartz San Diego Supercomputer Center Phylogenetics
The CIPRES Science Gateway: Enabling High-Impact Science for Phylogenetics Researchers with Limited Resources Mark A. Miller
 The CIPRES Science Gateway: Enabling High-Impact Science for Phylogenetics Researchers with Limited Resources Mark A. Miller San Diego Supercomputer Center Phylogenetics is the study of the diversification
The CIPRES Science Gateway: Enabling High-Impact Science for Phylogenetics Researchers with Limited Resources Mark A. Miller San Diego Supercomputer Center Phylogenetics is the study of the diversification
CPSC 340: Machine Learning and Data Mining. Feature Selection Fall 2016
 CPSC 34: Machine Learning and Data Mining Feature Selection Fall 26 Assignment 3: Admin Solutions will be posted after class Wednesday. Extra office hours Thursday: :3-2 and 4:3-6 in X836. Midterm Friday:
CPSC 34: Machine Learning and Data Mining Feature Selection Fall 26 Assignment 3: Admin Solutions will be posted after class Wednesday. Extra office hours Thursday: :3-2 and 4:3-6 in X836. Midterm Friday:
A. Using the data provided above, calculate the sampling variance and standard error for S for each week s data.
 WILD 502 Lab 1 Estimating Survival when Animal Fates are Known Today s lab will give you hands-on experience with estimating survival rates using logistic regression to estimate the parameters in a variety
WILD 502 Lab 1 Estimating Survival when Animal Fates are Known Today s lab will give you hands-on experience with estimating survival rates using logistic regression to estimate the parameters in a variety
Robust Signal-Structure Reconstruction
 Robust Signal-Structure Reconstruction V. Chetty 1, D. Hayden 2, J. Gonçalves 2, and S. Warnick 1 1 Information and Decision Algorithms Laboratories, Brigham Young University 2 Control Group, Department
Robust Signal-Structure Reconstruction V. Chetty 1, D. Hayden 2, J. Gonçalves 2, and S. Warnick 1 1 Information and Decision Algorithms Laboratories, Brigham Young University 2 Control Group, Department
Construction of a distance tree using clustering with the Unweighted Pair Group Method with Arithmatic Mean (UPGMA).
 Construction of a distance tree using clustering with the Unweighted Pair Group Method with Arithmatic Mean (UPGMA). The UPGMA is the simplest method of tree construction. It was originally developed for
Construction of a distance tree using clustering with the Unweighted Pair Group Method with Arithmatic Mean (UPGMA). The UPGMA is the simplest method of tree construction. It was originally developed for
Chapter 3. Bootstrap. 3.1 Introduction. 3.2 The general idea
 Chapter 3 Bootstrap 3.1 Introduction The estimation of parameters in probability distributions is a basic problem in statistics that one tends to encounter already during the very first course on the subject.
Chapter 3 Bootstrap 3.1 Introduction The estimation of parameters in probability distributions is a basic problem in statistics that one tends to encounter already during the very first course on the subject.
CPSC 340: Machine Learning and Data Mining. Feature Selection Fall 2017
 CPSC 340: Machine Learning and Data Mining Feature Selection Fall 2017 Assignment 2: Admin 1 late day to hand in tonight, 2 for Wednesday, answers posted Thursday. Extra office hours Thursday at 4pm (ICICS
CPSC 340: Machine Learning and Data Mining Feature Selection Fall 2017 Assignment 2: Admin 1 late day to hand in tonight, 2 for Wednesday, answers posted Thursday. Extra office hours Thursday at 4pm (ICICS
Lecture 20: Clustering and Evolution
 Lecture 20: Clustering and Evolution Study Chapter 10.4 10.8 11/12/2013 Comp 465 Fall 2013 1 Clique Graphs A clique is a graph where every vertex is connected via an edge to every other vertex A clique
Lecture 20: Clustering and Evolution Study Chapter 10.4 10.8 11/12/2013 Comp 465 Fall 2013 1 Clique Graphs A clique is a graph where every vertex is connected via an edge to every other vertex A clique
BIR pipeline steps and subsequent output files description STEP 1: BLAST search
 Lifeportal (Brief description) The Lifeportal at University of Oslo (https://lifeportal.uio.no) is a Galaxy based life sciences portal lifeportal.uio.no under the UiO tools section for phylogenomic analysis,
Lifeportal (Brief description) The Lifeportal at University of Oslo (https://lifeportal.uio.no) is a Galaxy based life sciences portal lifeportal.uio.no under the UiO tools section for phylogenomic analysis,
Notes on Simulations in SAS Studio
 Notes on Simulations in SAS Studio If you are not careful about simulations in SAS Studio, you can run into problems. In particular, SAS Studio has a limited amount of memory that you can use to write
Notes on Simulations in SAS Studio If you are not careful about simulations in SAS Studio, you can run into problems. In particular, SAS Studio has a limited amount of memory that you can use to write
