Supporting Online Material. Nuclear Pore Complex Structure and Dynamics. revealed by Cryoelectron Tomography
|
|
|
- Rosamond Parsons
- 5 years ago
- Views:
Transcription
1 1 Supporting Online Material Nuclear Pore Complex Structure and Dynamics revealed by Cryoelectron Tomography Martin Beck, Friedrich Förster, Mary Ecke, Jürgen M. Plitzko, Frauke Melchior, Günther Gerisch, Wolfgang Baumeister* and Ohad Medalia* Materials and methods Preparation of the nucleus-enriched fraction Dictyostelium discoideum AX2 cells of an exponentially growing suspension culture were washed with 17 mm phosphate buffer (adjusted to ph 7.5 with KOH) and resuspended in 50 mm Tris ph 7.6, 25 mm KCl, 5mM MgCl 2, 50 mm sucrose, 1 mm DTT and protease inhibitor cocktail (Roche) to a final density of approximately 10 8 cells/ml. The cells were lysed by passing them through a 5 µm polycarbonate filter (Whatman, Nuclepore). Nuclei were spun down (5 min at 500g) and washed in the same buffer to obtain an enriched fraction. All steps were carried out on ice. For nuclear import assays the enriched nuclei were resuspended in cytosolic supernatant, which was prepared by spinning the lysate for 5 min at 20,000 g. Nuclear import assay A volume of 30 µl total cell lysate or a nucleus-enriched fraction supplemented with cytosolic supernatant were used for the transport reactions. Transport mixtures with a total volume of 32 µl contained 50 µg/ml FITC-BSA-NLS and an energy regenerating system (1.5 mm ATP, 5 mm creatine phosphate, 35 U/ml creatine phosphate kinase). Energy depletion was achieved
2 2 by pre-incubation of the lysate with apyrase (100 U/mL) and omitting the components of the energy regenerating system. NLS-dependence was shown using a high concentration of FITC- BSA (up to 375 µg/ml) without NLS. Dependence on soluble cytosolic proteins was shown by resuspension of nucleus-enriched pellets in buffer instead of cytosolic supernatant prior to the transport assay. The samples were incubated for 15 min on ice in the dark and then observed with a Zeiss Axioplan 2 light microscope. Quantification of light microscopy images was done by determining the mean fluorescence of individual nuclei with correction for background fluorescence. To confirm active nuclear import as an energy- and time dependent process the assay was carried out in the presence of various amounts of GMP-PNP (Fig. S1A) and for different time intervals (Fig. S1B). Cryo-electron tomography Enriched suspensions of nuclei were pipetted onto lacy carbon grids (Plano) and rapidly frozen in liquid ethane as described by (1). The data acquisition was done as described previously (2). In brief, the specimens were transferred into a Phillips CM 300 FEG microscope equipped with a Gatan post column GIF 2002 energy filter. Tilt series were collected, typically covering an angular range from -63 to 63, sampled with 1.5 tilt increments and at 15 µm underfocus. Therefore, the theoretical resolution limit as set by the contrast transfer function was 5.5 nm. The pixel size was 0.82 nm at the specimen level. Reconstructions were calculated by weighted backprojection (3). All surface rendered views were created with the Amira software package (TGS). An anisotropic denoising algorithm (4) was used to improve the visualisation shown in (Fig. 1E).
3 3 Image processing 3D alignment of NPCs Individual NPCs were located manually in the tomograms and the corresponding subvolumes were extracted for further processing. An iterative averaging algorithm, similar to approaches used in the field of single particle analysis (5) was used for structure determination. The angular and spatial alignment was performed by an improved procedure based on the method described by (6). The cross correlation function used for alignment accounts for the missing wedge in Fourier space (Förster et al, in preparation). Membrane-bound complexes represent highly anisotropic structures and consequently, their information content varies considerably as a function of spatial direction. Therefore, the cross-correlation function was normalized accordingly (7). Exclusion of particles possessing a low signal-to-noise-ratio (SNR) was done by thresholding based on the intensity of the obtained cross-correlation coefficients. Since the nuclear envelope surrounding the NPCs is highly flexible, the individual particles were masked close to the perimeter of the NPC prior to alignment. During the iterative process the position of the mask was adjusted due to the shifts determined. The orientations of the individual complexes were described by the Eulerian angles ϕ, ψ, and θ. Initially, ψ and θ were approximated by the orientation of the particles on the surface of the nucleus, the shape of which was assumed ellipsoid. An initial model for the NPC was generated using a random distribution of the polar angle ϕ. Eight-fold rotational symmetry along ϕ was detected and therefore imposed during the refinement process in order to improve the SNR. All computations were performed with data cubes of 64x64x64 voxels, each voxel corresponding to 35.2 nm 3 at the specimen level. Finally, five iteration cycles were carried out using 128x128x128 voxels, corresponding to 4.4 nm 3 each. In order to determine the resolution of the averaged structure, the aligned particles were divided into 2 stacks and averaged separately. Fourier-ring correlation (8) of both structures decreases to less than 0.5 below 9 nm (using the 2-σ-criterion the resolution is 6.5 nm, Fig. S3A). Because of the denser
4 4 environment in the nucleoplasm, different thresholds have been applied for the surface rendering of the nuclear basket and the core structure, consisting of the three rings, cytoplasmic filaments and the CP/T (see Fig. 2, 3, nuclear basket in brown, core in blue). The grey value of a subregion in the averaged structure corresponding to nucleoplasm was chosen as threshold level for the intensity of the nuclear basket, while the grey value of a subregion corresponding to solvent (cytoplasmic side) was chosen as threshold level for the intensity of the core structure. Database information The electron density map of the structure has been uploaded to the European Electron Microscopy Database (EMBL-ESI, accession number EMD-1097). Quantitative analysis of the CP/T Due to the missing wedge problem, the classification of particles derived from tomographic volumes by multivariate statistical analysis (9) was not successful. With NPCs active in transport, the highest variability is expected inside the central channel and indeed individual NPCs showed strong variations in this feature of the structure (Fig. S2). Determination of the centres of gravity and the occupied volume served as a classification criterion independent of the missing wedge. CP/Ts were extracted in silico by application of an ellipsoidal mask fitting into the central channel. The grey value of a subregion in the averaged structure corresponding to the solvent was chosen as threshold level for the intensity. The occupied voxels have been summed, and local centres of gravity were determined. The largest occupied volume that we detected (1180 voxels) is consistent with a sphere-shaped complex with a diameter of ~ 40 nm; however, reliable estimates regarding the size of cargo complexes cannot be deduced from such data. The cargo interacts with nucleoporins lining the central channel, and there could be more than one, bound cargo complex. The distribution of the
5 5 occupied volumes is broader than a normal distribution (Fig. 3A), which would indicate that the CP/T in different NPCs comprises essentially the same components. The observed mass is thus likely to represent an average of different structures, rather than a constitutive substructure within the central channel of the NPC, supporting the notion that the CP/T comprises, in part, cargo complexes arrested during translocation. In contrast to the occupied volumes, the distribution of the centers of gravity along the nucleocytoplasmic axis shows two distinct peaks (Fig. 3B). A double Gaussian function can be fitted to this distribution, indicating that two preferred positions of mass within the central channel exist, and that these are likely to correspond to different NPC states. Hence, the distribution of the centers of gravity along the nucleocytoplasmic axis provides a means for an objective classification of individual NPCs. The local minimum between the two peaks served as a criterion for separating the particles into two stacks (position marked white in Fig. S2B, right). The two classes consisted of 115 (LR-class, represented by ~9200 individual images) and 82 (CFclass, represented by ~6500 individual images) particles and have been averaged as described above. The spatial orientation of the particles within each class covered the entire angular spectrum (Fig. S3B). The resolutions of both structures according to the Fourier-ringcorrelation-0.5-criterion were about 10.5 nm (Fig. S3). Structures generated with an equal number of randomly chosen particles had significantly lower resolutions. Surface rendering of both classes was done as described above (see 3D alignment of NPCs).
6 6 Supporting figures Fig. S1. Active nuclear import into isolated Dictyostelium nuclei. (A) Dependence of nuclear import on a non-cleavable GTP analogue. Import reactions have been carried out with various amounts of GMP-PNP, which had an inhibitory effect on nuclear import. (B) Time course of nuclear import. Import reactions were carried out over the periods of time indicated.
7 7 Fig. S2. Visualisation of a nuclear envelope and variations of the CP/T in individual NPCs. (A) Gallery of NPCs sliced perpendicular to the nucleocytoplasmic axis at the lumenal spoke ring plane in comparison to the averaged structure (right). The CP/Ts are connected to the rings by internal filaments (arrows). (B) Gallery of NPCs sliced parallel to the nucleocytoplasmic axis in comparison to the averaged structure (right). The CP/Ts vary in size and shape and occupy diverse positions.
8 8 Fig. S3. Resolution determination and particle orientation. (A) Resolution determination by Fourier ring correlation (FRC). The FRC of the 2 stacks of aligned particles decreased to less than 0.5 below 9 nm (green dotted line). Using the 2-σ criterion (red line), the resolution was 6.5 nm. (B) Individual NPCs of the two classes cover the entire spectra of angular orientation. Plot of the Eulerian angles Phi and Psi against each other (CF-class in blue, LR-class in red). (C) Same as (A) but for the CF-class. The resolution was 10.5 nm. (D) Same as (A) but for the LR-class. The resolution was 10.5 nm.
9 9 Supporting references and notes 1. J. Dubochet et al., Q Rev Biophys 21, (May, 1988). 2. O. Medalia et al., Science 298, (Nov 8, 2002). 3. R. Hegerl, J Struct Biol 116, 30-4 (Oct, 1996). 4. A. S. Frangakis, R. Hegerl, J Struct Biol 135, (Sep, 2001). 5. M. Van Heel, Ultramicroscopy 21, (1987). 6. J. Walz et al., J Struct Biol 120, (Dec, 1997). 7. A. S. Frangakis et al., Proc Natl Acad Sci U S A 99, (Oct 29, 2002). 8. W. O. Saxton, W. Baumeister, J Microsc 127 (Pt 2), (Aug, 1982). 9. J. Frank, M. van Heel, J Mol Biol 161, (Oct 15, 1982).
Supplementary Materials
 Supplementary Materials Microtubule doublets isolated from sea urchin sperm were studied by cryoelectron tomography, producing a density map in which we can identify a number of nontubulin components,
Supplementary Materials Microtubule doublets isolated from sea urchin sperm were studied by cryoelectron tomography, producing a density map in which we can identify a number of nontubulin components,
Supporting information SI 0: TEM & STEM acquisition parameters, in 2D and 3D
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2015 Supporting information SI 0: TEM & STEM acquisition parameters, in 2D and 3D Electron tomography
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2015 Supporting information SI 0: TEM & STEM acquisition parameters, in 2D and 3D Electron tomography
Supporting Information
 Supporting Information Schweitzer et al. 10.1073/pnas.1608050113 SI Materials and Methods 26S Proteasome Purification and Characterization. Fresh human blood from a healthy donor was collected under medical
Supporting Information Schweitzer et al. 10.1073/pnas.1608050113 SI Materials and Methods 26S Proteasome Purification and Characterization. Fresh human blood from a healthy donor was collected under medical
II: Single particle cryoem - an averaging technique
 II: Single particle cryoem - an averaging technique Radiation damage limits the total electron dose that can be used to image biological sample. Thus, images of frozen hydrated macromolecules are very
II: Single particle cryoem - an averaging technique Radiation damage limits the total electron dose that can be used to image biological sample. Thus, images of frozen hydrated macromolecules are very
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12009 Supplementary Figure 1. Experimental tilt series of 104 projections with a tilt range of ±72.6 and equal slope increments, acquired from a Pt nanoparticle using HAADF- STEM (energy:
doi:10.1038/nature12009 Supplementary Figure 1. Experimental tilt series of 104 projections with a tilt range of ±72.6 and equal slope increments, acquired from a Pt nanoparticle using HAADF- STEM (energy:
Structural Information obtained
 Structural Information obtained from Electron Microscopy Christiane Schaffitzel, 09.05.2013 Including slides from John Briggs, Bettina Boettcher, Nicolas Boisset, Andy Hoenger, Michael Schatz, and more
Structural Information obtained from Electron Microscopy Christiane Schaffitzel, 09.05.2013 Including slides from John Briggs, Bettina Boettcher, Nicolas Boisset, Andy Hoenger, Michael Schatz, and more
doi: /
 Yiting Xie ; Anthony P. Reeves; Single 3D cell segmentation from optical CT microscope images. Proc. SPIE 934, Medical Imaging 214: Image Processing, 9343B (March 21, 214); doi:1.1117/12.243852. (214)
Yiting Xie ; Anthony P. Reeves; Single 3D cell segmentation from optical CT microscope images. Proc. SPIE 934, Medical Imaging 214: Image Processing, 9343B (March 21, 214); doi:1.1117/12.243852. (214)
The Electron Microscopy Data Bank and OME
 The Electron Microscopy Data Bank and OME Rich data, quality assessment, and cloud computing Christoph Best European Bioinformatics Institute, Cambridge, UK Transmossion Electron Microscope ADVANTAGES
The Electron Microscopy Data Bank and OME Rich data, quality assessment, and cloud computing Christoph Best European Bioinformatics Institute, Cambridge, UK Transmossion Electron Microscope ADVANTAGES
Quantifying Three-Dimensional Deformations of Migrating Fibroblasts
 45 Chapter 4 Quantifying Three-Dimensional Deformations of Migrating Fibroblasts This chapter presents the full-field displacements and tractions of 3T3 fibroblast cells during migration on polyacrylamide
45 Chapter 4 Quantifying Three-Dimensional Deformations of Migrating Fibroblasts This chapter presents the full-field displacements and tractions of 3T3 fibroblast cells during migration on polyacrylamide
Single Particle Reconstruction Techniques
 T H E U N I V E R S I T Y of T E X A S S C H O O L O F H E A L T H I N F O R M A T I O N S C I E N C E S A T H O U S T O N Single Particle Reconstruction Techniques For students of HI 6001-125 Computational
T H E U N I V E R S I T Y of T E X A S S C H O O L O F H E A L T H I N F O R M A T I O N S C I E N C E S A T H O U S T O N Single Particle Reconstruction Techniques For students of HI 6001-125 Computational
Supplementary Information. Structure of a RSC nucleosome Complex and Insights into Chromatin Remodeling
 Supplementary Information Structure of a RSC nucleosome Complex and Insights into Chromatin Remodeling Yuriy Chaban 1, Chukwudi Ezeokonkwo 1, Wen-Hsiang Chung 1, Fan Zhang 1, Roger D. Kornberg 2, Barbara
Supplementary Information Structure of a RSC nucleosome Complex and Insights into Chromatin Remodeling Yuriy Chaban 1, Chukwudi Ezeokonkwo 1, Wen-Hsiang Chung 1, Fan Zhang 1, Roger D. Kornberg 2, Barbara
THREE-DIMENSIONA L ELECTRON MICROSCOP Y OF MACROMOLECULAR ASSEMBLIE S. Visualization of Biological Molecules in Their Native Stat e.
 THREE-DIMENSIONA L ELECTRON MICROSCOP Y OF MACROMOLECULAR ASSEMBLIE S Visualization of Biological Molecules in Their Native Stat e Joachim Frank CHAPTER 1 Introduction 1 1 The Electron Microscope and
THREE-DIMENSIONA L ELECTRON MICROSCOP Y OF MACROMOLECULAR ASSEMBLIE S Visualization of Biological Molecules in Their Native Stat e Joachim Frank CHAPTER 1 Introduction 1 1 The Electron Microscope and
II: Single particle cryoem - an averaging technique
 II: Single particle cryoem - an averaging technique Radiation damage limits the total electron dose that can be used to image biological sample. Thus, images of frozen hydrated macromolecules are very
II: Single particle cryoem - an averaging technique Radiation damage limits the total electron dose that can be used to image biological sample. Thus, images of frozen hydrated macromolecules are very
Denoising Cryotomograms with IMOD
 Denoising Cryotomograms with IMOD Reasons to Denoise Cryotomograms Easier segmentation of features Presentation Particle picking for subvolume averaging The high-resolution information that you hope to
Denoising Cryotomograms with IMOD Reasons to Denoise Cryotomograms Easier segmentation of features Presentation Particle picking for subvolume averaging The high-resolution information that you hope to
Image Alignment and Application of 2D Fast Rotational Matching in Single Particle cryo-electron Microscopy. Yao Cong. Baylor College of Medicine
 Image Alignment and Application of 2D Fast Rotational Matching in Single Particle cryo-electron Microscopy Yao Cong Baylor College of Medicine cryo-em & single particle analysis Single-particle electron
Image Alignment and Application of 2D Fast Rotational Matching in Single Particle cryo-electron Microscopy Yao Cong Baylor College of Medicine cryo-em & single particle analysis Single-particle electron
Supporting Information. High-Throughput, Algorithmic Determination of Nanoparticle Structure From Electron Microscopy Images
 Supporting Information High-Throughput, Algorithmic Determination of Nanoparticle Structure From Electron Microscopy Images Christine R. Laramy, 1, Keith A. Brown, 2, Matthew N. O Brien, 2 and Chad. A.
Supporting Information High-Throughput, Algorithmic Determination of Nanoparticle Structure From Electron Microscopy Images Christine R. Laramy, 1, Keith A. Brown, 2, Matthew N. O Brien, 2 and Chad. A.
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature10934 Supplementary Methods Mathematical implementation of the EST method. The EST method begins with padding each projection with zeros (that is, embedding
SUPPLEMENTARY INFORMATION doi:10.1038/nature10934 Supplementary Methods Mathematical implementation of the EST method. The EST method begins with padding each projection with zeros (that is, embedding
Three-dimensional structure and flexibility of a membrane-coating module of the nuclear pore complex
 CORRECTION NOTICE Nat. Struct. Mol. Biol. advance online publication, doi:10.1038/nsmb.1618 (7 June 2009) Three-dimensional structure and flexibility of a membrane-coating module of the nuclear pore complex
CORRECTION NOTICE Nat. Struct. Mol. Biol. advance online publication, doi:10.1038/nsmb.1618 (7 June 2009) Three-dimensional structure and flexibility of a membrane-coating module of the nuclear pore complex
Cover Page. The handle holds various files of this Leiden University dissertation
 Cover Page The handle http://hdl.handle.net/1887/48877 holds various files of this Leiden University dissertation Author: Li, Y. Title: A new method to reconstruct the structure from crystal images Issue
Cover Page The handle http://hdl.handle.net/1887/48877 holds various files of this Leiden University dissertation Author: Li, Y. Title: A new method to reconstruct the structure from crystal images Issue
3. Image formation, Fourier analysis and CTF theory. Paula da Fonseca
 3. Image formation, Fourier analysis and CTF theory Paula da Fonseca EM course 2017 - Agenda - Overview of: Introduction to Fourier analysis o o o o Sine waves Fourier transform (simple examples of 1D
3. Image formation, Fourier analysis and CTF theory Paula da Fonseca EM course 2017 - Agenda - Overview of: Introduction to Fourier analysis o o o o Sine waves Fourier transform (simple examples of 1D
Lecture 8. Conical tilt reconstruction
 Lecture 8 Conical tilt reconstruction Central section theorem Euler angles Conical tilt series Missing cone artefact Early D studies and negative staining problems Perspectives and new trends Tilt series
Lecture 8 Conical tilt reconstruction Central section theorem Euler angles Conical tilt series Missing cone artefact Early D studies and negative staining problems Perspectives and new trends Tilt series
Image processing in electron tomography
 A. Méndez-Vilas and J. Díaz (Eds.) Image processing in electron tomography J.J. Fernández 1,2, J.I. Agulleiro 2, J.R. Bilbao-Castro 2, A. Martínez 2, I. García 2, F.J. Chichón 1, J. Martín-Benito 1, J.L.
A. Méndez-Vilas and J. Díaz (Eds.) Image processing in electron tomography J.J. Fernández 1,2, J.I. Agulleiro 2, J.R. Bilbao-Castro 2, A. Martínez 2, I. García 2, F.J. Chichón 1, J. Martín-Benito 1, J.L.
Comparison of fiber orientation analysis methods in Avizo
 Comparison of fiber orientation analysis methods in Avizo More info about this article: http://www.ndt.net/?id=20865 Abstract Rémi Blanc 1, Peter Westenberger 2 1 FEI, 3 Impasse Rudolf Diesel, Bât A, 33708
Comparison of fiber orientation analysis methods in Avizo More info about this article: http://www.ndt.net/?id=20865 Abstract Rémi Blanc 1, Peter Westenberger 2 1 FEI, 3 Impasse Rudolf Diesel, Bât A, 33708
Supporting Information: FIB-SEM Tomography Probes the Meso-Scale Pore Space of an Individual Catalytic Cracking Particle
 Supporting Information: FIB-SEM Tomography Probes the Meso-Scale Pore Space of an Individual Catalytic Cracking Particle D.A.Matthijs de Winter, Florian Meirer, Bert M. Weckhuysen*, Inorganic Chemistry
Supporting Information: FIB-SEM Tomography Probes the Meso-Scale Pore Space of an Individual Catalytic Cracking Particle D.A.Matthijs de Winter, Florian Meirer, Bert M. Weckhuysen*, Inorganic Chemistry
Initial Model Generation
 Initial Model Generation Workshop on Advanced Topics in EM Structure Determination The Scripps Research Institute La Jolla, November 2007 The issue: Structures of the IP3 receptor as determined by single
Initial Model Generation Workshop on Advanced Topics in EM Structure Determination The Scripps Research Institute La Jolla, November 2007 The issue: Structures of the IP3 receptor as determined by single
3DEM > GENERAL METHODS. SINGLE PARTICLE RECONSTRUCTION (SPA): Introduction
 3DEM > GENERAL METHODS SINGLE PARTICLE RECONSTRUCTION (SPA): Introduction SINGLE PARTICLE RECONSTRUCTION > INTRODUCTION General principle of single particle reconstruction o 2D projection images of a 3D
3DEM > GENERAL METHODS SINGLE PARTICLE RECONSTRUCTION (SPA): Introduction SINGLE PARTICLE RECONSTRUCTION > INTRODUCTION General principle of single particle reconstruction o 2D projection images of a 3D
2, the coefficient of variation R 2, and properties of the photon counts traces
 Supplementary Figure 1 Quality control of FCS traces. (a) Typical trace that passes the quality control (QC) according to the parameters shown in f. The QC is based on thresholds applied to fitting parameters
Supplementary Figure 1 Quality control of FCS traces. (a) Typical trace that passes the quality control (QC) according to the parameters shown in f. The QC is based on thresholds applied to fitting parameters
Alignment and Other Challenges in Reconstructing Cryotomograms with IMOD
 Alignment and Other Challenges in Reconstructing Cryotomograms with IMOD Challenges in Cryotomography Alignment, alignment, alignment It can be hard to get fiducials onto/in the sample The low SNR makes
Alignment and Other Challenges in Reconstructing Cryotomograms with IMOD Challenges in Cryotomography Alignment, alignment, alignment It can be hard to get fiducials onto/in the sample The low SNR makes
From cryo-em images of protein complexes to high-resolution 3D structures
 From cryo-em images of protein complexes to high-resolution 3D structures Rouslan Efremov www.cryo-em.be 2018/01/23 Gent What you should do in order for us to make more rapid progress is to make the electron
From cryo-em images of protein complexes to high-resolution 3D structures Rouslan Efremov www.cryo-em.be 2018/01/23 Gent What you should do in order for us to make more rapid progress is to make the electron
The same X-ray transparent in situ pouch cell design and sample holder plates were used for both 2D
 Electronic Supplementary Material (ESI) for Energy & Environmental Science. This journal is The Royal Society of Chemistry 2014 Supplementary Information Experimental Materials and Electrochemistry The
Electronic Supplementary Material (ESI) for Energy & Environmental Science. This journal is The Royal Society of Chemistry 2014 Supplementary Information Experimental Materials and Electrochemistry The
T H E J O U R N A L O F C E L L B I O L O G Y
 Supplemental material Courtois et al., http://www.jcb.org/cgi/content/full/jcb.201202135/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Measurement of distribution and size of MTOCs in the
Supplemental material Courtois et al., http://www.jcb.org/cgi/content/full/jcb.201202135/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Measurement of distribution and size of MTOCs in the
Batch Processing of Tomograms with IMOD
 Batch Processing of Tomograms with IMOD Batch Processing Can Be Useful for a Wide Range of Tilt Series Routine plastic section tilt series can be fully reconstructed automatically with 80-95% success rate
Batch Processing of Tomograms with IMOD Batch Processing Can Be Useful for a Wide Range of Tilt Series Routine plastic section tilt series can be fully reconstructed automatically with 80-95% success rate
2D Image Alignment and Classification
 Structural Biology from Cells to Atoms Optical microscopy D Image Alignment and Classification Yao Cong cryotomography cryomicroscopy 8nm 00 nm 50nm 1nm 5nm Crystallography 0.1nm 0.35nm 0.6nm 0.3nm Shanghai
Structural Biology from Cells to Atoms Optical microscopy D Image Alignment and Classification Yao Cong cryotomography cryomicroscopy 8nm 00 nm 50nm 1nm 5nm Crystallography 0.1nm 0.35nm 0.6nm 0.3nm Shanghai
Determination of the image orientation
 Lecture E. Orlova Determination of the image orientation Images of molecules in vitreous ice Images: shadow, projection Euler angles Methods of orientation determination: Common lines in Fourier space
Lecture E. Orlova Determination of the image orientation Images of molecules in vitreous ice Images: shadow, projection Euler angles Methods of orientation determination: Common lines in Fourier space
STEM electron tomography in the Scanning Electron Microscope
 Journal of Physics: Conference Series PAPER OPEN ACCESS STEM electron tomography in the Scanning Electron Microscope To cite this article: M Ferroni et al 2015 J. Phys.: Conf. Ser. 644 012012 Recent citations
Journal of Physics: Conference Series PAPER OPEN ACCESS STEM electron tomography in the Scanning Electron Microscope To cite this article: M Ferroni et al 2015 J. Phys.: Conf. Ser. 644 012012 Recent citations
EMBO Practical Course on Image Processing for Cryo EM 1-11 September 2015
 EMBO Practical Course on Image Processing for Cryo EM 1-11 September 2015 Practical 4: Optional part for experienced IMOD users - Reconstructing a cryo tomogram and sub-tomogram averaging of GroEL IMOD
EMBO Practical Course on Image Processing for Cryo EM 1-11 September 2015 Practical 4: Optional part for experienced IMOD users - Reconstructing a cryo tomogram and sub-tomogram averaging of GroEL IMOD
3D Energy Dispersive Spectroscopy Elemental Tomography in the Scanning Transmission Electron Microscope
 3D Energy Dispersive Spectroscopy Elemental Tomography in the Scanning Transmission Electron Microscope Brian Van Devener Topics 1.Introduction to EDS in the STEM 2.Extending EDS into three dimensions
3D Energy Dispersive Spectroscopy Elemental Tomography in the Scanning Transmission Electron Microscope Brian Van Devener Topics 1.Introduction to EDS in the STEM 2.Extending EDS into three dimensions
Digital Image Processing
 Digital Image Processing Part 9: Representation and Description AASS Learning Systems Lab, Dep. Teknik Room T1209 (Fr, 11-12 o'clock) achim.lilienthal@oru.se Course Book Chapter 11 2011-05-17 Contents
Digital Image Processing Part 9: Representation and Description AASS Learning Systems Lab, Dep. Teknik Room T1209 (Fr, 11-12 o'clock) achim.lilienthal@oru.se Course Book Chapter 11 2011-05-17 Contents
SUPPLEMENTARY INFORMATION
 Supplementary Information Compact spectrometer based on a disordered photonic chip Brandon Redding, Seng Fatt Liew, Raktim Sarma, Hui Cao* Department of Applied Physics, Yale University, New Haven, CT
Supplementary Information Compact spectrometer based on a disordered photonic chip Brandon Redding, Seng Fatt Liew, Raktim Sarma, Hui Cao* Department of Applied Physics, Yale University, New Haven, CT
12/7/2012. Biomolecular structure. Diffraction, X-ray crystallography, light- and electron microscopy. CD spectroscopy, mass spectrometry
 phase difference at a given distance constructive/destructive interference Biomolecular structure. Diffraction, X-ray crystallography, light- and electron microscopy. CD spectroscopy, mass spectrometry
phase difference at a given distance constructive/destructive interference Biomolecular structure. Diffraction, X-ray crystallography, light- and electron microscopy. CD spectroscopy, mass spectrometry
Random Conical tilt 3D reconstruction
 Random Conical tilt 3D reconstruction Central section theorem Euler angles Principle of conical tilt series Missing cone artefact Multivariate statistical analysis Early 3D studies and negative staining
Random Conical tilt 3D reconstruction Central section theorem Euler angles Principle of conical tilt series Missing cone artefact Multivariate statistical analysis Early 3D studies and negative staining
X-Ray fluorescence and Raman spectroscopy
 X-Ray fluorescence and Raman spectroscopy Advanced physics laboratory (nd part) 4CFU Catalini Letizia, De Angelis Giulia Vittoria, Piselli Verdiana Abstract In this paper we report about two different
X-Ray fluorescence and Raman spectroscopy Advanced physics laboratory (nd part) 4CFU Catalini Letizia, De Angelis Giulia Vittoria, Piselli Verdiana Abstract In this paper we report about two different
Likelihood-based optimization of cryo-em data. Sjors Scheres National Center for Biotechnology CSIC Madrid, Spain
 Likelihood-based optimization of cryo-em data Sjors Scheres National Center for Biotechnology CSIC Madrid, Spain Life based on molecular machines DNA replication Protein synthesis Dynein motion Molecular
Likelihood-based optimization of cryo-em data Sjors Scheres National Center for Biotechnology CSIC Madrid, Spain Life based on molecular machines DNA replication Protein synthesis Dynein motion Molecular
Random Conical tilt 3D reconstruction
 Sali A, Glaeser R, Earnest T, Baumeister W. (00) From Words to literature in structural proteomics. Nature (98): 1-. Random Conical tilt D reconstruction Central section theorem Euler angles Principle
Sali A, Glaeser R, Earnest T, Baumeister W. (00) From Words to literature in structural proteomics. Nature (98): 1-. Random Conical tilt D reconstruction Central section theorem Euler angles Principle
Chapter 1 Introduction
 Chapter 1 Introduction 1.1 Structural biology, cryo-em and image processing Structural biology is a branch of life science which focuses on the structures of biological macromolecules, investigating what
Chapter 1 Introduction 1.1 Structural biology, cryo-em and image processing Structural biology is a branch of life science which focuses on the structures of biological macromolecules, investigating what
Supplementary Materials for
 www.advances.sciencemag.org/cgi/content/full/1/3/e1400199/dc1 Supplementary Materials for Life and death of a single catalytic cracking particle Florian Meirer, Sam Kalirai, Darius Morris, Santosh Soparawalla,
www.advances.sciencemag.org/cgi/content/full/1/3/e1400199/dc1 Supplementary Materials for Life and death of a single catalytic cracking particle Florian Meirer, Sam Kalirai, Darius Morris, Santosh Soparawalla,
Enhanced material contrast by dual-energy microct imaging
 Enhanced material contrast by dual-energy microct imaging Method note Page 1 of 12 2 Method note: Dual-energy microct analysis 1. Introduction 1.1. The basis for dual energy imaging Micro-computed tomography
Enhanced material contrast by dual-energy microct imaging Method note Page 1 of 12 2 Method note: Dual-energy microct analysis 1. Introduction 1.1. The basis for dual energy imaging Micro-computed tomography
Structural Basis for RNA Processing by the Human. RISC-Loading Complex
 Supplementary Information Structural Basis for RNA Processing by the Human RISC-Loading Complex Hong-Wei Wang, Cameron Noland, Bunpote Siridechadilok, David W. Taylor, Enbo Ma, Karin Felderer, Jennifer
Supplementary Information Structural Basis for RNA Processing by the Human RISC-Loading Complex Hong-Wei Wang, Cameron Noland, Bunpote Siridechadilok, David W. Taylor, Enbo Ma, Karin Felderer, Jennifer
Digital Volume Correlation for Materials Characterization
 19 th World Conference on Non-Destructive Testing 2016 Digital Volume Correlation for Materials Characterization Enrico QUINTANA, Phillip REU, Edward JIMENEZ, Kyle THOMPSON, Sharlotte KRAMER Sandia National
19 th World Conference on Non-Destructive Testing 2016 Digital Volume Correlation for Materials Characterization Enrico QUINTANA, Phillip REU, Edward JIMENEZ, Kyle THOMPSON, Sharlotte KRAMER Sandia National
WCIF COLOCALIS ATION PLUGINS
 WCIF COLOCALIS ATION PLUGINS Colocalisation Test Background When a coefficient is calculated for two images, it is often unclear quite what this means, in particular for intermediate values. This raises
WCIF COLOCALIS ATION PLUGINS Colocalisation Test Background When a coefficient is calculated for two images, it is often unclear quite what this means, in particular for intermediate values. This raises
Automatic Partiicle Tracking Software USE ER MANUAL Update: May 2015
 Automatic Particle Tracking Software USER MANUAL Update: May 2015 File Menu The micrograph below shows the panel displayed when a movie is opened, including a playback menu where most of the parameters
Automatic Particle Tracking Software USER MANUAL Update: May 2015 File Menu The micrograph below shows the panel displayed when a movie is opened, including a playback menu where most of the parameters
18th World Conference on Nondestructive Testing, April 2012, Durban, South Africa
 18th World Conference on Nondestructive Testing, 16-20 April 2012, Durban, South Africa TOWARDS ESTABLISHMENT OF STANDARDIZED PRACTICE FOR ASSESSMENT OF SPATIAL RESOLUTION AND CONTRAST OF INTERNATIONAL
18th World Conference on Nondestructive Testing, 16-20 April 2012, Durban, South Africa TOWARDS ESTABLISHMENT OF STANDARDIZED PRACTICE FOR ASSESSMENT OF SPATIAL RESOLUTION AND CONTRAST OF INTERNATIONAL
SuRVoS Workbench. Super-Region Volume Segmentation. Imanol Luengo
 SuRVoS Workbench Super-Region Volume Segmentation Imanol Luengo Index - The project - What is SuRVoS - SuRVoS Overview - What can it do - Overview of the internals - Current state & Limitations - Future
SuRVoS Workbench Super-Region Volume Segmentation Imanol Luengo Index - The project - What is SuRVoS - SuRVoS Overview - What can it do - Overview of the internals - Current state & Limitations - Future
Digital Core study of Wanaea and Perseus Core Fragments:
 Digital Core study of Wanaea and Perseus Core Fragments: Summary for Woodside Energy Mark A. Knackstedt,2, A. Ghous 2, C. H. Arns, H. Averdunk, F. Bauget, A. Sakellariou, T.J. Senden, A.P. Sheppard,R.
Digital Core study of Wanaea and Perseus Core Fragments: Summary for Woodside Energy Mark A. Knackstedt,2, A. Ghous 2, C. H. Arns, H. Averdunk, F. Bauget, A. Sakellariou, T.J. Senden, A.P. Sheppard,R.
Presentation and analysis of multidimensional data sets
 Presentation and analysis of multidimensional data sets Overview 1. 3D data visualisation Multidimensional images Data pre-processing Visualisation methods Multidimensional images wavelength time 3D image
Presentation and analysis of multidimensional data sets Overview 1. 3D data visualisation Multidimensional images Data pre-processing Visualisation methods Multidimensional images wavelength time 3D image
Experiment Guide - Trypan Blue Image Analysis. Technical Note. Content
 Experiment Guide - Trypan Blue Image Analysis The purpose of this document is to guide the user through the Trypan Blue Image analysis. It does not contain any procedures of setting up a Trypan Blue assay
Experiment Guide - Trypan Blue Image Analysis The purpose of this document is to guide the user through the Trypan Blue Image analysis. It does not contain any procedures of setting up a Trypan Blue assay
Using a multipoint interferometer to measure the orbital angular momentum of light
 CHAPTER 3 Using a multipoint interferometer to measure the orbital angular momentum of light Recently it was shown that the orbital angular momentum of light can be measured using a multipoint interferometer,
CHAPTER 3 Using a multipoint interferometer to measure the orbital angular momentum of light Recently it was shown that the orbital angular momentum of light can be measured using a multipoint interferometer,
9 length of contour = no. of horizontal and vertical components + ( 2 no. of diagonal components) diameter of boundary B
 8. Boundary Descriptor 8.. Some Simple Descriptors length of contour : simplest descriptor - chain-coded curve 9 length of contour no. of horiontal and vertical components ( no. of diagonal components
8. Boundary Descriptor 8.. Some Simple Descriptors length of contour : simplest descriptor - chain-coded curve 9 length of contour no. of horiontal and vertical components ( no. of diagonal components
45 µm polystyrene bead embedded in scattering tissue phantom. (a,b) raw images under oblique
 Phase gradient microscopy in thick tissue with oblique back-illumination Tim N Ford, Kengyeh K Chu & Jerome Mertz Supplementary Figure 1: Comparison of added versus subtracted raw OBM images 45 µm polystyrene
Phase gradient microscopy in thick tissue with oblique back-illumination Tim N Ford, Kengyeh K Chu & Jerome Mertz Supplementary Figure 1: Comparison of added versus subtracted raw OBM images 45 µm polystyrene
Determination of Three-Dimensional Voxel Sensitivity for Two- and Three-Headed Coincidence Imaging
 IEEE TRANSACTIONS ON NUCLEAR SCIENCE, VOL. 50, NO. 3, JUNE 2003 405 Determination of Three-Dimensional Voxel Sensitivity for Two- and Three-Headed Coincidence Imaging Edward J. Soares, Kevin W. Germino,
IEEE TRANSACTIONS ON NUCLEAR SCIENCE, VOL. 50, NO. 3, JUNE 2003 405 Determination of Three-Dimensional Voxel Sensitivity for Two- and Three-Headed Coincidence Imaging Edward J. Soares, Kevin W. Germino,
Lecture 8 Object Descriptors
 Lecture 8 Object Descriptors Azadeh Fakhrzadeh Centre for Image Analysis Swedish University of Agricultural Sciences Uppsala University 2 Reading instructions Chapter 11.1 11.4 in G-W Azadeh Fakhrzadeh
Lecture 8 Object Descriptors Azadeh Fakhrzadeh Centre for Image Analysis Swedish University of Agricultural Sciences Uppsala University 2 Reading instructions Chapter 11.1 11.4 in G-W Azadeh Fakhrzadeh
Effect of OTF attenuation on single-slice SR-SIM reconstruction
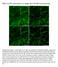 Effect of OTF attenuation on single-slice SR-SIM reconstruction Supplementary Figure 1: Actin filaments in U2OS cells, labelled with Phalloidin-Atto488, measured on a DeltaVision OMX, excited at 488 nm
Effect of OTF attenuation on single-slice SR-SIM reconstruction Supplementary Figure 1: Actin filaments in U2OS cells, labelled with Phalloidin-Atto488, measured on a DeltaVision OMX, excited at 488 nm
A novel 3D wavelet-based filter for visualizing features in noisy
 Journal of Microscopy, Vol. 219, Pt 2 August 2005, pp. 43 49 Received 28 January 2005; accepted 11 May 2005 A novel 3D wavelet-based filter for visualizing features in noisy Blackwell Publishing, Ltd.
Journal of Microscopy, Vol. 219, Pt 2 August 2005, pp. 43 49 Received 28 January 2005; accepted 11 May 2005 A novel 3D wavelet-based filter for visualizing features in noisy Blackwell Publishing, Ltd.
Experiment 8 Wave Optics
 Physics 263 Experiment 8 Wave Optics In this laboratory, we will perform two experiments on wave optics. 1 Double Slit Interference In two-slit interference, light falls on an opaque screen with two closely
Physics 263 Experiment 8 Wave Optics In this laboratory, we will perform two experiments on wave optics. 1 Double Slit Interference In two-slit interference, light falls on an opaque screen with two closely
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Schematic demonstrating the preferred orientation sampling problem in single-particle cryo-em. Specimens sticking to the air-water interface or a substrate support of a cryo-em grid
Supplementary Figure 1 Schematic demonstrating the preferred orientation sampling problem in single-particle cryo-em. Specimens sticking to the air-water interface or a substrate support of a cryo-em grid
Cryo-electron microscopy Cryo-EM. Garry Taylor
 Cryo-electron microscopy Cryo-EM Garry Taylor www.st-andrews.ac.uk/~glt2/bl3301 Electron has a wavelength de Broglie relationship: m v = h / λ or λ = h / mv Accelerate e - in a field of potential V, it
Cryo-electron microscopy Cryo-EM Garry Taylor www.st-andrews.ac.uk/~glt2/bl3301 Electron has a wavelength de Broglie relationship: m v = h / λ or λ = h / mv Accelerate e - in a field of potential V, it
DETECTION AND ROBUST ESTIMATION OF CYLINDER FEATURES IN POINT CLOUDS INTRODUCTION
 DETECTION AND ROBUST ESTIMATION OF CYLINDER FEATURES IN POINT CLOUDS Yun-Ting Su James Bethel Geomatics Engineering School of Civil Engineering Purdue University 550 Stadium Mall Drive, West Lafayette,
DETECTION AND ROBUST ESTIMATION OF CYLINDER FEATURES IN POINT CLOUDS Yun-Ting Su James Bethel Geomatics Engineering School of Civil Engineering Purdue University 550 Stadium Mall Drive, West Lafayette,
Filtering Images. Contents
 Image Processing and Data Visualization with MATLAB Filtering Images Hansrudi Noser June 8-9, 010 UZH, Multimedia and Robotics Summer School Noise Smoothing Filters Sigmoid Filters Gradient Filters Contents
Image Processing and Data Visualization with MATLAB Filtering Images Hansrudi Noser June 8-9, 010 UZH, Multimedia and Robotics Summer School Noise Smoothing Filters Sigmoid Filters Gradient Filters Contents
SUPPLEMENTARY FILE S1: 3D AIRWAY TUBE RECONSTRUCTION AND CELL-BASED MECHANICAL MODEL. RELATED TO FIGURE 1, FIGURE 7, AND STAR METHODS.
 SUPPLEMENTARY FILE S1: 3D AIRWAY TUBE RECONSTRUCTION AND CELL-BASED MECHANICAL MODEL. RELATED TO FIGURE 1, FIGURE 7, AND STAR METHODS. 1. 3D AIRWAY TUBE RECONSTRUCTION. RELATED TO FIGURE 1 AND STAR METHODS
SUPPLEMENTARY FILE S1: 3D AIRWAY TUBE RECONSTRUCTION AND CELL-BASED MECHANICAL MODEL. RELATED TO FIGURE 1, FIGURE 7, AND STAR METHODS. 1. 3D AIRWAY TUBE RECONSTRUCTION. RELATED TO FIGURE 1 AND STAR METHODS
X-RAY crystallography [1], [2] and nuclear magnetic. Computational Approaches for Automatic Structural Analysis of Large Bio-molecular Complexes
![X-RAY crystallography [1], [2] and nuclear magnetic. Computational Approaches for Automatic Structural Analysis of Large Bio-molecular Complexes X-RAY crystallography [1], [2] and nuclear magnetic. Computational Approaches for Automatic Structural Analysis of Large Bio-molecular Complexes](/thumbs/84/89164539.jpg) IEEE/ACM TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS 1 Computational Approaches for Automatic Structural Analysis of Large Bio-molecular Complexes Zeyun Yu, Student Member, IEEE, Chandrajit
IEEE/ACM TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS 1 Computational Approaches for Automatic Structural Analysis of Large Bio-molecular Complexes Zeyun Yu, Student Member, IEEE, Chandrajit
Single-particle electron microscopy (cryo-electron microscopy) CS/CME/BioE/Biophys/BMI 279 Nov. 16 and 28, 2017 Ron Dror
 Single-particle electron microscopy (cryo-electron microscopy) CS/CME/BioE/Biophys/BMI 279 Nov. 16 and 28, 2017 Ron Dror 1 Last month s Nobel Prize in Chemistry Awarded to Jacques Dubochet, Joachim Frank
Single-particle electron microscopy (cryo-electron microscopy) CS/CME/BioE/Biophys/BMI 279 Nov. 16 and 28, 2017 Ron Dror 1 Last month s Nobel Prize in Chemistry Awarded to Jacques Dubochet, Joachim Frank
Math D Printing Group Final Report
 Math 4020 3D Printing Group Final Report Gayan Abeynanda Brandon Oubre Jeremy Tillay Dustin Wright Wednesday 29 th April, 2015 Introduction 3D printers are a growing technology, but there is currently
Math 4020 3D Printing Group Final Report Gayan Abeynanda Brandon Oubre Jeremy Tillay Dustin Wright Wednesday 29 th April, 2015 Introduction 3D printers are a growing technology, but there is currently
Fluorescence Tomography Source Reconstruction and Analysis
 TECHNICAL NOTE Pre-clinical in vivo imaging Fluorescence Tomography Source Reconstruction and Analysis Note: This Technical Note is part of a series for Fluorescence Imaging Tomography (FLIT). The user
TECHNICAL NOTE Pre-clinical in vivo imaging Fluorescence Tomography Source Reconstruction and Analysis Note: This Technical Note is part of a series for Fluorescence Imaging Tomography (FLIT). The user
Image Reconstruction from Multiple Projections ECE 6258 Class project
 Image Reconstruction from Multiple Projections ECE 658 Class project Introduction: The ability to reconstruct an object from multiple angular projections is a powerful tool. What this procedure gives people
Image Reconstruction from Multiple Projections ECE 658 Class project Introduction: The ability to reconstruct an object from multiple angular projections is a powerful tool. What this procedure gives people
Compressed Sensing for Electron Tomography
 University of Maryland, College Park Department of Mathematics February 10, 2015 1/33 Outline I Introduction 1 Introduction 2 3 4 2/33 1 Introduction 2 3 4 3/33 Tomography Introduction Tomography - Producing
University of Maryland, College Park Department of Mathematics February 10, 2015 1/33 Outline I Introduction 1 Introduction 2 3 4 2/33 1 Introduction 2 3 4 3/33 Tomography Introduction Tomography - Producing
Introductory Guide to Light Microscopy - Biomedical Confocal Microscopy
 Introductory Guide to Light Microscopy - Biomedical Confocal Microscopy 7 May 2007 Michael Hooker Microscopy Facility Michael Chua microscopy@unc.edu 843-3268 6007 Thurston Bowles Wendy Salmon wendy_salmon@med.unc.edu
Introductory Guide to Light Microscopy - Biomedical Confocal Microscopy 7 May 2007 Michael Hooker Microscopy Facility Michael Chua microscopy@unc.edu 843-3268 6007 Thurston Bowles Wendy Salmon wendy_salmon@med.unc.edu
Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging
 1 CS 9 Final Project Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging Feiyu Chen Department of Electrical Engineering ABSTRACT Subject motion is a significant
1 CS 9 Final Project Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging Feiyu Chen Department of Electrical Engineering ABSTRACT Subject motion is a significant
BMB/Bi/Ch 173 Winter 2017
 BMB/Bi/Ch 173 Winter 2017 Homework Set 3.2 Assigned 1/26/2017, Due 1/31/17 by 10:30am TA - Sara Weaver sjweaver [a] Caltech.edu Office hours Broad 3rd floor kitchen - Friday 1/27 1:30pm-2:30pm, Monday
BMB/Bi/Ch 173 Winter 2017 Homework Set 3.2 Assigned 1/26/2017, Due 1/31/17 by 10:30am TA - Sara Weaver sjweaver [a] Caltech.edu Office hours Broad 3rd floor kitchen - Friday 1/27 1:30pm-2:30pm, Monday
HIGH-SPEED THEE-DIMENSIONAL TOMOGRAPHIC IMAGING OF FRAGMENTS AND PRECISE STATISTICS FROM AN AUTOMATED ANALYSIS
 23 RD INTERNATIONAL SYMPOSIUM ON BALLISTICS TARRAGONA, SPAIN 16-20 APRIL 2007 HIGH-SPEED THEE-DIMENSIONAL TOMOGRAPHIC IMAGING OF FRAGMENTS AND PRECISE STATISTICS FROM AN AUTOMATED ANALYSIS P. Helberg 1,
23 RD INTERNATIONAL SYMPOSIUM ON BALLISTICS TARRAGONA, SPAIN 16-20 APRIL 2007 HIGH-SPEED THEE-DIMENSIONAL TOMOGRAPHIC IMAGING OF FRAGMENTS AND PRECISE STATISTICS FROM AN AUTOMATED ANALYSIS P. Helberg 1,
Difficult Problems & EMAN2 Introduction
 Difficult Problems & EMAN2 Introduction Steve Ludtke National Center for Macromolecular Imaging Biochemistry & Molecular Biology Baylor College of Medicine sludtke@bcm.edu Initial Model Bias? EMAN2 'monte
Difficult Problems & EMAN2 Introduction Steve Ludtke National Center for Macromolecular Imaging Biochemistry & Molecular Biology Baylor College of Medicine sludtke@bcm.edu Initial Model Bias? EMAN2 'monte
ADVANCED RECONSTRUCTION FOR ELECTRON MICROSCOPY
 1 ADVANCED RECONSTRUCTION FOR ELECTRON MICROSCOPY SUHAS SREEHARI S. V. VENKATAKRISHNAN (VENKAT) CHARLES A. BOUMAN PURDUE UNIVERSITY AUGUST 15, 2014 2 OUTLINE 1. Overview of MBIR 2. Previous work 3. Leading
1 ADVANCED RECONSTRUCTION FOR ELECTRON MICROSCOPY SUHAS SREEHARI S. V. VENKATAKRISHNAN (VENKAT) CHARLES A. BOUMAN PURDUE UNIVERSITY AUGUST 15, 2014 2 OUTLINE 1. Overview of MBIR 2. Previous work 3. Leading
3D-Modeling with Microscopy Image Browser (im_browser) (pdf version) Ilya Belevich, Darshan Kumar, Helena Vihinen
 3D-Modeling with Microscopy Image Browser (im_browser) (pdf version) Ilya Belevich, Darshan Kumar, Helena Vihinen Dataset: Huh7.tif (original), obtained by Serial Block Face-SEM (Gatan 3View) Huh7_crop.tif
3D-Modeling with Microscopy Image Browser (im_browser) (pdf version) Ilya Belevich, Darshan Kumar, Helena Vihinen Dataset: Huh7.tif (original), obtained by Serial Block Face-SEM (Gatan 3View) Huh7_crop.tif
Classification of Hyperspectral Breast Images for Cancer Detection. Sander Parawira December 4, 2009
 1 Introduction Classification of Hyperspectral Breast Images for Cancer Detection Sander Parawira December 4, 2009 parawira@stanford.edu In 2009 approximately one out of eight women has breast cancer.
1 Introduction Classification of Hyperspectral Breast Images for Cancer Detection Sander Parawira December 4, 2009 parawira@stanford.edu In 2009 approximately one out of eight women has breast cancer.
Deformation of granular texture media studied by X-ray CT & 3D DIC at the continuous and microstructure scales
 Deformation of granular texture media studied by X-ray CT & 3D DIC at the continuous and microstructure scales N. Lenoir, S.A. Hall, J. Desrues, G. Viggiani and P. Bésuelle (Laboratoire 3S-R) Michel Bornert
Deformation of granular texture media studied by X-ray CT & 3D DIC at the continuous and microstructure scales N. Lenoir, S.A. Hall, J. Desrues, G. Viggiani and P. Bésuelle (Laboratoire 3S-R) Michel Bornert
Nonbias-Limited Tracking of Spherical Particles, Enabling Nanometer Resolution at Low Magnification
 Nonbias-Limited Tracking of Spherical Particles, Enabling Nanometer Resolution at Low Magnification Marijn T. J. van Loenhout, Jacob Kerssemakers, Iwijn De Vlaminck, and Cees Dekker Department of Bionanoscience,
Nonbias-Limited Tracking of Spherical Particles, Enabling Nanometer Resolution at Low Magnification Marijn T. J. van Loenhout, Jacob Kerssemakers, Iwijn De Vlaminck, and Cees Dekker Department of Bionanoscience,
Algorithm User Guide:
 Algorithm User Guide: Membrane Quantification Use the Aperio algorithms to adjust (tune) the parameters until the quantitative results are sufficiently accurate for the purpose for which you intend to
Algorithm User Guide: Membrane Quantification Use the Aperio algorithms to adjust (tune) the parameters until the quantitative results are sufficiently accurate for the purpose for which you intend to
Image Acquisition Systems
 Image Acquisition Systems Goals and Terminology Conventional Radiography Axial Tomography Computer Axial Tomography (CAT) Magnetic Resonance Imaging (MRI) PET, SPECT Ultrasound Microscopy Imaging ITCS
Image Acquisition Systems Goals and Terminology Conventional Radiography Axial Tomography Computer Axial Tomography (CAT) Magnetic Resonance Imaging (MRI) PET, SPECT Ultrasound Microscopy Imaging ITCS
Analysis of Planar Anisotropy of Fibre Systems by Using 2D Fourier Transform
 Maroš Tunák, Aleš Linka Technical University in Liberec Faculty of Textile Engineering Department of Textile Materials Studentská 2, 461 17 Liberec 1, Czech Republic E-mail: maros.tunak@tul.cz ales.linka@tul.cz
Maroš Tunák, Aleš Linka Technical University in Liberec Faculty of Textile Engineering Department of Textile Materials Studentská 2, 461 17 Liberec 1, Czech Republic E-mail: maros.tunak@tul.cz ales.linka@tul.cz
Raster Image Correlation Spectroscopy and Number and Brightness Analysis
 Raster Image Correlation Spectroscopy and Number and Brightness Analysis 15 Principles of Fluorescence Course Paolo Annibale, PhD On behalf of Dr. Enrico Gratton lfd Raster Image Correlation Spectroscopy
Raster Image Correlation Spectroscopy and Number and Brightness Analysis 15 Principles of Fluorescence Course Paolo Annibale, PhD On behalf of Dr. Enrico Gratton lfd Raster Image Correlation Spectroscopy
APPROACHES IN QUANTITATIVE, MULTI-DIMENSIONAL MICROSCOPY
 APPROACHES IN QUANTITATIVE, MULTI-DIMENSIONAL MICROSCOPY Profs. Zvi Kam and Benjamin Geiger Department of Molecular Cell Biology The Weizmann Institute of Science Rehovot, Israel In this presentation we
APPROACHES IN QUANTITATIVE, MULTI-DIMENSIONAL MICROSCOPY Profs. Zvi Kam and Benjamin Geiger Department of Molecular Cell Biology The Weizmann Institute of Science Rehovot, Israel In this presentation we
CT Reconstruction with Good-Orientation and Layer Separation for Multilayer Objects
 17th World Conference on Nondestructive Testing, 25-28 Oct 2008, Shanghai, China CT Reconstruction with Good-Orientation and Layer Separation for Multilayer Objects Tong LIU 1, Brian Stephan WONG 2, Tai
17th World Conference on Nondestructive Testing, 25-28 Oct 2008, Shanghai, China CT Reconstruction with Good-Orientation and Layer Separation for Multilayer Objects Tong LIU 1, Brian Stephan WONG 2, Tai
Chapter 3 Image Registration. Chapter 3 Image Registration
 Chapter 3 Image Registration Distributed Algorithms for Introduction (1) Definition: Image Registration Input: 2 images of the same scene but taken from different perspectives Goal: Identify transformation
Chapter 3 Image Registration Distributed Algorithms for Introduction (1) Definition: Image Registration Input: 2 images of the same scene but taken from different perspectives Goal: Identify transformation
Independent Resolution Test of
 Independent Resolution Test of as conducted and published by Dr. Adam Puche, PhD University of Maryland June 2005 as presented by (formerly Thales Optem Inc.) www.qioptiqimaging.com Independent Resolution
Independent Resolution Test of as conducted and published by Dr. Adam Puche, PhD University of Maryland June 2005 as presented by (formerly Thales Optem Inc.) www.qioptiqimaging.com Independent Resolution
Counting Particles or Cells Using IMAQ Vision
 Application Note 107 Counting Particles or Cells Using IMAQ Vision John Hanks Introduction To count objects, you use a common image processing technique called particle analysis, often referred to as blob
Application Note 107 Counting Particles or Cells Using IMAQ Vision John Hanks Introduction To count objects, you use a common image processing technique called particle analysis, often referred to as blob
Virtual Frap User Guide
 Virtual Frap User Guide http://wiki.vcell.uchc.edu/twiki/bin/view/vcell/vfrap Center for Cell Analysis and Modeling University of Connecticut Health Center 2010-1 - 1 Introduction Flourescence Photobleaching
Virtual Frap User Guide http://wiki.vcell.uchc.edu/twiki/bin/view/vcell/vfrap Center for Cell Analysis and Modeling University of Connecticut Health Center 2010-1 - 1 Introduction Flourescence Photobleaching
Multivariate Calibration Quick Guide
 Last Updated: 06.06.2007 Table Of Contents 1. HOW TO CREATE CALIBRATION MODELS...1 1.1. Introduction into Multivariate Calibration Modelling... 1 1.1.1. Preparing Data... 1 1.2. Step 1: Calibration Wizard
Last Updated: 06.06.2007 Table Of Contents 1. HOW TO CREATE CALIBRATION MODELS...1 1.1. Introduction into Multivariate Calibration Modelling... 1 1.1.1. Preparing Data... 1 1.2. Step 1: Calibration Wizard
A NEW APPROACH FOR 3D SEGMENTATION OF CELLULAR TOMOGRAMS OBTAINED USING THREE-DIMENSIONAL ELECTRON MICROSCOPY
 A NEW APPROACH FOR 3D SEGMENTATION OF CELLULAR TOMOGRAMS OBTAINED USING THREE-DIMENSIONAL ELECTRON MICROSCOPY A. Bartesaghi and G. Sapiro University of Minnesota Electrical and Computer Engineering Department
A NEW APPROACH FOR 3D SEGMENTATION OF CELLULAR TOMOGRAMS OBTAINED USING THREE-DIMENSIONAL ELECTRON MICROSCOPY A. Bartesaghi and G. Sapiro University of Minnesota Electrical and Computer Engineering Department
C101-E137 TALK LETTER. Vol. 14
 C101-E137 TALK LETTER Vol. 14 Diffuse Reflectance Measurement of Powder Samples and Kubelka-Munk Transformation ------- 02 Application: Measuring Food Color ------- 08 Q&A: What effect does the scan speed
C101-E137 TALK LETTER Vol. 14 Diffuse Reflectance Measurement of Powder Samples and Kubelka-Munk Transformation ------- 02 Application: Measuring Food Color ------- 08 Q&A: What effect does the scan speed
generator graphical interface for generating the IHRSR script
 IHRSR Programs generator graphical interface for generating the IHRSR script This program (generator) is invoked in the directory where you will want the IHRSR script written. The output file created by
IHRSR Programs generator graphical interface for generating the IHRSR script This program (generator) is invoked in the directory where you will want the IHRSR script written. The output file created by
SPCM Software Runs Online-FLIM at 10 Images per Second
 SPCM Software Runs Online-FLIM at 10 Images per Second Abstract: Version 9.72 SPCM software of the bh TCSPC/FLIM systems displays fluorescence lifetime images at a rate of 10 images per second. The calculation
SPCM Software Runs Online-FLIM at 10 Images per Second Abstract: Version 9.72 SPCM software of the bh TCSPC/FLIM systems displays fluorescence lifetime images at a rate of 10 images per second. The calculation
