Learning Tensor-Based Features for Whole-Brain fmri Classification
|
|
|
- Allison Townsend
- 5 years ago
- Views:
Transcription
1 Learning Tensor-Based Features for Whole-Brain fmri Classification Xiaonan Song, Lingnan Meng, Qiquan Shi, and Haiping Lu Department of Computer Science, Hong Kong Baptist University Abstract This paper presents a novel tensor-based feature learning approach for whole-brain fmri classification. Whole-brain fmri data have high exploratory power, but they are challenging to deal with due to large numbers of voxels. A critical step for fmri classification is dimensionality reduction, via feature selection or feature extraction. Most current approaches perform voxel selection based on feature selection methods. In contrast, feature extraction methods, such as principal component analysis (PCA), have limited usage on whole brain due to the small sample size problem and limited interpretability. To address these issues, we propose to directly extract features from natural tensor (rather than vector) representations of whole-brain fmri using multilinear PCA (MPCA), and map MPCA bases to voxels for interpretability. Specifically, we extract low-dimensional tensors by MPCA, and then select a number of MPCA features according to the captured variance or mutual information as the input to SVM. To provide interpretability, we construct a mapping from the selected MPCA bases to raw voxels for localizing discriminating regions. Quantitative evaluations on challenging multiclass tasks demonstrate the superior performance of our proposed methods against the state-of-the-art, while qualitative analysis on localized discriminating regions shows the spatial coherence and interpretability of our mapping. 1 Introduction Over the past decades, functional Magnetic Resonance Imaging (fmri) has emerged as a powerful instrument to collect vast quantities of data for measuring brain activities. It becomes a popular tool in applications such as brain state encoding/decoding and brain disease detection, including Alzheimer s disease, Mild Cognitive Impairment, and Autism Spectrum Disorder [2,14]. Most existing studies on fmri classification restrict the analysis to specific brain regions of interest (ROIs). However, ROI analysis is labor-intensive, subject to human error, and requires the assumption that a functionally active brain region will be within an anatomically standardized index [10]. In contrast, whole-brain fmri data have higher exploratory power and lower bias (with no prior user-dependent hypothesis/selection of spatial voxels) [5,17], and recent works reported promising results on whole-brain-based classification [1,16,17]. Inspired by these works, this paper focuses on whole-brain fmri classification. Corresponding author.
2 2 X. Song, L. Meng, Q. Shi, and H. Lu It is challenging to analyze all voxels in the whole brain. The number of whole-brain voxels usually far exceeds the number of observations available in practice, leading to overfitting [17]. We need to perform dimensionality reduction first, through either feature selection or feature extraction [12]. Feature selection methods are more popular for fmri classification, partly due to their good interpretability. There are two main approaches: univariate and multivariate feature selection [14]. For the univariate approach, mutual information (MI) is a popular choice [3,15], e.g., Chou et al. [3] select informative fmri voxels with high MI values individually for brain state decoding and report good improvement in classification accuracy. In contrast, the multivariate methods consider interactions between multiple features, e.g., Ryali et al. [17] and Kampa et al. [7] present sparse optimization frameworks for whole-brain fmri feature selection and demonstrate the effectiveness of logistic regression (LR) with the elastic net penalty, which outperforms LR with l 1 -norm regularization and recursive feature elimination [16], and serves as the state-of-the-art. The other dimensionality reduction approach is feature extraction. Principal component analysis (PCA) is arguably the most popular linear feature extraction method. To apply PCA to whole-brain fmri, we need to concatenate all voxels into a very high-dimensional vector, making the small sample size problem more severe. Moreover, though individual PCA bases can be well interpreted [6], a group of PCA bases are seldom interpreted together effectively [12,13]. On the other hand, multilinear feature extraction methods, such as the multilinear PCA (MPCA) [9], are getting popular recently. They represent multidimensional data as tensors rather than vectors, with three key benefits: preserved multidimensional structure, lower computational demand, and less parameters to estimate. For example, for 3D volumes, a PCA basis needs = 1, 048, 576 parameters, while an MPCA basis needs only = 320 parameters [8]. fmri data are multidimensional so it is more intuitive to analyze them using tensor representations [1,8]. In this paper, we propose a novel tensor-based feature learning approach via MPCA for whole-brain fmri classification and a new mapping scheme to localize discriminating regions based on MPCA features. We perform evaluations on a challenging multiclass fmri dataset [11]. Our contributions are twofold: Our methods directly extract features from tensor representations of fmri using MPCA for the three key benefits mentioned above. The extracted MPCA features are then selected according to variance or mutual information to be fed into the Support Vector Machine (SVM). Superior performance on both binary and multiclass tasks is achieved without requiring vectorization or a priori identification of localized ROIs. Our mapping scheme localizes discriminating regions in the voxel space via MPCA bases for interpretability. It is different from the scheme in [12] of mapping the coefficients of the optimal hyperplane in linear SVM, which tends to be noisy and fragmented, as pointed out in [4]. We can obtain spatial maps with good spatial coherence and good interpretability for neuroscience.
3 Learning Tensor-Based Features for Whole-Brain fmri Classification 3 2 Methods Our proposed methods use the fmri data represented by the mean percent signal change (PSC) over the time dimension [11] as input features and model them directly as third-order tensors (3D data). We use MPCA to learn multilinear bases from these tensorial input to obtain low-dimensional tensorial MPCA features. We then select the most informative MPCA features to form feature vectors for the SVM classifier. We present the key steps of our methods in detail below. Notations and Basic Operations. Following [8], we denote vectors by lowercase boldface letters, e.g., x; matrices by uppercase boldface, e.g., X; and tensors by calligraphic letters, e.g., X. An index is denoted with a lowercase letter, spanning the range from 1 to the uppercase letter of the index, e.g., i = 1,..., I. We denote an Nth-order tensor as A R I1 I N and their elements with indices in parentheses A(i 1,..., i N ). The n-mode index is denoted with i n, n = 1,..., N. The n-mode product of a tensor A by a matrix U R Jn In, is written as B = A n U, with its entries obtained as [8]: B(i 1,..., i n 1, j n, i n+1,..., i N ) = i n A(i 1,..., i N )U(j n, i n ), j n = 1,..., J n. The scalar product of two tensors A, B R I1 I2 I N A, B = i 1 is defined as: (1) i N A(i 1,..., i N )B(i 1,..., i N ). (2) A rank-one tensor U equals to the outer product of N vectors [8]: U = u (1) u (N), where U(i 1,..., i N ) = u (1) (i 1 ) u (N) (i N ). (3) MPCA Feature Extraction. MPCA [9] is an unsupervised learning method to learn features directly from tensorial representations of multidimensional data. Thus, we represent our M training fmri samples as third-order tensors {X 1,..., X M R I1 I2 I3 } as input to MPCA. MPCA then extracts low-dimensional tensor features {Y 1,..., Y M R P1 P2 P3 } through three (N = 3) projection matrices {U (n) R In Pn, n = 1, 2, 3} as follows: Y m = X m 1 U (1)T 2 U (2)T 3 U (3)T, m = 1,..., M, (4) where P n < I n. In this way, the tensor dimensions are reduced from I 1 I 2 I 3 to P 1 P 2 P 3. The solutions for the projection matrices {U (n) } are obtained via maximizing the total tensor scatter Ψ Y = M m=1 Y m Ȳ 2 F, where Ȳ = 1 M M m=1 Y m is the mean tensor feature and F is the Frobenius norm [9]. This problem is solved through an iterative alternating projection method in [9]. Each iteration involves N modewise eigendecompositions to get the n-mode eigenvalues and eigenvectors. We denote the i n th n-mode eigenvalue as λ (n) i n. There are two parameters to set in MPCA. One is Q for determining the tensor subspace dimensions {P 1, P 2, P 3 }. Specifically, the first P n eigenvectors
4 4 X. Song, L. Meng, Q. Shi, and H. Lu Figure 1. Illustration of three selected eigentensors (MPCA features), where each row corresponds to an eigentensor with the third (depth) dimension concatenated. are kept in the n-mode so that the same (or similar) amount of variances is kept in each mode: Q (1) = Q (2) = Q (3) = Q, where Q (n) is the ratio of variances kept in the n-mode defined as Q (n) = P n i n=1 λ(n) i n / I n i n=1 λ(n) i n, and λ (n) i n is the i n th n-mode eigenvalue in the full projection [9]. The second parameter is the maximum number of iterations K, which can be safely set to 1 following [9]. MPCA Feature Selection. The MPCA projection matrices {U (n), n = 1, 2, 3} can be viewed as P 1 P 2 P 3 eigentensors [9] using (3): U p1p 2p 3 = u (1) p 1 u (2) p 2 u (3) p 3 R I1 I2 I3, p n = 1,..., P n, (5) where u (n) p n is the p n th column of U (n). Each eigentensor U p1p 2p 3 is an MPCA feature and it can be mapped to the voxel space. Figure 1 illustrates three eigentensors capturing the most variance of the fmri data studied in this paper. Each eigentensor is shown in a row by concatenating the third dimension. It is rich in structure because it is a rank-one tensor. Since our objective is wholebrain fmri classification, it will be beneficial to select the P most informative (rather than all) features to be fed into a classifier such as the SVM [9]. Therefore, we further perform feature selection based on an importance score using either the variance or the MI criterion. We arrange the entries in {Y m } into feature vectors {y m } according to the importance score in descending order. Only the first P entries of {y m } are selected as SVM input. We can determine the optimal value for P via cross-validation. For convenience, we denote the eigentensor corresponding to the pth selected feature as U p so the pth feature y p can be written as y p = X, U p using (2). The variance is an unsupervised criterion. We obtain the variance S p1p 2p 3 captured by the eigentensor U p1p 2p 3 using a scatter measure as S p1p 2p 3 = M [ Ym (p 1, p 2, p 3 ) Ȳ(p 1, p 2, p 3 ) ] 2. (6) m=1 The MI is a criterion to quantify statistical dependence between two discrete random variables A and B (for example) as [3]: MI(A; B) = p(a, b) p(a, b)log p(a)p(b), (7) a A b B where p(a, b) is the joint probability distribution, and p(a) and p(b) are the marginal probability distribution. In feature selection, we can use MI in a supervised way to measure the relevancy between a feature Y(p 1, p 2, p 3 ) and the class label c. A higher MI indicates a greater dependency or relevancy between them.
5 Learning Tensor-Based Features for Whole-Brain fmri Classification 5 Table 1. Details of the four classification tasks for experimental evaluation. #Class #Sample Semantic categories (classes) Animals (animal+insect) / tools (tool+furniture) [7] Animal/insect/tool/vegetable [7] Animal/insect/tool/vegetable/building/vehicle Animal/insect/tool/vegetable/building/vehicle/buildingpart/clothing Mapping for Interpretability. It is often useful to localize regions in the original voxel space of the brain for interpretation. Good features for classification are expected to be closely related to discriminating regions. Since y p = X, U p, we can view y p as a weighted summation of the voxels in X, where the weights are contained in U p. Therefore, we propose a scheme to map the selected MPCA features (the eigentensors) to the voxel space. We perform a weighted aggregation of the selected eigentensors first and then determine the D most informative voxels to produce a spatial map M by choosing an appropriate threshold T (depending on D): M = P p=1 w p U p > T, where w p is the weight for the pth eigentensor, and denotes the absolute value (magnitude). Note that M is actually a low-rank tensor (rank P ) since it is a summation of P rank-one tensors {U p } [8]. 3 Experiments and Discussions Data. We choose a challenging multiclass dataset, the CMU Science 2008 fmri data (CMU2008) [11]. It aims to predict brain activities associated with the meanings of nouns. The data acquisition experiments had 9 subjects viewing 60 different word-picture stimuli from 12 semantic categories, with 5 exemplars per category and 6 runs per stimulus. The acquisition matrix was with 23 slices, with the numbers of brain voxels for 9 subjects ranging from 19,750 to 21,764. The mean PSC values over time are extracted as input fmri features. 1 Multiclass Tasks. We study four classification tasks: the binary (2-class) and 4-class tasks with the same settings as [7], and two additional, more challenging, 6-class and 8-class tasks. Table 1 summarizes the details. Algorithms. 2 We evaluate seven feature selection/extraction algorithms in Table 2 on the four tasks above: the MI-based univariate feature selection (MI) [3], variance-based univariate feature selection (Var), LR with the elastic net penalty (LR+ENet) for multivariate feature selection [5,7]; PCA-based feature extraction followed by MI-based and variance-based feature selection (PCA- MI and PCA-Var), and the proposed methods of MPCA with MI-based and variance-based feature selection (MPCA-MI and MPCA-Var) We have also tested MPCA alone and MPCA with Lasso-based feature selection. They have similar accuracy as MPCA-MI/Var, but using much more features. In addition, replacing PCA with its popular extension, kernel PCA, gives only slightly better results than PCA-MI/Var.
6 6 X. Song, L. Meng, Q. Shi, and H. Lu Table 2. The classification accuracy in percentage (Acc) and average numbers of features selected (#Features) by five competing methods and the proposed two methods for the four tasks. The top two results in accuracy are highlighted in bold font. Task 2-Class 4-Class 6-Class 8-Class Method Acc #Features Acc #Features Acc #Features Acc #Features MI Var LR+ENet PCA-MI PCA-Var MPCA-MI MPCA-Var Experimental Settings. We follow [7] to arrange testing, validation, and training sets in the format of (1 : 1 : 4) for the six runs in all the experiments. Following [3], we use the SVM classifier with the linear kernel to classify selected features for all methods except LR+ENet which serves as a classifier itself [7,17]. We use the average classification accuracy as the evaluation metric. Algorithm Settings. Parameters for LR+ENet are set following [7] and the number of selected features is determined according to the weight matrix in LR [17]. Other methods use the validation set to determine the number of selected features with the same steps as in [3]. For our MPCA-MI and MPCA-Var methods, we set the parameter Q = 80 in MPCA to report the results. Empirical studies to be shown in Fig. 2(a) show that the classification performance is not sensitive to Q as long as it is not too small (e.g., for Q 70). Classification Accuracy. As shown in Table 2, our proposed methods, MPCA-MI and MPCA-Var achieve the top two overall accuracy (highlighted in bold font), with MPCA-Var achieving the best results on 2-class and 4-class tasks and MPCA-MI achieving the best results on 6-class and 8-class tasks. MPCA- Var and MPCA-MI outperform the state-of-the-art (LR+ENet) by an average of 1.82% and 1.86%, respectively. Though inferior to our methods, LR+ENet indeed outperforms the other existing methods on the whole. In particular, LR+ENet achieves the second best result on 6-class task, slightly better than MPCA-Var. PCA gives the worst results. Number of Selected Features. The number of features selected varies for different methods in Table 2. It fluctuates drastically (from 629 to 3,136) for LR+ENet. It is stable for the Var method but exceeds 5,000. It increases monotonically with the class number for the MI method. PCA can extract only (M 1) features, e.g., M = 150 for the 6-class task. MPCA-Var and MPCA-MI use fewer features in general than MI, Var, and LR+ENet, and the number is relatively stable in contrast. Parameter Sensitivity. Figure 2(a) plots the average accuracy against the Q values for each task. The four curves share a similar trend. The accuracy increases monotonically with Q till Q = 70 and then remains almost constant
7 Classification Accuracy (%) Number of Features Learning Tensor-Based Features for Whole-Brain fmri Classification Class 20 4-Class 6-Class 8-Class Value of Parameter Q (a) MPCA MPCA-MI MPCA-Var Value of Parameter Q (b) Figure 2. Sensitivity against Q: (a) average accuracy of MPCA-Var (MPCA-MI has similar trends), and (b) average number of extracted (for MPCA) and selected (for MPCA-MI/MPCA-Var) features on the 2-class task (other tasks share similar trends). (a) Subject 1 (b) Subject 5 Figure 3. Discriminating regions localized by MPCA-MI for a 2-class task. for Q 70. Thus we choose Q = 80 to report the results. In addition, as shown in Fig. 2(b), the Q value has a greater effect on the number of features, which affects the efficiency in turn. The number of features extracted by MPCA increases almost exponentially with Q, while that by MPCA-MI or MPCA-Var increases with Q at a much slower rate for Q > 60. Mapping and Interpretation. Since raw fmri data are not provided in the CMU2008, we overlay the regions localized by our mapping scheme on a properly scaled and cropped version of the MRI template in mri.mat (Matlab R2013b). We set the weight w p = 1/p to give higher weights to MPCA features with higher importance scores. The regions localized with D = 2000 voxels are highlighted in red in Fig. 3 for the 9th-11th slices of two subjects for MPCA-MI on the 2-class task. The localized regions are spatially coherent and largely consistent between different subjects. Moreover, the localized discriminating regions of Subject 5 have significant overlap with the interpretable regions of the same subject depicted in Fig. 3(B) of [11], indicating good interpretability. 4 Conclusion In this paper, we propose to learn features directly from tensor representations of whole-brain fmri data via MPCA for classification. We use a variance-based or an MI-based criterion to select the most informative MPCA features for SVM classification. In addition, we propose a novel scheme to localize discriminating regions by mapping the selected MPCA features to the raw voxel space. Experimental results on challenging multiclass tasks show that our methods outperform
8 8 X. Song, L. Meng, Q. Shi, and H. Lu the state-of-the-art methods. Furthermore, the proposed mapping scheme can localize discriminating regions that are spatially coherent and consistent cross subjects, with good potential for neuroscience interpretation. Acknowledgments. We thank the support of Hong Kong Research Grants Council (under Grant and the Hong Kong PhD Fellowship Scheme). References 1. Batmanghelich, N., Dong, A., et al.: Regularized tensor factorization for multimodality medical image classification. In: MICCAI. pp (2011) 2. Chen, M., et al.: Survey of encoding and decoding of visual stimulus via fmri: An image analysis perspective. Brain Imaging and Behavior 8(1), 7 23 (2014) 3. Chou, C.A., et al.: Voxel selection framework in multi-voxel pattern analysis of fmri data for prediction of neural response to visual stimuli. IEEE Transactions on Medical Imaging 33(4), (2014) 4. Cuingnet, R., Rosso, C., et al.: Spatially regularized SVM for the detection of brain areas associated with stroke outcome. In: MICCAI, pp (2010) 5. Ecker, C., et al.: Investigating the predictive value of whole-brain structural MR scans in autism: A pattern classification approach. NeuroImage 49(1), (2010) 6. Irimia, A., Van Horn, J.D.: Systematic network lesioning reveals the core white matter scaffold of the human brain. Frontiers in Human Neuroscience 8, 1 14 (2014) 7. Kampa, K., Mehta, S., et al.: Sparse optimization in feature selection: application in neuroimaging. Journal of Global Optimization 59(2-3), (2014) 8. Lu, H., Plataniotis, K.N., Venetsanopoulos, A.: Multilinear Subspace Learning: Dimensionality Reduction of Multidimensional Data. CRC Press (2013) 9. Lu, H., Plataniotis, K.N., et al.: MPCA: Multilinear principal component analysis of tensor objects. IEEE Trans. Neural Networks 19(1), (2008) 10. McKeown, M.J., et al.: Local linear discriminant analysis (LLDA) for group and region of interest (ROI)-based fmri analysis. NeuroImage 37(3), (2007) 11. Mitchell, T.M., Shinkareva, S.V., et al.: Predicting human brain activity associated with the meanings of nouns. Science 320(5880), (2008) 12. Mourão-Miranda, J., et al.: Classifying brain states and determining the discriminating activation patterns: Support vector machine on functional MRI data. NeuroImage 28(4), (2005) 13. Mourão-Miranda, J., et al.: The impact of temporal compression and space selection on SVM analysis of single-subject and multi-subject fmri data. NeuroImage 33(4), (2006) 14. Mwangi, B., Tian, T.S., Soares, J.C.: A review of feature reduction techniques in neuroimaging. Neuroinformatics 12(2), (2013) 15. Rasmussen, P.M., et al.: Model sparsity and brain pattern interpretation of classification models in neuroimaging. Pattern Recognition 45(6), (2012) 16. Retico, A., Bosco, P., et al.: Predictive models based on support vector machines: Whole-brain versus regional analysis of structural MRI in the alzheimer s disease. Journal of Neuroimaging pp (2014) 17. Ryali, S., Supekar, K., Abrams, D.A., Menon, V.: Sparse logistic regression for whole-brain classification of fmri data. NeuroImage 51(2), (2010)
Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging
 1 CS 9 Final Project Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging Feiyu Chen Department of Electrical Engineering ABSTRACT Subject motion is a significant
1 CS 9 Final Project Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging Feiyu Chen Department of Electrical Engineering ABSTRACT Subject motion is a significant
CS 229 Final Project Report Learning to Decode Cognitive States of Rat using Functional Magnetic Resonance Imaging Time Series
 CS 229 Final Project Report Learning to Decode Cognitive States of Rat using Functional Magnetic Resonance Imaging Time Series Jingyuan Chen //Department of Electrical Engineering, cjy2010@stanford.edu//
CS 229 Final Project Report Learning to Decode Cognitive States of Rat using Functional Magnetic Resonance Imaging Time Series Jingyuan Chen //Department of Electrical Engineering, cjy2010@stanford.edu//
Voxel selection algorithms for fmri
 Voxel selection algorithms for fmri Henryk Blasinski December 14, 2012 1 Introduction Functional Magnetic Resonance Imaging (fmri) is a technique to measure and image the Blood- Oxygen Level Dependent
Voxel selection algorithms for fmri Henryk Blasinski December 14, 2012 1 Introduction Functional Magnetic Resonance Imaging (fmri) is a technique to measure and image the Blood- Oxygen Level Dependent
MULTIVARIATE ANALYSES WITH fmri DATA
 MULTIVARIATE ANALYSES WITH fmri DATA Sudhir Shankar Raman Translational Neuromodeling Unit (TNU) Institute for Biomedical Engineering University of Zurich & ETH Zurich Motivation Modelling Concepts Learning
MULTIVARIATE ANALYSES WITH fmri DATA Sudhir Shankar Raman Translational Neuromodeling Unit (TNU) Institute for Biomedical Engineering University of Zurich & ETH Zurich Motivation Modelling Concepts Learning
Feature Selection for fmri Classification
 Feature Selection for fmri Classification Chuang Wu Program of Computational Biology Carnegie Mellon University Pittsburgh, PA 15213 chuangw@andrew.cmu.edu Abstract The functional Magnetic Resonance Imaging
Feature Selection for fmri Classification Chuang Wu Program of Computational Biology Carnegie Mellon University Pittsburgh, PA 15213 chuangw@andrew.cmu.edu Abstract The functional Magnetic Resonance Imaging
Interpreting predictive models in terms of anatomically labelled regions
 Interpreting predictive models in terms of anatomically labelled regions Data Feature extraction/selection Model specification Accuracy and significance Model interpretation 2 Data Feature extraction/selection
Interpreting predictive models in terms of anatomically labelled regions Data Feature extraction/selection Model specification Accuracy and significance Model interpretation 2 Data Feature extraction/selection
Applying Supervised Learning
 Applying Supervised Learning When to Consider Supervised Learning A supervised learning algorithm takes a known set of input data (the training set) and known responses to the data (output), and trains
Applying Supervised Learning When to Consider Supervised Learning A supervised learning algorithm takes a known set of input data (the training set) and known responses to the data (output), and trains
MultiVariate Bayesian (MVB) decoding of brain images
 MultiVariate Bayesian (MVB) decoding of brain images Alexa Morcom Edinburgh SPM course 2015 With thanks to J. Daunizeau, K. Brodersen for slides stimulus behaviour encoding of sensorial or cognitive state?
MultiVariate Bayesian (MVB) decoding of brain images Alexa Morcom Edinburgh SPM course 2015 With thanks to J. Daunizeau, K. Brodersen for slides stimulus behaviour encoding of sensorial or cognitive state?
Multivariate pattern classification
 Multivariate pattern classification Thomas Wolbers Space & Ageing Laboratory (www.sal.mvm.ed.ac.uk) Centre for Cognitive and Neural Systems & Centre for Cognitive Ageing and Cognitive Epidemiology Outline
Multivariate pattern classification Thomas Wolbers Space & Ageing Laboratory (www.sal.mvm.ed.ac.uk) Centre for Cognitive and Neural Systems & Centre for Cognitive Ageing and Cognitive Epidemiology Outline
Multivariate Pattern Classification. Thomas Wolbers Space and Aging Laboratory Centre for Cognitive and Neural Systems
 Multivariate Pattern Classification Thomas Wolbers Space and Aging Laboratory Centre for Cognitive and Neural Systems Outline WHY PATTERN CLASSIFICATION? PROCESSING STREAM PREPROCESSING / FEATURE REDUCTION
Multivariate Pattern Classification Thomas Wolbers Space and Aging Laboratory Centre for Cognitive and Neural Systems Outline WHY PATTERN CLASSIFICATION? PROCESSING STREAM PREPROCESSING / FEATURE REDUCTION
Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques
 Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques Sea Chen Department of Biomedical Engineering Advisors: Dr. Charles A. Bouman and Dr. Mark J. Lowe S. Chen Final Exam October
Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques Sea Chen Department of Biomedical Engineering Advisors: Dr. Charles A. Bouman and Dr. Mark J. Lowe S. Chen Final Exam October
FACE RECOGNITION USING SUPPORT VECTOR MACHINES
 FACE RECOGNITION USING SUPPORT VECTOR MACHINES Ashwin Swaminathan ashwins@umd.edu ENEE633: Statistical and Neural Pattern Recognition Instructor : Prof. Rama Chellappa Project 2, Part (b) 1. INTRODUCTION
FACE RECOGNITION USING SUPPORT VECTOR MACHINES Ashwin Swaminathan ashwins@umd.edu ENEE633: Statistical and Neural Pattern Recognition Instructor : Prof. Rama Chellappa Project 2, Part (b) 1. INTRODUCTION
An efficient face recognition algorithm based on multi-kernel regularization learning
 Acta Technica 61, No. 4A/2016, 75 84 c 2017 Institute of Thermomechanics CAS, v.v.i. An efficient face recognition algorithm based on multi-kernel regularization learning Bi Rongrong 1 Abstract. A novel
Acta Technica 61, No. 4A/2016, 75 84 c 2017 Institute of Thermomechanics CAS, v.v.i. An efficient face recognition algorithm based on multi-kernel regularization learning Bi Rongrong 1 Abstract. A novel
Second revision: Supplementary Material Linking brain-wide multivoxel activation patterns to behaviour: examples from language and math
 Second revision: Supplementary Material Linking brain-wide multivoxel activation patterns to behaviour: examples from language and math Rajeev D. S. Raizada, Feng Ming Tsao, Huei-Mei Liu, Ian D. Holloway,
Second revision: Supplementary Material Linking brain-wide multivoxel activation patterns to behaviour: examples from language and math Rajeev D. S. Raizada, Feng Ming Tsao, Huei-Mei Liu, Ian D. Holloway,
Quick Manual for Sparse Logistic Regression ToolBox ver1.2.1
 Quick Manual for Sparse Logistic Regression ToolBox ver1.2.1 By Okito Yamashita Updated in Feb.2011 First version : Sep.2009 The Sparse Logistic Regression toolbox (SLR toolbox hereafter) is a suite of
Quick Manual for Sparse Logistic Regression ToolBox ver1.2.1 By Okito Yamashita Updated in Feb.2011 First version : Sep.2009 The Sparse Logistic Regression toolbox (SLR toolbox hereafter) is a suite of
CS6375: Machine Learning Gautam Kunapuli. Mid-Term Review
 Gautam Kunapuli Machine Learning Data is identically and independently distributed Goal is to learn a function that maps to Data is generated using an unknown function Learn a hypothesis that minimizes
Gautam Kunapuli Machine Learning Data is identically and independently distributed Goal is to learn a function that maps to Data is generated using an unknown function Learn a hypothesis that minimizes
Preface to the Second Edition. Preface to the First Edition. 1 Introduction 1
 Preface to the Second Edition Preface to the First Edition vii xi 1 Introduction 1 2 Overview of Supervised Learning 9 2.1 Introduction... 9 2.2 Variable Types and Terminology... 9 2.3 Two Simple Approaches
Preface to the Second Edition Preface to the First Edition vii xi 1 Introduction 1 2 Overview of Supervised Learning 9 2.1 Introduction... 9 2.2 Variable Types and Terminology... 9 2.3 Two Simple Approaches
Mapping of Hierarchical Activation in the Visual Cortex Suman Chakravartula, Denise Jones, Guillaume Leseur CS229 Final Project Report. Autumn 2008.
 Mapping of Hierarchical Activation in the Visual Cortex Suman Chakravartula, Denise Jones, Guillaume Leseur CS229 Final Project Report. Autumn 2008. Introduction There is much that is unknown regarding
Mapping of Hierarchical Activation in the Visual Cortex Suman Chakravartula, Denise Jones, Guillaume Leseur CS229 Final Project Report. Autumn 2008. Introduction There is much that is unknown regarding
Pattern Recognition for Neuroimaging Data
 Pattern Recognition for Neuroimaging Data Edinburgh, SPM course April 2013 C. Phillips, Cyclotron Research Centre, ULg, Belgium http://www.cyclotron.ulg.ac.be Overview Introduction Univariate & multivariate
Pattern Recognition for Neuroimaging Data Edinburgh, SPM course April 2013 C. Phillips, Cyclotron Research Centre, ULg, Belgium http://www.cyclotron.ulg.ac.be Overview Introduction Univariate & multivariate
Segmenting Lesions in Multiple Sclerosis Patients James Chen, Jason Su
 Segmenting Lesions in Multiple Sclerosis Patients James Chen, Jason Su Radiologists and researchers spend countless hours tediously segmenting white matter lesions to diagnose and study brain diseases.
Segmenting Lesions in Multiple Sclerosis Patients James Chen, Jason Su Radiologists and researchers spend countless hours tediously segmenting white matter lesions to diagnose and study brain diseases.
Linear Discriminant Analysis for 3D Face Recognition System
 Linear Discriminant Analysis for 3D Face Recognition System 3.1 Introduction Face recognition and verification have been at the top of the research agenda of the computer vision community in recent times.
Linear Discriminant Analysis for 3D Face Recognition System 3.1 Introduction Face recognition and verification have been at the top of the research agenda of the computer vision community in recent times.
Supplementary Figure 1
 Supplementary Figure 1 BOLD and CBV functional maps showing EPI versus line-scanning FLASH fmri. A. Colored BOLD and CBV functional maps are shown in the highlighted window (green frame) of the raw EPI
Supplementary Figure 1 BOLD and CBV functional maps showing EPI versus line-scanning FLASH fmri. A. Colored BOLD and CBV functional maps are shown in the highlighted window (green frame) of the raw EPI
Learning Common Features from fmri Data of Multiple Subjects John Ramish Advised by Prof. Tom Mitchell 8/10/04
 Learning Common Features from fmri Data of Multiple Subjects John Ramish Advised by Prof. Tom Mitchell 8/10/04 Abstract Functional Magnetic Resonance Imaging (fmri), a brain imaging technique, has allowed
Learning Common Features from fmri Data of Multiple Subjects John Ramish Advised by Prof. Tom Mitchell 8/10/04 Abstract Functional Magnetic Resonance Imaging (fmri), a brain imaging technique, has allowed
Supplementary Figure 1. Decoding results broken down for different ROIs
 Supplementary Figure 1 Decoding results broken down for different ROIs Decoding results for areas V1, V2, V3, and V1 V3 combined. (a) Decoded and presented orientations are strongly correlated in areas
Supplementary Figure 1 Decoding results broken down for different ROIs Decoding results for areas V1, V2, V3, and V1 V3 combined. (a) Decoded and presented orientations are strongly correlated in areas
Introduction to Neuroimaging Janaina Mourao-Miranda
 Introduction to Neuroimaging Janaina Mourao-Miranda Neuroimaging techniques have changed the way neuroscientists address questions about functional anatomy, especially in relation to behavior and clinical
Introduction to Neuroimaging Janaina Mourao-Miranda Neuroimaging techniques have changed the way neuroscientists address questions about functional anatomy, especially in relation to behavior and clinical
Statistical Analysis of Metabolomics Data. Xiuxia Du Department of Bioinformatics & Genomics University of North Carolina at Charlotte
 Statistical Analysis of Metabolomics Data Xiuxia Du Department of Bioinformatics & Genomics University of North Carolina at Charlotte Outline Introduction Data pre-treatment 1. Normalization 2. Centering,
Statistical Analysis of Metabolomics Data Xiuxia Du Department of Bioinformatics & Genomics University of North Carolina at Charlotte Outline Introduction Data pre-treatment 1. Normalization 2. Centering,
Cognitive States Detection in fmri Data Analysis using incremental PCA
 Department of Computer Engineering Cognitive States Detection in fmri Data Analysis using incremental PCA Hoang Trong Minh Tuan, Yonggwan Won*, Hyung-Jeong Yang International Conference on Computational
Department of Computer Engineering Cognitive States Detection in fmri Data Analysis using incremental PCA Hoang Trong Minh Tuan, Yonggwan Won*, Hyung-Jeong Yang International Conference on Computational
Large-Scale Lasso and Elastic-Net Regularized Generalized Linear Models
 Large-Scale Lasso and Elastic-Net Regularized Generalized Linear Models DB Tsai Steven Hillion Outline Introduction Linear / Nonlinear Classification Feature Engineering - Polynomial Expansion Big-data
Large-Scale Lasso and Elastic-Net Regularized Generalized Linear Models DB Tsai Steven Hillion Outline Introduction Linear / Nonlinear Classification Feature Engineering - Polynomial Expansion Big-data
Resting state network estimation in individual subjects
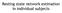 Resting state network estimation in individual subjects Data 3T NIL(21,17,10), Havard-MGH(692) Young adult fmri BOLD Method Machine learning algorithm MLP DR LDA Network image Correlation Spatial Temporal
Resting state network estimation in individual subjects Data 3T NIL(21,17,10), Havard-MGH(692) Young adult fmri BOLD Method Machine learning algorithm MLP DR LDA Network image Correlation Spatial Temporal
Clustering and Visualisation of Data
 Clustering and Visualisation of Data Hiroshi Shimodaira January-March 28 Cluster analysis aims to partition a data set into meaningful or useful groups, based on distances between data points. In some
Clustering and Visualisation of Data Hiroshi Shimodaira January-March 28 Cluster analysis aims to partition a data set into meaningful or useful groups, based on distances between data points. In some
A Comparative Study of Locality Preserving Projection and Principle Component Analysis on Classification Performance Using Logistic Regression
 Journal of Data Analysis and Information Processing, 2016, 4, 55-63 Published Online May 2016 in SciRes. http://www.scirp.org/journal/jdaip http://dx.doi.org/10.4236/jdaip.2016.42005 A Comparative Study
Journal of Data Analysis and Information Processing, 2016, 4, 55-63 Published Online May 2016 in SciRes. http://www.scirp.org/journal/jdaip http://dx.doi.org/10.4236/jdaip.2016.42005 A Comparative Study
Detecting Burnscar from Hyperspectral Imagery via Sparse Representation with Low-Rank Interference
 Detecting Burnscar from Hyperspectral Imagery via Sparse Representation with Low-Rank Interference Minh Dao 1, Xiang Xiang 1, Bulent Ayhan 2, Chiman Kwan 2, Trac D. Tran 1 Johns Hopkins Univeristy, 3400
Detecting Burnscar from Hyperspectral Imagery via Sparse Representation with Low-Rank Interference Minh Dao 1, Xiang Xiang 1, Bulent Ayhan 2, Chiman Kwan 2, Trac D. Tran 1 Johns Hopkins Univeristy, 3400
Toward a rigorous statistical framework for brain mapping
 Toward a rigorous statistical framework for brain mapping Bertrand Thirion, bertrand.thirion@inria.fr 1 Cognitive neuroscience How are cognitive activities affected or controlled by neural circuits in
Toward a rigorous statistical framework for brain mapping Bertrand Thirion, bertrand.thirion@inria.fr 1 Cognitive neuroscience How are cognitive activities affected or controlled by neural circuits in
RADAR DATA CUBE ANALYSIS FOR FALL DETECTION. Baris Erol and Moeness G. Amin
 RADAR DATA CUBE ANALYSIS FOR FALL DETECTION Baris Erol and Moeness G. Amin Center for Advanced Communications, Villanova University, Villanova, PA, 19085, USA ABSTRACT In recent years, radar has been employed
RADAR DATA CUBE ANALYSIS FOR FALL DETECTION Baris Erol and Moeness G. Amin Center for Advanced Communications, Villanova University, Villanova, PA, 19085, USA ABSTRACT In recent years, radar has been employed
Statistical Analysis of Neuroimaging Data. Phebe Kemmer BIOS 516 Sept 24, 2015
 Statistical Analysis of Neuroimaging Data Phebe Kemmer BIOS 516 Sept 24, 2015 Review from last time Structural Imaging modalities MRI, CAT, DTI (diffusion tensor imaging) Functional Imaging modalities
Statistical Analysis of Neuroimaging Data Phebe Kemmer BIOS 516 Sept 24, 2015 Review from last time Structural Imaging modalities MRI, CAT, DTI (diffusion tensor imaging) Functional Imaging modalities
Automatic Generation of Training Data for Brain Tissue Classification from MRI
 MICCAI-2002 1 Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. Cocosco, Alex P. Zijdenbos, and Alan C. Evans McConnell Brain Imaging Centre, Montreal Neurological
MICCAI-2002 1 Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. Cocosco, Alex P. Zijdenbos, and Alan C. Evans McConnell Brain Imaging Centre, Montreal Neurological
Regularized Tensor Factorizations & Higher-Order Principal Components Analysis
 Regularized Tensor Factorizations & Higher-Order Principal Components Analysis Genevera I. Allen Department of Statistics, Rice University, Department of Pediatrics-Neurology, Baylor College of Medicine,
Regularized Tensor Factorizations & Higher-Order Principal Components Analysis Genevera I. Allen Department of Statistics, Rice University, Department of Pediatrics-Neurology, Baylor College of Medicine,
Application of Support Vector Machine Algorithm in Spam Filtering
 Application of Support Vector Machine Algorithm in E-Mail Spam Filtering Julia Bluszcz, Daria Fitisova, Alexander Hamann, Alexey Trifonov, Advisor: Patrick Jähnichen Abstract The problem of spam classification
Application of Support Vector Machine Algorithm in E-Mail Spam Filtering Julia Bluszcz, Daria Fitisova, Alexander Hamann, Alexey Trifonov, Advisor: Patrick Jähnichen Abstract The problem of spam classification
Predict Outcomes and Reveal Relationships in Categorical Data
 PASW Categories 18 Specifications Predict Outcomes and Reveal Relationships in Categorical Data Unleash the full potential of your data through predictive analysis, statistical learning, perceptual mapping,
PASW Categories 18 Specifications Predict Outcomes and Reveal Relationships in Categorical Data Unleash the full potential of your data through predictive analysis, statistical learning, perceptual mapping,
Learning to Recognize Faces in Realistic Conditions
 000 001 002 003 004 005 006 007 008 009 010 011 012 013 014 015 016 017 018 019 020 021 022 023 024 025 026 027 028 029 030 031 032 033 034 035 036 037 038 039 040 041 042 043 044 045 046 047 048 049 050
000 001 002 003 004 005 006 007 008 009 010 011 012 013 014 015 016 017 018 019 020 021 022 023 024 025 026 027 028 029 030 031 032 033 034 035 036 037 038 039 040 041 042 043 044 045 046 047 048 049 050
A NEURAL NETWORK BASED IMAGING SYSTEM FOR fmri ANALYSIS IMPLEMENTING WAVELET METHOD
 6th WSEAS International Conference on CIRCUITS, SYSTEMS, ELECTRONICS,CONTROL & SIGNAL PROCESSING, Cairo, Egypt, Dec 29-31, 2007 454 A NEURAL NETWORK BASED IMAGING SYSTEM FOR fmri ANALYSIS IMPLEMENTING
6th WSEAS International Conference on CIRCUITS, SYSTEMS, ELECTRONICS,CONTROL & SIGNAL PROCESSING, Cairo, Egypt, Dec 29-31, 2007 454 A NEURAL NETWORK BASED IMAGING SYSTEM FOR fmri ANALYSIS IMPLEMENTING
Link Prediction for Social Network
 Link Prediction for Social Network Ning Lin Computer Science and Engineering University of California, San Diego Email: nil016@eng.ucsd.edu Abstract Friendship recommendation has become an important issue
Link Prediction for Social Network Ning Lin Computer Science and Engineering University of California, San Diego Email: nil016@eng.ucsd.edu Abstract Friendship recommendation has become an important issue
A Reduced-Dimension fmri! Shared Response Model
 A Reduced-Dimension fmri! Shared Response Model Po-Hsuan! (Cameron)! Chen 1! Janice! Chen 2! Yaara! Yeshurun 2! Uri! Hasson 2! James! Haxby 3! Peter! Ramadge 1! 1 Department of Electrical Engineering,
A Reduced-Dimension fmri! Shared Response Model Po-Hsuan! (Cameron)! Chen 1! Janice! Chen 2! Yaara! Yeshurun 2! Uri! Hasson 2! James! Haxby 3! Peter! Ramadge 1! 1 Department of Electrical Engineering,
Week 7 Picturing Network. Vahe and Bethany
 Week 7 Picturing Network Vahe and Bethany Freeman (2005) - Graphic Techniques for Exploring Social Network Data The two main goals of analyzing social network data are identification of cohesive groups
Week 7 Picturing Network Vahe and Bethany Freeman (2005) - Graphic Techniques for Exploring Social Network Data The two main goals of analyzing social network data are identification of cohesive groups
Version. Getting Started: An fmri-cpca Tutorial
 Version 11 Getting Started: An fmri-cpca Tutorial 2 Table of Contents Table of Contents... 2 Introduction... 3 Definition of fmri-cpca Data... 3 Purpose of this tutorial... 3 Prerequisites... 4 Used Terms
Version 11 Getting Started: An fmri-cpca Tutorial 2 Table of Contents Table of Contents... 2 Introduction... 3 Definition of fmri-cpca Data... 3 Purpose of this tutorial... 3 Prerequisites... 4 Used Terms
Machine Learning / Jan 27, 2010
 Revisiting Logistic Regression & Naïve Bayes Aarti Singh Machine Learning 10-701/15-781 Jan 27, 2010 Generative and Discriminative Classifiers Training classifiers involves learning a mapping f: X -> Y,
Revisiting Logistic Regression & Naïve Bayes Aarti Singh Machine Learning 10-701/15-781 Jan 27, 2010 Generative and Discriminative Classifiers Training classifiers involves learning a mapping f: X -> Y,
Random projection for non-gaussian mixture models
 Random projection for non-gaussian mixture models Győző Gidófalvi Department of Computer Science and Engineering University of California, San Diego La Jolla, CA 92037 gyozo@cs.ucsd.edu Abstract Recently,
Random projection for non-gaussian mixture models Győző Gidófalvi Department of Computer Science and Engineering University of California, San Diego La Jolla, CA 92037 gyozo@cs.ucsd.edu Abstract Recently,
Predicting the Word from Brain Activity
 CS771A Semester Project 25th November, 2016 Aman Garg Apurv Gupta Ayushya Agarwal Gopichand Kotana Kushal Kumar 14073 14124 14168 14249 14346 Abstract In this project, our objective was to predict the
CS771A Semester Project 25th November, 2016 Aman Garg Apurv Gupta Ayushya Agarwal Gopichand Kotana Kushal Kumar 14073 14124 14168 14249 14346 Abstract In this project, our objective was to predict the
INDEPENDENT COMPONENT ANALYSIS APPLIED TO fmri DATA: A GENERATIVE MODEL FOR VALIDATING RESULTS
 INDEPENDENT COMPONENT ANALYSIS APPLIED TO fmri DATA: A GENERATIVE MODEL FOR VALIDATING RESULTS V. Calhoun 1,2, T. Adali, 2 and G. Pearlson 1 1 Johns Hopkins University Division of Psychiatric Neuro-Imaging,
INDEPENDENT COMPONENT ANALYSIS APPLIED TO fmri DATA: A GENERATIVE MODEL FOR VALIDATING RESULTS V. Calhoun 1,2, T. Adali, 2 and G. Pearlson 1 1 Johns Hopkins University Division of Psychiatric Neuro-Imaging,
Limitations of Matrix Completion via Trace Norm Minimization
 Limitations of Matrix Completion via Trace Norm Minimization ABSTRACT Xiaoxiao Shi Computer Science Department University of Illinois at Chicago xiaoxiao@cs.uic.edu In recent years, compressive sensing
Limitations of Matrix Completion via Trace Norm Minimization ABSTRACT Xiaoxiao Shi Computer Science Department University of Illinois at Chicago xiaoxiao@cs.uic.edu In recent years, compressive sensing
Fast or furious? - User analysis of SF Express Inc
 CS 229 PROJECT, DEC. 2017 1 Fast or furious? - User analysis of SF Express Inc Gege Wen@gegewen, Yiyuan Zhang@yiyuan12, Kezhen Zhao@zkz I. MOTIVATION The motivation of this project is to predict the likelihood
CS 229 PROJECT, DEC. 2017 1 Fast or furious? - User analysis of SF Express Inc Gege Wen@gegewen, Yiyuan Zhang@yiyuan12, Kezhen Zhao@zkz I. MOTIVATION The motivation of this project is to predict the likelihood
CHAPTER 6 MODIFIED FUZZY TECHNIQUES BASED IMAGE SEGMENTATION
 CHAPTER 6 MODIFIED FUZZY TECHNIQUES BASED IMAGE SEGMENTATION 6.1 INTRODUCTION Fuzzy logic based computational techniques are becoming increasingly important in the medical image analysis arena. The significant
CHAPTER 6 MODIFIED FUZZY TECHNIQUES BASED IMAGE SEGMENTATION 6.1 INTRODUCTION Fuzzy logic based computational techniques are becoming increasingly important in the medical image analysis arena. The significant
COSC160: Detection and Classification. Jeremy Bolton, PhD Assistant Teaching Professor
 COSC160: Detection and Classification Jeremy Bolton, PhD Assistant Teaching Professor Outline I. Problem I. Strategies II. Features for training III. Using spatial information? IV. Reducing dimensionality
COSC160: Detection and Classification Jeremy Bolton, PhD Assistant Teaching Professor Outline I. Problem I. Strategies II. Features for training III. Using spatial information? IV. Reducing dimensionality
CIE L*a*b* color model
 CIE L*a*b* color model To further strengthen the correlation between the color model and human perception, we apply the following non-linear transformation: with where (X n,y n,z n ) are the tristimulus
CIE L*a*b* color model To further strengthen the correlation between the color model and human perception, we apply the following non-linear transformation: with where (X n,y n,z n ) are the tristimulus
The Curse of Dimensionality
 The Curse of Dimensionality ACAS 2002 p1/66 Curse of Dimensionality The basic idea of the curse of dimensionality is that high dimensional data is difficult to work with for several reasons: Adding more
The Curse of Dimensionality ACAS 2002 p1/66 Curse of Dimensionality The basic idea of the curse of dimensionality is that high dimensional data is difficult to work with for several reasons: Adding more
Supplementary Data. in residuals voxel time-series exhibiting high variance, for example, large sinuses.
 Supplementary Data Supplementary Materials and Methods Step-by-step description of principal component-orthogonalization technique Below is a step-by-step description of the principal component (PC)-orthogonalization
Supplementary Data Supplementary Materials and Methods Step-by-step description of principal component-orthogonalization technique Below is a step-by-step description of the principal component (PC)-orthogonalization
Linear Discriminant Analysis in Ottoman Alphabet Character Recognition
 Linear Discriminant Analysis in Ottoman Alphabet Character Recognition ZEYNEB KURT, H. IREM TURKMEN, M. ELIF KARSLIGIL Department of Computer Engineering, Yildiz Technical University, 34349 Besiktas /
Linear Discriminant Analysis in Ottoman Alphabet Character Recognition ZEYNEB KURT, H. IREM TURKMEN, M. ELIF KARSLIGIL Department of Computer Engineering, Yildiz Technical University, 34349 Besiktas /
I How does the formulation (5) serve the purpose of the composite parameterization
 Supplemental Material to Identifying Alzheimer s Disease-Related Brain Regions from Multi-Modality Neuroimaging Data using Sparse Composite Linear Discrimination Analysis I How does the formulation (5)
Supplemental Material to Identifying Alzheimer s Disease-Related Brain Regions from Multi-Modality Neuroimaging Data using Sparse Composite Linear Discrimination Analysis I How does the formulation (5)
COMP 551 Applied Machine Learning Lecture 16: Deep Learning
 COMP 551 Applied Machine Learning Lecture 16: Deep Learning Instructor: Ryan Lowe (ryan.lowe@cs.mcgill.ca) Slides mostly by: Class web page: www.cs.mcgill.ca/~hvanho2/comp551 Unless otherwise noted, all
COMP 551 Applied Machine Learning Lecture 16: Deep Learning Instructor: Ryan Lowe (ryan.lowe@cs.mcgill.ca) Slides mostly by: Class web page: www.cs.mcgill.ca/~hvanho2/comp551 Unless otherwise noted, all
MIT Samberg Center Cambridge, MA, USA. May 30 th June 2 nd, by C. Rea, R.S. Granetz MIT Plasma Science and Fusion Center, Cambridge, MA, USA
 Exploratory Machine Learning studies for disruption prediction on DIII-D by C. Rea, R.S. Granetz MIT Plasma Science and Fusion Center, Cambridge, MA, USA Presented at the 2 nd IAEA Technical Meeting on
Exploratory Machine Learning studies for disruption prediction on DIII-D by C. Rea, R.S. Granetz MIT Plasma Science and Fusion Center, Cambridge, MA, USA Presented at the 2 nd IAEA Technical Meeting on
Contents Machine Learning concepts 4 Learning Algorithm 4 Predictive Model (Model) 4 Model, Classification 4 Model, Regression 4 Representation
 Contents Machine Learning concepts 4 Learning Algorithm 4 Predictive Model (Model) 4 Model, Classification 4 Model, Regression 4 Representation Learning 4 Supervised Learning 4 Unsupervised Learning 4
Contents Machine Learning concepts 4 Learning Algorithm 4 Predictive Model (Model) 4 Model, Classification 4 Model, Regression 4 Representation Learning 4 Supervised Learning 4 Unsupervised Learning 4
Variable Selection 6.783, Biomedical Decision Support
 6.783, Biomedical Decision Support (lrosasco@mit.edu) Department of Brain and Cognitive Science- MIT November 2, 2009 About this class Why selecting variables Approaches to variable selection Sparsity-based
6.783, Biomedical Decision Support (lrosasco@mit.edu) Department of Brain and Cognitive Science- MIT November 2, 2009 About this class Why selecting variables Approaches to variable selection Sparsity-based
Improving Positron Emission Tomography Imaging with Machine Learning David Fan-Chung Hsu CS 229 Fall
 Improving Positron Emission Tomography Imaging with Machine Learning David Fan-Chung Hsu (fcdh@stanford.edu), CS 229 Fall 2014-15 1. Introduction and Motivation High- resolution Positron Emission Tomography
Improving Positron Emission Tomography Imaging with Machine Learning David Fan-Chung Hsu (fcdh@stanford.edu), CS 229 Fall 2014-15 1. Introduction and Motivation High- resolution Positron Emission Tomography
Facial Expression Classification with Random Filters Feature Extraction
 Facial Expression Classification with Random Filters Feature Extraction Mengye Ren Facial Monkey mren@cs.toronto.edu Zhi Hao Luo It s Me lzh@cs.toronto.edu I. ABSTRACT In our work, we attempted to tackle
Facial Expression Classification with Random Filters Feature Extraction Mengye Ren Facial Monkey mren@cs.toronto.edu Zhi Hao Luo It s Me lzh@cs.toronto.edu I. ABSTRACT In our work, we attempted to tackle
Lecture on Modeling Tools for Clustering & Regression
 Lecture on Modeling Tools for Clustering & Regression CS 590.21 Analysis and Modeling of Brain Networks Department of Computer Science University of Crete Data Clustering Overview Organizing data into
Lecture on Modeling Tools for Clustering & Regression CS 590.21 Analysis and Modeling of Brain Networks Department of Computer Science University of Crete Data Clustering Overview Organizing data into
Context-sensitive Classification Forests for Segmentation of Brain Tumor Tissues
 Context-sensitive Classification Forests for Segmentation of Brain Tumor Tissues D. Zikic, B. Glocker, E. Konukoglu, J. Shotton, A. Criminisi, D. H. Ye, C. Demiralp 3, O. M. Thomas 4,5, T. Das 4, R. Jena
Context-sensitive Classification Forests for Segmentation of Brain Tumor Tissues D. Zikic, B. Glocker, E. Konukoglu, J. Shotton, A. Criminisi, D. H. Ye, C. Demiralp 3, O. M. Thomas 4,5, T. Das 4, R. Jena
An independent component analysis based tool for exploring functional connections in the brain
 An independent component analysis based tool for exploring functional connections in the brain S. M. Rolfe a, L. Finney b, R. F. Tungaraza b, J. Guan b, L.G. Shapiro b, J. F. Brinkely b, A. Poliakov c,
An independent component analysis based tool for exploring functional connections in the brain S. M. Rolfe a, L. Finney b, R. F. Tungaraza b, J. Guan b, L.G. Shapiro b, J. F. Brinkely b, A. Poliakov c,
LOCAL APPEARANCE BASED FACE RECOGNITION USING DISCRETE COSINE TRANSFORM
 LOCAL APPEARANCE BASED FACE RECOGNITION USING DISCRETE COSINE TRANSFORM Hazim Kemal Ekenel, Rainer Stiefelhagen Interactive Systems Labs, University of Karlsruhe Am Fasanengarten 5, 76131, Karlsruhe, Germany
LOCAL APPEARANCE BASED FACE RECOGNITION USING DISCRETE COSINE TRANSFORM Hazim Kemal Ekenel, Rainer Stiefelhagen Interactive Systems Labs, University of Karlsruhe Am Fasanengarten 5, 76131, Karlsruhe, Germany
Multi-voxel pattern analysis: Decoding Mental States from fmri Activity Patterns
 Multi-voxel pattern analysis: Decoding Mental States from fmri Activity Patterns Artwork by Leon Zernitsky Jesse Rissman NITP Summer Program 2012 Part 1 of 2 Goals of Multi-voxel Pattern Analysis Decoding
Multi-voxel pattern analysis: Decoding Mental States from fmri Activity Patterns Artwork by Leon Zernitsky Jesse Rissman NITP Summer Program 2012 Part 1 of 2 Goals of Multi-voxel Pattern Analysis Decoding
Dimension Reduction for Big Data Analysis. Dan Shen. Department of Mathematics & Statistics University of South Florida.
 Dimension Reduction for Big Data Analysis Dan Shen Department of Mathematics & Statistics University of South Florida danshen@usf.edu October 24, 2014 1 Outline Multiscale weighted PCA for Image Analysis
Dimension Reduction for Big Data Analysis Dan Shen Department of Mathematics & Statistics University of South Florida danshen@usf.edu October 24, 2014 1 Outline Multiscale weighted PCA for Image Analysis
Facial Expression Recognition Using Non-negative Matrix Factorization
 Facial Expression Recognition Using Non-negative Matrix Factorization Symeon Nikitidis, Anastasios Tefas and Ioannis Pitas Artificial Intelligence & Information Analysis Lab Department of Informatics Aristotle,
Facial Expression Recognition Using Non-negative Matrix Factorization Symeon Nikitidis, Anastasios Tefas and Ioannis Pitas Artificial Intelligence & Information Analysis Lab Department of Informatics Aristotle,
Learning from High Dimensional fmri Data using Random Projections
 Learning from High Dimensional fmri Data using Random Projections Author: Madhu Advani December 16, 011 Introduction The term the Curse of Dimensionality refers to the difficulty of organizing and applying
Learning from High Dimensional fmri Data using Random Projections Author: Madhu Advani December 16, 011 Introduction The term the Curse of Dimensionality refers to the difficulty of organizing and applying
Multiresponse Sparse Regression with Application to Multidimensional Scaling
 Multiresponse Sparse Regression with Application to Multidimensional Scaling Timo Similä and Jarkko Tikka Helsinki University of Technology, Laboratory of Computer and Information Science P.O. Box 54,
Multiresponse Sparse Regression with Application to Multidimensional Scaling Timo Similä and Jarkko Tikka Helsinki University of Technology, Laboratory of Computer and Information Science P.O. Box 54,
SGN (4 cr) Chapter 10
 SGN-41006 (4 cr) Chapter 10 Feature Selection and Extraction Jussi Tohka & Jari Niemi Department of Signal Processing Tampere University of Technology February 18, 2014 J. Tohka & J. Niemi (TUT-SGN) SGN-41006
SGN-41006 (4 cr) Chapter 10 Feature Selection and Extraction Jussi Tohka & Jari Niemi Department of Signal Processing Tampere University of Technology February 18, 2014 J. Tohka & J. Niemi (TUT-SGN) SGN-41006
Feature Selection Using Principal Feature Analysis
 Feature Selection Using Principal Feature Analysis Ira Cohen Qi Tian Xiang Sean Zhou Thomas S. Huang Beckman Institute for Advanced Science and Technology University of Illinois at Urbana-Champaign Urbana,
Feature Selection Using Principal Feature Analysis Ira Cohen Qi Tian Xiang Sean Zhou Thomas S. Huang Beckman Institute for Advanced Science and Technology University of Illinois at Urbana-Champaign Urbana,
A New Multi Fractal Dimension Method for Face Recognition with Fewer Features under Expression Variations
 A New Multi Fractal Dimension Method for Face Recognition with Fewer Features under Expression Variations Maksud Ahamad Assistant Professor, Computer Science & Engineering Department, Ideal Institute of
A New Multi Fractal Dimension Method for Face Recognition with Fewer Features under Expression Variations Maksud Ahamad Assistant Professor, Computer Science & Engineering Department, Ideal Institute of
Weighted-CS for reconstruction of highly under-sampled dynamic MRI sequences
 Weighted- for reconstruction of highly under-sampled dynamic MRI sequences Dornoosh Zonoobi and Ashraf A. Kassim Dept. Electrical and Computer Engineering National University of Singapore, Singapore E-mail:
Weighted- for reconstruction of highly under-sampled dynamic MRI sequences Dornoosh Zonoobi and Ashraf A. Kassim Dept. Electrical and Computer Engineering National University of Singapore, Singapore E-mail:
Feature Selection. CE-725: Statistical Pattern Recognition Sharif University of Technology Spring Soleymani
 Feature Selection CE-725: Statistical Pattern Recognition Sharif University of Technology Spring 2013 Soleymani Outline Dimensionality reduction Feature selection vs. feature extraction Filter univariate
Feature Selection CE-725: Statistical Pattern Recognition Sharif University of Technology Spring 2013 Soleymani Outline Dimensionality reduction Feature selection vs. feature extraction Filter univariate
D.A. Karras 1, S.A. Karkanis 2, D. Iakovidis 2, D. E. Maroulis 2 and B.G. Mertzios , Greece,
 NIMIA-SC2001-2001 NATO Advanced Study Institute on Neural Networks for Instrumentation, Measurement, and Related Industrial Applications: Study Cases Crema, Italy, 9-20 October 2001 Support Vector Machines
NIMIA-SC2001-2001 NATO Advanced Study Institute on Neural Networks for Instrumentation, Measurement, and Related Industrial Applications: Study Cases Crema, Italy, 9-20 October 2001 Support Vector Machines
CS229 Project: Classification of Motor Tasks Based on Functional Neuroimaging
 CS229 Project: Classification of Motor Tasks Based on Functional Neuroimaging Gerald Brantner Mechanical Engineering, Stanford University, Stanford, CA 9435, USA. geraldb@stanford.edu Georg Schorpp Management
CS229 Project: Classification of Motor Tasks Based on Functional Neuroimaging Gerald Brantner Mechanical Engineering, Stanford University, Stanford, CA 9435, USA. geraldb@stanford.edu Georg Schorpp Management
Chapter 3 Set Redundancy in Magnetic Resonance Brain Images
 16 Chapter 3 Set Redundancy in Magnetic Resonance Brain Images 3.1 MRI (magnetic resonance imaging) MRI is a technique of measuring physical structure within the human anatomy. Our proposed research focuses
16 Chapter 3 Set Redundancy in Magnetic Resonance Brain Images 3.1 MRI (magnetic resonance imaging) MRI is a technique of measuring physical structure within the human anatomy. Our proposed research focuses
Basic fmri Design and Analysis. Preprocessing
 Basic fmri Design and Analysis Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial filtering
Basic fmri Design and Analysis Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial filtering
Classification of Whole Brain fmri Activation Patterns. Serdar Kemal Balcı
 Classification of Whole Brain fmri Activation Patterns by Serdar Kemal Balcı Submitted to the Department of Electrical Engineering and Computer Science in partial fulfillment of the requirements for the
Classification of Whole Brain fmri Activation Patterns by Serdar Kemal Balcı Submitted to the Department of Electrical Engineering and Computer Science in partial fulfillment of the requirements for the
GENDER CLASSIFICATION USING SUPPORT VECTOR MACHINES
 GENDER CLASSIFICATION USING SUPPORT VECTOR MACHINES Ashwin Swaminathan ashwins@umd.edu ENEE633: Statistical and Neural Pattern Recognition Instructor : Prof. Rama Chellappa Project 2, Part (a) 1. INTRODUCTION
GENDER CLASSIFICATION USING SUPPORT VECTOR MACHINES Ashwin Swaminathan ashwins@umd.edu ENEE633: Statistical and Neural Pattern Recognition Instructor : Prof. Rama Chellappa Project 2, Part (a) 1. INTRODUCTION
Cluster Analysis. Mu-Chun Su. Department of Computer Science and Information Engineering National Central University 2003/3/11 1
 Cluster Analysis Mu-Chun Su Department of Computer Science and Information Engineering National Central University 2003/3/11 1 Introduction Cluster analysis is the formal study of algorithms and methods
Cluster Analysis Mu-Chun Su Department of Computer Science and Information Engineering National Central University 2003/3/11 1 Introduction Cluster analysis is the formal study of algorithms and methods
CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS
 CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS CHAPTER 4 CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS 4.1 Introduction Optical character recognition is one of
CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS CHAPTER 4 CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS 4.1 Introduction Optical character recognition is one of
Segmentation of MR Images of a Beating Heart
 Segmentation of MR Images of a Beating Heart Avinash Ravichandran Abstract Heart Arrhythmia is currently treated using invasive procedures. In order use to non invasive procedures accurate imaging modalities
Segmentation of MR Images of a Beating Heart Avinash Ravichandran Abstract Heart Arrhythmia is currently treated using invasive procedures. In order use to non invasive procedures accurate imaging modalities
IMPROVED FACE RECOGNITION USING ICP TECHNIQUES INCAMERA SURVEILLANCE SYSTEMS. Kirthiga, M.E-Communication system, PREC, Thanjavur
 IMPROVED FACE RECOGNITION USING ICP TECHNIQUES INCAMERA SURVEILLANCE SYSTEMS Kirthiga, M.E-Communication system, PREC, Thanjavur R.Kannan,Assistant professor,prec Abstract: Face Recognition is important
IMPROVED FACE RECOGNITION USING ICP TECHNIQUES INCAMERA SURVEILLANCE SYSTEMS Kirthiga, M.E-Communication system, PREC, Thanjavur R.Kannan,Assistant professor,prec Abstract: Face Recognition is important
Rapid growth of massive datasets
 Overview Rapid growth of massive datasets E.g., Online activity, Science, Sensor networks Data Distributed Clusters are Pervasive Data Distributed Computing Mature Methods for Common Problems e.g., classification,
Overview Rapid growth of massive datasets E.g., Online activity, Science, Sensor networks Data Distributed Clusters are Pervasive Data Distributed Computing Mature Methods for Common Problems e.g., classification,
Evaluation Metrics. (Classifiers) CS229 Section Anand Avati
 Evaluation Metrics (Classifiers) CS Section Anand Avati Topics Why? Binary classifiers Metrics Rank view Thresholding Confusion Matrix Point metrics: Accuracy, Precision, Recall / Sensitivity, Specificity,
Evaluation Metrics (Classifiers) CS Section Anand Avati Topics Why? Binary classifiers Metrics Rank view Thresholding Confusion Matrix Point metrics: Accuracy, Precision, Recall / Sensitivity, Specificity,
Basic Introduction to Data Analysis. Block Design Demonstration. Robert Savoy
 Basic Introduction to Data Analysis Block Design Demonstration Robert Savoy Sample Block Design Experiment Demonstration Use of Visual and Motor Task Separability of Responses Combined Visual and Motor
Basic Introduction to Data Analysis Block Design Demonstration Robert Savoy Sample Block Design Experiment Demonstration Use of Visual and Motor Task Separability of Responses Combined Visual and Motor
A Sparse and Locally Shift Invariant Feature Extractor Applied to Document Images
 A Sparse and Locally Shift Invariant Feature Extractor Applied to Document Images Marc Aurelio Ranzato Yann LeCun Courant Institute of Mathematical Sciences New York University - New York, NY 10003 Abstract
A Sparse and Locally Shift Invariant Feature Extractor Applied to Document Images Marc Aurelio Ranzato Yann LeCun Courant Institute of Mathematical Sciences New York University - New York, NY 10003 Abstract
Forward Feature Selection Using Residual Mutual Information
 Forward Feature Selection Using Residual Mutual Information Erik Schaffernicht, Christoph Möller, Klaus Debes and Horst-Michael Gross Ilmenau University of Technology - Neuroinformatics and Cognitive Robotics
Forward Feature Selection Using Residual Mutual Information Erik Schaffernicht, Christoph Möller, Klaus Debes and Horst-Michael Gross Ilmenau University of Technology - Neuroinformatics and Cognitive Robotics
Learning Algorithms for Medical Image Analysis. Matteo Santoro slipguru
 Learning Algorithms for Medical Image Analysis Matteo Santoro slipguru santoro@disi.unige.it June 8, 2010 Outline 1. learning-based strategies for quantitative image analysis 2. automatic annotation of
Learning Algorithms for Medical Image Analysis Matteo Santoro slipguru santoro@disi.unige.it June 8, 2010 Outline 1. learning-based strategies for quantitative image analysis 2. automatic annotation of
Dimensionality Reduction of fmri Data using PCA
 Dimensionality Reduction of fmri Data using PCA Elio Andia ECE-699 Learning from Data George Mason University Fairfax VA, USA INTRODUCTION The aim of this project was to implement PCA as a dimensionality
Dimensionality Reduction of fmri Data using PCA Elio Andia ECE-699 Learning from Data George Mason University Fairfax VA, USA INTRODUCTION The aim of this project was to implement PCA as a dimensionality
Robust Face Recognition via Sparse Representation
 Robust Face Recognition via Sparse Representation Panqu Wang Department of Electrical and Computer Engineering University of California, San Diego La Jolla, CA 92092 pawang@ucsd.edu Can Xu Department of
Robust Face Recognition via Sparse Representation Panqu Wang Department of Electrical and Computer Engineering University of California, San Diego La Jolla, CA 92092 pawang@ucsd.edu Can Xu Department of
Introduction to Pattern Recognition Part II. Selim Aksoy Bilkent University Department of Computer Engineering
 Introduction to Pattern Recognition Part II Selim Aksoy Bilkent University Department of Computer Engineering saksoy@cs.bilkent.edu.tr RETINA Pattern Recognition Tutorial, Summer 2005 Overview Statistical
Introduction to Pattern Recognition Part II Selim Aksoy Bilkent University Department of Computer Engineering saksoy@cs.bilkent.edu.tr RETINA Pattern Recognition Tutorial, Summer 2005 Overview Statistical
10/14/2017. Dejan Sarka. Anomaly Detection. Sponsors
 Dejan Sarka Anomaly Detection Sponsors About me SQL Server MVP (17 years) and MCT (20 years) 25 years working with SQL Server Authoring 16 th book Authoring many courses, articles Agenda Introduction Simple
Dejan Sarka Anomaly Detection Sponsors About me SQL Server MVP (17 years) and MCT (20 years) 25 years working with SQL Server Authoring 16 th book Authoring many courses, articles Agenda Introduction Simple
Feature Selection Using Modified-MCA Based Scoring Metric for Classification
 2011 International Conference on Information Communication and Management IPCSIT vol.16 (2011) (2011) IACSIT Press, Singapore Feature Selection Using Modified-MCA Based Scoring Metric for Classification
2011 International Conference on Information Communication and Management IPCSIT vol.16 (2011) (2011) IACSIT Press, Singapore Feature Selection Using Modified-MCA Based Scoring Metric for Classification
CPSC 340: Machine Learning and Data Mining. Principal Component Analysis Fall 2016
 CPSC 340: Machine Learning and Data Mining Principal Component Analysis Fall 2016 A2/Midterm: Admin Grades/solutions will be posted after class. Assignment 4: Posted, due November 14. Extra office hours:
CPSC 340: Machine Learning and Data Mining Principal Component Analysis Fall 2016 A2/Midterm: Admin Grades/solutions will be posted after class. Assignment 4: Posted, due November 14. Extra office hours:
