matter: Supplementary 2-3D mass spectrometry
|
|
|
- Jasmin Kelly
- 6 years ago
- Views:
Transcription
1 matter: Supplementar 2-3D mass spectrometr imaging case stud Klie A. Bemis June 13, 2018 Contents 1 Introduction Analing large 3D MSI eperiments with Cardinal and matter 1 3 Comparisons with alternative approaches Using bigmemor Using ff Evaluating performance between approaches An R script for comparing performance Session info Introduction The first half of this vignette demonstrates the usefulness of matter for working with large mass spectrometr imaging (MSI) eperiments in Cardinal. Cardinal is an R package for importing, pre-processing, visualiation, and statistical analsis of mass spectrometr imaging eperiments. Cardinal version 1.8 supports using matter as a backend for larger-thanmemor datasets. More information is available at The second half of this vignette presents an in-depth comparison in the performance between matter, bigmemor, and ff on real eperimental datasets that are larger than memor. 2 Analing large 3D MSI eperiments with Cardinal and matter This eample will use one of the benchmark 3D MSI eperiments from Oetjen et al. [1]. We will use the 3D microbial timecourse eperiment, which is comprised of interacting microbes at 3 time points, with a total of 17,672 piels and 40,299 features. The data is stored in
2 imml format [2]. The ".imml" XML file with eperimental metadata is 30.5 MB, and the ".ibd" binar file with the m/ values and spectral intensities is 2.85 GB. This is one of the smaller datasets, making it a good place to begin. Due to the various offsets in imml ibd files, the cannot be attached as simpl as bigmemor or ff files. These packages have strict requirements on the format of their data, for maimum computational effienc. matter takes a different approach with more fleibilit, which allows use of imml s domain-specific binar file format directl, and with minimal memor footprint, at the cost potentiall slower computational performance in some situations. > librar(matter) > librar(cardinal) > path <- "~/Documents/Datasets/3D-MSI/3D_Timecourse/" > file <- "Microbe_Interaction_3D_Timecourse_LP.imML" We load the dataset in Cardinal with the readmsidata function and the argument attach.onl=true. In Cardinal version 1.8, this automaticall uses matter. > msi <- readmsidata(paste0(path, file), attach.onl=true) The data is attached as a matter_mat matri. The matri metadata takes up approimatel 14.9 KB in memor, and points to 2.8 GB on disk. The dataset was stored so that the first time point t = 4 corresponds to = 1, 2, 3, 4, 5, 6, the second time point t = 8 corresponds to = 7, 8, 9, 10, 11, 12, and the third time point t = 11 corresponds to = 13, 14, 15, 16, 17, 18. We will reparamaterie the coordinates below to make it easier to work with the dataset. We also remove duplicated coordinates caused b converting the -dimension to sequential integers. > msi <- msi[,!duplicated(coord(msi))] > msi$sample <- factor(sappl(msi$, function() { if ( %in% 1:6 ) { 1 } else if ( %in% 7:12 ) { 2 } else if ( %in% 13:18 ) { 3 } }), labels=c("t = 4", "t = 8", "t = 11")) > msi$ <- sappl(msi$, function() { if ( %in% 1:6 ) { } else if ( %in% 7:12 ) { -6 } else if ( %in% 13:18 ) { -12 } }) > msi$ <- mappl(function(, t) { switch(as.integer(t), -30, -15, ) }, msi$, msi$sample) 2
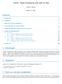
3 matter: Supplementar 2-3D mass spectrometr imaging case stud > varmetadata(msi)[c("","","","sample"),"labeltpe"] <- "dim" > protocoldata(msi) <- AnnotatedDataFrame( data=data.frame(row.names=samplenames(msi))) > msi <- regeneratepositions(msi) > validobject(msi) We can plot 3D molecular ion images using the image3d method. > image3d(msi, ~ * *, m=262, theta=-55, contrast="suppress", laout=c(3,1)) Now we perform principal components analsis using the PCA method of Cardinal. > pca <- PCA(msi, ncomp=2, method="irlba", center=true) > pdata(msi)[,c("pc1","pc2")] <- pca$scores[["ncomp = 2"]] > fdata(msi)[,c("pc1","pc2")] <- pca$loadings[["ncomp = 2"]] We plot the first three principal components. > image3d(msi, PC1 ~ * *, theta=-55, col.regions=risk.colors(100), laout=c(3,1)) > image3d(msi, PC2 ~ * *, theta=-55, col.regions=risk.colors(100), laout=c(3,1)) m/ = t=4 m/ = t=8 PC1 t=8 (b) PC1 scores PC2 t = 11 PC2 t=8 (a) m/ 262 PC2 t=4 PC1 t = 11 PC1 t=4 m/ = t = 11 (c) PC2 scores Figure 1: Plotting an ion image and the first 2 principal components for the 3D microbial time course eperiment 3 Comparisons with alternative approaches We will now illustrate the steps necessar for performing a principal components analsis using similar packages bigmemor or ff, and compare their performance with matter s. 3.1 Using bigmemor First we load bigmemor and create a blank to this new matri. filebacked.big.matri. We will cop the data We must use matter to read the data and convert it to a filebacked.big.matri, because the mass spectra are stored in a binar file incompatible with bigmemor. Note that the original data elements are 32-bit floats: 3
4 > head(atomdata(idata(msi))) However, if we want to use bigalgebra for matri multiplication, we must change the data tpe from a 32-bit float to a 64-bit double. While bigmemor provides efficient matri algebra routines in the bigalgebra package, the do not work with 32-bit floats; onl 64-bit doubles are supported. (Note that bigmemor itself does support 32-bit float matrices; onl bigalgebra s native routines don t.) > librar(bigmemor) > librar(bigalgebra) > backingfile <- paste0(epname, ".bin") > backingpath <- "~/Documents/Temporar/" > descriptorfile <- paste0(epname, ".desc") > msi.bm <- filebacked.big.matri(nrow=ncol(msi), ncol=nrow(msi), backingfile=backingfile, backingpath=backingpath, descriptorfile=descriptorfile, tpe="double") Furthermore, we must transpose the matri while converting it. This is because bioinformatics data is traditionall stored and manipulated using an P N matri, while most statistical functions in R epect a N P matri (where N is the number of samples and P is the number of features). bigmemor does not currentl support virtuall transposing a matri, so it must be transposed now in order to perform PCA later. > for ( i in seq_len(ncol(idata(msi))) ) msi.bm[i,] <- idata(msi)[,i] Lastl, bigmemor does not support virtuall scaling and centering rows or columns of a matri. PCA should tpicall be performed on a centered data matri. Although bigmemor supports arithmetic and algebra for big.matri objects through the bigalgebra package, a new big.matri with the transformed data is created as output. When the file must alread double in sie (conversion from float to double) to accomodate matri algebra, duplicating the matri again simpl to scale it seems an unacceptable waste of storage space. Therefore, we implement implicit centering of the data matri in a custom matri multiplication function which can be passed to the irlba function. > ct.mult.bm <- function(a, B, center = ct) { if ( is.vector(a) ) { A <- t(a) cbind((a %*% B)[]) - (sum(a) * ct) } else if ( is.vector(b) ) { B <- as.matri(b) cbind((a %*% B)[]) - sum(b * ct) } } Lastl, we calculate the mean of each column, and perform PCA using singular value decomposition via irlba with the custom multiplication function. 4
5 > librar(biganaltics) > librar(irlba) > ct <- appl(msi.bm, 2, mean) > pca.out <- irlba(msi.bm, nu=0, nv=2, mult=ct.mult.bm) > fdata(msi)[,c("pc1","pc2")] <- pca.out$v 3.2 Using ff We must again convert the dataset to a format compatible with ff. However, ff supports virtuall transposing matrices, so we can keep the same virtual data laout as with matter. While ff presents its own problems with matri multiplication, the do not etend to the data tpe of the data elements, so we can keep the data as 32-bit floats, saving storage space as compared to bigmemor. > librar(ff) > msi.ff <- ff(dim=c(nrow(msi), ncol(msi)), filename=paste0(backingpath, backingfile), vmode="single") > for ( i in seq_len(ncol(idata(msi))) ) msi.ff[,i] <- idata(msi)[,i] > msi.ff <- vt(msi.ff) However, the main ff package offers little in terms of arithmetic or algebraic operations that can be performed on ff objects. The ffbase package supplements and implements much of the arithmetic functionalit missing from ff. However, it also creates brand new on-disk data files as the output rather than supporting virtual scaling and centering of matrices. Therefore, we again perform implicit centering during matri multiplication using a custom matri multiplication function. Unfortunatel, ffbase does not implement matri multiplication for ff matrices. Fortunatel, the package bootsvd provides a function that performs matri multiplication of ff objects. It is also worth noting that ff, ffbase, and bootsvd are each maintained b different developers. > librar(ffbase) > librar(bootsvd) > ct.mult.ff <- function(a, B, center = ct) { if ( is.vector(a) ) { A <- t(a) cbind(ffmatrimult(a, B)[]) - (sum(a) * ct) } else if ( is.vector(b) ) { B <- as.matri(b) cbind(ffmatrimult(a, B)[]) - sum(b * ct) } } Now we can calculate the mean of each column, and perform PCA using singular value decomposition via irlba with the custom multiplication function. 5
6 > ct <- as.vector(ffappl(x=msi.ff, MARGIN=2, AFUN=mean, RETURN=TRUE)[]) > pca.out <- irlba(msi.ff, nu=0, nv=2, mult=ct.mult.ff) > fdata(msi)[,c("pc1","pc2")] <- pca.out$v 3.3 Evaluating performance between approaches Principal components analsis Dataset Sie Method Mem. used Mem. overhead Time 3D Microbial Time Course 2.9 GB matter 228 MB 50 MB 13 min, 6 sec bigmemor 330 MB 141 MB 53 sec ff 278 MB 85 MB 19 min, 48 sec 3D Oral Squamous Cell Carcinoma 25.4 GB matter 977 MB 668 MB 2 hr, 7 min, 9 sec bigmemor 408 MB 266 MB 2 hr, 28 min, 2 sec 3D Mouse Pancreas 26.4 GB matter 628 MB 370 MB 2 hr, 12 min, 46 sec bigmemor 303 MB 164 MB 2 hr, 52 min, 10 sec 3D Mouse Kidne 41.8 GB matter 1.5 GB 1074 MB 3 hr, 22 min, 23 sec bigmemor 617 MB 431 MB 4 hr, 29 min, 23 sec Table 1: Performance comparison of matter, bigmemor, and ff in calculating the first two principal components of real datasets Memor overhead is the maimum memor used during the eecution minus the memor in use upon completion. Some cells are missing because the analsis could not be performed with ff. Table 1 shows the amount of time and memor matter, bigmemor, and ff used when performing PCA on each of the 3D MSI datasets. For three of the datasets, the number of the elements in the dataset eceeded that maimum sie of an ff object. The total length of an ff object is stored as a 32-bit integer, so it can at most reference data elements. Conversel, matter and bigmemor store the total length of an object as a 64-bit double, allowing up to data elements. While bigmemor was dramaticall faster than either matter or ff for the smallest 2.8 GB dataset, for the larger datasets that actuall eceeded available memor, matter was faster. It should be noted that the memor consumption reported for bigmemor is likel erroneous. The analses were timed using the profmem function of matter, which wraps R s basic garbage collector call. The memor use will therefore be accurate for objects that use R s garbage collector, but inaccurate for objects that do not. The bigmemor package uses mmap on Uni sstems, which is not controlled b R. In fact, sstem memor consumption under bigmemor was dramaticall more than matter, but the authors were not able to measure this from R. For datasets which eceed available computer memor, matter appears to outperform both bigmemor and ff. Additionall, working with these datasets in bigmemor and ff either required additional steps that were either not required with matter, or, for some datasets, were simpl impossible. 6
7 File conversion Dataset Sie Method Mem. used Mem. overhead Time 3D Microbial Time Course 2.9 GB bigmemor 235 MB 46 MB 7 min, 12 sec ff 224 MB 32 MB 1 min, 16 sec 3D Oral Squamous Cell Carcinoma 25.4 GB bigmemor 1.4 GB 954 MB 51 min, 3 sec 3D Mouse Pancreas 26.4 GB bigmemor 630 MB 256 MB 1 hr, 31 min, 30 sec 3D Mouse Kidne 41.8 GB bigmemor 1.5 GB 781 MB 1 hr, 47 min, 58 sec Table 2: Time and memor used converting the dataset to a file compatible with bigmemor and/or ff Memor overhead is the maimum memor used during the eecution minus the memor in use upon completion. Some cells are missing because the conversion could not be performed for ff. Table 2 shows the amount of time and memor it took to convert the data to files compatible with bigmemor and ff. For ff, this was onl possible for one dataset. The time for file conversion (not included in the timing from Table 1) represents a substantial proportion of the total time it would take to anale these datasets. In addition, conversion to bigmemor required doubling the file sie if matri multiplication was to be performed using bigalgebra. In summar, matter often allows working with datasets without file conversion, which is often preferred for the sake of reproducibilit, and tpicall requires less effort even after file conversion. 4 An R script for comparing performance > librar(matter) > datapath <- "~/Documents/Datasets/3D-MSI/" > require(cardinal) > # file <- "3D_Timecourse/Microbe_Interaction_3D_Timecourse_LP.imML" > # epname <- "3D_Timecourse" > > # file <- "3D_OSCC/3D_OSCC.imML" > # epname <- "3D_OSCC" > > # file <- "3D_Mouse_Pancreas/3D_Mouse_Pancreas.imML" > # epname <- "3D_Mouse_Pancreas" > > file <- "3DMouseKidne/3DMouseKidne.imML" > epname <- "3DMouseKidne" > msi <- readmsidata(paste0(datapath, file), attach.onl=true) > msi.prof[[epname]][["matter"]] <- profmem({ pca.out <- PCA(msi, ncomp=2, method="irlba", center=true) pdata(msi)[,c("pc1","pc2")] <- pca.out$scores[[1]] 7
8 fdata(msi)[,c("pc1","pc2")] <- pca.out$loadings[[1]] }) > require(bigmemor) > require(bigalgebra) > backingfile <- paste0(epname, ".bin") > backingpath <- "~/Documents/Temporar/" > descriptorfile <- paste0(epname, ".desc") > msi.bm <- filebacked.big.matri(nrow=ncol(msi), ncol=nrow(msi), backingfile=backingfile, backingpath=backingpath, descriptorfile=descriptorfile, tpe="double") > msi.prof[[epname]][["convert.bigmemor"]] <- profmem({ for ( i in seq_len(ncol(idata(msi))) ) msi.bm[i,] <- idata(msi)[,i] }) > ct.mult.bm <- function(a, B, center = ct) { if ( is.vector(a) ) { A <- t(a) cbind((a %*% B)[]) - (sum(a) * ct) } else if ( is.vector(b) ) { B <- as.matri(b) cbind((a %*% B)[]) - sum(b * ct) } } > require(biganaltics) > require(irlba) > msi.prof[[epname]][["bigmemor"]] <- profmem({ ct <- appl(msi.bm, 2, mean) pca.out <- irlba(msi.bm, nu=0, nv=2, mult=ct.mult.bm) }) > file.remove(paste0(backingpath, backingfile)) > require(ff) > librar(ffbase) > require(bootsvd) > msi.ff <- ff(dim=c(nrow(msi), ncol(msi)), filename=paste0(backingpath, backingfile), vmode="single") > msi.prof[[epname]][["convert.ff"]] <- profmem({ 8
9 for ( i in seq_len(ncol(idata(msi))) ) msi.ff[,i] <- idata(msi)[,i] }) > msi.ff <- vt(msi.ff) > ct.mult.ff <- function(a, B, center = ct) { if ( is.vector(a) ) { A <- t(a) cbind(ffmatrimult(a, B)[]) - (sum(a) * ct) } else if ( is.vector(b) ) { B <- as.matri(b) cbind(ffmatrimult(a, B)[]) - sum(b * ct) } } > msi.prof[[epname]][["ff"]] <- profmem({ ct <- as.vector(ffappl(x=msi.ff, MARGIN=2, AFUN=mean, RETURN=TRUE)[]) pca.out <- irlba(msi.ff, nu=0, nv=2, mult=ct.mult.ff) }) > file.remove(paste0(backingpath, backingfile)) > print(msi.prof) $`3D_Timecourse` $`3D_Timecourse`$matter $`3D_Timecourse`$convert.bigmemor $`3D_Timecourse`$bigmemor $`3D_Timecourse`$convert.ff $`3D_Timecourse`$ff $`3D_OSCC` $`3D_OSCC`$matter 9
10 $`3D_OSCC`$convert.bigmemor $`3D_OSCC`$bigmemor $`3DMouseKidne` $`3DMouseKidne`$matter $`3DMouseKidne`$convert.bigmemor $`3DMouseKidne`$bigmemor $`3D_Mouse_Pancreas` $`3D_Mouse_Pancreas`$matter $`3D_Mouse_Pancreas`$convert.bigmemor $`3D_Mouse_Pancreas`$bigmemor Session info R version Patched ( r74699), 86_64-pc-linu-gnu Locale: LC_CTYPE=en_US.UTF-8, LC_NUMERIC=C, LC_TIME=en_US.UTF-8, LC_COLLATE=C, LC_MONETARY=en_US.UTF-8, LC_MESSAGES=en_US.UTF-8, LC_PAPER=en_US.UTF-8, LC_NAME=C, LC_ADDRESS=C, LC_TELEPHONE=C, LC_MEASUREMENT=en_US.UTF-8, LC_IDENTIFICATION=C Running under: Ubuntu LTS 10
11 Matri products: default BLAS: /home/biocbuild/bbs-3.8-bioc/r/lib/librblas.so LAPACK: /home/biocbuild/bbs-3.8-bioc/r/lib/librlapack.so Base packages: base, datasets, grdevices, graphics, methods, stats, utils Other packages: DBI 1.0.0, biglm 0.9-1, matter Loaded via a namespace (and not attached): BiocGenerics , BiocStle 2.9.3, Matri , Rcpp , backports 1.1.2, compiler 3.5.0, digest , evaluate , grid 3.5.0, htmltools 0.3.6, irlba 2.3.2, knitr 1.20, lattice , magrittr 1.5, parallel 3.5.0, rmarkdown 1.10, rprojroot 1.3-2, stringi 1.2.3, stringr 1.3.1, tools 3.5.0, aml References [1] Janina Oetjen, Kirill Veselkov, Jeramie Watrous, James S McKenie, Michael Becker, Lena Hauberg-Lotte, Jan Hendrik Kobarg, Nicole Strittmatter, Anna K Mró, Franiska Hoffmann, Dennis Trede, Andrew Palmer, Stefan Schiffler, Klaus Steinhorst, Michaela Aichler, Robert Goldin, Orlando Guntinas-Lichius, Ferdinand von Eggeling, Herbert Thiele, Kathrin Maedler, Ael Walch, Peter Maass, Pieter C Dorrestein, Zoltan Takats, and Theodore Aleandrov. Benchmark datasets for 3D MALDI- and DESI-imaging mass spectrometr. GigaScience, 4(1):2105 8, Ma [2] T. Schramm, A. Hester, I. Klinkert, J. P. Both, R. M. Heeren, A. Brunelle, O. Laprévote, N. Desbenoit, F. Robbe M, M. Stoeckli, B. Spengler, and A. Römpp. imml A common data format for the fleible echange and processing of mass spectrometr imaging data. Journal of Proteomics, 75:5106,
Cardinal design and development
 Kylie A. Bemis October 30, 2017 Contents 1 Introduction.............................. 2 2 Design overview........................... 2 3 iset: high-throughput imaging experiments............ 3 3.1 SImageSet:
Kylie A. Bemis October 30, 2017 Contents 1 Introduction.............................. 2 2 Design overview........................... 2 3 iset: high-throughput imaging experiments............ 3 3.1 SImageSet:
Image Analysis with beadarray
 Mike Smith October 30, 2017 Introduction From version 2.0 beadarray provides more flexibility in the processing of array images and the extraction of bead intensities than its predecessor. In the past
Mike Smith October 30, 2017 Introduction From version 2.0 beadarray provides more flexibility in the processing of array images and the extraction of bead intensities than its predecessor. In the past
matter: Rapid prototyping with data on disk
 matter: Rapid prototyping with data on disk Kylie A. Bemis March 17, 2017 Contents 1 Introduction 1 2 Installation 1 3 Basic use and data manipulation 1 4 Linear regression for on-disk datasets 5 5 Principal
matter: Rapid prototyping with data on disk Kylie A. Bemis March 17, 2017 Contents 1 Introduction 1 2 Installation 1 3 Basic use and data manipulation 1 4 Linear regression for on-disk datasets 5 5 Principal
Advanced analysis using bayseq; generic distribution definitions
 Advanced analysis using bayseq; generic distribution definitions Thomas J Hardcastle October 30, 2017 1 Generic Prior Distributions bayseq now offers complete user-specification of underlying distributions
Advanced analysis using bayseq; generic distribution definitions Thomas J Hardcastle October 30, 2017 1 Generic Prior Distributions bayseq now offers complete user-specification of underlying distributions
rhdf5 - HDF5 interface for R
 Bernd Fischer October 30, 2017 Contents 1 Introduction 1 2 Installation of the HDF5 package 2 3 High level R -HDF5 functions 2 31 Creating an HDF5 file and group hierarchy 2 32 Writing and reading objects
Bernd Fischer October 30, 2017 Contents 1 Introduction 1 2 Installation of the HDF5 package 2 3 High level R -HDF5 functions 2 31 Creating an HDF5 file and group hierarchy 2 32 Writing and reading objects
survsnp: Power and Sample Size Calculations for SNP Association Studies with Censored Time to Event Outcomes
 survsnp: Power and Sample Size Calculations for SNP Association Studies with Censored Time to Event Outcomes Kouros Owzar Zhiguo Li Nancy Cox Sin-Ho Jung Chanhee Yi June 29, 2016 1 Introduction This vignette
survsnp: Power and Sample Size Calculations for SNP Association Studies with Censored Time to Event Outcomes Kouros Owzar Zhiguo Li Nancy Cox Sin-Ho Jung Chanhee Yi June 29, 2016 1 Introduction This vignette
riboseqr Introduction Getting Data Workflow Example Thomas J. Hardcastle, Betty Y.W. Chung October 30, 2017
 riboseqr Thomas J. Hardcastle, Betty Y.W. Chung October 30, 2017 Introduction Ribosome profiling extracts those parts of a coding sequence currently bound by a ribosome (and thus, are likely to be undergoing
riboseqr Thomas J. Hardcastle, Betty Y.W. Chung October 30, 2017 Introduction Ribosome profiling extracts those parts of a coding sequence currently bound by a ribosome (and thus, are likely to be undergoing
segmentseq: methods for detecting methylation loci and differential methylation
 segmentseq: methods for detecting methylation loci and differential methylation Thomas J. Hardcastle October 30, 2018 1 Introduction This vignette introduces analysis methods for data from high-throughput
segmentseq: methods for detecting methylation loci and differential methylation Thomas J. Hardcastle October 30, 2018 1 Introduction This vignette introduces analysis methods for data from high-throughput
An Overview of the S4Vectors package
 Patrick Aboyoun, Michael Lawrence, Hervé Pagès Edited: February 2018; Compiled: June 7, 2018 Contents 1 Introduction.............................. 1 2 Vector-like and list-like objects...................
Patrick Aboyoun, Michael Lawrence, Hervé Pagès Edited: February 2018; Compiled: June 7, 2018 Contents 1 Introduction.............................. 1 2 Vector-like and list-like objects...................
PROPER: PROspective Power Evaluation for RNAseq
 PROPER: PROspective Power Evaluation for RNAseq Hao Wu [1em]Department of Biostatistics and Bioinformatics Emory University Atlanta, GA 303022 [1em] hao.wu@emory.edu October 30, 2017 Contents 1 Introduction..............................
PROPER: PROspective Power Evaluation for RNAseq Hao Wu [1em]Department of Biostatistics and Bioinformatics Emory University Atlanta, GA 303022 [1em] hao.wu@emory.edu October 30, 2017 Contents 1 Introduction..............................
How to Use pkgdeptools
 How to Use pkgdeptools Seth Falcon February 10, 2018 1 Introduction The pkgdeptools package provides tools for computing and analyzing dependency relationships among R packages. With it, you can build
How to Use pkgdeptools Seth Falcon February 10, 2018 1 Introduction The pkgdeptools package provides tools for computing and analyzing dependency relationships among R packages. With it, you can build
Creating IGV HTML reports with tracktables
 Thomas Carroll 1 [1em] 1 Bioinformatics Core, MRC Clinical Sciences Centre; thomas.carroll (at)imperial.ac.uk June 13, 2018 Contents 1 The tracktables package.................... 1 2 Creating IGV sessions
Thomas Carroll 1 [1em] 1 Bioinformatics Core, MRC Clinical Sciences Centre; thomas.carroll (at)imperial.ac.uk June 13, 2018 Contents 1 The tracktables package.................... 1 2 Creating IGV sessions
Visualisation, transformations and arithmetic operations for grouped genomic intervals
 ## Warning: replacing previous import ggplot2::position by BiocGenerics::Position when loading soggi Visualisation, transformations and arithmetic operations for grouped genomic intervals Thomas Carroll
## Warning: replacing previous import ggplot2::position by BiocGenerics::Position when loading soggi Visualisation, transformations and arithmetic operations for grouped genomic intervals Thomas Carroll
MIGSA: Getting pbcmc datasets
 MIGSA: Getting pbcmc datasets Juan C Rodriguez Universidad Católica de Córdoba Universidad Nacional de Córdoba Cristóbal Fresno Instituto Nacional de Medicina Genómica Andrea S Llera Fundación Instituto
MIGSA: Getting pbcmc datasets Juan C Rodriguez Universidad Católica de Córdoba Universidad Nacional de Córdoba Cristóbal Fresno Instituto Nacional de Medicina Genómica Andrea S Llera Fundación Instituto
Working with aligned nucleotides (WORK- IN-PROGRESS!)
 Working with aligned nucleotides (WORK- IN-PROGRESS!) Hervé Pagès Last modified: January 2014; Compiled: November 17, 2017 Contents 1 Introduction.............................. 1 2 Load the aligned reads
Working with aligned nucleotides (WORK- IN-PROGRESS!) Hervé Pagès Last modified: January 2014; Compiled: November 17, 2017 Contents 1 Introduction.............................. 1 2 Load the aligned reads
Introduction to the Codelink package
 Introduction to the Codelink package Diego Diez October 30, 2018 1 Introduction This package implements methods to facilitate the preprocessing and analysis of Codelink microarrays. Codelink is a microarray
Introduction to the Codelink package Diego Diez October 30, 2018 1 Introduction This package implements methods to facilitate the preprocessing and analysis of Codelink microarrays. Codelink is a microarray
matter: Rapid prototyping with data on disk
 Kylie A. Bemis November 1, 2017 Contents 1 Introduction.............................. 1 2 Installation............................... 2 3 Basic use and data manipulation.................. 2 4 Linear regression
Kylie A. Bemis November 1, 2017 Contents 1 Introduction.............................. 1 2 Installation............................... 2 3 Basic use and data manipulation.................. 2 4 Linear regression
Introduction to BatchtoolsParam
 Nitesh Turaga 1, Martin Morgan 2 Edited: March 22, 2018; Compiled: January 4, 2019 1 Nitesh.Turaga@ RoswellPark.org 2 Martin.Morgan@ RoswellPark.org Contents 1 Introduction..............................
Nitesh Turaga 1, Martin Morgan 2 Edited: March 22, 2018; Compiled: January 4, 2019 1 Nitesh.Turaga@ RoswellPark.org 2 Martin.Morgan@ RoswellPark.org Contents 1 Introduction..............................
mocluster: Integrative clustering using multiple
 mocluster: Integrative clustering using multiple omics data Chen Meng Modified: June 03, 2015. Compiled: October 30, 2018. Contents 1 mocluster overview......................... 1 2 Run mocluster............................
mocluster: Integrative clustering using multiple omics data Chen Meng Modified: June 03, 2015. Compiled: October 30, 2018. Contents 1 mocluster overview......................... 1 2 Run mocluster............................
The analysis of rtpcr data
 The analysis of rtpcr data Jitao David Zhang, Markus Ruschhaupt October 30, 2017 With the help of this document, an analysis of rtpcr data can be performed. For this, the user has to specify several parameters
The analysis of rtpcr data Jitao David Zhang, Markus Ruschhaupt October 30, 2017 With the help of this document, an analysis of rtpcr data can be performed. For this, the user has to specify several parameters
Analysis of two-way cell-based assays
 Analysis of two-way cell-based assays Lígia Brás, Michael Boutros and Wolfgang Huber April 16, 2015 Contents 1 Introduction 1 2 Assembling the data 2 2.1 Reading the raw intensity files..................
Analysis of two-way cell-based assays Lígia Brás, Michael Boutros and Wolfgang Huber April 16, 2015 Contents 1 Introduction 1 2 Assembling the data 2 2.1 Reading the raw intensity files..................
RMassBank for XCMS. Erik Müller. January 4, Introduction 2. 2 Input files LC/MS data Additional Workflow-Methods 2
 RMassBank for XCMS Erik Müller January 4, 2019 Contents 1 Introduction 2 2 Input files 2 2.1 LC/MS data........................... 2 3 Additional Workflow-Methods 2 3.1 Options..............................
RMassBank for XCMS Erik Müller January 4, 2019 Contents 1 Introduction 2 2 Input files 2 2.1 LC/MS data........................... 2 3 Additional Workflow-Methods 2 3.1 Options..............................
HowTo: Querying online Data
 HowTo: Querying online Data Jeff Gentry and Robert Gentleman November 12, 2017 1 Overview This article demonstrates how you can make use of the tools that have been provided for on-line querying of data
HowTo: Querying online Data Jeff Gentry and Robert Gentleman November 12, 2017 1 Overview This article demonstrates how you can make use of the tools that have been provided for on-line querying of data
How To Use GOstats and Category to do Hypergeometric testing with unsupported model organisms
 How To Use GOstats and Category to do Hypergeometric testing with unsupported model organisms M. Carlson October 30, 2017 1 Introduction This vignette is meant as an extension of what already exists in
How To Use GOstats and Category to do Hypergeometric testing with unsupported model organisms M. Carlson October 30, 2017 1 Introduction This vignette is meant as an extension of what already exists in
DupChecker: a bioconductor package for checking high-throughput genomic data redundancy in meta-analysis
 DupChecker: a bioconductor package for checking high-throughput genomic data redundancy in meta-analysis Quanhu Sheng, Yu Shyr, Xi Chen Center for Quantitative Sciences, Vanderbilt University, Nashville,
DupChecker: a bioconductor package for checking high-throughput genomic data redundancy in meta-analysis Quanhu Sheng, Yu Shyr, Xi Chen Center for Quantitative Sciences, Vanderbilt University, Nashville,
Creating a New Annotation Package using SQLForge
 Creating a New Annotation Package using SQLForge Marc Carlson, HervÃľ PagÃĺs, Nianhua Li April 30, 2018 1 Introduction The AnnotationForge package provides a series of functions that can be used to build
Creating a New Annotation Package using SQLForge Marc Carlson, HervÃľ PagÃĺs, Nianhua Li April 30, 2018 1 Introduction The AnnotationForge package provides a series of functions that can be used to build
An Introduction to ShortRead
 Martin Morgan Modified: 21 October, 2013. Compiled: April 30, 2018 > library("shortread") The ShortRead package provides functionality for working with FASTQ files from high throughput sequence analysis.
Martin Morgan Modified: 21 October, 2013. Compiled: April 30, 2018 > library("shortread") The ShortRead package provides functionality for working with FASTQ files from high throughput sequence analysis.
MMDiff2: Statistical Testing for ChIP-Seq Data Sets
 MMDiff2: Statistical Testing for ChIP-Seq Data Sets Gabriele Schwikert 1 and David Kuo 2 [1em] 1 The Informatics Forum, University of Edinburgh and The Wellcome
MMDiff2: Statistical Testing for ChIP-Seq Data Sets Gabriele Schwikert 1 and David Kuo 2 [1em] 1 The Informatics Forum, University of Edinburgh and The Wellcome
Triform: peak finding in ChIP-Seq enrichment profiles for transcription factors
 Triform: peak finding in ChIP-Seq enrichment profiles for transcription factors Karl Kornacker * and Tony Håndstad October 30, 2018 A guide for using the Triform algorithm to predict transcription factor
Triform: peak finding in ChIP-Seq enrichment profiles for transcription factors Karl Kornacker * and Tony Håndstad October 30, 2018 A guide for using the Triform algorithm to predict transcription factor
NacoStringQCPro. Dorothee Nickles, Thomas Sandmann, Robert Ziman, Richard Bourgon. Modified: April 10, Compiled: April 24, 2017.
 NacoStringQCPro Dorothee Nickles, Thomas Sandmann, Robert Ziman, Richard Bourgon Modified: April 10, 2017. Compiled: April 24, 2017 Contents 1 Scope 1 2 Quick start 1 3 RccSet loaded with NanoString data
NacoStringQCPro Dorothee Nickles, Thomas Sandmann, Robert Ziman, Richard Bourgon Modified: April 10, 2017. Compiled: April 24, 2017 Contents 1 Scope 1 2 Quick start 1 3 RccSet loaded with NanoString data
Bioconductor: Annotation Package Overview
 Bioconductor: Annotation Package Overview April 30, 2018 1 Overview In its current state the basic purpose of annotate is to supply interface routines that support user actions that rely on the different
Bioconductor: Annotation Package Overview April 30, 2018 1 Overview In its current state the basic purpose of annotate is to supply interface routines that support user actions that rely on the different
Count outlier detection using Cook s distance
 Count outlier detection using Cook s distance Michael Love August 9, 2014 1 Run DE analysis with and without outlier removal The following vignette produces the Supplemental Figure of the effect of replacing
Count outlier detection using Cook s distance Michael Love August 9, 2014 1 Run DE analysis with and without outlier removal The following vignette produces the Supplemental Figure of the effect of replacing
bayseq: Empirical Bayesian analysis of patterns of differential expression in count data
 bayseq: Empirical Bayesian analysis of patterns of differential expression in count data Thomas J. Hardcastle October 30, 2017 1 Introduction This vignette is intended to give a rapid introduction to the
bayseq: Empirical Bayesian analysis of patterns of differential expression in count data Thomas J. Hardcastle October 30, 2017 1 Introduction This vignette is intended to give a rapid introduction to the
segmentseq: methods for detecting methylation loci and differential methylation
 segmentseq: methods for detecting methylation loci and differential methylation Thomas J. Hardcastle October 13, 2015 1 Introduction This vignette introduces analysis methods for data from high-throughput
segmentseq: methods for detecting methylation loci and differential methylation Thomas J. Hardcastle October 13, 2015 1 Introduction This vignette introduces analysis methods for data from high-throughput
A complete analysis of peptide microarray binding data using the pepstat framework
 A complete analysis of peptide microarray binding data using the pepstat framework Greg Imholte, Renan Sauteraud, Mike Jiang and Raphael Gottardo April 30, 2018 This document present a full analysis, from
A complete analysis of peptide microarray binding data using the pepstat framework Greg Imholte, Renan Sauteraud, Mike Jiang and Raphael Gottardo April 30, 2018 This document present a full analysis, from
How to use cghmcr. October 30, 2017
 How to use cghmcr Jianhua Zhang Bin Feng October 30, 2017 1 Overview Copy number data (arraycgh or SNP) can be used to identify genomic regions (Regions Of Interest or ROI) showing gains or losses that
How to use cghmcr Jianhua Zhang Bin Feng October 30, 2017 1 Overview Copy number data (arraycgh or SNP) can be used to identify genomic regions (Regions Of Interest or ROI) showing gains or losses that
MultipleAlignment Objects
 MultipleAlignment Objects Marc Carlson Bioconductor Core Team Fred Hutchinson Cancer Research Center Seattle, WA April 30, 2018 Contents 1 Introduction 1 2 Creation and masking 1 3 Analytic utilities 7
MultipleAlignment Objects Marc Carlson Bioconductor Core Team Fred Hutchinson Cancer Research Center Seattle, WA April 30, 2018 Contents 1 Introduction 1 2 Creation and masking 1 3 Analytic utilities 7
Fitting Baselines to Spectra
 Fitting Baselines to Spectra Claudia Beleites DIA Raman Spectroscopy Group, University of Trieste/Italy (2005 2008) Spectroscopy Imaging, IPHT, Jena/Germany (2008 2016) ÖPV, JKI,
Fitting Baselines to Spectra Claudia Beleites DIA Raman Spectroscopy Group, University of Trieste/Italy (2005 2008) Spectroscopy Imaging, IPHT, Jena/Germany (2008 2016) ÖPV, JKI,
Shrinkage of logarithmic fold changes
 Shrinkage of logarithmic fold changes Michael Love August 9, 2014 1 Comparing the posterior distribution for two genes First, we run a DE analysis on the Bottomly et al. dataset, once with shrunken LFCs
Shrinkage of logarithmic fold changes Michael Love August 9, 2014 1 Comparing the posterior distribution for two genes First, we run a DE analysis on the Bottomly et al. dataset, once with shrunken LFCs
Design Issues in Matrix package Development
 Design Issues in Matrix package Development Martin Maechler and Douglas Bates R Core Development Team maechler@stat.math.ethz.ch, bates@r-project.org Spring 2008 (typeset on November 16, 2017) Abstract
Design Issues in Matrix package Development Martin Maechler and Douglas Bates R Core Development Team maechler@stat.math.ethz.ch, bates@r-project.org Spring 2008 (typeset on November 16, 2017) Abstract
Analyse RT PCR data with ddct
 Analyse RT PCR data with ddct Jitao David Zhang, Rudolf Biczok and Markus Ruschhaupt October 30, 2017 Abstract Quantitative real time PCR (qrt PCR or RT PCR for short) is a laboratory technique based on
Analyse RT PCR data with ddct Jitao David Zhang, Rudolf Biczok and Markus Ruschhaupt October 30, 2017 Abstract Quantitative real time PCR (qrt PCR or RT PCR for short) is a laboratory technique based on
Design Issues in Matrix package Development
 Design Issues in Matrix package Development Martin Maechler and Douglas Bates R Core Development Team maechler@stat.math.ethz.ch, bates@r-project.org Spring 2008 (typeset on October 26, 2018) Abstract
Design Issues in Matrix package Development Martin Maechler and Douglas Bates R Core Development Team maechler@stat.math.ethz.ch, bates@r-project.org Spring 2008 (typeset on October 26, 2018) Abstract
Getting started with flowstats
 Getting started with flowstats F. Hahne, N. Gopalakrishnan October, 7 Abstract flowstats is a collection of algorithms for the statistical analysis of flow cytometry data. So far, the focus is on automated
Getting started with flowstats F. Hahne, N. Gopalakrishnan October, 7 Abstract flowstats is a collection of algorithms for the statistical analysis of flow cytometry data. So far, the focus is on automated
The bumphunter user s guide
 The bumphunter user s guide Kasper Daniel Hansen khansen@jhsph.edu Martin Aryee aryee.martin@mgh.harvard.edu Rafael A. Irizarry rafa@jhu.edu Modified: November 23, 2012. Compiled: April 30, 2018 Introduction
The bumphunter user s guide Kasper Daniel Hansen khansen@jhsph.edu Martin Aryee aryee.martin@mgh.harvard.edu Rafael A. Irizarry rafa@jhu.edu Modified: November 23, 2012. Compiled: April 30, 2018 Introduction
Analysis of screens with enhancer and suppressor controls
 Analysis of screens with enhancer and suppressor controls Lígia Brás, Michael Boutros and Wolfgang Huber April 16, 2015 Contents 1 Introduction 1 2 Assembling the data 2 2.1 Reading the raw intensity files..................
Analysis of screens with enhancer and suppressor controls Lígia Brás, Michael Boutros and Wolfgang Huber April 16, 2015 Contents 1 Introduction 1 2 Assembling the data 2 2.1 Reading the raw intensity files..................
INSPEcT - INference of Synthesis, Processing and degradation rates in Time-course analysis
 INSPEcT - INference of Synthesis, Processing and degradation rates in Time-course analysis Stefano de Pretis October 30, 2017 Contents 1 Introduction.............................. 1 2 Quantification of
INSPEcT - INference of Synthesis, Processing and degradation rates in Time-course analysis Stefano de Pretis October 30, 2017 Contents 1 Introduction.............................. 1 2 Quantification of
MetCirc: Navigating mass spectral similarity in high-resolution MS/MS metabolomics data
 MetCirc: Navigating mass spectral similarity in high-resolution MS/MS metabolomics data Thomas Naake and Emmanuel Gaquerel Plant Defense Metabolism Centre for Organismal Studies August 30, 2018 1 Introduction
MetCirc: Navigating mass spectral similarity in high-resolution MS/MS metabolomics data Thomas Naake and Emmanuel Gaquerel Plant Defense Metabolism Centre for Organismal Studies August 30, 2018 1 Introduction
Overview of ensemblvep
 Overview of ensemblvep Lori Shepherd and Valerie Obenchain October 30, 2017 Contents 1 Introduction 1 2 Results as R objects 1 3 Write results to a file 4 4 Configuring runtime options 5 5 sessioninfo()
Overview of ensemblvep Lori Shepherd and Valerie Obenchain October 30, 2017 Contents 1 Introduction 1 2 Results as R objects 1 3 Write results to a file 4 4 Configuring runtime options 5 5 sessioninfo()
Introduction to GenomicFiles
 Valerie Obenchain, Michael Love, Martin Morgan Last modified: October 2014; Compiled: October 30, 2018 Contents 1 Introduction.............................. 1 2 Quick Start..............................
Valerie Obenchain, Michael Love, Martin Morgan Last modified: October 2014; Compiled: October 30, 2018 Contents 1 Introduction.............................. 1 2 Quick Start..............................
CQN (Conditional Quantile Normalization)
 CQN (Conditional Quantile Normalization) Kasper Daniel Hansen khansen@jhsph.edu Zhijin Wu zhijin_wu@brown.edu Modified: August 8, 2012. Compiled: April 30, 2018 Introduction This package contains the CQN
CQN (Conditional Quantile Normalization) Kasper Daniel Hansen khansen@jhsph.edu Zhijin Wu zhijin_wu@brown.edu Modified: August 8, 2012. Compiled: April 30, 2018 Introduction This package contains the CQN
Using metama for differential gene expression analysis from multiple studies
 Using metama for differential gene expression analysis from multiple studies Guillemette Marot and Rémi Bruyère Modified: January 28, 2015. Compiled: January 28, 2015 Abstract This vignette illustrates
Using metama for differential gene expression analysis from multiple studies Guillemette Marot and Rémi Bruyère Modified: January 28, 2015. Compiled: January 28, 2015 Abstract This vignette illustrates
Comparing Least Squares Calculations
 Comparing Least Squares Calculations Douglas Bates R Development Core Team Douglas.Bates@R-project.org November 16, 2017 Abstract Many statistics methods require one or more least squares problems to be
Comparing Least Squares Calculations Douglas Bates R Development Core Team Douglas.Bates@R-project.org November 16, 2017 Abstract Many statistics methods require one or more least squares problems to be
An Introduction to GenomeInfoDb
 Martin Morgan, Hervé Pagès, Marc Carlson, Sonali Arora Modified: 17 January, 2014. Compiled: January 21, 2018 Contents 1 Introduction.............................. 2 2 Functionality for all existing organisms...............
Martin Morgan, Hervé Pagès, Marc Carlson, Sonali Arora Modified: 17 January, 2014. Compiled: January 21, 2018 Contents 1 Introduction.............................. 2 2 Functionality for all existing organisms...............
SCnorm: robust normalization of single-cell RNA-seq data
 SCnorm: robust normalization of single-cell RNA-seq data Rhonda Bacher and Christina Kendziorski February 18, 2019 Contents 1 Introduction.............................. 1 2 Run SCnorm.............................
SCnorm: robust normalization of single-cell RNA-seq data Rhonda Bacher and Christina Kendziorski February 18, 2019 Contents 1 Introduction.............................. 1 2 Run SCnorm.............................
Introduction: microarray quality assessment with arrayqualitymetrics
 : microarray quality assessment with arrayqualitymetrics Audrey Kauffmann, Wolfgang Huber October 30, 2017 Contents 1 Basic use............................... 3 1.1 Affymetrix data - before preprocessing.............
: microarray quality assessment with arrayqualitymetrics Audrey Kauffmann, Wolfgang Huber October 30, 2017 Contents 1 Basic use............................... 3 1.1 Affymetrix data - before preprocessing.............
Overview of ensemblvep
 Overview of ensemblvep Lori Shepherd and Valerie Obenchain October 30, 2018 Contents 1 Introduction 1 2 Results as R objects 1 3 Write results to a file 4 4 Configuring runtime options 5 5 sessioninfo()
Overview of ensemblvep Lori Shepherd and Valerie Obenchain October 30, 2018 Contents 1 Introduction 1 2 Results as R objects 1 3 Write results to a file 4 4 Configuring runtime options 5 5 sessioninfo()
Raman Spectra of Chondrocytes in Cartilage: hyperspec s chondro data set
 Raman Spectra of Chondrocytes in Cartilage: hyperspec s chondro data set Claudia Beleites CENMAT and DI3, University of Trieste Spectroscopy Imaging, IPHT Jena e.v. February 13,
Raman Spectra of Chondrocytes in Cartilage: hyperspec s chondro data set Claudia Beleites CENMAT and DI3, University of Trieste Spectroscopy Imaging, IPHT Jena e.v. February 13,
Bioconductor L A T E X Style 2.0
 Andrzej Oleś 1, Martin Morgan 2, and Wolfgang Huber 1 1 European Molecular Biology Laboratory, Heidelberg, Germany 2 Roswell Park Cancer Institute, Buffalo, NY Abstract Package November 29, 2017 This vignette
Andrzej Oleś 1, Martin Morgan 2, and Wolfgang Huber 1 1 European Molecular Biology Laboratory, Heidelberg, Germany 2 Roswell Park Cancer Institute, Buffalo, NY Abstract Package November 29, 2017 This vignette
Creating a New Annotation Package using SQLForge
 Creating a New Annotation Package using SQLForge Marc Carlson, Herve Pages, Nianhua Li November 19, 2013 1 Introduction The AnnotationForge package provides a series of functions that can be used to build
Creating a New Annotation Package using SQLForge Marc Carlson, Herve Pages, Nianhua Li November 19, 2013 1 Introduction The AnnotationForge package provides a series of functions that can be used to build
highscreen: High-Throughput Screening for Plate Based Assays
 highscreen: High-Throughput Screening for Plate Based Assays Ivo D. Shterev Cliburn Chan Gregory D. Sempowski August 22, 2018 1 Introduction This vignette describes the use of the R extension package highscreen
highscreen: High-Throughput Screening for Plate Based Assays Ivo D. Shterev Cliburn Chan Gregory D. Sempowski August 22, 2018 1 Introduction This vignette describes the use of the R extension package highscreen
GxE.scan. October 30, 2018
 GxE.scan October 30, 2018 Overview GxE.scan can process a GWAS scan using the snp.logistic, additive.test, snp.score or snp.matched functions, whereas snp.scan.logistic only calls snp.logistic. GxE.scan
GxE.scan October 30, 2018 Overview GxE.scan can process a GWAS scan using the snp.logistic, additive.test, snp.score or snp.matched functions, whereas snp.scan.logistic only calls snp.logistic. GxE.scan
Classification of Breast Cancer Clinical Stage with Gene Expression Data
 Classification of Breast Cancer Clinical Stage with Gene Expression Data Zhu Wang Connecticut Children s Medical Center University of Connecticut School of Medicine zwang@connecticutchildrens.org July
Classification of Breast Cancer Clinical Stage with Gene Expression Data Zhu Wang Connecticut Children s Medical Center University of Connecticut School of Medicine zwang@connecticutchildrens.org July
INSPEcT - INference of Synthesis, Processing and degradation rates in Time-course analysis
 INSPEcT - INference of Synthesis, Processing and degradation rates in Time-course analysis Stefano de Pretis April 7, 20 Contents 1 Introduction 1 2 Quantification of Exon and Intron features 3 3 Time-course
INSPEcT - INference of Synthesis, Processing and degradation rates in Time-course analysis Stefano de Pretis April 7, 20 Contents 1 Introduction 1 2 Quantification of Exon and Intron features 3 3 Time-course
Simulation of molecular regulatory networks with graphical models
 Simulation of molecular regulatory networks with graphical models Inma Tur 1 inma.tur@upf.edu Alberto Roverato 2 alberto.roverato@unibo.it Robert Castelo 1 robert.castelo@upf.edu 1 Universitat Pompeu Fabra,
Simulation of molecular regulatory networks with graphical models Inma Tur 1 inma.tur@upf.edu Alberto Roverato 2 alberto.roverato@unibo.it Robert Castelo 1 robert.castelo@upf.edu 1 Universitat Pompeu Fabra,
How to use CNTools. Overview. Algorithms. Jianhua Zhang. April 14, 2011
 How to use CNTools Jianhua Zhang April 14, 2011 Overview Studies have shown that genomic alterations measured as DNA copy number variations invariably occur across chromosomal regions that span over several
How to use CNTools Jianhua Zhang April 14, 2011 Overview Studies have shown that genomic alterations measured as DNA copy number variations invariably occur across chromosomal regions that span over several
Introduction to QDNAseq
 Ilari Scheinin October 30, 2017 Contents 1 Running QDNAseq.......................... 1 1.1 Bin annotations.......................... 1 1.2 Processing bam files....................... 2 1.3 Downstream analyses......................
Ilari Scheinin October 30, 2017 Contents 1 Running QDNAseq.......................... 1 1.1 Bin annotations.......................... 1 1.2 Processing bam files....................... 2 1.3 Downstream analyses......................
Bioconductor L A T E X Style 2.0
 Andrzej Oleś 1, Martin Morgan 2, and Wolfgang Huber 1 1 European Molecular Biology Laboratory, Heidelberg, Germany 2 Roswell Park Cancer Institute, Buffalo, NY Abstract Package November 23, 2016 This vignette
Andrzej Oleś 1, Martin Morgan 2, and Wolfgang Huber 1 1 European Molecular Biology Laboratory, Heidelberg, Germany 2 Roswell Park Cancer Institute, Buffalo, NY Abstract Package November 23, 2016 This vignette
ssviz: A small RNA-seq visualizer and analysis toolkit
 ssviz: A small RNA-seq visualizer and analysis toolkit Diana HP Low Institute of Molecular and Cell Biology Agency for Science, Technology and Research (A*STAR), Singapore dlow@imcb.a-star.edu.sg August
ssviz: A small RNA-seq visualizer and analysis toolkit Diana HP Low Institute of Molecular and Cell Biology Agency for Science, Technology and Research (A*STAR), Singapore dlow@imcb.a-star.edu.sg August
A wrapper to query DGIdb using R
 A wrapper to query DGIdb using R Thomas Thurnherr 1, Franziska Singer, Daniel J. Stekhoven, Niko Beerenwinkel 1 thomas.thurnherr@gmail.com Abstract February 16, 2018 Annotation and interpretation of DNA
A wrapper to query DGIdb using R Thomas Thurnherr 1, Franziska Singer, Daniel J. Stekhoven, Niko Beerenwinkel 1 thomas.thurnherr@gmail.com Abstract February 16, 2018 Annotation and interpretation of DNA
SIBER User Manual. Pan Tong and Kevin R Coombes. May 27, Introduction 1
 SIBER User Manual Pan Tong and Kevin R Coombes May 27, 2015 Contents 1 Introduction 1 2 Using SIBER 1 2.1 A Quick Example........................................... 1 2.2 Dealing With RNAseq Normalization................................
SIBER User Manual Pan Tong and Kevin R Coombes May 27, 2015 Contents 1 Introduction 1 2 Using SIBER 1 2.1 A Quick Example........................................... 1 2.2 Dealing With RNAseq Normalization................................
Biobase development and the new eset
 Biobase development and the new eset Martin T. Morgan * 7 August, 2006 Revised 4 September, 2006 featuredata slot. Revised 20 April 2007 minor wording changes; verbose and other arguments passed through
Biobase development and the new eset Martin T. Morgan * 7 August, 2006 Revised 4 September, 2006 featuredata slot. Revised 20 April 2007 minor wording changes; verbose and other arguments passed through
An introduction to gaucho
 An introduction to gaucho Alex Murison Alexander.Murison@icr.ac.uk Christopher P Wardell Christopher.Wardell@icr.ac.uk October 30, 2017 This vignette serves as an introduction to the R package gaucho.
An introduction to gaucho Alex Murison Alexander.Murison@icr.ac.uk Christopher P Wardell Christopher.Wardell@icr.ac.uk October 30, 2017 This vignette serves as an introduction to the R package gaucho.
SimBindProfiles: Similar Binding Profiles, identifies common and unique regions in array genome tiling array data
 SimBindProfiles: Similar Binding Profiles, identifies common and unique regions in array genome tiling array data Bettina Fischer October 30, 2018 Contents 1 Introduction 1 2 Reading data and normalisation
SimBindProfiles: Similar Binding Profiles, identifies common and unique regions in array genome tiling array data Bettina Fischer October 30, 2018 Contents 1 Introduction 1 2 Reading data and normalisation
Sparse Model Matrices
 Sparse Model Matrices Martin Maechler R Core Development Team maechler@r-project.org July 2007, 2008 (typeset on April 6, 2018) Introduction Model matrices in the very widely used (generalized) linear
Sparse Model Matrices Martin Maechler R Core Development Team maechler@r-project.org July 2007, 2008 (typeset on April 6, 2018) Introduction Model matrices in the very widely used (generalized) linear
Introduction: microarray quality assessment with arrayqualitymetrics
 Introduction: microarray quality assessment with arrayqualitymetrics Audrey Kauffmann, Wolfgang Huber April 4, 2013 Contents 1 Basic use 3 1.1 Affymetrix data - before preprocessing.......................
Introduction: microarray quality assessment with arrayqualitymetrics Audrey Kauffmann, Wolfgang Huber April 4, 2013 Contents 1 Basic use 3 1.1 Affymetrix data - before preprocessing.......................
Comparing methods for differential expression analysis of RNAseq data with the compcoder
 Comparing methods for differential expression analysis of RNAseq data with the compcoder package Charlotte Soneson October 30, 2017 Contents 1 Introduction.............................. 2 2 The compdata
Comparing methods for differential expression analysis of RNAseq data with the compcoder package Charlotte Soneson October 30, 2017 Contents 1 Introduction.............................. 2 2 The compdata
Flexible Logo plots of symbols and alphanumeric strings using Logolas
 Flexible Logo plots of symbols and alphanumeric strings using Logolas Kushal K Dey [1em] Stephens Lab, Dept. of Statistics, The University of Chicago Correspondending Email: kkdey@uchicago.edu Abstract
Flexible Logo plots of symbols and alphanumeric strings using Logolas Kushal K Dey [1em] Stephens Lab, Dept. of Statistics, The University of Chicago Correspondending Email: kkdey@uchicago.edu Abstract
Seminar III: R/Bioconductor
 Leonardo Collado Torres lcollado@lcg.unam.mx Bachelor in Genomic Sciences www.lcg.unam.mx/~lcollado/ August - December, 2009 1 / 25 Class outline Working with HTS data: a simulated case study Intro R for
Leonardo Collado Torres lcollado@lcg.unam.mx Bachelor in Genomic Sciences www.lcg.unam.mx/~lcollado/ August - December, 2009 1 / 25 Class outline Working with HTS data: a simulated case study Intro R for
Classes. Compiling Methods. Code Generation for Objects. Implementing Objects. Methods. Fields
 Classes Implementing Objects Components fields/instance variables values differ from to usuall mutable methods values shared b all s of a class usuall immutable component visibilit: public/private/protected
Classes Implementing Objects Components fields/instance variables values differ from to usuall mutable methods values shared b all s of a class usuall immutable component visibilit: public/private/protected
Flexible Logo plots of symbols and alphanumeric strings using Logolas
 Flexible Logo plots of symbols and alphanumeric strings using Logolas Kushal K Dey Stephens Lab, Dept. of Statistics, The University of Chicago Correspondending Email: kkdey@uchicago.edu April 24, 2017
Flexible Logo plots of symbols and alphanumeric strings using Logolas Kushal K Dey Stephens Lab, Dept. of Statistics, The University of Chicago Correspondending Email: kkdey@uchicago.edu April 24, 2017
Calibration of Quinine Fluorescence Emission Vignette for the Data Set flu of the R package hyperspec
 Calibration of Quinine Fluorescence Emission Vignette for the Data Set flu of the R package hyperspec Claudia Beleites CENMAT and DI3, University of Trieste Spectroscopy Imaging,
Calibration of Quinine Fluorescence Emission Vignette for the Data Set flu of the R package hyperspec Claudia Beleites CENMAT and DI3, University of Trieste Spectroscopy Imaging,
INVESTIGATION OF K-MEANS AND FUZZY K-MEANS CLUSTERING FOR THE ANALYSIS OF MASS SPECTROMETRY IMAGING DATA
 INVESTIGATION OF K-MEANS AND FUZZY K-MEANS CLUSTERING FOR THE ANALYSIS OF MASS SPECTROMETRY IMAGING DATA A Thesis Presented to The Academic Faculty by Sanaiya Sarkari In Partial Fulfillment of the Requirements
INVESTIGATION OF K-MEANS AND FUZZY K-MEANS CLUSTERING FOR THE ANALYSIS OF MASS SPECTROMETRY IMAGING DATA A Thesis Presented to The Academic Faculty by Sanaiya Sarkari In Partial Fulfillment of the Requirements
An Introduction to Bioconductor s ExpressionSet Class
 An Introduction to Bioconductor s ExpressionSet Class Seth Falcon, Martin Morgan, and Robert Gentleman 6 October, 2006; revised 9 February, 2007 1 Introduction Biobase is part of the Bioconductor project,
An Introduction to Bioconductor s ExpressionSet Class Seth Falcon, Martin Morgan, and Robert Gentleman 6 October, 2006; revised 9 February, 2007 1 Introduction Biobase is part of the Bioconductor project,
Clustering time series gene expression data with TMixClust
 4 Session info.............................. 15 ## Warning: latex2 is deprecated. ## Use latex instead. ## See help("deprecated") Clustering series gene expression data with TMixClust Abstract Monica Golumbeanu
4 Session info.............................. 15 ## Warning: latex2 is deprecated. ## Use latex instead. ## See help("deprecated") Clustering series gene expression data with TMixClust Abstract Monica Golumbeanu
A Parser for mzxml, mzdata and mzml files
 A Parser for mzxml, mzdata and mzml files Bernd Fischer Steffen Neumann Laurent Gatto December 23, 2012 Contents 1 Introduction 1 2 Mass spectrometry raw data 2 2.1 Spectral data access.........................
A Parser for mzxml, mzdata and mzml files Bernd Fischer Steffen Neumann Laurent Gatto December 23, 2012 Contents 1 Introduction 1 2 Mass spectrometry raw data 2 2.1 Spectral data access.........................
Introduction to the TSSi package: Identification of Transcription Start Sites
 Introduction to the TSSi package: Identification of Transcription Start Sites Julian Gehring, Clemens Kreutz October 30, 2017 Along with the advances in high-throughput sequencing, the detection of transcription
Introduction to the TSSi package: Identification of Transcription Start Sites Julian Gehring, Clemens Kreutz October 30, 2017 Along with the advances in high-throughput sequencing, the detection of transcription
Basic4Cseq: an R/Bioconductor package for the analysis of 4C-seq data
 Basic4Cseq: an R/Bioconductor package for the analysis of 4C-seq data Carolin Walter October 30, 2017 Contents 1 Introduction 1 1.1 Loading the package...................................... 2 1.2 Provided
Basic4Cseq: an R/Bioconductor package for the analysis of 4C-seq data Carolin Walter October 30, 2017 Contents 1 Introduction 1 1.1 Loading the package...................................... 2 1.2 Provided
The Command Pattern in R
 The Command Pattern in R Michael Lawrence September 2, 2012 Contents 1 Introduction 1 2 Example pipelines 2 3 sessioninfo 8 1 Introduction Command pattern is a design pattern used in object-oriented programming,
The Command Pattern in R Michael Lawrence September 2, 2012 Contents 1 Introduction 1 2 Example pipelines 2 3 sessioninfo 8 1 Introduction Command pattern is a design pattern used in object-oriented programming,
An Introduction to the methylumi package
 An Introduction to the methylumi package Sean Davis and Sven Bilke October 13, 2014 1 Introduction Gene expression patterns are very important in understanding any biologic system. The regulation of gene
An Introduction to the methylumi package Sean Davis and Sven Bilke October 13, 2014 1 Introduction Gene expression patterns are very important in understanding any biologic system. The regulation of gene
MALDIquantForeign: Import/Export routines for MALDIquant
 MALDIquantForeign: Import/Export routines for MALDIquant Sebastian Gibb December 4, 2017 Abstract MALDIquantForeign provides routines for importing/exporting different file formats into/from MALDIquant.
MALDIquantForeign: Import/Export routines for MALDIquant Sebastian Gibb December 4, 2017 Abstract MALDIquantForeign provides routines for importing/exporting different file formats into/from MALDIquant.
The rtracklayer package
 The rtracklayer package Michael Lawrence January 22, 2018 Contents 1 Introduction 2 2 Gene expression and microrna target sites 2 2.1 Creating a target site track..................... 2 2.1.1 Constructing
The rtracklayer package Michael Lawrence January 22, 2018 Contents 1 Introduction 2 2 Gene expression and microrna target sites 2 2.1 Creating a target site track..................... 2 2.1.1 Constructing
RDAVIDWebService: a versatile R interface to DAVID
 RDAVIDWebService: a versatile R interface to DAVID Cristóbal Fresno 1,2 and Elmer A. Fernández 1,2 1 Bio-science Data Mining Group, Catholic University of Córdoba, Córdoba, Argentina. 2 CONICET, Argentina
RDAVIDWebService: a versatile R interface to DAVID Cristóbal Fresno 1,2 and Elmer A. Fernández 1,2 1 Bio-science Data Mining Group, Catholic University of Córdoba, Córdoba, Argentina. 2 CONICET, Argentina
To 3D or not to 3D? Why GPUs Are Critical for 3D Mass Spectrometry Imaging Eri Rubin SagivTech Ltd.
 To 3D or not to 3D? Why GPUs Are Critical for 3D Mass Spectrometry Imaging Eri Rubin SagivTech Ltd. Established in 2009 and headquartered in Israel Core domain expertise: GPU Computing and Computer Vision
To 3D or not to 3D? Why GPUs Are Critical for 3D Mass Spectrometry Imaging Eri Rubin SagivTech Ltd. Established in 2009 and headquartered in Israel Core domain expertise: GPU Computing and Computer Vision
Introduction to R (BaRC Hot Topics)
 Introduction to R (BaRC Hot Topics) George Bell September 30, 2011 This document accompanies the slides from BaRC s Introduction to R and shows the use of some simple commands. See the accompanying slides
Introduction to R (BaRC Hot Topics) George Bell September 30, 2011 This document accompanies the slides from BaRC s Introduction to R and shows the use of some simple commands. See the accompanying slides
M 3 D: Statistical Testing for RRBS Data Sets
 Tom Mayo Modified: 5 September, 2016. Compiled: October 30, 2018 Contents 1 Introduction 1 2 Analysis Pipeline 2 2.1 Reading in Data. 2 2.2 Computing the MMD, and coverage-mmd. 5 2.3 Generating p-values..
Tom Mayo Modified: 5 September, 2016. Compiled: October 30, 2018 Contents 1 Introduction 1 2 Analysis Pipeline 2 2.1 Reading in Data. 2 2.2 Computing the MMD, and coverage-mmd. 5 2.3 Generating p-values..
RAPIDR. Kitty Lo. November 20, Intended use of RAPIDR 1. 2 Create binned counts file from BAMs Masking... 1
 RAPIDR Kitty Lo November 20, 2014 Contents 1 Intended use of RAPIDR 1 2 Create binned counts file from BAMs 1 2.1 Masking.................................................... 1 3 Build the reference 2 3.1
RAPIDR Kitty Lo November 20, 2014 Contents 1 Intended use of RAPIDR 1 2 Create binned counts file from BAMs 1 2.1 Masking.................................................... 1 3 Build the reference 2 3.1
Using the dorng package
 Using the dorng package dorng package Version 1.6 Renaud Gaujoux March 5, 2014 Contents Introduction............. 1 1 The %dorng% operator...... 3 1.1 How it works......... 3 1.2 Seeding computations.....
Using the dorng package dorng package Version 1.6 Renaud Gaujoux March 5, 2014 Contents Introduction............. 1 1 The %dorng% operator...... 3 1.1 How it works......... 3 1.2 Seeding computations.....
MALDIquant: Quantitative Analysis of Mass Spectrometry Data
 MALDIquant: Quantitative Analysis of Mass Spectrometry Data Sebastian Gibb November 12, 2017 Abstract MALDIquant provides a complete analysis pipeline for MALDI- TOF and other 2D mass spectrometry data.
MALDIquant: Quantitative Analysis of Mass Spectrometry Data Sebastian Gibb November 12, 2017 Abstract MALDIquant provides a complete analysis pipeline for MALDI- TOF and other 2D mass spectrometry data.
Package bigmemory. January 11, 2018
 Version 4.5.33 Package bigmemory January 11, 2018 Title Manage Massive Matrices with Shared Memory and Memory-Mapped Files Author Michael J. Kane , John W. Emerson ,
Version 4.5.33 Package bigmemory January 11, 2018 Title Manage Massive Matrices with Shared Memory and Memory-Mapped Files Author Michael J. Kane , John W. Emerson ,
bcseq Fast Sequence Alignment for High-throughput shrna and CRISPR Screens
 bcseq Fast Sequence Alignment for High-throughput shrna and CRISPR Screens Jiaxing Lin Jeremy Gresham Tongrong Wang So Young Kim James Alvarez Jeffrey S. Damrauer Scott Floyd Joshua Granek Andrew Allen
bcseq Fast Sequence Alignment for High-throughput shrna and CRISPR Screens Jiaxing Lin Jeremy Gresham Tongrong Wang So Young Kim James Alvarez Jeffrey S. Damrauer Scott Floyd Joshua Granek Andrew Allen
