A Divide and Conquer Framework for Distributed Graph Clustering
|
|
|
- Moses Marshall
- 5 years ago
- Views:
Transcription
1 Wenzhuo Yang Department of Mechanical Engineering National University of Singapore Singapore Huan Xu Department of Mechanical Engineering National University of Singapore Singapore Abstract Graph clustering is about identifying clusters of closely connected nodes and is a fundamental technique of data analysis with many applications including community detection VLSI network partitioning collaborative filtering etc. In order to improve the scalability of existing graph clustering algorithms we propose a novel divide and conquer framework for graph clustering and establish theoretical guarantees of exact recovery of the clusters. One additional advantage of the proposed framework is that it can identify small clusters the size of the smallest cluster can be of size o( n) in contrast to Ω( n) required by standard methods. Extensive experiments on synthetic and real-world datasets demonstrate the efficiency and effectiveness of our framework. 1. Introduction Graph clustering is a widely-used tool for exploiting the relationships between nodes connected in a graph. Such graph usually consists of a large number of nodes e.g. the friendship graph on Facebook the co-author graph in bibliographic records. The pairwise connection between two nodes indicates their similarity or affinity whereas the disconnection represents their dissimilarity or difference. The goal of graph clustering is to partition the nodes into clusters so that the nodes within the same cluster have more connections than those in different clusters in other words the closely connected nodes are grouped together. Various kinds of graph models have been proposed and studied in literature (e.g. Holland et al. 1983; Watts & Strogatz 1998; Goldenberg et al. 2010) among which the stochastic block model (Holland et al. 1983) also known Proceedings of the 32 nd International Conference on Machine Learning Lille France JMLR: W&CP volume 37. Copyright 2015 by the author(s). as the planted partition model (Condon & arp 2001) is arguably the most natural one for clustering. The stochastic block model assumes that the nodes are partitioned into several clusters and the probability of the connection between each pair of nodes depends on their cluster membership i.e. an edge is generated with probability p if they are in the same cluster or with probability q if they are in different clusters with p > q and the goal is to recover the ground truth clusters from the observed graph. Many graph clustering algorithms have been analyzed under this model. One of the most popular approaches is spectral clustering (Luxburg 2007). Rohe et al. (2011) analyzed the performance of spectral clustering under the stochastic block model and provided an upper bound of the misclassified nodes. Another approach developed recently and has attracted great attention is based on convex optimization using trace-norm and l 1 norm as a surrogate of the rank and sparsity functions respectively (e.g. Oymak & Hassibi 2011; Jalali et al. 2011; Chen et al. 2014b; Ames & Vavasis 2011; Chen et al. 2014a). The main advantages of the trace-norm based clustering are their theoretically guarantees for the exact recovery of the ground-truth clustering and their outstanding empirical performance. Indeed one of them proposed by Chen et al. (2014b) can provably recover the ground-truth clustering even in the face of outlier nodes (nodes that are not in any cluster) and can be used to solve planted clique and planted coloring problems. However both spectral clustering and trace-norm based clustering algorithms have difficulties tackling very large graphs. Indeed for a graph with n nodes both approaches need to compute the spectral decomposition of a n n matrix which requires O(n 3 ) operations and O(n 2 ) storage. Hence it is impractical to apply them even on mediumsized graphs with nodes on a desktop PC. How to reduce the memory and computational cost of these methods to improve their scalability is undoubtedly a significant problem as real-world applications often involve graphs with tens of thousands to even millions of nodes (Günnemann et al. 2010; Macropol & Singh 2010; Whang et al. 2012). To address this one approach is to perform
2 a multilevel clustering that reduces the size of the graph by collapsing vertices and edges partitions the smaller graph and then uncoarsens to recover clusters in the original graph (arypis & umar 1998a;b; Yan et al. 2009; Chen et al. 2011; Liu et al. 2013). Another approach is to apply the Nyström method to approximate the similarity matrix for acceleration (Fowlkes et al. 2004; Drineas & Mahoney 2005). While these approaches are empirically evaluated by the experiments on synthetic or realworld datasets it is not clear whether any theoretical guarantees on exact recovery of the ground truth can be obtained. Moreover it is hard to extend them to improve the scalability of trace-norm based clustering algorithms. The goal of this paper is to make trace-norm based clustering algorithms applicable for clustering large graphs. Specifically we develop and analyze a novel divide and conquer framework for improving the scalability of a wide range of graph clustering algorithms including spectral clustering and trace-norm based clustering. Our framework divides the original graph into several easily handled subgraphs executes a selected graph clustering algorithm on the subgraphs in parallel and then combines the results using a new algorithm called graph clustering with high confidence nodes. Our analysis provides sufficient conditions on when the proposed framework can exactly recover the ground-truth clustering. Moreover applying our framework on trace-norm based clustering algorithms offers additional benefits for identifying small clusters the size of the smallest cluster can in fact be smaller than O( n). Notations. We use lower-case boldface letters to denote column vectors and upper-case boldface letters to denote matrices. The minimum of an empty set is defined as +. Three matrix norms are used: M 2 is the spectral norm M is the nuclear norm and M 1 is the element-wise l 1 norm. We use [r] to denote the set {1 2 r} for integer r and S to denote the cardinality of set S. For a set of matrix indices Ω we let P Ω (X) be the matrix by setting the entries outside Ω to zero. 2. Divide and Conquer for Graph Clustering We first provide a brief introduction about the generalized stochastic blockmodel (Chen et al. 2014b) which is an extension of the standard stochastic block model (Holland et al. 1983; Condon & arp 2001) and then present our graph clustering framework in details Generalized Stochastic Blockmodel (GSBM) We consider a random graph with n nodes. These n nodes are divided into two sets V 1 and V 2 which have n 1 inlier nodes and n 2 outlier nodes respectively. The n 1 inlier nodes are partitioned into r disjoint clusters which we will refer to as the true clusters. The sizes of these r clusters are 1 r. Let = min i [r] i i.e. the minimum cluster size. The edges of the graph are generated as follows: For each pair of nodes i j that belong to the same cluster edge (i j) exists with probability at least p; for each pair of nodes that are in different clusters the edge exists with probability at most q. The n 2 outlier nodes are loosely connected to other nodes namely edge (i j) exists with probability at most q for outlier node i and each node j i. The adjacency matrix of the graph is denoted by A. Our goal is to recover from A the adjacency matrix Y of the ground-truth clusters i.e. 1) Yii = 1 if node i is an inlier node and Yii = 0 otherwise 2) for i j Yij = 1 if i j are in the same cluster and equals 0 otherwise Intuition Before formally presenting the proposed framework we explain its intuition first. Our framework involves three steps: 1) dividing step: uniformly randomly partition the n nodes into m groups and then construct the corresponding m subgraphs of A 2) clustering step: recover clusters in each subgraph in parallel using some graph clustering algorithm and 3) combining step: construct the final clustering by combining the recovered clusters in the subgraphs. We need to answer four questions to make the above procedure concrete: 1) How does m the number of subgraphs affect the performance of this framework? 2) How to choose the graph clustering algorithm used in the clustering step? 3) How to combine the recovered clusters from the subgraphs? 4) Under what conditions can we recover the ground-truth clustering via this framework? Question 1 2 and 4 will be studied in Section 3. We now focus on Question 3. For convenience we suppose that the nodes in the graph are labeled by consecutive integers {1 n}. We denote the m subgraphs by {g 1 g m } and the set of the clusters recovered in subgraph g i by S gi. To illustrate our main idea we take the graph shown in Figure 1 containing 12 nodes and two clusters as an example. Suppose that this graph is divided into two subgraphs g 1 and g 2 in the dividing step as shown in Figure 1(b) and the clusters in each subgraph are recovered by a certain algorithm i.e. S g1 = {{1 2 3} {7 8 9}} and = {{4 5 6} { }} in the clustering step. S g2 Now we discuss how to combine these clusters. For clusters U and V which may or may not belong to the same subgraph let U V represent that the nodes in U and V belong to the same cluster and U V denote otherwise. Under the GSBM we observe that for the bipartite graph generated by nodes in U and in V the expected edge density is at least p if U V while it is at most q if U V. Therefore if the edge density is greater than some threshold t where q t p we have U V w.h.p. In our
3 (a) (c) (b) Figure 1. A simple graph with 12 nodes and two clusters YEL- LOW and PURPLE. (a) The observed graph. (b) Two subgraphs generated in the dividing step. (c) The fused graph. (d) The final recovered clusters. example the edge density corresponding to {7 8 9} and { } is 0.56 while the edge density corresponding to {7 8 9} and {4 5 6} is 0.11 so select t = 0.3 and we conclude that w.h.p {7 8 9} { } and {7 8 9} {4 5 6}. Of course estimating and relationship based on edge density is not always correct and may even lead to results contradicting each other. Therefore instead of naively merging clusters based on the edge density we build a new (meta-)graph whose nodes correspond to the clusters in S g1 S g2 and edges are formed according to the edge density between two clusters i.e. an edge is constructed between them if their edge density is greater than threshold t. We call this graph fused graph and call its nodes super nodes as shown in Figure 1(c). Then the final clustering result can be obtained by performing a certain graph clustering algorithm on this fused graph which is shown in Figure 1(d). There is an interesting property of these super nodes: each edge/no-edge connecting with a super node is very likely to be correct because it is estimated by averaging many edges in the original graph (recall that a super-node corresponds to a cluster in a subgraph). We call a node with such property a high confidence node formally defined as follows: Definition 1. For a graph with n nodes node i is called a high confidence node if 1) for each node j in the same cluster as i edge (i j) exists with probability at least τ 1 and 2) for each node j in different clusters from i edge (i j) exists with probability at most 1 τ 2 where τ 1 τ 2 satisfies min{τ 1 τ 2 } 0.8 and 1 min{τ 1 τ 2 } c τ n for some constant c τ. Node i is an ordinary node if it is a not high confident. To cluster the fused graph we used a variant of the tracenorm based optimization approach for graph clustering. In particular since we know that the super-nodes are of high confidence we put more weights on the penalty of the edge disagreement related to high confidence nodes: min YS Y + c A P A C cs 1 + c A c P A c CcS 1 + cc PCS 1 s.t. Y + S = A 0 Y 1 (d) (1) where c A c A c c C c C c are parameters whose values are determined later A {(i j) : A ij = 1} and C {(i j) : i or j is a high confidence node}. The solution Y is the clustering relationship we recover and S is the disagreement between the observation and the recovered clustering. Thus when we set c C large Y and A are less likely to disagree on edges/no-edges connecting to high confidence nodes. We analyze this formulation in Section 3 and show that it offers important benefits Algorithm Based on the intuitive idea discussed in the previous section we design the following framework shown in Algorithm 1 for distributed graph clustering. Clearly if a clus- Algorithm 1 Large Graph Clustering Input: Graph clustering algorithm A observed adjacency matrix A and parameter m l t T. Procedure: 1) Uniformly randomly partition the n nodes into m groups and then extract the corresponding m subgraphs {g 1 g m }; 2) Find clusters in each subgraph in parallel by applying A. Denote the set of the recovered clusters in g i by S gi ; 3) Build the fused graph G based on {S g1 S gm } by Algorithm 2; 4) Find clusters in G by solving (1). Denote the set of the recovered clusters by S G ; 5) Merge {S g1 S gm } based on S G. ter from a subgraph is too small it may not be truly high confident. Hence we break up the clusters with size less than some threshold T into single nodes so that only the large clusters form high confidence nodes. Moreover as we will show in the next section high confidence nodes can assist in achieving the exact recovery of clustering so we divide each large cluster recovered in subgraphs into l parts to generate more high confidence nodes for more help for recovering clusters in the fused graph. 3. Theoretical Analysis In this section we establish the conditions under which the proposed framework provably recovers the ground truth. Indeed even when the clusters in the subgraphs are partially recovered we can still recover the correct clustering relationship of the entire graph. We first investigate the theoretical guarantee for the weighted clustering shown in Equation (1). Based on this we then provide the performance guarantees of Algorithm 1. Along the way we also provide answers to Question 1 and 2 mentioned in the previous section.
4 Algorithm 2 Build a Fused Graph Input: Observed adjacency matrix A the sets of the recovered clusters S g1 S gm and parameter l t T. Procedure: 1) Break up small clusters: For each g {g 1 g m } and each U S g if U < T let S g := (S g \ U) {{k} for each k U}. Otherwise uniformly randomly partition the nodes in U into l clusters U 1 U l and let S g := (S g \ U) {U 1 U l }; 2) Create super nodes: Create a corresponding super node V for each cluster U in m i=1 S g i. This relationship is represented by V U; 3) Build the fused graph G: For super nodes V i and V j where V i U i and V j U j edge E ij that connects them is generated as follows: If U i = U j = 1 let E ij := A uv where u U i and v U j. Otherwise we compute u U Ê(V i V j ) := i v U j A uv u U i v U j 1. Let E ij := 1 if Ê(V i V j ) t or E ij := 0 otherwise; 4) Construct the set of high confidence nodes: H {V i : U i > 1 for V i V}; 5) Return G and H Graph Clustering with High Confidence Nodes Let us consider a graph with high confidence nodes: more specifically the graph is generated according to the GSBM which contains n nodes and r clusters and its ith cluster contains s i high confidence nodes. Let s r i=1 s i i.e. the total number of the high confidence nodes is the size of the smallest cluster and be the minimum size of the clusters containing at least one ordinary node. We have the following theorem: c Theorem 1. Let λ = 0 c 1 t A = λ max{n s } log n t t c A c = λ 1 t and c c C = 0 log n where t satisfies 1 4 p q t 3 4 p q then Y is the unique optimal solution of (1) with high probability if c log n and { } p q (n s) log n log n c 1 max p(1 q) (2) hold for some universal constants c c 0 and c 1. Furthermore if there exist no outliers in the graph and λ = c 0 max{ max i{ j i (i si)}} log n (2) can be replaced by a milder condition p q max i { j i ( i s i )} log n log n c 1 max p(1 q). Remark. This theorem makes no assumptions on how the confidence nodes are distributed over different clusters. Consider an extreme case: fix r = O(1) and let the r clusters have equal size n r and the nodes in the first r 2 clusters are all highly confident while the nodes in the remaining r 2 clusters are all ordinary. By an information-theoretic argument (Chen et al. 2014b;a) one can show that for any algorithm to correctly recover the clusters with probability at least 3 4 the inequality p q c 1 (3) p(1 q) n should hold for some universal constant c. Theorem 1 shows that in this case p q should satisfy p q r log n c 1 p(1 q) n This matches the condition (3) up to a logarithmic factor and thus cannot be substantially improved unless additional assumptions are made Performance Guarantee of Algorithm 1 Graph clustering algorithm A used to find clusters in subgraphs is crucial to our framework. Clearly if A generates many misclassified nodes it is impossible for Algorithm 1 to correctly recover the true clusters. We now characterize two sets of algorithms used in our framework. Definition 2 (Workable Algorithm). For vector λ R r and non-empty set I [r] A is (λ I)-workable if the followings hold with probability at least 1 n 2 : when (p q) is in a set C parameterized by (n 1 r λ I) A is able to recover a set of clusters R C satisfying 1) i I C i R C so that C i is a subset of the ith cluster and C i λ i i and 2) C min i I ρλ i i for any C R C \ i I C i for some constant ρ < 1. Example 1. Most of the trace-norm based graph clustering algorithms (e.g. Oymak & Hassibi 2011; Ames & Vavasis 2011; Chen et al. 2014b; Jalali et al. 2011) are workable. For example the algorithm proposed by Chen et al. (2014b) has a performance guarantee that when p q c max{ n p(1 q) log 2 n } for universal constant c its output is the same as the true adjacency matrix Y with probability at least 1 n 8 which means that it is (1 [r])- workable. Ailon et al. (2013) provided a refined analysis of this algorithm showing that small clusters do not hinder recovery of sufficiently large ones under some mild conditions i.e. it is (λ [r])-workable where λ i = 1 if the ith cluster is relatively large or 0 otherwise. Definition 3 (Pseudo-workable Algorithm). For vector λ ϵ R r and non-empty set I [r] A is (λ I ϵ)- pseudo-workable if the followings hold with probability at
5 least 1 n 2 : when (p q) is in a set C parameterized by (n 1 r λ I ϵ) A is able to recover a set of clusters R C satisfying 1) i I C i R C so that C i contains at least λ i i nodes in the ith cluster and at most ϵ i i nodes not in the ith cluster and 2) C min i I ρλ i i for any C R C \ i I C i for some constant ρ < 1. Example 2. Rohe et al. (2011) analyzed the performance of spectral clustering under the standard stochastic blockmodel and provided an upper bound of the number of misclassified nodes. For clarity we here present a specialized version of their theorem as shown in Proposition 1 where the clusters are equal-sized from which one can easily observe that spectral clustering is indeed (1 ϵ [r] ϵ)- pseudo-workable where ϵ = cr4 log 2 n n 1. Proposition 1. Suppose that the graph has n nodes with r equal-sized clusters and satisfies that the probability of a connection between two nodes in the same cluster is p and the probability of a connection between two nodes in different clusters is q. If p and q do not vary as n and r = O(n 1 4 log 1 n) then the size of the set of the misclassified nodes M after running spectral clustering on this graph is upper bounded by M cr 3 log 2 n for some constant c. We now present our main theorems which establish the performance guarantee of Algorithm 1 when A used to find clusters in subgraphs is either workable or pseudoworkable respectively. Theorem 2. If A is (λ I)-workable t ( 1 4 p q 3 4 p q) and T min i I λ i i ( 3ρ+ρ2 2(1+ρ)m 1+3ρ 2(1+ρ)m ) then when (p q) belongs to the set C(n/m 1 /m r /m λ I) c 3 log n m p q min l l 1 max 1 ρ 4(1+ρ) {c 2 log n and (1 + ρ)ml log n (1 + 3ρ) min i I λ i i { (n c 1 p(1 q) max i I λ i i ) log n S(m l) log n S(m l) }} with probability at least 1 max{(mr) 10 n 1 } the true clusters can be recovered by Algorithm 1 where S(m l) = min{min i I:λi 1{ml + (1 λ i ) i } min i I c i } l = { } 1 max (1+3ρ) mini I λ i i 4 2(1+ρ)m log n 1 and c 1 c 2 c 3 are universal constants. Remark. Our framework can guarantee exact recovery even when each cluster in the subgraphs is partially recovered. This happens especially when trace-norm based clustering algorithms are applied on a sparse graph where p q < 0.2 as shown in the experiments in (Chen et al. 2014b) or spectral clustering runs with a relatively large cluster number. Parameter T and m are well defined when ρ < 1 and = Ω(log 3 n). Recall that m controls the number of the separated subgraphs. From the bound related to (p q) we observe that increasing m leads to a smaller C which increases the difficulty of recovering the true clusters in subgraphs but a larger S(m l) which means that the difficulty of recovering clusters in the fused graph is reduced. Theorem 3. Suppose that A is (λ I ϵ)-pseudo-workable t ( 1 4 p q + φ ( 3 4 p + 1 T 4q)(1 φ)) min i I λ i i ( 3ρ+ρ2 2(1+ρ)m 1+3ρ 2(1+ρ)m ) then when (p q) belongs to the set C(n/m 1 /m r /m λ I ϵ) c 3 log n m 1 ρ 4(1+ρ) log n and (1 + ρ)ml log p q max {c n ϵ i 2 72 max (1 + 3ρ) min i I λ i i i I λ i { (n c 1 p(1 q) max i I λ }} i i ) log n log n S(m l) S(m l) with probability at least 1 max{(mr) 10 n 1 } the clusters recovered by Algorithm 1 contain at most i I ϵ i i misclassified nodes where l min { max { 1 4 S(m l) = min min max (1+3ρ) mini I λ i i i I:λ i 1 } } 2(1+ρ)m log n 1 p q 72 min i I λi ϵ i min i I c i j I ml + [(1 λ i) i ϵ j j 1 j Ij i ϵ j j ] + φ = 18 max i I ϵ i l λ i and c 1 c 2 c 3 are universal constants. Remark. Theorem 3 states the error-tolerant property of Algorithm 1. In particular even when A does not guarantee exact recovery the total number of the misclassified nodes in the clusters recovered by Algorithm 1 is still small as long as the recovered clusters in subgraphs do not contain too many misclassified nodes. We now specialize our main theorem. For simplicity we set l = 1 in Algorithm 1. The following corollaries consider the case where A is the trace-norm based clustering algorithm proposed by Chen et al. (2014b) i.e. A solves the optimization problem shown in (1) with no high confidence nodes i.e. C =. We call this algorithm CSX following author acronym in the sequel. Corollary 1. If A is CSX t ( 1 4 p q 3 4 p q) 0 mini i T 2m c 3 log n m 1 4 log n and { } p(1 q)mn log n m log n p q max c 1 c 2 (4) where c 1 c 2 c 3 are universal constants then Algorithm 1 recovers the true clusters with probability at least 1 max{(mr) 10 n 1 }.
6 Remark. Speedup is not a free lunch : Increasing m reduces the computational time but requires stronger conditions to guarantee exact recovery i.e. the lower bound of p q is m times greater than the one shown in (Chen et al. 2014b). In practice p q can be smaller than (4) while exact recovery is still achieved since the exact recovery of clusters in subgraphs is not necessary as stated in Theorem 1. More discussion can be found in Section 4. Our framework also offers benefits to dealing with small clusters. Most of graph clustering algorithms (Giesen & Mitsche 2005; Shamir & Tsur 2007; Chen et al. 2014b; Oymak & Hassibi 2011; Ames 2014; Chaudhuri et al. 2012) require = Ω( n) but our algorithm requires a much weaker condition where can be o( n). Corollary 2. If A is CSX then there exist universal constants c 0 c 1 c 2 c 3 c 4 such that the following holds with probability at least 1 max{(mr) 10 n 1 }: Let u = c 1 p(1 q)mn p q p(1 q)mn log 2 n and l = c 2. p q if for all i [r] either i u or i l and if t ( 1 4 p q 3 4 p q) 0 T min i u i 2m { 1 c 0 log n m min 4 log n c (p q) 2 } min i u i 4 log n and max } p(1 q) {c i l i log n 3 p q c 2 p(1 q) log n 3 (p q) 2 Algorithm 1 recovers the true clusters. Remark. We may observe that if the graph only has O(1) small clusters whose sizes have the same order of magnitude then there exists a constant c so that = c log 3 n satisfies the conditions above. Note that the iterative algorithm proposed by Ailon et al. (2013) can recover almost all clusters via a so-called peeling strategy. But it has computational complexity O(n 3 ) which becomes unaffordable when the number of nodes is large say more than In contrast as we show in Section 3.3 the computational complexity of our algorithm is much lower. We now address clustering partially observed graph. The graph is first generated by the GSBM and then A ij is set to? (unknown) with probability 1 µ for each (i j) independently. Clearly if we construct a new graph by setting A ij = 0 when A ij =? we can convert it into the fully observed case. Based on Corollary 1 we have Corollary 3. If A is CSX t ( 1 4 µp µq 3 4 µp µq) 0 T min i i 2m c 3 log n m 1 4 log n and } pmn log n m log n µ(p q) max {c 1 c 2 where c 1 c 2 c 3 are universal constants then Algorithm 1 recovers the true clusters with probability at least 1 (mr) Complexity The computational cost of our graph clustering framework can be much less than the trace-norm based clustering e.g. (Oymak & Hassibi 2011; Ames & Vavasis 2011; Chen et al. 2014b; Jalali et al. 2011) and spectral clustering (Luxburg 2007). Theorem 4. If A is (λ I)-workable or (λ I ϵ)- pseudo-workable and has computational complexity O(f(n)) and memory complexity O(g(n)) then the computational and memory complexity of Algorithm 1 are O ( f( n m )m + (mrl + n i I λ i i ) 3) and O ( g( n m )m + (mrl + n i I λ i i ) 2) respectively. When we use CSX for A and Algorithm 1 runs on a machine with B cores we obtain the following corollary. Corollary 4. If A is CSX then when the conditions in Corollary 1 are satisfied the computational and memory complexity for Algorithm 1 running ( in parallel on a machine with B cores are O ( Bn O 2 m ) + (mrl)2 respectively. ) n 3 Bm + (mrl) 3 2 and Clearly when r l n our algorithm is approximately Bm 2 times faster than applying A on the entire graph. 4. Experiments We now investigate the performance of Algorithm 1 on a variety of simulated and real-world datasets and compare it with other algorithms e.g. spectral clustering (Luxburg 2007) Metis (arypis & umar 1998b) CSX (Chen et al. 2014b) and ESCG that is recently proposed by Liu et al. (2013) and is designed for large graph clustering based on spectral clustering. Except for Metis implemented by arypis & umar (1998b) we implement the other algorithms in Python. The experiments are conducted on a desktop PC with an i7 3.4GHz CPU and 4GB memory Simulations on Synthetic Data The simulated graphs are generated based on the standard stochastic blockmodel with n nodes r clusters and probabilities p and q. We select CSX the trace-norm based algorithm proposed by Chen et al. (2014b) as A. Given the output refined adjacency matrices generated by A or the clustering algorithm shown in (1) the clusters are constructed by simply finding the connected components. In the simulations we repeat each test 10 times and use the averaged ratio of non-exact recovery (RNER) and the wallclock time to measure the performance of each algorithm.
7 p Tra c e-no rm DC: m = DC: m = 6 DC: m = q Running time (s) Tra c e-no rm DC: m= 4 DC: m = 6 DC: m = 8 Figure 2. Comparison between DC and Trace-norm when n = 2000 and m = (Left) Each curve presents the pairs of p and q for which a method succeeds. (Right) The running time. RNER is the fraction of attempts where the output clustering is not exactly the same as the the ground truth. We set l = 1 t = (p + q)/2 and T = 10 in the following experiments. When p and q are unknown we use Algorithm 2 in Chen et al. (2014b) to estimate t. One can also apply the cross validation to determine t such that the incluster edge density of the recovered clusters is maximized. Parameter λ and c C in (1) are set to 1.6 and 5 respectively. For simplicity we use DC Trace-norm and Spectral to denote the proposed Algorithm 1 CSX and spectral clustering respectively. For small graphs with around 1000 to 2000 nodes Trace-norm has been extensively compared with Spectral SLIN (Sibson 1973) etc. (Chen et al. 2014b) and typically outperforms the others. Hence in the first two experiments we mainly compare the performance of DC and Trace-norm. The first experiment compares the performance of DC and Trace-norm when m varies. Corollary 1 is established based on analyzing the conditions when the exact recovery of clustering for each subgraph is achieved which leads to a lower bound of p q that is m times larger than that of Trace-norm. This lower bound is conservative since Theorem 1 shows that the it is possible to achieve exact recovery when A correctly classifies at least λ i i nodes in the ith cluster which means that empirically DC can do better than predicted by Corollary 1. Figure 2 shows the comparison results. It can be observed that p q is approximately 0.2 for Trace-norm and 0.25 for DC when m = 4. Clearly 0.25 for DC is less than 0.4 predicted by Corollary 1. Figure 2 also shows their running time. We observe that DC is almost 7 times faster than Trace-norm when m = 4. Therefore DC is able to recover clusters much faster that Trace-norm without much loss of performance. The second experiment compares the performance of DC and Trace-norm when small clusters exist. Figures 3(a) and (b) plot the ratio of exact recovery and running time of each algorithm against the number of clusters and the size of the smallest cluster respectively. As shown in Corollary 2 DC requires a weaker condition on than Trace-norm which implies that DC will perform better than Trace-norm in the face of small clusters. The simulation results empirically R NE R R NE R DC Tra c e-no rm Number of clusters Size of the smallest cluster DC Tra c e-no rm Running time (s) (a) Running time (s) (b) DC Tra c e-no rm Number of clusters Size of the smallest cluster DC Tra ce -norm Figure 3. The comparison between DC and Trace-norm (n = 1000 p = 0.8 q = 0.2). (a) Nearly equal-sized clusters: the sizes of the first r 1 clusters are [ n ] while the size of the rth r cluster is n (r 1)[ n ]. (b) Small clusters: there exist three r clusters whose sizes are and respectively m n = 4000 q = p n = q = p m Running time (s) m Figure 4. (Left) The fraction of successes for different m and p. (Right) The average running time. validate this conclusion from which we observe that Tracenorm fails when is less than 50 but DC still succeeds. Moreover the running time of DC is again much shorter than that of Trace-norm. Interestingly we observe form Figure 3(b) that the running time of Trace-norm has a sharp rise when is about 40 because it appears more iterations needed to converge. In the third experiment we investigate the performance of DC for different values of p q and m. We repeat each test 10 times. Figure 4 displays the fraction of successes when p m vary and the corresponding averaged running time for different m when n = Obviously when the gap between p and q becomes larger one can increase m to achieve more speedup without affecting the clustering performance. This agrees with our theoretic result shown in Corollary 1 and 4. Figure 5 shows the fraction of successes when p q vary. We observe that in this case when p q 0.15 DC works pretty well. In real applications when the gap between p and q is large enough one can try to apply DC with proper m instead of performing Trace-norm or Spectral on the entire graph.
8 Table 1. Running time (second unit) of different clustering algorithms ( - means out of memory ). n DC Trace-norm Spectral ESCG Table 2. Running time (second unit) of different plant clique algorithms ( - means out of memory ). n DC (Ames & Vavasis 2014) (Ames 2014) q q n = 4000 m = p n = m = p Figure 5. The fraction of successes for different p and q. In the fourth experiment we compare the running time of DC with other methods. Table 1 shows the running times of four algorithms i.e. DC Trace-norm Spectral and ESCG. The simulated graphs are generated with different n but fixed p q r (p = 0.8 q = 0.2 r = 5). All these methods can recover the true clusters in this setting. Obviously DC has the least running time when n Indeed when n is relatively largetrace-norm and Spectral break down due to high memory and computational demand. Since the plant clique model is a special case of the GSBM we also compare DC with two recently proposed plant clique algorithms as shown in Table 2. The graphs are generated with p = 1.0 q = 0.2 and r = 5. The computational advantage of the proposed DC algorithm is obvious Real-world Datasets We now evaluate DC on three real-world datasets namely MNIST (LeCun et al. 1995) Arxiv 1 and DBLP 2. For MNIST samples were randomly selected from the digit images with label 0 1 or 7 and the graph was generated by connecting each sample to its 1000-nearest neighbors where the distance measure is the Euclidean distance. For Arxiv we used the texts from the years to build the graph whose nodes and edges represent texts and Table 3. The performance of different algorithms on the real world datasets MNIST Arxiv and DBLP which is measured by computing the in-cluster and cross-cluster edge densities. MNIST (10000 nodes edges) Algorithm DC Metis Spectral ESCG In-cluster (10 2 ) Cross-cluster (10 2 ) Arxiv (12480 nodes edges) Algorithm DC Metis Spectral ESCG In-cluster (10 4 ) Cross-cluster (10 4 ) DBLP (24599 nodes edges) Algorithm DC Metis Spectral ESCG In-cluster (10 4 ) Cross-cluster (10 4 ) citations respectively. For DBLP we randomly selected a subset from it and constructed a co-author-graph where each node corresponds to an author and each edge corresponds to a co-authorship between two authors. The setting of Algorithm 1 is different from that in the simulations. We choose Spectral or Metis as A depending on whose result has a higher in-cluster edge density. Given the output of (1) performed on the fuse graph we apply Metis to classify the nodes into a specific number of clusters. Due to the memory limitation for two of the baseline algorithms namely Spectral and Metis we apply a similar algorithm as Algorithm 1 but replacing (1) with Spectral and Metis respectively. For fair comparison all the algorithms are forced to partition the nodes of MNIST into 3 clusters and partition the nodes of Arxiv and DBLP into 15 clusters. The performance of each algorithm is measured by computing the in-cluster and cross-cluster edge densities (Table 3). When the graph is dense (e.g. MNIST) DC and ESCG have similar performance. But when the graph is sparse (e.g. DBLP) Spectral and ESCG tend to classify most of the nodes into one big cluster so that the corresponding in-cluster densities are low. In general our algorithm DC outperforms all three baselines on these datasets.
9 Acknowledgments This work is partially supported by the Ministry of Education of Singapore AcRF Tier Two grants R and R and A*STAR Public Sector Funding R References Ailon N. Chen Y. and Xu H. Breaking the small cluster barrier of graph clustering. In ICML Ames Brendan P. W. Guaranteed clustering and biclustering via semidefinite programming. Mathematical Programming 143(1-2): Ames Brendan P. W. and Vavasis Stephen A. Nuclear norm minimization for the planted clique and biclique problems. Mathematical Programming 129(1):69 89 September Ames Brendan P. W. and Vavasis Stephen A. Convex optimization for the planted k-disjoint-clique problem. Mathematical Programming 143(1-2): Chaudhuri. Graham F. C. and Tsiatas A. Spectral clustering of graphs with general degrees in the extended planted partition model. In COLT Chen W. Song Y. Bai H. Lin C. and Chang E. Y. Parallel spectral clustering in distributed systems. IEEE Transactions on Pattern Analysis and Machine Intelligence (TPAMI) 33(3): Chen Y. Lim S. H. and Xu H. Weighted graph clustering with non-uniform uncertainties. In ICML 2014a. Chen Y. Sanghavi S. and Xu H. Improved graph clustering. IEEE Transactions on Information Theory 60(10): b. Condon A. and arp R. Algorithms for graph partitioning on the planted partition model. Random Structures Algorithms 18(2): Drineas P. and Mahoney M. W. On the Nyström method for approximating a gram matrix for improved kernelbased learning. Journal of Machine Learning Research 6: Fowlkes C. Belongie S. Chung F. and Malik J. Spectral grouping using the Nyström method. IEEE Transactions on Pattern Analysis and Machine Intelligence (TPAMI) 26(2): Giesen J. and Mitsche D. Reconstructing many partitions using spectral techniques. Fundamentals of Computation Theory pp Goldenberg A. Zheng A. X. Fienberg S. E. and Airoldi E. M. A survey of statistical network models. Foundations and Trends in Machine Learning 2(2): Günnemann S. Färber I. Boden B. and Seidl T. Subspace clustering meets dense subgraph mining: A synthesis of two paradigms. In ICDM Holland P. Laskey. and Leinhardt S. Stochastic blockmodels: Some first steps. Social Networks 5: Jalali A. Chen Y. Sanghavi S. and Xu H. Clustering partially observed graphs via convex optimization. In ICML arypis G. and umar V. A fast and high quality multilevel scheme for partitioning irregular graphs. SIAM Journal on Scientific Computing 20(1): a. arypis G. and umar V. Multilevel k-way partitioning scheme for irregular graphs. Journal of Parallel and Distributed Computing 48: b. LeCun Y. Jackel L. Bottou L. Brunot A. Cortes C. Denker J. Drucker H. Guyon I. Müller U. Säckinger E. Simard P. and Vapnik V. Comparison of learning algorithms for handwritten digit recognition. In International Conference on Artificial Neural Networks pp Liu J. Wang C. Danilevsky M. and Han J. Large-scale spectral clustering on graphs. In IJCAI Luxburg U. A tutorial on spectral clustering. Statistics and Computing 17(4): Macropol. and Singh A. Scalable discovery of best clusters on large graphs. Proceedings of the VLDB Endowment 3(1-2): Oymak S. and Hassibi B. Finding dense clusters via low rank + sparse decomposition. Arxiv preprint Rohe arl Chatterjee Sourav and Yu Bin. Spectral clustering and the high-dimensional stochastic blockmodel. The Annals of Statistics 39(4): Shamir R. and Tsur D. Improved algorithms for the random cluster graph model. Random Structures Algorithms 31(4): Sibson R. Slink: an optimally efficient algorithm for the single-link cluster method. The Computer Journal 16 (1): Watts D. J. and Strogatz S. H. Collective dynamics of small-world networks. Nature 393(6684):
10 Whang J. J. Sui X. and Dhillon I. S. Scalable and memory-efficient clustering of large-scale social networks. In ICDM Yan D. Huang L. and Jordan M. I. Fast approximate spectral clustering. In DD pp
Spectral Clustering and Community Detection in Labeled Graphs
 Spectral Clustering and Community Detection in Labeled Graphs Brandon Fain, Stavros Sintos, Nisarg Raval Machine Learning (CompSci 571D / STA 561D) December 7, 2015 {btfain, nisarg, ssintos} at cs.duke.edu
Spectral Clustering and Community Detection in Labeled Graphs Brandon Fain, Stavros Sintos, Nisarg Raval Machine Learning (CompSci 571D / STA 561D) December 7, 2015 {btfain, nisarg, ssintos} at cs.duke.edu
Limitations of Matrix Completion via Trace Norm Minimization
 Limitations of Matrix Completion via Trace Norm Minimization ABSTRACT Xiaoxiao Shi Computer Science Department University of Illinois at Chicago xiaoxiao@cs.uic.edu In recent years, compressive sensing
Limitations of Matrix Completion via Trace Norm Minimization ABSTRACT Xiaoxiao Shi Computer Science Department University of Illinois at Chicago xiaoxiao@cs.uic.edu In recent years, compressive sensing
Learning to Match. Jun Xu, Zhengdong Lu, Tianqi Chen, Hang Li
 Learning to Match Jun Xu, Zhengdong Lu, Tianqi Chen, Hang Li 1. Introduction The main tasks in many applications can be formalized as matching between heterogeneous objects, including search, recommendation,
Learning to Match Jun Xu, Zhengdong Lu, Tianqi Chen, Hang Li 1. Introduction The main tasks in many applications can be formalized as matching between heterogeneous objects, including search, recommendation,
Scalable Clustering of Signed Networks Using Balance Normalized Cut
 Scalable Clustering of Signed Networks Using Balance Normalized Cut Kai-Yang Chiang,, Inderjit S. Dhillon The 21st ACM International Conference on Information and Knowledge Management (CIKM 2012) Oct.
Scalable Clustering of Signed Networks Using Balance Normalized Cut Kai-Yang Chiang,, Inderjit S. Dhillon The 21st ACM International Conference on Information and Knowledge Management (CIKM 2012) Oct.
Mathematical and Algorithmic Foundations Linear Programming and Matchings
 Adavnced Algorithms Lectures Mathematical and Algorithmic Foundations Linear Programming and Matchings Paul G. Spirakis Department of Computer Science University of Patras and Liverpool Paul G. Spirakis
Adavnced Algorithms Lectures Mathematical and Algorithmic Foundations Linear Programming and Matchings Paul G. Spirakis Department of Computer Science University of Patras and Liverpool Paul G. Spirakis
On the Relationships between Zero Forcing Numbers and Certain Graph Coverings
 On the Relationships between Zero Forcing Numbers and Certain Graph Coverings Fatemeh Alinaghipour Taklimi, Shaun Fallat 1,, Karen Meagher 2 Department of Mathematics and Statistics, University of Regina,
On the Relationships between Zero Forcing Numbers and Certain Graph Coverings Fatemeh Alinaghipour Taklimi, Shaun Fallat 1,, Karen Meagher 2 Department of Mathematics and Statistics, University of Regina,
Bipartite graphs unique perfect matching.
 Generation of graphs Bipartite graphs unique perfect matching. In this section, we assume G = (V, E) bipartite connected graph. The following theorem states that if G has unique perfect matching, then
Generation of graphs Bipartite graphs unique perfect matching. In this section, we assume G = (V, E) bipartite connected graph. The following theorem states that if G has unique perfect matching, then
Clustering. SC4/SM4 Data Mining and Machine Learning, Hilary Term 2017 Dino Sejdinovic
 Clustering SC4/SM4 Data Mining and Machine Learning, Hilary Term 2017 Dino Sejdinovic Clustering is one of the fundamental and ubiquitous tasks in exploratory data analysis a first intuition about the
Clustering SC4/SM4 Data Mining and Machine Learning, Hilary Term 2017 Dino Sejdinovic Clustering is one of the fundamental and ubiquitous tasks in exploratory data analysis a first intuition about the
Non-exhaustive, Overlapping k-means
 Non-exhaustive, Overlapping k-means J. J. Whang, I. S. Dhilon, and D. F. Gleich Teresa Lebair University of Maryland, Baltimore County October 29th, 2015 Teresa Lebair UMBC 1/38 Outline Introduction NEO-K-Means
Non-exhaustive, Overlapping k-means J. J. Whang, I. S. Dhilon, and D. F. Gleich Teresa Lebair University of Maryland, Baltimore County October 29th, 2015 Teresa Lebair UMBC 1/38 Outline Introduction NEO-K-Means
Spectral Methods for Network Community Detection and Graph Partitioning
 Spectral Methods for Network Community Detection and Graph Partitioning M. E. J. Newman Department of Physics, University of Michigan Presenters: Yunqi Guo Xueyin Yu Yuanqi Li 1 Outline: Community Detection
Spectral Methods for Network Community Detection and Graph Partitioning M. E. J. Newman Department of Physics, University of Michigan Presenters: Yunqi Guo Xueyin Yu Yuanqi Li 1 Outline: Community Detection
Detecting Burnscar from Hyperspectral Imagery via Sparse Representation with Low-Rank Interference
 Detecting Burnscar from Hyperspectral Imagery via Sparse Representation with Low-Rank Interference Minh Dao 1, Xiang Xiang 1, Bulent Ayhan 2, Chiman Kwan 2, Trac D. Tran 1 Johns Hopkins Univeristy, 3400
Detecting Burnscar from Hyperspectral Imagery via Sparse Representation with Low-Rank Interference Minh Dao 1, Xiang Xiang 1, Bulent Ayhan 2, Chiman Kwan 2, Trac D. Tran 1 Johns Hopkins Univeristy, 3400
An Empirical Analysis of Communities in Real-World Networks
 An Empirical Analysis of Communities in Real-World Networks Chuan Sheng Foo Computer Science Department Stanford University csfoo@cs.stanford.edu ABSTRACT Little work has been done on the characterization
An Empirical Analysis of Communities in Real-World Networks Chuan Sheng Foo Computer Science Department Stanford University csfoo@cs.stanford.edu ABSTRACT Little work has been done on the characterization
c 2004 Society for Industrial and Applied Mathematics
 SIAM J. MATRIX ANAL. APPL. Vol. 26, No. 2, pp. 390 399 c 2004 Society for Industrial and Applied Mathematics HERMITIAN MATRICES, EIGENVALUE MULTIPLICITIES, AND EIGENVECTOR COMPONENTS CHARLES R. JOHNSON
SIAM J. MATRIX ANAL. APPL. Vol. 26, No. 2, pp. 390 399 c 2004 Society for Industrial and Applied Mathematics HERMITIAN MATRICES, EIGENVALUE MULTIPLICITIES, AND EIGENVECTOR COMPONENTS CHARLES R. JOHNSON
A Geometric Analysis of Subspace Clustering with Outliers
 A Geometric Analysis of Subspace Clustering with Outliers Mahdi Soltanolkotabi and Emmanuel Candés Stanford University Fundamental Tool in Data Mining : PCA Fundamental Tool in Data Mining : PCA Subspace
A Geometric Analysis of Subspace Clustering with Outliers Mahdi Soltanolkotabi and Emmanuel Candés Stanford University Fundamental Tool in Data Mining : PCA Fundamental Tool in Data Mining : PCA Subspace
Disjoint directed cycles
 Disjoint directed cycles Noga Alon Abstract It is shown that there exists a positive ɛ so that for any integer k, every directed graph with minimum outdegree at least k contains at least ɛk vertex disjoint
Disjoint directed cycles Noga Alon Abstract It is shown that there exists a positive ɛ so that for any integer k, every directed graph with minimum outdegree at least k contains at least ɛk vertex disjoint
Module 11. Directed Graphs. Contents
 Module 11 Directed Graphs Contents 11.1 Basic concepts......................... 256 Underlying graph of a digraph................ 257 Out-degrees and in-degrees.................. 258 Isomorphism..........................
Module 11 Directed Graphs Contents 11.1 Basic concepts......................... 256 Underlying graph of a digraph................ 257 Out-degrees and in-degrees.................. 258 Isomorphism..........................
Efficient Semi-supervised Spectral Co-clustering with Constraints
 2010 IEEE International Conference on Data Mining Efficient Semi-supervised Spectral Co-clustering with Constraints Xiaoxiao Shi, Wei Fan, Philip S. Yu Department of Computer Science, University of Illinois
2010 IEEE International Conference on Data Mining Efficient Semi-supervised Spectral Co-clustering with Constraints Xiaoxiao Shi, Wei Fan, Philip S. Yu Department of Computer Science, University of Illinois
Lecture 2 September 3
 EE 381V: Large Scale Optimization Fall 2012 Lecture 2 September 3 Lecturer: Caramanis & Sanghavi Scribe: Hongbo Si, Qiaoyang Ye 2.1 Overview of the last Lecture The focus of the last lecture was to give
EE 381V: Large Scale Optimization Fall 2012 Lecture 2 September 3 Lecturer: Caramanis & Sanghavi Scribe: Hongbo Si, Qiaoyang Ye 2.1 Overview of the last Lecture The focus of the last lecture was to give
A noninformative Bayesian approach to small area estimation
 A noninformative Bayesian approach to small area estimation Glen Meeden School of Statistics University of Minnesota Minneapolis, MN 55455 glen@stat.umn.edu September 2001 Revised May 2002 Research supported
A noninformative Bayesian approach to small area estimation Glen Meeden School of Statistics University of Minnesota Minneapolis, MN 55455 glen@stat.umn.edu September 2001 Revised May 2002 Research supported
General properties of staircase and convex dual feasible functions
 General properties of staircase and convex dual feasible functions JÜRGEN RIETZ, CLÁUDIO ALVES, J. M. VALÉRIO de CARVALHO Centro de Investigação Algoritmi da Universidade do Minho, Escola de Engenharia
General properties of staircase and convex dual feasible functions JÜRGEN RIETZ, CLÁUDIO ALVES, J. M. VALÉRIO de CARVALHO Centro de Investigação Algoritmi da Universidade do Minho, Escola de Engenharia
CHAPTER 6 IDENTIFICATION OF CLUSTERS USING VISUAL VALIDATION VAT ALGORITHM
 96 CHAPTER 6 IDENTIFICATION OF CLUSTERS USING VISUAL VALIDATION VAT ALGORITHM Clustering is the process of combining a set of relevant information in the same group. In this process KM algorithm plays
96 CHAPTER 6 IDENTIFICATION OF CLUSTERS USING VISUAL VALIDATION VAT ALGORITHM Clustering is the process of combining a set of relevant information in the same group. In this process KM algorithm plays
Leveraging Set Relations in Exact Set Similarity Join
 Leveraging Set Relations in Exact Set Similarity Join Xubo Wang, Lu Qin, Xuemin Lin, Ying Zhang, and Lijun Chang University of New South Wales, Australia University of Technology Sydney, Australia {xwang,lxue,ljchang}@cse.unsw.edu.au,
Leveraging Set Relations in Exact Set Similarity Join Xubo Wang, Lu Qin, Xuemin Lin, Ying Zhang, and Lijun Chang University of New South Wales, Australia University of Technology Sydney, Australia {xwang,lxue,ljchang}@cse.unsw.edu.au,
Advanced Algorithms Class Notes for Monday, October 23, 2012 Min Ye, Mingfu Shao, and Bernard Moret
 Advanced Algorithms Class Notes for Monday, October 23, 2012 Min Ye, Mingfu Shao, and Bernard Moret Greedy Algorithms (continued) The best known application where the greedy algorithm is optimal is surely
Advanced Algorithms Class Notes for Monday, October 23, 2012 Min Ye, Mingfu Shao, and Bernard Moret Greedy Algorithms (continued) The best known application where the greedy algorithm is optimal is surely
Random projection for non-gaussian mixture models
 Random projection for non-gaussian mixture models Győző Gidófalvi Department of Computer Science and Engineering University of California, San Diego La Jolla, CA 92037 gyozo@cs.ucsd.edu Abstract Recently,
Random projection for non-gaussian mixture models Győző Gidófalvi Department of Computer Science and Engineering University of California, San Diego La Jolla, CA 92037 gyozo@cs.ucsd.edu Abstract Recently,
Discrete mathematics , Fall Instructor: prof. János Pach
 Discrete mathematics 2016-2017, Fall Instructor: prof. János Pach - covered material - Lecture 1. Counting problems To read: [Lov]: 1.2. Sets, 1.3. Number of subsets, 1.5. Sequences, 1.6. Permutations,
Discrete mathematics 2016-2017, Fall Instructor: prof. János Pach - covered material - Lecture 1. Counting problems To read: [Lov]: 1.2. Sets, 1.3. Number of subsets, 1.5. Sequences, 1.6. Permutations,
K s,t -saturated bipartite graphs
 K s,t -saturated bipartite graphs Wenying Gan Dániel Korándi Benny Sudakov Abstract An n-by-n bipartite graph is H-saturated if the addition of any missing edge between its two parts creates a new copy
K s,t -saturated bipartite graphs Wenying Gan Dániel Korándi Benny Sudakov Abstract An n-by-n bipartite graph is H-saturated if the addition of any missing edge between its two parts creates a new copy
Effectiveness of Sparse Features: An Application of Sparse PCA
 000 001 002 003 004 005 006 007 008 009 010 011 012 013 014 015 016 017 018 019 020 021 022 023 024 025 026 027 028 029 030 031 032 033 034 035 036 037 038 039 040 041 042 043 044 045 046 047 048 049 050
000 001 002 003 004 005 006 007 008 009 010 011 012 013 014 015 016 017 018 019 020 021 022 023 024 025 026 027 028 029 030 031 032 033 034 035 036 037 038 039 040 041 042 043 044 045 046 047 048 049 050
The strong chromatic number of a graph
 The strong chromatic number of a graph Noga Alon Abstract It is shown that there is an absolute constant c with the following property: For any two graphs G 1 = (V, E 1 ) and G 2 = (V, E 2 ) on the same
The strong chromatic number of a graph Noga Alon Abstract It is shown that there is an absolute constant c with the following property: For any two graphs G 1 = (V, E 1 ) and G 2 = (V, E 2 ) on the same
Outlier Pursuit: Robust PCA and Collaborative Filtering
 Outlier Pursuit: Robust PCA and Collaborative Filtering Huan Xu Dept. of Mechanical Engineering & Dept. of Mathematics National University of Singapore Joint w/ Constantine Caramanis, Yudong Chen, Sujay
Outlier Pursuit: Robust PCA and Collaborative Filtering Huan Xu Dept. of Mechanical Engineering & Dept. of Mathematics National University of Singapore Joint w/ Constantine Caramanis, Yudong Chen, Sujay
Unsupervised Learning and Clustering
 Unsupervised Learning and Clustering Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Spring 2008 CS 551, Spring 2008 c 2008, Selim Aksoy (Bilkent University)
Unsupervised Learning and Clustering Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Spring 2008 CS 551, Spring 2008 c 2008, Selim Aksoy (Bilkent University)
Big Data Analytics. Special Topics for Computer Science CSE CSE Feb 11
 Big Data Analytics Special Topics for Computer Science CSE 4095-001 CSE 5095-005 Feb 11 Fei Wang Associate Professor Department of Computer Science and Engineering fei_wang@uconn.edu Clustering II Spectral
Big Data Analytics Special Topics for Computer Science CSE 4095-001 CSE 5095-005 Feb 11 Fei Wang Associate Professor Department of Computer Science and Engineering fei_wang@uconn.edu Clustering II Spectral
An efficient algorithm for rank-1 sparse PCA
 An efficient algorithm for rank- sparse PCA Yunlong He Georgia Institute of Technology School of Mathematics heyunlong@gatech.edu Renato Monteiro Georgia Institute of Technology School of Industrial &
An efficient algorithm for rank- sparse PCA Yunlong He Georgia Institute of Technology School of Mathematics heyunlong@gatech.edu Renato Monteiro Georgia Institute of Technology School of Industrial &
On Covering a Graph Optimally with Induced Subgraphs
 On Covering a Graph Optimally with Induced Subgraphs Shripad Thite April 1, 006 Abstract We consider the problem of covering a graph with a given number of induced subgraphs so that the maximum number
On Covering a Graph Optimally with Induced Subgraphs Shripad Thite April 1, 006 Abstract We consider the problem of covering a graph with a given number of induced subgraphs so that the maximum number
3 No-Wait Job Shops with Variable Processing Times
 3 No-Wait Job Shops with Variable Processing Times In this chapter we assume that, on top of the classical no-wait job shop setting, we are given a set of processing times for each operation. We may select
3 No-Wait Job Shops with Variable Processing Times In this chapter we assume that, on top of the classical no-wait job shop setting, we are given a set of processing times for each operation. We may select
Dynamic Clustering of Data with Modified K-Means Algorithm
 2012 International Conference on Information and Computer Networks (ICICN 2012) IPCSIT vol. 27 (2012) (2012) IACSIT Press, Singapore Dynamic Clustering of Data with Modified K-Means Algorithm Ahamed Shafeeq
2012 International Conference on Information and Computer Networks (ICICN 2012) IPCSIT vol. 27 (2012) (2012) IACSIT Press, Singapore Dynamic Clustering of Data with Modified K-Means Algorithm Ahamed Shafeeq
Exact Recovery in Stochastic Block Models
 Eact Recovery in Stochastic Block Models Ben Eysenbach May 11, 2016 1 Introduction Graphs are a natural way of capturing interactions between objects, such as phone calls between friends, binding between
Eact Recovery in Stochastic Block Models Ben Eysenbach May 11, 2016 1 Introduction Graphs are a natural way of capturing interactions between objects, such as phone calls between friends, binding between
E-Companion: On Styles in Product Design: An Analysis of US. Design Patents
 E-Companion: On Styles in Product Design: An Analysis of US Design Patents 1 PART A: FORMALIZING THE DEFINITION OF STYLES A.1 Styles as categories of designs of similar form Our task involves categorizing
E-Companion: On Styles in Product Design: An Analysis of US Design Patents 1 PART A: FORMALIZING THE DEFINITION OF STYLES A.1 Styles as categories of designs of similar form Our task involves categorizing
IEEE TRANSACTIONS ON INFORMATION THEORY, VOL. 53, NO. 2, FEBRUARY /$ IEEE
 IEEE TRANSACTIONS ON INFORMATION THEORY, VOL. 53, NO. 2, FEBRUARY 2007 599 Results on Punctured Low-Density Parity-Check Codes and Improved Iterative Decoding Techniques Hossein Pishro-Nik, Member, IEEE,
IEEE TRANSACTIONS ON INFORMATION THEORY, VOL. 53, NO. 2, FEBRUARY 2007 599 Results on Punctured Low-Density Parity-Check Codes and Improved Iterative Decoding Techniques Hossein Pishro-Nik, Member, IEEE,
Unsupervised Learning and Clustering
 Unsupervised Learning and Clustering Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Spring 2009 CS 551, Spring 2009 c 2009, Selim Aksoy (Bilkent University)
Unsupervised Learning and Clustering Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Spring 2009 CS 551, Spring 2009 c 2009, Selim Aksoy (Bilkent University)
Multi-label Classification. Jingzhou Liu Dec
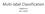 Multi-label Classification Jingzhou Liu Dec. 6 2016 Introduction Multi-class problem, Training data (x $, y $ ) ( ), x $ X R., y $ Y = 1,2,, L Learn a mapping f: X Y Each instance x $ is associated with
Multi-label Classification Jingzhou Liu Dec. 6 2016 Introduction Multi-class problem, Training data (x $, y $ ) ( ), x $ X R., y $ Y = 1,2,, L Learn a mapping f: X Y Each instance x $ is associated with
An efficient algorithm for sparse PCA
 An efficient algorithm for sparse PCA Yunlong He Georgia Institute of Technology School of Mathematics heyunlong@gatech.edu Renato D.C. Monteiro Georgia Institute of Technology School of Industrial & System
An efficient algorithm for sparse PCA Yunlong He Georgia Institute of Technology School of Mathematics heyunlong@gatech.edu Renato D.C. Monteiro Georgia Institute of Technology School of Industrial & System
Restricted edge connectivity and restricted connectivity of graphs
 Restricted edge connectivity and restricted connectivity of graphs Litao Guo School of Applied Mathematics Xiamen University of Technology Xiamen Fujian 361024 P.R.China ltguo2012@126.com Xiaofeng Guo
Restricted edge connectivity and restricted connectivity of graphs Litao Guo School of Applied Mathematics Xiamen University of Technology Xiamen Fujian 361024 P.R.China ltguo2012@126.com Xiaofeng Guo
Treewidth and graph minors
 Treewidth and graph minors Lectures 9 and 10, December 29, 2011, January 5, 2012 We shall touch upon the theory of Graph Minors by Robertson and Seymour. This theory gives a very general condition under
Treewidth and graph minors Lectures 9 and 10, December 29, 2011, January 5, 2012 We shall touch upon the theory of Graph Minors by Robertson and Seymour. This theory gives a very general condition under
Notes for Lecture 24
 U.C. Berkeley CS170: Intro to CS Theory Handout N24 Professor Luca Trevisan December 4, 2001 Notes for Lecture 24 1 Some NP-complete Numerical Problems 1.1 Subset Sum The Subset Sum problem is defined
U.C. Berkeley CS170: Intro to CS Theory Handout N24 Professor Luca Trevisan December 4, 2001 Notes for Lecture 24 1 Some NP-complete Numerical Problems 1.1 Subset Sum The Subset Sum problem is defined
Comparing the strength of query types in property testing: The case of testing k-colorability
 Comparing the strength of query types in property testing: The case of testing k-colorability Ido Ben-Eliezer Tali Kaufman Michael Krivelevich Dana Ron Abstract We study the power of four query models
Comparing the strength of query types in property testing: The case of testing k-colorability Ido Ben-Eliezer Tali Kaufman Michael Krivelevich Dana Ron Abstract We study the power of four query models
Vertical decomposition of a lattice using clique separators
 Vertical decomposition of a lattice using clique separators Anne Berry, Romain Pogorelcnik, Alain Sigayret LIMOS UMR CNRS 6158 Ensemble Scientifique des Cézeaux Université Blaise Pascal, F-63 173 Aubière,
Vertical decomposition of a lattice using clique separators Anne Berry, Romain Pogorelcnik, Alain Sigayret LIMOS UMR CNRS 6158 Ensemble Scientifique des Cézeaux Université Blaise Pascal, F-63 173 Aubière,
Graph Based Image Segmentation
 AUTOMATYKA 2011 Tom 15 Zeszyt 3 Anna Fabijañska* Graph Based Image Segmentation 1. Introduction Image segmentation is one of the fundamental problems in machine vision. In general it aims at extracting
AUTOMATYKA 2011 Tom 15 Zeszyt 3 Anna Fabijañska* Graph Based Image Segmentation 1. Introduction Image segmentation is one of the fundamental problems in machine vision. In general it aims at extracting
Faster parameterized algorithms for Minimum Fill-In
 Faster parameterized algorithms for Minimum Fill-In Hans L. Bodlaender Pinar Heggernes Yngve Villanger Technical Report UU-CS-2008-042 December 2008 Department of Information and Computing Sciences Utrecht
Faster parameterized algorithms for Minimum Fill-In Hans L. Bodlaender Pinar Heggernes Yngve Villanger Technical Report UU-CS-2008-042 December 2008 Department of Information and Computing Sciences Utrecht
A NOTE ON THE NUMBER OF DOMINATING SETS OF A GRAPH
 A NOTE ON THE NUMBER OF DOMINATING SETS OF A GRAPH STEPHAN WAGNER Abstract. In a recent article by Bród and Skupień, sharp upper and lower bounds for the number of dominating sets in a tree were determined.
A NOTE ON THE NUMBER OF DOMINATING SETS OF A GRAPH STEPHAN WAGNER Abstract. In a recent article by Bród and Skupień, sharp upper and lower bounds for the number of dominating sets in a tree were determined.
6 Randomized rounding of semidefinite programs
 6 Randomized rounding of semidefinite programs We now turn to a new tool which gives substantially improved performance guarantees for some problems We now show how nonlinear programming relaxations can
6 Randomized rounding of semidefinite programs We now turn to a new tool which gives substantially improved performance guarantees for some problems We now show how nonlinear programming relaxations can
Foundations of Computing
 Foundations of Computing Darmstadt University of Technology Dept. Computer Science Winter Term 2005 / 2006 Copyright c 2004 by Matthias Müller-Hannemann and Karsten Weihe All rights reserved http://www.algo.informatik.tu-darmstadt.de/
Foundations of Computing Darmstadt University of Technology Dept. Computer Science Winter Term 2005 / 2006 Copyright c 2004 by Matthias Müller-Hannemann and Karsten Weihe All rights reserved http://www.algo.informatik.tu-darmstadt.de/
A Review of K-mean Algorithm
 A Review of K-mean Algorithm Jyoti Yadav #1, Monika Sharma *2 1 PG Student, CSE Department, M.D.U Rohtak, Haryana, India 2 Assistant Professor, IT Department, M.D.U Rohtak, Haryana, India Abstract Cluster
A Review of K-mean Algorithm Jyoti Yadav #1, Monika Sharma *2 1 PG Student, CSE Department, M.D.U Rohtak, Haryana, India 2 Assistant Professor, IT Department, M.D.U Rohtak, Haryana, India Abstract Cluster
1. Introduction. performance of numerical methods. complexity bounds. structural convex optimization. course goals and topics
 1. Introduction EE 546, Univ of Washington, Spring 2016 performance of numerical methods complexity bounds structural convex optimization course goals and topics 1 1 Some course info Welcome to EE 546!
1. Introduction EE 546, Univ of Washington, Spring 2016 performance of numerical methods complexity bounds structural convex optimization course goals and topics 1 1 Some course info Welcome to EE 546!
Semi-Supervised Clustering with Partial Background Information
 Semi-Supervised Clustering with Partial Background Information Jing Gao Pang-Ning Tan Haibin Cheng Abstract Incorporating background knowledge into unsupervised clustering algorithms has been the subject
Semi-Supervised Clustering with Partial Background Information Jing Gao Pang-Ning Tan Haibin Cheng Abstract Incorporating background knowledge into unsupervised clustering algorithms has been the subject
Hierarchical Multi level Approach to graph clustering
 Hierarchical Multi level Approach to graph clustering by: Neda Shahidi neda@cs.utexas.edu Cesar mantilla, cesar.mantilla@mail.utexas.edu Advisor: Dr. Inderjit Dhillon Introduction Data sets can be presented
Hierarchical Multi level Approach to graph clustering by: Neda Shahidi neda@cs.utexas.edu Cesar mantilla, cesar.mantilla@mail.utexas.edu Advisor: Dr. Inderjit Dhillon Introduction Data sets can be presented
Robust Face Recognition via Sparse Representation
 Robust Face Recognition via Sparse Representation Panqu Wang Department of Electrical and Computer Engineering University of California, San Diego La Jolla, CA 92092 pawang@ucsd.edu Can Xu Department of
Robust Face Recognition via Sparse Representation Panqu Wang Department of Electrical and Computer Engineering University of California, San Diego La Jolla, CA 92092 pawang@ucsd.edu Can Xu Department of
Graph Laplacian Kernels for Object Classification from a Single Example
 Graph Laplacian Kernels for Object Classification from a Single Example Hong Chang & Dit-Yan Yeung Department of Computer Science, Hong Kong University of Science and Technology {hongch,dyyeung}@cs.ust.hk
Graph Laplacian Kernels for Object Classification from a Single Example Hong Chang & Dit-Yan Yeung Department of Computer Science, Hong Kong University of Science and Technology {hongch,dyyeung}@cs.ust.hk
/ Approximation Algorithms Lecturer: Michael Dinitz Topic: Linear Programming Date: 2/24/15 Scribe: Runze Tang
 600.469 / 600.669 Approximation Algorithms Lecturer: Michael Dinitz Topic: Linear Programming Date: 2/24/15 Scribe: Runze Tang 9.1 Linear Programming Suppose we are trying to approximate a minimization
600.469 / 600.669 Approximation Algorithms Lecturer: Michael Dinitz Topic: Linear Programming Date: 2/24/15 Scribe: Runze Tang 9.1 Linear Programming Suppose we are trying to approximate a minimization
Fast Efficient Clustering Algorithm for Balanced Data
 Vol. 5, No. 6, 214 Fast Efficient Clustering Algorithm for Balanced Data Adel A. Sewisy Faculty of Computer and Information, Assiut University M. H. Marghny Faculty of Computer and Information, Assiut
Vol. 5, No. 6, 214 Fast Efficient Clustering Algorithm for Balanced Data Adel A. Sewisy Faculty of Computer and Information, Assiut University M. H. Marghny Faculty of Computer and Information, Assiut
RECOVERY OF PARTIALLY OBSERVED DATA APPEARING IN CLUSTERS. Sunrita Poddar, Mathews Jacob
 RECOVERY OF PARTIALLY OBSERVED DATA APPEARING IN CLUSTERS Sunrita Poddar, Mathews Jacob Department of Electrical and Computer Engineering The University of Iowa, IA, USA ABSTRACT We propose a matrix completion
RECOVERY OF PARTIALLY OBSERVED DATA APPEARING IN CLUSTERS Sunrita Poddar, Mathews Jacob Department of Electrical and Computer Engineering The University of Iowa, IA, USA ABSTRACT We propose a matrix completion
CHAPTER 6 MODIFIED FUZZY TECHNIQUES BASED IMAGE SEGMENTATION
 CHAPTER 6 MODIFIED FUZZY TECHNIQUES BASED IMAGE SEGMENTATION 6.1 INTRODUCTION Fuzzy logic based computational techniques are becoming increasingly important in the medical image analysis arena. The significant
CHAPTER 6 MODIFIED FUZZY TECHNIQUES BASED IMAGE SEGMENTATION 6.1 INTRODUCTION Fuzzy logic based computational techniques are becoming increasingly important in the medical image analysis arena. The significant
Faster parameterized algorithms for Minimum Fill-In
 Faster parameterized algorithms for Minimum Fill-In Hans L. Bodlaender Pinar Heggernes Yngve Villanger Abstract We present two parameterized algorithms for the Minimum Fill-In problem, also known as Chordal
Faster parameterized algorithms for Minimum Fill-In Hans L. Bodlaender Pinar Heggernes Yngve Villanger Abstract We present two parameterized algorithms for the Minimum Fill-In problem, also known as Chordal
Complete Cototal Domination
 Chapter 5 Complete Cototal Domination Number of a Graph Published in Journal of Scientific Research Vol. () (2011), 547-555 (Bangladesh). 64 ABSTRACT Let G = (V,E) be a graph. A dominating set D V is said
Chapter 5 Complete Cototal Domination Number of a Graph Published in Journal of Scientific Research Vol. () (2011), 547-555 (Bangladesh). 64 ABSTRACT Let G = (V,E) be a graph. A dominating set D V is said
Online Stochastic Matching CMSC 858F: Algorithmic Game Theory Fall 2010
 Online Stochastic Matching CMSC 858F: Algorithmic Game Theory Fall 2010 Barna Saha, Vahid Liaghat Abstract This summary is mostly based on the work of Saberi et al. [1] on online stochastic matching problem
Online Stochastic Matching CMSC 858F: Algorithmic Game Theory Fall 2010 Barna Saha, Vahid Liaghat Abstract This summary is mostly based on the work of Saberi et al. [1] on online stochastic matching problem
arxiv: v1 [math.co] 28 Sep 2010
![arxiv: v1 [math.co] 28 Sep 2010 arxiv: v1 [math.co] 28 Sep 2010](/thumbs/85/91865330.jpg) Densities of Minor-Closed Graph Families David Eppstein Computer Science Department University of California, Irvine Irvine, California, USA arxiv:1009.5633v1 [math.co] 28 Sep 2010 September 3, 2018 Abstract
Densities of Minor-Closed Graph Families David Eppstein Computer Science Department University of California, Irvine Irvine, California, USA arxiv:1009.5633v1 [math.co] 28 Sep 2010 September 3, 2018 Abstract
Clustering: Classic Methods and Modern Views
 Clustering: Classic Methods and Modern Views Marina Meilă University of Washington mmp@stat.washington.edu June 22, 2015 Lorentz Center Workshop on Clusters, Games and Axioms Outline Paradigms for clustering
Clustering: Classic Methods and Modern Views Marina Meilă University of Washington mmp@stat.washington.edu June 22, 2015 Lorentz Center Workshop on Clusters, Games and Axioms Outline Paradigms for clustering
On the Max Coloring Problem
 On the Max Coloring Problem Leah Epstein Asaf Levin May 22, 2010 Abstract We consider max coloring on hereditary graph classes. The problem is defined as follows. Given a graph G = (V, E) and positive
On the Max Coloring Problem Leah Epstein Asaf Levin May 22, 2010 Abstract We consider max coloring on hereditary graph classes. The problem is defined as follows. Given a graph G = (V, E) and positive
Improving Image Segmentation Quality Via Graph Theory
 International Symposium on Computers & Informatics (ISCI 05) Improving Image Segmentation Quality Via Graph Theory Xiangxiang Li, Songhao Zhu School of Automatic, Nanjing University of Post and Telecommunications,
International Symposium on Computers & Informatics (ISCI 05) Improving Image Segmentation Quality Via Graph Theory Xiangxiang Li, Songhao Zhu School of Automatic, Nanjing University of Post and Telecommunications,
High Dimensional Indexing by Clustering
 Yufei Tao ITEE University of Queensland Recall that, our discussion so far has assumed that the dimensionality d is moderately high, such that it can be regarded as a constant. This means that d should
Yufei Tao ITEE University of Queensland Recall that, our discussion so far has assumed that the dimensionality d is moderately high, such that it can be regarded as a constant. This means that d should
On the packing chromatic number of some lattices
 On the packing chromatic number of some lattices Arthur S. Finbow Department of Mathematics and Computing Science Saint Mary s University Halifax, Canada BH C art.finbow@stmarys.ca Douglas F. Rall Department
On the packing chromatic number of some lattices Arthur S. Finbow Department of Mathematics and Computing Science Saint Mary s University Halifax, Canada BH C art.finbow@stmarys.ca Douglas F. Rall Department
6. Lecture notes on matroid intersection
 Massachusetts Institute of Technology 18.453: Combinatorial Optimization Michel X. Goemans May 2, 2017 6. Lecture notes on matroid intersection One nice feature about matroids is that a simple greedy algorithm
Massachusetts Institute of Technology 18.453: Combinatorial Optimization Michel X. Goemans May 2, 2017 6. Lecture notes on matroid intersection One nice feature about matroids is that a simple greedy algorithm
Finding Exemplars from Pairwise Dissimilarities via Simultaneous Sparse Recovery
 Finding Exemplars from Pairwise Dissimilarities via Simultaneous Sparse Recovery Ehsan Elhamifar EECS Department University of California, Berkeley Guillermo Sapiro ECE, CS Department Duke University René
Finding Exemplars from Pairwise Dissimilarities via Simultaneous Sparse Recovery Ehsan Elhamifar EECS Department University of California, Berkeley Guillermo Sapiro ECE, CS Department Duke University René
Spectral Clustering X I AO ZE N G + E L HA M TA BA S SI CS E CL A S S P R ESENTATION MA RCH 1 6,
 Spectral Clustering XIAO ZENG + ELHAM TABASSI CSE 902 CLASS PRESENTATION MARCH 16, 2017 1 Presentation based on 1. Von Luxburg, Ulrike. "A tutorial on spectral clustering." Statistics and computing 17.4
Spectral Clustering XIAO ZENG + ELHAM TABASSI CSE 902 CLASS PRESENTATION MARCH 16, 2017 1 Presentation based on 1. Von Luxburg, Ulrike. "A tutorial on spectral clustering." Statistics and computing 17.4
A Fuzzy C-means Clustering Algorithm Based on Pseudo-nearest-neighbor Intervals for Incomplete Data
 Journal of Computational Information Systems 11: 6 (2015) 2139 2146 Available at http://www.jofcis.com A Fuzzy C-means Clustering Algorithm Based on Pseudo-nearest-neighbor Intervals for Incomplete Data
Journal of Computational Information Systems 11: 6 (2015) 2139 2146 Available at http://www.jofcis.com A Fuzzy C-means Clustering Algorithm Based on Pseudo-nearest-neighbor Intervals for Incomplete Data
SPARSE COMPONENT ANALYSIS FOR BLIND SOURCE SEPARATION WITH LESS SENSORS THAN SOURCES. Yuanqing Li, Andrzej Cichocki and Shun-ichi Amari
 SPARSE COMPONENT ANALYSIS FOR BLIND SOURCE SEPARATION WITH LESS SENSORS THAN SOURCES Yuanqing Li, Andrzej Cichocki and Shun-ichi Amari Laboratory for Advanced Brain Signal Processing Laboratory for Mathematical
SPARSE COMPONENT ANALYSIS FOR BLIND SOURCE SEPARATION WITH LESS SENSORS THAN SOURCES Yuanqing Li, Andrzej Cichocki and Shun-ichi Amari Laboratory for Advanced Brain Signal Processing Laboratory for Mathematical
Popular matchings with two-sided preferences and one-sided ties
 Popular matchings with two-sided preferences and one-sided ties Ágnes Cseh 1, Chien-Chung Huang 2, and Telikepalli Kavitha 3 1 TU Berlin, Germany. cseh@math.tu-berlin.de 2 Chalmers University, Sweden.
Popular matchings with two-sided preferences and one-sided ties Ágnes Cseh 1, Chien-Chung Huang 2, and Telikepalli Kavitha 3 1 TU Berlin, Germany. cseh@math.tu-berlin.de 2 Chalmers University, Sweden.
Application of Spectral Clustering Algorithm
 1/27 Application of Spectral Clustering Algorithm Danielle Middlebrooks dmiddle1@math.umd.edu Advisor: Kasso Okoudjou kasso@umd.edu Department of Mathematics University of Maryland- College Park Advance
1/27 Application of Spectral Clustering Algorithm Danielle Middlebrooks dmiddle1@math.umd.edu Advisor: Kasso Okoudjou kasso@umd.edu Department of Mathematics University of Maryland- College Park Advance
Cluster Analysis. Mu-Chun Su. Department of Computer Science and Information Engineering National Central University 2003/3/11 1
 Cluster Analysis Mu-Chun Su Department of Computer Science and Information Engineering National Central University 2003/3/11 1 Introduction Cluster analysis is the formal study of algorithms and methods
Cluster Analysis Mu-Chun Su Department of Computer Science and Information Engineering National Central University 2003/3/11 1 Introduction Cluster analysis is the formal study of algorithms and methods
NP-Hardness. We start by defining types of problem, and then move on to defining the polynomial-time reductions.
 CS 787: Advanced Algorithms NP-Hardness Instructor: Dieter van Melkebeek We review the concept of polynomial-time reductions, define various classes of problems including NP-complete, and show that 3-SAT
CS 787: Advanced Algorithms NP-Hardness Instructor: Dieter van Melkebeek We review the concept of polynomial-time reductions, define various classes of problems including NP-complete, and show that 3-SAT
Music Recommendation with Implicit Feedback and Side Information
 Music Recommendation with Implicit Feedback and Side Information Shengbo Guo Yahoo! Labs shengbo@yahoo-inc.com Behrouz Behmardi Criteo b.behmardi@criteo.com Gary Chen Vobile gary.chen@vobileinc.com Abstract
Music Recommendation with Implicit Feedback and Side Information Shengbo Guo Yahoo! Labs shengbo@yahoo-inc.com Behrouz Behmardi Criteo b.behmardi@criteo.com Gary Chen Vobile gary.chen@vobileinc.com Abstract
Domination Cover Pebbling: Structural Results
 Domination Cover Pebbling: Structural Results arxiv:math.co/0509564 v 3 Sep 005 Nathaniel G. Watson Department of Mathematics Washington University at St. Louis Carl R. Yerger Department of Mathematics
Domination Cover Pebbling: Structural Results arxiv:math.co/0509564 v 3 Sep 005 Nathaniel G. Watson Department of Mathematics Washington University at St. Louis Carl R. Yerger Department of Mathematics
Robust Principal Component Analysis (RPCA)
 Robust Principal Component Analysis (RPCA) & Matrix decomposition: into low-rank and sparse components Zhenfang Hu 2010.4.1 reference [1] Chandrasekharan, V., Sanghavi, S., Parillo, P., Wilsky, A.: Ranksparsity
Robust Principal Component Analysis (RPCA) & Matrix decomposition: into low-rank and sparse components Zhenfang Hu 2010.4.1 reference [1] Chandrasekharan, V., Sanghavi, S., Parillo, P., Wilsky, A.: Ranksparsity
Semidefinite Programs for Exact Recovery of a Hidden Community (and Many Communities)
 Semidefinite Programs for Exact Recovery of a Hidden Community (and Many Communities) Bruce Hajek 1 Yihong Wu 1 Jiaming Xu 2 1 University of Illinois at Urbana-Champaign 2 Simons Insitute, UC Berkeley
Semidefinite Programs for Exact Recovery of a Hidden Community (and Many Communities) Bruce Hajek 1 Yihong Wu 1 Jiaming Xu 2 1 University of Illinois at Urbana-Champaign 2 Simons Insitute, UC Berkeley
Spanning trees and orientations of graphs
 Journal of Combinatorics Volume 1, Number 2, 101 111, 2010 Spanning trees and orientations of graphs Carsten Thomassen A conjecture of Merino and Welsh says that the number of spanning trees τ(g) of a
Journal of Combinatorics Volume 1, Number 2, 101 111, 2010 Spanning trees and orientations of graphs Carsten Thomassen A conjecture of Merino and Welsh says that the number of spanning trees τ(g) of a
Using graph theoretic measures to predict the performance of associative memory models
 Using graph theoretic measures to predict the performance of associative memory models Lee Calcraft, Rod Adams, Weiliang Chen and Neil Davey School of Computer Science, University of Hertfordshire College
Using graph theoretic measures to predict the performance of associative memory models Lee Calcraft, Rod Adams, Weiliang Chen and Neil Davey School of Computer Science, University of Hertfordshire College
Rank Measures for Ordering
 Rank Measures for Ordering Jin Huang and Charles X. Ling Department of Computer Science The University of Western Ontario London, Ontario, Canada N6A 5B7 email: fjhuang33, clingg@csd.uwo.ca Abstract. Many
Rank Measures for Ordering Jin Huang and Charles X. Ling Department of Computer Science The University of Western Ontario London, Ontario, Canada N6A 5B7 email: fjhuang33, clingg@csd.uwo.ca Abstract. Many
2. Department of Electronic Engineering and Computer Science, Case Western Reserve University
 Chapter MINING HIGH-DIMENSIONAL DATA Wei Wang 1 and Jiong Yang 2 1. Department of Computer Science, University of North Carolina at Chapel Hill 2. Department of Electronic Engineering and Computer Science,
Chapter MINING HIGH-DIMENSIONAL DATA Wei Wang 1 and Jiong Yang 2 1. Department of Computer Science, University of North Carolina at Chapel Hill 2. Department of Electronic Engineering and Computer Science,
Comparative Study of Subspace Clustering Algorithms
 Comparative Study of Subspace Clustering Algorithms S.Chitra Nayagam, Asst Prof., Dept of Computer Applications, Don Bosco College, Panjim, Goa. Abstract-A cluster is a collection of data objects that
Comparative Study of Subspace Clustering Algorithms S.Chitra Nayagam, Asst Prof., Dept of Computer Applications, Don Bosco College, Panjim, Goa. Abstract-A cluster is a collection of data objects that
On Dimensionality Reduction of Massive Graphs for Indexing and Retrieval
 On Dimensionality Reduction of Massive Graphs for Indexing and Retrieval Charu C. Aggarwal 1, Haixun Wang # IBM T. J. Watson Research Center Hawthorne, NY 153, USA 1 charu@us.ibm.com # Microsoft Research
On Dimensionality Reduction of Massive Graphs for Indexing and Retrieval Charu C. Aggarwal 1, Haixun Wang # IBM T. J. Watson Research Center Hawthorne, NY 153, USA 1 charu@us.ibm.com # Microsoft Research
Visual Representations for Machine Learning
 Visual Representations for Machine Learning Spectral Clustering and Channel Representations Lecture 1 Spectral Clustering: introduction and confusion Michael Felsberg Klas Nordberg The Spectral Clustering
Visual Representations for Machine Learning Spectral Clustering and Channel Representations Lecture 1 Spectral Clustering: introduction and confusion Michael Felsberg Klas Nordberg The Spectral Clustering
Discharging and reducible configurations
 Discharging and reducible configurations Zdeněk Dvořák March 24, 2018 Suppose we want to show that graphs from some hereditary class G are k- colorable. Clearly, we can restrict our attention to graphs
Discharging and reducible configurations Zdeněk Dvořák March 24, 2018 Suppose we want to show that graphs from some hereditary class G are k- colorable. Clearly, we can restrict our attention to graphs
On the Min-Max 2-Cluster Editing Problem
 JOURNAL OF INFORMATION SCIENCE AND ENGINEERING 29, 1109-1120 (2013) On the Min-Max 2-Cluster Editing Problem LI-HSUAN CHEN 1, MAW-SHANG CHANG 2, CHUN-CHIEH WANG 1 AND BANG YE WU 1,* 1 Department of Computer
JOURNAL OF INFORMATION SCIENCE AND ENGINEERING 29, 1109-1120 (2013) On the Min-Max 2-Cluster Editing Problem LI-HSUAN CHEN 1, MAW-SHANG CHANG 2, CHUN-CHIEH WANG 1 AND BANG YE WU 1,* 1 Department of Computer
Graph Theory S 1 I 2 I 1 S 2 I 1 I 2
 Graph Theory S I I S S I I S Graphs Definition A graph G is a pair consisting of a vertex set V (G), and an edge set E(G) ( ) V (G). x and y are the endpoints of edge e = {x, y}. They are called adjacent
Graph Theory S I I S S I I S Graphs Definition A graph G is a pair consisting of a vertex set V (G), and an edge set E(G) ( ) V (G). x and y are the endpoints of edge e = {x, y}. They are called adjacent
Trace Ratio Criterion for Feature Selection
 Proceedings of the Twenty-Third AAAI Conference on Artificial Intelligence (2008) Trace Ratio Criterion for Feature Selection Feiping Nie 1, Shiming Xiang 1, Yangqing Jia 1, Changshui Zhang 1 and Shuicheng
Proceedings of the Twenty-Third AAAI Conference on Artificial Intelligence (2008) Trace Ratio Criterion for Feature Selection Feiping Nie 1, Shiming Xiang 1, Yangqing Jia 1, Changshui Zhang 1 and Shuicheng
A Model of Machine Learning Based on User Preference of Attributes
 1 A Model of Machine Learning Based on User Preference of Attributes Yiyu Yao 1, Yan Zhao 1, Jue Wang 2 and Suqing Han 2 1 Department of Computer Science, University of Regina, Regina, Saskatchewan, Canada
1 A Model of Machine Learning Based on User Preference of Attributes Yiyu Yao 1, Yan Zhao 1, Jue Wang 2 and Suqing Han 2 1 Department of Computer Science, University of Regina, Regina, Saskatchewan, Canada
Bilevel Sparse Coding
 Adobe Research 345 Park Ave, San Jose, CA Mar 15, 2013 Outline 1 2 The learning model The learning algorithm 3 4 Sparse Modeling Many types of sensory data, e.g., images and audio, are in high-dimensional
Adobe Research 345 Park Ave, San Jose, CA Mar 15, 2013 Outline 1 2 The learning model The learning algorithm 3 4 Sparse Modeling Many types of sensory data, e.g., images and audio, are in high-dimensional
Recognizing Interval Bigraphs by Forbidden Patterns
 Recognizing Interval Bigraphs by Forbidden Patterns Arash Rafiey Simon Fraser University, Vancouver, Canada, and Indiana State University, IN, USA arashr@sfu.ca, arash.rafiey@indstate.edu Abstract Let
Recognizing Interval Bigraphs by Forbidden Patterns Arash Rafiey Simon Fraser University, Vancouver, Canada, and Indiana State University, IN, USA arashr@sfu.ca, arash.rafiey@indstate.edu Abstract Let
Superconcentrators of depth 2 and 3; odd levels help (rarely)
 Superconcentrators of depth 2 and 3; odd levels help (rarely) Noga Alon Bellcore, Morristown, NJ, 07960, USA and Department of Mathematics Raymond and Beverly Sackler Faculty of Exact Sciences Tel Aviv
Superconcentrators of depth 2 and 3; odd levels help (rarely) Noga Alon Bellcore, Morristown, NJ, 07960, USA and Department of Mathematics Raymond and Beverly Sackler Faculty of Exact Sciences Tel Aviv
Math 302 Introduction to Proofs via Number Theory. Robert Jewett (with small modifications by B. Ćurgus)
 Math 30 Introduction to Proofs via Number Theory Robert Jewett (with small modifications by B. Ćurgus) March 30, 009 Contents 1 The Integers 3 1.1 Axioms of Z...................................... 3 1.
Math 30 Introduction to Proofs via Number Theory Robert Jewett (with small modifications by B. Ćurgus) March 30, 009 Contents 1 The Integers 3 1.1 Axioms of Z...................................... 3 1.
What Does Robustness Say About Algorithms?
 What Does Robustness Say About Algorithms? Ankur Moitra (MIT) ICML 2017 Tutorial, August 6 th Let me tell you a story about the tension between sharp thresholds and robustness THE STOCHASTIC BLOCK MODEL
What Does Robustness Say About Algorithms? Ankur Moitra (MIT) ICML 2017 Tutorial, August 6 th Let me tell you a story about the tension between sharp thresholds and robustness THE STOCHASTIC BLOCK MODEL
