Biclustering Algorithms for Gene Expression Analysis
|
|
|
- Gervase Booth
- 5 years ago
- Views:
Transcription
1 Biclustering Algorithms for Gene Expression Analysis T. M. Murali August 19, 2008
2 Problems with Hierarchical Clustering It is a global clustering algorithm. Considers all genes to be equally important for all samples.
3 Problems with Hierarchical Clustering It is a global clustering algorithm. Considers all genes to be equally important for all samples. What if only a subset of the genes are co-expressed across only a subset of the samples? What if different subsets of the genes are co-expressed for different subsets of samples?
4 Example: Roberts et al. (Science 2000)
5 Example: Alizadeh et al. (Nature 2000)
6 Example: Alizadeh et al. (Nature 2000)
7 Biclustering A bicluster is a subset of genes and a subset of samples with the property that the selected genes are co-expressed only in the selected samples.
8 Biclustering A bicluster is a subset of genes and a subset of samples with the property that the selected genes are co-expressed only in the selected samples. By selecting samples and genes, a bicluster represents condition-specific patterns of expression. Issues in biclustering:
9 Biclustering A bicluster is a subset of genes and a subset of samples with the property that the selected genes are co-expressed only in the selected samples. By selecting samples and genes, a bicluster represents condition-specific patterns of expression. Issues in biclustering: How do we measure the degree of co-expression of a subset of genes in a subset of samples? How many biclusters should we compute? How do we compare two different sets of biclusters?
10 History of Biclustering Block clustering: Hartigan 1972, recursively partition matrix into blocks. Biclustering formulated in the context of gene expression data by Cheng and Church, ISMB Since 2000, a number of papers have been published on biclustering. Finding statistically-significant biclustering: Sharon, Tanay, and Shamir, ISMB Iterative signature algorithm: Bergmann, Ihmels, and Barkai, Phys Review E 2003 Two surveys of biclustering: Madeira and Oliveira, IEEE/ACM Transactions on Computational Biology and Bioinformatics, Tanay, Sharan, and Shamir, Handbook of Computational Molecular Biology, 2006.
11 Biclustering: Cheng and Church Defined the score of a bicluster to be its mean squared residue. Developed an iterative algorithm for computing biclusters with residue less than δ (specified by the user) by addition and deletion of genes and samples. To find multiple biclusters, they erase the values in the previously-computed biclusters and continue.
12 Mean Squared Residue A = matrix of gene expression values, a ij = value in the ith row and jth column of A. I = subset of genes/rows, J = subset of conditions/columns. A IJ = submatrix of A containing the rows in I and the columns in J. The mean squared residue of A IJ is H IJ = 1 I J i I,j J (a ij a ij a Ij a IJ ) 2, where a ij = average of values in A IJ along row i, a Ij = average of values in A IJ along column j and a IJ = average of all values in A IJ.
13 Examples of Mean Squared Residue Constant matrix: H IJ = 1 I J i I,j J (a ij a ij a Ij a IJ ) 2
14 Examples of Mean Squared Residue H IJ = 1 I J Constant matrix: 0. i I,j J (a ij a ij a Ij a IJ ) 2
15 Examples of Mean Squared Residue H IJ = 1 I J Constant matrix: 0. Single element: i I,j J (a ij a ij a Ij a IJ ) 2
16 Examples of Mean Squared Residue H IJ = 1 I J Constant matrix: 0. Single element: 0. i I,j J (a ij a ij a Ij a IJ ) 2
17 Examples of Mean Squared Residue H IJ = 1 I J Constant matrix: 0. Single element: 0. i I,j J (a ij a ij a Ij a IJ ) 2 Matrix with elements chosen randomly from the interval [a, b] has expected mean squared residue (b a) 2 /12.
18 Computational Problem Given a matrix A and a parameter δ find the largest submatrix of A with mean squared residue less than δ.
19 Computational Problem Given a matrix A and a parameter δ find the largest submatrix of A with mean squared residue less than δ. Why find the largest submatrix?
20 Computational Problem Given a matrix A and a parameter δ find the largest submatrix of A with mean squared residue less than δ. Why find the largest submatrix? How do we measure size of a submatrix?
21 Computational Problem Given a matrix A and a parameter δ find the largest submatrix of A with mean squared residue less than δ. Why find the largest submatrix? How do we measure size of a submatrix? Perimeter: maximise I + J. Square: maximise I = J. Area: maximise I J.
22 Computational Problem Given a matrix A and a parameter δ find the largest submatrix of A with mean squared residue less than δ. Why find the largest submatrix? How do we measure size of a submatrix? Perimeter: maximise I + J. Square: maximise I = J. Area: maximise I J. How hard is the problem?
23 Computational Problem Given a matrix A and a parameter δ find the largest submatrix of A with mean squared residue less than δ. Why find the largest submatrix? How do we measure size of a submatrix? Perimeter: maximise I + J. Square: maximise I = J. Area: maximise I J. How hard is the problem? Perimeter: maximise I + J, can be solved in polynomial time but is inappropriate when #rows >> #columns.
24 Computational Problem Given a matrix A and a parameter δ find the largest submatrix of A with mean squared residue less than δ. Why find the largest submatrix? How do we measure size of a submatrix? Perimeter: maximise I + J. Square: maximise I = J. Area: maximise I J. How hard is the problem? Perimeter: maximise I + J, can be solved in polynomial time but is inappropriate when #rows >> #columns. Square: maximise I = J, NP-Hard. Area: maximise I J, also NP-Hard (proven after the Cheng and Church paper).
25 Algorithms Since the problems are computationally intractable, use heuristics to find biclusters of large size. Basic idea: add/delete a row/column until mean squared residue does not decrease. Delete a row/column if its deletion improves mean squared residue. Add a row/column if its addition improves mean squared residue. Add some tricks to allow deletion/addition of multiple rows/columns so that it is not necessary to recompute mean squared residue after each change.
26 Biclustering: Sharan, Tanay, and Shamir Graph model: each gene is a node, each sample is a node.
27 Biclustering: Sharan, Tanay, and Shamir Graph model: each gene is a node, each sample is a node. Bipartite graph: connect a gene to a sample if the gene responds in that sample, i.e., if the expression value is not between -1 and 1 after standardisation.
28 Biclustering: Sharan, Tanay, and Shamir Graph model: each gene is a node, each sample is a node. Bipartite graph: connect a gene to a sample if the gene responds in that sample, i.e., if the expression value is not between -1 and 1 after standardisation.
29 Biclustering: Sharan, Tanay, and Shamir Graph model: each gene is a node, each sample is a node. Bipartite graph: connect a gene to a sample if the gene responds in that sample, i.e., if the expression value is not between -1 and 1 after standardisation. Bicluster clique; weight of a bicluster is the sum of the weights of its edges.
30 Computing Cliques Compute cliques with many edges or large weight.
31 Computing Cliques Compute cliques with many edges or large weight. Problem is NP-complete.
32 Computing Cliques Compute cliques with many edges or large weight. Problem is NP-complete. Assume each gene responds only in k samples.
33 Computing Cliques Compute cliques with many edges or large weight. Problem is NP-complete. Assume each gene responds only in k samples. Any clique has at most k samples in it.
34 Computing Cliques Compute cliques with many edges or large weight. Problem is NP-complete. Assume each gene responds only in k samples. Any clique has at most k samples in it. Algorithm: 1. For every gene, consider all subsets of neighbouring samples. 2. For each subset, find all the genes connected to all the samples in the subset. 3. Keep track of the clique of largest weight.
35 Computing Cliques Compute cliques with many edges or large weight. Problem is NP-complete. Assume each gene responds only in k samples. Any clique has at most k samples in it. Algorithm: 1. For every gene, consider all subsets of neighbouring samples. 2. For each subset, find all the genes connected to all the samples in the subset. 3. Keep track of the clique of largest weight. Running time is O(n2 k ), since each gene has at most O(2 k ) subsets of neighbouring samples. How do we obtain this running time? Exercise.
36 Computing Cliques Compute cliques with many edges or large weight. Problem is NP-complete. Assume each gene responds only in k samples. Any clique has at most k samples in it. Algorithm: 1. For every gene, consider all subsets of neighbouring samples. 2. For each subset, find all the genes connected to all the samples in the subset. 3. Keep track of the clique of largest weight. Running time is O(n2 k ), since each gene has at most O(2 k ) subsets of neighbouring samples. How do we obtain this running time? Exercise. Add some heuristics: iteratively add/delete node that improves the weight the most until no modification is possible.
37 Assessing Edge Weights Goal: assess statistical significance of a bicluster in random data; assign edge weights so that weight of a bicluster is equal to its statistical signficance. U = set of genes, V = set of conditions, E is the set of edges. Simple model: assume each edge occurs with probability p = E /( U V ). Given a bicluster H = (U, V, E ), the probability p(h) of observing a bicluster at least as dense as H is the probability that if we select each of the U V possible edges with probability p, we will select E or more edges:
38 Assessing Edge Weights Goal: assess statistical significance of a bicluster in random data; assign edge weights so that weight of a bicluster is equal to its statistical signficance. U = set of genes, V = set of conditions, E is the set of edges. Simple model: assume each edge occurs with probability p = E /( U V ). Given a bicluster H = (U, V, E ), the probability p(h) of observing a bicluster at least as dense as H is the probability that if we select each of the U V possible edges with probability p, we will select E or more edges: ( U V ) p i (1 p) U V i. i E i U V
39 Assessing Edge Weights Continued p(h) = ( U V ) E i U V i p i (1 p) U V i. If p 1/2, p(h) 2 U V p E (1 p) U V E. To minimise p(h), maximise log p(h) = U V E log p ( U V E ) log(1 p). Assign each edge in the graph a positive weight 1 log p and each edge not in the graph a negative weight 1 log(1 p).
Biclustering with δ-pcluster John Tantalo. 1. Introduction
 Biclustering with δ-pcluster John Tantalo 1. Introduction The subject of biclustering is chiefly concerned with locating submatrices of gene expression data that exhibit shared trends between genes. That
Biclustering with δ-pcluster John Tantalo 1. Introduction The subject of biclustering is chiefly concerned with locating submatrices of gene expression data that exhibit shared trends between genes. That
Biclustering Bioinformatics Data Sets. A Possibilistic Approach
 Possibilistic algorithm Bioinformatics Data Sets: A Possibilistic Approach Dept Computer and Information Sciences, University of Genova ITALY EMFCSC Erice 20/4/2007 Bioinformatics Data Sets Outline Introduction
Possibilistic algorithm Bioinformatics Data Sets: A Possibilistic Approach Dept Computer and Information Sciences, University of Genova ITALY EMFCSC Erice 20/4/2007 Bioinformatics Data Sets Outline Introduction
2. Background. 2.1 Clustering
 2. Background 2.1 Clustering Clustering involves the unsupervised classification of data items into different groups or clusters. Unsupervised classificaiton is basically a learning task in which learning
2. Background 2.1 Clustering Clustering involves the unsupervised classification of data items into different groups or clusters. Unsupervised classificaiton is basically a learning task in which learning
e-ccc-biclustering: Related work on biclustering algorithms for time series gene expression data
 : Related work on biclustering algorithms for time series gene expression data Sara C. Madeira 1,2,3, Arlindo L. Oliveira 1,2 1 Knowledge Discovery and Bioinformatics (KDBIO) group, INESC-ID, Lisbon, Portugal
: Related work on biclustering algorithms for time series gene expression data Sara C. Madeira 1,2,3, Arlindo L. Oliveira 1,2 1 Knowledge Discovery and Bioinformatics (KDBIO) group, INESC-ID, Lisbon, Portugal
DNA chips and other techniques measure the expression level of a large number of genes, perhaps all
 INESC-ID TECHNICAL REPORT 1/2004, JANUARY 2004 1 Biclustering Algorithms for Biological Data Analysis: A Survey* Sara C. Madeira and Arlindo L. Oliveira Abstract A large number of clustering approaches
INESC-ID TECHNICAL REPORT 1/2004, JANUARY 2004 1 Biclustering Algorithms for Biological Data Analysis: A Survey* Sara C. Madeira and Arlindo L. Oliveira Abstract A large number of clustering approaches
Mean Square Residue Biclustering with Missing Data and Row Inversions
 Mean Square Residue Biclustering with Missing Data and Row Inversions Stefan Gremalschi a, Gulsah Altun b, Irina Astrovskaya a, and Alexander Zelikovsky a a Department of Computer Science, Georgia State
Mean Square Residue Biclustering with Missing Data and Row Inversions Stefan Gremalschi a, Gulsah Altun b, Irina Astrovskaya a, and Alexander Zelikovsky a a Department of Computer Science, Georgia State
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 15: Microarray clustering http://compbio.pbworks.com/f/wood2.gif Some slides were adapted from Dr. Shaojie Zhang (University of Central Florida) Microarray
EECS730: Introduction to Bioinformatics Lecture 15: Microarray clustering http://compbio.pbworks.com/f/wood2.gif Some slides were adapted from Dr. Shaojie Zhang (University of Central Florida) Microarray
Order Preserving Clustering by Finding Frequent Orders in Gene Expression Data
 Order Preserving Clustering by Finding Frequent Orders in Gene Expression Data Li Teng and Laiwan Chan Department of Computer Science and Engineering, The Chinese University of Hong Kong, Hong Kong Abstract.
Order Preserving Clustering by Finding Frequent Orders in Gene Expression Data Li Teng and Laiwan Chan Department of Computer Science and Engineering, The Chinese University of Hong Kong, Hong Kong Abstract.
Biclustering algorithms ISA and SAMBA
 Biclustering algorithms ISA and SAMBA Slides with Amos Tanay 1 Biclustering Clusters: global. partition of genes according to common exp pattern across all conditions conditions Genes have multiple functions
Biclustering algorithms ISA and SAMBA Slides with Amos Tanay 1 Biclustering Clusters: global. partition of genes according to common exp pattern across all conditions conditions Genes have multiple functions
CS Introduction to Data Mining Instructor: Abdullah Mueen
 CS 591.03 Introduction to Data Mining Instructor: Abdullah Mueen LECTURE 8: ADVANCED CLUSTERING (FUZZY AND CO -CLUSTERING) Review: Basic Cluster Analysis Methods (Chap. 10) Cluster Analysis: Basic Concepts
CS 591.03 Introduction to Data Mining Instructor: Abdullah Mueen LECTURE 8: ADVANCED CLUSTERING (FUZZY AND CO -CLUSTERING) Review: Basic Cluster Analysis Methods (Chap. 10) Cluster Analysis: Basic Concepts
Optimal Web Page Category for Web Personalization Using Biclustering Approach
 Optimal Web Page Category for Web Personalization Using Biclustering Approach P. S. Raja Department of Computer Science, Periyar University, Salem, Tamil Nadu 636011, India. psraja5@gmail.com Abstract
Optimal Web Page Category for Web Personalization Using Biclustering Approach P. S. Raja Department of Computer Science, Periyar University, Salem, Tamil Nadu 636011, India. psraja5@gmail.com Abstract
Iterative Signature Algorithm for the Analysis of Large-Scale Gene Expression Data. By S. Bergmann, J. Ihmels, N. Barkai
 Iterative Signature Algorithm for the Analysis of Large-Scale Gene Expression Data By S. Bergmann, J. Ihmels, N. Barkai Reasoning Both clustering and Singular Value Decomposition(SVD) are useful tools
Iterative Signature Algorithm for the Analysis of Large-Scale Gene Expression Data By S. Bergmann, J. Ihmels, N. Barkai Reasoning Both clustering and Singular Value Decomposition(SVD) are useful tools
DNA chips and other techniques measure the expression
 24 IEEE TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS, VOL. 1, NO. 1, JANUARY-MARCH 2004 Biclustering Algorithms for Biological Data Analysis: A Survey Sara C. Madeira and Arlindo L. Oliveira
24 IEEE TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS, VOL. 1, NO. 1, JANUARY-MARCH 2004 Biclustering Algorithms for Biological Data Analysis: A Survey Sara C. Madeira and Arlindo L. Oliveira
Brain Inspired Cognitive Systems August 29 September 1, 2004 University of Stirling, Scotland, UK
 Brain Inspired Cognitive Systems August 29 September 1, 2004 University of Stirling, Scotland, UK A NEW BICLUSTERING TECHNIQUE BASED ON CROSSING MINIMIZATION Ahsan Abdullah Center for Agro-Informatics
Brain Inspired Cognitive Systems August 29 September 1, 2004 University of Stirling, Scotland, UK A NEW BICLUSTERING TECHNIQUE BASED ON CROSSING MINIMIZATION Ahsan Abdullah Center for Agro-Informatics
BIMAX. Lecture 11: December 31, Introduction Model An Incremental Algorithm.
 Analysis of DNA Chips and Gene Networks Spring Semester, 2009 Lecturer: Ron Shamir BIMAX 11.1 Introduction. Lecture 11: December 31, 2009 Scribe: Boris Kostenko In the course we have already seen different
Analysis of DNA Chips and Gene Networks Spring Semester, 2009 Lecturer: Ron Shamir BIMAX 11.1 Introduction. Lecture 11: December 31, 2009 Scribe: Boris Kostenko In the course we have already seen different
Clustering. RNA-seq: What is it good for? Finding Similarly Expressed Genes. Data... And Lots of It!
 RNA-seq: What is it good for? Clustering High-throughput RNA sequencing experiments (RNA-seq) offer the ability to measure simultaneously the expression level of thousands of genes in a single experiment!
RNA-seq: What is it good for? Clustering High-throughput RNA sequencing experiments (RNA-seq) offer the ability to measure simultaneously the expression level of thousands of genes in a single experiment!
Exploratory data analysis for microarrays
 Exploratory data analysis for microarrays Jörg Rahnenführer Computational Biology and Applied Algorithmics Max Planck Institute for Informatics D-66123 Saarbrücken Germany NGFN - Courses in Practical DNA
Exploratory data analysis for microarrays Jörg Rahnenführer Computational Biology and Applied Algorithmics Max Planck Institute for Informatics D-66123 Saarbrücken Germany NGFN - Courses in Practical DNA
Mining Deterministic Biclusters in Gene Expression Data
 Mining Deterministic Biclusters in Gene Expression Data Zonghong Zhang 1 Alvin Teo 1 BengChinOoi 1,2 Kian-Lee Tan 1,2 1 Department of Computer Science National University of Singapore 2 Singapore-MIT-Alliance
Mining Deterministic Biclusters in Gene Expression Data Zonghong Zhang 1 Alvin Teo 1 BengChinOoi 1,2 Kian-Lee Tan 1,2 1 Department of Computer Science National University of Singapore 2 Singapore-MIT-Alliance
Lecture 5: May 24, 2007
 Analysis of Gene Expression Data Spring Semester, 2007 Lecture 5: May 24, 2007 Lecturer: Ron Shamir Scribe: Shelly Mahlev and Shaul Karni 1 5.1 Introduction As we have seen in previous lectures, the technology
Analysis of Gene Expression Data Spring Semester, 2007 Lecture 5: May 24, 2007 Lecturer: Ron Shamir Scribe: Shelly Mahlev and Shaul Karni 1 5.1 Introduction As we have seen in previous lectures, the technology
A Scalable Framework for Discovering Coherent Co-clusters in Noisy Data
 A Scalable Framework for Discovering Coherent Co-clusters in Noisy Data Meghana Deodhar, Gunjan Gupta, Joydeep Ghosh deodhar,ggupta,ghosh@ece.utexas.edu Department of ECE, University of Texas at Austin,
A Scalable Framework for Discovering Coherent Co-clusters in Noisy Data Meghana Deodhar, Gunjan Gupta, Joydeep Ghosh deodhar,ggupta,ghosh@ece.utexas.edu Department of ECE, University of Texas at Austin,
COMBINATORIAL AND NONLINEAR OPTIMIZATION METHODS WITH APPLICATIONS IN DATA CLUSTERING, BICLUSTERING AND POWER SYSTEMS
 COMBINATORIAL AND NONLINEAR OPTIMIZATION METHODS WITH APPLICATIONS IN DATA CLUSTERING, BICLUSTERING AND POWER SYSTEMS By NENG FAN A DISSERTATION PRESENTED TO THE GRADUATE SCHOOL OF THE UNIVERSITY OF FLORIDA
COMBINATORIAL AND NONLINEAR OPTIMIZATION METHODS WITH APPLICATIONS IN DATA CLUSTERING, BICLUSTERING AND POWER SYSTEMS By NENG FAN A DISSERTATION PRESENTED TO THE GRADUATE SCHOOL OF THE UNIVERSITY OF FLORIDA
Reflexive Regular Equivalence for Bipartite Data
 Reflexive Regular Equivalence for Bipartite Data Aaron Gerow 1, Mingyang Zhou 2, Stan Matwin 1, and Feng Shi 3 1 Faculty of Computer Science, Dalhousie University, Halifax, NS, Canada 2 Department of Computer
Reflexive Regular Equivalence for Bipartite Data Aaron Gerow 1, Mingyang Zhou 2, Stan Matwin 1, and Feng Shi 3 1 Faculty of Computer Science, Dalhousie University, Halifax, NS, Canada 2 Department of Computer
Microarray data analysis
 Microarray data analysis Computational Biology IST Technical University of Lisbon Ana Teresa Freitas 016/017 Microarrays Rows represent genes Columns represent samples Many problems may be solved using
Microarray data analysis Computational Biology IST Technical University of Lisbon Ana Teresa Freitas 016/017 Microarrays Rows represent genes Columns represent samples Many problems may be solved using
Plaid models, biclustering, clustering on subsets of attributes, feature selection in clustering, et al.
 Plaid models, biclustering, clustering on subsets of attributes, feature selection in clustering, et al. Ramón Díaz-Uriarte rdiaz@cnio.es http://bioinfo.cnio.es/ rdiaz Unidad de Bioinformática Centro Nacional
Plaid models, biclustering, clustering on subsets of attributes, feature selection in clustering, et al. Ramón Díaz-Uriarte rdiaz@cnio.es http://bioinfo.cnio.es/ rdiaz Unidad de Bioinformática Centro Nacional
Unsupervised Learning. Unsupervised Learning. What is Clustering? Unsupervised Learning I Clustering 9/7/2017. Clustering
 Unsupervised Learning Clustering Centroid models (K-mean) Connectivity models (hierarchical clustering) Density models (DBSCAN) Graph-based models Subspace models (Biclustering) Feature extraction techniques
Unsupervised Learning Clustering Centroid models (K-mean) Connectivity models (hierarchical clustering) Density models (DBSCAN) Graph-based models Subspace models (Biclustering) Feature extraction techniques
9/29/13. Outline Data mining tasks. Clustering algorithms. Applications of clustering in biology
 9/9/ I9 Introduction to Bioinformatics, Clustering algorithms Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Outline Data mining tasks Predictive tasks vs descriptive tasks Example
9/9/ I9 Introduction to Bioinformatics, Clustering algorithms Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Outline Data mining tasks Predictive tasks vs descriptive tasks Example
2. Department of Electronic Engineering and Computer Science, Case Western Reserve University
 Chapter MINING HIGH-DIMENSIONAL DATA Wei Wang 1 and Jiong Yang 2 1. Department of Computer Science, University of North Carolina at Chapel Hill 2. Department of Electronic Engineering and Computer Science,
Chapter MINING HIGH-DIMENSIONAL DATA Wei Wang 1 and Jiong Yang 2 1. Department of Computer Science, University of North Carolina at Chapel Hill 2. Department of Electronic Engineering and Computer Science,
Biclustering for Microarray Data: A Short and Comprehensive Tutorial
 Biclustering for Microarray Data: A Short and Comprehensive Tutorial 1 Arabinda Panda, 2 Satchidananda Dehuri 1 Department of Computer Science, Modern Engineering & Management Studies, Balasore 2 Department
Biclustering for Microarray Data: A Short and Comprehensive Tutorial 1 Arabinda Panda, 2 Satchidananda Dehuri 1 Department of Computer Science, Modern Engineering & Management Studies, Balasore 2 Department
Nearest-Biclusters Collaborative Filtering
 Nearest-Biclusters Collaborative Filtering Panagiotis Symeonidis Alexandros Nanopoulos Apostolos Papadopoulos Yannis Manolopoulos Aristotle University, Department of Informatics, Thessaloniki 54124, Greece
Nearest-Biclusters Collaborative Filtering Panagiotis Symeonidis Alexandros Nanopoulos Apostolos Papadopoulos Yannis Manolopoulos Aristotle University, Department of Informatics, Thessaloniki 54124, Greece
Gene expression & Clustering (Chapter 10)
 Gene expression & Clustering (Chapter 10) Determining gene function Sequence comparison tells us if a gene is similar to another gene, e.g., in a new species Dynamic programming Approximate pattern matching
Gene expression & Clustering (Chapter 10) Determining gene function Sequence comparison tells us if a gene is similar to another gene, e.g., in a new species Dynamic programming Approximate pattern matching
Deposited on: 21 March 2012
 Filippone, M., Masulli, F., Rovetta, S., Mitra, S., and Banka, H. (2006) Possibilistic approach to biclustering: an application to oligonucleotide microarray data analysis. Lecture Notes in Computer Science,
Filippone, M., Masulli, F., Rovetta, S., Mitra, S., and Banka, H. (2006) Possibilistic approach to biclustering: an application to oligonucleotide microarray data analysis. Lecture Notes in Computer Science,
Social Data Management Communities
 Social Data Management Communities Antoine Amarilli 1, Silviu Maniu 2 January 9th, 2018 1 Télécom ParisTech 2 Université Paris-Sud 1/20 Table of contents Communities in Graphs 2/20 Graph Communities Communities
Social Data Management Communities Antoine Amarilli 1, Silviu Maniu 2 January 9th, 2018 1 Télécom ParisTech 2 Université Paris-Sud 1/20 Table of contents Communities in Graphs 2/20 Graph Communities Communities
Clustering. Robert M. Haralick. Computer Science, Graduate Center City University of New York
 Clustering Robert M. Haralick Computer Science, Graduate Center City University of New York Outline K-means 1 K-means 2 3 4 5 Clustering K-means The purpose of clustering is to determine the similarity
Clustering Robert M. Haralick Computer Science, Graduate Center City University of New York Outline K-means 1 K-means 2 3 4 5 Clustering K-means The purpose of clustering is to determine the similarity
Application of Hierarchical Clustering to Find Expression Modules in Cancer
 Application of Hierarchical Clustering to Find Expression Modules in Cancer T. M. Murali August 18, 2008 Innovative Application of Hierarchical Clustering A module map showing conditional activity of expression
Application of Hierarchical Clustering to Find Expression Modules in Cancer T. M. Murali August 18, 2008 Innovative Application of Hierarchical Clustering A module map showing conditional activity of expression
Co-clustering or Biclustering
 References: Co-clustering or Biclustering A. Anagnostopoulos, A. Dasgupta and R. Kumar: Approximation Algorithms for co-clustering, PODS 2008. K. Puolamaki. S. Hanhijarvi and G. Garriga: An approximation
References: Co-clustering or Biclustering A. Anagnostopoulos, A. Dasgupta and R. Kumar: Approximation Algorithms for co-clustering, PODS 2008. K. Puolamaki. S. Hanhijarvi and G. Garriga: An approximation
Controlling and visualizing the precision-recall tradeoff for external performance indices
 Controlling and visualizing the precision-recall tradeoff for external performance indices Blaise Hanczar 1 and Mohamed Nadif 2 1 IBISC, University of Paris-Saclay, Univ. Evry, Evry, France blaise.hanczar@ibisc.univ-evry.fr
Controlling and visualizing the precision-recall tradeoff for external performance indices Blaise Hanczar 1 and Mohamed Nadif 2 1 IBISC, University of Paris-Saclay, Univ. Evry, Evry, France blaise.hanczar@ibisc.univ-evry.fr
BIOINFORMATICS. Discovering statistically significant biclusters in gene expression data. Amos Tanay, Roded Sharan and Ron Shamir
 BIOINFORMATICS Electronic edition http://www.bioinformatics.oupjournals.org VOLUME 18 NUMBER Suppl. 1 JULY 2002 PAGES S136 S144 Discovering statistically significant biclusters in gene expression data
BIOINFORMATICS Electronic edition http://www.bioinformatics.oupjournals.org VOLUME 18 NUMBER Suppl. 1 JULY 2002 PAGES S136 S144 Discovering statistically significant biclusters in gene expression data
On the Approximability of Modularity Clustering
 On the Approximability of Modularity Clustering Newman s Community Finding Approach for Social Nets Bhaskar DasGupta Department of Computer Science University of Illinois at Chicago Chicago, IL 60607,
On the Approximability of Modularity Clustering Newman s Community Finding Approach for Social Nets Bhaskar DasGupta Department of Computer Science University of Illinois at Chicago Chicago, IL 60607,
Solving problems on graph algorithms
 Solving problems on graph algorithms Workshop Organized by: ACM Unit, Indian Statistical Institute, Kolkata. Tutorial-3 Date: 06.07.2017 Let G = (V, E) be an undirected graph. For a vertex v V, G {v} is
Solving problems on graph algorithms Workshop Organized by: ACM Unit, Indian Statistical Institute, Kolkata. Tutorial-3 Date: 06.07.2017 Let G = (V, E) be an undirected graph. For a vertex v V, G {v} is
Set Cover with Almost Consecutive Ones Property
 Set Cover with Almost Consecutive Ones Property 2004; Mecke, Wagner Entry author: Michael Dom INDEX TERMS: Covering Set problem, data reduction rules, enumerative algorithm. SYNONYMS: Hitting Set PROBLEM
Set Cover with Almost Consecutive Ones Property 2004; Mecke, Wagner Entry author: Michael Dom INDEX TERMS: Covering Set problem, data reduction rules, enumerative algorithm. SYNONYMS: Hitting Set PROBLEM
Analysis of Biological Networks. 1. Clustering 2. Random Walks 3. Finding paths
 Analysis of Biological Networks 1. Clustering 2. Random Walks 3. Finding paths Problem 1: Graph Clustering Finding dense subgraphs Applications Identification of novel pathways, complexes, other modules?
Analysis of Biological Networks 1. Clustering 2. Random Walks 3. Finding paths Problem 1: Graph Clustering Finding dense subgraphs Applications Identification of novel pathways, complexes, other modules?
FISA: Fast Iterative Signature Algorithm for the analysis of large-scale gene expression data
 FISA: Fast Iterative Signature Algorithm for the analysis of large-scale gene expression data Seema Aggarwal Department of Computer Science University of Delhi saggarwal@mh.du.ac.in and Neelima Gupta Department
FISA: Fast Iterative Signature Algorithm for the analysis of large-scale gene expression data Seema Aggarwal Department of Computer Science University of Delhi saggarwal@mh.du.ac.in and Neelima Gupta Department
Statistical Methods and Optimization in Data Mining
 Statistical Methods and Optimization in Data Mining Eloísa Macedo 1, Adelaide Freitas 2 1 University of Aveiro, Aveiro, Portugal; macedo@ua.pt 2 University of Aveiro, Aveiro, Portugal; adelaide@ua.pt The
Statistical Methods and Optimization in Data Mining Eloísa Macedo 1, Adelaide Freitas 2 1 University of Aveiro, Aveiro, Portugal; macedo@ua.pt 2 University of Aveiro, Aveiro, Portugal; adelaide@ua.pt The
Numerical Algorithms
 Chapter 10 Slide 464 Numerical Algorithms Slide 465 Numerical Algorithms In textbook do: Matrix multiplication Solving a system of linear equations Slide 466 Matrices A Review An n m matrix Column a 0,0
Chapter 10 Slide 464 Numerical Algorithms Slide 465 Numerical Algorithms In textbook do: Matrix multiplication Solving a system of linear equations Slide 466 Matrices A Review An n m matrix Column a 0,0
CSC 8301 Design and Analysis of Algorithms: Exhaustive Search
 CSC 8301 Design and Analysis of Algorithms: Exhaustive Search Professor Henry Carter Fall 2016 Recap Brute force is the use of iterative checking or solving a problem by its definition The straightforward
CSC 8301 Design and Analysis of Algorithms: Exhaustive Search Professor Henry Carter Fall 2016 Recap Brute force is the use of iterative checking or solving a problem by its definition The straightforward
Identifying network modules
 Network biology minicourse (part 3) Algorithmic challenges in genomics Identifying network modules Roded Sharan School of Computer Science, Tel Aviv University Gene/Protein Modules A module is a set of
Network biology minicourse (part 3) Algorithmic challenges in genomics Identifying network modules Roded Sharan School of Computer Science, Tel Aviv University Gene/Protein Modules A module is a set of
Biclustering Algorithms: A Survey
 Biclustering Algorithms: A Survey Amos Tanay Λ Roded Sharan y Ron Shamir Λ May 2004 Abstract Analysis of large scale geonomics data, notably gene expression, has initially focused on clustering methods.
Biclustering Algorithms: A Survey Amos Tanay Λ Roded Sharan y Ron Shamir Λ May 2004 Abstract Analysis of large scale geonomics data, notably gene expression, has initially focused on clustering methods.
Triclustering in Gene Expression Data Analysis: A Selected Survey
 Triclustering in Gene Expression Data Analysis: A Selected Survey P. Mahanta, H. A. Ahmed Dept of Comp Sc and Engg Tezpur University Napaam -784028, India Email: priyakshi@tezu.ernet.in, hasin@tezu.ernet.in
Triclustering in Gene Expression Data Analysis: A Selected Survey P. Mahanta, H. A. Ahmed Dept of Comp Sc and Engg Tezpur University Napaam -784028, India Email: priyakshi@tezu.ernet.in, hasin@tezu.ernet.in
Basics of Network Analysis
 Basics of Network Analysis Hiroki Sayama sayama@binghamton.edu Graph = Network G(V, E): graph (network) V: vertices (nodes), E: edges (links) 1 Nodes = 1, 2, 3, 4, 5 2 3 Links = 12, 13, 15, 23,
Basics of Network Analysis Hiroki Sayama sayama@binghamton.edu Graph = Network G(V, E): graph (network) V: vertices (nodes), E: edges (links) 1 Nodes = 1, 2, 3, 4, 5 2 3 Links = 12, 13, 15, 23,
Olmo S. Zavala Romero. Clustering Hierarchical Distance Group Dist. K-means. Center of Atmospheric Sciences, UNAM.
 Center of Atmospheric Sciences, UNAM November 16, 2016 Cluster Analisis Cluster analysis or clustering is the task of grouping a set of objects in such a way that objects in the same group (called a cluster)
Center of Atmospheric Sciences, UNAM November 16, 2016 Cluster Analisis Cluster analysis or clustering is the task of grouping a set of objects in such a way that objects in the same group (called a cluster)
Mixture Models and the EM Algorithm
 Mixture Models and the EM Algorithm Padhraic Smyth, Department of Computer Science University of California, Irvine c 2017 1 Finite Mixture Models Say we have a data set D = {x 1,..., x N } where x i is
Mixture Models and the EM Algorithm Padhraic Smyth, Department of Computer Science University of California, Irvine c 2017 1 Finite Mixture Models Say we have a data set D = {x 1,..., x N } where x i is
Exploiting Parallelism to Support Scalable Hierarchical Clustering
 Exploiting Parallelism to Support Scalable Hierarchical Clustering Rebecca Cathey, Eric Jensen, Steven Beitzel, Ophir Frieder, David Grossman Information Retrieval Laboratory http://ir.iit.edu Background
Exploiting Parallelism to Support Scalable Hierarchical Clustering Rebecca Cathey, Eric Jensen, Steven Beitzel, Ophir Frieder, David Grossman Information Retrieval Laboratory http://ir.iit.edu Background
Machine learning - HT Clustering
 Machine learning - HT 2016 10. Clustering Varun Kanade University of Oxford March 4, 2016 Announcements Practical Next Week - No submission Final Exam: Pick up on Monday Material covered next week is not
Machine learning - HT 2016 10. Clustering Varun Kanade University of Oxford March 4, 2016 Announcements Practical Next Week - No submission Final Exam: Pick up on Monday Material covered next week is not
Unsupervised Learning : Clustering
 Unsupervised Learning : Clustering Things to be Addressed Traditional Learning Models. Cluster Analysis K-means Clustering Algorithm Drawbacks of traditional clustering algorithms. Clustering as a complex
Unsupervised Learning : Clustering Things to be Addressed Traditional Learning Models. Cluster Analysis K-means Clustering Algorithm Drawbacks of traditional clustering algorithms. Clustering as a complex
Package superbiclust
 Type Package Package superbiclust February 20, 2015 Title Generating Robust Biclusters from a Bicluster Set (Ensemble Biclustering) Version 1.1 Date 2012-08-23 Author Maintainer
Type Package Package superbiclust February 20, 2015 Title Generating Robust Biclusters from a Bicluster Set (Ensemble Biclustering) Version 1.1 Date 2012-08-23 Author Maintainer
Use of biclustering for missing value imputation in gene expression data
 ORIGINAL RESEARCH Use of biclustering for missing value imputation in gene expression data K.O. Cheng, N.F. Law, W.C. Siu Department of Electronic and Information Engineering, The Hong Kong Polytechnic
ORIGINAL RESEARCH Use of biclustering for missing value imputation in gene expression data K.O. Cheng, N.F. Law, W.C. Siu Department of Electronic and Information Engineering, The Hong Kong Polytechnic
BiRange:An Efficient Framework for Biclustering of Gene Expression Data Using Range Bipartite Graph
 American Journal of Bioinformatics Research 212, 2(4): 4-46 DOI: 1.5923/j.bioinformatics.21224.2 BiRange:An Efficient Framework for Biclustering of Gene Expression Data Using Range Bipartite Graph Suvendu
American Journal of Bioinformatics Research 212, 2(4): 4-46 DOI: 1.5923/j.bioinformatics.21224.2 BiRange:An Efficient Framework for Biclustering of Gene Expression Data Using Range Bipartite Graph Suvendu
Maximizing edge-ratio is NP-complete
 Maximizing edge-ratio is NP-complete Steven D Noble, Pierre Hansen and Nenad Mladenović February 7, 01 Abstract Given a graph G and a bipartition of its vertices, the edge-ratio is the minimum for both
Maximizing edge-ratio is NP-complete Steven D Noble, Pierre Hansen and Nenad Mladenović February 7, 01 Abstract Given a graph G and a bipartition of its vertices, the edge-ratio is the minimum for both
Markov Network Structure Learning
 Markov Network Structure Learning Sargur Srihari srihari@cedar.buffalo.edu 1 Topics Structure Learning as model selection Structure Learning using Independence Tests Score based Learning: Hypothesis Spaces
Markov Network Structure Learning Sargur Srihari srihari@cedar.buffalo.edu 1 Topics Structure Learning as model selection Structure Learning using Independence Tests Score based Learning: Hypothesis Spaces
Lesson 3. Prof. Enza Messina
 Lesson 3 Prof. Enza Messina Clustering techniques are generally classified into these classes: PARTITIONING ALGORITHMS Directly divides data points into some prespecified number of clusters without a hierarchical
Lesson 3 Prof. Enza Messina Clustering techniques are generally classified into these classes: PARTITIONING ALGORITHMS Directly divides data points into some prespecified number of clusters without a hierarchical
Clustering Lecture 9: Other Topics. Jing Gao SUNY Buffalo
 Clustering Lecture 9: Other Topics Jing Gao SUNY Buffalo 1 Basics Outline Motivation, definition, evaluation Methods Partitional Hierarchical Density-based Miture model Spectral methods Advanced topics
Clustering Lecture 9: Other Topics Jing Gao SUNY Buffalo 1 Basics Outline Motivation, definition, evaluation Methods Partitional Hierarchical Density-based Miture model Spectral methods Advanced topics
Exploratory data analysis for microarrays
 Exploratory data analysis for microarrays Adrian Alexa Computational Biology and Applied Algorithmics Max Planck Institute for Informatics D-66123 Saarbrücken slides by Jörg Rahnenführer NGFN - Courses
Exploratory data analysis for microarrays Adrian Alexa Computational Biology and Applied Algorithmics Max Planck Institute for Informatics D-66123 Saarbrücken slides by Jörg Rahnenführer NGFN - Courses
Multivariate Analysis (slides 9)
 Multivariate Analysis (slides 9) Today we consider k-means clustering. We will address the question of selecting the appropriate number of clusters. Properties and limitations of the algorithm will be
Multivariate Analysis (slides 9) Today we consider k-means clustering. We will address the question of selecting the appropriate number of clusters. Properties and limitations of the algorithm will be
Exploratory data analysis for microarrays
 Exploratory data analysis for microarrays Jörg Rahnenführer Computational Biology and Applied Algorithmics Max Planck Institute for Informatics D-66123 Saarbrücken Germany NGFN - Courses in Practical DNA
Exploratory data analysis for microarrays Jörg Rahnenführer Computational Biology and Applied Algorithmics Max Planck Institute for Informatics D-66123 Saarbrücken Germany NGFN - Courses in Practical DNA
Data Clustering. Algorithmic Thinking Luay Nakhleh Department of Computer Science Rice University
 Data Clustering Algorithmic Thinking Luay Nakhleh Department of Computer Science Rice University Data clustering is the task of partitioning a set of objects into groups such that the similarity of objects
Data Clustering Algorithmic Thinking Luay Nakhleh Department of Computer Science Rice University Data clustering is the task of partitioning a set of objects into groups such that the similarity of objects
Gene Expression Data Knowledge Discovery using Global and Local Clustering
 HTTPS://SITES.GOOGLE.COM/SITE/JOURNALOFCOMPUTING/ 116 Gene Expression Data Knowledge Discovery using Global and Local Clustering Swathi. H Abstract To understand complex biological systems, the research
HTTPS://SITES.GOOGLE.COM/SITE/JOURNALOFCOMPUTING/ 116 Gene Expression Data Knowledge Discovery using Global and Local Clustering Swathi. H Abstract To understand complex biological systems, the research
CS242: Probabilistic Graphical Models Lecture 3: Factor Graphs & Variable Elimination
 CS242: Probabilistic Graphical Models Lecture 3: Factor Graphs & Variable Elimination Instructor: Erik Sudderth Brown University Computer Science September 11, 2014 Some figures and materials courtesy
CS242: Probabilistic Graphical Models Lecture 3: Factor Graphs & Variable Elimination Instructor: Erik Sudderth Brown University Computer Science September 11, 2014 Some figures and materials courtesy
Biclustering1 has proved of great value for finding interesting patterns in
 D a t a M i n i n g f o r B i o i n f o r m a t i c s MicroCluster: Efficient Deterministic Biclustering of Microarray Data Lizhuang Zhao and Mohammed J. Zaki, Rensselaer Polytechnic Institute Biclustering1
D a t a M i n i n g f o r B i o i n f o r m a t i c s MicroCluster: Efficient Deterministic Biclustering of Microarray Data Lizhuang Zhao and Mohammed J. Zaki, Rensselaer Polytechnic Institute Biclustering1
CS 231A CA Session: Problem Set 4 Review. Kevin Chen May 13, 2016
 CS 231A CA Session: Problem Set 4 Review Kevin Chen May 13, 2016 PS4 Outline Problem 1: Viewpoint estimation Problem 2: Segmentation Meanshift segmentation Normalized cut Problem 1: Viewpoint Estimation
CS 231A CA Session: Problem Set 4 Review Kevin Chen May 13, 2016 PS4 Outline Problem 1: Viewpoint estimation Problem 2: Segmentation Meanshift segmentation Normalized cut Problem 1: Viewpoint Estimation
Automatic Cluster Number Selection using a Split and Merge K-Means Approach
 Automatic Cluster Number Selection using a Split and Merge K-Means Approach Markus Muhr and Michael Granitzer 31st August 2009 The Know-Center is partner of Austria's Competence Center Program COMET. Agenda
Automatic Cluster Number Selection using a Split and Merge K-Means Approach Markus Muhr and Michael Granitzer 31st August 2009 The Know-Center is partner of Austria's Competence Center Program COMET. Agenda
Biomedical Data Exploration with Spectral Methods and Canonical Correlation. Outline. Diversity of Data. Nataliya Sokolovska
 Biomedical Data Exploration with Spectral Methods and Canonical Correlation Nataliya Sokolovska University Paris 6 INSERM, team NutriOmics, ICAN Outline Diversity of Data and Computational Challenges Data
Biomedical Data Exploration with Spectral Methods and Canonical Correlation Nataliya Sokolovska University Paris 6 INSERM, team NutriOmics, ICAN Outline Diversity of Data and Computational Challenges Data
Analysis of Biological Networks: Protein modules Color Coding
 Analysis of Biological Networks: Protein modules Color Coding Lecturer: Eithan Hirsh Scribe: Shelly Mahleb and Benny Davidovich Lecture 6, November 30, 2006 1 Introduction In this lecture we present: Color
Analysis of Biological Networks: Protein modules Color Coding Lecturer: Eithan Hirsh Scribe: Shelly Mahleb and Benny Davidovich Lecture 6, November 30, 2006 1 Introduction In this lecture we present: Color
CSE 417 Network Flows (pt 3) Modeling with Min Cuts
 CSE 417 Network Flows (pt 3) Modeling with Min Cuts Reminders > HW6 is due on Friday start early bug fixed on line 33 of OptimalLineup.java: > change true to false Review of last two lectures > Defined
CSE 417 Network Flows (pt 3) Modeling with Min Cuts Reminders > HW6 is due on Friday start early bug fixed on line 33 of OptimalLineup.java: > change true to false Review of last two lectures > Defined
BOOLEAN MATRIX FACTORISATIONS IN DATA MINING (AND ELSEWHERE) Pauli Miettinen 15 April 2013
 BOOLEAN MATRIX FACTORISATIONS IN DATA MINING (AND ELSEWHERE) Pauli Miettinen 15 April 2013 In the sleepy days when the provinces of France were still quietly provincial, matrices with Boolean entries were
BOOLEAN MATRIX FACTORISATIONS IN DATA MINING (AND ELSEWHERE) Pauli Miettinen 15 April 2013 In the sleepy days when the provinces of France were still quietly provincial, matrices with Boolean entries were
Progress Report: Collaborative Filtering Using Bregman Co-clustering
 Progress Report: Collaborative Filtering Using Bregman Co-clustering Wei Tang, Srivatsan Ramanujam, and Andrew Dreher April 4, 2008 1 Introduction Analytics are becoming increasingly important for business
Progress Report: Collaborative Filtering Using Bregman Co-clustering Wei Tang, Srivatsan Ramanujam, and Andrew Dreher April 4, 2008 1 Introduction Analytics are becoming increasingly important for business
Unsupervised Learning. Presenter: Anil Sharma, PhD Scholar, IIIT-Delhi
 Unsupervised Learning Presenter: Anil Sharma, PhD Scholar, IIIT-Delhi Content Motivation Introduction Applications Types of clustering Clustering criterion functions Distance functions Normalization Which
Unsupervised Learning Presenter: Anil Sharma, PhD Scholar, IIIT-Delhi Content Motivation Introduction Applications Types of clustering Clustering criterion functions Distance functions Normalization Which
Cluster Analysis. Ying Shen, SSE, Tongji University
 Cluster Analysis Ying Shen, SSE, Tongji University Cluster analysis Cluster analysis groups data objects based only on the attributes in the data. The main objective is that The objects within a group
Cluster Analysis Ying Shen, SSE, Tongji University Cluster analysis Cluster analysis groups data objects based only on the attributes in the data. The main objective is that The objects within a group
Correspondence Clustering: An Approach to Cluster Multiple Related Spatial Datasets
 Correspondence Clustering: An Approach to Cluster Multiple Related Spatial Datasets Vadeerat Rinsurongkawong, and Christoph F. Eick Department of Computer Science, University of Houston, Houston, TX 77204-3010
Correspondence Clustering: An Approach to Cluster Multiple Related Spatial Datasets Vadeerat Rinsurongkawong, and Christoph F. Eick Department of Computer Science, University of Houston, Houston, TX 77204-3010
Dense Matrix Algorithms
 Dense Matrix Algorithms Ananth Grama, Anshul Gupta, George Karypis, and Vipin Kumar To accompany the text Introduction to Parallel Computing, Addison Wesley, 2003. Topic Overview Matrix-Vector Multiplication
Dense Matrix Algorithms Ananth Grama, Anshul Gupta, George Karypis, and Vipin Kumar To accompany the text Introduction to Parallel Computing, Addison Wesley, 2003. Topic Overview Matrix-Vector Multiplication
Data Clustering. Danushka Bollegala
 Data Clustering Danushka Bollegala Outline Why cluster data? Clustering as unsupervised learning Clustering algorithms k-means, k-medoids agglomerative clustering Brown s clustering Spectral clustering
Data Clustering Danushka Bollegala Outline Why cluster data? Clustering as unsupervised learning Clustering algorithms k-means, k-medoids agglomerative clustering Brown s clustering Spectral clustering
Efficient Semi-supervised Spectral Co-clustering with Constraints
 2010 IEEE International Conference on Data Mining Efficient Semi-supervised Spectral Co-clustering with Constraints Xiaoxiao Shi, Wei Fan, Philip S. Yu Department of Computer Science, University of Illinois
2010 IEEE International Conference on Data Mining Efficient Semi-supervised Spectral Co-clustering with Constraints Xiaoxiao Shi, Wei Fan, Philip S. Yu Department of Computer Science, University of Illinois
The Simplex Algorithm for LP, and an Open Problem
 The Simplex Algorithm for LP, and an Open Problem Linear Programming: General Formulation Inputs: real-valued m x n matrix A, and vectors c in R n and b in R m Output: n-dimensional vector x There is one
The Simplex Algorithm for LP, and an Open Problem Linear Programming: General Formulation Inputs: real-valued m x n matrix A, and vectors c in R n and b in R m Output: n-dimensional vector x There is one
Optimal Movement-Assisted Sensor Deployment and Its Extensions in Wireless Sensor Networks
 Optimal Movement-Assisted Sensor Deployment and Its Extensions in Wireless Sensor Networks Jie Wu and Shuhui Yang Department of Computer Science and Engineering Florida Atlantic University Boca Raton,
Optimal Movement-Assisted Sensor Deployment and Its Extensions in Wireless Sensor Networks Jie Wu and Shuhui Yang Department of Computer Science and Engineering Florida Atlantic University Boca Raton,
Unsupervised and Semi-Supervised Learning
 Unsupervised and Semi-Supervised Learning Nataliya Sokolovska Sorbonne University Paris, France Master 2 BIM January, 25, 2019 Outline Patients Stratification and Methods of Personalized Medicine An application:
Unsupervised and Semi-Supervised Learning Nataliya Sokolovska Sorbonne University Paris, France Master 2 BIM January, 25, 2019 Outline Patients Stratification and Methods of Personalized Medicine An application:
Data Science and Statistics in Research: unlocking the power of your data Session 3.4: Clustering
 Data Science and Statistics in Research: unlocking the power of your data Session 3.4: Clustering 1/ 1 OUTLINE 2/ 1 Overview 3/ 1 CLUSTERING Clustering is a statistical technique which creates groupings
Data Science and Statistics in Research: unlocking the power of your data Session 3.4: Clustering 1/ 1 OUTLINE 2/ 1 Overview 3/ 1 CLUSTERING Clustering is a statistical technique which creates groupings
1 Undirected Vertex Geography UVG
 Geography Start with a chip sitting on a vertex v of a graph or digraph G. A move consists of moving the chip to a neighbouring vertex. In edge geography, moving the chip from x to y deletes the edge (x,
Geography Start with a chip sitting on a vertex v of a graph or digraph G. A move consists of moving the chip to a neighbouring vertex. In edge geography, moving the chip from x to y deletes the edge (x,
2.3 Algorithms Using Map-Reduce
 28 CHAPTER 2. MAP-REDUCE AND THE NEW SOFTWARE STACK one becomes available. The Master must also inform each Reduce task that the location of its input from that Map task has changed. Dealing with a failure
28 CHAPTER 2. MAP-REDUCE AND THE NEW SOFTWARE STACK one becomes available. The Master must also inform each Reduce task that the location of its input from that Map task has changed. Dealing with a failure
Compression, Clustering and Pattern Discovery in Very High Dimensional Discrete-Attribute Datasets
 IEEE TRANSACTIONS ON KNOWLEDGE AND DATA ENGINEERING 1 Compression, Clustering and Pattern Discovery in Very High Dimensional Discrete-Attribute Datasets Mehmet Koyutürk, Ananth Grama, and Naren Ramakrishnan
IEEE TRANSACTIONS ON KNOWLEDGE AND DATA ENGINEERING 1 Compression, Clustering and Pattern Discovery in Very High Dimensional Discrete-Attribute Datasets Mehmet Koyutürk, Ananth Grama, and Naren Ramakrishnan
Visual Representations for Machine Learning
 Visual Representations for Machine Learning Spectral Clustering and Channel Representations Lecture 1 Spectral Clustering: introduction and confusion Michael Felsberg Klas Nordberg The Spectral Clustering
Visual Representations for Machine Learning Spectral Clustering and Channel Representations Lecture 1 Spectral Clustering: introduction and confusion Michael Felsberg Klas Nordberg The Spectral Clustering
ViTraM: VIsualization of TRAnscriptional Modules
 ViTraM: VIsualization of TRAnscriptional Modules Version 1.0 June 1st, 2009 Hong Sun, Karen Lemmens, Tim Van den Bulcke, Kristof Engelen, Bart De Moor and Kathleen Marchal KULeuven, Belgium 1 Contents
ViTraM: VIsualization of TRAnscriptional Modules Version 1.0 June 1st, 2009 Hong Sun, Karen Lemmens, Tim Van den Bulcke, Kristof Engelen, Bart De Moor and Kathleen Marchal KULeuven, Belgium 1 Contents
Gene Clustering & Classification
 BINF, Introduction to Computational Biology Gene Clustering & Classification Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Introduction to Gene Clustering
BINF, Introduction to Computational Biology Gene Clustering & Classification Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Introduction to Gene Clustering
motifs In the context of networks, the term motif may refer to di erent notions. Subgraph motifs Coloured motifs { }
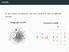 motifs In the context of networks, the term motif may refer to di erent notions. Subgraph motifs Coloured motifs G M { } 2 subgraph motifs 3 motifs Find interesting patterns in a network. 4 motifs Find
motifs In the context of networks, the term motif may refer to di erent notions. Subgraph motifs Coloured motifs G M { } 2 subgraph motifs 3 motifs Find interesting patterns in a network. 4 motifs Find
Rectangular Partitioning
 Rectangular Partitioning Joe Forsmann and Rock Hymas Introduction/Abstract We will look at a problem that I (Rock) had to solve in the course of my work. Given a set of non-overlapping rectangles each
Rectangular Partitioning Joe Forsmann and Rock Hymas Introduction/Abstract We will look at a problem that I (Rock) had to solve in the course of my work. Given a set of non-overlapping rectangles each
Analysis of Biological Networks: Network Modules Identication
 Analysis of Biological Networks: Network Modules Identication Lecturer: Roded Sharan Scribe: Regina Ring and Constantin Radchenko Lecture 4, March 25, 2009 In this lecture we complete the discussion of
Analysis of Biological Networks: Network Modules Identication Lecturer: Roded Sharan Scribe: Regina Ring and Constantin Radchenko Lecture 4, March 25, 2009 In this lecture we complete the discussion of
Performance Analysis of Isoperimetric Algorithm Applied to CT Images
 Performance Analysis of Isoperimetric Algorithm Applied to CT Images Priyanka Kumar Electronics and Telecome. Engg Maharashtra Institute of Technology, Pune, India Dr. Arti Khaparde Electronics and Telecome.
Performance Analysis of Isoperimetric Algorithm Applied to CT Images Priyanka Kumar Electronics and Telecome. Engg Maharashtra Institute of Technology, Pune, India Dr. Arti Khaparde Electronics and Telecome.
CAD Algorithms. Circuit Partitioning
 CAD Algorithms Partitioning Mohammad Tehranipoor ECE Department 13 October 2008 1 Circuit Partitioning Partitioning: The process of decomposing a circuit/system into smaller subcircuits/subsystems, which
CAD Algorithms Partitioning Mohammad Tehranipoor ECE Department 13 October 2008 1 Circuit Partitioning Partitioning: The process of decomposing a circuit/system into smaller subcircuits/subsystems, which
Modularity CMSC 858L
 Modularity CMSC 858L Module-detection for Function Prediction Biological networks generally modular (Hartwell+, 1999) We can try to find the modules within a network. Once we find modules, we can look
Modularity CMSC 858L Module-detection for Function Prediction Biological networks generally modular (Hartwell+, 1999) We can try to find the modules within a network. Once we find modules, we can look
Clustering and Visualisation of Data
 Clustering and Visualisation of Data Hiroshi Shimodaira January-March 28 Cluster analysis aims to partition a data set into meaningful or useful groups, based on distances between data points. In some
Clustering and Visualisation of Data Hiroshi Shimodaira January-March 28 Cluster analysis aims to partition a data set into meaningful or useful groups, based on distances between data points. In some
Advanced Algorithmics (6EAP) Project proposals. Jaak Vilo 2011 Spring
 Advanced Algorithmics (6EAP) Project proposals Jaak Vilo 2011 Spring Jaak Vilo 1 Key info Project = 1-2- 3 person teams Deadline: May 26th Delay: 1 day = 20% of max points Prerequisite for exam ExpectaIons:
Advanced Algorithmics (6EAP) Project proposals Jaak Vilo 2011 Spring Jaak Vilo 1 Key info Project = 1-2- 3 person teams Deadline: May 26th Delay: 1 day = 20% of max points Prerequisite for exam ExpectaIons:
Distance based tree reconstruction. Hierarchical clustering (UPGMA) Neighbor-Joining (NJ)
 Distance based tree reconstruction Hierarchical clustering (UPGMA) Neighbor-Joining (NJ) All organisms have evolved from a common ancestor. Infer the evolutionary tree (tree topology and edge lengths)
Distance based tree reconstruction Hierarchical clustering (UPGMA) Neighbor-Joining (NJ) All organisms have evolved from a common ancestor. Infer the evolutionary tree (tree topology and edge lengths)
