ASAP_2.0 (Automatic Software for ASL Processing) USER S MANUAL
|
|
|
- Gabriella Dixon
- 5 years ago
- Views:
Transcription
1 ASAP_2.0 (Automatic Software for ASL Processing) USER S MANUAL ASAP was developed as part of the COST Action "Arterial Spin Labelling Initiative in Dementia (AID)" by: Department of Neuroimaging, Institute of Psychiatry at King's College London (UK). Laboratorio de Análisis de Imagen Médica y Biometría (LAIMBIO), Universidad Rey Juan Carlos (Spain). Referencing: If you employ ASAP in your research please reference the paper below Mato Abad V, García-Polo P, O'Daly O, Hernández-Tamames JA, Zelaya F (2016). ASAP (Automatic Software for ASL Processing): A toolbox for processing Arterial Spin Labeling images. Magn Reson Imaging, 34(3): ASAP_2.0 is a beta version. Please assist us by reporting errors or questions to virginia.mato@urjc.es June 2017
2 1. Introduction ASAP processing toolbox is written in MATLAB and includes all the steps needed for preparing the resting-state ASL (Arterial Spin Labelling) maps for a statistical analysis. This toolbox includes different analysis strategies depending on the ASL image readout and the type of structural scans available from each subject (T2-weighted and/or T1- weighted high-resolution scans). The Graphical User Interface (GUI) can automatically process several ASL datasets by accessing both SPM-12 and FSL software, which are two of the most widely available image processing platforms for structural and functional MRI. These data can be setup manually through the GUI fields (which have been programmed to mimic the SPM file handling menus) or by advanced options that include a batch mode. The GUI also provides options to visualize the results and for different file format. There are different pipelines for the ASAP processing steps, depending on: (1) The reference volume used for the co-registration with the structural high-resolution scans: The ASL image The Proton Density (PD) acquisition. (2) The mode to do the co-registration: Upsampling the ASL to high-resolution. Downsampling the structural scan to the ASL resolution. Basic steps: 1. Rough skull-stripping of the resting state ASL map using the FSL Brain Extraction Tool (Bet) for noisy ASL maps using a conservative threshold. 2. Co-registration of [ASL scan <-> skull-stripped structural scan] (upsampling or downsampling mode) using SPM- 8 by to ways: 2.1. Using the ASL as source image Using the PD as source image and moving the ASL during co-register. 3. Skull-stripping of the structural scan by two different options: the FSL bet tool (applied to the T2 or T1 weighted scan) OR with a brain mask created from SPM segmentation of the T1-weighted high resolution scan. 4. Partial Volume Correction (PVC) of the ASL maps. In the current version, two different methods are available for PVC: (1) The linear regression method described by Asllani 1 and (2) The method based on a previous PET 1 Asllani I. et al.; Regression algorithm correcting for partial volume effects in arterial spin labeling MRI. Magnetic Resonance in Medicine 60: (2008). 2
3 study 2 (hereafter PET s correction) that assumes perfusion of WM is globally 40% of that of GM for correction resting cerebral blood flow. This method has been later applied in several ASL studies Skull-stripping of the co-registered and partial volume corrected ASL maps using the structural brain mask obtained on (4) above. 6. MNI Normalization of the structural scan and ASL maps obtained in (6) to the standard MNI space. 7. Smoothing of the final, spatially normalised CBF maps by a smoothing kernel chosen by the user. The software also offers additional facilities for rapid extraction of CBF values from anatomically or functionally defined Regions of Interest (ROIs). In batch mode, the software can simultaneously extract mean, median and maximum values from several ROIs in several CBF maps. The output is generated in text files that can easily be incorporated into statistical analysis packages such as SPSS, etc. IMPORTANT: Check Section 5 for important recommendations for a proper use of the different options available in the toolbox. 2 Leenders K.L. et al; Cerebral blood flow, blood volume and oxygen utilization. Normal values and effect of age. Brain : a journal of neurology 113:27-47 (1990). 3 Chen Y. et al.; Voxel-level comparison of arterial spin-labeled perfusion MRI and FDG-PET in Alzheimer disease. Neurology 77(22): (2011). Du A.T. et al.; Hypoperfusion in frontotemporal dementia and Alzheimer disease by arterial spin labeling MRI. Neurology 67(7): (2006). Johnson N.A., et al. Pattern of cerebral hypoperfusion in Alzheimer disease and mild cognitive impairment measured with arterial spinlabeling MR imaging: initial experience. Radiology 234(3): (2005). 3
4 2. Requirements ASAP runs on Unix systems (Linux, Mac OS) with MATLAB software installed (including the Tools for NIfTI and ANALYZE image package 4 ). ASAP access both FSL software and Statistical Parametric Mapping (SPM-12) software, so the toolbox requires you have: SPM-12 installed and added to your MATLAB path. FSL installed and added at your path (FSLDIR environment variable)* 3. Installation 1. Download the package ASAP_2.0.zip 2. Unzip the package: unzip ASAP_2.0.zip 3. Open MATLAB and add the ASAP folder to the MATLAB path ( File->Set path menu or addpath function). 4. Run ASL_GUI in the MATLAB console. * If you have problems with fsl path, try to add the FSLDIR environment variable in your MATLAB console: setenv('fsldir','fslabsolutepath ) 4 Tools for NIfTI and ANALYZE image, by Jimmy Shen: 4
5 4. User Guide Figure shows the ASAP processing GUI. Next are described all the GUI options and parameters. Input Files. For each subject: Structural: High-resolution structural scan. File format admitted: [.dcm.hdr.hdr.gz.img.img.gz.nii.nii.gz]. Before select files, make sure the checkbox NIFTI/ANALYZE or DICOM is in the right file format. For DICOM files, you must select all the series DICOM structural files and, after that, the output directory for the converted nifti files. In this directory, the nifti file will be saved under /<Patient_ID>/<Series_description>. Default=NIFTI/ANALYZE. T1-weighted: Check when structural scan is T1-weighted (default). T2-weighted: Check when structural scan is T2-weighted. ASL: ASL scan: CBF map or difference image (control tag image). File format admitted: [.dcm.hdr.hdr.gz.img.img.gz.nii.nii.gz]. Before select files, make sure the checkbox 5
6 NIFTI/ANALYZE or DICOM is in the right file format. For DICOM files, you must select all the series DICOM ASL files and, after that, the output directory for the converted nifti files. In this directory, the nifti file will be saved under /<Patient_ID>/<Series_description>. If the ASL series include the PD DICOM files, the PD will be also converted to nifti format and the Use PD for Coregistration will be auto selected. Default=NIFTI/ANALYZE. If you select Use PD for coregistration, the Proton Density scan is also required. File format admitted for PD: [.dcm.hdr.hdr.gz.img.img.gz.nii.nii.gz]. Before select files, make sure the checkbox NIFTI/ANALYZE or DICOM is in the right file format. For DICOM files, you must select all the series DICOM PD files and, after that, the output directory for the converted nifti files. In this directory, the nifti file will be saved under /<Patient_ID>/<Series_description>. Default=NIFTI/ ANALYZE. Check Use PD for Co-registration for using the PD scan for co-registration with the structural scan instead of the ASL volume. Output Directories. For each subject: Structural: Directory for structural output files. By default, it is set as the same as the input structural file. ASL: Directory for ASL output files. By default, it is set as the same as the input ASL file. PD: Directory for PD output files. By default, it is set as the same as the input PD file. Options. Processing parameters: Coregistration method: Upsampling: Check for upsampling the ASL to high-resolution (default) by interpolation. Downsampling: Check for downsampling the structural scan to low-resolution. ASL rough skull strip: Check for ASL rough skull-stripping using the FSL Brain Extraction Tool (Bet). Useful for noisy ASL maps. Default = 1. -f [0-1]: Fractional intensity threshold. Default = 0.2. Structural scan skull strip: Check for skull-stripping of the structural scan. Default = 1 SPM segmentation mask: Check for skull-stripping using the brain mask created with SPM segmentation output volumes. Default = 1 for T1-weighted scans and 0 for T2-weighted scans Use CSF: Check for includes the cerebrospinal fluid (CSF) map in the brain mask. Default = 1 6
7 FSL bet: Check for skull-stripping using FSL bet tool. Default = 0 for T1-weighted scans and 1 for T2-weighted scans -f [0-1]: Fractional intensity threshold. Default = 0.4 for T1-weighted scans and 0.35 for T2-weighted scans -r: Approximate head radius (mm not voxels). Default = 75. Useful for neck cleanup Partial Volume Correction: Check for apply partial volume correction to the ASL maps. The PVC of the ASL maps is only available when working in the low-resolution ASL space. Two different methods are now available: Asllani: A linear regression method described in reference [1] (see page 3). This algorithm estimates the ASL maps for grey matter (GM) and white matter (WM) independently. Kernel size: The regression-kernel size. Options are: [3x3x1, 5x5x1, 7x7x1, 9x9x1, 11x11x1] PET s correction: This method assumes that all contributions to perfusion are from brain tissue and that CSF has no contribution. ASL intensities are corrected according to the equation: I corr=i uncorr/(p GM+0.4*P WM), where I corr and I uncorr are the corrected and uncorrected intensities, the 0.4 factor is the global ratio between WM and GM and PGM and PWM are the probabilities of GM and WM, respectively. Normalization: Check for normalization to MNI space the output files. The resolution of normalized files is 2x2x2 with 91x109x91 dimensions. Smooth: Check for smooth final ASL maps. Default = 0 FWHM value. Kernel width in mm. Default = [6, 6, 6]. Output files: Format of output files [.hdr/.img.nii]. Default =.nii Quick checkreg: Check for display a quick checkreg of final ASL map in the web browser. Open SPM progress window: Opens the SPM interactive window to following the progress Actions: Next Sub.: Click to save current subject s dataset and start with the parameters for a new subject. Run: Click to start processing. Cancel: Click to reset and cancel all the inserted data (for all subjects). Quit: Exit. Advanced options: Load batch files: Load input files from text files. First, select the text file containing the input structural scans and then, the text file containing input ASL maps. If Use PD for 7
8 coregistration is checked, select in third option the text file with the input PD scans. For NIFTI/ANALYZE format, the text file will contain the absolute path of each file in a separate line. For DICOM format, the text file will contain the absolute path of the directory that contains all the DICOM files for each subject in a separate line. Important: Each text file must contain each filename or directory path in a separate line. Make sure that the subject file order is the same in all (two or three) files and that there is not blank lines at the end of files. For more than one ASL acquisition per session, repeat the structural file path as many times as ASL scans in the structural text file. Using this option, it is necessary to fill the Options panel parameters (the same options will apply for all subjects). After that, click the Run button to start the process. Output files will be saved in the same directory as input files. ROI Statistics: Open ROI GUI for statistics in ASL maps. Process For each subject: 1. Reorient to the AC-PC plane (Anterior Commissure - Posterior Commissure) the structural, PD* and ASL files. Output filenames: structural_reo, PD_reo* and ASL_reo. 2. If [ASL rough skull strip] = 1, run bet command on ASL image for a rough skull-stripping, using the input fractional threshold. This step is useful for noisy ASLs maps in upsampling mode for better co-registration with the structural scan. 3. If [Structural scan skull strip] = 1: 3.1. If [SPM segmentation mask] = 1, run tissue segmentation task of SPM-12 software on structural scan. Then, create the skull-stripped brain volume and mask with segmentation task outputs: grey matter (GM, c1) and white matter (WM, c2) maps. If [Use CSF] = 1, create the CSF (c3) map and include it in the skull-stripped brain volume and mask. Output filenames: c1structural_reo, c2structural_reo, c3structural_reo***, y_structural_reo, structural_reo_ stripped and structural_reo_stripped_mask If [FSL bet] = 1, run bet command on structural scan, using the input fractional threshold - f, the input head radius r and the m option to generate the brain mask. Output filenames: structural_reo_stripped and structural_reo_stripped_mask. 4. For co-registration: 4.1. Upsampling mode: If [Use PD for coregistration] = 1, Co-register the PD map to the skull-stripped 8
9 structural volume using the SPM-12 coreg function and moving the ASL map in the process. Output filenames: rpd_reo** and rasl_reo** If [Use PD for coregistration] = 0, Co-register the ASL map to the skull-stripped structural volume using the SPM-12 coreg function. Output filenames: rasl_reo** Downsampling mode: If [Use PD for coregistration] = 1, Co-register the skull-stripped structural volume (step 3) to the PD map using the SPM-12 coreg function and moving the ASL map in the process. Output filenames: rc1structural_reo**, rc2structural_reo**, rc3structural_reo***, rstructural_reo_stripped** and rstructural_reo_stripped_mask** If [Use PD for coregistration] = 0, Co-register the skull-stripped structural volume (step 3) to the ASL map using the SPM-12 coreg function. Output filenames: rc1structural_reo**, rc2structural_reo**, rc3structural_reo***, rstructural_reo _stripped** and rstructural_reo_stripped_mask**. 5. If [Partial volume correction] = 1, check is the structural scan is already segmented in GM, WM and CSF. If not, run tissue segmentation task of SPM-12 (see step 3.1) If Asllani method is selected run the linear regression method described in reference [1] using the selected kernel size. Four different files will be created: partial GM ASL, partial WM ASL and their respective masks: partial_gm_asl*(gm_map>0.05) and partial_wm_asl* (WM_map>0.05). Output filenames: ASL_reo_CBF_asllani_gm, ASL_reo_CBF_asllani _gm_mask, ASL_reo_CBF_asllani_wm and ASL_reo_CBF_asllani_wm_mask If PET s correction is selected, run the method described above. In this case, only the partial GM ASL file is estimated. Output filenames: ASL_reo_CBF_pet_gm. 6. Multiply the co-registered ASL file map by the skull-stripped mask (step 3) using the SPM-12 imcalc function. Output filenames: ASL_reo_stripped. 7. If [Normalization] = 1, normalise to MNI the ASL co-registered and skull-stripped map (step 6), the PD* and the structural output files using the SPM-12 normalise function. The resolution of normalized files is 2x2x2 with 91x109x91 dimensions. Output filenames: wasl_reo_stripped, wpd_reo*, wc1structural_reo, wc2structural_reo, wc3structural_reo***, wstructural_reo_stripped, wstructural_reo_stripped_mask and files created in step5 (if selected) with prefix w. 8. If [smooth] = 1, smooth the final output ASL map (step 7) using the SPM-12 smooth function and input FWHM value. Output filenames: swasl_reo_stripped and files created in step 5 (if selected) with prefix s. 9
10 9. If [quick checkreg] = 1, run FSL slicesdir command for display the output ASL map (step 7) in the web browser. * Only if [Use PD for coregistration] = 1 ** Depending on the co-registration mode, the r prefix will be in the structural (down-sampled) or ASL (upsampled) files for the subsequent steps *** Optional 10
11 ROI GUI ROI GUI offers extraction of CBF values from anatomically or functionally defined Regions of Interest. Options Save results in text file: Save the statistical results in a text file. Default = 1. Select files: Click to select first the ROI input files and, then, the CBF source files. Load list files: Click to read the input files from text files. First, select the text file with the absolute path of ROI files (.img/.nii) and then, the text file with the absolute path of source images. These files must contain each filename in a separate line. Run: Click to start processing. Quit: Exit Process For each ROI: 1. Calculates mean, median and max values for each source file. 2. Save the statistical results in a.mat file (filename: 'ROIresults_mmddyy_HHMM.mat). This file is saved in the current directory. The.mat file has 3 matrixes: ROI_mean, ROI_median and ROI_max with the ROI analysis results. Columns contain the results of each input ROI and rows contain the results of each source file. 3. If [save results in text file] = 1, the statistical results for each ROI are saved in individually files. This file is saved in the same directory that the ROI file (filename: <ROIfilename>_ROIstatistics.txt). Files will not be overwritten; new statistics over the same ROI will be appended at the end of the text file. 11
12 5. Recommendations 1. For T2-weighted structural scan is recommended the FSL options, due to the tissue segmentation in GM, WM and CSF does not usually work well with T2-weighted contrast images. 2. For T1-weighted structural scan is recommended the SPM-12 options. 3. The PVC of the ASL maps is only available when working in the low-resolution ASL space. The recommended kernel size for Asllani method for PVC is [5x5x1]. 4. Also, for PVC is highly recommended use the T1-weighted as structural scan, because high quality GM and WM maps are needed. 5. Option ASL rough skull strip is only recommended for noisy ASL maps, for better co-registration with the structural scan. It is also suggested check the skull-stripped ASL map in order to not deleting brain tissue. 6. The toolbox helps to repeating analysis by avoiding some processing steps. Useful, for example, for applying different partial volume corrections on the same input data: If input file contains the suffix _reo the toolbox accepts that the file is already reoriented and skips this step. Checking for GM, WM and CSF maps (c1*, c2* and c3* files) in the same directory that the structural scan. If they exist, the toolbox does not apply the SPM-12 segmentation step, using these files instead. Checking for structural skull-stripped and brain mask files (*_stripped and *_stripped_mask files) in the same directory that the structural scan. If they exist, the toolbox does not apply the skull strip step and uses these files instead. Checking for co-registered structural files (only in downsamplig mode). If they exist, the toolbox does not apply the co-registration of structural scan to the ASL and uses these files instead. 12
13 6. Example An example of the ASAP results is shown. The CBF map and the T1-weihgted 3D structural scan compose the dataset: Structural input data: T1file (1mm, 256x256x196) ASL input data: CBFfile
14 Selected options for analysis: Upsampling mode Rough skull-strip for CBF map T1-weighted tissue segmentation T1-weighted skull-strip using the GM+WM+CSF maps MNI normalization Next page shows the results for: Normalized and skull-stripped CBF map Normalized and skull-stripped T1-weighted scan (SPM- 12) Normalized tissue maps: GM and WM 14
15 Normalized and skull-stripped CBF: wrcbffile_reo_stripped T1 normalized and skull-stripped: wt1file_reo_stripped Normalized GM map: wc1t1file_reo Normalized WM map: wc2t1file_reo 15
Correction of Partial Volume Effects in Arterial Spin Labeling MRI
 Correction of Partial Volume Effects in Arterial Spin Labeling MRI By: Tracy Ssali Supervisors: Dr. Keith St. Lawrence and Udunna Anazodo Medical Biophysics 3970Z Six Week Project April 13 th 2012 Introduction
Correction of Partial Volume Effects in Arterial Spin Labeling MRI By: Tracy Ssali Supervisors: Dr. Keith St. Lawrence and Udunna Anazodo Medical Biophysics 3970Z Six Week Project April 13 th 2012 Introduction
MRI Segmentation MIDAS, 2007, 2010
 MRI Segmentation MIDAS, 2007, 2010 Lawrence O. Hall, Dmitry Goldgof, Yuhua Gu, Prodip Hore Dept. of Computer Science & Engineering University of South Florida CONTENTS: 1. Introduction... 1 2. Installing
MRI Segmentation MIDAS, 2007, 2010 Lawrence O. Hall, Dmitry Goldgof, Yuhua Gu, Prodip Hore Dept. of Computer Science & Engineering University of South Florida CONTENTS: 1. Introduction... 1 2. Installing
ANALYSIS OF FUNCTIONAL MAGNETIC RESONANCE IMAGING DATA USING SPM99: VOXEL-BASED MORPHOMETRY DONNA ROSE ADDIS
 Donna Rose Addis, TWRI, May 2004 1 ANALYSIS OF FUNCTIONAL MAGNETIC RESONANCE IMAGING DATA USING SPM99: VOXEL-BASED MORPHOMETRY DONNA ROSE ADDIS DEPT. OF PSYCHOLOGY, UNIVERSITY OF TORONTO TORONTO WESTERN
Donna Rose Addis, TWRI, May 2004 1 ANALYSIS OF FUNCTIONAL MAGNETIC RESONANCE IMAGING DATA USING SPM99: VOXEL-BASED MORPHOMETRY DONNA ROSE ADDIS DEPT. OF PSYCHOLOGY, UNIVERSITY OF TORONTO TORONTO WESTERN
GLM for fmri data analysis Lab Exercise 1
 GLM for fmri data analysis Lab Exercise 1 March 15, 2013 Medical Image Processing Lab Medical Image Processing Lab GLM for fmri data analysis Outline 1 Getting Started 2 AUDIO 1 st level Preprocessing
GLM for fmri data analysis Lab Exercise 1 March 15, 2013 Medical Image Processing Lab Medical Image Processing Lab GLM for fmri data analysis Outline 1 Getting Started 2 AUDIO 1 st level Preprocessing
A Spatio-temporal Denoising Approach based on Total Variation Regularization for Arterial Spin Labeling
 A Spatio-temporal Denoising Approach based on Total Variation Regularization for Arterial Spin Labeling Cagdas Ulas 1,2, Stephan Kaczmarz 3, Christine Preibisch 3, Jonathan I Sperl 2, Marion I Menzel 2,
A Spatio-temporal Denoising Approach based on Total Variation Regularization for Arterial Spin Labeling Cagdas Ulas 1,2, Stephan Kaczmarz 3, Christine Preibisch 3, Jonathan I Sperl 2, Marion I Menzel 2,
Automated MR Image Analysis Pipelines
 Automated MR Image Analysis Pipelines Andy Simmons Centre for Neuroimaging Sciences, Kings College London Institute of Psychiatry. NIHR Biomedical Research Centre for Mental Health at IoP & SLAM. Neuroimaging
Automated MR Image Analysis Pipelines Andy Simmons Centre for Neuroimaging Sciences, Kings College London Institute of Psychiatry. NIHR Biomedical Research Centre for Mental Health at IoP & SLAM. Neuroimaging
Supplementary methods
 Supplementary methods This section provides additional technical details on the sample, the applied imaging and analysis steps and methods. Structural imaging Trained radiographers placed all participants
Supplementary methods This section provides additional technical details on the sample, the applied imaging and analysis steps and methods. Structural imaging Trained radiographers placed all participants
Single Subject Demo Data Instructions 1) click "New" and answer "No" to the "spatially preprocess" question.
 (1) conn - Functional connectivity toolbox v1.0 Single Subject Demo Data Instructions 1) click "New" and answer "No" to the "spatially preprocess" question. 2) in "Basic" enter "1" subject, "6" seconds
(1) conn - Functional connectivity toolbox v1.0 Single Subject Demo Data Instructions 1) click "New" and answer "No" to the "spatially preprocess" question. 2) in "Basic" enter "1" subject, "6" seconds
FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING. Francesca Pizzorni Ferrarese
 FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING Francesca Pizzorni Ferrarese Pipeline overview WM and GM Segmentation Registration Data reconstruction Tractography
FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING Francesca Pizzorni Ferrarese Pipeline overview WM and GM Segmentation Registration Data reconstruction Tractography
Normalization for clinical data
 Normalization for clinical data Christopher Rorden, Leonardo Bonilha, Julius Fridriksson, Benjamin Bender, Hans-Otto Karnath (2012) Agespecific CT and MRI templates for spatial normalization. NeuroImage
Normalization for clinical data Christopher Rorden, Leonardo Bonilha, Julius Fridriksson, Benjamin Bender, Hans-Otto Karnath (2012) Agespecific CT and MRI templates for spatial normalization. NeuroImage
Pre-processing of ASL data T CT
 Wed October 2, 2013 Image Processing Pre-processing: motion correction, denoising, outlier detection Alessandra Bertoldo Pre-processing of ASL data T CT C T C Single TI ASL T T T T C CCC average Pre-processing
Wed October 2, 2013 Image Processing Pre-processing: motion correction, denoising, outlier detection Alessandra Bertoldo Pre-processing of ASL data T CT C T C Single TI ASL T T T T C CCC average Pre-processing
Measuring baseline whole-brain perfusion on GE 3.0T using arterial spin labeling (ASL) MRI
 Measuring baseline whole-brain perfusion on GE 3.0T using arterial spin labeling (ASL) MRI Revision date: 09/15/2008 Overview This document describes the procedure for measuring baseline whole-brain perfusion
Measuring baseline whole-brain perfusion on GE 3.0T using arterial spin labeling (ASL) MRI Revision date: 09/15/2008 Overview This document describes the procedure for measuring baseline whole-brain perfusion
IS MRI Based Structure a Mediator for Lead s Effect on Cognitive Function?
 IS MRI Based Structure a Mediator for Lead s Effect on Cognitive Function? Brian Caffo, Sining Chen, Brian Schwartz Department of Biostatistics and Environmental Health Sciences Johns Hopkins University
IS MRI Based Structure a Mediator for Lead s Effect on Cognitive Function? Brian Caffo, Sining Chen, Brian Schwartz Department of Biostatistics and Environmental Health Sciences Johns Hopkins University
Introduction to Neuroimaging Janaina Mourao-Miranda
 Introduction to Neuroimaging Janaina Mourao-Miranda Neuroimaging techniques have changed the way neuroscientists address questions about functional anatomy, especially in relation to behavior and clinical
Introduction to Neuroimaging Janaina Mourao-Miranda Neuroimaging techniques have changed the way neuroscientists address questions about functional anatomy, especially in relation to behavior and clinical
Introduction to fmri. Pre-processing
 Introduction to fmri Pre-processing Tibor Auer Department of Psychology Research Fellow in MRI Data Types Anatomical data: T 1 -weighted, 3D, 1/subject or session - (ME)MPRAGE/FLASH sequence, undistorted
Introduction to fmri Pre-processing Tibor Auer Department of Psychology Research Fellow in MRI Data Types Anatomical data: T 1 -weighted, 3D, 1/subject or session - (ME)MPRAGE/FLASH sequence, undistorted
PANDA Manual Zaixu Cui & Suyu Zhong & Gaolang Gong
 PANDA Manual Zaixu Cui & Suyu Zhong & Gaolang Gong National Key Laboratory of Cognitive Neuroscience and Learning Beijing Normal University, China Contents Overview Setup Files/Directories selection Preparing
PANDA Manual Zaixu Cui & Suyu Zhong & Gaolang Gong National Key Laboratory of Cognitive Neuroscience and Learning Beijing Normal University, China Contents Overview Setup Files/Directories selection Preparing
fmri Basics: Spatial Pre-processing Workshop
 fmri Basics: Spatial Pre-processing Workshop Starting a VNC session: Most of your fmri analysis will be done on the central Linux machines accessed via a VNC (Virtual Network Computing) server. This is
fmri Basics: Spatial Pre-processing Workshop Starting a VNC session: Most of your fmri analysis will be done on the central Linux machines accessed via a VNC (Virtual Network Computing) server. This is
Math in image processing
 Math in image processing Math in image processing Nyquist theorem Math in image processing Discrete Fourier Transformation Math in image processing Image enhancement: scaling Math in image processing Image
Math in image processing Math in image processing Nyquist theorem Math in image processing Discrete Fourier Transformation Math in image processing Image enhancement: scaling Math in image processing Image
Methods for data preprocessing
 Methods for data preprocessing John Ashburner Wellcome Trust Centre for Neuroimaging, 12 Queen Square, London, UK. Overview Voxel-Based Morphometry Morphometry in general Volumetrics VBM preprocessing
Methods for data preprocessing John Ashburner Wellcome Trust Centre for Neuroimaging, 12 Queen Square, London, UK. Overview Voxel-Based Morphometry Morphometry in general Volumetrics VBM preprocessing
Table of Contents. IntroLab < SPMLabs < Dynevor TWiki
 Table of Contents Lab 1: Introduction to SPM and data checking...1 Goals of this Lab...1 Prerequisites...1 An SPM Installation...1 SPM Defaults...2 L/R Brain Orientation...2 Memory Use for Data Processing...2
Table of Contents Lab 1: Introduction to SPM and data checking...1 Goals of this Lab...1 Prerequisites...1 An SPM Installation...1 SPM Defaults...2 L/R Brain Orientation...2 Memory Use for Data Processing...2
PRANA. Project Review and Analysis. March (Pre-release and still under development!)
 PRANA Project Review and Analysis March 2010- (Pre-release and still under development!) A.A. Maudsley Contents: PRANA... 1 1. Introduction... 2 1.1. Examples... 2 2. Subject/Study Selection and Filter...
PRANA Project Review and Analysis March 2010- (Pre-release and still under development!) A.A. Maudsley Contents: PRANA... 1 1. Introduction... 2 1.1. Examples... 2 2. Subject/Study Selection and Filter...
An Introduction To Automatic Tissue Classification Of Brain MRI. Colm Elliott Mar 2014
 An Introduction To Automatic Tissue Classification Of Brain MRI Colm Elliott Mar 2014 Tissue Classification Tissue classification is part of many processing pipelines. We often want to classify each voxel
An Introduction To Automatic Tissue Classification Of Brain MRI Colm Elliott Mar 2014 Tissue Classification Tissue classification is part of many processing pipelines. We often want to classify each voxel
Preprocessing of fmri Data in SPM 12 - Lab 1
 Preprocessing of fmri Data in SPM 12 - Lab 1 Index Goals of this Lab Preprocessing Overview MATLAB, SPM, Data Setup Preprocessing I: Checking Motion Correction Preprocessing II: Coregistration Preprocessing
Preprocessing of fmri Data in SPM 12 - Lab 1 Index Goals of this Lab Preprocessing Overview MATLAB, SPM, Data Setup Preprocessing I: Checking Motion Correction Preprocessing II: Coregistration Preprocessing
MARS: Multiple Atlases Robust Segmentation
 Software Release (1.0.1) Last updated: April 30, 2014. MARS: Multiple Atlases Robust Segmentation Guorong Wu, Minjeong Kim, Gerard Sanroma, and Dinggang Shen {grwu, mjkim, gerard_sanroma, dgshen}@med.unc.edu
Software Release (1.0.1) Last updated: April 30, 2014. MARS: Multiple Atlases Robust Segmentation Guorong Wu, Minjeong Kim, Gerard Sanroma, and Dinggang Shen {grwu, mjkim, gerard_sanroma, dgshen}@med.unc.edu
Functional MRI data preprocessing. Cyril Pernet, PhD
 Functional MRI data preprocessing Cyril Pernet, PhD Data have been acquired, what s s next? time No matter the design, multiple volumes (made from multiple slices) have been acquired in time. Before getting
Functional MRI data preprocessing Cyril Pernet, PhD Data have been acquired, what s s next? time No matter the design, multiple volumes (made from multiple slices) have been acquired in time. Before getting
Computational Neuroanatomy
 Computational Neuroanatomy John Ashburner john@fil.ion.ucl.ac.uk Smoothing Motion Correction Between Modality Co-registration Spatial Normalisation Segmentation Morphometry Overview fmri time-series kernel
Computational Neuroanatomy John Ashburner john@fil.ion.ucl.ac.uk Smoothing Motion Correction Between Modality Co-registration Spatial Normalisation Segmentation Morphometry Overview fmri time-series kernel
LST: A lesion segmentation tool for SPM
 LST: A lesion segmentation tool for SPM Manual/Documentation for version 2.0.15 June 2017 Paul Schmidt Lucie Wink Contents 1 Getting started 3 1.1 License................................. 3 1.2 Installation...............................
LST: A lesion segmentation tool for SPM Manual/Documentation for version 2.0.15 June 2017 Paul Schmidt Lucie Wink Contents 1 Getting started 3 1.1 License................................. 3 1.2 Installation...............................
Neuroimaging and mathematical modelling Lesson 2: Voxel Based Morphometry
 Neuroimaging and mathematical modelling Lesson 2: Voxel Based Morphometry Nivedita Agarwal, MD Nivedita.agarwal@apss.tn.it Nivedita.agarwal@unitn.it Volume and surface morphometry Brain volume White matter
Neuroimaging and mathematical modelling Lesson 2: Voxel Based Morphometry Nivedita Agarwal, MD Nivedita.agarwal@apss.tn.it Nivedita.agarwal@unitn.it Volume and surface morphometry Brain volume White matter
SPM Introduction. SPM : Overview. SPM: Preprocessing SPM! SPM: Preprocessing. Scott Peltier. FMRI Laboratory University of Michigan
 SPM Introduction Scott Peltier FMRI Laboratory University of Michigan! Slides adapted from T. Nichols SPM! SPM : Overview Library of MATLAB and C functions Graphical user interface Four main components:
SPM Introduction Scott Peltier FMRI Laboratory University of Michigan! Slides adapted from T. Nichols SPM! SPM : Overview Library of MATLAB and C functions Graphical user interface Four main components:
VBM Tutorial. 1 Getting Started. John Ashburner. March 12, 2015
 VBM Tutorial John Ashburner March 12, 2015 1 Getting Started The data provided are a selection of T1-weighted scans from the freely available IXI dataset 1. The overall plan will be to Start up SPM. Check
VBM Tutorial John Ashburner March 12, 2015 1 Getting Started The data provided are a selection of T1-weighted scans from the freely available IXI dataset 1. The overall plan will be to Start up SPM. Check
SPM Introduction SPM! Scott Peltier. FMRI Laboratory University of Michigan. Software to perform computation, manipulation and display of imaging data
 SPM Introduction Scott Peltier FMRI Laboratory University of Michigan Slides adapted from T. Nichols SPM! Software to perform computation, manipulation and display of imaging data 1 1 SPM : Overview Library
SPM Introduction Scott Peltier FMRI Laboratory University of Michigan Slides adapted from T. Nichols SPM! Software to perform computation, manipulation and display of imaging data 1 1 SPM : Overview Library
Structural Segmentation
 Structural Segmentation FAST tissue-type segmentation FIRST sub-cortical structure segmentation FSL-VBM voxelwise grey-matter density analysis SIENA atrophy analysis FAST FMRIB s Automated Segmentation
Structural Segmentation FAST tissue-type segmentation FIRST sub-cortical structure segmentation FSL-VBM voxelwise grey-matter density analysis SIENA atrophy analysis FAST FMRIB s Automated Segmentation
A User s Guide to Graphical-Model-based Multivariate Analysis
 A User s Guide to Graphical-Model-based Multivariate Analysis 1. Introduction Rong Chen December 2006 Graphical-Model-based Multivariate Analysis (GAMMA) is a Bayesian data mining software for structural
A User s Guide to Graphical-Model-based Multivariate Analysis 1. Introduction Rong Chen December 2006 Graphical-Model-based Multivariate Analysis (GAMMA) is a Bayesian data mining software for structural
Neuroimaging Group Pipeline Quick Start Manual
 Centre for Healthy Brain Ageing (CHeBA) Neuroimaging Group Pipeline Quick Start Manual UBO Detector UBO Detector is a cluster-based white matter hyperintensity (WMH) extraction pipeline based on k-nearest
Centre for Healthy Brain Ageing (CHeBA) Neuroimaging Group Pipeline Quick Start Manual UBO Detector UBO Detector is a cluster-based white matter hyperintensity (WMH) extraction pipeline based on k-nearest
Processing math: 100% Intensity Normalization
 Intensity Normalization Overall Pipeline 2/21 Intensity normalization Conventional MRI intensites (T1-w, T2-w, PD, FLAIR) are acquired in arbitrary units Images are not comparable across scanners, subjects,
Intensity Normalization Overall Pipeline 2/21 Intensity normalization Conventional MRI intensites (T1-w, T2-w, PD, FLAIR) are acquired in arbitrary units Images are not comparable across scanners, subjects,
EMSegmenter Tutorial (Advanced Mode)
 EMSegmenter Tutorial (Advanced Mode) Dominique Belhachemi Section of Biomedical Image Analysis Department of Radiology University of Pennsylvania 1/65 Overview The goal of this tutorial is to apply the
EMSegmenter Tutorial (Advanced Mode) Dominique Belhachemi Section of Biomedical Image Analysis Department of Radiology University of Pennsylvania 1/65 Overview The goal of this tutorial is to apply the
A Model-Independent, Multi-Image Approach to MR Inhomogeneity Correction
 Tina Memo No. 2007-003 Published in Proc. MIUA 2007 A Model-Independent, Multi-Image Approach to MR Inhomogeneity Correction P. A. Bromiley and N.A. Thacker Last updated 13 / 4 / 2007 Imaging Science and
Tina Memo No. 2007-003 Published in Proc. MIUA 2007 A Model-Independent, Multi-Image Approach to MR Inhomogeneity Correction P. A. Bromiley and N.A. Thacker Last updated 13 / 4 / 2007 Imaging Science and
Structural Segmentation
 Structural Segmentation FAST tissue-type segmentation FIRST sub-cortical structure segmentation FSL-VBM voxelwise grey-matter density analysis SIENA atrophy analysis FAST FMRIB s Automated Segmentation
Structural Segmentation FAST tissue-type segmentation FIRST sub-cortical structure segmentation FSL-VBM voxelwise grey-matter density analysis SIENA atrophy analysis FAST FMRIB s Automated Segmentation
GLIRT: Groupwise and Longitudinal Image Registration Toolbox
 Software Release (1.0.1) Last updated: March. 30, 2011. GLIRT: Groupwise and Longitudinal Image Registration Toolbox Guorong Wu 1, Qian Wang 1,2, Hongjun Jia 1, and Dinggang Shen 1 1 Image Display, Enhancement,
Software Release (1.0.1) Last updated: March. 30, 2011. GLIRT: Groupwise and Longitudinal Image Registration Toolbox Guorong Wu 1, Qian Wang 1,2, Hongjun Jia 1, and Dinggang Shen 1 1 Image Display, Enhancement,
BDP: BrainSuite Diffusion Pipeline. Chitresh Bhushan
 BDP: BrainSuite Diffusion Pipeline Chitresh Bhushan Why diffusion MRI? T 2 weighted MPRAGE FA map Fiber track Quantify microstructural tissue characteristics Structural connectivity Connectome Clinical
BDP: BrainSuite Diffusion Pipeline Chitresh Bhushan Why diffusion MRI? T 2 weighted MPRAGE FA map Fiber track Quantify microstructural tissue characteristics Structural connectivity Connectome Clinical
C8: May 5, 2011 Version
 C8 Documentation Timothy J. Herron Human Cognitive Neurophysiology Laboratory, David L. Woods, Director Neurology, Research Service, United States Veterans Affairs Martinez, California, USA; 1. Overview
C8 Documentation Timothy J. Herron Human Cognitive Neurophysiology Laboratory, David L. Woods, Director Neurology, Research Service, United States Veterans Affairs Martinez, California, USA; 1. Overview
n o r d i c B r a i n E x Tutorial DSC Module
 m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial DSC Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
m a k i n g f u n c t i o n a l M R I e a s y n o r d i c B r a i n E x Tutorial DSC Module Please note that this tutorial is for the latest released nordicbrainex. If you are using an older version please
Measuring baseline whole-brain perfusion on GE 3.0T using arterial spin labeling (ASL) MRI
 Measuring baseline whole-brain perfusion on GE 3.0T using arterial spin labeling (ASL) MRI Revision date: 11/20/2006 Overview This document describes the procedure for measuring baseline whole-brain perfusion
Measuring baseline whole-brain perfusion on GE 3.0T using arterial spin labeling (ASL) MRI Revision date: 11/20/2006 Overview This document describes the procedure for measuring baseline whole-brain perfusion
This Time. fmri Data analysis
 This Time Reslice example Spatial Normalization Noise in fmri Methods for estimating and correcting for physiologic noise SPM Example Spatial Normalization: Remind ourselves what a typical functional image
This Time Reslice example Spatial Normalization Noise in fmri Methods for estimating and correcting for physiologic noise SPM Example Spatial Normalization: Remind ourselves what a typical functional image
Playing with data from lab
 Playing with data from lab Getting data off the scanner From the Patient Browser, select the folder for the study you want (or within that study, the set of images you want), and then from the Transfer
Playing with data from lab Getting data off the scanner From the Patient Browser, select the folder for the study you want (or within that study, the set of images you want), and then from the Transfer
Preprocessing II: Between Subjects John Ashburner
 Preprocessing II: Between Subjects John Ashburner Pre-processing Overview Statistics or whatever fmri time-series Anatomical MRI Template Smoothed Estimate Spatial Norm Motion Correct Smooth Coregister
Preprocessing II: Between Subjects John Ashburner Pre-processing Overview Statistics or whatever fmri time-series Anatomical MRI Template Smoothed Estimate Spatial Norm Motion Correct Smooth Coregister
The organization of the human cerebral cortex estimated by intrinsic functional connectivity
 1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
Quantitative MRI of the Brain: Investigation of Cerebral Gray and White Matter Diseases
 Quantities Measured by MR - Quantitative MRI of the Brain: Investigation of Cerebral Gray and White Matter Diseases Static parameters (influenced by molecular environment): T, T* (transverse relaxation)
Quantities Measured by MR - Quantitative MRI of the Brain: Investigation of Cerebral Gray and White Matter Diseases Static parameters (influenced by molecular environment): T, T* (transverse relaxation)
BDP: BrainSuite Diffusion Pipeline. Chitresh Bhushan
 BDP: BrainSuite Diffusion Pipeline Chitresh Bhushan Why diffusion MRI? T 2 weighted MPRAGE FA map Fiber track Quantify microstructural tissue characteristics Structural connectivity Connectome Clinical
BDP: BrainSuite Diffusion Pipeline Chitresh Bhushan Why diffusion MRI? T 2 weighted MPRAGE FA map Fiber track Quantify microstructural tissue characteristics Structural connectivity Connectome Clinical
Evaluation of multiple voxel-based morphometry approaches and applications in the analysis of white matter changes in temporal lobe epilepsy
 Evaluation of multiple voxel-based morphometry approaches and applications in the analysis of white matter changes in temporal lobe epilepsy Wenjing Li a, Huiguang He a, Jingjing Lu b, Bin Lv a, Meng Li
Evaluation of multiple voxel-based morphometry approaches and applications in the analysis of white matter changes in temporal lobe epilepsy Wenjing Li a, Huiguang He a, Jingjing Lu b, Bin Lv a, Meng Li
Neuroimage Processing
 Neuroimage Processing Instructor: Moo K. Chung mkchung@wisc.edu Lecture 2-3. General Linear Models (GLM) Voxel-based Morphometry (VBM) September 11, 2009 What is GLM The general linear model (GLM) is a
Neuroimage Processing Instructor: Moo K. Chung mkchung@wisc.edu Lecture 2-3. General Linear Models (GLM) Voxel-based Morphometry (VBM) September 11, 2009 What is GLM The general linear model (GLM) is a
MIDAS Processing of Segmentation Data for Brain Lesions
 MIDAS Processing of Segmentation Data for Brain Lesions A. A. Maudsley 4/6/2012 For studies that contain lesions, the MIDAS processing has two steps that benefit from a modified processing pipeline to
MIDAS Processing of Segmentation Data for Brain Lesions A. A. Maudsley 4/6/2012 For studies that contain lesions, the MIDAS processing has two steps that benefit from a modified processing pipeline to
Automatic Registration-Based Segmentation for Neonatal Brains Using ANTs and Atropos
 Automatic Registration-Based Segmentation for Neonatal Brains Using ANTs and Atropos Jue Wu and Brian Avants Penn Image Computing and Science Lab, University of Pennsylvania, Philadelphia, USA Abstract.
Automatic Registration-Based Segmentation for Neonatal Brains Using ANTs and Atropos Jue Wu and Brian Avants Penn Image Computing and Science Lab, University of Pennsylvania, Philadelphia, USA Abstract.
fmri Preprocessing & Noise Modeling
 Translational Neuromodeling Unit fmri Preprocessing & Noise Modeling Lars Kasper September 25 th / October 17 th, 2015 MR-Technology Group & Translational Neuromodeling Unit An SPM Tutorial Institute for
Translational Neuromodeling Unit fmri Preprocessing & Noise Modeling Lars Kasper September 25 th / October 17 th, 2015 MR-Technology Group & Translational Neuromodeling Unit An SPM Tutorial Institute for
MB-EPI PCASL. Release Notes for Version February 2015
 MB-EPI PCASL Release Notes for Version 1.0 20 February 2015 1 Background High-resolution arterial spin labeling (ASL) imaging is highly desirable in both neuroscience research and clinical applications
MB-EPI PCASL Release Notes for Version 1.0 20 February 2015 1 Background High-resolution arterial spin labeling (ASL) imaging is highly desirable in both neuroscience research and clinical applications
Analysis of fmri data within Brainvisa Example with the Saccades database
 Analysis of fmri data within Brainvisa Example with the Saccades database 18/11/2009 Note : All the sentences in italic correspond to informations relative to the specific dataset under study TP participants
Analysis of fmri data within Brainvisa Example with the Saccades database 18/11/2009 Note : All the sentences in italic correspond to informations relative to the specific dataset under study TP participants
The University of Chicago. Center for EPR Imaging in Vivo Physiology. Image Registration. Boris Epel
 The University of Chicago Center for EPR Imaging in Vivo Physiology Image Registration Boris Epel Imaging Methods are Complimentary CT MRI EPRI High resolution anatomic images Quantitative Poor soft tissue
The University of Chicago Center for EPR Imaging in Vivo Physiology Image Registration Boris Epel Imaging Methods are Complimentary CT MRI EPRI High resolution anatomic images Quantitative Poor soft tissue
CONN fmri functional connectivity toolbox
 CONN fmri functional connectivity toolbox Gabrieli Lab. McGovern Institute for Brain Research Massachusetts Institute of Technology http://www.nitrc.org/projects/conn Susan Whitfield-Gabrieli Alfonso Nieto-Castanon
CONN fmri functional connectivity toolbox Gabrieli Lab. McGovern Institute for Brain Research Massachusetts Institute of Technology http://www.nitrc.org/projects/conn Susan Whitfield-Gabrieli Alfonso Nieto-Castanon
Parameter estimation in arterial spin labeling MRI: comparing the four phase model and the buxton model with fourier transform
 Original Article Parameter estimation in arterial spin labeling MRI: comparing the four phase model and the buxton model with fourier transform Jim Ji 1, Vincent Pham 1, Xiao-Ping Zhu 2, Ka-loh Li 2 1
Original Article Parameter estimation in arterial spin labeling MRI: comparing the four phase model and the buxton model with fourier transform Jim Ji 1, Vincent Pham 1, Xiao-Ping Zhu 2, Ka-loh Li 2 1
Existing Packages. Overview. Siemens. Michael A. Chappell
 Existing Packages Michael A. Chappell michael.chappell@eng.ox.ac.uk www.ibme.ox.ac.uk/qubic Institute of Biomedical Engineering & Oxford Centre for Functional MRI of the Brain University of Oxford. Scanner
Existing Packages Michael A. Chappell michael.chappell@eng.ox.ac.uk www.ibme.ox.ac.uk/qubic Institute of Biomedical Engineering & Oxford Centre for Functional MRI of the Brain University of Oxford. Scanner
FSL Pre-Processing Pipeline
 The Art and Pitfalls of fmri Preprocessing FSL Pre-Processing Pipeline Mark Jenkinson FMRIB Centre, University of Oxford FSL Pre-Processing Pipeline Standard pre-processing: Task fmri Resting-state fmri
The Art and Pitfalls of fmri Preprocessing FSL Pre-Processing Pipeline Mark Jenkinson FMRIB Centre, University of Oxford FSL Pre-Processing Pipeline Standard pre-processing: Task fmri Resting-state fmri
Appendix E1. Supplementary Methods. MR Image Acquisition. MR Image Analysis
 RSNA, 2015 10.1148/radiol.2015150532 Appendix E1 Supplementary Methods MR Image Acquisition By using a 1.5-T system (Avanto, Siemens Medical, Erlangen, Germany) under a program of regular maintenance (no
RSNA, 2015 10.1148/radiol.2015150532 Appendix E1 Supplementary Methods MR Image Acquisition By using a 1.5-T system (Avanto, Siemens Medical, Erlangen, Germany) under a program of regular maintenance (no
2. Creating Field Maps Using the Field Map GUI (Version 2.0) in SPM5
 1. Introduction This manual describes how to use the Field Map Toolbox Version 2.0 for creating unwrapped field maps that can be used to do geometric distortion correction of EPI images in SPM5. 1. 1.
1. Introduction This manual describes how to use the Field Map Toolbox Version 2.0 for creating unwrapped field maps that can be used to do geometric distortion correction of EPI images in SPM5. 1. 1.
PBSI: A symmetric probabilistic extension of the Boundary Shift Integral
 PBSI: A symmetric probabilistic extension of the Boundary Shift Integral Christian Ledig 1, Robin Wolz 1, Paul Aljabar 1,2, Jyrki Lötjönen 3, Daniel Rueckert 1 1 Department of Computing, Imperial College
PBSI: A symmetric probabilistic extension of the Boundary Shift Integral Christian Ledig 1, Robin Wolz 1, Paul Aljabar 1,2, Jyrki Lötjönen 3, Daniel Rueckert 1 1 Department of Computing, Imperial College
Fmri Spatial Processing
 Educational Course: Fmri Spatial Processing Ray Razlighi Jun. 8, 2014 Spatial Processing Spatial Re-alignment Geometric distortion correction Spatial Normalization Smoothing Why, When, How, Which Why is
Educational Course: Fmri Spatial Processing Ray Razlighi Jun. 8, 2014 Spatial Processing Spatial Re-alignment Geometric distortion correction Spatial Normalization Smoothing Why, When, How, Which Why is
Basic fmri Design and Analysis. Preprocessing
 Basic fmri Design and Analysis Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial filtering
Basic fmri Design and Analysis Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial filtering
Functional MRI in Clinical Research and Practice Preprocessing
 Functional MRI in Clinical Research and Practice Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization
Functional MRI in Clinical Research and Practice Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization
腦部結構影像 標準化 組織分割 體素型態 本週課程內容. Analysis Softwares. A Course of MRI
 本週課程內容 腦部結構影像 A Course of MRI 盧家鋒助理教授國立陽明大學物理治療暨輔助科技學系 alvin4016@ym.edu.tw 腦部結構影像 空間標準化 (Spatial normalization) 均勻度校正 (Bias correction) 組織分割 (Segmentation) 體素形態學分析 (Voxel-based morphometry, VBM) 影像平滑化
本週課程內容 腦部結構影像 A Course of MRI 盧家鋒助理教授國立陽明大學物理治療暨輔助科技學系 alvin4016@ym.edu.tw 腦部結構影像 空間標準化 (Spatial normalization) 均勻度校正 (Bias correction) 組織分割 (Segmentation) 體素形態學分析 (Voxel-based morphometry, VBM) 影像平滑化
QUANTITATION OF THE PREMATURE INFANT BRAIN VOLUME FROM MR IMAGES USING WATERSHED TRANSFORM AND BAYESIAN SEGMENTATION
 QUANTITATION OF THE PREMATURE INFANT BRAIN VOLUME FROM MR IMAGES USING WATERSHED TRANSFORM AND BAYESIAN SEGMENTATION Merisaari Harri 1;2, Teräs Mika 2, Alhoniemi Esa 1, Parkkola Riitta 2;3, Nevalainen
QUANTITATION OF THE PREMATURE INFANT BRAIN VOLUME FROM MR IMAGES USING WATERSHED TRANSFORM AND BAYESIAN SEGMENTATION Merisaari Harri 1;2, Teräs Mika 2, Alhoniemi Esa 1, Parkkola Riitta 2;3, Nevalainen
Surface-based Analysis: Inter-subject Registration and Smoothing
 Surface-based Analysis: Inter-subject Registration and Smoothing Outline Exploratory Spatial Analysis Coordinate Systems 3D (Volumetric) 2D (Surface-based) Inter-subject registration Volume-based Surface-based
Surface-based Analysis: Inter-subject Registration and Smoothing Outline Exploratory Spatial Analysis Coordinate Systems 3D (Volumetric) 2D (Surface-based) Inter-subject registration Volume-based Surface-based
SPM8 for Basic and Clinical Investigators. Preprocessing. fmri Preprocessing
 SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
Statistical Analysis of Neuroimaging Data. Phebe Kemmer BIOS 516 Sept 24, 2015
 Statistical Analysis of Neuroimaging Data Phebe Kemmer BIOS 516 Sept 24, 2015 Review from last time Structural Imaging modalities MRI, CAT, DTI (diffusion tensor imaging) Functional Imaging modalities
Statistical Analysis of Neuroimaging Data Phebe Kemmer BIOS 516 Sept 24, 2015 Review from last time Structural Imaging modalities MRI, CAT, DTI (diffusion tensor imaging) Functional Imaging modalities
Introductory Concepts for Voxel-Based Statistical Analysis
 Introductory Concepts for Voxel-Based Statistical Analysis John Kornak University of California, San Francisco Department of Radiology and Biomedical Imaging Department of Epidemiology and Biostatistics
Introductory Concepts for Voxel-Based Statistical Analysis John Kornak University of California, San Francisco Department of Radiology and Biomedical Imaging Department of Epidemiology and Biostatistics
Cocozza S., et al. : ALTERATIONS OF FUNCTIONAL CONNECTIVITY OF THE MOTOR CORTEX IN FABRY'S DISEASE: AN RS-FMRI STUDY
 ALTERATIONS OF FUNCTIONAL CONNECTIVITY OF THE MOTOR CORTEX IN FABRY'S DISEASE: AN RS-FMRI STUDY SUPPLEMENTARY MATERIALS Sirio Cocozza, MD 1*, Antonio Pisani, MD, PhD 2, Gaia Olivo, MD 1, Francesco Saccà,
ALTERATIONS OF FUNCTIONAL CONNECTIVITY OF THE MOTOR CORTEX IN FABRY'S DISEASE: AN RS-FMRI STUDY SUPPLEMENTARY MATERIALS Sirio Cocozza, MD 1*, Antonio Pisani, MD, PhD 2, Gaia Olivo, MD 1, Francesco Saccà,
Advanced MRI Techniques (and Applications)
 Advanced MRI Techniques (and Applications) Jeffry R. Alger, PhD Department of Neurology Ahmanson-Lovelace Brain Mapping Center Brain Research Institute Jonsson Comprehensive Cancer Center University of
Advanced MRI Techniques (and Applications) Jeffry R. Alger, PhD Department of Neurology Ahmanson-Lovelace Brain Mapping Center Brain Research Institute Jonsson Comprehensive Cancer Center University of
DynaConn Users Guide. Dynamic Function Connectivity Graphical User Interface. Last saved by John Esquivel Last update: 3/27/14 4:26 PM
 DynaConn Users Guide Dynamic Function Connectivity Graphical User Interface Last saved by John Esquivel Last update: 3/27/14 4:26 PM Table Of Contents Chapter 1 Introduction... 1 1.1 Introduction to this
DynaConn Users Guide Dynamic Function Connectivity Graphical User Interface Last saved by John Esquivel Last update: 3/27/14 4:26 PM Table Of Contents Chapter 1 Introduction... 1 1.1 Introduction to this
SISCOM (Subtraction Ictal SPECT CO-registered to MRI)
 SISCOM (Subtraction Ictal SPECT CO-registered to MRI) Introduction A method for advanced imaging of epilepsy patients has been developed with Analyze at the Mayo Foundation which uses a combination of
SISCOM (Subtraction Ictal SPECT CO-registered to MRI) Introduction A method for advanced imaging of epilepsy patients has been developed with Analyze at the Mayo Foundation which uses a combination of
CS/NEUR125 Brains, Minds, and Machines. Due: Wednesday, April 5
 CS/NEUR125 Brains, Minds, and Machines Lab 8: Using fmri to Discover Language Areas in the Brain Due: Wednesday, April 5 In this lab, you will analyze fmri data from an experiment that was designed to
CS/NEUR125 Brains, Minds, and Machines Lab 8: Using fmri to Discover Language Areas in the Brain Due: Wednesday, April 5 In this lab, you will analyze fmri data from an experiment that was designed to
Automatic Generation of Training Data for Brain Tissue Classification from MRI
 Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. COCOSCO, Alex P. ZIJDENBOS, and Alan C. EVANS http://www.bic.mni.mcgill.ca/users/crisco/ McConnell Brain Imaging
Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. COCOSCO, Alex P. ZIJDENBOS, and Alan C. EVANS http://www.bic.mni.mcgill.ca/users/crisco/ McConnell Brain Imaging
Supplementary Figure 1
 Supplementary Figure 1 BOLD and CBV functional maps showing EPI versus line-scanning FLASH fmri. A. Colored BOLD and CBV functional maps are shown in the highlighted window (green frame) of the raw EPI
Supplementary Figure 1 BOLD and CBV functional maps showing EPI versus line-scanning FLASH fmri. A. Colored BOLD and CBV functional maps are shown in the highlighted window (green frame) of the raw EPI
CP Generalize Concepts in Abstract Multi-dimensional Image Model Component Semantics. David Clunie.
 CP-1390 - Generalize Concepts in Abstract Multi-dimensional Image Model Semantics Page 1 STATUS Date of Last Update Person Assigned Submitter Name Submission Date Assigned 2014/06/09 David Clunie mailto:dclunie@dclunie.com
CP-1390 - Generalize Concepts in Abstract Multi-dimensional Image Model Semantics Page 1 STATUS Date of Last Update Person Assigned Submitter Name Submission Date Assigned 2014/06/09 David Clunie mailto:dclunie@dclunie.com
A novel dementia diagnosis strategy on arterial spin labeling magnetic resonance images via pixel-wise partial volume correction and ranking
 Multimed Tools Appl (216) 75:267 29 DOI 1.17/s1142-14-2395-2 A novel dementia diagnosis strategy on arterial spin labeling magnetic resonance images via pixel-wise partial volume correction and ranking
Multimed Tools Appl (216) 75:267 29 DOI 1.17/s1142-14-2395-2 A novel dementia diagnosis strategy on arterial spin labeling magnetic resonance images via pixel-wise partial volume correction and ranking
EMSegment Tutorial. How to Define and Fine-Tune Automatic Brain Compartment Segmentation and the Detection of White Matter Hyperintensities
 EMSegment Tutorial How to Define and Fine-Tune Automatic Brain Compartment Segmentation and the Detection of White Matter Hyperintensities This documentation serves as a tutorial to learn to customize
EMSegment Tutorial How to Define and Fine-Tune Automatic Brain Compartment Segmentation and the Detection of White Matter Hyperintensities This documentation serves as a tutorial to learn to customize
Image Registration + Other Stuff
 Image Registration + Other Stuff John Ashburner Pre-processing Overview fmri time-series Motion Correct Anatomical MRI Coregister m11 m 21 m 31 m12 m13 m14 m 22 m 23 m 24 m 32 m 33 m 34 1 Template Estimate
Image Registration + Other Stuff John Ashburner Pre-processing Overview fmri time-series Motion Correct Anatomical MRI Coregister m11 m 21 m 31 m12 m13 m14 m 22 m 23 m 24 m 32 m 33 m 34 1 Template Estimate
EPI Data Are Acquired Serially. EPI Data Are Acquired Serially 10/23/2011. Functional Connectivity Preprocessing. fmri Preprocessing
 Functional Connectivity Preprocessing Geometric distortion Head motion Geometric distortion Head motion EPI Data Are Acquired Serially EPI Data Are Acquired Serially descending 1 EPI Data Are Acquired
Functional Connectivity Preprocessing Geometric distortion Head motion Geometric distortion Head motion EPI Data Are Acquired Serially EPI Data Are Acquired Serially descending 1 EPI Data Are Acquired
Brain Extraction, Registration & EPI Distortion Correction
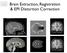 Brain Extraction, Registration & EPI Distortion Correction What use is Registration? Some common uses of registration: Combining across individuals in group studies: including fmri & diffusion Quantifying
Brain Extraction, Registration & EPI Distortion Correction What use is Registration? Some common uses of registration: Combining across individuals in group studies: including fmri & diffusion Quantifying
Spatial Filtering Methods in MEG. Part 3: Template Normalization and Group Analysis"
 Spatial Filtering Methods in MEG Part 3: Template Normalization and Group Analysis" Douglas Cheyne, PhD" Program in Neurosciences and Mental Health" Hospital for Sick Children Research Institute " &" Department
Spatial Filtering Methods in MEG Part 3: Template Normalization and Group Analysis" Douglas Cheyne, PhD" Program in Neurosciences and Mental Health" Hospital for Sick Children Research Institute " &" Department
SPM8 for Basic and Clinical Investigators. Preprocessing
 SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
White Matter Lesion Segmentation (WMLS) Manual
 White Matter Lesion Segmentation (WMLS) Manual 1. Introduction White matter lesions (WMLs) are brain abnormalities that appear in different brain diseases, such as multiple sclerosis (MS), head injury,
White Matter Lesion Segmentation (WMLS) Manual 1. Introduction White matter lesions (WMLs) are brain abnormalities that appear in different brain diseases, such as multiple sclerosis (MS), head injury,
A Graph Theoretical Network Analysis Toolbox
 A Graph Theoretical Network Analysis Toolbox Reference Manual for GRETNA (v2.0) June 2017 National Key Laboratory of Cognitive Neuroscience and Learning Beijing Key Laboratory of Brain Imaging and Connectomics
A Graph Theoretical Network Analysis Toolbox Reference Manual for GRETNA (v2.0) June 2017 National Key Laboratory of Cognitive Neuroscience and Learning Beijing Key Laboratory of Brain Imaging and Connectomics
COMETS v User s Manual
 COMETS v. 2.0 A Matlab toolbox for simulating electric field due to tdcs User s Manual Manual Written by Chany Lee Contact: cometstool@gmail.com Quick Guide Operation conditions Microsoft Windows environment
COMETS v. 2.0 A Matlab toolbox for simulating electric field due to tdcs User s Manual Manual Written by Chany Lee Contact: cometstool@gmail.com Quick Guide Operation conditions Microsoft Windows environment
Multimodal Imaging Brain Connectivity Analysis (MIBCA)
 Multimodal Imaging Brain Connectivity Analysis (MIBCA) Andre Santos Ribeiro, Luis Miguel Lacerda, Hugo Ferreira April 23, 2015 Abstract In recent years, connectivity studies using neuroimaging data have
Multimodal Imaging Brain Connectivity Analysis (MIBCA) Andre Santos Ribeiro, Luis Miguel Lacerda, Hugo Ferreira April 23, 2015 Abstract In recent years, connectivity studies using neuroimaging data have
MARS: Multiple Atlases Robust Segmentation
 Software Release (1.0.1) Last updated: April 30, 2014. MARS: Multiple Atlases Robust Segmentation Guorong Wu, Minjeong Kim, Gerard Sanroma, and Dinggang Shen {grwu, mjkim, gerard_sanroma, dgshen}@med.unc.edu
Software Release (1.0.1) Last updated: April 30, 2014. MARS: Multiple Atlases Robust Segmentation Guorong Wu, Minjeong Kim, Gerard Sanroma, and Dinggang Shen {grwu, mjkim, gerard_sanroma, dgshen}@med.unc.edu
Introduction to MRI data processing with FSL. Anna Blazejewska
 Introduction to MRI data processing with FSL Anna Blazejewska FSL = FMRIB Software Library FMRIB = Functional Magnetic Resonance Imaging of the Brain @ Oxford since 2000, last stable FSL 5.0, free! for
Introduction to MRI data processing with FSL Anna Blazejewska FSL = FMRIB Software Library FMRIB = Functional Magnetic Resonance Imaging of the Brain @ Oxford since 2000, last stable FSL 5.0, free! for
Resting State fmri Data Processing. Resting State fmri Data Processing 2018/8/21. Data Processing of Resting-State fmri: DPARSF DPARSF DPARSF DPABI
 DPARSF Data of Resting-State fmri: DPARSF Chao-Gan YAN, Ph.D. 严超赣 ycg.yan@gmail.com http://rfmri.org Institute of Psychology, Chinese Academy of Sciences 1 (Yan and Zang, 2010) 2 DPARSF Data Assistant
DPARSF Data of Resting-State fmri: DPARSF Chao-Gan YAN, Ph.D. 严超赣 ycg.yan@gmail.com http://rfmri.org Institute of Psychology, Chinese Academy of Sciences 1 (Yan and Zang, 2010) 2 DPARSF Data Assistant
Group (Level 2) fmri Data Analysis - Lab 4
 Group (Level 2) fmri Data Analysis - Lab 4 Index Goals of this Lab Before Getting Started The Chosen Ten Checking Data Quality Create a Mean Anatomical of the Group Group Analysis: One-Sample T-Test Examine
Group (Level 2) fmri Data Analysis - Lab 4 Index Goals of this Lab Before Getting Started The Chosen Ten Checking Data Quality Create a Mean Anatomical of the Group Group Analysis: One-Sample T-Test Examine
SPM Course! Single Subject Analysis
 SPM Course! Single Subject Analysis Practical Session Dr. Jakob Heinzle & Dr. Frederike Petzschner & Dr. Lionel Rigoux Hands up: Who has programming experience with Matlab? Who has analyzed an fmri experiment
SPM Course! Single Subject Analysis Practical Session Dr. Jakob Heinzle & Dr. Frederike Petzschner & Dr. Lionel Rigoux Hands up: Who has programming experience with Matlab? Who has analyzed an fmri experiment
Effect of age and dementia on topology of brain functional networks. Paul McCarthy, Luba Benuskova, Liz Franz University of Otago, New Zealand
 Effect of age and dementia on topology of brain functional networks Paul McCarthy, Luba Benuskova, Liz Franz University of Otago, New Zealand 1 Structural changes in aging brain Age-related changes in
Effect of age and dementia on topology of brain functional networks Paul McCarthy, Luba Benuskova, Liz Franz University of Otago, New Zealand 1 Structural changes in aging brain Age-related changes in
MIDAS Signal Calibration
 Contents MIDAS Signal Calibration A. Maudsley, 2004-2007 Introduction... 1 The Calibration Phantom... 2 Calibration Method... 4 Additional Considerations... 11 Acknowledgements... 12 Introduction The MR
Contents MIDAS Signal Calibration A. Maudsley, 2004-2007 Introduction... 1 The Calibration Phantom... 2 Calibration Method... 4 Additional Considerations... 11 Acknowledgements... 12 Introduction The MR
User s Guide Neuroimage Processing ToolKit (NPTK) Version.1.7 (beta) fmri Registration Software Pipeline for Functional Localization
 User s Guide Neuroimage Processing ToolKit (NPTK) Version.1.7 (beta) fmri Registration Software Pipeline for Functional Localization Software Written by Ali Gholipour SIP Lab, UTD, 2005-2007 Revision 1.7
User s Guide Neuroimage Processing ToolKit (NPTK) Version.1.7 (beta) fmri Registration Software Pipeline for Functional Localization Software Written by Ali Gholipour SIP Lab, UTD, 2005-2007 Revision 1.7
