Approaches to Efficient Multiple Sequence Alignment and Protein Search
|
|
|
- Steven Franklin
- 5 years ago
- Views:
Transcription
1 Approaches to Efficient Multiple Sequence Alignment and Protein Search Thesis statements of the PhD dissertation Adrienn Szabó Supervisor: István Miklós Eötvös Loránd University Faculty of Informatics Ph.D. School of Computer Science Erzsébet Csuhaj-Varjú D.Sc. Ph.D. Program: Basics and methodology of Informatics János Demetrovics D.Sc. Budapest, 2016.
2 Introduction The science of biology offers many great opportunities to a data scientist and a software developer to take part in the challenges of making sense of the gargantuous amount of data being gathered constantly. In my dissertation I focus on two topics: the NP-hard problem of aligning biological sequences [?], and searching protein families in a large database. The latter is more close to a traditional data mining task performed over biological data, while sequence alignment is a very challenging and active research subfield within bioinformatics in itself. After brief introductions to bioinformatics and data mining of the first chapter, Chapter 2 explores previously existing solutions and techniques for aligning proteins and searching protein databases, by reviewing the related scientific literature. Chapter 3 and Chapter 4 introduce new results in the topic of multiple sequence alignment: of the two, the first is based on [RA 1], and it describes how we implemented a new variant of cornercutting methods; while the second introduces a new representation for storing and carrying out statistical computations over a set of multiple sequence alignment paths. The related publications are [ER 1] and [ER 2]. Chapter 5 shows how we tackled the problem of finding a needle in a haystack within a biological context: the task was to find protein families exhibiting some pre-defined complex features in a large protein database. The main goal of the research-and-development projects behind my thesis was to discover new methodologies and techniques that could aid biologist researchers in their daily work in regards of acquiring new insights by automating the process of extracting useful information of the large amounts of data. The developed solutions were tested on real datasets and validated in practice. All the related software codes are accessible as open source projects, and I put great emphasis on re-usability in designing the related codebases. New Results 1. An efficient corner-cutting algorithm for multiple sequence alignment: Reticular Alignment Usually, corner-cutting methods define a compact, mostly convex part of the dynamic programming table for narrowing down the search for the best scored multiple alignment. We introduced a new progressive alignment method called Reticular Alignment, which obtains a set of optimal and suboptimal alignments at each step of the progressive alignment procedure. This set of alignments is 1
3 represented by a DAG network (a directed acyclic graph) of alignment columns. The novelty of our algorithm, introduced in [RA 1] is that we do not restrict the search for solutions to a compact part of the dynamic programming table, but instead, we use a special data structure (see Fig. 1) for both representing the alignments and aligning the set of alignments against another set of alignments. The common parts of the alignments are represented only once, and aligned only once, thus saving a large amount of memory and running time. Figure 1: A small example of a reticular alignment network: it shows three different alignments of the sequences ALLGVGQ and AVGQ This novel corner-cutting approach allows the efficient search of the space of multiple sequence alignments for better alignments. The method has a parameter, t, which affects how much of the alignment space is explored. The Reticular Alignment method can be combined with any scoring scheme, and in this way, we were able to infer the relative importance of sophisticated scoring schemes versus more exhaustive searches in the alignment space. The conclusion is that it is important to increase the search space for finding high-scored alignments. The accuracy of the final alignments was tested on the BAliBASE database [?]. We compared our method with the most popular alignment tools: ClustalW, MAFFT, and FSA. For a comparison of alignment accuracy, see Figure 2, and for more details, refer to Figure 3.2 in Chapter 3 of the dissertation. We found that combining sophisticated scoring schemes with the Reticular Alignment progressive alignment approach yielded a method whose accuracy is comparable to that of cutting-edge alignment methods, Clustal, MAFFT and FSA. It should be noted that this procedure of explicitly tracing the possible alignment paths as a means of a very strict and selective corner cutting method can not be easily enhanced, unless some dependencies between the alignment columns would be taken into account, but that would require much more data and intelligent algorithms to be trained on it. 2
4 Figure 2: Comparison of alignment software on BAliBASE v2.01 Refs Increasing the reticular threshold parameter in the Reticular Alignment method does not guarantee a better alignment The Reticular Alignment method usually could find more accurate alignments when itstparameter was increased to explore more of the space of possible alignments; although we did find some interesting cases when the final MSA score decreased with a higher value oft, see Figure 3. There can be be two fundamentally different reasons for why a widened search may result in worse final alignments: either the better scored alignments are less accurate (the wider search found better scored alignments, but these alignments happen to be differing more from the BAliBASE benchmark) or because of the local decisions at the guide tree nodes it may happen that more noise (meaning wrong sub-alignments) are propagated upwards, and a locally seemingly better alignment can be worse at the final alignment at the root. For a detailed explanation, see Fig. 4. 3
5 Figure 3: Dependency of alignment accuracy (all-column SP) on the reticular threshold Alignment accuracy achieved by RetAlign for different reticular threshold values. Accuracy here is measured as the mean all-column SP score on BAliBASE v2.01 Reference sets 1 5. RetAlign was run with sequence weighting on and pairwise indel scoring. 3. A new representation for storing and carrying out statistical computations over a set of multiple sequence alignment paths Representing a set of alignments as a DAG (directed acyclic graph) over alignment columns as nodes provides a way to quickly compute a distribution of alignment paths in the space of possible multiple sequence alignments, and other quantities of interest averaged over alignments [ER 1]. Joining a large collection of alignment paths in our experiments, up to a couple of thousands of them into a DAG network allows for downstream inference to be averaged over a substantive sample of alignments. Due to interchanges and crossovers at the common alignment columns, the number of alignments encoded in the network is usually a couple of orders of magnitude larger than the number of original alignment paths used to generate the DAG, so the effective sample size is greatly increased. It is also possible to weight each alignment according to a more reliable estimate of the posterior probability, rather than analysing only a small set of individual samples. Note, that many standard algorithms that were designed to work on single alignments for 4
6 Figure 4: Explaining how the internal score might decrease with the reticular threshold. In this particular example s a is the score of the best alignment at internal node v a, with t 1 as the threshold value. At an upper level, namely, at node v b, an x b,1 -network is generated (score: s b,1 ) based on the x a,1 -network. Then the best alignment at the root has score s r,1. After increasing the value of t to t 2, the best alignment at node v a is still the same, but the network of suboptimal alignments to consider at higher levels is the widerx a,2 > x a,1 network. From this network, it is possible to find a better scored alignment at node v b, with score s b,2 > s b,1. If s b,2 x b,2 > s b,1 then the x b,2 -network might be so different from x b,1, that they do not contain any common alignments. Therefore, the best alignment at the root obtained from thex b,2 -network might have a scores r,2 < s r,1. example, forward-backward algorithms for HMMs can be fairly easily adapted so that they can handle alignment DAGs as well. In [ER 1] and [ER 2], we presented a general framework for dealing with alignment uncertainty: on the one hand we explained the statistical background, while on the other hand, we tested our ideas on real data, by implementing the framework in Java as part of the WeaveAlign package. 4. A new protocol for finding re-occurring motifs in disordered regions of proteins Suspecting a role in the development of Alzheimer s disease, we were looking for examples of non-position specific conservation of an amino acid pattern (HD) within a certain distance of 5
7 transmembrane (TM) domains of TM proteins. I worked with the UniProt/SwissProt database, containing over half a million of protein sequences and their annotations. The proposed protocol consist of the following, individually parameterizable steps, or levels: 1. Filtering transmembrane proteins 2. Cutting TM and extracellular fragments 3. Clustering protein fragments 4. Multiple sequence alignment of protein fragments and tree building for each cluster 5. Finding subtrees of the cluster-trees that include the HD pattern 6. Visualization and analysis of results I wrote two Java packages, the first of which (ProteinSearch) contains classes to handle the SwissProt files and perform the filtering, cutting and clustering of proteins; while the second (PhyTreeSearch) gives an easy-to-use solution to the problem of searching subtrees of a large evolutionary tree that adhere to some pre-defined properties (in terms of containing an amino acid pattern in the sequences, for example). Scripts and configuration files for running the Java programs according to the above pipeline can be found in a separate, third code repository (APP-HD-pattern-runner). By running the pipeline, I managed to find a handful of small sets of proteins that could be interesting to look at from this point on, a biologist needs to decide if they are worth to be further analyzed. This project is still work-in-progress after proving the biological relevance of the findings, we plan to submit a methodology paper to BMC Bioinformatics in Summary, availability of software Ideally, all software tools used and developed during a research process should be open software. Usually at least some new code fragments or scripts have to be written to carry out new research experiments. Our Java software packages runnable jar files and java sources for multiple sequence alignment methods of Chapters 3 and 4 are openly available at the following URLs: In Chapter 5, while implementing a pipeline for filtering protein fragments, I used two complementing methodologies / tools to ensure easy reproducibility of the final results: 6
8 I shared my working environment through a Docker virtual image I published all the software codes (java program sources and scripts, along with example configuration files) on GitHub, as open source projects under Apache License and Unilicense 2 :
9 Author s related publications [RA 1] Adrienn Szabó, Ádám Novák, István Miklós, and Jotun Hein: Reticular alignment: A progressive corner-cutting method for multiple sequence alignment BMC Bioinformatics (IF: 3.028), vol 11, page 570, 2010 URL: DOI: / [ER 1] Joseph L. Herman, Ádám Novák, Rune Lyngsø, Adrienn Szabó, István Miklós, and Jotun Hein: Efficient representation of uncertainty in multiple sequence alignments using directed acyclic graphs BMC Bioinformatics (IF: 2.67), vol 16, number 1, 2015 URL: DOI: /s [ER 2] Joseph L. Herman, Adrienn Szabó, István Miklós, and Jotun Hein: Approximate statistical alignment by iterative sampling of substitution matrices arxiv, 2015 URL: 8
PROTEIN MULTIPLE ALIGNMENT MOTIVATION: BACKGROUND: Marina Sirota
 Marina Sirota MOTIVATION: PROTEIN MULTIPLE ALIGNMENT To study evolution on the genetic level across a wide range of organisms, biologists need accurate tools for multiple sequence alignment of protein
Marina Sirota MOTIVATION: PROTEIN MULTIPLE ALIGNMENT To study evolution on the genetic level across a wide range of organisms, biologists need accurate tools for multiple sequence alignment of protein
Lecture 5: Multiple sequence alignment
 Lecture 5: Multiple sequence alignment Introduction to Computational Biology Teresa Przytycka, PhD (with some additions by Martin Vingron) Why do we need multiple sequence alignment Pairwise sequence alignment
Lecture 5: Multiple sequence alignment Introduction to Computational Biology Teresa Przytycka, PhD (with some additions by Martin Vingron) Why do we need multiple sequence alignment Pairwise sequence alignment
Principles of Bioinformatics. BIO540/STA569/CSI660 Fall 2010
 Principles of Bioinformatics BIO540/STA569/CSI660 Fall 2010 Lecture 11 Multiple Sequence Alignment I Administrivia Administrivia The midterm examination will be Monday, October 18 th, in class. Closed
Principles of Bioinformatics BIO540/STA569/CSI660 Fall 2010 Lecture 11 Multiple Sequence Alignment I Administrivia Administrivia The midterm examination will be Monday, October 18 th, in class. Closed
Multiple sequence alignment. November 20, 2018
 Multiple sequence alignment November 20, 2018 Why do multiple alignment? Gain insight into evolutionary history Can assess time of divergence by looking at the number of mutations needed to change one
Multiple sequence alignment November 20, 2018 Why do multiple alignment? Gain insight into evolutionary history Can assess time of divergence by looking at the number of mutations needed to change one
GLOBEX Bioinformatics (Summer 2015) Multiple Sequence Alignment
 GLOBEX Bioinformatics (Summer 2015) Multiple Sequence Alignment Scoring Dynamic Programming algorithms Heuristic algorithms CLUSTAL W Courtesy of jalview Motivations Collective (or aggregate) statistic
GLOBEX Bioinformatics (Summer 2015) Multiple Sequence Alignment Scoring Dynamic Programming algorithms Heuristic algorithms CLUSTAL W Courtesy of jalview Motivations Collective (or aggregate) statistic
BLAST, Profile, and PSI-BLAST
 BLAST, Profile, and PSI-BLAST Jianlin Cheng, PhD School of Electrical Engineering and Computer Science University of Central Florida 26 Free for academic use Copyright @ Jianlin Cheng & original sources
BLAST, Profile, and PSI-BLAST Jianlin Cheng, PhD School of Electrical Engineering and Computer Science University of Central Florida 26 Free for academic use Copyright @ Jianlin Cheng & original sources
CISC 889 Bioinformatics (Spring 2003) Multiple Sequence Alignment
 CISC 889 Bioinformatics (Spring 2003) Multiple Sequence Alignment Courtesy of jalview 1 Motivations Collective statistic Protein families Identification and representation of conserved sequence features
CISC 889 Bioinformatics (Spring 2003) Multiple Sequence Alignment Courtesy of jalview 1 Motivations Collective statistic Protein families Identification and representation of conserved sequence features
Comparison and Evaluation of Multiple Sequence Alignment Tools In Bininformatics
 IJCSNS International Journal of Computer Science and Network Security, VOL.9 No.7, July 2009 51 Comparison and Evaluation of Multiple Sequence Alignment Tools In Bininformatics Asieh Sedaghatinia, Dr Rodziah
IJCSNS International Journal of Computer Science and Network Security, VOL.9 No.7, July 2009 51 Comparison and Evaluation of Multiple Sequence Alignment Tools In Bininformatics Asieh Sedaghatinia, Dr Rodziah
Lab 4: Multiple Sequence Alignment (MSA)
 Lab 4: Multiple Sequence Alignment (MSA) The objective of this lab is to become familiar with the features of several multiple alignment and visualization tools, including the data input and output, basic
Lab 4: Multiple Sequence Alignment (MSA) The objective of this lab is to become familiar with the features of several multiple alignment and visualization tools, including the data input and output, basic
Designing parallel algorithms for constructing large phylogenetic trees on Blue Waters
 Designing parallel algorithms for constructing large phylogenetic trees on Blue Waters Erin Molloy University of Illinois at Urbana Champaign General Allocation (PI: Tandy Warnow) Exploratory Allocation
Designing parallel algorithms for constructing large phylogenetic trees on Blue Waters Erin Molloy University of Illinois at Urbana Champaign General Allocation (PI: Tandy Warnow) Exploratory Allocation
Profiles and Multiple Alignments. COMP 571 Luay Nakhleh, Rice University
 Profiles and Multiple Alignments COMP 571 Luay Nakhleh, Rice University Outline Profiles and sequence logos Profile hidden Markov models Aligning profiles Multiple sequence alignment by gradual sequence
Profiles and Multiple Alignments COMP 571 Luay Nakhleh, Rice University Outline Profiles and sequence logos Profile hidden Markov models Aligning profiles Multiple sequence alignment by gradual sequence
Multiple Sequence Alignment (MSA)
 I519 Introduction to Bioinformatics, Fall 2013 Multiple Sequence Alignment (MSA) Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Outline Multiple sequence alignment (MSA) Generalize
I519 Introduction to Bioinformatics, Fall 2013 Multiple Sequence Alignment (MSA) Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Outline Multiple sequence alignment (MSA) Generalize
Approaches to Efficient Multiple Sequence Alignment and Protein Search. Adrienn Szabó
 Approaches to Efficient Multiple Sequence Alignment and Protein Search Adrienn Szabó Supervisor: István Miklós Eötvös Loránd University, Budapest Faculty of Informatics PhD School of Computer Science Erzsébet
Approaches to Efficient Multiple Sequence Alignment and Protein Search Adrienn Szabó Supervisor: István Miklós Eötvös Loránd University, Budapest Faculty of Informatics PhD School of Computer Science Erzsébet
Multiple Sequence Alignment Based on Profile Alignment of Intermediate Sequences
 Multiple Sequence Alignment Based on Profile Alignment of Intermediate Sequences Yue Lu and Sing-Hoi Sze RECOMB 2007 Presented by: Wanxing Xu March 6, 2008 Content Biology Motivation Computation Problem
Multiple Sequence Alignment Based on Profile Alignment of Intermediate Sequences Yue Lu and Sing-Hoi Sze RECOMB 2007 Presented by: Wanxing Xu March 6, 2008 Content Biology Motivation Computation Problem
Basic Local Alignment Search Tool (BLAST)
 BLAST 26.04.2018 Basic Local Alignment Search Tool (BLAST) BLAST (Altshul-1990) is an heuristic Pairwise Alignment composed by six-steps that search for local similarities. The most used access point to
BLAST 26.04.2018 Basic Local Alignment Search Tool (BLAST) BLAST (Altshul-1990) is an heuristic Pairwise Alignment composed by six-steps that search for local similarities. The most used access point to
Data Mining Technologies for Bioinformatics Sequences
 Data Mining Technologies for Bioinformatics Sequences Deepak Garg Computer Science and Engineering Department Thapar Institute of Engineering & Tecnology, Patiala Abstract Main tool used for sequence alignment
Data Mining Technologies for Bioinformatics Sequences Deepak Garg Computer Science and Engineering Department Thapar Institute of Engineering & Tecnology, Patiala Abstract Main tool used for sequence alignment
Giri Narasimhan. CAP 5510: Introduction to Bioinformatics. ECS 254; Phone: x3748
 CAP 5510: Introduction to Bioinformatics Giri Narasimhan ECS 254; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs07.html 1/30/07 CAP5510 1 BLAST & FASTA FASTA [Lipman, Pearson 85, 88]
CAP 5510: Introduction to Bioinformatics Giri Narasimhan ECS 254; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs07.html 1/30/07 CAP5510 1 BLAST & FASTA FASTA [Lipman, Pearson 85, 88]
Bioinformatics explained: Smith-Waterman
 Bioinformatics Explained Bioinformatics explained: Smith-Waterman May 1, 2007 CLC bio Gustav Wieds Vej 10 8000 Aarhus C Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com info@clcbio.com
Bioinformatics Explained Bioinformatics explained: Smith-Waterman May 1, 2007 CLC bio Gustav Wieds Vej 10 8000 Aarhus C Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com info@clcbio.com
Multiple alignment. Multiple alignment. Computational complexity. Multiple sequence alignment. Consensus. 2 1 sec sec sec
 Introduction to Bioinformatics Iosif Vaisman Email: ivaisman@gmu.edu VTISCTGSSSNIGAG-NHVKWYQQLPG VTISCTGTSSNIGS--ITVNWYQQLPG LRLSCSSSGFIFSS--YAMYWVRQAPG LSLTCTVSGTSFDD--YYSTWVRQPPG PEVTCVVVDVSHEDPQVKFNWYVDG--
Introduction to Bioinformatics Iosif Vaisman Email: ivaisman@gmu.edu VTISCTGSSSNIGAG-NHVKWYQQLPG VTISCTGTSSNIGS--ITVNWYQQLPG LRLSCSSSGFIFSS--YAMYWVRQAPG LSLTCTVSGTSFDD--YYSTWVRQPPG PEVTCVVVDVSHEDPQVKFNWYVDG--
A New Approach For Tree Alignment Based on Local Re-Optimization
 A New Approach For Tree Alignment Based on Local Re-Optimization Feng Yue and Jijun Tang Department of Computer Science and Engineering University of South Carolina Columbia, SC 29063, USA yuef, jtang
A New Approach For Tree Alignment Based on Local Re-Optimization Feng Yue and Jijun Tang Department of Computer Science and Engineering University of South Carolina Columbia, SC 29063, USA yuef, jtang
Chapter 6. Multiple sequence alignment (week 10)
 Course organization Introduction ( Week 1,2) Part I: Algorithms for Sequence Analysis (Week 1-11) Chapter 1-3, Models and theories» Probability theory and Statistics (Week 3)» Algorithm complexity analysis
Course organization Introduction ( Week 1,2) Part I: Algorithms for Sequence Analysis (Week 1-11) Chapter 1-3, Models and theories» Probability theory and Statistics (Week 3)» Algorithm complexity analysis
Special course in Computer Science: Advanced Text Algorithms
 Special course in Computer Science: Advanced Text Algorithms Lecture 8: Multiple alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
Special course in Computer Science: Advanced Text Algorithms Lecture 8: Multiple alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
3.4 Multiple sequence alignment
 3.4 Multiple sequence alignment Why produce a multiple sequence alignment? Using more than two sequences results in a more convincing alignment by revealing conserved regions in ALL of the sequences Aligned
3.4 Multiple sequence alignment Why produce a multiple sequence alignment? Using more than two sequences results in a more convincing alignment by revealing conserved regions in ALL of the sequences Aligned
Comparison of Phylogenetic Trees of Multiple Protein Sequence Alignment Methods
 Comparison of Phylogenetic Trees of Multiple Protein Sequence Alignment Methods Khaddouja Boujenfa, Nadia Essoussi, and Mohamed Limam International Science Index, Computer and Information Engineering waset.org/publication/482
Comparison of Phylogenetic Trees of Multiple Protein Sequence Alignment Methods Khaddouja Boujenfa, Nadia Essoussi, and Mohamed Limam International Science Index, Computer and Information Engineering waset.org/publication/482
Bioinformatics for Biologists
 Bioinformatics for Biologists Sequence Analysis: Part I. Pairwise alignment and database searching Fran Lewitter, Ph.D. Director Bioinformatics & Research Computing Whitehead Institute Topics to Cover
Bioinformatics for Biologists Sequence Analysis: Part I. Pairwise alignment and database searching Fran Lewitter, Ph.D. Director Bioinformatics & Research Computing Whitehead Institute Topics to Cover
HIDDEN MARKOV MODELS AND SEQUENCE ALIGNMENT
 HIDDEN MARKOV MODELS AND SEQUENCE ALIGNMENT - Swarbhanu Chatterjee. Hidden Markov models are a sophisticated and flexible statistical tool for the study of protein models. Using HMMs to analyze proteins
HIDDEN MARKOV MODELS AND SEQUENCE ALIGNMENT - Swarbhanu Chatterjee. Hidden Markov models are a sophisticated and flexible statistical tool for the study of protein models. Using HMMs to analyze proteins
Multiple sequence alignment. November 2, 2017
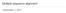 Multiple sequence alignment November 2, 2017 Why do multiple alignment? Gain insight into evolutionary history Can assess time of divergence by looking at the number of mutations needed to change one sequence
Multiple sequence alignment November 2, 2017 Why do multiple alignment? Gain insight into evolutionary history Can assess time of divergence by looking at the number of mutations needed to change one sequence
Multiple Sequence Alignment. Mark Whitsitt - NCSA
 Multiple Sequence Alignment Mark Whitsitt - NCSA What is a Multiple Sequence Alignment (MA)? GMHGTVYANYAVDSSDLLLAFGVRFDDRVTGKLEAFASRAKIVHIDIDSAEIGKNKQPHV GMHGTVYANYAVEHSDLLLAFGVRFDDRVTGKLEAFASRAKIVHIDIDSAEIGKNKTPHV
Multiple Sequence Alignment Mark Whitsitt - NCSA What is a Multiple Sequence Alignment (MA)? GMHGTVYANYAVDSSDLLLAFGVRFDDRVTGKLEAFASRAKIVHIDIDSAEIGKNKQPHV GMHGTVYANYAVEHSDLLLAFGVRFDDRVTGKLEAFASRAKIVHIDIDSAEIGKNKTPHV
PoMSA: An Efficient and Precise Position-based Multiple Sequence Alignment Technique
 PoMSA: An Efficient and Precise Position-based Multiple Sequence Alignment Technique Sara Shehab 1,a, Sameh Shohdy 1,b, Arabi E. Keshk 1,c 1 Department of Computer Science, Faculty of Computers and Information,
PoMSA: An Efficient and Precise Position-based Multiple Sequence Alignment Technique Sara Shehab 1,a, Sameh Shohdy 1,b, Arabi E. Keshk 1,c 1 Department of Computer Science, Faculty of Computers and Information,
Thesis book. Extension of strongly typed object-oriented systems using metaprograms István Zólyomi
 Thesis book Extension of strongly typed object-oriented systems using metaprograms István Zólyomi Supervisor: Dr. Zoltán Porkoláb Eötvös Loránd University Faculty of Informatics Department of Programming
Thesis book Extension of strongly typed object-oriented systems using metaprograms István Zólyomi Supervisor: Dr. Zoltán Porkoláb Eötvös Loránd University Faculty of Informatics Department of Programming
Clustering of Proteins
 Melroy Saldanha saldanha@stanford.edu CS 273 Project Report Clustering of Proteins Introduction Numerous genome-sequencing projects have led to a huge growth in the size of protein databases. Manual annotation
Melroy Saldanha saldanha@stanford.edu CS 273 Project Report Clustering of Proteins Introduction Numerous genome-sequencing projects have led to a huge growth in the size of protein databases. Manual annotation
MULTIPLE SEQUENCE ALIGNMENT SOLUTIONS AND APPLICATIONS
 MULTIPLE SEQUENCE ALIGNMENT SOLUTIONS AND APPLICATIONS By XU ZHANG A DISSERTATION PRESENTED TO THE GRADUATE SCHOOL OF THE UNIVERSITY OF FLORIDA IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE
MULTIPLE SEQUENCE ALIGNMENT SOLUTIONS AND APPLICATIONS By XU ZHANG A DISSERTATION PRESENTED TO THE GRADUATE SCHOOL OF THE UNIVERSITY OF FLORIDA IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE
Today s Lecture. Multiple sequence alignment. Improved scoring of pairwise alignments. Affine gap penalties Profiles
 Today s Lecture Multiple sequence alignment Improved scoring of pairwise alignments Affine gap penalties Profiles 1 The Edit Graph for a Pair of Sequences G A C G T T G A A T G A C C C A C A T G A C G
Today s Lecture Multiple sequence alignment Improved scoring of pairwise alignments Affine gap penalties Profiles 1 The Edit Graph for a Pair of Sequences G A C G T T G A A T G A C C C A C A T G A C G
Long Read RNA-seq Mapper
 UNIVERSITY OF ZAGREB FACULTY OF ELECTRICAL ENGENEERING AND COMPUTING MASTER THESIS no. 1005 Long Read RNA-seq Mapper Josip Marić Zagreb, February 2015. Table of Contents 1. Introduction... 1 2. RNA Sequencing...
UNIVERSITY OF ZAGREB FACULTY OF ELECTRICAL ENGENEERING AND COMPUTING MASTER THESIS no. 1005 Long Read RNA-seq Mapper Josip Marić Zagreb, February 2015. Table of Contents 1. Introduction... 1 2. RNA Sequencing...
C E N T R. Introduction to bioinformatics 2007 E B I O I N F O R M A T I C S V U F O R I N T. Lecture 13 G R A T I V. Iterative homology searching,
 C E N T R E F O R I N T E G R A T I V E B I O I N F O R M A T I C S V U Introduction to bioinformatics 2007 Lecture 13 Iterative homology searching, PSI (Position Specific Iterated) BLAST basic idea use
C E N T R E F O R I N T E G R A T I V E B I O I N F O R M A T I C S V U Introduction to bioinformatics 2007 Lecture 13 Iterative homology searching, PSI (Position Specific Iterated) BLAST basic idea use
Project Report on. De novo Peptide Sequencing. Course: Math 574 Gaurav Kulkarni Washington State University
 Project Report on De novo Peptide Sequencing Course: Math 574 Gaurav Kulkarni Washington State University Introduction Protein is the fundamental building block of one s body. Many biological processes
Project Report on De novo Peptide Sequencing Course: Math 574 Gaurav Kulkarni Washington State University Introduction Protein is the fundamental building block of one s body. Many biological processes
Lecture 5: Markov models
 Master s course Bioinformatics Data Analysis and Tools Lecture 5: Markov models Centre for Integrative Bioinformatics Problem in biology Data and patterns are often not clear cut When we want to make a
Master s course Bioinformatics Data Analysis and Tools Lecture 5: Markov models Centre for Integrative Bioinformatics Problem in biology Data and patterns are often not clear cut When we want to make a
9/29/13. Outline Data mining tasks. Clustering algorithms. Applications of clustering in biology
 9/9/ I9 Introduction to Bioinformatics, Clustering algorithms Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Outline Data mining tasks Predictive tasks vs descriptive tasks Example
9/9/ I9 Introduction to Bioinformatics, Clustering algorithms Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Outline Data mining tasks Predictive tasks vs descriptive tasks Example
Chapter 8 Multiple sequence alignment. Chaochun Wei Spring 2018
 1896 1920 1987 2006 Chapter 8 Multiple sequence alignment Chaochun Wei Spring 2018 Contents 1. Reading materials 2. Multiple sequence alignment basic algorithms and tools how to improve multiple alignment
1896 1920 1987 2006 Chapter 8 Multiple sequence alignment Chaochun Wei Spring 2018 Contents 1. Reading materials 2. Multiple sequence alignment basic algorithms and tools how to improve multiple alignment
INVESTIGATION STUDY: AN INTENSIVE ANALYSIS FOR MSA LEADING METHODS
 INVESTIGATION STUDY: AN INTENSIVE ANALYSIS FOR MSA LEADING METHODS MUHANNAD A. ABU-HASHEM, NUR'AINI ABDUL RASHID, ROSNI ABDULLAH, AWSAN A. HASAN AND ATHEER A. ABDULRAZZAQ School of Computer Sciences, Universiti
INVESTIGATION STUDY: AN INTENSIVE ANALYSIS FOR MSA LEADING METHODS MUHANNAD A. ABU-HASHEM, NUR'AINI ABDUL RASHID, ROSNI ABDULLAH, AWSAN A. HASAN AND ATHEER A. ABDULRAZZAQ School of Computer Sciences, Universiti
1. R. Durbin, S. Eddy, A. Krogh und G. Mitchison: Biological sequence analysis, Cambridge, 1998
 7 Multiple Sequence Alignment The exposition was prepared by Clemens Gröpl, based on earlier versions by Daniel Huson, Knut Reinert, and Gunnar Klau. It is based on the following sources, which are all
7 Multiple Sequence Alignment The exposition was prepared by Clemens Gröpl, based on earlier versions by Daniel Huson, Knut Reinert, and Gunnar Klau. It is based on the following sources, which are all
Multiple Sequence Alignment. With thanks to Eric Stone and Steffen Heber, North Carolina State University
 Multiple Sequence Alignment With thanks to Eric Stone and Steffen Heber, North Carolina State University Definition: Multiple sequence alignment Given a set of sequences, a multiple sequence alignment
Multiple Sequence Alignment With thanks to Eric Stone and Steffen Heber, North Carolina State University Definition: Multiple sequence alignment Given a set of sequences, a multiple sequence alignment
Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014
 Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014 Dynamic programming is a group of mathematical methods used to sequentially split a complicated problem into
Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014 Dynamic programming is a group of mathematical methods used to sequentially split a complicated problem into
Sequence alignment algorithms
 Sequence alignment algorithms Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 23 rd 27 After this lecture, you can decide when to use local and global sequence alignments
Sequence alignment algorithms Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 23 rd 27 After this lecture, you can decide when to use local and global sequence alignments
Efficient Inference in Phylogenetic InDel Trees
 Efficient Inference in Phylogenetic InDel Trees Alexandre Bouchard-Côté Michael I. Jordan Dan Klein Computer Science Division, Department of Statistics University of California at Berkeley Berkeley, CA
Efficient Inference in Phylogenetic InDel Trees Alexandre Bouchard-Côté Michael I. Jordan Dan Klein Computer Science Division, Department of Statistics University of California at Berkeley Berkeley, CA
CS313 Exercise 4 Cover Page Fall 2017
 CS313 Exercise 4 Cover Page Fall 2017 Due by the start of class on Thursday, October 12, 2017. Name(s): In the TIME column, please estimate the time you spent on the parts of this exercise. Please try
CS313 Exercise 4 Cover Page Fall 2017 Due by the start of class on Thursday, October 12, 2017. Name(s): In the TIME column, please estimate the time you spent on the parts of this exercise. Please try
Software review. Shopping in the genome market with EnsMart
 Shopping in the genome market with EnsMart Keywords: genome databases, human genome, comparative genomics, data mining, open source software Abstract Life scientists who work with the supermarket of genome
Shopping in the genome market with EnsMart Keywords: genome databases, human genome, comparative genomics, data mining, open source software Abstract Life scientists who work with the supermarket of genome
Geneious 5.6 Quickstart Manual. Biomatters Ltd
 Geneious 5.6 Quickstart Manual Biomatters Ltd October 15, 2012 2 Introduction This quickstart manual will guide you through the features of Geneious 5.6 s interface and help you orient yourself. You should
Geneious 5.6 Quickstart Manual Biomatters Ltd October 15, 2012 2 Introduction This quickstart manual will guide you through the features of Geneious 5.6 s interface and help you orient yourself. You should
A Modified Vector Space Model for Protein Retrieval
 IJCSNS International Journal of Computer Science and Network Security, VOL.7 No.9, September 27 85 A Modified Vector Space Model for Protein Retrieval Mohammed Said Abual-Rub, Rosni Abdullah and Nur'Aini
IJCSNS International Journal of Computer Science and Network Security, VOL.7 No.9, September 27 85 A Modified Vector Space Model for Protein Retrieval Mohammed Said Abual-Rub, Rosni Abdullah and Nur'Aini
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 06: Multiple Sequence Alignment https://upload.wikimedia.org/wikipedia/commons/thumb/7/79/rplp0_90_clustalw_aln.gif/575px-rplp0_90_clustalw_aln.gif Slides
EECS730: Introduction to Bioinformatics Lecture 06: Multiple Sequence Alignment https://upload.wikimedia.org/wikipedia/commons/thumb/7/79/rplp0_90_clustalw_aln.gif/575px-rplp0_90_clustalw_aln.gif Slides
Bio3D: Interactive Tools for Structural Bioinformatics.
 Bio3D: Interactive Tools for Structural Bioinformatics http://thegrantlab.org/bio3d/ What is Bio3D A freely distributed and widely used R package for structural bioinformatics. Provides a large number
Bio3D: Interactive Tools for Structural Bioinformatics http://thegrantlab.org/bio3d/ What is Bio3D A freely distributed and widely used R package for structural bioinformatics. Provides a large number
Comparison of FP tree and Apriori Algorithm
 International Journal of Engineering Research and Development e-issn: 2278-067X, p-issn: 2278-800X, www.ijerd.com Volume 10, Issue 6 (June 2014), PP.78-82 Comparison of FP tree and Apriori Algorithm Prashasti
International Journal of Engineering Research and Development e-issn: 2278-067X, p-issn: 2278-800X, www.ijerd.com Volume 10, Issue 6 (June 2014), PP.78-82 Comparison of FP tree and Apriori Algorithm Prashasti
Mapping Sequence Conservation onto Structures with Chimera
 This page: www.rbvi.ucsf.edu/chimera/data/tutorials/systems/outline.html Chimera in BP205A BP205A syllabus Mapping Sequence Conservation onto Structures with Chimera Case 1: You already have a structure
This page: www.rbvi.ucsf.edu/chimera/data/tutorials/systems/outline.html Chimera in BP205A BP205A syllabus Mapping Sequence Conservation onto Structures with Chimera Case 1: You already have a structure
CS 581. Tandy Warnow
 CS 581 Tandy Warnow This week Maximum parsimony: solving it on small datasets Maximum Likelihood optimization problem Felsenstein s pruning algorithm Bayesian MCMC methods Research opportunities Maximum
CS 581 Tandy Warnow This week Maximum parsimony: solving it on small datasets Maximum Likelihood optimization problem Felsenstein s pruning algorithm Bayesian MCMC methods Research opportunities Maximum
Multiple Sequence Alignment Using Reconfigurable Computing
 Multiple Sequence Alignment Using Reconfigurable Computing Carlos R. Erig Lima, Heitor S. Lopes, Maiko R. Moroz, and Ramon M. Menezes Bioinformatics Laboratory, Federal University of Technology Paraná
Multiple Sequence Alignment Using Reconfigurable Computing Carlos R. Erig Lima, Heitor S. Lopes, Maiko R. Moroz, and Ramon M. Menezes Bioinformatics Laboratory, Federal University of Technology Paraná
Stephen Scott.
 1 / 33 sscott@cse.unl.edu 2 / 33 Start with a set of sequences In each column, residues are homolgous Residues occupy similar positions in 3D structure Residues diverge from a common ancestral residue
1 / 33 sscott@cse.unl.edu 2 / 33 Start with a set of sequences In each column, residues are homolgous Residues occupy similar positions in 3D structure Residues diverge from a common ancestral residue
srna Detection Results
 srna Detection Results Summary: This tutorial explains how to work with the output obtained from the srna Detection module of Oasis. srna detection is the first analysis module of Oasis, and it examines
srna Detection Results Summary: This tutorial explains how to work with the output obtained from the srna Detection module of Oasis. srna detection is the first analysis module of Oasis, and it examines
Order Preserving Triclustering Algorithm. (Version1.0)
 Order Preserving Triclustering Algorithm User Manual (Version1.0) Alain B. Tchagang alain.tchagang@nrc-cnrc.gc.ca Ziying Liu ziying.liu@nrc-cnrc.gc.ca Sieu Phan sieu.phan@nrc-cnrc.gc.ca Fazel Famili fazel.famili@nrc-cnrc.gc.ca
Order Preserving Triclustering Algorithm User Manual (Version1.0) Alain B. Tchagang alain.tchagang@nrc-cnrc.gc.ca Ziying Liu ziying.liu@nrc-cnrc.gc.ca Sieu Phan sieu.phan@nrc-cnrc.gc.ca Fazel Famili fazel.famili@nrc-cnrc.gc.ca
Dependency detection with Bayesian Networks
 Dependency detection with Bayesian Networks M V Vikhreva Faculty of Computational Mathematics and Cybernetics, Lomonosov Moscow State University, Leninskie Gory, Moscow, 119991 Supervisor: A G Dyakonov
Dependency detection with Bayesian Networks M V Vikhreva Faculty of Computational Mathematics and Cybernetics, Lomonosov Moscow State University, Leninskie Gory, Moscow, 119991 Supervisor: A G Dyakonov
Introduction to Computer Science
 DM534 Introduction to Computer Science Clustering and Feature Spaces Richard Roettger: About Me Computer Science (Technical University of Munich and thesis at the ICSI at the University of California at
DM534 Introduction to Computer Science Clustering and Feature Spaces Richard Roettger: About Me Computer Science (Technical University of Munich and thesis at the ICSI at the University of California at
Text Mining. Representation of Text Documents
 Data Mining is typically concerned with the detection of patterns in numeric data, but very often important (e.g., critical to business) information is stored in the form of text. Unlike numeric data,
Data Mining is typically concerned with the detection of patterns in numeric data, but very often important (e.g., critical to business) information is stored in the form of text. Unlike numeric data,
Biology 644: Bioinformatics
 Find the best alignment between 2 sequences with lengths n and m, respectively Best alignment is very dependent upon the substitution matrix and gap penalties The Global Alignment Problem tries to find
Find the best alignment between 2 sequences with lengths n and m, respectively Best alignment is very dependent upon the substitution matrix and gap penalties The Global Alignment Problem tries to find
Reconstructing long sequences from overlapping sequence fragment. Searching databases for related sequences and subsequences
 SEQUENCE ALIGNMENT ALGORITHMS 1 Why compare sequences? Reconstructing long sequences from overlapping sequence fragment Searching databases for related sequences and subsequences Storing, retrieving and
SEQUENCE ALIGNMENT ALGORITHMS 1 Why compare sequences? Reconstructing long sequences from overlapping sequence fragment Searching databases for related sequences and subsequences Storing, retrieving and
Environmental Sample Classification E.S.C., Josh Katz and Kurt Zimmer
 Environmental Sample Classification E.S.C., Josh Katz and Kurt Zimmer Goal: The task we were given for the bioinformatics capstone class was to construct an interface for the Pipas lab that integrated
Environmental Sample Classification E.S.C., Josh Katz and Kurt Zimmer Goal: The task we were given for the bioinformatics capstone class was to construct an interface for the Pipas lab that integrated
PERFORMANCE OF GRID COMPUTING FOR DISTRIBUTED NEURAL NETWORK. Submitted By:Mohnish Malviya & Suny Shekher Pankaj [CSE,7 TH SEM]
![PERFORMANCE OF GRID COMPUTING FOR DISTRIBUTED NEURAL NETWORK. Submitted By:Mohnish Malviya & Suny Shekher Pankaj [CSE,7 TH SEM] PERFORMANCE OF GRID COMPUTING FOR DISTRIBUTED NEURAL NETWORK. Submitted By:Mohnish Malviya & Suny Shekher Pankaj [CSE,7 TH SEM]](/thumbs/95/123911651.jpg) PERFORMANCE OF GRID COMPUTING FOR DISTRIBUTED NEURAL NETWORK Submitted By:Mohnish Malviya & Suny Shekher Pankaj [CSE,7 TH SEM] All Saints` College Of Technology, Gandhi Nagar, Bhopal. Abstract: In this
PERFORMANCE OF GRID COMPUTING FOR DISTRIBUTED NEURAL NETWORK Submitted By:Mohnish Malviya & Suny Shekher Pankaj [CSE,7 TH SEM] All Saints` College Of Technology, Gandhi Nagar, Bhopal. Abstract: In this
User Manual. Ver. 3.0 March 19, 2012
 User Manual Ver. 3.0 March 19, 2012 Table of Contents 1. Introduction... 2 1.1 Rationale... 2 1.2 Software Work-Flow... 3 1.3 New in GenomeGems 3.0... 4 2. Software Description... 5 2.1 Key Features...
User Manual Ver. 3.0 March 19, 2012 Table of Contents 1. Introduction... 2 1.1 Rationale... 2 1.2 Software Work-Flow... 3 1.3 New in GenomeGems 3.0... 4 2. Software Description... 5 2.1 Key Features...
Taxonomically Clustering Organisms Based on the Profiles of Gene Sequences Using PCA
 Journal of Computer Science 2 (3): 292-296, 2006 ISSN 1549-3636 2006 Science Publications Taxonomically Clustering Organisms Based on the Profiles of Gene Sequences Using PCA 1 E.Ramaraj and 2 M.Punithavalli
Journal of Computer Science 2 (3): 292-296, 2006 ISSN 1549-3636 2006 Science Publications Taxonomically Clustering Organisms Based on the Profiles of Gene Sequences Using PCA 1 E.Ramaraj and 2 M.Punithavalli
ProDA User Manual. Version 1.0. Algorithm developed by Tu Minh Phuong, Chuong B. Do, Robert C. Edgar, and Serafim Batzoglou.
 ProDA User Manual Version 1.0 Algorithm developed by Tu Minh Phuong, Chuong B. Do, Robert C. Edgar, and Serafim Batzoglou. Software written by Tu Minh Phuong and Chuong B. Do. Manual written by Tu Minh
ProDA User Manual Version 1.0 Algorithm developed by Tu Minh Phuong, Chuong B. Do, Robert C. Edgar, and Serafim Batzoglou. Software written by Tu Minh Phuong and Chuong B. Do. Manual written by Tu Minh
Integrating Probabilistic Reasoning with Constraint Satisfaction
 Integrating Probabilistic Reasoning with Constraint Satisfaction IJCAI Tutorial #7 Instructor: Eric I. Hsu July 17, 2011 http://www.cs.toronto.edu/~eihsu/tutorial7 Getting Started Discursive Remarks. Organizational
Integrating Probabilistic Reasoning with Constraint Satisfaction IJCAI Tutorial #7 Instructor: Eric I. Hsu July 17, 2011 http://www.cs.toronto.edu/~eihsu/tutorial7 Getting Started Discursive Remarks. Organizational
Introduction to Bioinformatics Online Course: IBT
 Introduction to Bioinformatics Online Course: IBT Multiple Sequence Alignment Building Multiple Sequence Alignment Lec2 Choosing the Right Sequences Choosing the Right Sequences Before you build your alignment,
Introduction to Bioinformatics Online Course: IBT Multiple Sequence Alignment Building Multiple Sequence Alignment Lec2 Choosing the Right Sequences Choosing the Right Sequences Before you build your alignment,
Technical University of Munich. Exercise 8: Neural Networks
 Technical University of Munich Chair for Bioinformatics and Computational Biology Protein Prediction I for Computer Scientists SoSe 2018 Prof. B. Rost M. Bernhofer, M. Heinzinger, D. Nechaev, L. Richter
Technical University of Munich Chair for Bioinformatics and Computational Biology Protein Prediction I for Computer Scientists SoSe 2018 Prof. B. Rost M. Bernhofer, M. Heinzinger, D. Nechaev, L. Richter
Phylogenetics. Introduction to Bioinformatics Dortmund, Lectures: Sven Rahmann. Exercises: Udo Feldkamp, Michael Wurst
 Phylogenetics Introduction to Bioinformatics Dortmund, 16.-20.07.2007 Lectures: Sven Rahmann Exercises: Udo Feldkamp, Michael Wurst 1 Phylogenetics phylum = tree phylogenetics: reconstruction of evolutionary
Phylogenetics Introduction to Bioinformatics Dortmund, 16.-20.07.2007 Lectures: Sven Rahmann Exercises: Udo Feldkamp, Michael Wurst 1 Phylogenetics phylum = tree phylogenetics: reconstruction of evolutionary
A Study On Pair-Wise Local Alignment Of Protein Sequence For Identifying The Structural Similarity
 A Study On Pair-Wise Local Alignment Of Protein Sequence For Identifying The Structural Similarity G. Pratyusha, Department of Computer Science & Engineering, V.R.Siddhartha Engineering College(Autonomous)
A Study On Pair-Wise Local Alignment Of Protein Sequence For Identifying The Structural Similarity G. Pratyusha, Department of Computer Science & Engineering, V.R.Siddhartha Engineering College(Autonomous)
PyMod 2. User s Guide. PyMod 2 Documention (Last updated: 7/11/2016)
 PyMod 2 User s Guide PyMod 2 Documention (Last updated: 7/11/2016) http://schubert.bio.uniroma1.it/pymod/index.html Department of Biochemical Sciences A. Rossi Fanelli, Sapienza University of Rome, Italy
PyMod 2 User s Guide PyMod 2 Documention (Last updated: 7/11/2016) http://schubert.bio.uniroma1.it/pymod/index.html Department of Biochemical Sciences A. Rossi Fanelli, Sapienza University of Rome, Italy
Multiple Sequence Alignment II
 Multiple Sequence Alignment II Lectures 20 Dec 5, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20 Johnson Hall (JHN) 022 1 Outline
Multiple Sequence Alignment II Lectures 20 Dec 5, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20 Johnson Hall (JHN) 022 1 Outline
Skewed-Associative Caches: CS752 Final Project
 Skewed-Associative Caches: CS752 Final Project Professor Sohi Corey Halpin Scot Kronenfeld Johannes Zeppenfeld 13 December 2002 Abstract As the gap between microprocessor performance and memory performance
Skewed-Associative Caches: CS752 Final Project Professor Sohi Corey Halpin Scot Kronenfeld Johannes Zeppenfeld 13 December 2002 Abstract As the gap between microprocessor performance and memory performance
Fractal Compression. Related Topic Report. Henry Xiao. Queen s University. Kingston, Ontario, Canada. April 2004
 Fractal Compression Related Topic Report By Henry Xiao Queen s University Kingston, Ontario, Canada April 2004 Fractal Introduction Fractal is first introduced in geometry field. The birth of fractal geometry
Fractal Compression Related Topic Report By Henry Xiao Queen s University Kingston, Ontario, Canada April 2004 Fractal Introduction Fractal is first introduced in geometry field. The birth of fractal geometry
Heuristic methods for pairwise alignment:
 Bi03c_1 Unit 03c: Heuristic methods for pairwise alignment: k-tuple-methods k-tuple-methods for alignment of pairs of sequences Bi03c_2 dynamic programming is too slow for large databases Use heuristic
Bi03c_1 Unit 03c: Heuristic methods for pairwise alignment: k-tuple-methods k-tuple-methods for alignment of pairs of sequences Bi03c_2 dynamic programming is too slow for large databases Use heuristic
National Centre for Text Mining NaCTeM. e-science and data mining workshop
 National Centre for Text Mining NaCTeM e-science and data mining workshop John Keane Co-Director, NaCTeM john.keane@manchester.ac.uk School of Informatics, University of Manchester What is text mining?
National Centre for Text Mining NaCTeM e-science and data mining workshop John Keane Co-Director, NaCTeM john.keane@manchester.ac.uk School of Informatics, University of Manchester What is text mining?
Network Traffic Classification by Common Subsequence Finding
 Network Traffic Classification by Common Subsequence Finding Krzysztof Fabjański and Tomasz Kruk NASK, The Research Division, Wąwozowa 18, 2-796 Warszawa, Poland {krzysztof.fabjanski,tomasz.kruk}@nask.pl
Network Traffic Classification by Common Subsequence Finding Krzysztof Fabjański and Tomasz Kruk NASK, The Research Division, Wąwozowa 18, 2-796 Warszawa, Poland {krzysztof.fabjanski,tomasz.kruk}@nask.pl
Salvador Capella-Gutiérrez, Jose M. Silla-Martínez and Toni Gabaldón
 trimal: a tool for automated alignment trimming in large-scale phylogenetics analyses Salvador Capella-Gutiérrez, Jose M. Silla-Martínez and Toni Gabaldón Version 1.2b Index of contents 1. General features
trimal: a tool for automated alignment trimming in large-scale phylogenetics analyses Salvador Capella-Gutiérrez, Jose M. Silla-Martínez and Toni Gabaldón Version 1.2b Index of contents 1. General features
Finding data. HMMER Answer key
 Finding data HMMER Answer key HMMER input is prepared using VectorBase ClustalW, which runs a Java application for the graphical representation of the results. If you get an error message that blocks this
Finding data HMMER Answer key HMMER input is prepared using VectorBase ClustalW, which runs a Java application for the graphical representation of the results. If you get an error message that blocks this
mpmorfsdb: A database of Molecular Recognition Features (MoRFs) in membrane proteins. Introduction
 mpmorfsdb: A database of Molecular Recognition Features (MoRFs) in membrane proteins. Introduction Molecular Recognition Features (MoRFs) are short, intrinsically disordered regions in proteins that undergo
mpmorfsdb: A database of Molecular Recognition Features (MoRFs) in membrane proteins. Introduction Molecular Recognition Features (MoRFs) are short, intrinsically disordered regions in proteins that undergo
MetAmp: a tool for Meta-Amplicon analysis User Manual
 November 12, 2014 MetAmp: a tool for Meta-Amplicon analysis User Manual Ilya Y. Zhbannikov 1, Janet E. Williams 1, James A. Foster 1,2,3 3 Institute for Bioinformatics and Evolutionary Studies, University
November 12, 2014 MetAmp: a tool for Meta-Amplicon analysis User Manual Ilya Y. Zhbannikov 1, Janet E. Williams 1, James A. Foster 1,2,3 3 Institute for Bioinformatics and Evolutionary Studies, University
PROJECT PERIODIC REPORT
 PROJECT PERIODIC REPORT Grant Agreement number: 257403 Project acronym: CUBIST Project title: Combining and Uniting Business Intelligence and Semantic Technologies Funding Scheme: STREP Date of latest
PROJECT PERIODIC REPORT Grant Agreement number: 257403 Project acronym: CUBIST Project title: Combining and Uniting Business Intelligence and Semantic Technologies Funding Scheme: STREP Date of latest
On Generalizations and Improvements to the Shannon-Fano Code
 Acta Technica Jaurinensis Vol. 10, No.1, pp. 1-12, 2017 DOI: 10.14513/actatechjaur.v10.n1.405 Available online at acta.sze.hu On Generalizations and Improvements to the Shannon-Fano Code D. Várkonyi 1,
Acta Technica Jaurinensis Vol. 10, No.1, pp. 1-12, 2017 DOI: 10.14513/actatechjaur.v10.n1.405 Available online at acta.sze.hu On Generalizations and Improvements to the Shannon-Fano Code D. Várkonyi 1,
Comparison of Sequence Similarity Measures for Distant Evolutionary Relationships
 Comparison of Sequence Similarity Measures for Distant Evolutionary Relationships Abhishek Majumdar, Peter Z. Revesz Department of Computer Science and Engineering, University of Nebraska-Lincoln, Lincoln,
Comparison of Sequence Similarity Measures for Distant Evolutionary Relationships Abhishek Majumdar, Peter Z. Revesz Department of Computer Science and Engineering, University of Nebraska-Lincoln, Lincoln,
Evaluating the Effect of Perturbations in Reconstructing Network Topologies
 DSC 2 Working Papers (Draft Versions) http://www.ci.tuwien.ac.at/conferences/dsc-2/ Evaluating the Effect of Perturbations in Reconstructing Network Topologies Florian Markowetz and Rainer Spang Max-Planck-Institute
DSC 2 Working Papers (Draft Versions) http://www.ci.tuwien.ac.at/conferences/dsc-2/ Evaluating the Effect of Perturbations in Reconstructing Network Topologies Florian Markowetz and Rainer Spang Max-Planck-Institute
Arbiter: the Evaluation Tool in the Contests of the China NOI
 Olympiads in Informatics, 2013, Vol. 7, 180 185 180 2013 Vilnius University Arbiter: the Evaluation Tool in the Contests of the China NOI Qiyang ZHAO 1, Fan WANG 1, Baolin YIN 1,HuiSUN 2 1 School of Computer
Olympiads in Informatics, 2013, Vol. 7, 180 185 180 2013 Vilnius University Arbiter: the Evaluation Tool in the Contests of the China NOI Qiyang ZHAO 1, Fan WANG 1, Baolin YIN 1,HuiSUN 2 1 School of Computer
Biochemistry 324 Bioinformatics. Multiple Sequence Alignment (MSA)
 Biochemistry 324 Bioinformatics Multiple Sequence Alignment (MSA) Big- Οh notation Greek omicron symbol Ο The Big-Oh notation indicates the complexity of an algorithm in terms of execution speed and storage
Biochemistry 324 Bioinformatics Multiple Sequence Alignment (MSA) Big- Οh notation Greek omicron symbol Ο The Big-Oh notation indicates the complexity of an algorithm in terms of execution speed and storage
Computational Molecular Biology
 Computational Molecular Biology Erwin M. Bakker Lecture 3, mainly from material by R. Shamir [2] and H.J. Hoogeboom [4]. 1 Pairwise Sequence Alignment Biological Motivation Algorithmic Aspect Recursive
Computational Molecular Biology Erwin M. Bakker Lecture 3, mainly from material by R. Shamir [2] and H.J. Hoogeboom [4]. 1 Pairwise Sequence Alignment Biological Motivation Algorithmic Aspect Recursive
Parsimony-Based Approaches to Inferring Phylogenetic Trees
 Parsimony-Based Approaches to Inferring Phylogenetic Trees BMI/CS 576 www.biostat.wisc.edu/bmi576.html Mark Craven craven@biostat.wisc.edu Fall 0 Phylogenetic tree approaches! three general types! distance:
Parsimony-Based Approaches to Inferring Phylogenetic Trees BMI/CS 576 www.biostat.wisc.edu/bmi576.html Mark Craven craven@biostat.wisc.edu Fall 0 Phylogenetic tree approaches! three general types! distance:
Enhancing Internet Search Engines to Achieve Concept-based Retrieval
 Enhancing Internet Search Engines to Achieve Concept-based Retrieval Fenghua Lu 1, Thomas Johnsten 2, Vijay Raghavan 1 and Dennis Traylor 3 1 Center for Advanced Computer Studies University of Southwestern
Enhancing Internet Search Engines to Achieve Concept-based Retrieval Fenghua Lu 1, Thomas Johnsten 2, Vijay Raghavan 1 and Dennis Traylor 3 1 Center for Advanced Computer Studies University of Southwestern
Two Phase Evolutionary Method for Multiple Sequence Alignments
 The First International Symposium on Optimization and Systems Biology (OSB 07) Beijing, China, August 8 10, 2007 Copyright 2007 ORSC & APORC pp. 309 323 Two Phase Evolutionary Method for Multiple Sequence
The First International Symposium on Optimization and Systems Biology (OSB 07) Beijing, China, August 8 10, 2007 Copyright 2007 ORSC & APORC pp. 309 323 Two Phase Evolutionary Method for Multiple Sequence
Efficient, Scalable, and Provenance-Aware Management of Linked Data
 Efficient, Scalable, and Provenance-Aware Management of Linked Data Marcin Wylot 1 Motivation and objectives of the research The proliferation of heterogeneous Linked Data on the Web requires data management
Efficient, Scalable, and Provenance-Aware Management of Linked Data Marcin Wylot 1 Motivation and objectives of the research The proliferation of heterogeneous Linked Data on the Web requires data management
FREQUENT PATTERN MINING IN BIG DATA USING MAVEN PLUGIN. School of Computing, SASTRA University, Thanjavur , India
 Volume 115 No. 7 2017, 105-110 ISSN: 1311-8080 (printed version); ISSN: 1314-3395 (on-line version) url: http://www.ijpam.eu ijpam.eu FREQUENT PATTERN MINING IN BIG DATA USING MAVEN PLUGIN Balaji.N 1,
Volume 115 No. 7 2017, 105-110 ISSN: 1311-8080 (printed version); ISSN: 1314-3395 (on-line version) url: http://www.ijpam.eu ijpam.eu FREQUENT PATTERN MINING IN BIG DATA USING MAVEN PLUGIN Balaji.N 1,
Dynamic Programming: Sequence alignment. CS 466 Saurabh Sinha
 Dynamic Programming: Sequence alignment CS 466 Saurabh Sinha DNA Sequence Comparison: First Success Story Finding sequence similarities with genes of known function is a common approach to infer a newly
Dynamic Programming: Sequence alignment CS 466 Saurabh Sinha DNA Sequence Comparison: First Success Story Finding sequence similarities with genes of known function is a common approach to infer a newly
Biological Sequence Matching Using Fuzzy Logic
 International Journal of Scientific & Engineering Research Volume 2, Issue 7, July-2011 1 Biological Sequence Matching Using Fuzzy Logic Nivit Gill, Shailendra Singh Abstract: Sequence alignment is the
International Journal of Scientific & Engineering Research Volume 2, Issue 7, July-2011 1 Biological Sequence Matching Using Fuzzy Logic Nivit Gill, Shailendra Singh Abstract: Sequence alignment is the
Sequence alignment theory and applications Session 3: BLAST algorithm
 Sequence alignment theory and applications Session 3: BLAST algorithm Introduction to Bioinformatics online course : IBT Sonal Henson Learning Objectives Understand the principles of the BLAST algorithm
Sequence alignment theory and applications Session 3: BLAST algorithm Introduction to Bioinformatics online course : IBT Sonal Henson Learning Objectives Understand the principles of the BLAST algorithm
Research Article Aligning Sequences by Minimum Description Length
 Hindawi Publishing Corporation EURASIP Journal on Bioinformatics and Systems Biology Volume 2007, Article ID 72936, 14 pages doi:10.1155/2007/72936 Research Article Aligning Sequences by Minimum Description
Hindawi Publishing Corporation EURASIP Journal on Bioinformatics and Systems Biology Volume 2007, Article ID 72936, 14 pages doi:10.1155/2007/72936 Research Article Aligning Sequences by Minimum Description
