EMSegment Tutorial. How to Define and Fine-Tune Automatic Brain Compartment Segmentation and the Detection of White Matter Hyperintensities
|
|
|
- Stanley Arnold
- 5 years ago
- Views:
Transcription
1 EMSegment Tutorial How to Define and Fine-Tune Automatic Brain Compartment Segmentation and the Detection of White Matter Hyperintensities This documentation serves as a tutorial to learn to customize the 3D Slicer EMSegment module for White Matter Hyperintensity and Deep Gray Matter segmentation based on a sample case. The strategies described here can also be applied to different segmentation problems. Istvan Csapo, Kilian Pohl, Charles R.G. Guttmann Center for Neurological Imaging and Surgical Planning Laboratory Brigham & Women's Hospital Harvard Medical School icsapo@bwh.harvard.edu pohl@csail.mit.edu guttmann@bwh.harvard.edu Version 1 (3D Slicer 2.6) Last update: January 4,
2 T a b l e o f C o n t e n t s HOW TO USE THIS TUTORIAL...3 General Usage... 3 Glossary... 3 Sample Case... 3 SEGMENTATION Gray Matter, White Matter, and CSF Segmentation General Strategy Files Used Loading the Subject and Atlas Images Setting the EMSegment Parameters Setting Up the Class Hierarchy (Class Tab) Class Distributions EM Algorithm Settings Running EMSegment Results Saving the Results and EMSegment Parameters Adding the Deep Gray Matter Subclass General Strategy Files Used Loading the DEEP GM Atlas Image Modifying the Class Hierarchy (Class Tab) Running EMSegment Adding the White Matter Hyperintensity Segmentation General Strategy REFERENCES
3 H o w t o U s e T h i s T u t o r i a l General Usage This guide assumes a basic knowledge of 3D Slicer, including, but not limited to loading volumes and label maps; loading scenes; saving volumes; and registering images. For further information on these topics, please refer to the 3D Slicer website ( Glossary The terminology used in reference to general Slicer components is the same as found on the Slicer website. CGM - cortical gray matter CSF - cerebrospinal fluid DEEPGM - deep gray matter (caudate, putamen, thalamus) ECC - extra cranial content (inverse of ICC: skull, eyes, skin, background, etc.) GM - gray matter (including CGM and DEEPGM) ICC - intra-cranial cavity (GM, WM, and CSF) NAWM - normal appearing white matter WM - white matter (including both NAWM and WMH) WMH - white matter hyperintensity Sample Case The tutorial describes general strategies for customizing the EMSegment module. However, it also includes the specifics for a sample case. The two input channels for the sample case are MPRAGE and FLAIR. Different channels may be used, but the strategies described here are specific to the aforementioned two channels. All the necessary files for the sample case are included in EMSegmentTutorial_v1.zip and can be download from Instructions and parameter settings for the sample case appear in this textbox style throughout the tutorial. Note: The parameters used for the sample case produce good results; however, these might not be the best results. Sample Case Files The EMSegmentTutorial_v1.zip contains the following files: Subject Files: o subject001_mprage.nii.gz - MPRAGE (176x256 in plane resolution, 176 slices, 1x1x1mm 3 voxel) o subject001_rflair.nii.gz - FLAIR registered to MPRAGE Atlas Files (registered to MPRAGE): o subject001_icbmcsf.nii.gz - probabilistic atlas of the CSF o subject001_icbmdeepgm.nii.gz - probabilistic atlas of the deep GM structures (caudate, putamen, thalamus) o subject001_icbmgray.nii.gz - probabilistic atlas of the GM o subject001_icbmwhite.nii.gz - probabilistic atlas of the WM Label Maps: o subject001_ecc.nii.gz - ECC label map (inverse of ICC) XML Files: o subject001_scene01_images.xml - scene file containing all the images o subject001_scene01.xml - scene file with all the images and settings for GM, WM, and CSF segmentation Results: o subject001_scene01_seg.nii.gz Atlas The tutorial uses the ICBM probabilistic atlases (available at The atlases are registered to the MPRAGE image using the FMRIB Linear Registration Tool (FLIRT, applied with 9 degrees of freedom. ECC The ECC label map was generated by inverting the ICC label map. The ICC was automatically generated from the subject s T2 image and then manually corrected by an expert user. 3
4 File Formats All images and label maps are in NifTI-1 format ( Use Slicer s Menu Window -> Data Panel -> Add Volume -> Generic Readers to read NifTI images. Loading Scenes There are some scene files included with the sample case. Opening these files takes care of loading the images, setting the EMSegment parameters, and setting the labels/colors in one easy step. It is recommended to start with these scene files when starting a new segmentation problem and modify the parameters as seen necessary. 4
5 S e g m e n t a t i o n EMSegment combines information from the input channels, and optionally the probabilistic atlases, to create a segmentation label map that delineates the subclasses defined in the class hierarchy. For a more detailed description of the EMSegment algorithm, refer to [1]. 1 GRAY MATTER, WHITE MATTER, AND CSF SEGMENTATION This section describes the process of segmenting the intra-cranial cavity (ICC) into gray matter (GM), white matter (WM), and cerebrospinal fluid (CSF). 1.1 General Strategy 1. Load the subject and the atlas images. 2. Set the EMSegment parameters. 3. Set up the class hierarchy (see Figure 1.1). 4. Build the intensity distribution for each subclass. 5. Run EMSegment. 6. Refine the parameters and the class distributions and rerun EMSegment until satisfactory segmentation is achieved. 7. Save the parameters into a scene file this file can then be used to segment datasets with the same characteristics. Figure 1.1: The class hierarchy for GM, WM, and CSF segmentation (HEAD and ICC are super classes). 1.2 Files Used Subject Files: o subject001_mprage.nii.gz o subject001_rflair.nii.gz Atlas Files (registered to MPRAGE): o subject001_icbmcsf.nii.gz o subject001_icbmgray.nii.gz o subject001_icbmwhite.nii.gz Label Maps: o subject001_ecc.nii.gz 5
6 Figure 1.2: Sample slices from the MPRAGE and FLAIR subject images and the GM, WM, and CSF probabilistic atlases. 1.3 Loading the Subject and Atlas Images Load the input channels (subject images) and the probabilistic atlas images into Slicer. Figure 1.3: Contents of the Current Scene window after loading the images. There are two ways to load the sample images into Slicer: 1. Automatically via Menu Window -> File Menu -> Open Scene: subject001_scene01_images.xml 2. Manually via Menu Window -> Data Panel -> Add Volume: subject001_mprage.nii.gz subject001_rflair.nii.gz subject001_ecc.nii.gz subject001_icbmcsf.nii.gz subject001_icbmgray.nii.gz subject001_icbmwhite.nii.gz Note: if you load subject001_scene01.xml, these and all other parameters for the GM, WM, and CSF segmentation will be set automatically. Refer to section 1.10 for saving the parameters into an XML file. 1.4 Setting the EMSegment Parameters Go to the EMSegment module: Menu Window -> Modules -> Segmentation -> EMSegment or Menu Window -> More -> EMSegment. 6
7 Figure 1.4: Opening the EMSegment module. Overview of the EMSegment Tabs: Help - Brief description of the module. EM - Guides the user through the general steps of the segmentation. This tab can also be used to set up the class hierarchy, although, the Class tab provides access to more advanced setups. Class - Parameters of the class hierarchy. CIM - Class Interaction Matrix (not modified in this tutorial). Setting - Parameters of the EM algorithm. Adding Input Channels (EM -> Step1 Tab) The intensity information from the input channels, combined with the optional probability atlases, is used to classify the image pixels into different classes. Only two channels are used in this tutorial; however, any number of channels can be used. To add the input channels go to the EM panel -> Step1 tab. The Volume List window shows the available loaded volumes. Figure 1.5: Adding MPRAGE and FLAIR as input channels. Highlight the image to be added in the Volume List window then click on the right arrow to add. Use the left arrow to remove images from the Input Greyscales window. Note: the input channels are referenced by the order they appear in (MPRAGE becomes channel 0, FLAIR becomes channel 1). Add subject001_mprage and subject001_rflair to the Input Greyscales window. MPRAGE should be added first and FLAIR should be added second. Note: Keep in mind the order of the input channels: MPRAGE becomes channel 0, FLAIR becomes channel 1. Tip: Slicer may sometimes become unstable and crash without a warning if certain parameters are not properly set. To avoid retyping the parameter values, periodically save the settings as described in section Setting the Number of Iterations, Class Settings, and Boundary Settings (EM -> Step2 Tab) Parameters: No. of classes - Defines the number of subclasses under the current super class. The classes are hierarchically organized. The top super class, by default, is the Head super class. Iterations - The number of maximum iterations the EM algorithm is to perform. Boundary Min, Max - Define the area of the image to be segmented. Tip: Increasing the number of iterations increases the running time and might not have much affect on the segmentation quality. Set the number of iterations to the lowest that produces good results. 7
8 Figure 1.6: Define general parameters. Set the following parameters: No. of Classes: 2 Iterations: 4 Boundary Min: 1, 1, 1 Boundary Max: 176, 256, Setting Up the Class Hierarchy (Class Tab) The class hierarchy determines the relationship between the different classes. The structure of the class hierarchy and the parameters of each class can be changed under the Class tab. See Figure 1.1 for a general overview of the hierarchy. Head Super Class The current class can be selected with the top button of the Class tab. First, choose the Head super class: Figure 1.7: Choosing the current class. Click on the top button in the class tab and select the desired class from the pull-down menu. Figure 1.8: Stop Parameters for the Super Class structure. Description of parameters for the Super Class structure: Name - Name of the super class. Class Probability - Expected proportion of the classes at the given level. The class probabilities for a level have to add up to 1. (e.g. Figure 1, level 3: GM: 0.6; WM: 0.3; CSF: 0.1). Number of Classes - Number of subclasses. Prob Data Weight - Weight of the probability atlas. Input Channel Weights - Weights of the input channels. Print Parameters - Intermediate results, such as the LabelMap, can be printed out at each iteration. Frequency defines how often these results will be printed out (-1 = only the last iteration; 0 = never; >0 = at each iteration). Stop Parameters o EMType - Defines after which criteria should the algorithm halt. 0 = fixed number of iterations (defined by EMMaxIter); 1 = when label map converges measure (calculate difference measure by comparing labelmaps between iterations); 2 = when weights calculated in the E-Step converge measure (calculate difference measure by comparing weights between iterations). o EMValue - Defines the threshold for stopping the algorithm based on the average difference (DifferenceMeasure/Number of Voxels) (e.g. if the algorithm should stop if the difference between 8
9 iterations is below 1%, then set EMValue to 0.01). Note: the algorithm halts if the number of iterations goes beyond EMMaxIter; EMValue is only activated if EMType >0. o EMMaxIter - The number of maximum iterations of the algorithm. o MFAType - Defined as EMType, but for MFA. o MFAValue - Defined as EMValue, but for MFA. o FAMaxIter - Should be MFAMaxIter but tcl displays it wrong. Definition as for EMMaxIter just applied to MFA. o BiasCalculation - At each EM iteration the algorithm not only computes a MFA, but also an inhomogeneity correction. This parameter defines after how many EM iterations should the inhomogeneity correction stop (default = -1, which means never stop). Figure 1.9: Parameters for the Head super class. Set the following parameters: Name: Head Class Probability: 1.0 Number of Classes: 2 Prob Data Weight: 1.0 Input Channel Weights: 1.0, 1.0 Stop Parameters EMMaxIter: 4 FAMaxIter: 2 The rest of the parameters can be left at their default settings. Extra Cranial Content (ECC) Subclass Change the current class to the first subclass of the Head super class. This subclass will represent the ECC. 9
10 Figure 1.10: Parameters for the Sub Class structure. Description of parameters for the Sub Class structure: Mean - Mean intensity value of the sample points. Covariance - Covariance matrix. Prob. - Expected proportion of the classes at the given level. The class probabilities for a level have to add up to 1. (e.g. Figure 1, level 2: ECC: 0.15; ICC: 0.85). Prob. Atlas - The probabilistic atlas to be used for the current subclass. Color/Label - Color and label of the current subclass in the resulting LabelMap. Prob Data Weight - Weight of the probabilistic atlas. Input Channel Weights - Weights of the input channels (channel 0, channel 1, etc.). The order of the channels here is the same as in the Input Greyscales There are two ways of sampling a distribution: Manually: To set the class distribution (Mean and Covariance), click on the Use Sample button (so it is lowered) and pick a few (5-20) samples (that represent the ECC) from the lower image panel of the Viewer window by pressing CTRL-Left Mouse Button over the desired voxels. The Mean and Covariance matrix should change with each sample. [Note: this can also be done from the EM Tab -> Step3 -> StepD panel by clicking on the Manual button and selecting sample points the same way.] Tip: See section 1.6 on how to visualize where the pixel under the cursor falls on the intensity distribution graph. Figure 1.11: Lower Image Panel of the Viewer window. Automatically: By clicking on the Auto button in the EM Tab -> Step3 -> StepD panel. [Note: not recommended.] 10
11 Figure 1.12: Parameters for the ECC subclass. Set the following parameters: Mean: , Covariance: , , Prob: 0.15 Prob Atlas: subject001_ecc Color/Label: Black=0 Prob Data Weight: 1 Input Channel Weight: 0.01, 0.01 Note: The first channel (channel 0) is the MPRAGE, the second (channel 1) is the FLAIR. Note: To exclude (set to label 0) all of the ECC, we want to put maximum weight on the ECC label map and little weight on the input channels (0 weight will give an error, thus we set it to 0.01). To set the label and color of the class click on the Color/Label button. A pop-up window will appear, where you can pick a color/label or define your own. Figure 1.13: Select a Color window with the predefined colors for the sample case. The default colors and labels are different. 11
12 The colors for the sample case are the following: Color = Gray Label = 4 RGB = Color = White Label = 8 RGB = Color = Csf Label = 5 RGB = Color = Black Label = 0 RGB = Note: If you loaded the scene file subject001_scene01.xml, these colors will be predefined. ICC Super Class Change the current class to the second subclass. Click on the Super Class button (so it is lowered) to make it a super class. This super class will include the GM, WM, and CSF subclasses. For a description of the parameters refer back to the Head super class section. Figure 1.14: Parameters for the ICC super class. Set the following parameters: Name: ICC Class Probability: 0.85 Number of Classes: 3 Prob Data Weight: 1.0 Input Channel Weights: 1.0, 1.0 Stop Parameters EMMaxIter: 4 FAMaxIter: 2 GM Subclass Change the current class to the first subclass under ICC. Using the method described in the ECC section, pick sample voxels to set the Mean and Covariance. 12
13 Figure 1.15: Parameters for the GM subclass. Set the following parameters: Mean: , Covariance: , , Prob: 0.6 Prob Atlas: subject001_icbmgray Color/Label: Gray=4 Prob Data Weight: 0.8 Input Channel Weight: 0.7, 0.3 WM Subclass Change the current class to the second subclass under ICC. Using the method described in the ECC section, pick sample voxels to set the Mean and Covariance. Figure 1.16: Parameters for the WM subclass. Set the following parameters: Mean: , Covariance: , , Prob: 0.3 Prob Atlas: subject001_icbmwhite Color/Label: White=8 Prob Data Weight: 0.08 Input Channel Weight: 0.95, 0.05 CSF Subclass Change the current class to the third subclass under ICC. Using the method described in the ECC section, pick sample voxels to set the Mean and Covariance. 13
14 Figure 1.17: Parameters for the CSF subclass. Set the following parameters: Mean: , Covariance: , , Prob: 0.1 Prob Atlas: subject001_icbmcsf Color/Label: Csf=5 Prob Data Weight: 0.2 Input Channel Weight: 1.0, 1.0 Subclass Probabilities The probabilities of subclasses within a super class should add up to 1. To get an overview of the subclasses of a certain super class, click on the Class Overview button at the bottom of the Class tab. A pop-up window will display some of the subclass parameters. These parameters can also be changed in this window. Figure 1.18: Class Overview window for the ICC super class. 1.6 Class Distributions The class distributions of all the subclasses can be displayed by clicking on the Class Distribution button at the bottom of the Class tab. In the pop-up window click on the buttons on the top and bottom left corners to set the input channels and click on the labeled buttons to display the distributions. Figure 1.19: Setting the Input Channels in the Class Distribution window. Click on the top left and bottom left buttons to set the input channels. 14
15 Figure 1.20: Class Distributions for the Sample Case. To change the graph settings, such as the range of the axis click on the graph and change the settings in the pop-up window. Tip: When picking the sample points for the distributions, a red line indicates in the Display Class Distribution window where the value of the pixel under the cursor falls. This can be a useful indicator of how the distribution would be affected by that pixel value. 1.7 EM Algorithm Settings Figure 1.21: Setting panel with default values. Description of parameters for the Setting tab: EM-Iterations - Same as EMMaxIter (obsolete field). Alpha - Influence of MFA on the definition of weights [0 = MFA is deactivated (good for high signal-to-noise ratio); >0 = importance of MFA, 1 = maximum importance (good for noisy images). Smoothing Width - When computing image inhomogeneity, a Gaussian filter is used to guarantee smoothness constraints of the field. This parameter determines the size of the filter (e.g. width = 20 means a 20x20x20 Gaussian filter). Smoothing Sigma - The variance of the filter. If sigma is large, the filter is similar to a box filter. If sigma is small, the filter relates to a median filter. Multi Threading - Use multiple threads. Printing Directory - The directory where intermediate results, as activated by Print Parameters, should be printed out. 15
16 Use Probability - 1 = incorporate the global probabilities (Prob.) when plotting distributions in the Class Distribution window; 0 = plot the distributions only considering mean and covariance. Training Samples - Maximum value of the atlas images. Intensity Class - Deactivated field. Run Remotely - Run on a remote machine. Generate Models - Generate 3D models from the subclasses. Set the following parameters: EM-Iterations: 4 Alpha: 0.3 Smoothing Width: 19 Smoothing Sigma: 9 Multi threading: On Use Probability: On Training Samples: 100 Intensity Class: None 1.8 Running EMSegment Once all the necessary parameters are set, run EMSegment by clicking on the Run button under the EM tab. Figure 1.22: Running EMSegment. Click on the Run button in the EM tab. The progress of the segmentation can be followed in the terminal window of Slicer. Figure 1.23: Slicer Terminal window messages. Upon successful segmentation the following message is displayed in the EM tab. Figure 1.24: Completion message. 16
17 1.9 Results Figure 1.25: Segmentation results MPRAGE, FLAIR, LabelMap and MPRAGE overlay, LabelMap with ICC outlined (white line). Although, the current settings provide good results the following problems are apparent: Deep gray matter structures, such as the caudate and the thalamus are not properly segmented. CSF is slightly underestimated. There is brain matter outside of the ICC (white line). Note: We can only define ECC as a probabilistic atlas and not as a mask. Therefore, some parts of the ECC are still classified as brain matter. These extra pixels have to be masked out with the ECC. Refer to the Slicer website on masking. In the next chapter we describe how to add a deep gray matter subclass to improve the segmentation of the caudate and the thalamus. 17
18 Ways to improve the segmentation: Depending on the weight of the probabilistic atlases, the class distributions have the largest impact on the outcome of the segmentation. If one of the classes is way over or underestimated, changing the class distributions should be first step. A particular distribution can be broadened, for example, by picking additional sample points that lie towards the edge of the distribution. Use the Display Class Distribution window (section 1.6) to see where the current pixel under the cursor falls and how the distributions look like. The weights of the input channels also have a large impact. Put more weights on the input channel that has the largest contrast for the particular subclass. Changing the relative probabilities of the subclasses within a super class has little effect, thus these probabilities should be used for fine-tuning Saving the Results and EMSegment Parameters See the Slicer tutorial on saving the segmentation results. To save the EMSegment parameters, go to the Setting tab and click on the Save Setting button. The XML file can then be reloaded to restore the saved parameters (Menu Window -> File Menu -> Open Scene). 2 ADDING THE DEEP GRAY MATTER SUBCLASS In this section we ll describe the strategy to further refine the segmentation by introducing the Deep Gray Matter subclass. Deep gray matter structures (caudate, putamen, thalamus) have different intensity distributions than cortical gray matter. These differences can be modeled by further dividing GM into subclasses and building unique distributions for each of these classes. Note: This section gives only an overview of the general strategy and it is up to the reader to refine the segmentation; therefore, we do not provide specific parameters. 2.1 General Strategy 1. Continue from the previous segmentation. 2. Set up the class hierarchy (add GM super class, and CGM and DEEPGM subclasses, see Figure 2.1). 3. Build the intensity distribution for the new subclasses. 4. Run EMSegment. 5. Refine the parameters and the class distributions and rerun EMSegment until satisfactory segmentation is achieved. 6. Save the parameters into a scene file. Figure 2.1: Class hierarchy for CGM, DEEPGM, WM, and CSF segmentation (HEAD, ICC, and GM are super classes). 2.2 Files Used Subject Files: o subject001_mprage.nii.gz o subject001_rflair.nii.gz Atlas Files (registered to MPRAGE): o subject001_icbmcsf.nii.gz o subject001_icbmgray.nii.gz o subject001_icbmdeepgm.nii.gz o subject001_icbmwhite.nii.gz 18
19 Label Maps: o subject001_ecc.nii.gz Figure 2.2: Sample slices from the MPRAGE and FLAIR subject images and the GM, WM, CSF, and DEEPGM probabilistic atlases. 2.3 Loading the DEEP GM Atlas Image Load the DEEP GM probabilistic atlas image into Slicer. Manually via Data Panel -> Add Volume: subject001_icbmdeepgm.nii.gz 2.4 Modifying the Class Hierarchy (Class Tab) GM Super Class Change the current class to the GM subclass. Click on the Super Class button (so it is lowered) to make it a super class. This super class will include the CGM and DEEPGM subclasses. For a description of the parameters refer back to the Head super class under section
20 Figure 2.3: Parameters for the GM super class. CGM Subclass Change the current class to the first subclass under ICC. Using the method described in the ECC section, pick sample voxels to set the Mean and Covariance. Figure 2.4: Parameters for the CGM subclass. DEEPGM Subclass Change the current class to the first subclass under ICC. Using the method described in the ECC section, pick sample voxels to set the Mean and Covariance. 20
21 Figure 2.5: Setting the parameters for the DEEPGM subclass. 2.5 Running EMSegment Run EMSegment by clicking on the Run button under the EM tab. Refine the parameters until the segmentation quality is satisfactory. 3 ADDING THE WHITE MATTER HYPERINTENSITY SEGMENTATION In this section we ll further refine the segmentation by dividing the White Matter into Normal Appearing White Matter (NAWM) and White Matter Hyperintensities (WMH). WMH is segmented in a separate step and then the results are combined with the previous segmentation. Note: This section gives only an overview of the general strategy and it is up to the reader to refine the segmentation; therefore, we do not provide specific parameters. 3.1 General Strategy Adding in the NAWM and WMH subclasses under the hierarchy used so far does not produce satisfactory results. Segmenting WMH and GM together in the same segmentation step tends to result in a considerable amount of GM misclassifications as WMH. To circumvent such misclassifications, the following strategy can be used: 1. Load the FLAIR image. 2. Set up the class hierarchy for the separate WM segmentation (WM super class, and NAWM and WMH subclasses, see Figure 3.1). 3. Build the intensity distribution for the new subclasses. 4. Run EMSegment. 5. Refine the parameters and the class distributions and rerun EMSegment until satisfactory segmentation is achieved. 6. Merge the WMH segmentation and the previous result to get the final segmentation that includes all tissue types. Note: This strategy works well for the sample dataset used; however, it might not be suitable for different datasets. It is used to demonstrate an approach that might lead to satisfactory results. 21
22 Figure 3.1: Class hierarchy for NAWM and WMH segmentation (SEGMENTATION 2) and merging the results with the previous segmentation. 22
23 R e f e r e n c e s 1. K.M. Pohl, S. Bouix, R. Kikinis, W.E.L. Grimson, Anatomical Guided Segmentation with Non-Stationary Tissue Class Distributions in an Expectation-Maximization Framework. In Proc. ISBI 2004: IEEE International Symposium on Biomedical Imaging: From Nano to Macro, Arlington, VA, USA, pp ,
EMSegmenter Tutorial (Advanced Mode)
 EMSegmenter Tutorial (Advanced Mode) Dominique Belhachemi Section of Biomedical Image Analysis Department of Radiology University of Pennsylvania 1/65 Overview The goal of this tutorial is to apply the
EMSegmenter Tutorial (Advanced Mode) Dominique Belhachemi Section of Biomedical Image Analysis Department of Radiology University of Pennsylvania 1/65 Overview The goal of this tutorial is to apply the
Slicer3 Training Tutorial Using EM Segmenter with Non- Human Primate Images
 Slicer3 Training Compendium Slicer3 Training Tutorial Using EM Segmenter with Non- Human Primate Images Vidya Rajagopalan Christopher Wyatt BioImaging Systems Lab Dept. of Electrical Engineering Virginia
Slicer3 Training Compendium Slicer3 Training Tutorial Using EM Segmenter with Non- Human Primate Images Vidya Rajagopalan Christopher Wyatt BioImaging Systems Lab Dept. of Electrical Engineering Virginia
An Introduction To Automatic Tissue Classification Of Brain MRI. Colm Elliott Mar 2014
 An Introduction To Automatic Tissue Classification Of Brain MRI Colm Elliott Mar 2014 Tissue Classification Tissue classification is part of many processing pipelines. We often want to classify each voxel
An Introduction To Automatic Tissue Classification Of Brain MRI Colm Elliott Mar 2014 Tissue Classification Tissue classification is part of many processing pipelines. We often want to classify each voxel
3D Visualization of FreeSurfer Data
 3D Visualization of FreeSurfer Data Sonia Pujol, Ph.D. Silas Mann, B.Sc. Randy Gollub, MD., Ph.D. Surgical Planning Laboratory Athinoula A. Martinos Center Harvard University Acknowledgements NIH U54EB005149
3D Visualization of FreeSurfer Data Sonia Pujol, Ph.D. Silas Mann, B.Sc. Randy Gollub, MD., Ph.D. Surgical Planning Laboratory Athinoula A. Martinos Center Harvard University Acknowledgements NIH U54EB005149
Norbert Schuff VA Medical Center and UCSF
 Norbert Schuff Medical Center and UCSF Norbert.schuff@ucsf.edu Medical Imaging Informatics N.Schuff Course # 170.03 Slide 1/67 Objective Learn the principle segmentation techniques Understand the role
Norbert Schuff Medical Center and UCSF Norbert.schuff@ucsf.edu Medical Imaging Informatics N.Schuff Course # 170.03 Slide 1/67 Objective Learn the principle segmentation techniques Understand the role
ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION
 ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION Abstract: MIP Project Report Spring 2013 Gaurav Mittal 201232644 This is a detailed report about the course project, which was to implement
ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION Abstract: MIP Project Report Spring 2013 Gaurav Mittal 201232644 This is a detailed report about the course project, which was to implement
Atlas Registration & Label Merging
 NA-MIC Slicer3 Tutorial Atlas Registration & Label Merging Dominik Meier, Ron Kikinis February 2010 Overview 1. Introduction 2. Prerequisites 3. Modules Used takes how long to do? 4. Loading Example Dataset
NA-MIC Slicer3 Tutorial Atlas Registration & Label Merging Dominik Meier, Ron Kikinis February 2010 Overview 1. Introduction 2. Prerequisites 3. Modules Used takes how long to do? 4. Loading Example Dataset
Stereoscopic Atlas of Intrinsic Brain Networks (SAIBN)
 Stereoscopic Atlas of Intrinsic Brain Networks (SAIBN) Gonzalo M. Rojas, Marcelo Gálvez, Daniel Margulies, Cameron Craddock, Xavier Castellanos, Michael Milham 1.- Introduction Stereoscopic Atlas of Intrinsic
Stereoscopic Atlas of Intrinsic Brain Networks (SAIBN) Gonzalo M. Rojas, Marcelo Gálvez, Daniel Margulies, Cameron Craddock, Xavier Castellanos, Michael Milham 1.- Introduction Stereoscopic Atlas of Intrinsic
MRI Segmentation MIDAS, 2007, 2010
 MRI Segmentation MIDAS, 2007, 2010 Lawrence O. Hall, Dmitry Goldgof, Yuhua Gu, Prodip Hore Dept. of Computer Science & Engineering University of South Florida CONTENTS: 1. Introduction... 1 2. Installing
MRI Segmentation MIDAS, 2007, 2010 Lawrence O. Hall, Dmitry Goldgof, Yuhua Gu, Prodip Hore Dept. of Computer Science & Engineering University of South Florida CONTENTS: 1. Introduction... 1 2. Installing
Data Loading & 3D Visualization
 Neuroimage Analysis Center Data Loading & 3D Visualization Sonia Pujol, Ph.D. Surgical Planning Laboratory Harvard Medical School Leonardo da Vinci (1452-1519), Virgin and Child Alte Pinakothek, München
Neuroimage Analysis Center Data Loading & 3D Visualization Sonia Pujol, Ph.D. Surgical Planning Laboratory Harvard Medical School Leonardo da Vinci (1452-1519), Virgin and Child Alte Pinakothek, München
Introduction. Welcome to 3D Slicer! Surgical Planning Laboratory -1- Brigham and Women s Hospital
 Introduction Welcome to 3D Slicer! -1- Overview of Training What is Slicer? Uses of Slicer Getting Slicer How to Use Slicer Loading Data Viewing Data Modifying Data Saving Data -2- What is Slicer? -3-
Introduction Welcome to 3D Slicer! -1- Overview of Training What is Slicer? Uses of Slicer Getting Slicer How to Use Slicer Loading Data Viewing Data Modifying Data Saving Data -2- What is Slicer? -3-
Histograms. h(r k ) = n k. p(r k )= n k /NM. Histogram: number of times intensity level rk appears in the image
 Histograms h(r k ) = n k Histogram: number of times intensity level rk appears in the image p(r k )= n k /NM normalized histogram also a probability of occurence 1 Histogram of Image Intensities Create
Histograms h(r k ) = n k Histogram: number of times intensity level rk appears in the image p(r k )= n k /NM normalized histogram also a probability of occurence 1 Histogram of Image Intensities Create
Slicer4Minute Tutorial. Sonia Pujol, Ph.D. Surgical Planning Laboratory Harvard Medical School
 Slicer4Minute Tutorial Sonia Pujol, Ph.D. Surgical Planning Laboratory Harvard Medical School 1 Slicer4 minute tutorial This tutorial is a 4-minute introduction to the 3D visualization capabilities of
Slicer4Minute Tutorial Sonia Pujol, Ph.D. Surgical Planning Laboratory Harvard Medical School 1 Slicer4 minute tutorial This tutorial is a 4-minute introduction to the 3D visualization capabilities of
Comparison Study of Clinical 3D MRI Brain Segmentation Evaluation
 Comparison Study of Clinical 3D MRI Brain Segmentation Evaluation Ting Song 1, Elsa D. Angelini 2, Brett D. Mensh 3, Andrew Laine 1 1 Heffner Biomedical Imaging Laboratory Department of Biomedical Engineering,
Comparison Study of Clinical 3D MRI Brain Segmentation Evaluation Ting Song 1, Elsa D. Angelini 2, Brett D. Mensh 3, Andrew Laine 1 1 Heffner Biomedical Imaging Laboratory Department of Biomedical Engineering,
Norbert Schuff Professor of Radiology VA Medical Center and UCSF
 Norbert Schuff Professor of Radiology Medical Center and UCSF Norbert.schuff@ucsf.edu 2010, N.Schuff Slide 1/67 Overview Definitions Role of Segmentation Segmentation methods Intensity based Shape based
Norbert Schuff Professor of Radiology Medical Center and UCSF Norbert.schuff@ucsf.edu 2010, N.Schuff Slide 1/67 Overview Definitions Role of Segmentation Segmentation methods Intensity based Shape based
Automatic Registration-Based Segmentation for Neonatal Brains Using ANTs and Atropos
 Automatic Registration-Based Segmentation for Neonatal Brains Using ANTs and Atropos Jue Wu and Brian Avants Penn Image Computing and Science Lab, University of Pennsylvania, Philadelphia, USA Abstract.
Automatic Registration-Based Segmentation for Neonatal Brains Using ANTs and Atropos Jue Wu and Brian Avants Penn Image Computing and Science Lab, University of Pennsylvania, Philadelphia, USA Abstract.
Ensemble registration: Combining groupwise registration and segmentation
 PURWANI, COOTES, TWINING: ENSEMBLE REGISTRATION 1 Ensemble registration: Combining groupwise registration and segmentation Sri Purwani 1,2 sri.purwani@postgrad.manchester.ac.uk Tim Cootes 1 t.cootes@manchester.ac.uk
PURWANI, COOTES, TWINING: ENSEMBLE REGISTRATION 1 Ensemble registration: Combining groupwise registration and segmentation Sri Purwani 1,2 sri.purwani@postgrad.manchester.ac.uk Tim Cootes 1 t.cootes@manchester.ac.uk
Tutorial BOLD Module
 m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial BOLD Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial BOLD Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
Adaptive thresholding. in CTAn
 Adaptive thresholding in CTAn Method note Page 1 of 8 2 Bruker-MicroCT method note:adaptive thresholding in CTAn Introduction 2D or 3D morphometric analysis of micro-ct images always requires binarization
Adaptive thresholding in CTAn Method note Page 1 of 8 2 Bruker-MicroCT method note:adaptive thresholding in CTAn Introduction 2D or 3D morphometric analysis of micro-ct images always requires binarization
mritc: A Package for MRI Tissue Classification
 mritc: A Package for MRI Tissue Classification Dai Feng 1 Luke Tierney 2 1 Merck Research Labratories 2 University of Iowa July 2010 Feng & Tierney (Merck & U of Iowa) MRI Tissue Classification July 2010
mritc: A Package for MRI Tissue Classification Dai Feng 1 Luke Tierney 2 1 Merck Research Labratories 2 University of Iowa July 2010 Feng & Tierney (Merck & U of Iowa) MRI Tissue Classification July 2010
MR IMAGE SEGMENTATION
 MR IMAGE SEGMENTATION Prepared by : Monil Shah What is Segmentation? Partitioning a region or regions of interest in images such that each region corresponds to one or more anatomic structures Classification
MR IMAGE SEGMENTATION Prepared by : Monil Shah What is Segmentation? Partitioning a region or regions of interest in images such that each region corresponds to one or more anatomic structures Classification
Performance Evaluation of the TINA Medical Image Segmentation Algorithm on Brainweb Simulated Images
 Tina Memo No. 2008-003 Internal Memo Performance Evaluation of the TINA Medical Image Segmentation Algorithm on Brainweb Simulated Images P. A. Bromiley Last updated 20 / 12 / 2007 Imaging Science and
Tina Memo No. 2008-003 Internal Memo Performance Evaluation of the TINA Medical Image Segmentation Algorithm on Brainweb Simulated Images P. A. Bromiley Last updated 20 / 12 / 2007 Imaging Science and
NA-MIC National Alliance for Medical Image Computing fmri Data Analysis
 NA-MIC fmri Data Analysis Sonia Pujol, Ph.D. Wendy Plesniak, Ph.D. Randy Gollub, M.D., Ph.D. Acknowledgments NIH U54EB005149 Neuroimage Analysis Center NIH P41RR013218 FIRST Biomedical Informatics Research
NA-MIC fmri Data Analysis Sonia Pujol, Ph.D. Wendy Plesniak, Ph.D. Randy Gollub, M.D., Ph.D. Acknowledgments NIH U54EB005149 Neuroimage Analysis Center NIH P41RR013218 FIRST Biomedical Informatics Research
Detecting White Matter Lesions in Lupus
 Slicer3 Training Compendium Detecting White Matter Lesions in Lupus Version 2.0 1/6/2009 H. Jeremy Bockholt Mark Scully -1- Learning objective Following this tutorial, you ll be able to load scans into
Slicer3 Training Compendium Detecting White Matter Lesions in Lupus Version 2.0 1/6/2009 H. Jeremy Bockholt Mark Scully -1- Learning objective Following this tutorial, you ll be able to load scans into
FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING. Francesca Pizzorni Ferrarese
 FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING Francesca Pizzorni Ferrarese Pipeline overview WM and GM Segmentation Registration Data reconstruction Tractography
FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING Francesca Pizzorni Ferrarese Pipeline overview WM and GM Segmentation Registration Data reconstruction Tractography
This exercise uses one anatomical data set (ANAT1) and two functional data sets (FUNC1 and FUNC2).
 Exploring Brain Anatomy This week s exercises will let you explore the anatomical organization of the brain to learn some of its basic properties, as well as the location of different structures. The human
Exploring Brain Anatomy This week s exercises will let you explore the anatomical organization of the brain to learn some of its basic properties, as well as the location of different structures. The human
Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge
 Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge Christian Wasserthal 1, Karin Engel 1, Karsten Rink 1 und André Brechmann
Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge Christian Wasserthal 1, Karin Engel 1, Karsten Rink 1 und André Brechmann
Slicer3 minute tutorial
 Slicer3 minute tutorial Sonia Pujol, Ph.D. Surgical Planning Laboratory Harvard Medical School -1- Slicer3 minute tutorial This tutorial is a short introduction to the advanced 3D visualization capabilities
Slicer3 minute tutorial Sonia Pujol, Ph.D. Surgical Planning Laboratory Harvard Medical School -1- Slicer3 minute tutorial This tutorial is a short introduction to the advanced 3D visualization capabilities
Automatic Optimization of Segmentation Algorithms Through Simultaneous Truth and Performance Level Estimation (STAPLE)
 Automatic Optimization of Segmentation Algorithms Through Simultaneous Truth and Performance Level Estimation (STAPLE) Mahnaz Maddah, Kelly H. Zou, William M. Wells, Ron Kikinis, and Simon K. Warfield
Automatic Optimization of Segmentation Algorithms Through Simultaneous Truth and Performance Level Estimation (STAPLE) Mahnaz Maddah, Kelly H. Zou, William M. Wells, Ron Kikinis, and Simon K. Warfield
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit
 Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
Correction of Partial Volume Effects in Arterial Spin Labeling MRI
 Correction of Partial Volume Effects in Arterial Spin Labeling MRI By: Tracy Ssali Supervisors: Dr. Keith St. Lawrence and Udunna Anazodo Medical Biophysics 3970Z Six Week Project April 13 th 2012 Introduction
Correction of Partial Volume Effects in Arterial Spin Labeling MRI By: Tracy Ssali Supervisors: Dr. Keith St. Lawrence and Udunna Anazodo Medical Biophysics 3970Z Six Week Project April 13 th 2012 Introduction
Multiple Sclerosis Brain MRI Segmentation Workflow deployment on the EGEE grid
 Multiple Sclerosis Brain MRI Segmentation Workflow deployment on the EGEE grid Erik Pernod 1, Jean-Christophe Souplet 1, Javier Rojas Balderrama 2, Diane Lingrand 2, Xavier Pennec 1 Speaker: Grégoire Malandain
Multiple Sclerosis Brain MRI Segmentation Workflow deployment on the EGEE grid Erik Pernod 1, Jean-Christophe Souplet 1, Javier Rojas Balderrama 2, Diane Lingrand 2, Xavier Pennec 1 Speaker: Grégoire Malandain
Learn Image Segmentation Basics with Hands-on Introduction to ITK-SNAP. RSNA 2016 Courses RCB22 and RCB54
 Learn Image Segmentation Basics with Hands-on Introduction to ITK-SNAP RSNA 2016 Courses RCB22 and RCB54 RCB22 Mon, Nov 28 10:30-12:00 PM, Room S401CD RCB54 Thu, Dec 1 2:30-4:30 PM, Room S401CD Presenters:
Learn Image Segmentation Basics with Hands-on Introduction to ITK-SNAP RSNA 2016 Courses RCB22 and RCB54 RCB22 Mon, Nov 28 10:30-12:00 PM, Room S401CD RCB54 Thu, Dec 1 2:30-4:30 PM, Room S401CD Presenters:
PRANA. Project Review and Analysis. March (Pre-release and still under development!)
 PRANA Project Review and Analysis March 2010- (Pre-release and still under development!) A.A. Maudsley Contents: PRANA... 1 1. Introduction... 2 1.1. Examples... 2 2. Subject/Study Selection and Filter...
PRANA Project Review and Analysis March 2010- (Pre-release and still under development!) A.A. Maudsley Contents: PRANA... 1 1. Introduction... 2 1.1. Examples... 2 2. Subject/Study Selection and Filter...
FSL Workshop Session 3 David Smith & John Clithero
 FSL Workshop 12.09.08 Session 3 David Smith & John Clithero What is MELODIC? Probabilistic ICA Improves upon standard ICA Allows for inference Avoids over-fitting Three stage process ( ppca ) 1.) Dimension
FSL Workshop 12.09.08 Session 3 David Smith & John Clithero What is MELODIC? Probabilistic ICA Improves upon standard ICA Allows for inference Avoids over-fitting Three stage process ( ppca ) 1.) Dimension
Segmenting Glioma in Multi-Modal Images using a Generative Model for Brain Lesion Segmentation
 Segmenting Glioma in Multi-Modal Images using a Generative Model for Brain Lesion Segmentation Bjoern H. Menze 1,2, Koen Van Leemput 3, Danial Lashkari 4 Marc-André Weber 5, Nicholas Ayache 2, and Polina
Segmenting Glioma in Multi-Modal Images using a Generative Model for Brain Lesion Segmentation Bjoern H. Menze 1,2, Koen Van Leemput 3, Danial Lashkari 4 Marc-André Weber 5, Nicholas Ayache 2, and Polina
NIH Public Access Author Manuscript Proc Soc Photo Opt Instrum Eng. Author manuscript; available in PMC 2014 October 07.
 NIH Public Access Author Manuscript Published in final edited form as: Proc Soc Photo Opt Instrum Eng. 2014 March 21; 9034: 903442. doi:10.1117/12.2042915. MRI Brain Tumor Segmentation and Necrosis Detection
NIH Public Access Author Manuscript Published in final edited form as: Proc Soc Photo Opt Instrum Eng. 2014 March 21; 9034: 903442. doi:10.1117/12.2042915. MRI Brain Tumor Segmentation and Necrosis Detection
ASAP_2.0 (Automatic Software for ASL Processing) USER S MANUAL
 ASAP_2.0 (Automatic Software for ASL Processing) USER S MANUAL ASAP was developed as part of the COST Action "Arterial Spin Labelling Initiative in Dementia (AID)" by: Department of Neuroimaging, Institute
ASAP_2.0 (Automatic Software for ASL Processing) USER S MANUAL ASAP was developed as part of the COST Action "Arterial Spin Labelling Initiative in Dementia (AID)" by: Department of Neuroimaging, Institute
An ITK Filter for Bayesian Segmentation: itkbayesianclassifierimagefilter
 An ITK Filter for Bayesian Segmentation: itkbayesianclassifierimagefilter John Melonakos 1, Karthik Krishnan 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332, USA {jmelonak,
An ITK Filter for Bayesian Segmentation: itkbayesianclassifierimagefilter John Melonakos 1, Karthik Krishnan 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332, USA {jmelonak,
White Matter Lesion Segmentation (WMLS) Manual
 White Matter Lesion Segmentation (WMLS) Manual 1. Introduction White matter lesions (WMLs) are brain abnormalities that appear in different brain diseases, such as multiple sclerosis (MS), head injury,
White Matter Lesion Segmentation (WMLS) Manual 1. Introduction White matter lesions (WMLs) are brain abnormalities that appear in different brain diseases, such as multiple sclerosis (MS), head injury,
Slicer3 Minute Tutorial
 Slicer3 Minute Tutorial Surgical Planning Laboratory Harvard Medical School Sonia Pujol, PhD Slicer3 Minute Tutorial This tutorial is a short introduction to the advanced 3D visualization capabilities
Slicer3 Minute Tutorial Surgical Planning Laboratory Harvard Medical School Sonia Pujol, PhD Slicer3 Minute Tutorial This tutorial is a short introduction to the advanced 3D visualization capabilities
Norbert Schuff Professor of Radiology VA Medical Center and UCSF
 Norbert Schuff Professor of Radiology Medical Center and UCSF Norbert.schuff@ucsf.edu Slide 1/67 Overview Definitions Role of Segmentation Segmentation methods Intensity based Shape based Texture based
Norbert Schuff Professor of Radiology Medical Center and UCSF Norbert.schuff@ucsf.edu Slide 1/67 Overview Definitions Role of Segmentation Segmentation methods Intensity based Shape based Texture based
MATERIALS PLUS Segmentation Measurement
 Example: Segmentation MATERIALS PLUS Segmentation is a method of image partitioning based on the intensity / gray scale range of its components. Since a phase is detected and its area is estimated on the
Example: Segmentation MATERIALS PLUS Segmentation is a method of image partitioning based on the intensity / gray scale range of its components. Since a phase is detected and its area is estimated on the
GLM for fmri data analysis Lab Exercise 1
 GLM for fmri data analysis Lab Exercise 1 March 15, 2013 Medical Image Processing Lab Medical Image Processing Lab GLM for fmri data analysis Outline 1 Getting Started 2 AUDIO 1 st level Preprocessing
GLM for fmri data analysis Lab Exercise 1 March 15, 2013 Medical Image Processing Lab Medical Image Processing Lab GLM for fmri data analysis Outline 1 Getting Started 2 AUDIO 1 st level Preprocessing
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit
 Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
Elastically Deforming a Three-Dimensional Atlas to Match Anatomical Brain Images
 University of Pennsylvania ScholarlyCommons Technical Reports (CIS) Department of Computer & Information Science May 1993 Elastically Deforming a Three-Dimensional Atlas to Match Anatomical Brain Images
University of Pennsylvania ScholarlyCommons Technical Reports (CIS) Department of Computer & Information Science May 1993 Elastically Deforming a Three-Dimensional Atlas to Match Anatomical Brain Images
EPILOG PREOP. Viewer Interpretation Guideline
 EPILOG PREOP Viewer Interpretation Guideline Starting the viewer Launch the viewer for a patient by clicking this icon. A new browser tab or window will open. Data loading To ensure smooth switching between
EPILOG PREOP Viewer Interpretation Guideline Starting the viewer Launch the viewer for a patient by clicking this icon. A new browser tab or window will open. Data loading To ensure smooth switching between
Automated MR Image Analysis Pipelines
 Automated MR Image Analysis Pipelines Andy Simmons Centre for Neuroimaging Sciences, Kings College London Institute of Psychiatry. NIHR Biomedical Research Centre for Mental Health at IoP & SLAM. Neuroimaging
Automated MR Image Analysis Pipelines Andy Simmons Centre for Neuroimaging Sciences, Kings College London Institute of Psychiatry. NIHR Biomedical Research Centre for Mental Health at IoP & SLAM. Neuroimaging
n o r d i c B r a i n E x Tutorial DTI Module
 m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial DTI Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial DTI Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
Structural Segmentation
 Structural Segmentation FAST tissue-type segmentation FIRST sub-cortical structure segmentation FSL-VBM voxelwise grey-matter density analysis SIENA atrophy analysis FAST FMRIB s Automated Segmentation
Structural Segmentation FAST tissue-type segmentation FIRST sub-cortical structure segmentation FSL-VBM voxelwise grey-matter density analysis SIENA atrophy analysis FAST FMRIB s Automated Segmentation
Segmenting Glioma in Multi-Modal Images using a Generative-Discriminative Model for Brain Lesion Segmentation
 Segmenting Glioma in Multi-Modal Images using a Generative-Discriminative Model for Brain Lesion Segmentation Bjoern H. Menze 1,2, Ezequiel Geremia 2, Nicholas Ayache 2, and Gabor Szekely 1 1 Computer
Segmenting Glioma in Multi-Modal Images using a Generative-Discriminative Model for Brain Lesion Segmentation Bjoern H. Menze 1,2, Ezequiel Geremia 2, Nicholas Ayache 2, and Gabor Szekely 1 1 Computer
Fuzzy Multi-channel Clustering with Individualized Spatial Priors for Segmenting Brain Lesions and Infarcts
 Fuzzy Multi-channel Clustering with Individualized Spatial Priors for Segmenting Brain Lesions and Infarcts Evangelia Zacharaki, Guray Erus, Anastasios Bezerianos, Christos Davatzikos To cite this version:
Fuzzy Multi-channel Clustering with Individualized Spatial Priors for Segmenting Brain Lesions and Infarcts Evangelia Zacharaki, Guray Erus, Anastasios Bezerianos, Christos Davatzikos To cite this version:
3D-Modeling with Microscopy Image Browser (im_browser) (pdf version) Ilya Belevich, Darshan Kumar, Helena Vihinen
 3D-Modeling with Microscopy Image Browser (im_browser) (pdf version) Ilya Belevich, Darshan Kumar, Helena Vihinen Dataset: Huh7.tif (original), obtained by Serial Block Face-SEM (Gatan 3View) Huh7_crop.tif
3D-Modeling with Microscopy Image Browser (im_browser) (pdf version) Ilya Belevich, Darshan Kumar, Helena Vihinen Dataset: Huh7.tif (original), obtained by Serial Block Face-SEM (Gatan 3View) Huh7_crop.tif
Medical Image Synthesis Methods and Applications
 MR Intensity Scale is Arbitrary This causes problems in most postprocessing methods Inconsistency or algorithm failure 11/5/2015 2 Joint Histogram 1.5 T GE SPGR 3 T Philips MPRAGE 11/5/2015 3 Problem With
MR Intensity Scale is Arbitrary This causes problems in most postprocessing methods Inconsistency or algorithm failure 11/5/2015 2 Joint Histogram 1.5 T GE SPGR 3 T Philips MPRAGE 11/5/2015 3 Problem With
Stroke Quantification Tool (Sonia) Ver User Manual
 Stroke Quantification Tool (Sonia) Ver. 1.0 User Manual English. 12/2016 Rev. 1.0 www.wakeup-stroke.eu 1 Table of Contents 1. Introduction...3 2. Installation...4 3. Data Import...5 4. Registration...7
Stroke Quantification Tool (Sonia) Ver. 1.0 User Manual English. 12/2016 Rev. 1.0 www.wakeup-stroke.eu 1 Table of Contents 1. Introduction...3 2. Installation...4 3. Data Import...5 4. Registration...7
Input tab. Insert the path to the DTI Image that you want to tract.
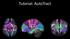 Tutorial: AutoTract Input tab Insert the path to the DTI Image that you want to tract. Input tab (optional)if you already have a WM/CSF mask you want to use to process the tracts, insert the path here.
Tutorial: AutoTract Input tab Insert the path to the DTI Image that you want to tract. Input tab (optional)if you already have a WM/CSF mask you want to use to process the tracts, insert the path here.
Contents... 1 Installation... 3
 Contents Contents... 1 Installation... 3 1 Prerequisites (check for.net framework 3.5)... 3 Install Doctor Eye... 3 Start Using Doctor Eye... 4 How to create a new user... 4 The Main Window... 4 Open a
Contents Contents... 1 Installation... 3 1 Prerequisites (check for.net framework 3.5)... 3 Install Doctor Eye... 3 Start Using Doctor Eye... 4 How to create a new user... 4 The Main Window... 4 Open a
syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets
 syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets Julien Gervais; Lisa Chuah Siemens Healthcare, Magnetic
syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets Julien Gervais; Lisa Chuah Siemens Healthcare, Magnetic
Structural Segmentation
 Structural Segmentation FAST tissue-type segmentation FIRST sub-cortical structure segmentation FSL-VBM voxelwise grey-matter density analysis SIENA atrophy analysis FAST FMRIB s Automated Segmentation
Structural Segmentation FAST tissue-type segmentation FIRST sub-cortical structure segmentation FSL-VBM voxelwise grey-matter density analysis SIENA atrophy analysis FAST FMRIB s Automated Segmentation
How to...create a Video VBOX Gauge in Inkscape. So you want to create your own gauge? How about a transparent background for those text elements?
 BASIC GAUGE CREATION The Video VBox setup software is capable of using many different image formats for gauge backgrounds, static images, or logos, including Bitmaps, JPEGs, or PNG s. When the software
BASIC GAUGE CREATION The Video VBox setup software is capable of using many different image formats for gauge backgrounds, static images, or logos, including Bitmaps, JPEGs, or PNG s. When the software
UGviewer: a medical image viewer
 Appendix A UGviewer: a medical image viewer As a complement to this master s thesis, an own medical image viewer was programmed. This piece of software lets the user visualize and compare images. Designing
Appendix A UGviewer: a medical image viewer As a complement to this master s thesis, an own medical image viewer was programmed. This piece of software lets the user visualize and compare images. Designing
Saturn User Manual. Rubén Cárdenes. 29th January 2010 Image Processing Laboratory, University of Valladolid. Abstract
 Saturn User Manual Rubén Cárdenes 29th January 2010 Image Processing Laboratory, University of Valladolid Abstract Saturn is a software package for DTI processing and visualization, provided with a graphic
Saturn User Manual Rubén Cárdenes 29th January 2010 Image Processing Laboratory, University of Valladolid Abstract Saturn is a software package for DTI processing and visualization, provided with a graphic
Tissue Tracking: Applications for Brain MRI Classification
 Tissue Tracking: Applications for Brain MRI Classification John Melonakos a and Yi Gao a and Allen Tannenbaum a a Georgia Institute of Technology, 414 Ferst Dr, Atlanta, GA, USA; ABSTRACT Bayesian classification
Tissue Tracking: Applications for Brain MRI Classification John Melonakos a and Yi Gao a and Allen Tannenbaum a a Georgia Institute of Technology, 414 Ferst Dr, Atlanta, GA, USA; ABSTRACT Bayesian classification
HOW TO SETUP AND RUN SHIVA...
 TABLE OF CONTENTS HOW TO SETUP AND RUN SHIVA... 1 REQUIREMENTS... 1 Java... 1 Architecture... 2 LAUNCHING SHIVA... 2 LOADING THE ATLAS... 2 USAGE INSTRUCTIONS... 2 FILES... 2 Image Volumes... 2 Label Indices...
TABLE OF CONTENTS HOW TO SETUP AND RUN SHIVA... 1 REQUIREMENTS... 1 Java... 1 Architecture... 2 LAUNCHING SHIVA... 2 LOADING THE ATLAS... 2 USAGE INSTRUCTIONS... 2 FILES... 2 Image Volumes... 2 Label Indices...
Supplementary methods
 Supplementary methods This section provides additional technical details on the sample, the applied imaging and analysis steps and methods. Structural imaging Trained radiographers placed all participants
Supplementary methods This section provides additional technical details on the sample, the applied imaging and analysis steps and methods. Structural imaging Trained radiographers placed all participants
Group (Level 2) fmri Data Analysis - Lab 4
 Group (Level 2) fmri Data Analysis - Lab 4 Index Goals of this Lab Before Getting Started The Chosen Ten Checking Data Quality Create a Mean Anatomical of the Group Group Analysis: One-Sample T-Test Examine
Group (Level 2) fmri Data Analysis - Lab 4 Index Goals of this Lab Before Getting Started The Chosen Ten Checking Data Quality Create a Mean Anatomical of the Group Group Analysis: One-Sample T-Test Examine
CS 664 Segmentation. Daniel Huttenlocher
 CS 664 Segmentation Daniel Huttenlocher Grouping Perceptual Organization Structural relationships between tokens Parallelism, symmetry, alignment Similarity of token properties Often strong psychophysical
CS 664 Segmentation Daniel Huttenlocher Grouping Perceptual Organization Structural relationships between tokens Parallelism, symmetry, alignment Similarity of token properties Often strong psychophysical
Modern Medical Image Analysis 8DC00 Exam
 Parts of answers are inside square brackets [... ]. These parts are optional. Answers can be written in Dutch or in English, as you prefer. You can use drawings and diagrams to support your textual answers.
Parts of answers are inside square brackets [... ]. These parts are optional. Answers can be written in Dutch or in English, as you prefer. You can use drawings and diagrams to support your textual answers.
Detecting White Matter Lesions in Lupus
 Slicer3 Training Compendium Detecting White Matter Lesions in Lupus Version 2.1 6/25/2009 H. Jeremy Bockholt Mark Scully -1- Learning objective This tutorial demonstrates an automated, multi-level method
Slicer3 Training Compendium Detecting White Matter Lesions in Lupus Version 2.1 6/25/2009 H. Jeremy Bockholt Mark Scully -1- Learning objective This tutorial demonstrates an automated, multi-level method
EXCEL 2007 TIP SHEET. Dialog Box Launcher these allow you to access additional features associated with a specific Group of buttons within a Ribbon.
 EXCEL 2007 TIP SHEET GLOSSARY AutoSum a function in Excel that adds the contents of a specified range of Cells; the AutoSum button appears on the Home ribbon as a. Dialog Box Launcher these allow you to
EXCEL 2007 TIP SHEET GLOSSARY AutoSum a function in Excel that adds the contents of a specified range of Cells; the AutoSum button appears on the Home ribbon as a. Dialog Box Launcher these allow you to
Structural MRI of Amygdala Tutorial: Observation, Segmentation, Quantification
 Structural MRI of Amygdala Tutorial: Observation, Segmentation, Quantification The FMRIB Software Library (FSL) is a powerful tool that allows users to observe the human brain in various planes and dimensions,
Structural MRI of Amygdala Tutorial: Observation, Segmentation, Quantification The FMRIB Software Library (FSL) is a powerful tool that allows users to observe the human brain in various planes and dimensions,
Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques
 Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques Sea Chen Department of Biomedical Engineering Advisors: Dr. Charles A. Bouman and Dr. Mark J. Lowe S. Chen Final Exam October
Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques Sea Chen Department of Biomedical Engineering Advisors: Dr. Charles A. Bouman and Dr. Mark J. Lowe S. Chen Final Exam October
Slicer3 Training Compendium. Slicer3 Training Tutorial ARCTIC (v1.2)
 Slicer3 Training Compendium Slicer3 Training Tutorial ARCTIC (v1.2) ( ThICkness (Automatic Regional Cortical University of North Carolina, Chapel Hill: Neuro Image Research and Analysis Lab Neurodevelopmental
Slicer3 Training Compendium Slicer3 Training Tutorial ARCTIC (v1.2) ( ThICkness (Automatic Regional Cortical University of North Carolina, Chapel Hill: Neuro Image Research and Analysis Lab Neurodevelopmental
Processing math: 100% Intensity Normalization
 Intensity Normalization Overall Pipeline 2/21 Intensity normalization Conventional MRI intensites (T1-w, T2-w, PD, FLAIR) are acquired in arbitrary units Images are not comparable across scanners, subjects,
Intensity Normalization Overall Pipeline 2/21 Intensity normalization Conventional MRI intensites (T1-w, T2-w, PD, FLAIR) are acquired in arbitrary units Images are not comparable across scanners, subjects,
Automatic Generation of Training Data for Brain Tissue Classification from MRI
 Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. COCOSCO, Alex P. ZIJDENBOS, and Alan C. EVANS http://www.bic.mni.mcgill.ca/users/crisco/ McConnell Brain Imaging
Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. COCOSCO, Alex P. ZIJDENBOS, and Alan C. EVANS http://www.bic.mni.mcgill.ca/users/crisco/ McConnell Brain Imaging
Modify Panel. Flatten Tab
 AFM Image Processing Most images will need some post acquisition processing. A typical procedure is to: i) modify the image by flattening, using a planefit, and possibly also a mask, ii) analyzing the
AFM Image Processing Most images will need some post acquisition processing. A typical procedure is to: i) modify the image by flattening, using a planefit, and possibly also a mask, ii) analyzing the
Segmentation of the neonatal brain for MR images A REVIEW
 Segmentation of the neonatal brain for MR images A REVIEW Tineke Hoppinga 19-5-2010 Inhoud Abbreviations... 4 Abstract... 5 Introduction... 6 Method... 8 Basic explanations of the methods... 8 Weisenfeld...
Segmentation of the neonatal brain for MR images A REVIEW Tineke Hoppinga 19-5-2010 Inhoud Abbreviations... 4 Abstract... 5 Introduction... 6 Method... 8 Basic explanations of the methods... 8 Weisenfeld...
Image Segmentation. Ross Whitaker SCI Institute, School of Computing University of Utah
 Image Segmentation Ross Whitaker SCI Institute, School of Computing University of Utah What is Segmentation? Partitioning images/volumes into meaningful pieces Partitioning problem Labels Isolating a specific
Image Segmentation Ross Whitaker SCI Institute, School of Computing University of Utah What is Segmentation? Partitioning images/volumes into meaningful pieces Partitioning problem Labels Isolating a specific
Neuroimage Processing
 Neuroimage Processing Instructor: Moo K. Chung mkchung@wisc.edu Lecture 2-3. General Linear Models (GLM) Voxel-based Morphometry (VBM) September 11, 2009 What is GLM The general linear model (GLM) is a
Neuroimage Processing Instructor: Moo K. Chung mkchung@wisc.edu Lecture 2-3. General Linear Models (GLM) Voxel-based Morphometry (VBM) September 11, 2009 What is GLM The general linear model (GLM) is a
RSNA, /rg
 RSNA, 2015 10.1148/rg.2015140320 Appendix As noted in the main article, DICOM image files cannot be used directly for 3D printing; further steps are necessary to make them readable by 3D printers. The
RSNA, 2015 10.1148/rg.2015140320 Appendix As noted in the main article, DICOM image files cannot be used directly for 3D printing; further steps are necessary to make them readable by 3D printers. The
Subcortical Structure Segmentation using Probabilistic Atlas Priors
 Subcortical Structure Segmentation using Probabilistic Atlas Priors Sylvain Gouttard a, Martin Styner a,b, Sarang Joshi c, Rachel G. Smith a, Heather Cody a, Guido Gerig a,b a Department of Psychiatry,
Subcortical Structure Segmentation using Probabilistic Atlas Priors Sylvain Gouttard a, Martin Styner a,b, Sarang Joshi c, Rachel G. Smith a, Heather Cody a, Guido Gerig a,b a Department of Psychiatry,
n o r d i c B r a i n E x Tutorial DSC Module
 m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial DSC Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial DSC Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
The organization of the human cerebral cortex estimated by intrinsic functional connectivity
 1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
Incorporating Non-rigid Registration into Expectation Maximization Algorithm to Segment MR Images
 Incorporating Non-rigid Registration into Expectation Maximization Algorithm to Segment MR Images Kilian M Pohl 1, William M. Wells 2, Alexandre Guimond 3, Kiyoto Kasai 4, Martha E. Shenton 2,4, Ron Kikinis
Incorporating Non-rigid Registration into Expectation Maximization Algorithm to Segment MR Images Kilian M Pohl 1, William M. Wells 2, Alexandre Guimond 3, Kiyoto Kasai 4, Martha E. Shenton 2,4, Ron Kikinis
CS 229 Midterm Review
 CS 229 Midterm Review Course Staff Fall 2018 11/2/2018 Outline Today: SVMs Kernels Tree Ensembles EM Algorithm / Mixture Models [ Focus on building intuition, less so on solving specific problems. Ask
CS 229 Midterm Review Course Staff Fall 2018 11/2/2018 Outline Today: SVMs Kernels Tree Ensembles EM Algorithm / Mixture Models [ Focus on building intuition, less so on solving specific problems. Ask
Analysis of fmri data within Brainvisa Example with the Saccades database
 Analysis of fmri data within Brainvisa Example with the Saccades database 18/11/2009 Note : All the sentences in italic correspond to informations relative to the specific dataset under study TP participants
Analysis of fmri data within Brainvisa Example with the Saccades database 18/11/2009 Note : All the sentences in italic correspond to informations relative to the specific dataset under study TP participants
Fmri Spatial Processing
 Educational Course: Fmri Spatial Processing Ray Razlighi Jun. 8, 2014 Spatial Processing Spatial Re-alignment Geometric distortion correction Spatial Normalization Smoothing Why, When, How, Which Why is
Educational Course: Fmri Spatial Processing Ray Razlighi Jun. 8, 2014 Spatial Processing Spatial Re-alignment Geometric distortion correction Spatial Normalization Smoothing Why, When, How, Which Why is
Image Segmentation. Ross Whitaker SCI Institute, School of Computing University of Utah
 Image Segmentation Ross Whitaker SCI Institute, School of Computing University of Utah What is Segmentation? Partitioning images/volumes into meaningful pieces Partitioning problem Labels Isolating a specific
Image Segmentation Ross Whitaker SCI Institute, School of Computing University of Utah What is Segmentation? Partitioning images/volumes into meaningful pieces Partitioning problem Labels Isolating a specific
Preview from Notesale.co.uk Page 2 of 61
 Modify a table Applying styles to tables; banding rows and columns; inserting total rows; removing styles from tables Filter and sort a table Filtering records; sorting data on multiple columns; changing
Modify a table Applying styles to tables; banding rows and columns; inserting total rows; removing styles from tables Filter and sort a table Filtering records; sorting data on multiple columns; changing
MARS: Multiple Atlases Robust Segmentation
 Software Release (1.0.1) Last updated: April 30, 2014. MARS: Multiple Atlases Robust Segmentation Guorong Wu, Minjeong Kim, Gerard Sanroma, and Dinggang Shen {grwu, mjkim, gerard_sanroma, dgshen}@med.unc.edu
Software Release (1.0.1) Last updated: April 30, 2014. MARS: Multiple Atlases Robust Segmentation Guorong Wu, Minjeong Kim, Gerard Sanroma, and Dinggang Shen {grwu, mjkim, gerard_sanroma, dgshen}@med.unc.edu
Multi-Sectional Views Textural Based SVM for MS Lesion Segmentation in Multi-Channels MRIs
 56 The Open Biomedical Engineering Journal, 2012, 6, 56-72 Open Access Multi-Sectional Views Textural Based SVM for MS Lesion Segmentation in Multi-Channels MRIs Bassem A. Abdullah*, Akmal A. Younis and
56 The Open Biomedical Engineering Journal, 2012, 6, 56-72 Open Access Multi-Sectional Views Textural Based SVM for MS Lesion Segmentation in Multi-Channels MRIs Bassem A. Abdullah*, Akmal A. Younis and
S-PLUS INSTRUCTIONS FOR CASE STUDIES IN THE STATISTICAL SLEUTH
 S-PLUS INSTRUCTIONS FOR CASE STUDIES IN THE STATISTICAL SLEUTH Dan Schafer January, 2002 This guide contains brief instructions for accomplishing the graphical and numerical analyses of the case studies
S-PLUS INSTRUCTIONS FOR CASE STUDIES IN THE STATISTICAL SLEUTH Dan Schafer January, 2002 This guide contains brief instructions for accomplishing the graphical and numerical analyses of the case studies
3D Slicer. NA-MIC National Alliance for Medical Image Computing 4 February 2011
 NA-MIC http://na-mic.org 3D Slicer 4 February 2011 Andrey Fedorov, PhD Steve Pieper, PhD Ron Kikinis, MD Surgical Planning Lab Brigham and Women's Hospital Acknowledgments Picture courtesy Kapur, Jakab,
NA-MIC http://na-mic.org 3D Slicer 4 February 2011 Andrey Fedorov, PhD Steve Pieper, PhD Ron Kikinis, MD Surgical Planning Lab Brigham and Women's Hospital Acknowledgments Picture courtesy Kapur, Jakab,
Measuring longitudinal brain changes in humans and small animal models. Christos Davatzikos
 Measuring longitudinal brain changes in humans and small animal models Christos Davatzikos Section of Biomedical Image Analysis University of Pennsylvania (Radiology) http://www.rad.upenn.edu/sbia Computational
Measuring longitudinal brain changes in humans and small animal models Christos Davatzikos Section of Biomedical Image Analysis University of Pennsylvania (Radiology) http://www.rad.upenn.edu/sbia Computational
BioEST ver. 1.5beta User Manual Chang-Hwan Im (Ph.D.)
 BioEST ver. 1.5beta User Manual Chang-Hwan Im (Ph.D.) http://www.bioest.com E-mail: ichism@elecmech.snu.ac.kr imxxx010@umn.edu 1. What is BioEST? BioEST is a special edition of SNUEEG(http://www.bioinverse.com),
BioEST ver. 1.5beta User Manual Chang-Hwan Im (Ph.D.) http://www.bioest.com E-mail: ichism@elecmech.snu.ac.kr imxxx010@umn.edu 1. What is BioEST? BioEST is a special edition of SNUEEG(http://www.bioinverse.com),
XnView 1.9. a ZOOMERS guide. Introduction...2 Browser Mode... 5 Image View Mode...15 Printing Image Editing...28 Configuration...
 XnView 1.9 a ZOOMERS guide Introduction...2 Browser Mode... 5 Image View Mode...15 Printing... 22 Image Editing...28 Configuration... 36 Written by Chorlton Workshop for hsbp Introduction This is a guide
XnView 1.9 a ZOOMERS guide Introduction...2 Browser Mode... 5 Image View Mode...15 Printing... 22 Image Editing...28 Configuration... 36 Written by Chorlton Workshop for hsbp Introduction This is a guide
Work with Shapes. Concepts CHAPTER. Concepts, page 3-1 Procedures, page 3-5
 3 CHAPTER Revised: November 15, 2011 Concepts, page 3-1, page 3-5 Concepts The Shapes Tool is Versatile, page 3-2 Guidelines for Shapes, page 3-2 Visual Density Transparent, Translucent, or Opaque?, page
3 CHAPTER Revised: November 15, 2011 Concepts, page 3-1, page 3-5 Concepts The Shapes Tool is Versatile, page 3-2 Guidelines for Shapes, page 3-2 Visual Density Transparent, Translucent, or Opaque?, page
Basal Ganglia Matching Tools 2- Tutorial
 Basal Ganglia Matching Tools 2- Tutorial Minimum system hardware requirements: - Processor: Pentium 4 or Athlon with speed of 1 GHz - RAM: 256 MB - Hard disk space: 40 MB. System requirements 1. Insert
Basal Ganglia Matching Tools 2- Tutorial Minimum system hardware requirements: - Processor: Pentium 4 or Athlon with speed of 1 GHz - RAM: 256 MB - Hard disk space: 40 MB. System requirements 1. Insert
FIFI-LS: Basic Cube Analysis using SOSPEX
 FIFI-LS: Basic Cube Analysis using SOSPEX Date: 1 Oct 2018 Revision: - CONTENTS 1 INTRODUCTION... 1 2 INGREDIENTS... 1 3 INSPECTING THE CUBE... 3 4 COMPARING TO A REFERENCE IMAGE... 5 5 REFERENCE VELOCITY
FIFI-LS: Basic Cube Analysis using SOSPEX Date: 1 Oct 2018 Revision: - CONTENTS 1 INTRODUCTION... 1 2 INGREDIENTS... 1 3 INSPECTING THE CUBE... 3 4 COMPARING TO A REFERENCE IMAGE... 5 5 REFERENCE VELOCITY
IDL Tutorial. Working with Images. Copyright 2008 ITT Visual Information Solutions All Rights Reserved
 IDL Tutorial Working with Images Copyright 2008 ITT Visual Information Solutions All Rights Reserved http://www.ittvis.com/ IDL is a registered trademark of ITT Visual Information Solutions for the computer
IDL Tutorial Working with Images Copyright 2008 ITT Visual Information Solutions All Rights Reserved http://www.ittvis.com/ IDL is a registered trademark of ITT Visual Information Solutions for the computer
