syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets
|
|
|
- Hugh Russell
- 5 years ago
- Views:
Transcription
1 syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets Julien Gervais; Lisa Chuah Siemens Healthcare, Magnetic Resonance, Erlangen, Germany syngo.mr Neuro 3D enables a streamlined post processing and visualization of functional BOLD and diffusion tensor images. For clinical insights into the use of fmri and DTI in the clinical settings, we would like to refer the interested reader to these case series [1, 2, 3]. 1A Within this article, we have described our preferred workflow and steps for the processing and analysis of BOLD fmri and diffusion tensor imaging (DTI) data. Obviously, it is not necessary to have both BOLD fmri and DTI datasets available for each patient, syngo.mr Neuro 3D can be used even if there is only BOLD fmri or DTI data for a patient. First step: Open the case Assign the complete case to syngo.mr Neuro 3D with a rightmouse click and selection of the option Assign with further selection of MR Neuro 3D (Fig. 1A). Once the case has been assigned, the status of the case changes from Scheduled to Ready (Fig. 1B). Open the case by double clicking on it. For faster loading, select only the relevant series (BOLD MOCO, GLM, Tensor, anatomical data) by multiselection with Ctrl key and left mouse clicks (Fig. 2). Follow this with a right mouse click and selection of the option Read only with further selection of MR Neuro 3D (Fig. 2). syngo.mr Neuro 3D can accept more than one BOLD series with the capability of visualizing activations for up to 4 different 1B 1 (1A) Case assignment to Neuro 3D with a right mouse click. (1B) When the case is ready, the Workflow Status changes from Scheduled to Ready. 52 MAGNETOM Flash (65) 2/2016
2 paradigms simultaneously. However, the corresponding GLM must also be loaded. For tractography analysis, only the TENSOR data is required. The raw diffusion series does not have to be loaded. syngo.mr Neuro 3D can accept data collected in different scan sessions (e.g., the patient anatomical and DTI data were collected in separate scan sessions from the BOLD data). 2 2 Multi-selection of the relevant datasets to be opened as read only for Neuro 3D. Second step: Alignment The BOLD and DTI data should be registered to a reference scan, usually the anatomical scan such as the MPRAGE. Accurate alignment is especially critical for the analysis of data acquired in different scan sessions. The alignment overview displays all available series overlaid on the reference scan. This is the starting point for registration (Fig. 3). A drop down menu in the Control Area on the far left enables the selection of different overlay palettes, such as Edges or Perfusion, to help visual assessment of co-registration accuracy (Fig. 4). In addition, a blending functionality is available. 3 3 The alignment overview with all available series overlaid on the reference scan. 4A 4B 4 Dropdown menu for different overlay palettes in the Control Area on the far left of the screen. These different overlay palettes can aid assessment of co-registration accuracy. The different examples shown are (4A) Advanced Perfusion (default) and (4B) Edges. MAGNETOM Flash (65) 2/
3 5 2 Image alignment can be performed automatically or manually. First, click on the Auto Alignment All button in the Control Area (Fig. 5, indicated by arrow) in order to perform automatic alignment for all series simultaneously. The algorithm works with an iterative approach: Every time you click on the registration button, the registration is further improved until a steady state is reached (Fig. 6). We recommend the individual verification of each and every series. To do this, click on the series in the target images. This opens the single series in a dedicated 3D MPR view with the functional data overlaid on the reference scan. For BOLD acquisitions, the raw series is also displayed together with motion correction graphs (rotation and translation) (Fig. 7). 5 Automatic alignment for all series to the reference series is performed by clicking on the Auto Alignment All button (indicated with arrow). For each segment, scroll through the slices to check the alignment. You can scroll using three alternatives: 6A 6 Improvements to the automatic coregistration with several iterations. (6A) Prior to application of alignment algorithm, (6B) after one iteration, (6C) after three iterations. 1. using the mouse wheel, 2. moving the mouse up and down while pressing the right mouse button, 3. pressing the keyboard arrows up and down. For the BOLD raw series, you can additionally scroll through time using two alternatives: 1. moving the mouse left and right while pressing the right mouse button, 2. pressing the keyboard arrows left and right. 6B 6C 54 MAGNETOM Flash (65) 2/2016
4 7 7 Verification of the alignment of a single series in a dedicated display. 8 8 Manual alignment is activated by pressing the Visual Alignment button which triggers the appearance of a compass guide. To modify the alignment, click on the Visual Alignment button in the Control Area (Fig. 8, indicated by arrow). This opens a compass guide in the MPR segments. For translation: Position the cursor in the desired MPR segment and drag the overlay using the left mouse button. For rotation: Position the cursor over the compass in the desired MPR seg- ment. The compass changes to white and you can drag it clockwise or counter clockwise using the left mouse button. The center of the rotation can be adjusted by dragging the center of the compass. Many users find the Edges palette useful for the verification of the alignment. After you have performed the manual alignment, deactivate the Visual Alignment button to avoid any further accidental modifications. Repeat this step for all the other series: Click in the target images list to select the next series. To go back to the overview display, click on the Alignment Overview button. When all the series are properly registered, processing and visualization can then be performed. Third step: fmri Click on the fmri step to perform visualization of BOLD activation areas. All BOLD paradigms are presented in the function list and the corresponding activation maps are displayed overlaid on the reference scan in a 3D MPR view and as a volume rendered image (Fig. 9). The visualization and processing tools offer the following functionalities: Hide or display dedicated activation maps by clicking the eye icon in the function list. 9 9 Visualization of BOLD activation maps within the fmri step. MAGNETOM Flash (65) 2/
5 10A 10 (10A) Clip plane functionality to better visualize BOLD activations within the VRT segment. (10B) fmri activations as displayed on a gray-scale VRT-like visualization within the VRT segment. 10B Assignment of color palettes to the activation maps. Different modes are available: Individual Mode to assign different color palettes to each activation map. Uniform Mode to assign the same color palette to all activation maps. Highlight Mode to assign a specific color palette to the activation map resulting from a primary task and another color palette to the secondary tasks. Adjustment of statistical threshold levels Optional interpolation and specification of cluster size. This can be found in the function properties menu (bottom right corner of each MPR segment) In the VRT segment: Clip plane for visualization of the BOLD activation specific to a plane (upper left corner). Several planes can be activated simultaneously. Split plane for visualization of BOLD activation in the whole imaging volume up to the specified anatomical plane (upper left corner) Brain mask for skull removal to better visualize brain surface cortical structures Synchronization of MPR and VRT segments: Clicking on the VRT automatically brings the MPR segments to the corresponding reference point. First, examine the activated areas for the different tasks in the MPR segments. If necessary, adjust the statistical threshold values for each paradigm. Highlight the corresponding paradigm with a click in the function list and adjust the threshold located below (slider bar or keyboard input). The colors of each activation map can be individually specified by using the drop down menus of available color palettes. Within the VRT segment, activate the clip plane and split plane functionalities to help 3D visualization (Fig. 10A). A list of important keyboard shortcuts is present in Appendix A. To translate a plane, position the cursor close to the plane, press and hold the right mouse button and move the mouse in the desired direction. To rotate a plane, position the cursor close to the spheres on its border, press and hold the left mouse button and move the mouse in the desired direction. Should a VRT-like gray-scale visualization be preferred (Fig. 10B), please do these following steps: 1. Click on Fused Volume Renderer (lower left corner) to deactivate the VRT. 2. Click on the wrench within Orthogonal MPR (lower left corner) and click on Orthogonal MPR Properties. This will activate a pop-up window. Activate the In-situ Display option. 3. To translate or rotate the different planes within the VRT segment, simply modify the reference lines within the MPR segments. 56 MAGNETOM Flash (65) 2/2016
6 Fourth step: Time Course Evaluation To perform a quality check of the activation maps, click on the Time Course Evaluation step to analyze signal-time curves. Select the desired paradigm in the function list by clicking on the corresponding white box. An eye icon will appear and the corresponding activation map is displayed. Timecourse analysis can be performed for individual voxels or volumes of interest. Select your desired tool by clicking on the appropriate icon fmri voxel or fmri VOI within the control area or in the upper right corner of each MPR segment (Fig. 11A, highlighted with squares). For single voxel analysis, select the fmri voxel tool and click on the appropriate voxels of interest within the MPR segments. Every click will create a voxel of a different color and the corresponding signal-time curve is displayed with the same color in the time-course segment. To modify the position of a voxel, first deactivate the fmri voxel tool and relocate the voxel with drag and drop. For volume analysis, select the fmri VOI tool and draw a volume over an activated area of interest (Fig. 11B). By default, the VOI is restricted to the voxels which are activated above threshold. You can change this behavior with a right click on the VOI and de-selection of shrink to activation option. To delete any voxel or volume of interest, select the corresponding object and choose the delete measurement option with a right mouse click. 11A 11 (11A) Options for tool selection. (11B) Display of activation maps with signal-time curves in the Time Course Evaluation step. 11B Fifth step: Tractography For visualization of diffusion tracts, click on the Tractography step. Tractography data for the complete imaging volume have been automatically generated. They are displayed overlaid on the reference scan in a 3D MPR view and as a volume rendered image (Fig. 12). If you have performed the fmri step as described above, the 12 MAGNETOM Flash (65) 2/
7 tractography data is simply added to the currently available MPRs and VRT. The tracts are generated using a deterministic approach with the FACT algorithm (Fiber Assignment by Continuous Tracking) [4]. Different parameters can be specified, for example seed points and angle threshold (see Appendix B). Specific settings can be saved to be applied for the initial automated whole volume tractography as well as for the more precise generation of refined tracts. Default settings are provided. You can save your own settings, for this, please refer to the user manual. The whole volume tractography may be useful for the immediate visualization of tracts displacement in the presence of tumors (Fig. 13). The whole volume tractography can be interactively refined through different options: volume of interest, volume of interest using activated voxels, planes, or logical combination (AND, OR ) of these (Fig. 14): 1. Tractography VOI: selective visualization of tracts which traverses the VOI. 2. Tractography VOI (Shrink to activation): selective visualization of tracts within the vicinity of fmri activated voxels in the VOI. 3. Tractography Plane: selective visualization of tracts which crosses the set plane. As there are many different approaches for tractography analysis, this article will focus on two use cases Visualization of whole volume tractography within the Tractography step (two clip planes have been used). 15A 15B 15C 15D 15 Visualization of the arcuate fasciculus in the tractography step. (15A) Initial whole brain tractography. (15B) Refined tracts after positioning of the VOI. (15C) Rendered Tracts with defined settings. (15D) Additional rendering with skull stripping. 58 MAGNETOM Flash (65) 2/2016
8 16A 16B 16C 16D 16 Visualization of the connecting pathways for the visual system in the tractography step. (16A) Initial whole brain tractography. (16B) After fmri VOI. (16C) After VOI 2, only tracts going through 1 and 2 are displayed. (16D) Rendered Tracts with defined settings. 1. Visualization of the arcuate fasciculus Click on the Interactive Mode icon (Fig. 14). Visualization of this very important tract can be done by placing a Tractography VOI in the inferior frontal lobe or in the temporal lobe. However, in cases of lesions or tumors potentially displacing important functional language regions, it may be helpful to use fmri activation voxels to guide placement of the VOI. Select the Tractography VOI or Tractography VOI (Shrink to activation) by clicking on the corresponding icon. Draw the VOI in any MPR segment. This creates a VOI labeled VOI_1 (Fig. 15B). The color of the VOI reflects its function: green for a Tractography VOI, purple for a Tractography VOI (Shrink to activation). You can move the VOI by dragging and moving it in the different MPR segments. The VRT is updated in real time. In the VRT segment, use the clip planes and split planes as described in the fmri step. Once you are satisfied with the rendering, hide the seed volumes, if desired, with a right click and further selection of Hide seed points. Save your result by clicking the Create and Save Diffusion Tract Bundle icon in the control area. This generates the finalized tract bundle using the settings highlighted in the drop down menu Settings (Fig. 15C). 2. Visualization of the connecting pathways for the visual system Click on the Interactive Mode icon. First, select the Tractography VOI (Shrink to activation) by clicking on the corresponding icon and draw a VOI encompassing the visual cortex on the axial plane. This creates a purple VOI labeled VOI_1 (Fig. 16B). Manipulate the shape of the VOI in the different planes to capture the visual cortex selectively for the left or right hemisphere. Second, select the Tractography VOI and draw a VOI next to the optic chiasm, usually best identified on the axial plane. This creates a green MAGNETOM Flash (65) 2/
9 17A 17B VOI labeled VOI_2 (Fig. 16C). By default, the logical combination A AND B is selected (Fig. 16C, arrow), and therefore only the tracts going through the VOI_1 and the VOI_2 are displayed. You can change the position and shape of both VOIs in all planes in order to best capture the connecting pathways. Use the clip planes and split planes in the VRT segment to fine-tune the rendering. Save your result by clicking the Create and Save Diffusion Tract Bundle icon in the control area. This generates the finalized tract bundle using the settings highlighted in the drop down menu Settings. Should a VRT-like gray-scale visualization be preferred (Figs. 17A, B), please refer to the end of the fmri section for the necessary steps. 17 Tractography displayed on a VRT-like gray-scale visualization with single seed point (17A) and multiple seed points (17B). Export: To export the processing results, a dedicated Export step is available. Please refer to the user manual. References 1 Benzinger TL, McKinstry III RC, Chen CI, Priatna A. Clinical applications of Diffusion Tensor Imaging. MAGNETOM Flash. 2007;37: Pillai JJ, Zaca D. Case series: Presurgical planning with fmri/dti. MAGNETOM Flash. 2011; 48: Bartsch AJ. Simultaneous Multi-Slice (SMS) Imaging for Pre-Surgical BOLD fmri and Diffusion Tractography: Case Illustrations. MAGNETOM Flash Special Simultaneous Multi-Slice Supplement. 2015; 63: Mori S, van Zijl PCM. Fiber tracking: Principles and strategies A technical review. NMR in Biomedicine. 2002;15: Appendix B2 60 MAGNETOM Flash (65) 2/2016
10 Clip Planes Tract Length Action Keyboard Shortcut larger range = more tracts smaller range = less tracts Left Clip Plane Shift+ L Right Clip Plane Shift + R Head (Dorsal) Clip Plane Shift + H Foot (Ventral) Clip Plane Shift + F Anterior Clip Plane Shift + A Posterior Clip Plane Shift + P Home all Clip Planes (Orientation Reset) Shift + 0 (zero) Show/Hide all Clip Planes Shift + 1 Orthogonal MPRs Action Keyboard Shortcut Action Increase offset of orthogonal MPRs Shift + Decrease offset of orthogonal MPRs Shift + Show/Hide orthogonal MPR frames Shift + 2 Show/Hide orthogonal planes Shift + 3 Others Action Keyboard Shortcut Show/Hide VRT Shift + 4 Show/Hide Split Plane Shift + 5 Toggle between Clip Plane and Split Plane Shift + s Appendix A: Important keyboard shortcuts for syngo.mr Neuro 3D It is defined as the length of the tracts that will be shown as tracts. Seed points per voxel length smaller value = less tracts value 1 = 1 3 = tract, value 2 = 2 3 = 8 tracts, value 3 = 3 3 = 27 tracts... Angle Threshold smaller value = less tracts higher value = more tracts higher value = more tracts Fiber tracking stops at a certain position if the resulting fiber tract would bend by more than Angle Threshold degrees at this position. FA Threshold smaller value = more tracts higher value = less tracts It is defined as the degree of Anisotropy: 0 = low anisotropy, 1 = very strong anisotropy; typical values are between 0.05 and 0.2. Fiber tracking stops at voxel postion with an FA value lower than the given FA threshold. TraceW Threshold smaller value = more tracts higher value = less tracts The trace-weighted image is obtained by taking the average of the individual diffusion weighted images from the DTI scan. This generates an image comparable to orthogonal DWI, yet is derived from the full diffusion sampling. The TraceW threshold value is set to 120 as default based on internal experience. Appendix B1 Contact Siemens Healthcare GmbH HC DI MR CRM AW Lisa Chuah, Ph.D. Global Segment Manager Neurology, Pediatrics and Orthopedics Postbox Erlangen Germany lisa.chuah@siemens.com Contact Siemens Healthcare GmbH HC DI MR CRM AW Julien Gervais Global Marketing Manager syngo.via for MR Postbox Erlangen Germany Julien.Gervais@siemens.com MAGNETOM Flash (65) 2/
Fiber Selection from Diffusion Tensor Data based on Boolean Operators
 Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhof 1, G. Greiner 2, M. Buchfelder 3, C. Nimsky 4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhof 1, G. Greiner 2, M. Buchfelder 3, C. Nimsky 4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
n o r d i c B r a i n E x Tutorial DTI Module
 m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial DTI Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial DTI Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
NEURO M203 & BIOMED M263 WINTER 2014
 NEURO M203 & BIOMED M263 WINTER 2014 MRI Lab 2: Neuroimaging Connectivity Lab In today s lab we will work with sample diffusion imaging data and the group averaged fmri data collected during your scanning
NEURO M203 & BIOMED M263 WINTER 2014 MRI Lab 2: Neuroimaging Connectivity Lab In today s lab we will work with sample diffusion imaging data and the group averaged fmri data collected during your scanning
BrainSuite Lab Exercises. presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014
 BrainSuite Lab Exercises presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014 1. Opening and Displaying an MRI Start BrainSuite Drag and drop the T1 image from the native space
BrainSuite Lab Exercises presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014 1. Opening and Displaying an MRI Start BrainSuite Drag and drop the T1 image from the native space
Tutorial Visualization and Interaction
 m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial Visualization and Interaction Please note that this tutorial is for the latest released nordicbrainex. If you are using
m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial Visualization and Interaction Please note that this tutorial is for the latest released nordicbrainex. If you are using
Saturn User Manual. Rubén Cárdenes. 29th January 2010 Image Processing Laboratory, University of Valladolid. Abstract
 Saturn User Manual Rubén Cárdenes 29th January 2010 Image Processing Laboratory, University of Valladolid Abstract Saturn is a software package for DTI processing and visualization, provided with a graphic
Saturn User Manual Rubén Cárdenes 29th January 2010 Image Processing Laboratory, University of Valladolid Abstract Saturn is a software package for DTI processing and visualization, provided with a graphic
fmri/dti analysis using Dynasuite
 fmri/dti analysis using Dynasuite Contents 1 Logging in 2 Finding patient session 3 Viewing and adjusting images 4 Checking brain segmentation 5 Checking image registration 6 Seeing fmri results 7 Saving
fmri/dti analysis using Dynasuite Contents 1 Logging in 2 Finding patient session 3 Viewing and adjusting images 4 Checking brain segmentation 5 Checking image registration 6 Seeing fmri results 7 Saving
Fiber Selection from Diffusion Tensor Data based on Boolean Operators
 Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhofl, G. Greiner 2, M. Buchfelder 3, C. Nimsky4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhofl, G. Greiner 2, M. Buchfelder 3, C. Nimsky4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
Evaluation of Local Filter Approaches for Diffusion Tensor based Fiber Tracking
 Evaluation of Local Filter Approaches for Diffusion Tensor based Fiber Tracking D. Merhof 1, M. Buchfelder 2, C. Nimsky 3 1 Visual Computing, University of Konstanz, Konstanz 2 Department of Neurosurgery,
Evaluation of Local Filter Approaches for Diffusion Tensor based Fiber Tracking D. Merhof 1, M. Buchfelder 2, C. Nimsky 3 1 Visual Computing, University of Konstanz, Konstanz 2 Department of Neurosurgery,
HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Generation of Hulls Encompassing Neuronal Pathways Based on Tetrahedralization and 3D Alpha Shapes
 Generation of Hulls Encompassing Neuronal Pathways Based on Tetrahedralization and 3D Alpha Shapes Dorit Merhof 1,2, Martin Meister 1, Ezgi Bingöl 1, Peter Hastreiter 1,2, Christopher Nimsky 2,3, Günther
Generation of Hulls Encompassing Neuronal Pathways Based on Tetrahedralization and 3D Alpha Shapes Dorit Merhof 1,2, Martin Meister 1, Ezgi Bingöl 1, Peter Hastreiter 1,2, Christopher Nimsky 2,3, Günther
DIFFUSION TENSOR IMAGING ANALYSIS. Using Analyze
 DIFFUSION TENSOR IMAGING ANALYSIS Using Analyze 2 Table of Contents 1. Introduction page 3 2. Loading DTI Data page 4 3. Computing DTI Maps page 5 4. Defining ROIs for Fiber Tracking page 6 5. Visualizing
DIFFUSION TENSOR IMAGING ANALYSIS Using Analyze 2 Table of Contents 1. Introduction page 3 2. Loading DTI Data page 4 3. Computing DTI Maps page 5 4. Defining ROIs for Fiber Tracking page 6 5. Visualizing
better images mean better results
 better images mean better results A better way for YOU and YOUR patient brought to you by Advanced Neuro analysis with access to studies wherever you need it Advanced Neuro from Invivo Advancements in
better images mean better results A better way for YOU and YOUR patient brought to you by Advanced Neuro analysis with access to studies wherever you need it Advanced Neuro from Invivo Advancements in
NeuroQLab A Software Assistant for Neurosurgical Planning and Quantitative Image Analysis
 NeuroQLab A Software Assistant for Neurosurgical Planning and Quantitative Image Analysis Florian Weiler 1, Jan Rexilius 2, Jan Klein 1, Horst K. Hahn 1 1 Fraunhofer MEVIS, Universitätsallee 29, 28359
NeuroQLab A Software Assistant for Neurosurgical Planning and Quantitative Image Analysis Florian Weiler 1, Jan Rexilius 2, Jan Klein 1, Horst K. Hahn 1 1 Fraunhofer MEVIS, Universitätsallee 29, 28359
n o r d i c B r a i n E x Tutorial DSC Module
 m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial DSC Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial DSC Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
syngo.pet&ct Oncology CT Tutorial
 www.siemens.com/healthcare syngo.pet&ct Oncology CT Tutorial Storyboard VA30 Answers for life. Table of contents 1 What's New in the Latest Upgrade of syngo.pet&ct Oncology? 5 2 Introduction 7 2.1 Prerequisites
www.siemens.com/healthcare syngo.pet&ct Oncology CT Tutorial Storyboard VA30 Answers for life. Table of contents 1 What's New in the Latest Upgrade of syngo.pet&ct Oncology? 5 2 Introduction 7 2.1 Prerequisites
The Advanced Neuro MR Processing & Visualization Solution
 The Advanced Neuro MR Processing & Visualization Solution Optimized Workflow Automated Processing Provides Answers Faster and More Efficiently The DynaSuite Neuro Workstation is a high performance advanced
The Advanced Neuro MR Processing & Visualization Solution Optimized Workflow Automated Processing Provides Answers Faster and More Efficiently The DynaSuite Neuro Workstation is a high performance advanced
Tutorial BOLD Module
 m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial BOLD Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial BOLD Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
A Workflow Optimized Software Platform for Multimodal Neurosurgical Planning and Monitoring
 A Workflow Optimized Software Platform for Multimodal Neurosurgical Planning and Monitoring Eine Workflow Optimierte Software Umgebung für Multimodale Neurochirurgische Planung und Verlaufskontrolle A
A Workflow Optimized Software Platform for Multimodal Neurosurgical Planning and Monitoring Eine Workflow Optimierte Software Umgebung für Multimodale Neurochirurgische Planung und Verlaufskontrolle A
Voxar 3D ColonMetrix. Reference Guide
 Voxar 3D ColonMetrix Reference Guide The software described in this document is furnished under a license, and may be used or copied only according to the terms of such license. Toshiba means, Toshiba
Voxar 3D ColonMetrix Reference Guide The software described in this document is furnished under a license, and may be used or copied only according to the terms of such license. Toshiba means, Toshiba
Diffusion Tensor Imaging and Reading Development
 Diffusion Tensor Imaging and Reading Development Bob Dougherty Stanford Institute for Reading and Learning Reading and Anatomy Every brain is different... Not all brains optimized for highly proficient
Diffusion Tensor Imaging and Reading Development Bob Dougherty Stanford Institute for Reading and Learning Reading and Anatomy Every brain is different... Not all brains optimized for highly proficient
ECE1778 Final Report MRI Visualizer
 ECE1778 Final Report MRI Visualizer David Qixiang Chen Alex Rodionov Word Count: 2408 Introduction We aim to develop a mobile phone/tablet based neurosurgical MRI visualization application with the goal
ECE1778 Final Report MRI Visualizer David Qixiang Chen Alex Rodionov Word Count: 2408 Introduction We aim to develop a mobile phone/tablet based neurosurgical MRI visualization application with the goal
Learn Image Segmentation Basics with Hands-on Introduction to ITK-SNAP. RSNA 2016 Courses RCB22 and RCB54
 Learn Image Segmentation Basics with Hands-on Introduction to ITK-SNAP RSNA 2016 Courses RCB22 and RCB54 RCB22 Mon, Nov 28 10:30-12:00 PM, Room S401CD RCB54 Thu, Dec 1 2:30-4:30 PM, Room S401CD Presenters:
Learn Image Segmentation Basics with Hands-on Introduction to ITK-SNAP RSNA 2016 Courses RCB22 and RCB54 RCB22 Mon, Nov 28 10:30-12:00 PM, Room S401CD RCB54 Thu, Dec 1 2:30-4:30 PM, Room S401CD Presenters:
Image and dataset registration in Dataviewer - Example of expanded clay
 Image and dataset registration in Dataviewer - Example of expanded clay Method note Page 1 of 6 2 Bruker microct method note: Dataviewer registration Introduction Starting from version 1.5.0.0, DataViewer
Image and dataset registration in Dataviewer - Example of expanded clay Method note Page 1 of 6 2 Bruker microct method note: Dataviewer registration Introduction Starting from version 1.5.0.0, DataViewer
This exercise uses one anatomical data set (ANAT1) and two functional data sets (FUNC1 and FUNC2).
 Exploring Brain Anatomy This week s exercises will let you explore the anatomical organization of the brain to learn some of its basic properties, as well as the location of different structures. The human
Exploring Brain Anatomy This week s exercises will let you explore the anatomical organization of the brain to learn some of its basic properties, as well as the location of different structures. The human
ThE ultimate, INTuITIVE Mr INTErFAcE
 ThE ultimate, INTuITIVE Mr INTErFAcE Empowering you to do more The revolutionary Toshiba M-power user interface takes Mr performance and flexibility to levels higher than ever before. M-power is able to
ThE ultimate, INTuITIVE Mr INTErFAcE Empowering you to do more The revolutionary Toshiba M-power user interface takes Mr performance and flexibility to levels higher than ever before. M-power is able to
3DMMVR REFERENCE MANUAL V 0.81
 3DMMVR REFERENCE MANUAL V 0.81 Page 1 of 30 Index: 1.0 System Requirements...5 1.1 System Processor...5 1.2 System RAM...5 1.3 Graphics Card...5 1.4 Operating System...5 2.0 Conventions...6 2.1 Typographic
3DMMVR REFERENCE MANUAL V 0.81 Page 1 of 30 Index: 1.0 System Requirements...5 1.1 System Processor...5 1.2 System RAM...5 1.3 Graphics Card...5 1.4 Operating System...5 2.0 Conventions...6 2.1 Typographic
Software Release Notes for nordicbrainex v2.3.2
 Software Release Notes for nordicbrainex v2.3.2 Revision 1 Date: April 20 th, 2017 Approved by: VP Development NOTE: The most current documentation for released products is available on http://www.nordicneurolab.com.
Software Release Notes for nordicbrainex v2.3.2 Revision 1 Date: April 20 th, 2017 Approved by: VP Development NOTE: The most current documentation for released products is available on http://www.nordicneurolab.com.
IMPAX Volume Viewing 3D Visualization & Segmentation
 Getting started guide IMPAX Volume Viewing 3D Visualization & Segmentation This guide outlines the basic steps to perform and manipulate a 3D reconstruction of volumetric image data using IMPAX Volume
Getting started guide IMPAX Volume Viewing 3D Visualization & Segmentation This guide outlines the basic steps to perform and manipulate a 3D reconstruction of volumetric image data using IMPAX Volume
Visage 7 Clinical Training Basic Features
 Visage 7 Clinical Training Basic Features Contents Overview... 4 Usage... 4 Client Server Architecture... 5 Client Login... 6 Study Browser... 7 Query Section... 8 Study Labels... 10 Query Labeled Studies...
Visage 7 Clinical Training Basic Features Contents Overview... 4 Usage... 4 Client Server Architecture... 5 Client Login... 6 Study Browser... 7 Query Section... 8 Study Labels... 10 Query Labeled Studies...
BrainMask. Quick Start
 BrainMask Quick Start Segmentation of the brain from three-dimensional MR images is a crucial preprocessing step in morphological and volumetric brain studies. BrainMask software implements a fully automatic
BrainMask Quick Start Segmentation of the brain from three-dimensional MR images is a crucial preprocessing step in morphological and volumetric brain studies. BrainMask software implements a fully automatic
Slicer4Minute Tutorial. Sonia Pujol, Ph.D. Surgical Planning Laboratory Harvard Medical School
 Slicer4Minute Tutorial Sonia Pujol, Ph.D. Surgical Planning Laboratory Harvard Medical School 1 Slicer4 minute tutorial This tutorial is a 4-minute introduction to the 3D visualization capabilities of
Slicer4Minute Tutorial Sonia Pujol, Ph.D. Surgical Planning Laboratory Harvard Medical School 1 Slicer4 minute tutorial This tutorial is a 4-minute introduction to the 3D visualization capabilities of
Working with PDF s. To open a recent file on the Start screen, double click on the file name.
 Working with PDF s Acrobat DC Start Screen (Home Tab) When Acrobat opens, the Acrobat Start screen (Home Tab) populates displaying a list of recently opened files. The search feature on the top of the
Working with PDF s Acrobat DC Start Screen (Home Tab) When Acrobat opens, the Acrobat Start screen (Home Tab) populates displaying a list of recently opened files. The search feature on the top of the
VXvue User Manual (For Human Use)
 VXvue User Manual (For Human Use) Page 2 of 90 Revision History Version Date Description 1.0 2012-03-20 Initial Release Page 3 of 90 Contents Safety and Regulatory... 8 Safety Notice... 8 1. Introduction...
VXvue User Manual (For Human Use) Page 2 of 90 Revision History Version Date Description 1.0 2012-03-20 Initial Release Page 3 of 90 Contents Safety and Regulatory... 8 Safety Notice... 8 1. Introduction...
Advanced Visual Medicine: Techniques for Visual Exploration & Analysis
 Advanced Visual Medicine: Techniques for Visual Exploration & Analysis Interactive Visualization of Multimodal Volume Data for Neurosurgical Planning Felix Ritter, MeVis Research Bremen Multimodal Neurosurgical
Advanced Visual Medicine: Techniques for Visual Exploration & Analysis Interactive Visualization of Multimodal Volume Data for Neurosurgical Planning Felix Ritter, MeVis Research Bremen Multimodal Neurosurgical
Ancient Cell Phone Tracing an Object and Drawing with Layers
 Ancient Cell Phone Tracing an Object and Drawing with Layers 1) Open Corel Draw. Create a blank 8.5 x 11 Document. 2) Go to the Import option and browse to the Graphics 1 > Lessons folder 3) Find the Cell
Ancient Cell Phone Tracing an Object and Drawing with Layers 1) Open Corel Draw. Create a blank 8.5 x 11 Document. 2) Go to the Import option and browse to the Graphics 1 > Lessons folder 3) Find the Cell
The organization of the human cerebral cortex estimated by intrinsic functional connectivity
 1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
icatvision Quick Reference
 icatvision Quick Reference Navigating the i-cat Interface This guide shows how to: View reconstructed images Use main features and tools to optimize an image. REMINDER Images are displayed as if you are
icatvision Quick Reference Navigating the i-cat Interface This guide shows how to: View reconstructed images Use main features and tools to optimize an image. REMINDER Images are displayed as if you are
BrainMask. Quick Start
 BrainMask Quick Start Segmentation of the brain from three-dimensional MR images is a crucial pre-processing step in morphological and volumetric brain studies. BrainMask software implements a fully automatic
BrainMask Quick Start Segmentation of the brain from three-dimensional MR images is a crucial pre-processing step in morphological and volumetric brain studies. BrainMask software implements a fully automatic
Slicer3 minute tutorial
 Slicer3 minute tutorial Sonia Pujol, Ph.D. Surgical Planning Laboratory Harvard Medical School -1- Slicer3 minute tutorial This tutorial is a short introduction to the advanced 3D visualization capabilities
Slicer3 minute tutorial Sonia Pujol, Ph.D. Surgical Planning Laboratory Harvard Medical School -1- Slicer3 minute tutorial This tutorial is a short introduction to the advanced 3D visualization capabilities
This is your imaging dashboard. There is a bevy of valuable information to read from here. Let s break it up.
 The Volume Strikes Back Press the (4D) button. This is your imaging dashboard. There is a bevy of valuable information to read from here. Let s break it up. In the Left GUI (Graphic User Interface), you
The Volume Strikes Back Press the (4D) button. This is your imaging dashboard. There is a bevy of valuable information to read from here. Let s break it up. In the Left GUI (Graphic User Interface), you
MITK-DI. A new Diffusion Imaging Component for MITK. Klaus Fritzsche, Hans-Peter Meinzer
 MITK-DI A new Diffusion Imaging Component for MITK Klaus Fritzsche, Hans-Peter Meinzer Division of Medical and Biological Informatics, DKFZ Heidelberg k.fritzsche@dkfz-heidelberg.de Abstract. Diffusion-MRI
MITK-DI A new Diffusion Imaging Component for MITK Klaus Fritzsche, Hans-Peter Meinzer Division of Medical and Biological Informatics, DKFZ Heidelberg k.fritzsche@dkfz-heidelberg.de Abstract. Diffusion-MRI
Software Release Notes for nordicbrainex v1.1.3
 Software Release Notes for nordicbrainex v1.1.3 Revision 1 Date: February 15 th 2013 Approved by: Sigvald Høyheim NOTE: The most current documentation for released products is available on http://www.nordicneurolab.com.
Software Release Notes for nordicbrainex v1.1.3 Revision 1 Date: February 15 th 2013 Approved by: Sigvald Høyheim NOTE: The most current documentation for released products is available on http://www.nordicneurolab.com.
RSNA, /rg
 RSNA, 2015 10.1148/rg.2015140320 Appendix As noted in the main article, DICOM image files cannot be used directly for 3D printing; further steps are necessary to make them readable by 3D printers. The
RSNA, 2015 10.1148/rg.2015140320 Appendix As noted in the main article, DICOM image files cannot be used directly for 3D printing; further steps are necessary to make them readable by 3D printers. The
Fluorescence Tomography Source Reconstruction and Analysis
 TECHNICAL NOTE Pre-clinical in vivo imaging Fluorescence Tomography Source Reconstruction and Analysis Note: This Technical Note is part of a series for Fluorescence Imaging Tomography (FLIT). The user
TECHNICAL NOTE Pre-clinical in vivo imaging Fluorescence Tomography Source Reconstruction and Analysis Note: This Technical Note is part of a series for Fluorescence Imaging Tomography (FLIT). The user
Stroke Quantification Tool (Sonia) Ver User Manual
 Stroke Quantification Tool (Sonia) Ver. 1.0 User Manual English. 12/2016 Rev. 1.0 www.wakeup-stroke.eu 1 Table of Contents 1. Introduction...3 2. Installation...4 3. Data Import...5 4. Registration...7
Stroke Quantification Tool (Sonia) Ver. 1.0 User Manual English. 12/2016 Rev. 1.0 www.wakeup-stroke.eu 1 Table of Contents 1. Introduction...3 2. Installation...4 3. Data Import...5 4. Registration...7
Complex Fiber Visualization
 Annales Mathematicae et Informaticae 34 (2007) pp. 103 109 http://www.ektf.hu/tanszek/matematika/ami Complex Fiber Visualization Henrietta Tomán a, Róbert Tornai b, Marianna Zichar c a Department of Computer
Annales Mathematicae et Informaticae 34 (2007) pp. 103 109 http://www.ektf.hu/tanszek/matematika/ami Complex Fiber Visualization Henrietta Tomán a, Róbert Tornai b, Marianna Zichar c a Department of Computer
BrainVoyager TM. Getting Started Guide. Version 3.0. for BV 21
 BrainVoyager TM Getting Started Guide Version 3.0 for BV 21 Rainer Goebel, Henk Jansma, Caroline Benjamins, Judith Eck, Hester Breman and Armin Heinecke Copyright 2018 Brain Innovation B.V. Contents About
BrainVoyager TM Getting Started Guide Version 3.0 for BV 21 Rainer Goebel, Henk Jansma, Caroline Benjamins, Judith Eck, Hester Breman and Armin Heinecke Copyright 2018 Brain Innovation B.V. Contents About
MITK-DI. A new Diffusion Imaging Component for MITK. Klaus Fritzsche, Hans-Peter Meinzer
 MITK-DI A new Diffusion Imaging Component for MITK Klaus Fritzsche, Hans-Peter Meinzer Division of Medical and Biological Informatics, DKFZ Heidelberg k.fritzsche@dkfz-heidelberg.de Abstract. Diffusion-MRI
MITK-DI A new Diffusion Imaging Component for MITK Klaus Fritzsche, Hans-Peter Meinzer Division of Medical and Biological Informatics, DKFZ Heidelberg k.fritzsche@dkfz-heidelberg.de Abstract. Diffusion-MRI
Automatic Quantification of DTI Parameters along Fiber Bundles
 Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein 1, Simon Hermann 1, Olaf Konrad 1, Horst K. Hahn 1, and Heinz-Otto Peitgen 1 1 MeVis Research, 28359 Bremen Email: klein@mevis.de
Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein 1, Simon Hermann 1, Olaf Konrad 1, Horst K. Hahn 1, and Heinz-Otto Peitgen 1 1 MeVis Research, 28359 Bremen Email: klein@mevis.de
Gamepad Controls. Figure 1: A diagram of an Xbox controller. Figure 2: A screenshot of the BodyViz Controller Panel. BodyViz 3 User Manual 1
 BodyViz User Manual Gamepad Controls The first step in becoming an expert BodyViz user is to get acquainted with the Xbox gamepad, also known as a controller, and the BodyViz Controller Panel. These can
BodyViz User Manual Gamepad Controls The first step in becoming an expert BodyViz user is to get acquainted with the Xbox gamepad, also known as a controller, and the BodyViz Controller Panel. These can
How to Measure Wedge. Purpose. Introduction. Tools Needed
 Purpose Optical Wedge Application (OWA) is an add-on analysis tool for measurement of optical wedges in either transmission or reflection. OWA can measure a single part or many parts simultaneously (e.g.
Purpose Optical Wedge Application (OWA) is an add-on analysis tool for measurement of optical wedges in either transmission or reflection. OWA can measure a single part or many parts simultaneously (e.g.
WAYLAND FREE PUBLIC LIBRARY 3D Design and Printing Tutorial: Create a Keychain
 WAYLAND FREE PUBLIC LIBRARY 3D Design and Printing Tutorial: Create a Keychain Welcome! In this tutorial we will be creating a 3D printed keychain. You will personalize this name tag with text to make
WAYLAND FREE PUBLIC LIBRARY 3D Design and Printing Tutorial: Create a Keychain Welcome! In this tutorial we will be creating a 3D printed keychain. You will personalize this name tag with text to make
SISCOM (Subtraction Ictal SPECT CO-registered to MRI)
 SISCOM (Subtraction Ictal SPECT CO-registered to MRI) Introduction A method for advanced imaging of epilepsy patients has been developed with Analyze at the Mayo Foundation which uses a combination of
SISCOM (Subtraction Ictal SPECT CO-registered to MRI) Introduction A method for advanced imaging of epilepsy patients has been developed with Analyze at the Mayo Foundation which uses a combination of
The Allen Human Brain Atlas offers three types of searches to allow a user to: (1) obtain gene expression data for specific genes (or probes) of
 Microarray Data MICROARRAY DATA Gene Search Boolean Syntax Differential Search Mouse Differential Search Search Results Gene Classification Correlative Search Download Search Results Data Visualization
Microarray Data MICROARRAY DATA Gene Search Boolean Syntax Differential Search Mouse Differential Search Search Results Gene Classification Correlative Search Download Search Results Data Visualization
Automatic Quantification of DTI Parameters along Fiber Bundles
 Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein, Simon Hermann, Olaf Konrad, Horst K. Hahn, Heinz-Otto Peitgen MeVis Research, 28359 Bremen Email: klein@mevis.de Abstract. We introduce
Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein, Simon Hermann, Olaf Konrad, Horst K. Hahn, Heinz-Otto Peitgen MeVis Research, 28359 Bremen Email: klein@mevis.de Abstract. We introduce
It is a good idea to practice View Control tools for 5 minutes at the start of every 3D session, before doing any other work.
 3D View Control Module Overview All the 2D view controls, such as Fit View, Zoom In and Out, Window Area, and Pan, can be used in 3D. As in 2D, elements to the left, right, above, or below can be excluded
3D View Control Module Overview All the 2D view controls, such as Fit View, Zoom In and Out, Window Area, and Pan, can be used in 3D. As in 2D, elements to the left, right, above, or below can be excluded
LIGHTCONVERSE TOOLS Interface Overview
 MANUAL 1 Contents Contents... 1 LIGHTCONVERSE TOOLS Interface Overview... 2 Tool Manager... 3 Mouse... 4 Mouse Control Operation:... 4 3D Space Area... 4 Modes... 5 Balance Calculator in Warehouse Mode...
MANUAL 1 Contents Contents... 1 LIGHTCONVERSE TOOLS Interface Overview... 2 Tool Manager... 3 Mouse... 4 Mouse Control Operation:... 4 3D Space Area... 4 Modes... 5 Balance Calculator in Warehouse Mode...
Correction of susceptibility artifacts in diffusion tensor data using non-linear registration
 Correction of susceptibility artifacts in diffusion tensor data using non-linear registration D. Merhof a,b,, G. Soza c, A. Stadlbauer b, G. Greiner a, C. Nimsky b a Computer Graphics Group, University
Correction of susceptibility artifacts in diffusion tensor data using non-linear registration D. Merhof a,b,, G. Soza c, A. Stadlbauer b, G. Greiner a, C. Nimsky b a Computer Graphics Group, University
Transform Introduction page 96 Spatial Transforms page 97
 Transform Introduction page 96 Spatial Transforms page 97 Pad page 97 Subregion page 101 Resize page 104 Shift page 109 1. Correcting Wraparound Using the Shift Tool page 109 Flip page 116 2. Flipping
Transform Introduction page 96 Spatial Transforms page 97 Pad page 97 Subregion page 101 Resize page 104 Shift page 109 1. Correcting Wraparound Using the Shift Tool page 109 Flip page 116 2. Flipping
CS/NEUR125 Brains, Minds, and Machines. Due: Wednesday, April 5
 CS/NEUR125 Brains, Minds, and Machines Lab 8: Using fmri to Discover Language Areas in the Brain Due: Wednesday, April 5 In this lab, you will analyze fmri data from an experiment that was designed to
CS/NEUR125 Brains, Minds, and Machines Lab 8: Using fmri to Discover Language Areas in the Brain Due: Wednesday, April 5 In this lab, you will analyze fmri data from an experiment that was designed to
FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING. Francesca Pizzorni Ferrarese
 FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING Francesca Pizzorni Ferrarese Pipeline overview WM and GM Segmentation Registration Data reconstruction Tractography
FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING Francesca Pizzorni Ferrarese Pipeline overview WM and GM Segmentation Registration Data reconstruction Tractography
Using TeraRecon intuition Viewer with STAT Quick Reference Guide
 Using TeraRecon intuition Viewer with STAT Quick Reference Guide 1. Launching intuition 2. 3D Assessment of Coronary Arteries for Stenosis 3. Time Volume Analysis (TVA) for Determination of Left Ventricular
Using TeraRecon intuition Viewer with STAT Quick Reference Guide 1. Launching intuition 2. 3D Assessment of Coronary Arteries for Stenosis 3. Time Volume Analysis (TVA) for Determination of Left Ventricular
GE Dynamic 13 C Acquisition
 GE Dynamic 13 C Acquisition In this example you will load a human prostate dynamic MRS data set (single slice, 24 time points) acquired on a GE 3T scanner. You will generate metabolite maps for pyruvate
GE Dynamic 13 C Acquisition In this example you will load a human prostate dynamic MRS data set (single slice, 24 time points) acquired on a GE 3T scanner. You will generate metabolite maps for pyruvate
1 Filter the search by entering search criteria; 2 Enter a range of dates in which to search. 3 You can filter the search by modality type.
 efilm / Managing Studies STUDY MANAGER How to use the study manager The Study Manager can search for four different types of exams: Local Exams: studies stored on your workstation s hard drive. Remote
efilm / Managing Studies STUDY MANAGER How to use the study manager The Study Manager can search for four different types of exams: Local Exams: studies stored on your workstation s hard drive. Remote
Autodesk Fusion 360 Training: The Future of Making Things Attendee Guide
 Autodesk Fusion 360 Training: The Future of Making Things Attendee Guide Abstract After completing this workshop, you will have a basic understanding of editing 3D models using Autodesk Fusion 360 TM to
Autodesk Fusion 360 Training: The Future of Making Things Attendee Guide Abstract After completing this workshop, you will have a basic understanding of editing 3D models using Autodesk Fusion 360 TM to
Rat 2D EPSI Dual Band Variable Flip Angle 13 C Dynamic Spectroscopy
 Rat 2D EPSI Dual Band Variable Flip Angle 13 C Dynamic Spectroscopy In this example you will load a dynamic MRS animal data set acquired on a GE 3T scanner. This data was acquired with an EPSI sequence
Rat 2D EPSI Dual Band Variable Flip Angle 13 C Dynamic Spectroscopy In this example you will load a dynamic MRS animal data set acquired on a GE 3T scanner. This data was acquired with an EPSI sequence
InSyTe FLECT/CT Application Note
 Summary This note is a quick guide for FLECT and CT image co-registration and fusion with VivoQuant (VQ3.0Patch1). Scope This application note is for users who process/analyze images acquired with InSyTe
Summary This note is a quick guide for FLECT and CT image co-registration and fusion with VivoQuant (VQ3.0Patch1). Scope This application note is for users who process/analyze images acquired with InSyTe
SPM99 fmri Data Analysis Workbook
 SPM99 fmri Data Analysis Workbook This file is a description of the steps needed to use SPM99 analyze a fmri data set from a single subject using a simple on/off activation paradigm. There are two parts
SPM99 fmri Data Analysis Workbook This file is a description of the steps needed to use SPM99 analyze a fmri data set from a single subject using a simple on/off activation paradigm. There are two parts
Overview of Adobe Fireworks
 Adobe Fireworks Overview of Adobe Fireworks In this guide, you ll learn how to do the following: Work with the Adobe Fireworks workspace: tools, Document windows, menus, and panels. Customize the workspace.
Adobe Fireworks Overview of Adobe Fireworks In this guide, you ll learn how to do the following: Work with the Adobe Fireworks workspace: tools, Document windows, menus, and panels. Customize the workspace.
Getting Started Guide
 Getting Started Guide Version 2.5 for BVQX 1.9 Rainer Goebel, Henk Jansma and Jochen Seitz Copyright 2007 Brain Innovation B.V. 2 Contents Preface...4 The Objects Tutorial...5 Scanning session information...5
Getting Started Guide Version 2.5 for BVQX 1.9 Rainer Goebel, Henk Jansma and Jochen Seitz Copyright 2007 Brain Innovation B.V. 2 Contents Preface...4 The Objects Tutorial...5 Scanning session information...5
User Guide for Sylvius 4 Online An Interactive Atlas and Visual Glossary of Human Neuroanatomy
 User Guide for Sylvius 4 Online An Interactive Atlas and Visual Glossary of Human Neuroanatomy S. Mark Williams & Leonard E. White Overview About Sylvius 4 Sylvius 4 provides a unique computer-based learning
User Guide for Sylvius 4 Online An Interactive Atlas and Visual Glossary of Human Neuroanatomy S. Mark Williams & Leonard E. White Overview About Sylvius 4 Sylvius 4 provides a unique computer-based learning
IMAGE STUDIO LITE. Tutorial Guide Featuring Image Studio Analysis Software Version 3.1
 IMAGE STUDIO LITE Tutorial Guide Featuring Image Studio Analysis Software Version 3.1 Notice The information contained in this document is subject to change without notice. LI-COR MAKES NO WARRANTY OF
IMAGE STUDIO LITE Tutorial Guide Featuring Image Studio Analysis Software Version 3.1 Notice The information contained in this document is subject to change without notice. LI-COR MAKES NO WARRANTY OF
Diffusion Mapping with FireVoxel Quick Start Guide
 Diffusion Mapping with FireVoxel Quick Start Guide Medical image analysis tool developed by Artem Mikheev and Henry Rusinek Radiology Department, NYU School of Medicine Original version prepared by Jinyu
Diffusion Mapping with FireVoxel Quick Start Guide Medical image analysis tool developed by Artem Mikheev and Henry Rusinek Radiology Department, NYU School of Medicine Original version prepared by Jinyu
MSI 2D Viewer Software Guide
 MSI 2D Viewer Software Guide Page:1 DISCLAIMER We have used reasonable effort to include accurate and up-to-date information in this manual; it does not, however, make any warranties, conditions or representations
MSI 2D Viewer Software Guide Page:1 DISCLAIMER We have used reasonable effort to include accurate and up-to-date information in this manual; it does not, however, make any warranties, conditions or representations
Table of Contents. IntroLab < SPMLabs < Dynevor TWiki
 Table of Contents Lab 1: Introduction to SPM and data checking...1 Goals of this Lab...1 Prerequisites...1 An SPM Installation...1 SPM Defaults...2 L/R Brain Orientation...2 Memory Use for Data Processing...2
Table of Contents Lab 1: Introduction to SPM and data checking...1 Goals of this Lab...1 Prerequisites...1 An SPM Installation...1 SPM Defaults...2 L/R Brain Orientation...2 Memory Use for Data Processing...2
SIVIC GUI Overview. SIVIC GUI Layout Overview
 SIVIC GUI Overview SIVIC GUI Layout Overview At the top of the SIVIC GUI is a row of buttons called the Toolbar. It is a quick interface for loading datasets, controlling how the mouse manipulates the
SIVIC GUI Overview SIVIC GUI Layout Overview At the top of the SIVIC GUI is a row of buttons called the Toolbar. It is a quick interface for loading datasets, controlling how the mouse manipulates the
FEA of Composites Classical Lamination Theory Example 1
 FEA of Composites Classical Lamination Theory Example 1 22.514 Instructor: Professor James Sherwood Author: Dimitri Soteropoulos Revised by Jacob Wardell Problem Description: A four layer [0/90] s graphite-epoxy
FEA of Composites Classical Lamination Theory Example 1 22.514 Instructor: Professor James Sherwood Author: Dimitri Soteropoulos Revised by Jacob Wardell Problem Description: A four layer [0/90] s graphite-epoxy
BrainSpace V2.1: Geometric mapping and diffusion-based software for cross-subject multimodality brain imaging informatics
 Brief Introduction to BrainSpace V2.1 BrainSpace V2.1: Geometric mapping and diffusion-based software for cross-subject multimodality brain imaging informatics Graphics and Imaging Laboratory Wayne State
Brief Introduction to BrainSpace V2.1 BrainSpace V2.1: Geometric mapping and diffusion-based software for cross-subject multimodality brain imaging informatics Graphics and Imaging Laboratory Wayne State
Digital Imaging and Communications in Medicine (DICOM) Supplement 189: Parametrical Blending Presentation State Storage
 5 Digital Imaging and Communications in edicine (DICO) Supplement 189: Parametrical Blending Presentation State Storage 10 15 Prepared by: DICO Standards Committee, Working Group 16 (R sub group Functional
5 Digital Imaging and Communications in edicine (DICO) Supplement 189: Parametrical Blending Presentation State Storage 10 15 Prepared by: DICO Standards Committee, Working Group 16 (R sub group Functional
Diffusion model fitting and tractography: A primer
 Diffusion model fitting and tractography: A primer Anastasia Yendiki HMS/MGH/MIT Athinoula A. Martinos Center for Biomedical Imaging 03/18/10 Why n how Diffusion model fitting and tractography 0/18 Why
Diffusion model fitting and tractography: A primer Anastasia Yendiki HMS/MGH/MIT Athinoula A. Martinos Center for Biomedical Imaging 03/18/10 Why n how Diffusion model fitting and tractography 0/18 Why
Caret Tutorial Contours and Sections April 17, 2007
 Caret Tutorial Contours and Sections April 17, 2007 John Harwell, Donna Dierker, and David C. Van Essen Washington University School of Medicine Department of Anatomy and Neurobiology Saint Louis, Missouri,
Caret Tutorial Contours and Sections April 17, 2007 John Harwell, Donna Dierker, and David C. Van Essen Washington University School of Medicine Department of Anatomy and Neurobiology Saint Louis, Missouri,
Dental System. Basic Course
 Dental System Basic Course This Course Booklet contains notes and workflows to help during your training and can be used later for reference and to help you train others. How to Perform a Workflow for
Dental System Basic Course This Course Booklet contains notes and workflows to help during your training and can be used later for reference and to help you train others. How to Perform a Workflow for
Guide to WB Annotations
 Guide to WB Annotations 04 May 2016 Annotations are a powerful new feature added to Workbench v1.2.0 (Released May 2016) for placing text and symbols within wb_view tabs and windows. They enable generation
Guide to WB Annotations 04 May 2016 Annotations are a powerful new feature added to Workbench v1.2.0 (Released May 2016) for placing text and symbols within wb_view tabs and windows. They enable generation
Quick Reference Guide
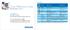 Key functions Quick Reference Guide Philips isite Enterprise Version 3.6 Enhance mouse cursor mode Function Wheel mouse 2-button mouse Adjust WW/WL Left click and drag Left click and drag Scroll/Cine Wheel
Key functions Quick Reference Guide Philips isite Enterprise Version 3.6 Enhance mouse cursor mode Function Wheel mouse 2-button mouse Adjust WW/WL Left click and drag Left click and drag Scroll/Cine Wheel
SPM Introduction. SPM : Overview. SPM: Preprocessing SPM! SPM: Preprocessing. Scott Peltier. FMRI Laboratory University of Michigan
 SPM Introduction Scott Peltier FMRI Laboratory University of Michigan! Slides adapted from T. Nichols SPM! SPM : Overview Library of MATLAB and C functions Graphical user interface Four main components:
SPM Introduction Scott Peltier FMRI Laboratory University of Michigan! Slides adapted from T. Nichols SPM! SPM : Overview Library of MATLAB and C functions Graphical user interface Four main components:
Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques
 Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques Sea Chen Department of Biomedical Engineering Advisors: Dr. Charles A. Bouman and Dr. Mark J. Lowe S. Chen Final Exam October
Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques Sea Chen Department of Biomedical Engineering Advisors: Dr. Charles A. Bouman and Dr. Mark J. Lowe S. Chen Final Exam October
Tekla Structures Analysis Guide. Product version 21.0 March Tekla Corporation
 Tekla Structures Analysis Guide Product version 21.0 March 2015 2015 Tekla Corporation Contents 1 Getting started with analysis... 7 1.1 What is an analysis model... 7 Analysis model objects...9 1.2 About
Tekla Structures Analysis Guide Product version 21.0 March 2015 2015 Tekla Corporation Contents 1 Getting started with analysis... 7 1.1 What is an analysis model... 7 Analysis model objects...9 1.2 About
RadiAnt DICOM Viewer. User manual. Version July 1, Copyright Medixant. All rights reserved.
 RadiAnt DICOM Viewer User manual Version 1.8.6 July 1, 2013 http://www.radiantviewer.com Copyright 2009-2013 Medixant. All rights reserved. Table of contents 3 Table of contents 1 Welcome to RadiAnt DICOM
RadiAnt DICOM Viewer User manual Version 1.8.6 July 1, 2013 http://www.radiantviewer.com Copyright 2009-2013 Medixant. All rights reserved. Table of contents 3 Table of contents 1 Welcome to RadiAnt DICOM
Warping & Blending AP
 Warping & Blending AP Operation about AP This AP provides three major functions including Warp, Edge Blending and Black Level. If the AP is already installed, please remove previous version before installing
Warping & Blending AP Operation about AP This AP provides three major functions including Warp, Edge Blending and Black Level. If the AP is already installed, please remove previous version before installing
SPM Introduction SPM! Scott Peltier. FMRI Laboratory University of Michigan. Software to perform computation, manipulation and display of imaging data
 SPM Introduction Scott Peltier FMRI Laboratory University of Michigan Slides adapted from T. Nichols SPM! Software to perform computation, manipulation and display of imaging data 1 1 SPM : Overview Library
SPM Introduction Scott Peltier FMRI Laboratory University of Michigan Slides adapted from T. Nichols SPM! Software to perform computation, manipulation and display of imaging data 1 1 SPM : Overview Library
ClinicalConnect TM eunity TM Training Guide
 ClinicalConnect TM eunity TM Training Guide October, 2013 Launch eunity TM from ClinicalConnect TM Search and select the patient whose record you wish to view. Navigate to the Radiology module in ClinicalConnect
ClinicalConnect TM eunity TM Training Guide October, 2013 Launch eunity TM from ClinicalConnect TM Search and select the patient whose record you wish to view. Navigate to the Radiology module in ClinicalConnect
This group is dedicated to Modeler tools for Layout s FiberFX hair and fur system. For the Layout interface and controls see FiberFX
 Fiber FX Click here to expand Table of Contents... FiberFX Strand Modeler Global Controls Fiber Tab Guides Tab Random Tab Gravity Tab Tools1 Tab Tools2 Tab Options Tab Strand Tool Strand Maker This group
Fiber FX Click here to expand Table of Contents... FiberFX Strand Modeler Global Controls Fiber Tab Guides Tab Random Tab Gravity Tab Tools1 Tab Tools2 Tab Options Tab Strand Tool Strand Maker This group
Vizit Pro User Manual
 Vizit Pro User Manual 1 Table of Contents Vizit Pro User Manual... 1 Using Vizit Pro... 3 The Vizit Pro User Interface... 3 Toolbars... 4 File Tab Toolbar... 4 Edit Tab Toolbar... 5 Annotations Tab Toolbar...
Vizit Pro User Manual 1 Table of Contents Vizit Pro User Manual... 1 Using Vizit Pro... 3 The Vizit Pro User Interface... 3 Toolbars... 4 File Tab Toolbar... 4 Edit Tab Toolbar... 5 Annotations Tab Toolbar...
Matrox MuraControl for Windows
 Matrox MuraControl for Windows User Guide (for software version 6.00) 20179-301-0600 2017.09.25 Contents About this user guide... 6 Using this guide... 6 More information... 6 Overview... 7 Supported Matrox
Matrox MuraControl for Windows User Guide (for software version 6.00) 20179-301-0600 2017.09.25 Contents About this user guide... 6 Using this guide... 6 More information... 6 Overview... 7 Supported Matrox
Selective Space Structures Manual
 Selective Space Structures Manual February 2017 CONTENTS 1 Contents 1 Overview and Concept 4 1.1 General Concept........................... 4 1.2 Modules................................ 6 2 The 3S Generator
Selective Space Structures Manual February 2017 CONTENTS 1 Contents 1 Overview and Concept 4 1.1 General Concept........................... 4 1.2 Modules................................ 6 2 The 3S Generator
Instructions for Display of Imaging Studies Using Stentor isite
 Instructions for Display of Imaging Studies Using Stentor isite Open http://isite.rad.tju.edu in Internet Explorer Log-In: Enter your user name (case Sensitive) and password (case sensitive) and press
Instructions for Display of Imaging Studies Using Stentor isite Open http://isite.rad.tju.edu in Internet Explorer Log-In: Enter your user name (case Sensitive) and password (case sensitive) and press
Correction of susceptibility artifacts in diffusion tensor data using non-linear registration
 Available online at www.sciencedirect.com Medical Image Analysis 11 (2007) 588 603 www.elsevier.com/locate/media Correction of susceptibility artifacts in diffusion tensor data using non-linear registration
Available online at www.sciencedirect.com Medical Image Analysis 11 (2007) 588 603 www.elsevier.com/locate/media Correction of susceptibility artifacts in diffusion tensor data using non-linear registration
Manual image registration in BrainVoyager QX Table of Contents
 Manual image registration in BrainVoyager QX Table of Contents Manual image registration in BrainVoyager QX......1 Performing manual alignment for functional to anatomical images......2 Step 1: preparation......2
Manual image registration in BrainVoyager QX Table of Contents Manual image registration in BrainVoyager QX......1 Performing manual alignment for functional to anatomical images......2 Step 1: preparation......2
Reconstruct Basics Patrick Parker Aug. 2017
 Reconstruct Basics Patrick Parker Aug. 2017 Trace Palette From the trace palette, you can select the color, name, and shape of the trace you want to make. Here s the Trace Palette: If the Trace Palette
Reconstruct Basics Patrick Parker Aug. 2017 Trace Palette From the trace palette, you can select the color, name, and shape of the trace you want to make. Here s the Trace Palette: If the Trace Palette
