BRB-ArrayTools Data Archive for Human Cancer Gene Expression: A Unique and Efficient Data Sharing Resource
|
|
|
- Alexander Farmer
- 6 years ago
- Views:
Transcription
1 SHORT REPORT BRB-ArrayTools Data Archive for Human Cancer Gene Expression: A Unique and Efficient Data Sharing Resource Yingdong Zhao and Richard Simon Biometric Research Branch, National Cancer Institute, National Institutes of Health, Rockville, Maryland, U.S.A. Abstract: The explosion of available microarray data on human cancer increases the urgency for developing methods for effectively sharing this data among clinical cancer investigators. Lack of a smooth interface between the databases and statistical analysis tools limits the potential benefits of sharing the publicly available microarray data. To facilitate the efficient sharing and use of publicly available microarray data among cancer investigators, we have built a BRB-ArrayTools Data Archive including over one hundred human cancer microarray projects for 28 cancer types. Expression array data and clinical descriptors have been imported into BRB-ArrayTools and are stored as BRB-ArrayTools project folders on the archive. The data archive can be accessed from: Our BRB-ArrayTools data archive and GEO importer represent ongoing efforts to provide effective tools for efficiently sharing and utilizing human cancer microarray data. Background Since its first appearance in the late 1990 s, microarray technology has been applied in almost all areas of biology and medicine. Gene expression experiments have been proven to be very powerful in cancer diagnosis and treatment [1,2]. The explosion of available microarray data on human cancer increases the urgency for developing methods for effectively sharing this data among cancer clinical investigators. As we know, data sharing not only allows the research community to validate original analyses, but also to enable new analyses, design new methodology and perform meta- analysis on data generated from different laboratories [3,4]. There are two major components to data sharing effectiveness. One is ease of access (deposit and download) to the publicly available data. This objective is closely related to the effectiveness of the database as an archive. The second component is how easily and effectively the archived data can be analyzed by the community. Unlike sequence data which is successfully shared by research community via public sequence databases and online search tools, DNA microarray expression data are much more complex. Multiple steps are involved in the data generation, many experimental platforms are widely used and data can be deposited with different levels of processing (for example, log intensity, log ratio, intensity, or probe set level vs. probe level). Microarray data analysis is also statistically complex. Currently, efforts to build publicly available gene expression databases have been successfully as archives only. For example, NCBI s Gene Expression Omnibus (GEO), EBI s ArrayExpress and the Stanford Microarray Database (SMD) support MIAME compliant data submissions and provide online resource for gene expression data browsing, query and retrieval. However, they provide only limited statistical analysis capabilities. Sophisticated statistical analysis software for microarray data analysis are available as stand-alone tools. Users often find it difficult to load the data into external analysis tools because of in-compatibility of the data format. Although more and more stand alone statistical packages have been designed specifically for gene expression data analysis, the user interfaces are not well designed to accommodate publicly available microarray data. Lack of a smooth interface between the databases and statistical analysis tools limits the potential benefits of sharing the publicly available microarray data. BRB-ArrayTools is a comprehensive state-of-the-art statistical analysis system for the analysis of microarray gene expression data [5]. The ArrayTools package is portable and not tied to any database. BRB-ArrayTools contains numerous analysis tools, including SAM, multivariate permutation Correspondence: Richard Simon. rsimon@mail.nih.gov Copyright in this article, its metadata, and any supplementary data is held by its author or authors. It is published under the Creative Commons Attribution By licence. For further information go to:
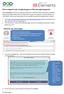
2 Zhao and Simon tests, classification analysis with complete cross-validation, survival prediction analysis and gene set enhancement analysis. It incorporates state-of-the-art statistical methods for analysis of microarray data but the methods are menu driven and the user interface is Excel based and accessible to biomedical scientists. BRB-ArrayTools is computationally very efficient, using compiled modules running transparently to the user. It can handle projects with up to 1000 arrays and 100,000 probe sets per array. It now has over 7000 registered users from about 2000 institutions in 68 countries around the world. Researchers have been using BRB-ArrayTools to do microarray analysis and published over 500 peer-reviewed papers. BRB-ArrayTools has been proven to be a very powerful tool for microarray analysis. To facilitate the efficient sharing and use of publicly available microarray data among cancer investigators, we have built a BRB-ArrayTools Data Archive for Human Cancer Human Gene Expression. Construction and Content Data archive content and infrastructure The data sets included are all from publicly available microarray data resources such as GEO, ArrayExpress and SMD [6 9]. We were not able to collect any datasets from the Oncomine site as it does not support download of raw data [10]. Our data archive currently includes 109 human cancer microarray projects for 28 cancer types, which include more than 6000 microarrays. Each project consists as least 10 samples, along with useful clinical and pathological phenotype information. The citations and the PDF files for the publication of each data set are also listed and linked to PUBMED. There is a README file attached to each record explaining the study purpose and data information. Expression array data and clinical descriptors have been imported into BRB-Array- Tools and are stored as BRB-ArrayTools project folders on the archive. Data access To analyze a dataset, simply download and unzip the file from DataArchive.html The unzipped archive contains a project worksheet which the user opens in Excel on a Microsoft Windows PC on which BRB-ArrayTools has been installed. To download BRB-ArrayTools, go to html Data deposition and management We update the data archive frequently to include new publicly available datasets from NCBI GEO, EBI ArrayExpress and Stanford MicroArray Database. We also encourage BRB-ArrayTools users to deposit their datasets directly. To deposit a dataset, users just need to create a README file, using the sample README file as a template and upload it with their BRB-ArrayTools project folder to brb@linus.nci.nih.gov. For inclusion in the Data Archive we require that the dataset be based on profiling human cancer tissue samples, contain more than 10 samples, and contain clinical phenotype information. Some degree of quality assessment on the data is provided in the process of creating BRB-ArrayTools projects from archived or submitted datasets. Enabling users to subject these datasets to their independent analyses provides some quality checks on the initial publications. Example We chose one lung cancer data set (Beers et al. [11]) as an example to illustrate use of the archive with BRB-ArrayTools to do comprehensive statistical analysis. 1) Click on the zip file link in the data archive web page and save the dataset on your computer. Extract the zip file into a new folder and click on the file named Project.xls to open it in Excel. The project file contains four sheets: experiment descriptors, gene annotations, filtered log intensity and gene identifiers. The experiment descriptors sheet contains clinical information associated with each of the arrays. Each row of the experiment descriptor sheet corresponds to an array and there is a column for each type of clinical descriptor. The descriptors are different for different projects, depending on what information was made available by the authors. For the Beers dataset, the experiment descriptors sheet is shown in Figure 1 and contains age, sex, stage, smoking history, survival time, survival censoring indicator, etc. The filtered log intensity worksheet for the current release of 10
3 BRB ArrayTools data archive for human cancer gene expression BRB-ArrayTools no longer shows the expression data because that data is saved in binary files in the project folder in order to avoid limitations of Excel on numbers of rows and columns. Any desired subset of the expression data can be displayed in the filtered log intensity sheet by clicking on Click to display the data on the top left of the sheet. 2) Data normalization needs to be done before analysis. We have created the projects with raw unnormalized data so that users can normalize as they wish. BRB-ArrayTools provides several normalization options. They are available under the filter and subset the data submenu of the BRB-ArrayTools menu on the Excel menu bar. The menu options available for dual label arrays are somewhat different than for single label arrays like Affymetrix GeneChips TM arrays but in creating the projects we have specified to BRB- ArrayTools what array platform was used. The Beers dataset was based on Affymetrix arrays and for this example we have selected using median over entire array and use median array as reference for normalization (Fig. 1 right side). 3) Suppose the user wants to build a classifier for predicting tumor stage based on gene expression profile. Clicking on the class prediction entry under the BRB-ArrayTools menu (Fig. 2) brings up the class prediction set-up dialog (Fig. 3). STAGE is chosen as Column defining class using the pull down menu. The pull-down menu will show labels for all columns in the experiment descriptors sheet. There are many options possible for class prediction but the default settings have been carefully selected to be useful for many applications. By default the program will develop predictive classifiers using six different methods including the compound covariate method of Radmacher et al. [12], diagonal linear discriminant analysis, nearest neighbor and nearest centroid classification and support vector machines. One part of developing a predictive classifier is selecting the genes to be included and BRB-ArrayTools provides several options for that also. The default is to select genes differentially expressed among classes at a significance level More complex methods Figure 1. Screen shot of experiment descriptor sheet and data normalization menu (right). 11
4 Zhao and Simon Figure 2. Screen shot of select class prediction in BRB-ArrayTools menu. Figure 3. Screen shot of the parameter settings dialog in class prediction function menu. 12
5 BRB ArrayTools data archive for human cancer gene expression like recursive feature elimination are also available. The options page of the class prediction set-up dialog is shown in Figure 4. There are several options for internal validation of the predictive classifier. The default is complete leave-one-out cross-validation. Complete means that for each loop of the cross-validation the informative genes are selected for the restricted training set not containing the omitted case and the six classifiers re-fitted using those genes. If the user wants to repeat the complete cross-validation with the class labels permuted to determine whether the crossvalidated prediction accuracy is better than chance, the box at the top right of the option dialog (Fig. 4) would be checked. Here all of the options can be left at their default values; the user needs only to have given the column defining the classes of interest and possibly the name to be used for the output file that will save the results of the analysis. 4) The class prediction analysis, including complete cross-validation, will usually be complete within a few minutes or less and results will be shown by the program automatically opening an html file containing the output (Fig. 5A and 5B). The first table shows performance of the classifiers during cross-validation for each of the prediction methods. Subsequent tables show cross-validated sensitivity, specificity, positive and negative predictive values for the classifiers and information about the genes included in the classifiers. The genes shown are those based on fitting the classifier to the full dataset, but the column labeled % CV support shows what proportion of the cross-validation loops contained each gene in the classifiers. The genes included in the classifier for the full dataset are sorted by t-value with a link to view the actual expression of significant genes. Parameters for the prediction rules for the linear classifiers are also shown on the html file. The details of the output file for this example can be seen at cancerinfo/example/classprediction.html Discussion We have also built a Gene Expression Omnibus (GEO) import tool for BRB-ArrayTools. GEO currently represents the largest single resource for publicly available gene expression data. This Figure 4. Screen shot of the options page of the class prediction set-up dialog. 13
6 Zhao and Simon A B Figure 5A and 5B. Screen shot of the output html fi le for class prediction results. importer allows the users to automatically import the GDS dataset file in the NCBI Gene Expression Omnibus database into BRB-ArrayTools. A GDS dataset is a text file in SOFT format that represents a curated collection of biologically and statistically comparable GEO Samples reassembled by NCBI staff. Currently there are more than 1500 GDS data sets in GEO database. To import the GEO dataset file into BRB-ArrayTools, the users only need to input the GDS number which can be retrieved from 14
7 BRB ArrayTools data archive for human cancer gene expression NCBI GEO website. The GEO importer therefore serves us as an efficient pipeline to collect the most recent publicly available data in the GEO database. Conclusions In conclusion, our BRB-ArrayTools data archive and GEO importer represent ongoing efforts to provide effective tools for efficiently sharing and utilizing human cancer microarray data. By combining BRB-ArrayTools with the Data Archive, cancer investigators can re-analyze or extend the analysis of publicly available data or combining the publicly available data with their own data set to do external validation analysis. The fact that the Data Archive includes clinical annotations of specimens in an experiment descriptor file ready for use in BRB-ArrayTools makes the resource especially valuable. [6] Edgar, R., Domrachev, M. and Lash, A.E Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res., 30: [7] Barrett, T., Troup, D.B., Wilhite, S.E., Ledoux, P., Rudnev, D., Evangelista, C., Kim, I.F., Soboleva, A., Tomashevsky, M. and Edgar, R NCBI GEO: mining tens of millions of expression profiles database and tools update. Nucleic Acids Res., 35:D760 D765. [8] Parkinson, H. et al ArrayExpress a public database of microarray experiments and gene expression profiles. Nucleic Acids Res., 35:D747 D750. [9] Demeter, J. et al The Stanford Microarray Database: implementation of new analysis tools and open source release of software. Nucleic Acids Res., 35:D766 D770. [10] Rhodes, D.R., Yu, J., Shanker, K., Deshpande, N., Varambally, R., Ghosh, D., Barrette, T., Pandey, A. and Chinnaiyan, A.M ONCOMINE: A Cancer Microarray Database and Data-Mining Platform. Neoplasia, 6:1 6. [11] Beer, D.G. et al Gene-expression profiles predict survival of patients with lung adenocarcinoma. Nat. Med., 8: [12] Radmacher, M.D., McShane, L.M. and Simon, R A paradigm for class prediction using gene expression profiles. Journal of Computational Biology, 9: Availability and Requirements The data archive can be accessed from: nci.nih.gov/~brb/dataarchive.html BRB Array- Tools must have been installed on the machine. Authors Contributions YZ and RS carried out design and implementation of the data archive. YZ and RS drafted the manuscript. Both authors read and approved the final manuscript. Acknowledgements We thank Michael Ngan for programming support in developing GEO importer. We also thank helps from NCBI GEO and EBI ArrayExpress technical support teams. References [1] Sandvik, A.K., Alsberg, B.K., Nørsett, K.G., Yadetie, F., Waldum, H.L. and Laegreid, A Gene expression analysis and clinical diagnosis. Clin. Chim. Acta., 363: [2] Chin, K.V., Alabanza, L., Fujii, K., Kudoh, K., Kita, T., Kikuchi, Y., Selvanayagam, Z.E., Wong, Y.F., Lin, Y. and Shih, W.C. Application of expression genomics for predicting treatment response in cancer. Ann. N. Y. Acad. Sci., 1058: [3] Geschwind, D.H Sharing gene expression data: an array of options. Nature Reviews: Neuroscience, 2: [4] Ball, C.A., Sherlock, G. and Brazma, A Funding high-throughput data sharing. Nature Biotech., 22: [5] Simon, R., Lam, A.P., Li, M.C., Ngan, M., Menenzes, S. and Zhao, Y Analysis of Gene Expression Data Using BRB-Array Tools. Cancer Informatics, 2:
Drug versus Disease (DrugVsDisease) package
 1 Introduction Drug versus Disease (DrugVsDisease) package The Drug versus Disease (DrugVsDisease) package provides a pipeline for the comparison of drug and disease gene expression profiles where negatively
1 Introduction Drug versus Disease (DrugVsDisease) package The Drug versus Disease (DrugVsDisease) package provides a pipeline for the comparison of drug and disease gene expression profiles where negatively
How to store and visualize RNA-seq data
 How to store and visualize RNA-seq data Gabriella Rustici Functional Genomics Group gabry@ebi.ac.uk EBI is an Outstation of the European Molecular Biology Laboratory. Talk summary How do we archive RNA-seq
How to store and visualize RNA-seq data Gabriella Rustici Functional Genomics Group gabry@ebi.ac.uk EBI is an Outstation of the European Molecular Biology Laboratory. Talk summary How do we archive RNA-seq
Tutorial - Analysis of Microarray Data. Microarray Core E Consortium for Functional Glycomics Funded by the NIGMS
 Tutorial - Analysis of Microarray Data Microarray Core E Consortium for Functional Glycomics Funded by the NIGMS Data Analysis introduction Warning: Microarray data analysis is a constantly evolving science.
Tutorial - Analysis of Microarray Data Microarray Core E Consortium for Functional Glycomics Funded by the NIGMS Data Analysis introduction Warning: Microarray data analysis is a constantly evolving science.
Radmacher, M, McShante, L, Simon, R (2002) A paradigm for Class Prediction Using Expression Profiles, J Computational Biol 9:
 Microarray Statistics Module 3: Clustering, comparison, prediction, and Go term analysis Johanna Hardin and Laura Hoopes Worksheet to be handed in the week after discussion Name Clustering algorithms:
Microarray Statistics Module 3: Clustering, comparison, prediction, and Go term analysis Johanna Hardin and Laura Hoopes Worksheet to be handed in the week after discussion Name Clustering algorithms:
Database and R Interfacing for Annotated Microarray Data
 DSC 2003 Working Papers (Draft Versions) http://www.ci.tuwien.ac.at/conferences/dsc-2003/ Database and R Interfacing for Annotated Microarray Data Michael Mader, Werner Mewes GSF Research Center, Institute
DSC 2003 Working Papers (Draft Versions) http://www.ci.tuwien.ac.at/conferences/dsc-2003/ Database and R Interfacing for Annotated Microarray Data Michael Mader, Werner Mewes GSF Research Center, Institute
/ Computational Genomics. Normalization
 10-810 /02-710 Computational Genomics Normalization Genes and Gene Expression Technology Display of Expression Information Yeast cell cycle expression Experiments (over time) baseline expression program
10-810 /02-710 Computational Genomics Normalization Genes and Gene Expression Technology Display of Expression Information Yeast cell cycle expression Experiments (over time) baseline expression program
ArrayExpress and Expression Atlas: Mining Functional Genomics data
 and Expression Atlas: Mining Functional Genomics data Gabriella Rustici, PhD Functional Genomics Team EBI-EMBL gabry@ebi.ac.uk What is functional genomics (FG)? The aim of FG is to understand the function
and Expression Atlas: Mining Functional Genomics data Gabriella Rustici, PhD Functional Genomics Team EBI-EMBL gabry@ebi.ac.uk What is functional genomics (FG)? The aim of FG is to understand the function
PROCEDURE HELP PREPARED BY RYAN MURPHY
 Module on Microarray Statistics for Biochemistry: Metabolomics & Regulation Part 2: Normalization of Microarray Data By Johanna Hardin and Laura Hoopes Instructions and worksheet to be handed in NAME Lecture/Discussion
Module on Microarray Statistics for Biochemistry: Metabolomics & Regulation Part 2: Normalization of Microarray Data By Johanna Hardin and Laura Hoopes Instructions and worksheet to be handed in NAME Lecture/Discussion
Import GEO Experiment into Partek Genomics Suite
 Import GEO Experiment into Partek Genomics Suite This tutorial will illustrate how to: Import a gene expression experiment from GEO SOFT files Specify annotations Import RAW data from GEO for gene expression
Import GEO Experiment into Partek Genomics Suite This tutorial will illustrate how to: Import a gene expression experiment from GEO SOFT files Specify annotations Import RAW data from GEO for gene expression
SEEK User Manual. Introduction
 SEEK User Manual Introduction SEEK is a computational gene co-expression search engine. It utilizes a vast human gene expression compendium to deliver fast, integrative, cross-platform co-expression analyses.
SEEK User Manual Introduction SEEK is a computational gene co-expression search engine. It utilizes a vast human gene expression compendium to deliver fast, integrative, cross-platform co-expression analyses.
Literature Databases
 Literature Databases Introduction to Bioinformatics Dortmund, 16.-20.07.2007 Lectures: Sven Rahmann Exercises: Udo Feldkamp, Michael Wurst 1 Overview 1. Databases 2. Publications in Science 3. PubMed and
Literature Databases Introduction to Bioinformatics Dortmund, 16.-20.07.2007 Lectures: Sven Rahmann Exercises: Udo Feldkamp, Michael Wurst 1 Overview 1. Databases 2. Publications in Science 3. PubMed and
Data Curation Profile Human Genomics
 Data Curation Profile Human Genomics Profile Author Profile Author Institution Name Contact J. Carlson N. Brown Purdue University J. Carlson, jrcarlso@purdue.edu Date of Creation October 27, 2009 Date
Data Curation Profile Human Genomics Profile Author Profile Author Institution Name Contact J. Carlson N. Brown Purdue University J. Carlson, jrcarlso@purdue.edu Date of Creation October 27, 2009 Date
SHARING YOUR RESEARCH DATA VIA
 SHARING YOUR RESEARCH DATA VIA SCHOLARBANK@NUS MEET OUR TEAM Gerrie Kow Head, Scholarly Communication NUS Libraries gerrie@nus.edu.sg Estella Ye Research Data Management Librarian NUS Libraries estella.ye@nus.edu.sg
SHARING YOUR RESEARCH DATA VIA SCHOLARBANK@NUS MEET OUR TEAM Gerrie Kow Head, Scholarly Communication NUS Libraries gerrie@nus.edu.sg Estella Ye Research Data Management Librarian NUS Libraries estella.ye@nus.edu.sg
Enabling Open Science: Data Discoverability, Access and Use. Jo McEntyre Head of Literature Services
 Enabling Open Science: Data Discoverability, Access and Use Jo McEntyre Head of Literature Services www.ebi.ac.uk About EMBL-EBI Part of the European Molecular Biology Laboratory International, non-profit
Enabling Open Science: Data Discoverability, Access and Use Jo McEntyre Head of Literature Services www.ebi.ac.uk About EMBL-EBI Part of the European Molecular Biology Laboratory International, non-profit
Ontrez Project Report National Center for Biomedical Ontology November, 2007
 Ontrez Project Report National Center for Biomedical Ontology November, 2007 Executive summary Currently, genomics data and data repositories in the public domain are expanding at an explosive pace. 1
Ontrez Project Report National Center for Biomedical Ontology November, 2007 Executive summary Currently, genomics data and data repositories in the public domain are expanding at an explosive pace. 1
Feed the Future Innovation Lab for Peanut (Peanut Innovation Lab) Data Management Plan Version:
 Feed the Future Innovation Lab for Peanut (Peanut Innovation Lab) Data Management Plan Version: 20180316 Peanut Innovation Lab Management Entity The University of Georgia, Athens, Georgia Feed the Future
Feed the Future Innovation Lab for Peanut (Peanut Innovation Lab) Data Management Plan Version: 20180316 Peanut Innovation Lab Management Entity The University of Georgia, Athens, Georgia Feed the Future
Gene signature selection to predict survival benefits from adjuvant chemotherapy in NSCLC patients
 1 Gene signature selection to predict survival benefits from adjuvant chemotherapy in NSCLC patients 1,2 Keyue Ding, Ph.D. Nov. 8, 2014 1 NCIC Clinical Trials Group, Kingston, Ontario, Canada 2 Dept. Public
1 Gene signature selection to predict survival benefits from adjuvant chemotherapy in NSCLC patients 1,2 Keyue Ding, Ph.D. Nov. 8, 2014 1 NCIC Clinical Trials Group, Kingston, Ontario, Canada 2 Dept. Public
Nancy Baker 1, Thomas Knudsen 2, Antony Williams 2
 SOFTWARE TOOL ARTICLE Abstract Sifter: a comprehensive front-end system to PubMed [version 1; referees: 2 approved] Nancy Baker 1, Thomas Knudsen 2, Antony Williams 2 1Leidos, Research Triangle Park, NC,
SOFTWARE TOOL ARTICLE Abstract Sifter: a comprehensive front-end system to PubMed [version 1; referees: 2 approved] Nancy Baker 1, Thomas Knudsen 2, Antony Williams 2 1Leidos, Research Triangle Park, NC,
A Novel Hyper-Active Algorithm to Estimate Missing Microarray Attributes
 Computer and Information Science; Vol. 8, No. 3; 2015 ISSN 1913-8989 E-ISSN 1913-8997 Published by Canadian Center of Science and Education A Novel Hyper-Active Algorithm to Estimate Missing Microarray
Computer and Information Science; Vol. 8, No. 3; 2015 ISSN 1913-8989 E-ISSN 1913-8997 Published by Canadian Center of Science and Education A Novel Hyper-Active Algorithm to Estimate Missing Microarray
NIH PUBLIC ACCESS Policy WORKSHOP April
 University of Michigan Deep Blue deepblue.lib.umich.edu 2018-04-18 NIH PUBLIC ACCESS Policy WORKSHOP April Rosenzweig, Merle http://hdl.handle.net/2027.42/143151 Everything you need to know 1 ALL ABOUT
University of Michigan Deep Blue deepblue.lib.umich.edu 2018-04-18 NIH PUBLIC ACCESS Policy WORKSHOP April Rosenzweig, Merle http://hdl.handle.net/2027.42/143151 Everything you need to know 1 ALL ABOUT
HRA Open User Guide for Awardees
 HRA Open User Guide for Awardees HRA Open is an web-based service sponsored by the Health Research Alliance (HRA) that promotes open access by linking your award information with PubMed Central so you
HRA Open User Guide for Awardees HRA Open is an web-based service sponsored by the Health Research Alliance (HRA) that promotes open access by linking your award information with PubMed Central so you
Data management: databases and standards. Practical Bioinformatics
 Data management: databases and standards 1 Motivation to submit your data into microarray database 2 Why would you publish your gene expression data in a standard form? Peer-review: review: If the data
Data management: databases and standards 1 Motivation to submit your data into microarray database 2 Why would you publish your gene expression data in a standard form? Peer-review: review: If the data
FEATURE EXTRACTION TECHNIQUES USING SUPPORT VECTOR MACHINES IN DISEASE PREDICTION
 FEATURE EXTRACTION TECHNIQUES USING SUPPORT VECTOR MACHINES IN DISEASE PREDICTION Sandeep Kaur 1, Dr. Sheetal Kalra 2 1,2 Computer Science Department, Guru Nanak Dev University RC, Jalandhar(India) ABSTRACT
FEATURE EXTRACTION TECHNIQUES USING SUPPORT VECTOR MACHINES IN DISEASE PREDICTION Sandeep Kaur 1, Dr. Sheetal Kalra 2 1,2 Computer Science Department, Guru Nanak Dev University RC, Jalandhar(India) ABSTRACT
Using open access literature to guide full-text query formulation. Heather A. Piwowar and Wendy W. Chapman. Background
 Using open access literature to guide full-text query formulation Heather A. Piwowar and Wendy W. Chapman Background Much scientific knowledge is contained in the details of the full-text biomedical literature.
Using open access literature to guide full-text query formulation Heather A. Piwowar and Wendy W. Chapman Background Much scientific knowledge is contained in the details of the full-text biomedical literature.
US Geo-Explorer User s Guide. Web:
 US Geo-Explorer User s Guide Web: http://usgeoexplorer.org Updated on October 26, 2016 TABLE OF CONTENTS Introduction... 3 1. System Interface... 5 2. Administrative Unit... 7 2.1 Region Selection... 7
US Geo-Explorer User s Guide Web: http://usgeoexplorer.org Updated on October 26, 2016 TABLE OF CONTENTS Introduction... 3 1. System Interface... 5 2. Administrative Unit... 7 2.1 Region Selection... 7
Bayesian Pathway Analysis (BPA) Tutorial
 Bayesian Pathway Analysis (BPA) Tutorial Step by Step to run BPA: 1-) Download latest version of BPAS from BPA website. Unzip it to an appropriate directory. You need to have JAVA Runtime engine and Matlab
Bayesian Pathway Analysis (BPA) Tutorial Step by Step to run BPA: 1-) Download latest version of BPAS from BPA website. Unzip it to an appropriate directory. You need to have JAVA Runtime engine and Matlab
How to use the rbsurv Package
 How to use the rbsurv Package HyungJun Cho, Sukwoo Kim, Soo-heang Eo, and Jaewoo Kang April 30, 2018 Contents 1 Introduction 1 2 Robust likelihood-based survival modeling 2 3 Algorithm 2 4 Example: Glioma
How to use the rbsurv Package HyungJun Cho, Sukwoo Kim, Soo-heang Eo, and Jaewoo Kang April 30, 2018 Contents 1 Introduction 1 2 Robust likelihood-based survival modeling 2 3 Algorithm 2 4 Example: Glioma
Tutorial. RNA-Seq Analysis of Breast Cancer Data. Sample to Insight. November 21, 2017
 RNA-Seq Analysis of Breast Cancer Data November 21, 2017 Sample to Insight QIAGEN Aarhus Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.qiagenbioinformatics.com AdvancedGenomicsSupport@qiagen.com
RNA-Seq Analysis of Breast Cancer Data November 21, 2017 Sample to Insight QIAGEN Aarhus Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.qiagenbioinformatics.com AdvancedGenomicsSupport@qiagen.com
Deposit guide for Nottingham eprints
 Deposit guide for Nottingham eprints Contents 1. Deposit process... 1 1.1. Create a PDF... 2 2. How to place an embargo using eprints... 3 3. How to use an existing eprint as the basis for one with updated
Deposit guide for Nottingham eprints Contents 1. Deposit process... 1 1.1. Create a PDF... 2 2. How to place an embargo using eprints... 3 3. How to use an existing eprint as the basis for one with updated
DATA-SHARING PLAN FOR MOORE FOUNDATION Coral resilience investigated in the field and via a sea anemone model system
 DATA-SHARING PLAN FOR MOORE FOUNDATION Coral resilience investigated in the field and via a sea anemone model system GENERAL PHILOSOPHY (Arthur Grossman, Steve Palumbi, and John Pringle) The three Principal
DATA-SHARING PLAN FOR MOORE FOUNDATION Coral resilience investigated in the field and via a sea anemone model system GENERAL PHILOSOPHY (Arthur Grossman, Steve Palumbi, and John Pringle) The three Principal
Submission Guidelines
 Submission Guidelines Clinical Trial Results invites the submission of phase I, II, and III clinical trials for publication in a brief print format, with full trials results online. We encourage the submission
Submission Guidelines Clinical Trial Results invites the submission of phase I, II, and III clinical trials for publication in a brief print format, with full trials results online. We encourage the submission
Accepted manuscript deposits via Symplectic Elements
 Accepted manuscript deposits via Symplectic Elements Contents Process... 1 Accessing Symplectic... 2 Depositing an Accepted Author Manuscript or OA published article... 3 1. Let s get started... 3 2. Tell
Accepted manuscript deposits via Symplectic Elements Contents Process... 1 Accessing Symplectic... 2 Depositing an Accepted Author Manuscript or OA published article... 3 1. Let s get started... 3 2. Tell
Maximizing Public Data Sources for Sequencing and GWAS
 Maximizing Public Data Sources for Sequencing and GWAS February 4, 2014 G Bryce Christensen Director of Services Questions during the presentation Use the Questions pane in your GoToWebinar window Agenda
Maximizing Public Data Sources for Sequencing and GWAS February 4, 2014 G Bryce Christensen Director of Services Questions during the presentation Use the Questions pane in your GoToWebinar window Agenda
Gene Survey: FAQ. Gene Survey: FAQ Tod Casasent DRAFT
 Gene Survey: FAQ Tod Casasent 2016-02-22-1245 DRAFT 1 What is this document? This document is intended for use by internal and external users of the Gene Survey package, results, and output. This document
Gene Survey: FAQ Tod Casasent 2016-02-22-1245 DRAFT 1 What is this document? This document is intended for use by internal and external users of the Gene Survey package, results, and output. This document
Supplementary Note-- Williams et al The Image Data Resource: A Bioimage Data Integration and Publication Platform
 Supplementary Note-- Williams et al The Image Data Resource: A Bioimage Data Integration and Publication Platform 1. Exploring the IDR This current IDR web user interface (WUI) is based on the open source
Supplementary Note-- Williams et al The Image Data Resource: A Bioimage Data Integration and Publication Platform 1. Exploring the IDR This current IDR web user interface (WUI) is based on the open source
Advancing code and data publication and peer review. Erika Pastrana, PhD Executive Editor, Nature Journals ALPSP_Sept 2018
 Advancing code and data publication and peer review Erika Pastrana, PhD Executive Editor, Nature Journals ALPSP_Sept 2018 2 Code related initiatives at Nature Research 1.1 Nature journals have been industry
Advancing code and data publication and peer review Erika Pastrana, PhD Executive Editor, Nature Journals ALPSP_Sept 2018 2 Code related initiatives at Nature Research 1.1 Nature journals have been industry
Making data publication a first class research output
 Making data publication a first class research output Andrew L. Hufton Managing Editor, Scientific Data https://www.nature.com/sdata/ Helping Researchers Publish, University of Cambridge, Oct 2017 Launched
Making data publication a first class research output Andrew L. Hufton Managing Editor, Scientific Data https://www.nature.com/sdata/ Helping Researchers Publish, University of Cambridge, Oct 2017 Launched
Adding Research Datasets to the UWA Research Repository
 University Library Adding Research Datasets to the UWA Research Repository Guide to Researchers What does UWA mean by Research Datasets? Research Data is defined as facts, observations or experiences on
University Library Adding Research Datasets to the UWA Research Repository Guide to Researchers What does UWA mean by Research Datasets? Research Data is defined as facts, observations or experiences on
DataSTORRE Deposit Guide
 DataSTORRE Deposit Guide Introduction DataStorre is an online digital repository of multi-disciplinary research datasets produced at the University of Stirling. University of Stirling researchers who have
DataSTORRE Deposit Guide Introduction DataStorre is an online digital repository of multi-disciplinary research datasets produced at the University of Stirling. University of Stirling researchers who have
Tutorial:OverRepresentation - OpenTutorials
 Tutorial:OverRepresentation From OpenTutorials Slideshow OverRepresentation (about 12 minutes) (http://opentutorials.rbvi.ucsf.edu/index.php?title=tutorial:overrepresentation& ce_slide=true&ce_style=cytoscape)
Tutorial:OverRepresentation From OpenTutorials Slideshow OverRepresentation (about 12 minutes) (http://opentutorials.rbvi.ucsf.edu/index.php?title=tutorial:overrepresentation& ce_slide=true&ce_style=cytoscape)
Database Repository and Tools
 Database Repository and Tools John Matese May 9, 2008 What is the Repository? Save and exchange retrieved and analyzed datafiles Perform datafile manipulations (averaging and annotations) Run specialized
Database Repository and Tools John Matese May 9, 2008 What is the Repository? Save and exchange retrieved and analyzed datafiles Perform datafile manipulations (averaging and annotations) Run specialized
Noise-based Feature Perturbation as a Selection Method for Microarray Data
 Noise-based Feature Perturbation as a Selection Method for Microarray Data Li Chen 1, Dmitry B. Goldgof 1, Lawrence O. Hall 1, and Steven A. Eschrich 2 1 Department of Computer Science and Engineering
Noise-based Feature Perturbation as a Selection Method for Microarray Data Li Chen 1, Dmitry B. Goldgof 1, Lawrence O. Hall 1, and Steven A. Eschrich 2 1 Department of Computer Science and Engineering
EBP. Accessing the Biomedical Literature for the Best Evidence
 Accessing the Biomedical Literature for the Best Evidence Structuring the search for information and evidence Basic search resources Starting the search EBP Lab / Practice: Simple searches Using PubMed
Accessing the Biomedical Literature for the Best Evidence Structuring the search for information and evidence Basic search resources Starting the search EBP Lab / Practice: Simple searches Using PubMed
Genetic Analysis. Page 1
 Genetic Analysis Page 1 Genetic Analysis Objectives: 1) Set up Case-Control Association analysis and the Basic Genetics Workflow 2) Use JMP tools to interact with and explore results 3) Learn advanced
Genetic Analysis Page 1 Genetic Analysis Objectives: 1) Set up Case-Control Association analysis and the Basic Genetics Workflow 2) Use JMP tools to interact with and explore results 3) Learn advanced
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Curtin JA, Fridlyand J, Kageshita T, et al. Distinct sets of
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Curtin JA, Fridlyand J, Kageshita T, et al. Distinct sets of
Biomedical Digital Libraries
 Biomedical Digital Libraries BioMed Central Resource review database: a review Judy F Burnham* Open Access Address: University of South Alabama Biomedical Library, 316 BLB, Mobile, AL 36688, USA Email:
Biomedical Digital Libraries BioMed Central Resource review database: a review Judy F Burnham* Open Access Address: University of South Alabama Biomedical Library, 316 BLB, Mobile, AL 36688, USA Email:
User guide for GEM-TREND
 User guide for GEM-TREND 1. Requirements for Using GEM-TREND GEM-TREND is implemented as a java applet which can be run in most common browsers and has been test with Internet Explorer 7.0, Internet Explorer
User guide for GEM-TREND 1. Requirements for Using GEM-TREND GEM-TREND is implemented as a java applet which can be run in most common browsers and has been test with Internet Explorer 7.0, Internet Explorer
Powering Knowledge Discovery. Insights from big data with Linguamatics I2E
 Powering Knowledge Discovery Insights from big data with Linguamatics I2E Gain actionable insights from unstructured data The world now generates an overwhelming amount of data, most of it written in natural
Powering Knowledge Discovery Insights from big data with Linguamatics I2E Gain actionable insights from unstructured data The world now generates an overwhelming amount of data, most of it written in natural
Methodology for spot quality evaluation
 Methodology for spot quality evaluation Semi-automatic pipeline in MAIA The general workflow of the semi-automatic pipeline analysis in MAIA is shown in Figure 1A, Manuscript. In Block 1 raw data, i.e..tif
Methodology for spot quality evaluation Semi-automatic pipeline in MAIA The general workflow of the semi-automatic pipeline analysis in MAIA is shown in Figure 1A, Manuscript. In Block 1 raw data, i.e..tif
Introduction to Systems Biology II: Lab
 Introduction to Systems Biology II: Lab Amin Emad NIH BD2K KnowEnG Center of Excellence in Big Data Computing Carl R. Woese Institute for Genomic Biology Department of Computer Science University of Illinois
Introduction to Systems Biology II: Lab Amin Emad NIH BD2K KnowEnG Center of Excellence in Big Data Computing Carl R. Woese Institute for Genomic Biology Department of Computer Science University of Illinois
Interactive comment on Data compilation on the biological response to ocean acidification: an update by Y. Yang et al.
 Earth Syst. Sci. Data Discuss., 8, C469 C477, 2016 www.earth-syst-sci-data-discuss.net/8/c469/2016/ Author(s) 2016. This work is distributed under the Creative Commons Attribute 3.0 License. Open Access
Earth Syst. Sci. Data Discuss., 8, C469 C477, 2016 www.earth-syst-sci-data-discuss.net/8/c469/2016/ Author(s) 2016. This work is distributed under the Creative Commons Attribute 3.0 License. Open Access
AMNH Gerstner Scholars in Bioinformatics & Computational Biology Application Instructions
 PURPOSE AMNH Gerstner Scholars in Bioinformatics & Computational Biology Application Instructions The seeks highly qualified applicants for its Gerstner postdoctoral fellowship program in Bioinformatics
PURPOSE AMNH Gerstner Scholars in Bioinformatics & Computational Biology Application Instructions The seeks highly qualified applicants for its Gerstner postdoctoral fellowship program in Bioinformatics
and the bringing cabig cancer data to the.net Developer and Microsoft Office User Communities
 and the bringing cabig cancer data to the.net Developer and Microsoft Office User Communities http://xl-cabig-client.sourceforge.net/ Robert Macura Tom Macura escience Workshop October 2005 Science Paradigms
and the bringing cabig cancer data to the.net Developer and Microsoft Office User Communities http://xl-cabig-client.sourceforge.net/ Robert Macura Tom Macura escience Workshop October 2005 Science Paradigms
Reproducible Research.. Why we love R & Bioconductor
 Reproducible Research.. Why we love R & Bioconductor Aedín Culhane aedin@jimmy.harvard.edu Boston Bioconductor Course, Oct 24/25 th http://bosbioc.wordpress.com/ My R Course Website http://bcb.dfci.harvard.edu/~aedin/
Reproducible Research.. Why we love R & Bioconductor Aedín Culhane aedin@jimmy.harvard.edu Boston Bioconductor Course, Oct 24/25 th http://bosbioc.wordpress.com/ My R Course Website http://bcb.dfci.harvard.edu/~aedin/
DMPonline / DaMaRO Workshop. Rewley House, Oxford 28 June DataStage. - a simple file management system. David Shotton
 DMPonline / DaMaRO Workshop Rewley House, Oxford 28 June 2013 DataStage - a simple file management system David Shotton Oxford e-research Centre and Department of Zoology University of Oxford, UK david.shotton@zoo.ox.ac.uk
DMPonline / DaMaRO Workshop Rewley House, Oxford 28 June 2013 DataStage - a simple file management system David Shotton Oxford e-research Centre and Department of Zoology University of Oxford, UK david.shotton@zoo.ox.ac.uk
Core Technology Development Team Meeting
 Core Technology Development Team Meeting To hear the meeting, you must call in Toll-free phone number: 1-866-740-1260 Access Code: 2201876 For international call in numbers, please visit: https://www.readytalk.com/account-administration/international-numbers
Core Technology Development Team Meeting To hear the meeting, you must call in Toll-free phone number: 1-866-740-1260 Access Code: 2201876 For international call in numbers, please visit: https://www.readytalk.com/account-administration/international-numbers
warwick.ac.uk/lib-publications
 Original citation: Zhao, Lei, Lim Choi Keung, Sarah Niukyun and Arvanitis, Theodoros N. (2016) A BioPortalbased terminology service for health data interoperability. In: Unifying the Applications and Foundations
Original citation: Zhao, Lei, Lim Choi Keung, Sarah Niukyun and Arvanitis, Theodoros N. (2016) A BioPortalbased terminology service for health data interoperability. In: Unifying the Applications and Foundations
SciVerse ScienceDirect. User Guide. October SciVerse ScienceDirect. Open to accelerate science
 SciVerse ScienceDirect User Guide October 2010 SciVerse ScienceDirect Open to accelerate science Welcome to SciVerse ScienceDirect: How to get the most from your subscription SciVerse ScienceDirect is
SciVerse ScienceDirect User Guide October 2010 SciVerse ScienceDirect Open to accelerate science Welcome to SciVerse ScienceDirect: How to get the most from your subscription SciVerse ScienceDirect is
ClinVar. Jennifer Lee, PhD, NCBI/NLM/NIH ClinVar
 ClinVar What is ClinVar ClinVar is a freely available, central archive for associating observed variation with supporting clinical and experimental evidence for a wide range of disorders. The database
ClinVar What is ClinVar ClinVar is a freely available, central archive for associating observed variation with supporting clinical and experimental evidence for a wide range of disorders. The database
The IEEE Metadata Standard for Supporting Big Data Management
 The IEEE Metadata Standard for Supporting Big Data Management Alex MH Kuo 1,2 (Ph.D) 1 School of Health Information Science University of Victoria, BC, Canada. 2 CEDAR, School of Medicine University of
The IEEE Metadata Standard for Supporting Big Data Management Alex MH Kuo 1,2 (Ph.D) 1 School of Health Information Science University of Victoria, BC, Canada. 2 CEDAR, School of Medicine University of
Categorized software tools: (this page is being updated and links will be restored ASAP. Click on one of the menu links for more information)
 Categorized software tools: (this page is being updated and links will be restored ASAP. Click on one of the menu links for more information) 1 / 5 For array design, fabrication and maintaining a database
Categorized software tools: (this page is being updated and links will be restored ASAP. Click on one of the menu links for more information) 1 / 5 For array design, fabrication and maintaining a database
ISO INTERNATIONAL STANDARD. Health informatics Genomic Sequence Variation Markup Language (GSVML)
 INTERNATIONAL STANDARD ISO 25720 First edition 2009-08-15 Health informatics Genomic Sequence Variation Markup Language (GSVML) Informatique de santé Langage de balisage de la variation de séquence génomique
INTERNATIONAL STANDARD ISO 25720 First edition 2009-08-15 Health informatics Genomic Sequence Variation Markup Language (GSVML) Informatique de santé Langage de balisage de la variation de séquence génomique
SAM-GS. Significance Analysis of Microarray for Gene Sets
 SAM-GS Significance Analysis of Microarray for Gene Sets He Gao, Stacey Fisher, Qi Liu, Sachin Bharadwaj, Shahab Jabbari, Irina Dinu, Yutaka Yasui Last modified on September 5, 2012 Table of Contents 1
SAM-GS Significance Analysis of Microarray for Gene Sets He Gao, Stacey Fisher, Qi Liu, Sachin Bharadwaj, Shahab Jabbari, Irina Dinu, Yutaka Yasui Last modified on September 5, 2012 Table of Contents 1
The Final Updates. Philippe Rocca-Serra Alejandra Gonzalez-Beltran, Susanna-Assunta Sansone, Oxford e-research Centre, University of Oxford, UK
 The Final Updates Supported by the NIH grant 1U24 AI117966-01 to UCSD PI, Co-Investigators at: Philippe Rocca-Serra Alejandra Gonzalez-Beltran, Susanna-Assunta Sansone, Oxford e-research Centre, University
The Final Updates Supported by the NIH grant 1U24 AI117966-01 to UCSD PI, Co-Investigators at: Philippe Rocca-Serra Alejandra Gonzalez-Beltran, Susanna-Assunta Sansone, Oxford e-research Centre, University
Importing and Merging Data Tutorial
 Importing and Merging Data Tutorial Release 1.0 Golden Helix, Inc. February 17, 2012 Contents 1. Overview 2 2. Import Pedigree Data 4 3. Import Phenotypic Data 6 4. Import Genetic Data 8 5. Import and
Importing and Merging Data Tutorial Release 1.0 Golden Helix, Inc. February 17, 2012 Contents 1. Overview 2 2. Import Pedigree Data 4 3. Import Phenotypic Data 6 4. Import Genetic Data 8 5. Import and
Title: Optimized multilayer perceptrons for molecular classification and diagnosis using genomic data
 Supplementary material for Manuscript BIOINF-2005-1602 Title: Optimized multilayer perceptrons for molecular classification and diagnosis using genomic data Appendix A. Testing K-Nearest Neighbor and Support
Supplementary material for Manuscript BIOINF-2005-1602 Title: Optimized multilayer perceptrons for molecular classification and diagnosis using genomic data Appendix A. Testing K-Nearest Neighbor and Support
On Demand Phenotype Ranking through Subspace Clustering
 On Demand Phenotype Ranking through Subspace Clustering Xiang Zhang, Wei Wang Department of Computer Science University of North Carolina at Chapel Hill Chapel Hill, NC 27599, USA {xiang, weiwang}@cs.unc.edu
On Demand Phenotype Ranking through Subspace Clustering Xiang Zhang, Wei Wang Department of Computer Science University of North Carolina at Chapel Hill Chapel Hill, NC 27599, USA {xiang, weiwang}@cs.unc.edu
Blast2GO Teaching Exercises
 Blast2GO Teaching Exercises Ana Conesa and Stefan Götz 2012 BioBam Bioinformatics S.L. Valencia, Spain Contents 1 Annotate 10 sequences with Blast2GO 2 2 Perform a complete annotation process with Blast2GO
Blast2GO Teaching Exercises Ana Conesa and Stefan Götz 2012 BioBam Bioinformatics S.L. Valencia, Spain Contents 1 Annotate 10 sequences with Blast2GO 2 2 Perform a complete annotation process with Blast2GO
National Center for Biotechnology Information and National Institute of Health Manuscript Submission Accounts Set Up
 Puerto Rico Clinical and Translational Research Consortium National Center for Biotechnology Information and National Institute of Health Manuscript Submission Accounts Set Up Prepared by the Evaluation
Puerto Rico Clinical and Translational Research Consortium National Center for Biotechnology Information and National Institute of Health Manuscript Submission Accounts Set Up Prepared by the Evaluation
Academic Editor Tutorial
 Academic Editor Tutorial Contents I. Assignments a. Receiving an invitation b. Responding to an invitation c. Primary review i. Cascading peer review II. Inviting additional reviewers a. Reviewer selection
Academic Editor Tutorial Contents I. Assignments a. Receiving an invitation b. Responding to an invitation c. Primary review i. Cascading peer review II. Inviting additional reviewers a. Reviewer selection
Class Prediction Methods Applied to Microarray Data for Classification
 Class Prediction Methods Applied to Microarray Data for Classification Fatima.S. Shukir The Department of Statistic, Iraqi Commission for Planning and Follow up Directorate Computers and Informatics (ICCI),
Class Prediction Methods Applied to Microarray Data for Classification Fatima.S. Shukir The Department of Statistic, Iraqi Commission for Planning and Follow up Directorate Computers and Informatics (ICCI),
Embracing Semantic Technology for Better Metadata Authoring in Biomedicine
 Embracing Semantic Technology for Better Metadata Authoring in Biomedicine Attila L. Egyedi, Martin J. O Connor, Marcos Martínez-Romero, Debra Willrett, Josef Hardi, John Graybeal, and Mark A. Musen Stanford
Embracing Semantic Technology for Better Metadata Authoring in Biomedicine Attila L. Egyedi, Martin J. O Connor, Marcos Martínez-Romero, Debra Willrett, Josef Hardi, John Graybeal, and Mark A. Musen Stanford
Once the data warehouse is assembled, its customers will likely
 Clinical Data Warehouse Development with Base SAS Software and Common Desktop Tools Patricia L. Gerend, Genentech, Inc., South San Francisco, California ABSTRACT By focusing on the information needed by
Clinical Data Warehouse Development with Base SAS Software and Common Desktop Tools Patricia L. Gerend, Genentech, Inc., South San Francisco, California ABSTRACT By focusing on the information needed by
SETTING UP AN HCS DATA ANALYSIS SYSTEM
 A WHITE PAPER FROM GENEDATA JANUARY 2010 SETTING UP AN HCS DATA ANALYSIS SYSTEM WHY YOU NEED ONE HOW TO CREATE ONE HOW IT WILL HELP HCS MARKET AND DATA ANALYSIS CHALLENGES High Content Screening (HCS)
A WHITE PAPER FROM GENEDATA JANUARY 2010 SETTING UP AN HCS DATA ANALYSIS SYSTEM WHY YOU NEED ONE HOW TO CREATE ONE HOW IT WILL HELP HCS MARKET AND DATA ANALYSIS CHALLENGES High Content Screening (HCS)
Automated Bioinformatics Analysis System on Chip ABASOC. version 1.1
 Automated Bioinformatics Analysis System on Chip ABASOC version 1.1 Phillip Winston Miller, Priyam Patel, Daniel L. Johnson, PhD. University of Tennessee Health Science Center Office of Research Molecular
Automated Bioinformatics Analysis System on Chip ABASOC version 1.1 Phillip Winston Miller, Priyam Patel, Daniel L. Johnson, PhD. University of Tennessee Health Science Center Office of Research Molecular
Introduction to The Cochrane Library
 Introduction to The Cochrane Library What is The Cochrane Library? A collection of databases that include high quality, independent, reliable evidence from Cochrane and other systematic reviews, clinical
Introduction to The Cochrane Library What is The Cochrane Library? A collection of databases that include high quality, independent, reliable evidence from Cochrane and other systematic reviews, clinical
Metadata Ingestion and Processinng
 biomedical and healthcare Data Discovery Index Ecosystem Ingestion and Processinng Jeffrey S. Grethe, Ph.D. 2017 BioCADDIE All Hands Meeting prototype Ingestion Indexing Repositories Ingestion ElasticSearch
biomedical and healthcare Data Discovery Index Ecosystem Ingestion and Processinng Jeffrey S. Grethe, Ph.D. 2017 BioCADDIE All Hands Meeting prototype Ingestion Indexing Repositories Ingestion ElasticSearch
Integrating TCGA Clinical Data Using Metadata-driven Tools and NLP
 Integrating TCGA Clinical Data Using Metadata-driven Tools and NLP Guoqian Jiang, MD, PhD, Mayo Clinic Guergana Savova, PhD, HMS/Boston Children's Hospital Rebecca Jacobson, MD,MS, University of Pittsburgh
Integrating TCGA Clinical Data Using Metadata-driven Tools and NLP Guoqian Jiang, MD, PhD, Mayo Clinic Guergana Savova, PhD, HMS/Boston Children's Hospital Rebecca Jacobson, MD,MS, University of Pittsburgh
SciVerse Scopus. 1. Scopus introduction and content coverage. 2. Scopus in comparison with Web of Science. 3. Basic functionalities of Scopus
 Prepared by: Jawad Sayadi Account Manager, United Kingdom Elsevier BV Radarweg 29 1043 NX Amsterdam The Netherlands J.Sayadi@elsevier.com SciVerse Scopus SciVerse Scopus 1. Scopus introduction and content
Prepared by: Jawad Sayadi Account Manager, United Kingdom Elsevier BV Radarweg 29 1043 NX Amsterdam The Netherlands J.Sayadi@elsevier.com SciVerse Scopus SciVerse Scopus 1. Scopus introduction and content
Introduction/Instructions
 Introduction/Instructions Registries (data banks) and repositories (tissue banks, usually with databases associated) all involve the collection and storage of information and/or biological specimens that
Introduction/Instructions Registries (data banks) and repositories (tissue banks, usually with databases associated) all involve the collection and storage of information and/or biological specimens that
Reproducible Workflows Biomedical Research. P Berlin, Germany
 Reproducible Workflows Biomedical Research P11 2018 Berlin, Germany Contributors Leslie McIntosh Research Data Alliance, U.S., Executive Director Oya Beyan Aachen University, Germany Anthony Juehne RDA,
Reproducible Workflows Biomedical Research P11 2018 Berlin, Germany Contributors Leslie McIntosh Research Data Alliance, U.S., Executive Director Oya Beyan Aachen University, Germany Anthony Juehne RDA,
Core Technology Development Team Meeting
 Core Technology Development Team Meeting To hear the meeting, you must call in Toll-free phone number: 1-866-740-1260 Access Code: 2201876 For international call in numbers, please visit: https://www.readytalk.com/account-administration/international-numbers
Core Technology Development Team Meeting To hear the meeting, you must call in Toll-free phone number: 1-866-740-1260 Access Code: 2201876 For international call in numbers, please visit: https://www.readytalk.com/account-administration/international-numbers
Subject. Dataset. Copy paste feature of the diagram. Importing the dataset. Copy paste feature into the diagram.
 Subject Copy paste feature into the diagram. When we define the data analysis process into Tanagra, it is possible to copy components (or entire branches of components) towards another location into the
Subject Copy paste feature into the diagram. When we define the data analysis process into Tanagra, it is possible to copy components (or entire branches of components) towards another location into the
How to deposit your accepted paper in ORA through Symplectic
 How to deposit your accepted paper in ORA through Symplectic Act on Acceptance: when you ve had a journal article or conference paper accepted for publication, deposit the accepted manuscript 1 into ORA
How to deposit your accepted paper in ORA through Symplectic Act on Acceptance: when you ve had a journal article or conference paper accepted for publication, deposit the accepted manuscript 1 into ORA
IsoGeneGUI Package Vignette
 IsoGeneGUI Package Vignette Setia Pramana, Dan Lin, Ziv Shkedy April 2, 2010 1 Introduction The IsoGene Graphical User Interface (IsoGeneGUI) is a user friendly interface of the IsoGene package which is
IsoGeneGUI Package Vignette Setia Pramana, Dan Lin, Ziv Shkedy April 2, 2010 1 Introduction The IsoGene Graphical User Interface (IsoGeneGUI) is a user friendly interface of the IsoGene package which is
Troubleshooting Guide for Search Settings. Why Are Some of My Papers Not Found? How Does the System Choose My Initial Settings?
 Solution home Elements Getting Started Troubleshooting Guide for Search Settings Modified on: Wed, 30 Mar, 2016 at 12:22 PM The purpose of this document is to assist Elements users in creating and curating
Solution home Elements Getting Started Troubleshooting Guide for Search Settings Modified on: Wed, 30 Mar, 2016 at 12:22 PM The purpose of this document is to assist Elements users in creating and curating
Paper CC06. a seed number for the random number generator, a prime number is recommended
 Paper CC06 %ArrayPerm: A SAS Macro for Permutation Analysis of Microarray Data Deqing Pei, M.S., Wei Liu, M.S., Cheng Cheng, Ph.D. St. Jude Children s Research Hospital, Memphis, TN ABSTRACT Microarray
Paper CC06 %ArrayPerm: A SAS Macro for Permutation Analysis of Microarray Data Deqing Pei, M.S., Wei Liu, M.S., Cheng Cheng, Ph.D. St. Jude Children s Research Hospital, Memphis, TN ABSTRACT Microarray
ClaNC: The Manual (v1.1)
 ClaNC: The Manual (v1.1) Alan R. Dabney June 23, 2008 Contents 1 Installation 3 1.1 The R programming language............................... 3 1.2 X11 with Mac OS X....................................
ClaNC: The Manual (v1.1) Alan R. Dabney June 23, 2008 Contents 1 Installation 3 1.1 The R programming language............................... 3 1.2 X11 with Mac OS X....................................
Gene Set Enrichment Analysis. GSEA User Guide
 Gene Set Enrichment Analysis GSEA User Guide 1 Software Copyright The Broad Institute SOFTWARE COPYRIGHT NOTICE AGREEMENT This software and its documentation are copyright 2009, 2010 by the Broad Institute/Massachusetts
Gene Set Enrichment Analysis GSEA User Guide 1 Software Copyright The Broad Institute SOFTWARE COPYRIGHT NOTICE AGREEMENT This software and its documentation are copyright 2009, 2010 by the Broad Institute/Massachusetts
Introducing the Springer Nature Data Support Services
 Introducing the Springer Nature Data Support Services 1 What motivates researchers to share data? 97% - to accelerate research and its applications 1 96% - increased visibility and discovery of their research
Introducing the Springer Nature Data Support Services 1 What motivates researchers to share data? 97% - to accelerate research and its applications 1 96% - increased visibility and discovery of their research
Prepare input data for CINdex
 1 Introduction Prepare input data for CINdex Genomic instability is known to be a fundamental trait in the development of tumors; and most human tumors exhibit this instability in structural and numerical
1 Introduction Prepare input data for CINdex Genomic instability is known to be a fundamental trait in the development of tumors; and most human tumors exhibit this instability in structural and numerical
Bioinformatics Hubs on the Web
 Bioinformatics Hubs on the Web Take a class The Galter Library teaches a related class called Bioinformatics Hubs on the Web. See our Classes schedule for the next available offering. If this class is
Bioinformatics Hubs on the Web Take a class The Galter Library teaches a related class called Bioinformatics Hubs on the Web. See our Classes schedule for the next available offering. If this class is
Open Access and DOAJ 과편협편집인워크숍 10:50-11:10. 한양대구리병원이현정
 Open Access and DOAJ 2017. 11. 30 과편협편집인워크숍 10:50-11:10 한양대구리병원이현정 caulis98@gmail.com 1 Quality Open Access Free and (almost) unrestricted access Adherence to the BOAI definition No Embargoes No Hybrid
Open Access and DOAJ 2017. 11. 30 과편협편집인워크숍 10:50-11:10 한양대구리병원이현정 caulis98@gmail.com 1 Quality Open Access Free and (almost) unrestricted access Adherence to the BOAI definition No Embargoes No Hybrid
Bioconductor tutorial
 Bioconductor tutorial Adapted by Alex Sanchez from tutorials by (1) Steffen Durinck, Robert Gentleman and Sandrine Dudoit (2) Laurent Gautier (3) Matt Ritchie (4) Jean Yang Outline The Bioconductor Project
Bioconductor tutorial Adapted by Alex Sanchez from tutorials by (1) Steffen Durinck, Robert Gentleman and Sandrine Dudoit (2) Laurent Gautier (3) Matt Ritchie (4) Jean Yang Outline The Bioconductor Project
SciMiner User s Manual
 SciMiner User s Manual Copyright 2008 Junguk Hur. All rights reserved. Bioinformatics Program University of Michigan Ann Arbor, MI 48109, USA Email: juhur@umich.edu Homepage: http://jdrf.neurology.med.umich.edu/sciminer/
SciMiner User s Manual Copyright 2008 Junguk Hur. All rights reserved. Bioinformatics Program University of Michigan Ann Arbor, MI 48109, USA Email: juhur@umich.edu Homepage: http://jdrf.neurology.med.umich.edu/sciminer/
How to deposit your accepted paper in ORA through Symplectic
 How to deposit your accepted paper in ORA through Symplectic Act on Acceptance: when you ve had a journal article or conference paper accepted for publication, deposit the accepted manuscript 1 into ORA
How to deposit your accepted paper in ORA through Symplectic Act on Acceptance: when you ve had a journal article or conference paper accepted for publication, deposit the accepted manuscript 1 into ORA
Building R objects from ArrayExpress datasets
 Building R objects from ArrayExpress datasets Audrey Kauffmann October 30, 2017 1 ArrayExpress database ArrayExpress is a public repository for transcriptomics and related data, which is aimed at storing
Building R objects from ArrayExpress datasets Audrey Kauffmann October 30, 2017 1 ArrayExpress database ArrayExpress is a public repository for transcriptomics and related data, which is aimed at storing
Using metama for differential gene expression analysis from multiple studies
 Using metama for differential gene expression analysis from multiple studies Guillemette Marot and Rémi Bruyère Modified: January 28, 2015. Compiled: January 28, 2015 Abstract This vignette illustrates
Using metama for differential gene expression analysis from multiple studies Guillemette Marot and Rémi Bruyère Modified: January 28, 2015. Compiled: January 28, 2015 Abstract This vignette illustrates
Windows. RNA-Seq Tutorial
 Windows RNA-Seq Tutorial 2017 Gene Codes Corporation Gene Codes Corporation 525 Avis Drive, Ann Arbor, MI 48108 USA 1.800.497.4939 (USA) +1.734.769.7249 (elsewhere) +1.734.769.7074 (fax) www.genecodes.com
Windows RNA-Seq Tutorial 2017 Gene Codes Corporation Gene Codes Corporation 525 Avis Drive, Ann Arbor, MI 48108 USA 1.800.497.4939 (USA) +1.734.769.7249 (elsewhere) +1.734.769.7074 (fax) www.genecodes.com
Queen s University Library. Research Data Management (RDM) Workflow
 Queen s University Library Research Data Management (RDM) Workflow Alexandra Cooper Jeff Moon Data Services, Open Scholarship Services Queen s University Library February 2018 Table of Contents RDM Planning...
Queen s University Library Research Data Management (RDM) Workflow Alexandra Cooper Jeff Moon Data Services, Open Scholarship Services Queen s University Library February 2018 Table of Contents RDM Planning...
