EVIDENCE FOR SHOWING GENE/PROTEIN NAME SUGGESTIONS IN BIOSCIENCE LITERATURE SEARCH INTERFACES
|
|
|
- Gillian Harris
- 6 years ago
- Views:
Transcription
1 EVIDENCE FOR SHOWING GENE/PROTEIN NAME SUGGESTIONS IN BIOSCIENCE LITERATURE SEARCH INTERFACES ANNA DIVOLI, MARTI A. HEARST, MICHAEL A. WOOLDRIDGE School of Information, UC Berkeley This paper reports on the results of two questionnaires asking biologists about the incorporation of text-extracted entity information, specifically gene and protein names, into bioscience literature search user interfaces. Among the findings are that study participants want to see gene/protein metadata in combination with organism information; that a significant proportion would like to see gene names grouped by type (synonym, homolog, etc.), and that most participants want to see information that the system is confident about immediately, and see less certain information after taking additional action. These results inform future interface designs. 1. Introduction Bioinformaticians have developed numerous algorithms for extracting entity and relation information from the bioscience literature, and have developed some very interesting user interfaces for showing this information. However, little research has been done on the usability of these systems and how to best incorporate such information into literature search and text mining interfaces. As part of an on-going project to build a highly usable literature search tool for bioscience researchers, we are carefully investigating what kinds of biological information searchers want to see, as well as how they want to see this information presented. We are interested in supporting biologists whose main tasks are biological (as opposed to database curators and bioinformaticians doing text mining) and who presumably do not want to spend a lot of time searching. We use methods from the field of Human Computer Interaction (HCI) for the careful design of search interfaces. We have already used these methods to develop a novel bioliterature search user interface whose focus is allowing users to search over and view figures and captions 5 (see That interface is based on the observation that many researchers, when assessing a research article, first look at the title, abstract, and figures. In this paper, we investigate whether or not bioscience literature searchers wish to see related term suggestions, in particular, gene and protein names, in
2 response to their queries. a This is one step in a larger investigation in which we plan to assess presentation of other results of text analysis, such as the entities corresponding to diseases, pathways, gene interactions, localization information, function information, and so on. When it comes to presenting users with the output of text mining programs, the interface designer is faced with an embarrassment of riches. There are many choices of entity and relationship information that can be displayed to the searcher. However, search user interface research suggests that users are quickly overwhelmed when presented with too many options and too much information. Therefore, our approach is to assess the usability of one feature at a time, see how participants respond, and then test out other features. We focus on gene names here because of their prominent role in the queries devised for the TREC Genomics track 6, and because of their focus in text mining efforts, as seen in the BioCreative text analysis competitions 7. Thus, this paper assesses one way in which the output of text mining can be useful for bioscience software tools. In the remainder of this paper, we first describe the user-centered design process and then discuss related work. We then report on the results of two questionnaires. The first asked participants a number of questions about how they search the bioscience literature, including questions about their use of gene names. Among the findings were that participants did indeed want to see suggestions of gene names as part of their search experience. The second questionnaire, building on these results, asked participants to assess several designs for presenting gene names in a search user interface. Finally, we conclude the paper with plans for acting on the results of this study. 2. The User-Centered Design Process We are following the method of user-centered design, which is standard practice in the field of Human-Computer Interaction (HCI) 11. This method focuses on making decisions about the design of a user interface based on feedback obtained from target users of the system, rather than coding first and evaluating later. First a needs assessment is performed in which the designers investigate who the users are, what their goals are, and what tasks they have to complete in order to achieve those goals. The next stage is a task analysis in which the designers characterize which steps the users need to take to complete their tasks, decide which user goals they will attempt to support, and then create scenarios which exemplify these tasks being executed by the target user population. a For the remainder of the paper, we will use the term gene name to refer to both gene and protein names.
3 Once the target user goals and tasks have been determined, design is done in a tight evaluation cycle consisting of mocking up prototypes, obtaining reactions from potential users, and revising the designs based on those reactions. This sequence of activities often needs to be repeated several times before a satisfactory design has been achieved. This is often referred to as discount usability testing, since useful results can be obtained with only a few participants. After a design is testing well in informal studies, formal experiments comparing different designs and measuring for statistically significant differences can be conducted. This iterative procedure is necessary because interface design is still more of an art than a science. There are usually several good solutions within the interface design space, and the task of the designers is to navigate through the design space until reaching some local optimum. The iterative process allows study participants to help the designers make decisions about which paths to explore in that space. Experienced designers often know how to start close to a good solution; less experienced designers need to do more work. Designing for an entirely novel interaction paradigm often requires more iteration and experimentation. 3. Research on Term Suggestions Usability An important class of query reformulation aids is automatically suggested term refinements and expansions. Spelling correction suggestions are query reformulation aids, but the phrase term expansion is usually applied to tools that suggest alternative wordings. Usability studies are generally positive as to the efficacy of term suggestions when users are not required to make relevance judgements and do not have to choose among too many terms. Those that produce negative results seem to stem from problems with the presentation interface 2. Interfaces that allow users to reformulate their query by selecting a single term (usually via a hyperlink) seem to fare better. Anick 1 describes the results of a large-scale investigation of the effects of incorporating related term suggestions into a major web search engine. The term suggestion tool, called Prisma, was placed within the Altavista search engine s results page. The number of feedback terms was limited to 12 to conserve space in the display and minimize cognitive load. In a large web-based study, 16% of users applied the Prisma feedback mechanism at least once on any given day. However, effectiveness when measured in the occurrence of search results clicks did not differ between the baseline and the Prisma groups. In a more recent study, Jansen et al. 9 analyzed 1.5M queries from a log taken in 2005 from the Dogpile.com metasearch engine. The interface for this engine shows suggested additional terms in a box on the righthand side under the heading
4 Are you looking for? Jansen et al. found that 8.4% of all queries were generated by the reformulation assistant provided by Dogpile. Thus, there is evidence that searchers use such term reformulations, although the benefits are as yet unproven. 4. Current Bioliterature Search Interfaces There are a number of innovative interfaces for analyzing the results of text analysis. The ihop system 8 converts the contents of PubMed abstracts into a network of information about genes and interactions, displaying sentences extracted from abstracts and annotated with entity information. The ChiliBot 3 system also shows extracted information in the form of relationships between genes, proteins, and keywords. TextPresso 10 uses an ontology to search over the full text of a collection of articles about C. elegans, extracting out sentences that contain entities and relations of interest. These systems have not been assessed in terms of usability of their interface or their features. The GoPubMed system 4 shows a wealth of information in search results over PubMed. Most prominent is a hierarchical display of a wide range of categories from the Gene Ontology and MeSH associated with the article. Users may sort search results by navigating in this hierarchy and selecting categories. This interface is compelling, but it is not clear which kinds of information are most useful to show, whether a hierarchy is the best way to show metadata information for grouping search results, and whether or not this is too much information to show. The goal of this paper is to make a start at determining which kinds of information searchers want to see, and how they want to select it. 5. First Questionnaire: Biological Information Preferences Both studies were administered in the form of an online questionnaire. For the first study, we recruited biosciences researchers from 7 research institutions via lists and personal contacts. The 38 participants were all from academic institutions (22 graduate students, 6 postdoctoral researchers, 5 faculty, and 5 others), and had a wide range of specialties, including systems biology, bioinformatics, genomics, biochemistry, cellular and evolutionary biology, microbiology, physiology and ecology. Figure 1 shows the percentage of time each participant uses computers for their work. A surprising 37% say they use computers for % of the time they are working, although only 6 participants listed bioinformatics as one of their fields. Participants were for the most part heavy users of literature search; 84% said they search biomedical literature either daily or weekly. We asked participants which existing literature search tools they use, and for
5 Figure 1. names. Statistics on computer use, search frequency, and percentage of queries that include gene what percent of their searches. 12 participants (32%) said they use PubMed 80% of the time or more; on average it was used 50% of the time. Google Scholar was used on average 25% of the time; all but 3 participants used it at least some of the time. 6 participants used Ovid at least 5% of the time. The other popular search engine mentioned was the ISI Web of Science, which 9 participants used; 2 said they used it more than 90% of the time. Also mentioned were BIOSIS (3 mentions), Connotea (1), PubMedCentral (1), Google web search (1), and bloglines (1). Figure 1 shows the responses to a question on what proportion of searches include gene names. 37% of the participants use gene names in % of their queries. Five participants do not use gene names in their queries; one of these
6 people noted that they use literature search in order to discover relevant genes. Next, participants answered two detailed questions about what kinds of information they would like to see associated with the gene name from their query. Table 1 shows the averaged scores for responses to the question When you search for genes/proteins, what type of related gene/protein names would you like a system to suggest? Participants selected choices from a Likert scale which spanned from 1 ( strongly do not want to see this ) to 5 ( extremely important to see this information ), with 3 indicating do not mind seeing this. (These results are for 33 participants, because the 5 participants who said they do not use gene names in their search were made to automatically skip these questions.) The table below also shows the number of participants who assigned either a 1 or a 2 score, indicating that they do not want to see this kind of information. Table 1. Averaged scores for responses to the question When you search for genes/proteins, what type of related gene/protein names would you like a system to suggest? 1 is strongly disagree, 5 is strongly agree. Related Information Type Avg. rating # (%) selecting 1 or 2 Gene s synonyms (5%) Gene s synonyms refined by organism (5%) Gene s homologs (13%) Genes from the same family: parents (18%) Genes from the same family: children (10%) Genes from the same family: siblings (24%) The next question, When you search for genes/proteins what other related information would you like a system to return? used the same rating scale as above. The results are shown in Table 2. Table 2. Averaged scores for responses to the question When you search for genes/proteins what other related information would you like a system to return? using same rating scale as above. Related Information Type Avg. rating # (%) selecting 1 or 2 Genes this gene interacts with (10%) Diseases this gene is associated with (16%) Chemicals/drugs this gene is associated with (21%) Localization information for this gene (8%) When asked for additional information of interest, people suggested: pathways (suggested 4 times), experimental modification, promoter information, lists of organisms for which the gene is sequenced, ability to limit searches to a tax-
7 onomic group, protein motifs, hypothesized or known functions, downstream effects and link to a model organism page. The results of this questionnaire suggest that not only are many biologists heavy users of literature search, but gene names figure prominently in a significant proportion of their searches. Furthermore, there is interest in seeing information associated with gene names. Not surprisingly, the more directly related the information is to the gene, the more participants viewed it favorably. 22 participants said they thought gene synonyms would be extremely useful (i.e., rated this choice with a score of 5). However, as the third columns of the tables show, a notable minority of participants expressed opposition to showing the additional information. In a box asking for general comments, two participants noted that for some kinds of searches, expansion information would be useful, but for others the extra information would be in the way. One participant suggested offering these options at the start of the search as a link to follow optionally. These responses reflect a common view among users of search systems: they do not want to see a cluttered display. This is further warning that one should proceed with caution when adding information to a search user interface. 6. Second Questionnaire: Gene/Protein Name Expansion Preferences 6.1. The Evaluated Designs To reproduce what users would see in a Web search interface, four designs were constructed using HTML and CSS, building upon the design used for our group s search engine. To constrain the participants evaluation of the designs and to focus them on a specific aspect of the interface, static screenshots of just the relevant portion of the search interface were used in the testing. Example interactions with the interface were conveyed using before and after screenshots of the designs. Limiting the testing to static screenshots decreased the development time required to set up the tests, since we did not need to anticipate the myriad potential interactions between the testers and a live interface. Figures 2 4 show the screenshots seen by the participants for Designs 1 4. Participants were told they were seeing what happened after they clicked on the indicated link, but not what happens to the search results after the new search is executed. Design 1, which served as the baseline for comparison with the other designs, showed a standard search engine interface with a text box and submit button in the page header. The gene term RAD23 was used as the example search term, with a results summary showing three results returned. Design 2 added a horizontal box between the search box and the text summary. The box listed possible expansion terms for the original RAD23 query
8 Design 1 Design 2 Figure 2. Designs 1 and 2 shown to participants in the second questionnaire. organized under four categories: synonyms, homologs, parents, and siblings. All the terms were hyperlinked. The after screenshot showed the result of clicking a hyperlinked term, which added that term to the query in the text box using an
9 Figure 3. Design 3 shown to participants in the second questionnaire. OR operator. Design 3 had a similar layout except that instead of having hyperlinked expansion terms, each expansion term was paired with a checkbox. The terms were organized beneath the same four categories. The after screenshot showed that by clicking a checkbox, a user could add the term to the original query. Design 4 showed a box of plain text expansion terms that were neither hyperlinked nor paired with checkboxes. In this design, each category term had an Add all to query link next to it for adding all of a category s terms at once. The after screenshot showed the result of clicking a hyperlink, with multiple terms ORed to the original query Results Nineteen people completed the questionnaire. Nine of those who filled out the first questionnaire and who indicated that (a) they were interested in seeing gene/protein names in search results and (b) they were willing to be contacted for a second questionnaire participated in this followup study. Ten additional participants were recruited by ing colleagues and asking them to forward the
10 Figure 4. Design 4 shown to participants in the second questionnaire. request to biologists. Thus, the results are biased towards people who are interested in search interfaces and their improvement. Again, participants were from several academic institutions (4 graduate students, 7 postdoctoral researchers, 3 faculty, and 5 other researchers). Their areas of interest/specialization included molecular toxicology, evolutionary genomics, chromosome biology, plant reproductive biology, cell signaling networks, and computational biology more generally. The distribution of usage of genes in searches was similar to that of the first questionnaire. One question asked the participants to rank-order the designs. There was a clear preference for the expansion terms over the baseline, which was the lowest ranked for 15 out of 19 participants. Table 3 shows the results, with Design 3 most favored, followed by Designs 4 and 2, which were similarly ranked. In the next phase of questions, one participant indicated they would not like to see gene names, and so automatically skipped the questions. Of the remaining 18 participants, when asked to indicate a preference for clicking on hyperlinks versus checkboxes for adding gene names to the query, 10 participants (56%) selected checkboxes and 6 (33%) selected hyperlinks (one suggested a select all
11 Table 3. Design Preferences. # participants who rated % participants who rated Avg. rating Design 1st or 2nd Design 1st or 2nd (1=low, 4=high) Design % 3.3 Design % 2.6 Design % 2.5 Design 1 0 0% 1.6 option above each group for the checkboxes). When asked to indicate whether or not they would like to see the organisms associated with each gene name, 16 out of 18 participants said they would like the organism information to be directly visible, showing the organism either alongside (11) or grouping the gene names by organism (5). Two were undecided. When asked how gene names should be organized in the display, 9 preferred them to be grouped under type (synonyms, homologs, etc). The other participants were split between preferences for showing the information grouped by organism name, grouped by more generic taxonomic information, or not grouped but shown alphabetically or by frequency of occurrence in the collection. Participants were also asked if they prefer to select each gene individually (2), whole groups of gene names with one click (3), or to have the option to chose either individual names or whole groups with one click (13). Finally, they were asked if they prefer the system to suggest only names that it is highly confident are related (8), include names that it is less confident about (0), or include names that it is less confident about under a show more link (8). In the open comments field, one participant stated that the system should allow the user to choose among these, and another wrote something we could not interpret. These attitudes echo the finding that high-scoring systems in the TREC genomics track 6 often used principled gene name expansion. 7. Conclusions and Future Work This study addresses the results of the first steps of user-centered design for development of a literature search interface for biologists. Our needs assessment has revealed a strong desire for the search system to suggest information closely related to gene names, and some interest in less closely related information as well. Our task analysis has revealed that most participants want to see organism names in conjunction with gene names, a majority of participants prefer to see term suggestions grouped by type, and participants are split in preference between single-click hyperlink interaction and checkbox-style interaction. The last point suggests that we experiment with hybrid designs in which only hyperlinks
12 are used, but an additional new hyperlink allows for selecting all items in a group. Another hybrid to evaluate would have checkboxes for the individual terms and a link that immediately adds all terms in the group and executes the query. The second questionnaire did not ask participants to choose between seeing information related to genes and other kinds of metadata such as disease names. Adding additional information will require a delicate balancing act between usefulness and clutter. Another design idea would allow users to collapse and expand term suggestions of different types; we intend to test that as well. Armed with these results, we have reason to be confident that the designs will be found usable. Our next steps will be to implement prototypes of these designs, ask participants to perform queries, and contrast the different interaction styles. Acknowledgements: We thank the survey participants for their contributions to this work. This research was supported in part by NSF DBI References 1. P. Anick. Using terminological feedback for web search refinement: a log-based study. Proceedings of SIGIR 2003, pages 88 95, P. Bruza, R. McArthur, and S. Dennis. Interactive Internet search: keyword, directory and query reformulation mechanisms compared. Proceedings of SIGIR 2000, pages , H. Chen and B.M. Sharp. Content-rich biological network constructed by mining PubMed abstracts. BMC Bioinformatics, 5(147), A. Doms and M. Schroeder. GoPubMed: exploring PubMed with the Gene Ontology. Nucleic Acids Research, 33(1):W783 W786, M.A. Hearst, A. Divoli, J. Ye, and M.A. Wooldridge. Exploring the efficacy of caption search for bioscience journal search interfaces. Biological, translational, and clinical language processing, pages 73 80, W. Hersh, A. Cohen, J. Yang, R.T. Bhupatiraju, P. Roberts, and M. Hearst. TREC 2005 Genomics Track Overview. the Fourteenth Text Retrieval Conference, L. Hirschman, A. Yeh, C. Blaschke, and A. Valencia. Overview of BioCreAtIvE: critical assessment of information extraction for biology. BMC Bioinformatics, 6(S1), R. Hoffmann and A. Valencia. A gene network for navigating the literature. Nature Genetics, 36(7):664, B.J. Jansen, A. Spink, and S. Koshman. Web searcher interaction with the Dogpile.com metasearch engine. Journal of the American Society for Information Science and Technology, 58(5): , H.M. Müller, E.E. Kenny, and P.W. Sternberg. Textpresso: an ontology-based information retrieval and extraction system for biological literature. PLoS Biol, 2(11):e309, B. Shneiderman and C. Plaisant. Designing the user interface: strategies for effective human-computer interaction, 4/E. Addison-Wesley, Reading, MA, 2004.
Exploring the Efficacy of Caption Search for Bioscience Journal Search Interfaces
 Exploring the Efficacy of Caption Search for Bioscience Journal Search Interfaces Marti A. Hearst, Anna Divoli, Jerry Ye School of Information, UC Berkeley Berkeley, CA 94720 {hearst,divoli,jerryye}@ischool.berkeley.edu
Exploring the Efficacy of Caption Search for Bioscience Journal Search Interfaces Marti A. Hearst, Anna Divoli, Jerry Ye School of Information, UC Berkeley Berkeley, CA 94720 {hearst,divoli,jerryye}@ischool.berkeley.edu
Full Text and Figure Display Improves Bioscience Literature Search
 Full Text and Figure Display Improves Bioscience Literature Search Anna Divoli, Michael A. Wooldridge, Marti A. Hearst* School of Information, University of California, Berkeley, California, United States
Full Text and Figure Display Improves Bioscience Literature Search Anna Divoli, Michael A. Wooldridge, Marti A. Hearst* School of Information, University of California, Berkeley, California, United States
TheHotMap.com: Enabling Flexible Interaction in Next-Generation Web Search Interfaces
 TheHotMap.com: Enabling Flexible Interaction in Next-Generation Web Search Interfaces Orland Hoeber Department of Computer Science Memorial University of Newfoundland St. John s, NL, Canada A1B 3X5 hoeber@cs.mun.ca
TheHotMap.com: Enabling Flexible Interaction in Next-Generation Web Search Interfaces Orland Hoeber Department of Computer Science Memorial University of Newfoundland St. John s, NL, Canada A1B 3X5 hoeber@cs.mun.ca
Information Retrieval, Information Extraction, and Text Mining Applications for Biology. Slides by Suleyman Cetintas & Luo Si
 Information Retrieval, Information Extraction, and Text Mining Applications for Biology Slides by Suleyman Cetintas & Luo Si 1 Outline Introduction Overview of Literature Data Sources PubMed, HighWire
Information Retrieval, Information Extraction, and Text Mining Applications for Biology Slides by Suleyman Cetintas & Luo Si 1 Outline Introduction Overview of Literature Data Sources PubMed, HighWire
Using Clusters on the Vivisimo Web Search Engine
 Using Clusters on the Vivisimo Web Search Engine Sherry Koshman and Amanda Spink School of Information Sciences University of Pittsburgh 135 N. Bellefield Ave., Pittsburgh, PA 15237 skoshman@sis.pitt.edu,
Using Clusters on the Vivisimo Web Search Engine Sherry Koshman and Amanda Spink School of Information Sciences University of Pittsburgh 135 N. Bellefield Ave., Pittsburgh, PA 15237 skoshman@sis.pitt.edu,
Query Modifications Patterns During Web Searching
 Bernard J. Jansen The Pennsylvania State University jjansen@ist.psu.edu Query Modifications Patterns During Web Searching Amanda Spink Queensland University of Technology ah.spink@qut.edu.au Bhuva Narayan
Bernard J. Jansen The Pennsylvania State University jjansen@ist.psu.edu Query Modifications Patterns During Web Searching Amanda Spink Queensland University of Technology ah.spink@qut.edu.au Bhuva Narayan
The LAILAPS Search Engine - A Feature Model for Relevance Ranking in Life Science Databases
 International Symposium on Integrative Bioinformatics 2010 The LAILAPS Search Engine - A Feature Model for Relevance Ranking in Life Science Databases M Lange, K Spies, C Colmsee, S Flemming, M Klapperstück,
International Symposium on Integrative Bioinformatics 2010 The LAILAPS Search Engine - A Feature Model for Relevance Ranking in Life Science Databases M Lange, K Spies, C Colmsee, S Flemming, M Klapperstück,
A Framework for BioCuration (part II)
 A Framework for BioCuration (part II) Text Mining for the BioCuration Workflow Workshop, 3rd International Biocuration Conference Friday, April 17, 2009 (Berlin) Martin Krallinger Spanish National Cancer
A Framework for BioCuration (part II) Text Mining for the BioCuration Workflow Workshop, 3rd International Biocuration Conference Friday, April 17, 2009 (Berlin) Martin Krallinger Spanish National Cancer
Web document summarisation: a task-oriented evaluation
 Web document summarisation: a task-oriented evaluation Ryen White whiter@dcs.gla.ac.uk Ian Ruthven igr@dcs.gla.ac.uk Joemon M. Jose jj@dcs.gla.ac.uk Abstract In this paper we present a query-biased summarisation
Web document summarisation: a task-oriented evaluation Ryen White whiter@dcs.gla.ac.uk Ian Ruthven igr@dcs.gla.ac.uk Joemon M. Jose jj@dcs.gla.ac.uk Abstract In this paper we present a query-biased summarisation
Usability Evaluation of Cell Phones for Early Adolescent Users
 Yassierli*, Melati Gilang Industrial Management Research Division, Faculty of Industrial Technology, Bandung Institute of Technology Jl. Ganesa 10 Bandung 40134 Indonesia ABSTRACT:. The increasing number
Yassierli*, Melati Gilang Industrial Management Research Division, Faculty of Industrial Technology, Bandung Institute of Technology Jl. Ganesa 10 Bandung 40134 Indonesia ABSTRACT:. The increasing number
Hyacinth Macaws for Seniors Survey Report
 Hyacinth Macaws for Seniors Survey Report http://stevenmoskowitz24.com/hyacinth_macaw/ Steven Moskowitz IM30930 Usability Testing Spring, 2015 March 24, 2015 TABLE OF CONTENTS Introduction...1 Executive
Hyacinth Macaws for Seniors Survey Report http://stevenmoskowitz24.com/hyacinth_macaw/ Steven Moskowitz IM30930 Usability Testing Spring, 2015 March 24, 2015 TABLE OF CONTENTS Introduction...1 Executive
Analyzing Web Multimedia Query Reformulation Behavior
 Analyzing Web Multimedia Query Reformulation Behavior Liang-Chun Jack Tseng Faculty of Science and Queensland University of Brisbane, QLD 4001, Australia ntjack.au@hotmail.com Dian Tjondronegoro Faculty
Analyzing Web Multimedia Query Reformulation Behavior Liang-Chun Jack Tseng Faculty of Science and Queensland University of Brisbane, QLD 4001, Australia ntjack.au@hotmail.com Dian Tjondronegoro Faculty
Databases available to ISU researchers:
 Databases available to ISU researchers: Table of Contents Web of Knowledge Overview 3 Web of Science 4 Cited Reference Searching 5 Secondary Cited Author Searching 8 Eliminating Self-Citations 9 Saving
Databases available to ISU researchers: Table of Contents Web of Knowledge Overview 3 Web of Science 4 Cited Reference Searching 5 Secondary Cited Author Searching 8 Eliminating Self-Citations 9 Saving
Measuring inter-annotator agreement in GO annotations
 Measuring inter-annotator agreement in GO annotations Camon EB, Barrell DG, Dimmer EC, Lee V, Magrane M, Maslen J, Binns ns D, Apweiler R. An evaluation of GO annotation retrieval for BioCreAtIvE and GOA.
Measuring inter-annotator agreement in GO annotations Camon EB, Barrell DG, Dimmer EC, Lee V, Magrane M, Maslen J, Binns ns D, Apweiler R. An evaluation of GO annotation retrieval for BioCreAtIvE and GOA.
Chapter 27 Introduction to Information Retrieval and Web Search
 Chapter 27 Introduction to Information Retrieval and Web Search Copyright 2011 Pearson Education, Inc. Publishing as Pearson Addison-Wesley Chapter 27 Outline Information Retrieval (IR) Concepts Retrieval
Chapter 27 Introduction to Information Retrieval and Web Search Copyright 2011 Pearson Education, Inc. Publishing as Pearson Addison-Wesley Chapter 27 Outline Information Retrieval (IR) Concepts Retrieval
UNIVERSITY OF BOLTON WEB PUBLISHER GUIDE JUNE 2016 / VERSION 1.0
 UNIVERSITY OF BOLTON WEB PUBLISHER GUIDE WWW.BOLTON.AC.UK/DIA JUNE 2016 / VERSION 1.0 This guide is for staff who have responsibility for webpages on the university website. All Web Publishers must adhere
UNIVERSITY OF BOLTON WEB PUBLISHER GUIDE WWW.BOLTON.AC.UK/DIA JUNE 2016 / VERSION 1.0 This guide is for staff who have responsibility for webpages on the university website. All Web Publishers must adhere
The information in the database is taken from a range of publication types including journals, books, meeting and patents.
 UNIVERSITY OF ULSTER LIBRARY BIOSIS Previews Backfile 1969-2008 Coverage The backfile of BIOSIS Previews, covering the years 1969 to 2008, is available on the ISI Web of Knowledge platform It is a major
UNIVERSITY OF ULSTER LIBRARY BIOSIS Previews Backfile 1969-2008 Coverage The backfile of BIOSIS Previews, covering the years 1969 to 2008, is available on the ISI Web of Knowledge platform It is a major
A User Study on Features Supporting Subjective Relevance for Information Retrieval Interfaces
 A user study on features supporting subjective relevance for information retrieval interfaces Lee, S.S., Theng, Y.L, Goh, H.L.D., & Foo, S. (2006). Proc. 9th International Conference of Asian Digital Libraries
A user study on features supporting subjective relevance for information retrieval interfaces Lee, S.S., Theng, Y.L, Goh, H.L.D., & Foo, S. (2006). Proc. 9th International Conference of Asian Digital Libraries
A Model for Interactive Web Information Retrieval
 A Model for Interactive Web Information Retrieval Orland Hoeber and Xue Dong Yang University of Regina, Regina, SK S4S 0A2, Canada {hoeber, yang}@uregina.ca Abstract. The interaction model supported by
A Model for Interactive Web Information Retrieval Orland Hoeber and Xue Dong Yang University of Regina, Regina, SK S4S 0A2, Canada {hoeber, yang}@uregina.ca Abstract. The interaction model supported by
Relevance Feedback and Query Reformulation. Lecture 10 CS 510 Information Retrieval on the Internet Thanks to Susan Price. Outline
 Relevance Feedback and Query Reformulation Lecture 10 CS 510 Information Retrieval on the Internet Thanks to Susan Price IR on the Internet, Spring 2010 1 Outline Query reformulation Sources of relevance
Relevance Feedback and Query Reformulation Lecture 10 CS 510 Information Retrieval on the Internet Thanks to Susan Price IR on the Internet, Spring 2010 1 Outline Query reformulation Sources of relevance
Overview. TREC Genomics Track Plenary. The central dogma of biology. At the intersection of digital biology and IR. Overview of this session
 Overview TREC Genomics Track Plenary William Hersh Track Chair Oregon Health & Science University hersh@ohsu.edu http://medir.ohsu.edu/~genomics Introductory comments Track history 2003 track Primary task
Overview TREC Genomics Track Plenary William Hersh Track Chair Oregon Health & Science University hersh@ohsu.edu http://medir.ohsu.edu/~genomics Introductory comments Track history 2003 track Primary task
Optimizing Query results using Middle Layers Based on Concept Hierarchies
 Optimizing Query results using Middle Layers Based on Concept Hierarchies G.Kumari,I.Gayathri Devi G.Kumari Asst.Professor, CSE Department I.Gayathri Devi Asst.Professor, CSE Department Pragati Engineering
Optimizing Query results using Middle Layers Based on Concept Hierarchies G.Kumari,I.Gayathri Devi G.Kumari Asst.Professor, CSE Department I.Gayathri Devi Asst.Professor, CSE Department Pragati Engineering
Finding and Exporting Data. BioMart
 September 2017 Finding and Exporting Data Not sure what tool to use to find and export data? BioMart is used to retrieve data for complex queries, involving a few or many genes or even complete genomes.
September 2017 Finding and Exporting Data Not sure what tool to use to find and export data? BioMart is used to retrieve data for complex queries, involving a few or many genes or even complete genomes.
UPA 2004 Presentation Page 1
 UPA 2004 Presentation Page 1 Thomas S. Tullis and Jacqueline N. Stetson Human Interface Design Department, Fidelity Center for Applied Technology Fidelity Investments 82 Devonshire St., V4A Boston, MA
UPA 2004 Presentation Page 1 Thomas S. Tullis and Jacqueline N. Stetson Human Interface Design Department, Fidelity Center for Applied Technology Fidelity Investments 82 Devonshire St., V4A Boston, MA
WEB SEARCH, FILTERING, AND TEXT MINING: TECHNOLOGY FOR A NEW ERA OF INFORMATION ACCESS
 1 WEB SEARCH, FILTERING, AND TEXT MINING: TECHNOLOGY FOR A NEW ERA OF INFORMATION ACCESS BRUCE CROFT NSF Center for Intelligent Information Retrieval, Computer Science Department, University of Massachusetts,
1 WEB SEARCH, FILTERING, AND TEXT MINING: TECHNOLOGY FOR A NEW ERA OF INFORMATION ACCESS BRUCE CROFT NSF Center for Intelligent Information Retrieval, Computer Science Department, University of Massachusetts,
Using open access literature to guide full-text query formulation. Heather A. Piwowar and Wendy W. Chapman. Background
 Using open access literature to guide full-text query formulation Heather A. Piwowar and Wendy W. Chapman Background Much scientific knowledge is contained in the details of the full-text biomedical literature.
Using open access literature to guide full-text query formulation Heather A. Piwowar and Wendy W. Chapman Background Much scientific knowledge is contained in the details of the full-text biomedical literature.
A Task-Based Evaluation of an Aggregated Search Interface
 A Task-Based Evaluation of an Aggregated Search Interface No Author Given No Institute Given Abstract. This paper presents a user study that evaluated the effectiveness of an aggregated search interface
A Task-Based Evaluation of an Aggregated Search Interface No Author Given No Institute Given Abstract. This paper presents a user study that evaluated the effectiveness of an aggregated search interface
CD 485 Computer Applications in Communication Disorders and Sciences MODULE 3
 CD 485 Computer Applications in Communication Disorders and Sciences MODULE 3 SECTION VII IDENTIFYING THE APPROPRIATE DATABASES JOURNAL ARTICLES THROUGH PUBMED, MEDLINE AND COMMUNICATION DISORDERS MULTISEARCH
CD 485 Computer Applications in Communication Disorders and Sciences MODULE 3 SECTION VII IDENTIFYING THE APPROPRIATE DATABASES JOURNAL ARTICLES THROUGH PUBMED, MEDLINE AND COMMUNICATION DISORDERS MULTISEARCH
Retrieval of Highly Related Documents Containing Gene-Disease Association
 Retrieval of Highly Related Documents Containing Gene-Disease Association K. Santhosh kumar 1, P. Sudhakar 2 Department of Computer Science & Engineering Annamalai University Annamalai Nagar, India. santhosh09539@gmail.com,
Retrieval of Highly Related Documents Containing Gene-Disease Association K. Santhosh kumar 1, P. Sudhakar 2 Department of Computer Science & Engineering Annamalai University Annamalai Nagar, India. santhosh09539@gmail.com,
General Arc of a Search. 1. Define information need, get vocabulary. 2. Choose information source
 General Arc of a Search 1. Define information need, get vocabulary 2. Choose information source 3. Decide on your search strategy (keyword/author, citation analysis, related item search, etc.) 4. Construct
General Arc of a Search 1. Define information need, get vocabulary 2. Choose information source 3. Decide on your search strategy (keyword/author, citation analysis, related item search, etc.) 4. Construct
Core Technology Development Team Meeting
 Core Technology Development Team Meeting To hear the meeting, you must call in Toll-free phone number: 1-866-740-1260 Access Code: 2201876 For international call in numbers, please visit: https://www.readytalk.com/account-administration/international-numbers
Core Technology Development Team Meeting To hear the meeting, you must call in Toll-free phone number: 1-866-740-1260 Access Code: 2201876 For international call in numbers, please visit: https://www.readytalk.com/account-administration/international-numbers
efip online Help Document
 efip online Help Document University of Delaware Computer and Information Sciences & Center for Bioinformatics and Computational Biology Newark, DE, USA December 2013 K K S I K K Table of Contents INTRODUCTION...
efip online Help Document University of Delaware Computer and Information Sciences & Center for Bioinformatics and Computational Biology Newark, DE, USA December 2013 K K S I K K Table of Contents INTRODUCTION...
Outline. Possible solutions. The basic problem. How? How? Relevance Feedback, Query Expansion, and Inputs to Ranking Beyond Similarity
 Outline Relevance Feedback, Query Expansion, and Inputs to Ranking Beyond Similarity Lecture 10 CS 410/510 Information Retrieval on the Internet Query reformulation Sources of relevance for feedback Using
Outline Relevance Feedback, Query Expansion, and Inputs to Ranking Beyond Similarity Lecture 10 CS 410/510 Information Retrieval on the Internet Query reformulation Sources of relevance for feedback Using
Sheffield University and the TREC 2004 Genomics Track: Query Expansion Using Synonymous Terms
 Sheffield University and the TREC 2004 Genomics Track: Query Expansion Using Synonymous Terms Yikun Guo, Henk Harkema, Rob Gaizauskas University of Sheffield, UK {guo, harkema, gaizauskas}@dcs.shef.ac.uk
Sheffield University and the TREC 2004 Genomics Track: Query Expansion Using Synonymous Terms Yikun Guo, Henk Harkema, Rob Gaizauskas University of Sheffield, UK {guo, harkema, gaizauskas}@dcs.shef.ac.uk
Relational Retrieval Using a Combination of Path-Constrained Random Walks
 Relational Retrieval Using a Combination of Path-Constrained Random Walks Ni Lao, William W. Cohen University 2010.9.22 Outline Relational Retrieval Problems Path-constrained random walks The need for
Relational Retrieval Using a Combination of Path-Constrained Random Walks Ni Lao, William W. Cohen University 2010.9.22 Outline Relational Retrieval Problems Path-constrained random walks The need for
Usability Report. Author: Stephen Varnado Version: 1.0 Date: November 24, 2014
 Usability Report Author: Stephen Varnado Version: 1.0 Date: November 24, 2014 2 Table of Contents Executive summary... 3 Introduction... 3 Methodology... 3 Usability test results... 4 Effectiveness ratings
Usability Report Author: Stephen Varnado Version: 1.0 Date: November 24, 2014 2 Table of Contents Executive summary... 3 Introduction... 3 Methodology... 3 Usability test results... 4 Effectiveness ratings
SciMiner User s Manual
 SciMiner User s Manual Copyright 2008 Junguk Hur. All rights reserved. Bioinformatics Program University of Michigan Ann Arbor, MI 48109, USA Email: juhur@umich.edu Homepage: http://jdrf.neurology.med.umich.edu/sciminer/
SciMiner User s Manual Copyright 2008 Junguk Hur. All rights reserved. Bioinformatics Program University of Michigan Ann Arbor, MI 48109, USA Email: juhur@umich.edu Homepage: http://jdrf.neurology.med.umich.edu/sciminer/
Document Retrieval using Predication Similarity
 Document Retrieval using Predication Similarity Kalpa Gunaratna 1 Kno.e.sis Center, Wright State University, Dayton, OH 45435 USA kalpa@knoesis.org Abstract. Document retrieval has been an important research
Document Retrieval using Predication Similarity Kalpa Gunaratna 1 Kno.e.sis Center, Wright State University, Dayton, OH 45435 USA kalpa@knoesis.org Abstract. Document retrieval has been an important research
5 Choosing keywords Initially choosing keywords Frequent and rare keywords Evaluating the competition rates of search
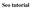 Seo tutorial Seo tutorial Introduction to seo... 4 1. General seo information... 5 1.1 History of search engines... 5 1.2 Common search engine principles... 6 2. Internal ranking factors... 8 2.1 Web page
Seo tutorial Seo tutorial Introduction to seo... 4 1. General seo information... 5 1.1 History of search engines... 5 1.2 Common search engine principles... 6 2. Internal ranking factors... 8 2.1 Web page
CABI Training Materials. Ovid Silver Platter (SP) platform. Simple Searching of CAB Abstracts and Global Health KNOWLEDGE FOR LIFE.
 CABI Training Materials Ovid Silver Platter (SP) platform Simple Searching of CAB Abstracts and Global Health www.cabi.org KNOWLEDGE FOR LIFE Contents The OvidSP Database Selection Screen... 3 The Ovid
CABI Training Materials Ovid Silver Platter (SP) platform Simple Searching of CAB Abstracts and Global Health www.cabi.org KNOWLEDGE FOR LIFE Contents The OvidSP Database Selection Screen... 3 The Ovid
Managing your metadata efficiently - a structured way to organise and frontload your analysis and submission data
 Paper TS06 Managing your metadata efficiently - a structured way to organise and frontload your analysis and submission data Kirsten Walther Langendorf, Novo Nordisk A/S, Copenhagen, Denmark Mikkel Traun,
Paper TS06 Managing your metadata efficiently - a structured way to organise and frontload your analysis and submission data Kirsten Walther Langendorf, Novo Nordisk A/S, Copenhagen, Denmark Mikkel Traun,
PPI Finder: A Mining Tool for Human Protein-Protein Interactions
 PPI Finder: A Mining Tool for Human Protein-Protein Interactions Min He 1,2., Yi Wang 1., Wei Li 1 * 1 Key Laboratory of Molecular and Developmental Biology, Institute of Genetics and Developmental Biology,
PPI Finder: A Mining Tool for Human Protein-Protein Interactions Min He 1,2., Yi Wang 1., Wei Li 1 * 1 Key Laboratory of Molecular and Developmental Biology, Institute of Genetics and Developmental Biology,
Surveys v Contents. User Guide March 11, 2008
 Surveys v8.3.0 User Guide March 11, 2008 Contents What Surveys does Creating survey questions Survey question types Likert Question Importing survey questions from a text file Setting up survey properties
Surveys v8.3.0 User Guide March 11, 2008 Contents What Surveys does Creating survey questions Survey question types Likert Question Importing survey questions from a text file Setting up survey properties
SEEK User Manual. Introduction
 SEEK User Manual Introduction SEEK is a computational gene co-expression search engine. It utilizes a vast human gene expression compendium to deliver fast, integrative, cross-platform co-expression analyses.
SEEK User Manual Introduction SEEK is a computational gene co-expression search engine. It utilizes a vast human gene expression compendium to deliver fast, integrative, cross-platform co-expression analyses.
Headings: Academic Libraries. Database Management. Database Searching. Electronic Information Resource Searching Evaluation. Web Portals.
 Erin R. Holmes. Reimagining the E-Research by Discipline Portal. A Master s Project for the M.S. in IS degree. April, 2014. 20 pages. Advisor: Emily King This project presents recommendations and wireframes
Erin R. Holmes. Reimagining the E-Research by Discipline Portal. A Master s Project for the M.S. in IS degree. April, 2014. 20 pages. Advisor: Emily King This project presents recommendations and wireframes
Improvement of Web Search Results using Genetic Algorithm on Word Sense Disambiguation
 Volume 3, No.5, May 24 International Journal of Advances in Computer Science and Technology Pooja Bassin et al., International Journal of Advances in Computer Science and Technology, 3(5), May 24, 33-336
Volume 3, No.5, May 24 International Journal of Advances in Computer Science and Technology Pooja Bassin et al., International Journal of Advances in Computer Science and Technology, 3(5), May 24, 33-336
Information Retrieval
 Information Retrieval CSC 375, Fall 2016 An information retrieval system will tend not to be used whenever it is more painful and troublesome for a customer to have information than for him not to have
Information Retrieval CSC 375, Fall 2016 An information retrieval system will tend not to be used whenever it is more painful and troublesome for a customer to have information than for him not to have
Text mining tools for semantically enriching the scientific literature
 Text mining tools for semantically enriching the scientific literature Sophia Ananiadou Director National Centre for Text Mining School of Computer Science University of Manchester Need for enriching the
Text mining tools for semantically enriching the scientific literature Sophia Ananiadou Director National Centre for Text Mining School of Computer Science University of Manchester Need for enriching the
Usability Report for Online Writing Portfolio
 Usability Report for Online Writing Portfolio October 30, 2012 WR 305.01 Written By: Kelsey Carper I pledge on my honor that I have not given or received any unauthorized assistance in the completion of
Usability Report for Online Writing Portfolio October 30, 2012 WR 305.01 Written By: Kelsey Carper I pledge on my honor that I have not given or received any unauthorized assistance in the completion of
The basics of searching biomedical databases. Francesca Frati, MLIS. Learning Outcomes. At the end of this workshop you will:
 The basics of searching biomedical databases Francesca Frati, MLIS Learning Outcomes At the end of this workshop you will: Be better able to formulate a clear search question Become more familiar with
The basics of searching biomedical databases Francesca Frati, MLIS Learning Outcomes At the end of this workshop you will: Be better able to formulate a clear search question Become more familiar with
University of Glasgow at CLEF 2013: Experiments in ehealth Task 3 with Terrier
 University of Glasgow at CLEF 2013: Experiments in ehealth Task 3 with Terrier Nut Limsopatham 1, Craig Macdonald 2, and Iadh Ounis 2 School of Computing Science University of Glasgow G12 8QQ, Glasgow,
University of Glasgow at CLEF 2013: Experiments in ehealth Task 3 with Terrier Nut Limsopatham 1, Craig Macdonald 2, and Iadh Ounis 2 School of Computing Science University of Glasgow G12 8QQ, Glasgow,
15/16 CSY2041 Quality and User-Centred Systems
 15/16 CSY2041 Quality and User-Centred Systems INTERACTION DESIGN 1 Heuristic evaluation and walkthroughs 2 1 Aims: Describe the key concepts associated with inspection methods. Explain how to do heuristic
15/16 CSY2041 Quality and User-Centred Systems INTERACTION DESIGN 1 Heuristic evaluation and walkthroughs 2 1 Aims: Describe the key concepts associated with inspection methods. Explain how to do heuristic
Contents. Access Content Library Scroll over the Services Tab and select Content Library.
 Job Aid: Content Library The purpose of this document is to provide step-by-step instructions on how to utilize features and locate items in the Content Library in AgentNet. The Content Library in AgentNet
Job Aid: Content Library The purpose of this document is to provide step-by-step instructions on how to utilize features and locate items in the Content Library in AgentNet. The Content Library in AgentNet
21. Search Models and UIs for IR
 21. Search Models and UIs for IR INFO 202-10 November 2008 Bob Glushko Plan for Today's Lecture The "Classical" Model of Search and the "Classical" UI for IR Web-based Search Best practices for UIs in
21. Search Models and UIs for IR INFO 202-10 November 2008 Bob Glushko Plan for Today's Lecture The "Classical" Model of Search and the "Classical" UI for IR Web-based Search Best practices for UIs in
 Joho, H. and Jose, J.M. (2006) A comparative study of the effectiveness of search result presentation on the web. Lecture Notes in Computer Science 3936:pp. 302-313. http://eprints.gla.ac.uk/3523/ A Comparative
Joho, H. and Jose, J.M. (2006) A comparative study of the effectiveness of search result presentation on the web. Lecture Notes in Computer Science 3936:pp. 302-313. http://eprints.gla.ac.uk/3523/ A Comparative
Enhancing E-Journal Access In A Digital Work Environment
 Enhancing e-journal access in a digital work environment Foo, S. (2006). Singapore Journal of Library & Information Management, 34, 31-40. Enhancing E-Journal Access In A Digital Work Environment Schubert
Enhancing e-journal access in a digital work environment Foo, S. (2006). Singapore Journal of Library & Information Management, 34, 31-40. Enhancing E-Journal Access In A Digital Work Environment Schubert
2) NCBI BLAST tutorial This is a users guide written by the education department at NCBI.
 Web resources -- Tour. page 1 of 8 This is a guided tour. Any homework is separate. In fact, this exercise is used for multiple classes and is publicly available to everyone. The entire tour will take
Web resources -- Tour. page 1 of 8 This is a guided tour. Any homework is separate. In fact, this exercise is used for multiple classes and is publicly available to everyone. The entire tour will take
Bioinformatics Hubs on the Web
 Bioinformatics Hubs on the Web Take a class The Galter Library teaches a related class called Bioinformatics Hubs on the Web. See our Classes schedule for the next available offering. If this class is
Bioinformatics Hubs on the Web Take a class The Galter Library teaches a related class called Bioinformatics Hubs on the Web. See our Classes schedule for the next available offering. If this class is
Getting Started with PsycTESTS on Ovid
 Getting Started with PsycTESTS on Ovid APA DATABASES & ELECTRONIC RESOURCES What is PsycTESTS? What will I find in it? PsycTESTS is a research database that provides information on tests that originated
Getting Started with PsycTESTS on Ovid APA DATABASES & ELECTRONIC RESOURCES What is PsycTESTS? What will I find in it? PsycTESTS is a research database that provides information on tests that originated
INTERNATIONAL JOURNAL OF PURE AND APPLIED RESEARCH IN ENGINEERING AND TECHNOLOGY
 INTERNATIONAL JOURNAL OF PURE AND APPLIED RESEARCH IN ENGINEERING AND TECHNOLOGY A PATH FOR HORIZING YOUR INNOVATIVE WORK REVIEW PAPER ON IMPLEMENTATION OF DOCUMENT ANNOTATION USING CONTENT AND QUERYING
INTERNATIONAL JOURNAL OF PURE AND APPLIED RESEARCH IN ENGINEERING AND TECHNOLOGY A PATH FOR HORIZING YOUR INNOVATIVE WORK REVIEW PAPER ON IMPLEMENTATION OF DOCUMENT ANNOTATION USING CONTENT AND QUERYING
Extracting Algorithms by Indexing and Mining Large Data Sets
 Extracting Algorithms by Indexing and Mining Large Data Sets Vinod Jadhav 1, Dr.Rekha Rathore 2 P.G. Student, Department of Computer Engineering, RKDF SOE Indore, University of RGPV, Bhopal, India Associate
Extracting Algorithms by Indexing and Mining Large Data Sets Vinod Jadhav 1, Dr.Rekha Rathore 2 P.G. Student, Department of Computer Engineering, RKDF SOE Indore, University of RGPV, Bhopal, India Associate
Federated Searching: User Perceptions, System Design, and Library Instruction
 Federated Searching: User Perceptions, System Design, and Library Instruction Rong Tang (Organizer & Presenter) Graduate School of Library and Information Science, Simmons College, 300 The Fenway, Boston,
Federated Searching: User Perceptions, System Design, and Library Instruction Rong Tang (Organizer & Presenter) Graduate School of Library and Information Science, Simmons College, 300 The Fenway, Boston,
Web Searcher Interactions with Multiple Federate Content Collections
 Web Searcher Interactions with Multiple Federate Content Collections Amanda Spink Faculty of IT Queensland University of Technology QLD 4001 Australia ah.spink@qut.edu.au Bernard J. Jansen School of IST
Web Searcher Interactions with Multiple Federate Content Collections Amanda Spink Faculty of IT Queensland University of Technology QLD 4001 Australia ah.spink@qut.edu.au Bernard J. Jansen School of IST
Web Design, 5 th Edition
 Planning a Successful Website: Part 2 Web Design, 5 th Edition Chapter Objectives Discuss the relationship between page length, content placement, and usability Complete Step : Specify the s navigation
Planning a Successful Website: Part 2 Web Design, 5 th Edition Chapter Objectives Discuss the relationship between page length, content placement, and usability Complete Step : Specify the s navigation
Web of Science Including BIOSIS Citation Index
 Web of Science Including BIOSIS Citation Index UCL Library Services, Gower St., London WC1E 6BT 020 7679 7792 E-mail: library@ucl.ac.uk http://www.ucl.ac.uk/library/ 1. What is Web of Science? Web of Science
Web of Science Including BIOSIS Citation Index UCL Library Services, Gower St., London WC1E 6BT 020 7679 7792 E-mail: library@ucl.ac.uk http://www.ucl.ac.uk/library/ 1. What is Web of Science? Web of Science
ithenticate User Guide Getting Started Folders Managing your Documents The Similarity Report Settings Account Information
 ithenticate User Guide Getting Started Folders Managing your Documents The Similarity Report Settings Account Information 1 Getting Started Whether you are a new user or a returning one, to access ithenticate
ithenticate User Guide Getting Started Folders Managing your Documents The Similarity Report Settings Account Information 1 Getting Started Whether you are a new user or a returning one, to access ithenticate
Connecting Library Instruction to Web Usability: The Key Role of Library Instruction to Change Students Web Behavior
 Connecting Library Instruction to Web Usability: The Key Role of Library Instruction to Change Students Web Behavior Yoo Young Lee, Eric Snajdr Indiana University Purdue University, USA yooylee@iupui.edu,
Connecting Library Instruction to Web Usability: The Key Role of Library Instruction to Change Students Web Behavior Yoo Young Lee, Eric Snajdr Indiana University Purdue University, USA yooylee@iupui.edu,
The user interactive task (IAT) in BioCreative Challenges BioCreative Workshop on Text Mining Applications April 7, 2014
 The user interactive task (IAT) in BioCreative Challenges BioCreative Workshop on Text Mining Applications April 7, 2014 N., PhD Research Associate Professor Protein Information Resource CBCB, University
The user interactive task (IAT) in BioCreative Challenges BioCreative Workshop on Text Mining Applications April 7, 2014 N., PhD Research Associate Professor Protein Information Resource CBCB, University
Powering Knowledge Discovery. Insights from big data with Linguamatics I2E
 Powering Knowledge Discovery Insights from big data with Linguamatics I2E Gain actionable insights from unstructured data The world now generates an overwhelming amount of data, most of it written in natural
Powering Knowledge Discovery Insights from big data with Linguamatics I2E Gain actionable insights from unstructured data The world now generates an overwhelming amount of data, most of it written in natural
Data Mining Technologies for Bioinformatics Sequences
 Data Mining Technologies for Bioinformatics Sequences Deepak Garg Computer Science and Engineering Department Thapar Institute of Engineering & Tecnology, Patiala Abstract Main tool used for sequence alignment
Data Mining Technologies for Bioinformatics Sequences Deepak Garg Computer Science and Engineering Department Thapar Institute of Engineering & Tecnology, Patiala Abstract Main tool used for sequence alignment
Evolutionary form design: the application of genetic algorithmic techniques to computer-aided product design
 Loughborough University Institutional Repository Evolutionary form design: the application of genetic algorithmic techniques to computer-aided product design This item was submitted to Loughborough University's
Loughborough University Institutional Repository Evolutionary form design: the application of genetic algorithmic techniques to computer-aided product design This item was submitted to Loughborough University's
A Granular Approach to Web Search Result Presentation
 A Granular Approach to Web Search Result Presentation Ryen W. White 1, Joemon M. Jose 1 & Ian Ruthven 2 1 Department of Computing Science, University of Glasgow, Glasgow. Scotland. 2 Department of Computer
A Granular Approach to Web Search Result Presentation Ryen W. White 1, Joemon M. Jose 1 & Ian Ruthven 2 1 Department of Computing Science, University of Glasgow, Glasgow. Scotland. 2 Department of Computer
EBSCO Discovery Service (EDS): a Research Study for NYU Libraries!!!!!!
 EBSCO Discovery Service (EDS): a Research Study for NYU Libraries Juliana Culbert, Houda El-Mimouni, Nadaleen Tempelman-Kluit NYU Bobst Library, UX Department Spring 2014 Table of Contents Executive Summary
EBSCO Discovery Service (EDS): a Research Study for NYU Libraries Juliana Culbert, Houda El-Mimouni, Nadaleen Tempelman-Kluit NYU Bobst Library, UX Department Spring 2014 Table of Contents Executive Summary
A Tagging Approach to Ontology Mapping
 A Tagging Approach to Ontology Mapping Colm Conroy 1, Declan O'Sullivan 1, Dave Lewis 1 1 Knowledge and Data Engineering Group, Trinity College Dublin {coconroy,declan.osullivan,dave.lewis}@cs.tcd.ie Abstract.
A Tagging Approach to Ontology Mapping Colm Conroy 1, Declan O'Sullivan 1, Dave Lewis 1 1 Knowledge and Data Engineering Group, Trinity College Dublin {coconroy,declan.osullivan,dave.lewis}@cs.tcd.ie Abstract.
Engagement Portal. Physician Engagement User Guide Press Ganey Associates, Inc.
 Engagement Portal Physician Engagement User Guide 2015 Press Ganey Associates, Inc. Contents Logging In... 3 Summary Dashboard... 4 Results For... 5 Filters... 6 Summary Page Engagement Tile... 7 Summary
Engagement Portal Physician Engagement User Guide 2015 Press Ganey Associates, Inc. Contents Logging In... 3 Summary Dashboard... 4 Results For... 5 Filters... 6 Summary Page Engagement Tile... 7 Summary
USER SEARCH INTERFACES. Design and Application
 USER SEARCH INTERFACES Design and Application KEEP IT SIMPLE Search is a means towards some other end, rather than a goal in itself. Search is a mentally intensive task. Task Example: You have a friend
USER SEARCH INTERFACES Design and Application KEEP IT SIMPLE Search is a means towards some other end, rather than a goal in itself. Search is a mentally intensive task. Task Example: You have a friend
Improving Interoperability of Text Mining Tools with BioC
 Improving Interoperability of Text Mining Tools with BioC Ritu Khare, Chih-Hsuan Wei, Yuqing Mao, Robert Leaman, Zhiyong Lu * National Center for Biotechnology Information, 8600 Rockville Pike, Bethesda,
Improving Interoperability of Text Mining Tools with BioC Ritu Khare, Chih-Hsuan Wei, Yuqing Mao, Robert Leaman, Zhiyong Lu * National Center for Biotechnology Information, 8600 Rockville Pike, Bethesda,
WEIGHTING QUERY TERMS USING WORDNET ONTOLOGY
 IJCSNS International Journal of Computer Science and Network Security, VOL.9 No.4, April 2009 349 WEIGHTING QUERY TERMS USING WORDNET ONTOLOGY Mohammed M. Sakre Mohammed M. Kouta Ali M. N. Allam Al Shorouk
IJCSNS International Journal of Computer Science and Network Security, VOL.9 No.4, April 2009 349 WEIGHTING QUERY TERMS USING WORDNET ONTOLOGY Mohammed M. Sakre Mohammed M. Kouta Ali M. N. Allam Al Shorouk
Explora - Basic Search
 Explora - Basic Search From the Explora home screen, you can quickly search for the information you need to complete research and classroom assignments. To create a Basic Search: 1. Enter your search terms
Explora - Basic Search From the Explora home screen, you can quickly search for the information you need to complete research and classroom assignments. To create a Basic Search: 1. Enter your search terms
TREC 2016 Dynamic Domain Track: Exploiting Passage Representation for Retrieval and Relevance Feedback
 RMIT @ TREC 2016 Dynamic Domain Track: Exploiting Passage Representation for Retrieval and Relevance Feedback Ameer Albahem ameer.albahem@rmit.edu.au Lawrence Cavedon lawrence.cavedon@rmit.edu.au Damiano
RMIT @ TREC 2016 Dynamic Domain Track: Exploiting Passage Representation for Retrieval and Relevance Feedback Ameer Albahem ameer.albahem@rmit.edu.au Lawrence Cavedon lawrence.cavedon@rmit.edu.au Damiano
RLIMS-P Website Help Document
 RLIMS-P Website Help Document Table of Contents Introduction... 1 RLIMS-P architecture... 2 RLIMS-P interface... 2 Login...2 Input page...3 Results Page...4 Text Evidence/Curation Page...9 URL: http://annotation.dbi.udel.edu/text_mining/rlimsp2/
RLIMS-P Website Help Document Table of Contents Introduction... 1 RLIMS-P architecture... 2 RLIMS-P interface... 2 Login...2 Input page...3 Results Page...4 Text Evidence/Curation Page...9 URL: http://annotation.dbi.udel.edu/text_mining/rlimsp2/
CACAO Training. Jim Hu and Suzi Aleksander Spring 2016
 CACAO Training Jim Hu and Suzi Aleksander Spring 2016 1 What is CACAO? Community Assessment of Community Annotation with Ontologies (CACAO) Annotation of gene function Competition Within a class Between
CACAO Training Jim Hu and Suzi Aleksander Spring 2016 1 What is CACAO? Community Assessment of Community Annotation with Ontologies (CACAO) Annotation of gene function Competition Within a class Between
User Testing Study: Collaborizm.com. Jessica Espejel, Megan Koontz, Lauren Restivo. LIS 644: Usability Theory and Practice
 User Testing Study: Collaborizm.com Jessica Espejel, Megan Koontz, Lauren Restivo LIS 644: Usability Theory and Practice TABLE OF CONTENTS EXECUTIVE SUMMARY... 3 INTRODUCTION... 4 METHODOLOGY... 5 FINDINGS
User Testing Study: Collaborizm.com Jessica Espejel, Megan Koontz, Lauren Restivo LIS 644: Usability Theory and Practice TABLE OF CONTENTS EXECUTIVE SUMMARY... 3 INTRODUCTION... 4 METHODOLOGY... 5 FINDINGS
Automatic Generation of Query Sessions using Text Segmentation
 Automatic Generation of Query Sessions using Text Segmentation Debasis Ganguly, Johannes Leveling, and Gareth J.F. Jones CNGL, School of Computing, Dublin City University, Dublin-9, Ireland {dganguly,
Automatic Generation of Query Sessions using Text Segmentation Debasis Ganguly, Johannes Leveling, and Gareth J.F. Jones CNGL, School of Computing, Dublin City University, Dublin-9, Ireland {dganguly,
GCE APPLIED ICT OCR UNIT 3. ICT Solutions for Individuals & Society. Student Workbook
 GCE APPLIED ICT OCR UNIT 3 ICT Solutions for Individuals & Society Student Workbook This is a mandatory AS unit. The World Wide Web allows individuals to access information on any topic. Easy access to
GCE APPLIED ICT OCR UNIT 3 ICT Solutions for Individuals & Society Student Workbook This is a mandatory AS unit. The World Wide Web allows individuals to access information on any topic. Easy access to
Incorporating Known Pathways into Gene Clustering Algorithms for Genetic Expression Data
 Incorporating Known Pathways into Gene Clustering Algorithms for Genetic Expression Data Ryan Atallah, John Ryan, David Aeschlimann December 14, 2013 Abstract In this project, we study the problem of classifying
Incorporating Known Pathways into Gene Clustering Algorithms for Genetic Expression Data Ryan Atallah, John Ryan, David Aeschlimann December 14, 2013 Abstract In this project, we study the problem of classifying
Effects of Advanced Search on User Performance and Search Efforts: A Case Study with Three Digital Libraries
 2012 45th Hawaii International Conference on System Sciences Effects of Advanced Search on User Performance and Search Efforts: A Case Study with Three Digital Libraries Xiangmin Zhang Wayne State University
2012 45th Hawaii International Conference on System Sciences Effects of Advanced Search on User Performance and Search Efforts: A Case Study with Three Digital Libraries Xiangmin Zhang Wayne State University
PubMed Assistant: A Biologist-Friendly Interface for Enhanced PubMed Search
 Bioinformatics (2006), accepted. PubMed Assistant: A Biologist-Friendly Interface for Enhanced PubMed Search Jing Ding Department of Electrical and Computer Engineering, Iowa State University, Ames, IA
Bioinformatics (2006), accepted. PubMed Assistant: A Biologist-Friendly Interface for Enhanced PubMed Search Jing Ding Department of Electrical and Computer Engineering, Iowa State University, Ames, IA
CSC9Y4 Programming Language Paradigms Spring 2013
 CSC9Y4 Programming Language Paradigms Spring 2013 Assignment: Programming Languages Report Date Due: 4pm Monday, April 15th 2013 Your report should be printed out, and placed in the box labelled CSC9Y4
CSC9Y4 Programming Language Paradigms Spring 2013 Assignment: Programming Languages Report Date Due: 4pm Monday, April 15th 2013 Your report should be printed out, and placed in the box labelled CSC9Y4
CLEF-IP 2009: Exploring Standard IR Techniques on Patent Retrieval
 DCU @ CLEF-IP 2009: Exploring Standard IR Techniques on Patent Retrieval Walid Magdy, Johannes Leveling, Gareth J.F. Jones Centre for Next Generation Localization School of Computing Dublin City University,
DCU @ CLEF-IP 2009: Exploring Standard IR Techniques on Patent Retrieval Walid Magdy, Johannes Leveling, Gareth J.F. Jones Centre for Next Generation Localization School of Computing Dublin City University,
CHAPTER 4 HUMAN FACTOR BASED USER INTERFACE DESIGN
 CHAPTER 4 HUMAN FACTOR BASED USER INTERFACE DESIGN 4.1 Introduction Today one of the most important concerns is how to use the system with effectiveness, efficiency and satisfaction. The ease or comfort
CHAPTER 4 HUMAN FACTOR BASED USER INTERFACE DESIGN 4.1 Introduction Today one of the most important concerns is how to use the system with effectiveness, efficiency and satisfaction. The ease or comfort
Extracting patient data from tables in clinical literature Case study on extraction of BMI, weight and number of patients
 Extracting patient data from tables in clinical literature Case study on extraction of BMI, weight and number of patients Nikola Milosevic 1, Cassie Gregson 2, Robert Hernandez 2 and Goran Nenadic 1,3
Extracting patient data from tables in clinical literature Case study on extraction of BMI, weight and number of patients Nikola Milosevic 1, Cassie Gregson 2, Robert Hernandez 2 and Goran Nenadic 1,3
Just as computer-based Web search has been a
 C O V E R F E A T U R E Deciphering Trends In Mobile Search Maryam Kamvar and Shumeet Baluja Google Understanding the needs of mobile search will help improve the user experience and increase the service
C O V E R F E A T U R E Deciphering Trends In Mobile Search Maryam Kamvar and Shumeet Baluja Google Understanding the needs of mobile search will help improve the user experience and increase the service
Cascading versus Indexed Menu Design
 February 2003, Vol. 5 Issue 1 Volume 5 Issue 1 Past Issues A-Z List Usability News is a free web newsletter that is produced by the Software Usability Research Laboratory (SURL) at Wichita State University.
February 2003, Vol. 5 Issue 1 Volume 5 Issue 1 Past Issues A-Z List Usability News is a free web newsletter that is produced by the Software Usability Research Laboratory (SURL) at Wichita State University.
Evaluating usability of screen designs with layout complexity
 Southern Cross University epublications@scu Southern Cross Business School 1995 Evaluating usability of screen designs with layout complexity Tim Comber Southern Cross University John R. Maltby Southern
Southern Cross University epublications@scu Southern Cross Business School 1995 Evaluating usability of screen designs with layout complexity Tim Comber Southern Cross University John R. Maltby Southern
Alternative Tools for Mining The Biomedical Literature
 Yale University From the SelectedWorks of Rolando Garcia-Milian May 14, 2014 Alternative Tools for Mining The Biomedical Literature Rolando Garcia-Milian, Yale University Available at: https://works.bepress.com/rolando_garciamilian/1/
Yale University From the SelectedWorks of Rolando Garcia-Milian May 14, 2014 Alternative Tools for Mining The Biomedical Literature Rolando Garcia-Milian, Yale University Available at: https://works.bepress.com/rolando_garciamilian/1/
Student Usability Project Recommendations Define Information Architecture for Library Technology
 Student Usability Project Recommendations Define Information Architecture for Library Technology Erika Rogers, Director, Honors Program, California Polytechnic State University, San Luis Obispo, CA. erogers@calpoly.edu
Student Usability Project Recommendations Define Information Architecture for Library Technology Erika Rogers, Director, Honors Program, California Polytechnic State University, San Luis Obispo, CA. erogers@calpoly.edu
e-scider: A tool to retrieve, prioritize and analyze the articles from PubMed database Sujit R. Tangadpalliwar 1, Rakesh Nimbalkar 2, Prabha Garg* 3
 e-scider: A tool to retrieve, prioritize and analyze the articles from PubMed database Sujit R. Tangadpalliwar 1, Rakesh Nimbalkar 2, Prabha Garg* 3 1 National Institute of Pharmaceutical Education and
e-scider: A tool to retrieve, prioritize and analyze the articles from PubMed database Sujit R. Tangadpalliwar 1, Rakesh Nimbalkar 2, Prabha Garg* 3 1 National Institute of Pharmaceutical Education and
Evaluating University course search
 Evaluating University course search - development of a new user-centred interface concept Johanna Bååth ME1301 Bachelor Thesis in Media Technology, 15 Credits Abstract Findwise is a Europe-wide producer
Evaluating University course search - development of a new user-centred interface concept Johanna Bååth ME1301 Bachelor Thesis in Media Technology, 15 Credits Abstract Findwise is a Europe-wide producer
Exploring the Impact of Information Visualization on Medical Information Seeking Using the Web
 Exploring the Impact of Information Visualization on Medical Information Seeking Using the Web Ketan K. Mane and Javed Mostafa School of Library and Information Science (SLIS) Indiana University, Bloomington,
Exploring the Impact of Information Visualization on Medical Information Seeking Using the Web Ketan K. Mane and Javed Mostafa School of Library and Information Science (SLIS) Indiana University, Bloomington,
