Generation of Hulls Encompassing Neuronal Pathways Based on Tetrahedralization and 3D Alpha Shapes
|
|
|
- Julie Penelope Heath
- 6 years ago
- Views:
Transcription
1 Generation of Hulls Encompassing Neuronal Pathways Based on Tetrahedralization and 3D Alpha Shapes Dorit Merhof 1,2, Martin Meister 1, Ezgi Bingöl 1, Peter Hastreiter 1,2, Christopher Nimsky 2,3, Günther Greiner 1 1 Computer Graphics Group, University of Erlangen-Nuremberg, Germany 2 Neurocenter, Dept. of Neurosurgery, University of Erlangen-Nuremberg, Germany 3 Dept. of Neurosurgery, University of Erlangen-Nuremberg, Germany dorit.merhof@informatik.uni-erlangen.de Abstract. Diffusion tensor imaging provides information about structure and location of white matter tracts within the human brain which is of particular interest for neurosurgery. The reconstruction of neuronal structures from diffusion tensor data is commonly solved by tracking algorithms based on streamline propagation. These approaches generate streamline bundles that approximate the course of neuronal fibers. For medical application, a 3D representation of streamline bundles provides valuable information for pre-operative planning. However, for intraoperative visualization, surfaces wrapping eloquent structures are required for integration into the OR microscope. In order to provide hulls tightly encompassing the neuronal structures obtained from fiber tracking, we propose an approach based on tetrahedralization. This technique reuses the sampling points derived from fiber tracking and therefore provides precise hulls which serve as basis for intra-operative visualization. 1 Introduction In recent years, diffusion tensor imaging (DTI) data has gained increasing interest due to its capability to reflect location and structure of fibrous tissue such as white matter in vivo. For this reason, DTI data is of high value in neurosurgery enhancing the information obtained from standard magnetic resonance imaging (MRI) data. For pre-operative planning as well as intra-operative visualization, tract systems such as the pyramidal tract, the optical tract or the corpus callosum are reconstructed. Commonly accepted techniques for fiber tract reconstruction from DTI data are fiber tracking algorithms. Respective tracking results indicate the location of white matter tracts within the human brain. In the context of neurosurgery, fiber bundles obtained from fiber tracking provide valuable information for diagnosis and therapy planning. However, for intra-operative visualization of fiber tract data, hulls tightly wrapping these structures are required. During surgery, the
2 Fig. 1. Microscope view: (a) tumor, (b) pyramidal tract in close neighborhood 309 boundary curves of the hulls are displayed in the focus plane of the OR microscope and provide a direct relation between tumor tissue and neuronal structures (Fig. 1). A first approach for wrapping fiber tracts [1] computes the centerline of the fiber bundle. In a second step, the center line is sampled equidistantly and planes perpendicular to the center line are considered. For each plane, the intersecting points of all fibers with the plane are computed. Finally, an ellipse encompassing all intersection points is defined for each plane and the ellipses of subsequent planes are connected using a triangular mesh. This approach provides hulls that fit the underlying fiber structure. However, the technique does not take into account branching fibers, requiring a splitting center line or a more sophisticated solution for defining ellipses and connecting them appropriately. In addition to that, the technique is restricted to elongated tract systems, where a centerline is well defined. For fiber tracts such as the corpus callosum encompassing fibers with significantly varying course and direction, the approach will fail. For this reason, we present a novel hull algorithm overcoming these drawbacks. In order to provide precise hulls, the technique takes advantage of the sampling points of the tracked fibers to guarantee high precision. In a first step, a tetrahedral mesh is constructed from the sampling points based on 3D Delaunay tetrahedralization. Since the tetrahedralization process results in the convex hull of the fiber tract, a variation of the 3D alpha shape algorithm has to be applied. As a result, the triangles on the surface of the remaining tetrahedral mesh describe a hull precisely encompassing the fiber tract. 2 Material All datasets used in this work were measured using a Siemens MR Magnetom Sonata Maestro Class 1.5 Tesla scanner. The specifications of the gradient system were a field strength of up to 40 mt/m (effective 69 mt/m) and a slew rate of up to 200 T/m/s (effective 346 T/m/s) for clinical application. DTI datasets were acquired using a field of view of 240 mm resulting in a voxel size of mm 3. For each of the six diffusion weighted datasets (gradient directions (±1,1,0), (±1,0,1) and (0,1,±1)) and the reference dataset,
3 310 Fig. 2. 3D Delaunay tetrahedralization of a corpus callosum: (left) convex hull, (middle) subset with alpha = 10, (right) semi-transparent hull for alpha = 5 displayed with tracked fibers sixty slices with no intersection gap and an acquisition matrix of pixels were measured. 3 Methods In a first step, fiber tracts were computed using a streamline-based tracking approach incorporating trilinear tensor interpolation and fourth order Runge- Kutta integration [2]. Fractional anisotropy was used as termination threshold for fiber propagation. Single tract systems were obtained by incorporating ROIs (regions of interest) defined by a medical expert into the tracking process. As a result, fiber tracts corresponding to specific function such as the pyramidal tract (motor), the optical tract (vision) and the corpus callosum (connection between the two hemispheres) are obtained. The point set comprising the sampling points of all fibers within the fiber tract is then used as input for the tetrahedralization algorithm. For the reconstruction of a tetrahedral mesh based on this point set, a 3D Delaunay [3] approach is applied. For points in general position, i.e. no geometric test is ambiguous, this tetrahedralization is uniquely defined and decomposes the convex hull of the point set into tetrahedra [3]. The tetrahedralization of a point set fulfills the 3D Delaunay criterion, if each sphere defined by the four points of a tetrahedron contains in its interior no other point of the point set. For implementation purposes, the Delaunay3D class of the Visualization ToolKit (VTK) [4] was used (Fig. 2). The output of the 3D Delaunay algorithm is a tetrahedral mesh filling the convex hull of the point set with volume elements. In order to obtain the subset of tetrahedra tightly enclosing the fiber tract, outer tetrahedra have to be removed in an iterative process. For this purpose, a variation of the 3D alpha shape algorithm is applied. The concept of alpha shapes [5] is a generalization of the convex hull, formalizing the intuitive notion of shape for spatial point set data. Depending on the alpha value, which is a real number greater than zero, the alpha shape of an object encompasses only those tetrahedra with smaller or equal
4 Fig. 3. Hulls obtained from tetrahedralization and 3D alpha shapes (alpha = 5): (blue) pyramidal tract, (green) optic tract, (right) combination with the fibers 311 circumsphere than a sphere with diameter alpha. For sufficiently large alpha, the alpha shape is identical to the convex hull which is the original tetrahedral mesh. For decreasing values of alpha, approaching the step size used for fiber tracking, the alpha shape shrinks and gradually reveals the shape of the fiber tract. When applying the alpha shape concept to the tetrahedral meshes obtained from 3D Delaunay tetrahedralization, holes may occur due to removal of inner tetrahedra. For this reason, a variation of the 3D alpha shape algorithm is used, where tetrahedra are only removed according to the alpha criterion, if they are on the surface of the current tetrahedral mesh. This is implemented by setting an alpha-flag for all tetrahedra which should be removed according to the alpha value, and a boundary-flag for all tetrahedra on the surface. In an iterative procedure, surface tetrahedra with valid alpha-flag are removed, and the boundary-flag of their neighbor elements is set. Sweeping through the tetrahedra data structure continues, as long as tetrahedra for removal are found. After applying the 3D alpha shape algorithm, a tetrahedral mesh remains which exactly corresponds to the intuitive shape of the fiber tract. The triangular hull mesh is constructed from the outer faces of the surface tetrahedra. 4 Results The novel technique for precise hull generation was applied to different tract systems, namely the pyramidal tract (motor), the optical tract (vision) and the corpus callosum (connection between the two hemispheres). For all tract systems, the algorithm succeeded to generate precise hulls following the shape of the fiber bundle (Figs. 2, 3). In comparison to the initial approach for wrapping fibers [1], the presented technique provides higher precision which is an essential feature for the intended
5 312 application. This is due to the fact, that points originating from fiber tracking are directly used for tetrahedralization and remain after application of 3D alpha shapes. Additionally, the algorithm is also able to wrap branching fiber tracts or tract systems with diverging fiber directions. With respect to computing times, the algorithm is more time consuming due to the reconstruction of the tetrahedral mesh. For a fiber tract comprising 7443 / / points (pyramidal tract / optical tract / corpus callosum), the 3D Delaunay tetrahedralization requires 2.3 / 8.5 / 80.2 seconds (on a PC equipped with a P4 3.0 GHz and 2 GB RAM). 5 Discussion We presented a novel method for computing hulls encompassing neuronal pathways. As an advantage over existing techniques, the approach is capable to wrap tract systems of arbitrary shape such as branching or winding fiber tracts. In addition to that, the resulting hulls tightly fit the underlying fiber structure since the hull mesh is composed from sampling points derived by fiber tracking. Overall, the presented technique is able to wrap fiber tracts of any shape, and at the same time provides maximum wrapping precision. For medical application, this is of high value in order to obtain a precise visualization denoting the localization of white matter tracts. Acknowledgments This work was supported by the Deutsche Forschungsgemeinschaft in the context of SFB 603, Project C9 and the Graduate Research Center 3D Image Analysis and Synthesis. We thank Frank Enders for contributions to the visualization framework. References 1. Enders F, Sauber N, Merhof D, Hastreiter P, Nimsky C, Stamminger M. Visualization of White Matter Tracts with Wrapped Streamlines. In: Proc. IEEE Visualization; Merhof D, Enders F, Vega F, Hastreiter P, Nimsky C, Stamminger M. Integrated Visualization of Diffusion Tensor Fiber Tracts and Anatomical Data. In: Proc. Simulation and Visualization; Delaunay B. Sur la sphère vide. Bulletin of Academy of Sciences of the USSR (VII) 1934; Schroeder W, Martin K, Lorensen B. The Visualisation ToolKit. Kitware; 2002., URL: 5. Edelsbrunner H, Mücke EP. Three-dimensional alpha shapes. ACM Transactions on Graphics 1994;13(1):43 72.
Fiber Selection from Diffusion Tensor Data based on Boolean Operators
 Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhof 1, G. Greiner 2, M. Buchfelder 3, C. Nimsky 4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhof 1, G. Greiner 2, M. Buchfelder 3, C. Nimsky 4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
Evaluation of Local Filter Approaches for Diffusion Tensor based Fiber Tracking
 Evaluation of Local Filter Approaches for Diffusion Tensor based Fiber Tracking D. Merhof 1, M. Buchfelder 2, C. Nimsky 3 1 Visual Computing, University of Konstanz, Konstanz 2 Department of Neurosurgery,
Evaluation of Local Filter Approaches for Diffusion Tensor based Fiber Tracking D. Merhof 1, M. Buchfelder 2, C. Nimsky 3 1 Visual Computing, University of Konstanz, Konstanz 2 Department of Neurosurgery,
Fiber Selection from Diffusion Tensor Data based on Boolean Operators
 Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhofl, G. Greiner 2, M. Buchfelder 3, C. Nimsky4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhofl, G. Greiner 2, M. Buchfelder 3, C. Nimsky4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
Automatic Quantification of DTI Parameters along Fiber Bundles
 Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein 1, Simon Hermann 1, Olaf Konrad 1, Horst K. Hahn 1, and Heinz-Otto Peitgen 1 1 MeVis Research, 28359 Bremen Email: klein@mevis.de
Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein 1, Simon Hermann 1, Olaf Konrad 1, Horst K. Hahn 1, and Heinz-Otto Peitgen 1 1 MeVis Research, 28359 Bremen Email: klein@mevis.de
Automatic Quantification of DTI Parameters along Fiber Bundles
 Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein, Simon Hermann, Olaf Konrad, Horst K. Hahn, Heinz-Otto Peitgen MeVis Research, 28359 Bremen Email: klein@mevis.de Abstract. We introduce
Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein, Simon Hermann, Olaf Konrad, Horst K. Hahn, Heinz-Otto Peitgen MeVis Research, 28359 Bremen Email: klein@mevis.de Abstract. We introduce
A Ray-based Approach for Boundary Estimation of Fiber Bundles Derived from Diffusion Tensor Imaging
 A Ray-based Approach for Boundary Estimation of Fiber Bundles Derived from Diffusion Tensor Imaging M. H. A. Bauer 1,3, S. Barbieri 2, J. Klein 2, J. Egger 1,3, D. Kuhnt 1, B. Freisleben 3, H.-K. Hahn
A Ray-based Approach for Boundary Estimation of Fiber Bundles Derived from Diffusion Tensor Imaging M. H. A. Bauer 1,3, S. Barbieri 2, J. Klein 2, J. Egger 1,3, D. Kuhnt 1, B. Freisleben 3, H.-K. Hahn
MITK-DI. A new Diffusion Imaging Component for MITK. Klaus Fritzsche, Hans-Peter Meinzer
 MITK-DI A new Diffusion Imaging Component for MITK Klaus Fritzsche, Hans-Peter Meinzer Division of Medical and Biological Informatics, DKFZ Heidelberg k.fritzsche@dkfz-heidelberg.de Abstract. Diffusion-MRI
MITK-DI A new Diffusion Imaging Component for MITK Klaus Fritzsche, Hans-Peter Meinzer Division of Medical and Biological Informatics, DKFZ Heidelberg k.fritzsche@dkfz-heidelberg.de Abstract. Diffusion-MRI
MITK-DI. A new Diffusion Imaging Component for MITK. Klaus Fritzsche, Hans-Peter Meinzer
 MITK-DI A new Diffusion Imaging Component for MITK Klaus Fritzsche, Hans-Peter Meinzer Division of Medical and Biological Informatics, DKFZ Heidelberg k.fritzsche@dkfz-heidelberg.de Abstract. Diffusion-MRI
MITK-DI A new Diffusion Imaging Component for MITK Klaus Fritzsche, Hans-Peter Meinzer Division of Medical and Biological Informatics, DKFZ Heidelberg k.fritzsche@dkfz-heidelberg.de Abstract. Diffusion-MRI
syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets
 syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets Julien Gervais; Lisa Chuah Siemens Healthcare, Magnetic
syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets Julien Gervais; Lisa Chuah Siemens Healthcare, Magnetic
Data Representation in Visualisation
 Data Representation in Visualisation Visualisation Lecture 4 Taku Komura Institute for Perception, Action & Behaviour School of Informatics Taku Komura Data Representation 1 Data Representation We have
Data Representation in Visualisation Visualisation Lecture 4 Taku Komura Institute for Perception, Action & Behaviour School of Informatics Taku Komura Data Representation 1 Data Representation We have
DIFFUSION TENSOR IMAGING ANALYSIS. Using Analyze
 DIFFUSION TENSOR IMAGING ANALYSIS Using Analyze 2 Table of Contents 1. Introduction page 3 2. Loading DTI Data page 4 3. Computing DTI Maps page 5 4. Defining ROIs for Fiber Tracking page 6 5. Visualizing
DIFFUSION TENSOR IMAGING ANALYSIS Using Analyze 2 Table of Contents 1. Introduction page 3 2. Loading DTI Data page 4 3. Computing DTI Maps page 5 4. Defining ROIs for Fiber Tracking page 6 5. Visualizing
HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Visualization of White Matter Tracts with Wrapped Streamlines
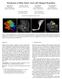 Visualization of White Matter Tracts with Wrapped Streamlines Frank Enders Natascha Sauber Dorit Merhof Peter Hastreiter Neurocenter, Dept. of Neurosurgery Neurocenter, Dept. of Neurosurgery Neurocenter,
Visualization of White Matter Tracts with Wrapped Streamlines Frank Enders Natascha Sauber Dorit Merhof Peter Hastreiter Neurocenter, Dept. of Neurosurgery Neurocenter, Dept. of Neurosurgery Neurocenter,
Contours & Implicit Modelling 4
 Brief Recap Contouring & Implicit Modelling Contouring Implicit Functions Visualisation Lecture 8 lecture 6 Marching Cubes lecture 3 visualisation of a Quadric toby.breckon@ed.ac.uk Computer Vision Lab.
Brief Recap Contouring & Implicit Modelling Contouring Implicit Functions Visualisation Lecture 8 lecture 6 Marching Cubes lecture 3 visualisation of a Quadric toby.breckon@ed.ac.uk Computer Vision Lab.
Diffusion Tensor Imaging and Reading Development
 Diffusion Tensor Imaging and Reading Development Bob Dougherty Stanford Institute for Reading and Learning Reading and Anatomy Every brain is different... Not all brains optimized for highly proficient
Diffusion Tensor Imaging and Reading Development Bob Dougherty Stanford Institute for Reading and Learning Reading and Anatomy Every brain is different... Not all brains optimized for highly proficient
Human Heart Coronary Arteries Segmentation
 Human Heart Coronary Arteries Segmentation Qian Huang Wright State University, Computer Science Department Abstract The volume information extracted from computed tomography angiogram (CTA) datasets makes
Human Heart Coronary Arteries Segmentation Qian Huang Wright State University, Computer Science Department Abstract The volume information extracted from computed tomography angiogram (CTA) datasets makes
Contours & Implicit Modelling 1
 Contouring & Implicit Modelling Visualisation Lecture 8 Institute for Perception, Action & Behaviour School of Informatics Contours & Implicit Modelling 1 Brief Recap Contouring Implicit Functions lecture
Contouring & Implicit Modelling Visualisation Lecture 8 Institute for Perception, Action & Behaviour School of Informatics Contours & Implicit Modelling 1 Brief Recap Contouring Implicit Functions lecture
3D Surface Reconstruction of the Brain based on Level Set Method
 3D Surface Reconstruction of the Brain based on Level Set Method Shijun Tang, Bill P. Buckles, and Kamesh Namuduri Department of Computer Science & Engineering Department of Electrical Engineering University
3D Surface Reconstruction of the Brain based on Level Set Method Shijun Tang, Bill P. Buckles, and Kamesh Namuduri Department of Computer Science & Engineering Department of Electrical Engineering University
Volume Illumination and Segmentation
 Volume Illumination and Segmentation Computer Animation and Visualisation Lecture 13 Institute for Perception, Action & Behaviour School of Informatics Overview Volume illumination Segmentation Volume
Volume Illumination and Segmentation Computer Animation and Visualisation Lecture 13 Institute for Perception, Action & Behaviour School of Informatics Overview Volume illumination Segmentation Volume
Outline of the presentation
 Surface Reconstruction Petra Surynková Charles University in Prague Faculty of Mathematics and Physics petra.surynkova@mff.cuni.cz Outline of the presentation My work up to now Surfaces of Building Practice
Surface Reconstruction Petra Surynková Charles University in Prague Faculty of Mathematics and Physics petra.surynkova@mff.cuni.cz Outline of the presentation My work up to now Surfaces of Building Practice
Correction of susceptibility artifacts in diffusion tensor data using non-linear registration
 Correction of susceptibility artifacts in diffusion tensor data using non-linear registration D. Merhof a,b,, G. Soza c, A. Stadlbauer b, G. Greiner a, C. Nimsky b a Computer Graphics Group, University
Correction of susceptibility artifacts in diffusion tensor data using non-linear registration D. Merhof a,b,, G. Soza c, A. Stadlbauer b, G. Greiner a, C. Nimsky b a Computer Graphics Group, University
n o r d i c B r a i n E x Tutorial DTI Module
 m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial DTI Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial DTI Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
Open Topology: A Toolkit for Brain Isosurface Correction
 Open Topology: A Toolkit for Brain Isosurface Correction Sylvain Jaume 1, Patrice Rondao 2, and Benoît Macq 2 1 National Institute of Research in Computer Science and Control, INRIA, France, sylvain@mit.edu,
Open Topology: A Toolkit for Brain Isosurface Correction Sylvain Jaume 1, Patrice Rondao 2, and Benoît Macq 2 1 National Institute of Research in Computer Science and Control, INRIA, France, sylvain@mit.edu,
Volume visualization. Volume visualization. Volume visualization methods. Sources of volume visualization. Sources of volume visualization
 Volume visualization Volume visualization Volumes are special cases of scalar data: regular 3D grids of scalars, typically interpreted as density values. Each data value is assumed to describe a cubic
Volume visualization Volume visualization Volumes are special cases of scalar data: regular 3D grids of scalars, typically interpreted as density values. Each data value is assumed to describe a cubic
The organization of the human cerebral cortex estimated by intrinsic functional connectivity
 1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
Advanced Visual Medicine: Techniques for Visual Exploration & Analysis
 Advanced Visual Medicine: Techniques for Visual Exploration & Analysis Interactive Visualization of Multimodal Volume Data for Neurosurgical Planning Felix Ritter, MeVis Research Bremen Multimodal Neurosurgical
Advanced Visual Medicine: Techniques for Visual Exploration & Analysis Interactive Visualization of Multimodal Volume Data for Neurosurgical Planning Felix Ritter, MeVis Research Bremen Multimodal Neurosurgical
Whole Body MRI Intensity Standardization
 Whole Body MRI Intensity Standardization Florian Jäger 1, László Nyúl 1, Bernd Frericks 2, Frank Wacker 2 and Joachim Hornegger 1 1 Institute of Pattern Recognition, University of Erlangen, {jaeger,nyul,hornegger}@informatik.uni-erlangen.de
Whole Body MRI Intensity Standardization Florian Jäger 1, László Nyúl 1, Bernd Frericks 2, Frank Wacker 2 and Joachim Hornegger 1 1 Institute of Pattern Recognition, University of Erlangen, {jaeger,nyul,hornegger}@informatik.uni-erlangen.de
NEURO M203 & BIOMED M263 WINTER 2014
 NEURO M203 & BIOMED M263 WINTER 2014 MRI Lab 2: Neuroimaging Connectivity Lab In today s lab we will work with sample diffusion imaging data and the group averaged fmri data collected during your scanning
NEURO M203 & BIOMED M263 WINTER 2014 MRI Lab 2: Neuroimaging Connectivity Lab In today s lab we will work with sample diffusion imaging data and the group averaged fmri data collected during your scanning
Correction of susceptibility artifacts in diffusion tensor data using non-linear registration
 Available online at www.sciencedirect.com Medical Image Analysis 11 (2007) 588 603 www.elsevier.com/locate/media Correction of susceptibility artifacts in diffusion tensor data using non-linear registration
Available online at www.sciencedirect.com Medical Image Analysis 11 (2007) 588 603 www.elsevier.com/locate/media Correction of susceptibility artifacts in diffusion tensor data using non-linear registration
Visualization of cross sectional data for morphogenetic studies
 Visualization of cross sectional data for morphogenetic studies G. Brunnett, M. Vančo, Ch. Haller, S. Washausen, H.-J. Kuhn, W. Knabe Abstract: We report on a visualization system that has been implemented
Visualization of cross sectional data for morphogenetic studies G. Brunnett, M. Vančo, Ch. Haller, S. Washausen, H.-J. Kuhn, W. Knabe Abstract: We report on a visualization system that has been implemented
coding of various parts showing different features, the possibility of rotation or of hiding covering parts of the object's surface to gain an insight
 Three-Dimensional Object Reconstruction from Layered Spatial Data Michael Dangl and Robert Sablatnig Vienna University of Technology, Institute of Computer Aided Automation, Pattern Recognition and Image
Three-Dimensional Object Reconstruction from Layered Spatial Data Michael Dangl and Robert Sablatnig Vienna University of Technology, Institute of Computer Aided Automation, Pattern Recognition and Image
Utilizing Salient Region Features for 3D Multi-Modality Medical Image Registration
 Utilizing Salient Region Features for 3D Multi-Modality Medical Image Registration Dieter Hahn 1, Gabriele Wolz 2, Yiyong Sun 3, Frank Sauer 3, Joachim Hornegger 1, Torsten Kuwert 2 and Chenyang Xu 3 1
Utilizing Salient Region Features for 3D Multi-Modality Medical Image Registration Dieter Hahn 1, Gabriele Wolz 2, Yiyong Sun 3, Frank Sauer 3, Joachim Hornegger 1, Torsten Kuwert 2 and Chenyang Xu 3 1
Complex Fiber Visualization
 Annales Mathematicae et Informaticae 34 (2007) pp. 103 109 http://www.ektf.hu/tanszek/matematika/ami Complex Fiber Visualization Henrietta Tomán a, Róbert Tornai b, Marianna Zichar c a Department of Computer
Annales Mathematicae et Informaticae 34 (2007) pp. 103 109 http://www.ektf.hu/tanszek/matematika/ami Complex Fiber Visualization Henrietta Tomán a, Róbert Tornai b, Marianna Zichar c a Department of Computer
3D Volume Mesh Generation of Human Organs Using Surface Geometries Created from the Visible Human Data Set
 3D Volume Mesh Generation of Human Organs Using Surface Geometries Created from the Visible Human Data Set John M. Sullivan, Jr., Ziji Wu, and Anand Kulkarni Worcester Polytechnic Institute Worcester,
3D Volume Mesh Generation of Human Organs Using Surface Geometries Created from the Visible Human Data Set John M. Sullivan, Jr., Ziji Wu, and Anand Kulkarni Worcester Polytechnic Institute Worcester,
Technische Universiteit Eindhoven Department of Mathematics and Computer Science. Master s Thesis HIERARCHICAL VISUALIZATION USING FIBER CLUSTERING
 Technische Universiteit Eindhoven Department of Mathematics and Computer Science Master s Thesis HIERARCHICAL VISUALIZATION USING FIBER CLUSTERING by Ing. B. Moberts Supervisors: Dr. A. Vilanova Prof.dr.ir.
Technische Universiteit Eindhoven Department of Mathematics and Computer Science Master s Thesis HIERARCHICAL VISUALIZATION USING FIBER CLUSTERING by Ing. B. Moberts Supervisors: Dr. A. Vilanova Prof.dr.ir.
Atelier 2 : Calcul Haute Performance et Sciences du Vivant Forum er juillet, Paris, France
 From Diffusion MR Image Analysis to Whole Brain Connectivity Simulation Jean-Philippe Thiran EPFL Lausanne, Switzerland EPFL - Lausanne HPC in life sciences at EPFL The Blue Brain project: create a biologically
From Diffusion MR Image Analysis to Whole Brain Connectivity Simulation Jean-Philippe Thiran EPFL Lausanne, Switzerland EPFL - Lausanne HPC in life sciences at EPFL The Blue Brain project: create a biologically
A Workflow Optimized Software Platform for Multimodal Neurosurgical Planning and Monitoring
 A Workflow Optimized Software Platform for Multimodal Neurosurgical Planning and Monitoring Eine Workflow Optimierte Software Umgebung für Multimodale Neurochirurgische Planung und Verlaufskontrolle A
A Workflow Optimized Software Platform for Multimodal Neurosurgical Planning and Monitoring Eine Workflow Optimierte Software Umgebung für Multimodale Neurochirurgische Planung und Verlaufskontrolle A
MITK Global Tractography
 MITK Global Tractography Peter F. Neher a, Bram Stieltjes b, Marco Reisert c, Ignaz Reicht a, Hans-Peter Meinzer a, Klaus H. Fritzsche a,b a German Cancer Research Center, Medical and Biological Informatics,
MITK Global Tractography Peter F. Neher a, Bram Stieltjes b, Marco Reisert c, Ignaz Reicht a, Hans-Peter Meinzer a, Klaus H. Fritzsche a,b a German Cancer Research Center, Medical and Biological Informatics,
3-D MRI Brain Scan Classification Using A Point Series Based Representation
 3-D MRI Brain Scan Classification Using A Point Series Based Representation Akadej Udomchaiporn 1, Frans Coenen 1, Marta García-Fiñana 2, and Vanessa Sluming 3 1 Department of Computer Science, University
3-D MRI Brain Scan Classification Using A Point Series Based Representation Akadej Udomchaiporn 1, Frans Coenen 1, Marta García-Fiñana 2, and Vanessa Sluming 3 1 Department of Computer Science, University
NeuroQLab A Software Assistant for Neurosurgical Planning and Quantitative Image Analysis
 NeuroQLab A Software Assistant for Neurosurgical Planning and Quantitative Image Analysis Florian Weiler 1, Jan Rexilius 2, Jan Klein 1, Horst K. Hahn 1 1 Fraunhofer MEVIS, Universitätsallee 29, 28359
NeuroQLab A Software Assistant for Neurosurgical Planning and Quantitative Image Analysis Florian Weiler 1, Jan Rexilius 2, Jan Klein 1, Horst K. Hahn 1 1 Fraunhofer MEVIS, Universitätsallee 29, 28359
Computational Medical Imaging Analysis Chapter 4: Image Visualization
 Computational Medical Imaging Analysis Chapter 4: Image Visualization Jun Zhang Laboratory for Computational Medical Imaging & Data Analysis Department of Computer Science University of Kentucky Lexington,
Computational Medical Imaging Analysis Chapter 4: Image Visualization Jun Zhang Laboratory for Computational Medical Imaging & Data Analysis Department of Computer Science University of Kentucky Lexington,
Saturn User Manual. Rubén Cárdenes. 29th January 2010 Image Processing Laboratory, University of Valladolid. Abstract
 Saturn User Manual Rubén Cárdenes 29th January 2010 Image Processing Laboratory, University of Valladolid Abstract Saturn is a software package for DTI processing and visualization, provided with a graphic
Saturn User Manual Rubén Cárdenes 29th January 2010 Image Processing Laboratory, University of Valladolid Abstract Saturn is a software package for DTI processing and visualization, provided with a graphic
Network connectivity via inference over curvature-regularizing line graphs
 Network connectivity via inference over curvature-regularizing line graphs Asian Conference on Computer Vision Maxwell D. Collins 1,2, Vikas Singh 2,1, Andrew L. Alexander 3 1 Department of Computer Sciences
Network connectivity via inference over curvature-regularizing line graphs Asian Conference on Computer Vision Maxwell D. Collins 1,2, Vikas Singh 2,1, Andrew L. Alexander 3 1 Department of Computer Sciences
A method of visualization of a brain neural pathway by using critical points and target regions
 Modelling in Medicine and Biology VII 309 A method of visualization of a brain neural pathway by using critical points and target regions A. Doi 1, H. Fujimura 2, M. Nagano 1, T. Inoue 3 & A. Ogawa 3 1
Modelling in Medicine and Biology VII 309 A method of visualization of a brain neural pathway by using critical points and target regions A. Doi 1, H. Fujimura 2, M. Nagano 1, T. Inoue 3 & A. Ogawa 3 1
A Study of Medical Image Analysis System
 Indian Journal of Science and Technology, Vol 8(25), DOI: 10.17485/ijst/2015/v8i25/80492, October 2015 ISSN (Print) : 0974-6846 ISSN (Online) : 0974-5645 A Study of Medical Image Analysis System Kim Tae-Eun
Indian Journal of Science and Technology, Vol 8(25), DOI: 10.17485/ijst/2015/v8i25/80492, October 2015 ISSN (Print) : 0974-6846 ISSN (Online) : 0974-5645 A Study of Medical Image Analysis System Kim Tae-Eun
Elastic registration of medical images using finite element meshes
 Elastic registration of medical images using finite element meshes Hartwig Grabowski Institute of Real-Time Computer Systems & Robotics, University of Karlsruhe, D-76128 Karlsruhe, Germany. Email: grabow@ira.uka.de
Elastic registration of medical images using finite element meshes Hartwig Grabowski Institute of Real-Time Computer Systems & Robotics, University of Karlsruhe, D-76128 Karlsruhe, Germany. Email: grabow@ira.uka.de
ISMI: A classification index for High Angular Resolution Diffusion Imaging
 ISMI: A classification index for High Angular Resolution Diffusion Imaging D. Röttger, D. Dudai D. Merhof and S. Müller Institute for Computational Visualistics, University of Koblenz-Landau, Germany Visual
ISMI: A classification index for High Angular Resolution Diffusion Imaging D. Röttger, D. Dudai D. Merhof and S. Müller Institute for Computational Visualistics, University of Koblenz-Landau, Germany Visual
A Constrained Delaunay Triangle Mesh Method for Three-Dimensional Unstructured Boundary Point Cloud
 International Journal of Computer Systems (ISSN: 2394-1065), Volume 03 Issue 02, February, 2016 Available at http://www.ijcsonline.com/ A Constrained Delaunay Triangle Mesh Method for Three-Dimensional
International Journal of Computer Systems (ISSN: 2394-1065), Volume 03 Issue 02, February, 2016 Available at http://www.ijcsonline.com/ A Constrained Delaunay Triangle Mesh Method for Three-Dimensional
TEMPLATE-BASED AUTOMATIC SEGMENTATION OF MASSETER USING PRIOR KNOWLEDGE
 TEMPLATE-BASED AUTOMATIC SEGMENTATION OF MASSETER USING PRIOR KNOWLEDGE H.P. Ng 1,, S.H. Ong 3, P.S. Goh 4, K.W.C. Foong 1, 5, W.L. Nowinski 1 NUS Graduate School for Integrative Sciences and Engineering,
TEMPLATE-BASED AUTOMATIC SEGMENTATION OF MASSETER USING PRIOR KNOWLEDGE H.P. Ng 1,, S.H. Ong 3, P.S. Goh 4, K.W.C. Foong 1, 5, W.L. Nowinski 1 NUS Graduate School for Integrative Sciences and Engineering,
A Method for Registering Diffusion Weighted Magnetic Resonance Images
 A Method for Registering Diffusion Weighted Magnetic Resonance Images Xiaodong Tao and James V. Miller GE Research, Niskayuna, New York, USA Abstract. Diffusion weighted magnetic resonance (DWMR or DW)
A Method for Registering Diffusion Weighted Magnetic Resonance Images Xiaodong Tao and James V. Miller GE Research, Niskayuna, New York, USA Abstract. Diffusion weighted magnetic resonance (DWMR or DW)
Diffusion Imaging Visualization
 Diffusion Imaging Visualization Thomas Schultz URL: http://cg.cs.uni-bonn.de/schultz/ E-Mail: schultz@cs.uni-bonn.de 1 Outline Introduction to Diffusion Imaging Basic techniques Glyph-based Visualization
Diffusion Imaging Visualization Thomas Schultz URL: http://cg.cs.uni-bonn.de/schultz/ E-Mail: schultz@cs.uni-bonn.de 1 Outline Introduction to Diffusion Imaging Basic techniques Glyph-based Visualization
Multimodal Visualization of DTI and fmri Data using Illustrative Methods
 Multimodal Visualization of DTI and fmri Data using Illustrative Methods Silvia Born 1, Werner Jainek 1, Mario Hlawitschka 2, Gerik Scheuermann 3, Christos Trantakis 4, Jürgen Meixensberger 4, Dirk Bartz
Multimodal Visualization of DTI and fmri Data using Illustrative Methods Silvia Born 1, Werner Jainek 1, Mario Hlawitschka 2, Gerik Scheuermann 3, Christos Trantakis 4, Jürgen Meixensberger 4, Dirk Bartz
Course Review. Computer Animation and Visualisation. Taku Komura
 Course Review Computer Animation and Visualisation Taku Komura Characters include Human models Virtual characters Animal models Representation of postures The body has a hierarchical structure Many types
Course Review Computer Animation and Visualisation Taku Komura Characters include Human models Virtual characters Animal models Representation of postures The body has a hierarchical structure Many types
Scientific Visualization Example exam questions with commented answers
 Scientific Visualization Example exam questions with commented answers The theoretical part of this course is evaluated by means of a multiple- choice exam. The questions cover the material mentioned during
Scientific Visualization Example exam questions with commented answers The theoretical part of this course is evaluated by means of a multiple- choice exam. The questions cover the material mentioned during
LAPLACIAN MESH SMOOTHING FOR TETRAHEDRA BASED VOLUME VISUALIZATION 1. INTRODUCTION
 JOURNAL OF MEDICAL INFORMATICS & TECHNOLOGIES Vol.4/2002, ISSN 642-6037 Rafał STĘGIERSKI *, Paweł MIKOŁAJCZAK * volume data,triangle mesh generation, mesh smoothing, marching tetrahedra LAPLACIAN MESH
JOURNAL OF MEDICAL INFORMATICS & TECHNOLOGIES Vol.4/2002, ISSN 642-6037 Rafał STĘGIERSKI *, Paweł MIKOŁAJCZAK * volume data,triangle mesh generation, mesh smoothing, marching tetrahedra LAPLACIAN MESH
A Design Toolbox to Generate Complex Phantoms for the Evaluation of Medical Image Processing Algorithms
 A Design Toolbox to Generate Complex Phantoms for the Evaluation of Medical Image Processing Algorithms Omar Hamo, Georg Nelles, Gudrun Wagenknecht Central Institute for Electronics, Research Center Juelich,
A Design Toolbox to Generate Complex Phantoms for the Evaluation of Medical Image Processing Algorithms Omar Hamo, Georg Nelles, Gudrun Wagenknecht Central Institute for Electronics, Research Center Juelich,
BrainSuite Lab Exercises. presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014
 BrainSuite Lab Exercises presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014 1. Opening and Displaying an MRI Start BrainSuite Drag and drop the T1 image from the native space
BrainSuite Lab Exercises presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014 1. Opening and Displaying an MRI Start BrainSuite Drag and drop the T1 image from the native space
Morphological Image Processing
 Morphological Image Processing Morphology Identification, analysis, and description of the structure of the smallest unit of words Theory and technique for the analysis and processing of geometric structures
Morphological Image Processing Morphology Identification, analysis, and description of the structure of the smallest unit of words Theory and technique for the analysis and processing of geometric structures
Regularization of Bending and Crossing White Matter Fibers in MRI Q-Ball Fields
 Regularization of Bending and Crossing White Matter Fibers in MRI Q-Ball Fields Hans-H. Ehricke 1, Kay-M. Otto 1 and Uwe Klose 2 1 Institute for Applied Computer Science (IACS), Stralsund University and
Regularization of Bending and Crossing White Matter Fibers in MRI Q-Ball Fields Hans-H. Ehricke 1, Kay-M. Otto 1 and Uwe Klose 2 1 Institute for Applied Computer Science (IACS), Stralsund University and
FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING. Francesca Pizzorni Ferrarese
 FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING Francesca Pizzorni Ferrarese Pipeline overview WM and GM Segmentation Registration Data reconstruction Tractography
FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING Francesca Pizzorni Ferrarese Pipeline overview WM and GM Segmentation Registration Data reconstruction Tractography
K-Means Clustering Using Localized Histogram Analysis
 K-Means Clustering Using Localized Histogram Analysis Michael Bryson University of South Carolina, Department of Computer Science Columbia, SC brysonm@cse.sc.edu Abstract. The first step required for many
K-Means Clustering Using Localized Histogram Analysis Michael Bryson University of South Carolina, Department of Computer Science Columbia, SC brysonm@cse.sc.edu Abstract. The first step required for many
Skåne University Hospital Lund, Lund, Sweden 2 Deparment of Numerical Analysis, Centre for Mathematical Sciences, Lund University, Lund, Sweden
 Volume Tracking: A New Method for Visualization of Intracardiac Blood Flow from Three-Dimensional, Time-Resolved, Three-Component Magnetic Resonance Velocity Mapping Appendix: Theory and Numerical Implementation
Volume Tracking: A New Method for Visualization of Intracardiac Blood Flow from Three-Dimensional, Time-Resolved, Three-Component Magnetic Resonance Velocity Mapping Appendix: Theory and Numerical Implementation
Level Set Evolution without Reinitilization
 Level Set Evolution without Reinitilization Outline Parametric active contour (snake) models. Concepts of Level set method and geometric active contours. A level set formulation without reinitialization.
Level Set Evolution without Reinitilization Outline Parametric active contour (snake) models. Concepts of Level set method and geometric active contours. A level set formulation without reinitialization.
Geometric Representations. Stelian Coros
 Geometric Representations Stelian Coros Geometric Representations Languages for describing shape Boundary representations Polygonal meshes Subdivision surfaces Implicit surfaces Volumetric models Parametric
Geometric Representations Stelian Coros Geometric Representations Languages for describing shape Boundary representations Polygonal meshes Subdivision surfaces Implicit surfaces Volumetric models Parametric
Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm
 Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm Lin Chen and Gudrun Wagenknecht Central Institute for Electronics, Research Center Jülich, Jülich, Germany Email: l.chen@fz-juelich.de
Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm Lin Chen and Gudrun Wagenknecht Central Institute for Electronics, Research Center Jülich, Jülich, Germany Email: l.chen@fz-juelich.de
Visualisation : Lecture 1. So what is visualisation? Visualisation
 So what is visualisation? UG4 / M.Sc. Course 2006 toby.breckon@ed.ac.uk Computer Vision Lab. Institute for Perception, Action & Behaviour Introducing 1 Application of interactive 3D computer graphics to
So what is visualisation? UG4 / M.Sc. Course 2006 toby.breckon@ed.ac.uk Computer Vision Lab. Institute for Perception, Action & Behaviour Introducing 1 Application of interactive 3D computer graphics to
Phantom-based evaluation of a semi-automatic segmentation algorithm for cerebral vascular structures in 3D ultrasound angiography (3D USA)
 Phantom-based evaluation of a semi-automatic segmentation algorithm for cerebral vascular structures in 3D ultrasound angiography (3D USA) C. Chalopin¹, K. Krissian², A. Müns 3, F. Arlt 3, J. Meixensberger³,
Phantom-based evaluation of a semi-automatic segmentation algorithm for cerebral vascular structures in 3D ultrasound angiography (3D USA) C. Chalopin¹, K. Krissian², A. Müns 3, F. Arlt 3, J. Meixensberger³,
Non linear Registration of Pre and Intraoperative Volume Data Based On Piecewise Linear Transformations
 Non linear Registration of Pre and Intraoperative Volume Data Based On Piecewise Linear Transformations C. Rezk Salama, P. Hastreiter, G. Greiner, T. Ertl University of Erlangen, Computer Graphics Group
Non linear Registration of Pre and Intraoperative Volume Data Based On Piecewise Linear Transformations C. Rezk Salama, P. Hastreiter, G. Greiner, T. Ertl University of Erlangen, Computer Graphics Group
Learning-based Neuroimage Registration
 Learning-based Neuroimage Registration Leonid Teverovskiy and Yanxi Liu 1 October 2004 CMU-CALD-04-108, CMU-RI-TR-04-59 School of Computer Science Carnegie Mellon University Pittsburgh, PA 15213 Abstract
Learning-based Neuroimage Registration Leonid Teverovskiy and Yanxi Liu 1 October 2004 CMU-CALD-04-108, CMU-RI-TR-04-59 School of Computer Science Carnegie Mellon University Pittsburgh, PA 15213 Abstract
HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Diffusion model fitting and tractography: A primer
 Diffusion model fitting and tractography: A primer Anastasia Yendiki HMS/MGH/MIT Athinoula A. Martinos Center for Biomedical Imaging 03/18/10 Why n how Diffusion model fitting and tractography 0/18 Why
Diffusion model fitting and tractography: A primer Anastasia Yendiki HMS/MGH/MIT Athinoula A. Martinos Center for Biomedical Imaging 03/18/10 Why n how Diffusion model fitting and tractography 0/18 Why
Indirect Volume Rendering
 Indirect Volume Rendering Visualization Torsten Möller Weiskopf/Machiraju/Möller Overview Contour tracing Marching cubes Marching tetrahedra Optimization octree-based range query Weiskopf/Machiraju/Möller
Indirect Volume Rendering Visualization Torsten Möller Weiskopf/Machiraju/Möller Overview Contour tracing Marching cubes Marching tetrahedra Optimization octree-based range query Weiskopf/Machiraju/Möller
Contouring and Isosurfaces. Ronald Peikert SciVis Contouring 2-1
 Contouring and Isosurfaces Ronald Peikert SciVis 2007 - Contouring 2-1 What are contours? Set of points where the scalar field s has a given value c: Examples in 2D: height contours on maps isobars on
Contouring and Isosurfaces Ronald Peikert SciVis 2007 - Contouring 2-1 What are contours? Set of points where the scalar field s has a given value c: Examples in 2D: height contours on maps isobars on
What is visualization? Why is it important?
 What is visualization? Why is it important? What does visualization do? What is the difference between scientific data and information data Visualization Pipeline Visualization Pipeline Overview Data acquisition
What is visualization? Why is it important? What does visualization do? What is the difference between scientific data and information data Visualization Pipeline Visualization Pipeline Overview Data acquisition
An Introduction to Flow Visualization (1) Christoph Garth
 An Introduction to Flow Visualization (1) Christoph Garth cgarth@ucdavis.edu Motivation What will I be talking about? Classical: Physical experiments to understand flow. 2 Motivation What will I be talking
An Introduction to Flow Visualization (1) Christoph Garth cgarth@ucdavis.edu Motivation What will I be talking about? Classical: Physical experiments to understand flow. 2 Motivation What will I be talking
ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION
 ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION Abstract: MIP Project Report Spring 2013 Gaurav Mittal 201232644 This is a detailed report about the course project, which was to implement
ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION Abstract: MIP Project Report Spring 2013 Gaurav Mittal 201232644 This is a detailed report about the course project, which was to implement
Chapter 3 Set Redundancy in Magnetic Resonance Brain Images
 16 Chapter 3 Set Redundancy in Magnetic Resonance Brain Images 3.1 MRI (magnetic resonance imaging) MRI is a technique of measuring physical structure within the human anatomy. Our proposed research focuses
16 Chapter 3 Set Redundancy in Magnetic Resonance Brain Images 3.1 MRI (magnetic resonance imaging) MRI is a technique of measuring physical structure within the human anatomy. Our proposed research focuses
Part I: Theoretical Background and Integration-Based Methods
 Large Vector Field Visualization: Theory and Practice Part I: Theoretical Background and Integration-Based Methods Christoph Garth Overview Foundations Time-Varying Vector Fields Numerical Integration
Large Vector Field Visualization: Theory and Practice Part I: Theoretical Background and Integration-Based Methods Christoph Garth Overview Foundations Time-Varying Vector Fields Numerical Integration
Scalar Algorithms: Contouring
 Scalar Algorithms: Contouring Computer Animation and Visualisation Lecture tkomura@inf.ed.ac.uk Institute for Perception, Action & Behaviour School of Informatics Contouring Scaler Data Last Lecture...
Scalar Algorithms: Contouring Computer Animation and Visualisation Lecture tkomura@inf.ed.ac.uk Institute for Perception, Action & Behaviour School of Informatics Contouring Scaler Data Last Lecture...
NIH Public Access Author Manuscript Proc Soc Photo Opt Instrum Eng. Author manuscript; available in PMC 2014 October 07.
 NIH Public Access Author Manuscript Published in final edited form as: Proc Soc Photo Opt Instrum Eng. 2014 March 21; 9034: 903442. doi:10.1117/12.2042915. MRI Brain Tumor Segmentation and Necrosis Detection
NIH Public Access Author Manuscript Published in final edited form as: Proc Soc Photo Opt Instrum Eng. 2014 March 21; 9034: 903442. doi:10.1117/12.2042915. MRI Brain Tumor Segmentation and Necrosis Detection
CSC Computer Graphics
 // CSC. Computer Graphics Lecture Kasun@dscs.sjp.ac.lk Department of Computer Science University of Sri Jayewardanepura Polygon Filling Scan-Line Polygon Fill Algorithm Span Flood-Fill Algorithm Inside-outside
// CSC. Computer Graphics Lecture Kasun@dscs.sjp.ac.lk Department of Computer Science University of Sri Jayewardanepura Polygon Filling Scan-Line Polygon Fill Algorithm Span Flood-Fill Algorithm Inside-outside
ECE1778 Final Report MRI Visualizer
 ECE1778 Final Report MRI Visualizer David Qixiang Chen Alex Rodionov Word Count: 2408 Introduction We aim to develop a mobile phone/tablet based neurosurgical MRI visualization application with the goal
ECE1778 Final Report MRI Visualizer David Qixiang Chen Alex Rodionov Word Count: 2408 Introduction We aim to develop a mobile phone/tablet based neurosurgical MRI visualization application with the goal
Iterative CT Reconstruction Using Curvelet-Based Regularization
 Iterative CT Reconstruction Using Curvelet-Based Regularization Haibo Wu 1,2, Andreas Maier 1, Joachim Hornegger 1,2 1 Pattern Recognition Lab (LME), Department of Computer Science, 2 Graduate School in
Iterative CT Reconstruction Using Curvelet-Based Regularization Haibo Wu 1,2, Andreas Maier 1, Joachim Hornegger 1,2 1 Pattern Recognition Lab (LME), Department of Computer Science, 2 Graduate School in
Computer Graphics Prof. Sukhendu Das Dept. of Computer Science and Engineering Indian Institute of Technology, Madras Lecture - 24 Solid Modelling
 Computer Graphics Prof. Sukhendu Das Dept. of Computer Science and Engineering Indian Institute of Technology, Madras Lecture - 24 Solid Modelling Welcome to the lectures on computer graphics. We have
Computer Graphics Prof. Sukhendu Das Dept. of Computer Science and Engineering Indian Institute of Technology, Madras Lecture - 24 Solid Modelling Welcome to the lectures on computer graphics. We have
Surface Mesh Generation
 Surface Mesh Generation J.-F. Remacle Université catholique de Louvain September 22, 2011 0 3D Model For the description of the mesh generation process, let us consider the CAD model of a propeller presented
Surface Mesh Generation J.-F. Remacle Université catholique de Louvain September 22, 2011 0 3D Model For the description of the mesh generation process, let us consider the CAD model of a propeller presented
Neural Network-Assisted Fiber Tracking of Synthetic and White Matter DT-MR Images
 Neural Network-Assisted Fiber Tracking of Synthetic and White Matter DT-MR Images L.M. San-José-Revuelta, M. Martín-Fernández and C. Alberola-López Abstract In this paper, a recently developed fiber tracking
Neural Network-Assisted Fiber Tracking of Synthetic and White Matter DT-MR Images L.M. San-José-Revuelta, M. Martín-Fernández and C. Alberola-López Abstract In this paper, a recently developed fiber tracking
3D VISUALIZATION OF SEGMENTED CRUCIATE LIGAMENTS 1. INTRODUCTION
 JOURNAL OF MEDICAL INFORMATICS & TECHNOLOGIES Vol. 10/006, ISSN 164-6037 Paweł BADURA * cruciate ligament, segmentation, fuzzy connectedness,3d visualization 3D VISUALIZATION OF SEGMENTED CRUCIATE LIGAMENTS
JOURNAL OF MEDICAL INFORMATICS & TECHNOLOGIES Vol. 10/006, ISSN 164-6037 Paweł BADURA * cruciate ligament, segmentation, fuzzy connectedness,3d visualization 3D VISUALIZATION OF SEGMENTED CRUCIATE LIGAMENTS
VOLUMETRIC HARMONIC MAP
 COMMUNICATIONS IN INFORMATION AND SYSTEMS c 2004 International Press Vol. 3, No. 3, pp. 191-202, March 2004 004 VOLUMETRIC HARMONIC MAP YALIN WANG, XIANFENG GU, AND SHING-TUNG YAU Abstract. We develop
COMMUNICATIONS IN INFORMATION AND SYSTEMS c 2004 International Press Vol. 3, No. 3, pp. 191-202, March 2004 004 VOLUMETRIC HARMONIC MAP YALIN WANG, XIANFENG GU, AND SHING-TUNG YAU Abstract. We develop
Introduction of a Quantitative Evaluation Method for White Matter Tractography using a HARDI-based Reference
 Introduction of a Quantitative Evaluation Method for White Matter Tractography using a HARDI-based Reference Peter F. Neher 1, Bram Stieltjes 2, Hans-Peter Meinzer 1, and Klaus H. Fritzsche 1,2, 1 German
Introduction of a Quantitative Evaluation Method for White Matter Tractography using a HARDI-based Reference Peter F. Neher 1, Bram Stieltjes 2, Hans-Peter Meinzer 1, and Klaus H. Fritzsche 1,2, 1 German
doi: /
 Yiting Xie ; Anthony P. Reeves; Single 3D cell segmentation from optical CT microscope images. Proc. SPIE 934, Medical Imaging 214: Image Processing, 9343B (March 21, 214); doi:1.1117/12.243852. (214)
Yiting Xie ; Anthony P. Reeves; Single 3D cell segmentation from optical CT microscope images. Proc. SPIE 934, Medical Imaging 214: Image Processing, 9343B (March 21, 214); doi:1.1117/12.243852. (214)
Detection of Unique Point Landmarks in HARDI Images of the Human Brain
 Detection of Unique Point Landmarks in HARDI Images of the Human Brain Henrik Skibbe and Marco Reisert Department of Radiology, University Medical Center Freiburg, Germany {henrik.skibbe, marco.reisert}@uniklinik-freiburg.de
Detection of Unique Point Landmarks in HARDI Images of the Human Brain Henrik Skibbe and Marco Reisert Department of Radiology, University Medical Center Freiburg, Germany {henrik.skibbe, marco.reisert}@uniklinik-freiburg.de
Generation of Triangle Meshes from Time-of-Flight Data for Surface Registration
 Generation of Triangle Meshes from Time-of-Flight Data for Surface Registration Thomas Kilgus, Thiago R. dos Santos, Alexander Seitel, Kwong Yung, Alfred M. Franz, Anja Groch, Ivo Wolf, Hans-Peter Meinzer,
Generation of Triangle Meshes from Time-of-Flight Data for Surface Registration Thomas Kilgus, Thiago R. dos Santos, Alexander Seitel, Kwong Yung, Alfred M. Franz, Anja Groch, Ivo Wolf, Hans-Peter Meinzer,
Isosurface Rendering. CSC 7443: Scientific Information Visualization
 Isosurface Rendering What is Isosurfacing? An isosurface is the 3D surface representing the locations of a constant scalar value within a volume A surface with the same scalar field value Isosurfaces form
Isosurface Rendering What is Isosurfacing? An isosurface is the 3D surface representing the locations of a constant scalar value within a volume A surface with the same scalar field value Isosurfaces form
Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging
 1 CS 9 Final Project Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging Feiyu Chen Department of Electrical Engineering ABSTRACT Subject motion is a significant
1 CS 9 Final Project Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging Feiyu Chen Department of Electrical Engineering ABSTRACT Subject motion is a significant
Visualization. Images are used to aid in understanding of data. Height Fields and Contours Scalar Fields Volume Rendering Vector Fields [chapter 26]
![Visualization. Images are used to aid in understanding of data. Height Fields and Contours Scalar Fields Volume Rendering Vector Fields [chapter 26] Visualization. Images are used to aid in understanding of data. Height Fields and Contours Scalar Fields Volume Rendering Vector Fields [chapter 26]](/thumbs/74/70771954.jpg) Visualization Images are used to aid in understanding of data Height Fields and Contours Scalar Fields Volume Rendering Vector Fields [chapter 26] Tumor SCI, Utah Scientific Visualization Visualize large
Visualization Images are used to aid in understanding of data Height Fields and Contours Scalar Fields Volume Rendering Vector Fields [chapter 26] Tumor SCI, Utah Scientific Visualization Visualize large
N. Hitschfeld. Blanco Encalada 2120, Santiago, CHILE.
 Generalization of modied octrees for geometric modeling N. Hitschfeld Dpto. Ciencias de la Computacion, Univ. de Chile Blanco Encalada 2120, Santiago, CHILE E-mail: nancy@dcc.uchile.cl Abstract. This paper
Generalization of modied octrees for geometric modeling N. Hitschfeld Dpto. Ciencias de la Computacion, Univ. de Chile Blanco Encalada 2120, Santiago, CHILE E-mail: nancy@dcc.uchile.cl Abstract. This paper
Correctness. The Powercrust Algorithm for Surface Reconstruction. Correctness. Correctness. Delaunay Triangulation. Tools - Voronoi Diagram
 Correctness The Powercrust Algorithm for Surface Reconstruction Nina Amenta Sunghee Choi Ravi Kolluri University of Texas at Austin Boundary of a solid Close to original surface Homeomorphic to original
Correctness The Powercrust Algorithm for Surface Reconstruction Nina Amenta Sunghee Choi Ravi Kolluri University of Texas at Austin Boundary of a solid Close to original surface Homeomorphic to original
A Method of Automated Landmark Generation for Automated 3D PDM Construction
 A Method of Automated Landmark Generation for Automated 3D PDM Construction A. D. Brett and C. J. Taylor Department of Medical Biophysics University of Manchester Manchester M13 9PT, Uk adb@sv1.smb.man.ac.uk
A Method of Automated Landmark Generation for Automated 3D PDM Construction A. D. Brett and C. J. Taylor Department of Medical Biophysics University of Manchester Manchester M13 9PT, Uk adb@sv1.smb.man.ac.uk
Lecture overview. Visualisatie BMT. Vector algorithms. Vector algorithms. Time animation. Time animation
 Visualisatie BMT Lecture overview Vector algorithms Tensor algorithms Modeling algorithms Algorithms - 2 Arjan Kok a.j.f.kok@tue.nl 1 2 Vector algorithms Vector 2 or 3 dimensional representation of direction
Visualisatie BMT Lecture overview Vector algorithms Tensor algorithms Modeling algorithms Algorithms - 2 Arjan Kok a.j.f.kok@tue.nl 1 2 Vector algorithms Vector 2 or 3 dimensional representation of direction
Chapter 11 Representation & Description
 Chain Codes Chain codes are used to represent a boundary by a connected sequence of straight-line segments of specified length and direction. The direction of each segment is coded by using a numbering
Chain Codes Chain codes are used to represent a boundary by a connected sequence of straight-line segments of specified length and direction. The direction of each segment is coded by using a numbering
