Interfacing Guide. Interfacing Guide. Version September, Page i
|
|
|
- Oliver Alexander
- 5 years ago
- Views:
Transcription
1 Version September, 2000 Page i
2 The software described in this book is furnished under a license agreement and may be used only in accordance with the terms of that agreement. Copyright Notice Micromass Ltd believes that the information in this publication is accurate. However the information is subject to change without notice and should not be construed as a contractual undertaking by Micromass Ltd. Despite the care which has been given to the preparation of this publication, Micromass Ltd accepts no responsibility for any loss or any other matter which may arise from any error or inaccuracy which may inadvertently have been included. Copyright (C) Micromass Ltd. All Rights Reserved. No part of this publication may be copied without the express written permission of Micromass Ltd. Trademarks Micromass is a registered trade mark of Micromass Limited (Reg. U.S. Pat. & Tm. Off.). Windows is a trademark of Microsoft Corporation. Other product names mentioned in this manual may be trademarks or registered trademarks of their respective companies and are hereby acknowledged. Page ii
3 Table of Contents Importing 5 Copying Data 5 Import Worksheet 6 To Import a Worksheet 6 Creating Import Files 7 Access 97 7 Excel 8 Notepad 8 Import Data 8 To Import Data 8 Creating Import Files 9 Access 97 9 Excel 9 Notepad 9 Exporting 10 Export to LIMS File 10 Copying to and from the Windows NT Clipboard 13 Export SEQUEST file 15 OpenLynx Batch Files 16 Paragraphs 16 Batch Block 17 Plate Block 17 Sample Block 18 Fields 18 Comments 18 File Format 19 OpenLynx Browser 23 Sections 23 Fields 23 Comments 23 Tabulated Sections 24 Free Format Text Sections 24 File Section Descriptions 25 MetaboLynx Browser 34 Sections 34 Fields 34 Comments 34 Tabulated Sections 35 Free Format Text Sections 35 File Section Descriptions 36 ProteinLynx Browser 46 Sections 46 Fields 46 Tabulated Sections 46 File Section Descriptions 47 Bio-Rad File Descriptions 53 File Formats 56 Sample List 56 An Example Generic Sample List File 59 AutoLynx 60 Status.ini 62 File Format 63 Page 1
4 Index 69 Page 2
5 Conventions The follows these typographic conventions This bold italic ALL CAPITALS Represents Anything you must type exactly as it appears Place holders for information you must provide. For example if you are asked to type filename, you would type the actual name for a file instead of the word shown in italic type. Directory names, filenames and acronyms. Keyboard Formats Key combinations and key sequences appear in the following formats. KEY1+KEY2 KEY1,KEY2 A plus sign (+) between key names means to press and hold down the first key while you press the second key. For example, "press ALT+ESC" means to press and hold down the ALT key and press the ESC key. Then release both keys. A comma sign (,) between key names means to press and release the keys one after the other. For example, "press ALT,F" means to press and release the ALT key and then press the F key. Page 3
6 About This Manual This manual is designed to explain how data can be imported into and exported from MassLynx for use in other applications. When you have read this manual you should be able to - Import data from Excel, Access and Notepad. Export quantify data to LIMS systems. Read ProteinLynx, OpenLynx and MetaboLynx report files. Read Chromatogram and Spectrum text files. This manual assumes that you have no previous knowledge of MassLynx. However this manual does assume that you are familiar with using the Microsoft Windows NT Graphical Environment and have the basic skills required to work with Windows NT software. If you have never used Microsoft Windows NT before we would suggest that you spend a short time reading Chapter 2 Learning the Basics in the Microsoft Windows NT Workstation Start Here guide supplied with your Windows NT software. This manual also assumes knowledge of Access, Excel, Notepad and LIMS systems. Page 4
7 Importing Copying Data There are several ways of importing data into a Sample List: Copying and pasting data from text editors, Access and Excel. Import Worksheet. Import Data. Data created in other Windows applications can be copied to the clipboard and pasted into the Sample List editor. 1. Press the button or select New from the File menu. If the previous Sample List has not been saved you will be prompted to save it. A Sample List with one default row will be displayed. 2. Add rows and columns to the Sample List so that it matches the number of rows and columns as the other Windows application. Note if this is not done data may be lost. 3. Select the relevant area in the other Windows application and copy it. 4. In the Sample List editor, position the cursor on the cell at the top left corner of the paste area and select Paste. Notes: If copying from a text editor e.g. Notepad fields must be tab delimited. When copying from an Access database the last record is not pasted and will have to be entered manually. Page 5
8 Import Worksheet Sample Lists can be created in a number of other packages and imported into MassLynx. MassLynx V3.0 and V3.1 allowed OpenLynx batch files and MassLynx V2.3 Sample List files to be imported. While these options are still supported, there are now several other file types that have been added. ACCESS 97 To Import a Worksheet Tab and Comma delimited text files Excel 97 and Excel 5.0, 6.0 and 7.0 files 1. Choose Import Worksheet from the MassLynx File menu. Figure 1 The Import Worksheet dialog 2. Locate the required file, or type in a file name, and press Open. Generic Batch files created in OpenLynx are the default file types. Click on the arrow, at the end of the Files of type box, to import a different file type. Page 6
9 Creating Import Files Access 97 The following is a list of instructions on how to create files suitable for importing into MassLynx. For all types of file fields must have the same name as in the Sample List (see Sample List on page 56) although they can be defined in any order. For Access 97 the data type must also match. The names correspond to the name in brackets on the Customize Field Display dialog. When the table is created it must be called ANALYSIS. It is recommended that the design view is used when creating a new table, this allows you to define the data type of the field. You must have the first column called Index as the primary key. Column headings must match those shown on page 56. Other columns can be present but they will not be imported into the Sample List. n To define the data type as a double 1. Select Number from the drop down list box in the Data Type column. 2. On the general page, at the bottom left of the screen, click on Field Size and select Double from the drop down list box. n To save in access 97 format The table can be saved as an access database by selecting Save from the File menu and can be imported into MassLynx in this format. Tables can also be saved as tab or comma delimited files for importing into MassLynx. n To save in tab or comma delimited format 1. Select Save As/Export from the File menu, select the To an External File or Database option and press OK. 2. Select the required directory from the browser displayed, select Text files (*.txt;*.csv;*.tab;*.asc) from the Save as type drop down list box and then press the Export button. 3. Make sure the Delimited option is selected and press the Next button. 4. Check the Include Field Names on First Row option, select the type of delimiter to use and press the Next button. 5. Enter the name to save the file as and press the Finish button. If files are saved as comma or tab delimited then they must be imported into MassLynx as comma or tab delimited files. Page 7
10 Excel You must have the first column called Index, other column headings must match those shown on page 56. Select the area containing the data to be imported, including the column headings, and name the area ANALYSIS. To do this select Define from the Name option on the Insert menu, type ANALYSIS and press OK. Leave all cells in General format For a text field containing only numeric data an apostrophe ( ) must be inserted in front of the number. If the file is to be saved as tab or comma delimited then Excel will only allow one sheet to be saved. If the current workbook contains more than one worksheet then each worksheet must be saved as a separate text file. Notepad Import Data You must have the first column called Index, other column headings must match those shown on page 56. Type in the field name/value and then a comma (or press tab for tab delimited files) and enter the next value. End each line with a carriage return. Text fields should be enclosed in quotes. To Import Data Sample List data can be created in a number of other packages and imported into MassLynx. The file types supported are: ACCESS 97 Tab and Comma delimited text files Excel 97 and Excel 5.0, 6.0 and 7.0 files 1. In MassLynx ensure that the correct number of rows and columns is displayed. If this is not done then data will be lost. 2. Choose Import Data from the MassLynx File menu. 3. Locate the required file, or type in a file name, and press Open. Excel 5.0 files are the default file types. Click on the arrow, at the end of the Files of type box, to import a different file type. Page 8
11 Creating Import Files Access 97 The following is a list of instructions on how to create files suitable for importing into MassLynx. For all types of file Fields must not have column headings. Fields must be in the same order as they are to appear in the MassLynx Sample List Excel When the table is created it must be called ANALYSIS. For Access 97 the data type of the column must match. It is recommended that the design view is used when creating a new table, this allows you to define the data type of the field. To define the data type as a double see Import Worksheet above. To save in tab or comma delimited format follow the instructions for Import Worksheet above, except for step 4 where the Include Field Names on First Row option should not be checked. If files are saved as comma or tab delimited then they must be imported into MassLynx as comma or tab delimited files. Notepad Select the area containing the data to be imported, including the column headings, and name the area ANALYSIS. To do this select Define from the Name option on the Insert menu, type ANALYSIS and press OK. Leave all cells in General format For a text field containing only numeric data an apostrophe ( ) must be inserted in front of the number. If the file is to be saved as tab or comma delimited then Excel will only allow one sheet to be saved. If the current workbook contains more than one worksheet then each worksheet must be saved as a separate text file. Type in the field name/value and then a comma (or press tab for tab delimited files) and enter the next value. End each line with a carriage return. Text fields should be enclosed in quotes. Page 9
12 Exporting Export to LIMS File There are several ways of exporting data from MassLynx: Export to LIMS. Copying Chromatogram and Spectrum peak lists. Export SEQUEST file. OpenLynx Batch files. OpenLynx Browser files. ProteinLynx Browser files. Quantification results can be written to a text file for use with LIMS systems. This can be performed automatically by selecting the Export Results to LIMS option on the Quantify Samples dialog. The results can also be exported from the Quantify window. Select Export to LIMS File from the File menu, select a file from the browser displayed or enter the name of a new one and press Save. If the selected file already exists, the user will be prompted to overwrite the existing file. The file generated will consist of three areas; the Header Section, the Samples Section and the Calibration section. The Header Section The header section contains the following four sections. Each shows the full path name of the file generated by or used to create the report and the date and time that the file was last modified. All fields are text. LIMS EXPORT FILE The LIMS file generated SAMPLELIST The Sample List file. QUANMETHOD The quantification method file. QUANCALIBRATION The quantify curve file. e.g. LIMS EXPORT FILE: C:\Masslynx\lims1 Tue Nov 10 11:33: Compound Section header fields The compound section will include an entry for each sample in the current sample list. The header will contain the File Name, Sample ID and any text entered in the SPARE_1 to SPARE_5 fields in the sample list. E.g. ASSAY01,ID,,,,, Page 10
13 Samples Section For each sample there will be one entry for each compound named in the compound box in the quantify method. Each entry will have the following fields, separated by a comma. The data type column shows what type of field the data was exported from. Field Description The compound number shown in the compound box in the quantification method. The text name of this compound. The scan at which the matching peak was found in the current sample datafile. The retention time of the matching peak. The relative retention time at which the matching peak was found. The area of the matching peak. The height of the matching peak. The response of the sample for this compound. The flags associated with the peak. The concentration of compound recorded for this sample. The blank subtracted concentration of the compound for this sample. The chromatogram trace used to locate peaks for this compound. The error between the expected concentration and the calculated concentration. The ordinal number of the compound in the quantification method that is used as the reference peak for this compound. The area of the reference peak used for this compound for this sample. The height of the reference peak The retention time of the reference peak. The modification date of the peak used to quantify this compound for this sample. This refers manual modification of the peak. The modification time of the peak. The modification text (modification comment) of the peak. The MassLynx user who altered the peak. The mass of the peak. The retention time the peak was expected at for this compound. Data Type int char Int Float Float Float Float Float Text Float Float Char Float Int Float Float float Text Text Text Text Float Float Page 11
14 Field Description The relative retention time the peak was expected at for this compound. The user factor associated with this compound. The user RF factor associated with this compound. The start retention time of the detected peak The end retention time of the detected peak Data Type Float Float Float Float Float The Calibration Section The calibration section will have a subsection for each calibration curve calculated for the current quantification calibration. Each subsection will contain information as displayed on the calibration graphs window. Where a line entry is inappropriate it will not be entered in the report file. Correlation coefficient: or Coefficient of Determination: Response Factor: or Calibration Curve: Response Type: Curve Type:, Origin:, and Weighting: E.g. Compound 1 name: I. Std Response Factor: Response type: External Std, Area Curve type: RF Compound 2 name: Parent Coefficient of Determination: 1 Calibration curve: * x Response type: Internal Std ( Ref 1 ), Area * ( IS Conc. / IS Area ) Curve type: Linear, Origin: Exclude, Weighting: 1/x, Axis trans: None Compound 3 name: Metabolite Coefficient of Determination: 1 Calibration curve: * x Response type: Internal Std ( Ref 1 ), Area * ( IS Conc. / IS Area ) Curve type: Linear, Origin: Exclude, Weighting: 1/x, Axis trans: None All fields are text. It is possible to parse the correlation coefficients out, but their format will depend on how the quantify methods are set up, so this will have to be done on a per application basis. Page 12
15 Copying to and from the Windows NT Clipboard The Windows NT Clipboard provides temporary storage for information that is being transferred between application programs (word processors, spreadsheets, MassLynx etc.). You can use the Clipboard to move data out of the Chromatogram and Spectrum windows as a text list. This data can then be pasted into reports written with a Windows compatible word processor. n To copy a chromatogram as a text list to the Clipboard 1. Display the required time range in a chromatogram window. 2. Press the Toolbar button or choose Copy Chromatogram List from the Chromatogram Edit menu. The section of the chromatogram on display will be transferred to the Clipboard as (time, intensity) pairs or (scan, intensity) pairs depending on the horizontal axis setting. 3. To read the information into another application, choose Paste from the other application s Edit menu. Copying the Chromatogram list to the clipboard gives the following format. Retention Time or Scan Number Retention Time or Scan Number Retention Time or Scan Number Retention Time or Scan Number Intensity Intensity Intensity Intensity The retention time or scan/intensity pairs are separated by a tab. n To copy integrated chromatogram peaks as a text list to the Clipboard 1. Display the required time range in a chromatogram window 2. Press the Toolbar button or choose Copy Detected Peaks from the Chromatogram Edit menu. The chromatogram peaks on display will be transferred to the Clipboard. The information transferred for each peak is the peak top, height, area, start, end, start height and end height. 3. To read the information into another application, choose Paste from the other application s Edit menu. Copying the Chromatogram list to the clipboard gives the following format. Retention Time Peak Height Peak Area Peak Start RT Peak End RT Peak Start Intensity Peak End Intensity Retention Time Peak Height Peak Area Peak Start RT Peak End RT Peak Start Intensity Peak End Intensity The fields are separated by a tab. Page 13
16 n To copy a spectrum as a text list to the Clipboard 1. Display the required mass range in a Spectrum window. 2. Press the Toolbar button or choose Copy Spectrum List from the Spectrum Edit menu. The section of the spectrum on display will be transferred to the Clipboard as (mass, intensity) pairs. 3. To read the information into another application, choose Paste from the other application s Edit menu. Copying the Spectrum list to the clipboard gives the following format. Mass Mass Mass Mass Intensity Intensity Intensity Intensity The mass/intensity pairs are separated by a tab. Page 14
17 Export SEQUEST file MassLynx has a facility to convert files into a format which can be used by the SEQUEST program. The SEQUEST program correlates uninterpreted tandem mass spectra of peptides with amino acid sequences from protein and nucleotide databases. It is written by Jimmy Eng and John Yates (University of Washington). Further details can be obtained via the Internet at n To Export a SEQUEST file 1. Display the relevant centered MS/MS data file select Export SEQUEST file from the File menu. Figure 2 Export SEQUEST dialog 2. The Precursor ion mass is picked up from the data file, if it was entered in the Function Editor, otherwise type in a value. 3. The Precursor charge state defaults to 2, change this as required. 4. Enter any known sequence information in the Partial sequence info box. 5. File Path and Destination is the location and filename that the file will be saved to. The file name is the original file name with the scan and function numbers appended to it. To change the destination press the Browse button and select a new destination from the dialog displayed, or type a new one in. 6. Press the OK button. Note: this option is only enabled if BioLynx is installed. The file produced is an ASCII text file with the following format. Precursor mass Charge Mass Intensity Mass Intensity Mass Intensity Mass Intensity The mass/intensity pairs are separated by a space. Page 15
18 OpenLynx Batch Files Paragraphs OpenLynx Batch files ( *.olb) are produced when samples are submitted from the OpenLynx Login program. When created they are stored in the Masslynx\Openlynx\BatchDB folder. When they have been acquired and processed they are moved to the Masslynx\Openlynx\BatchDB\Processed folder. The file will hold section blocks of data called paragraphs. A file will have a Batch paragraph and at least one Plate paragraph, followed by a paragraph for each sample to be analysed. Where there is more than one plate in an analysis, there will be a paragraph for each plate followed by sample paragraphs for each sample. A paragraph will always start with the name of the section surrounded by square parentheses on a line by itself. i.e. [SectionName]. e.g. [SectionName1]...Data Item Data Item 2 [SectionName2].Data Item 1.Data Item 2 A typical multi-plate analysis file would be arranged as follows [Batch].Data Item 1.Data Item 2 [Plate:1]...Data Item Data Item 2 [Sample:1.1]...Data Item Data Item 2 [Sample:1.n]...Data Item Data Item 2 [Plate:2]...Data Item Data Item 2 [Sample:2.1]...Data Item Data Item 2 [Sample:2.n]...Data Item Data Item 2 [Plate:m]...Data Item Data Item 2 Page 16
19 [Sample:m.1]...Data Item Data Item 2 [Sample:m.n]...Data Item Data Item 2 Batch Block The first paragraph in the file will contain information that is pertinent to the whole batch. MSMethod MSTune LCMethod EconomyScheduling PriorityScheduling AnalysisTime ProcessParameters Process UserName BatchID UserAddress SampleReportEnable NumberOfPlates Plate Block A file will contain a plate block for each plate which will contain information that is pertinent to the whole plate rather than a specific well. Within this plate block will be information concerning the plate description, plate ID and wells used. Where samples are not being presented in a plate this block can be used to describe the batch of samples loaded into an autosampler carousel. JobCode UserName Rows Columns Track NumberOfWells Page 17
20 Sample Block Fields For each sample to be analysed there will be a sample block that contains well position, and other information, specific to a certain sample. Well SampleDescription Conditions SampleID MSDataName SampleType MSInjectionVolume NumberOfMasses Mass1 NumberOfFractions NumberOfWaveLengths NumberOfConcs NumberOfFormulae NumberOfFactors NumberOfUserFields Sections will have a group of fields, each field will be on one line. The field name will be the first item on the line, followed by = and then the data item for that field. The type of data will depend on the field type. i.e. e.g. [SectionName] FieldName1=Data1 FieldName2=Data2... [Plate:3] JobCode=JB NumberOfWells=96 Comments A comment line is denoted by the first character being a semi-colon. A comment line can be any where in a section. Multiple line comments must have a semi-colon preceding each line. Page 18
21 File Format [Batch] When creating files manually the fields marked an (M) must be defined, all other fields are optional. Field/Section Name Data Type Description NumberOfPlates (M) Integer Number of plates in analysis. LabName Text Laboratory name. MSMethod Text The name of the MS method to be used. MSTune Text The name of the MS Tune file to be used. If not specified the current instrument tuning parameters will be used. LCMethod Text The name of the acquisition Inlet method to be used. InletPreRunMethod Text The name of the acquisition Inlet prerun method to be used. If not specified no prerun is performed. InletPostRunMethod Text The name of the acquisition Inlet postrun method to be used. If not specified no postrun is performed. InletSwitchMethod Text The name of the acquisition Inlet column switch method to be used. If not specified no switch method is performed. Process Text Path of External Program to be executed for sample processing. Directory is not necessary if file exists in current directory. ProcessParameters Text Path of Parameter File to be used by External Program. EconomyScheduling Integer None zero if batch is to be run as a Night-Time task. Otherwise will be scheduled normally. PriorityScheduling Integer None zero if batch is to be run as a Priority task. Otherwise will be scheduled normally. AnalysisTime Float Time of analysis for each sample in the batch. Equals zero if no time is specified. BatchID Text Name of batch. UserName Text Name of user submitting batch. UserAddress Text address to send batch results to. ReportScheme Text Name of the OpenLynx Browser report scheme to use when formatting output results. SampleReportEnable Boolean Indicates if per sample printing is required. 0 = False, 1 = True HPLCMethod Text Path of HPLC method parameter override file if one exists. Page 19
22 [Plate:N] Where N = Plate Number Field/Section Name Data Type Description PlateID Text Plate Identifier. A text description of the plate can be defined in files created externally. This is not generated in files created from the OpenLynx Login program. PlatePosition Text The description of the plate position, as co-ordinates for plate position in the autosampler bed. 1. X,Y. 2. Not used for a linear autosampler e.g. HP GC type. Where X and Y are integers. Origin Text The description of the start well position, as a pair of co-ordinates for well position in plate, or an absolute position WellAwellX(or wellxwella ) e.g. A1 JobCode Text Job or Batch Identifier. Where X is an integer and A is an alphabetical character. Or TL=Top Left, TR=Top Right, BL=Bottom Left, BR=Bottom Right UserName Text Name of operator performing analysis. Rows (M) Integer Number of rows on plate Columns (M) Integer Number of columns on plate Track (M) Text R=Rows, C=Columns, RS=Rows Snake, CS=Columns Snake NumberOfWells (M) Integer Number of wells used in plate or vials in an autosampler tray Page 20
23 [Sample:N:M] Where N = Plate Number, M = Well Number Field/Section Name Data Type Description Well (M) Text The description of the well position, as co-ordinates for well position in plate, or an absolute position. There are two formats for a Well. This field can be used to identify a vial position in a non XY autosampler. SampleType Integer Sample type 1. wellawelly (or wellywella ) e.g. B6, 6B or 2,6. 2. Absolute rack position. e.g. 18 Where X and Y are integers and A is an alphabetical character. 0 = Unknown, 1 = Standard, 2 = QC and 3 = Blank. NumberOfMasses Integer The number of masses defined. Mass1 Float 1 st mass to be used for targeting. This could be the monoisotopic mass or the mass with the greatest intensity, derived from the formula s isotope cluster. If the analysis being performed by the MS requires an expected molecular weight then this field is Mandatory unless a formula is supplied. Mass2 Float 2 nd Mass. MassN Float N th Mass. NumberOfFraction Triggers Integer The number of fractions defined. Only one fraction mass is currently supported by MassLynx. Fraction1 Float 1 St Mass to be monitored by Fraction Collection system. Fraction2 Float Second Fraction Mass. Not currently used. FractionN Float N th Fraction Mass NumberOfFormulae Integer The number of formulae defined. Formula1 Text The 1 st chemical composition, if used. Formula2 Text The 2 nd chemical composition FormulaN Text The N th chemical composition NumberOfWavelengths Integer The number of wavelengths defined. Wavelength1 Text The 1 st wavelength, if used. Wavelength2 Text The 2 nd wavelength Page 21
24 Field/Section Name Data Type Description WavelengthN Text The N th wavelength NumberOfConcs Integer The number of concentrations defined. Conc1 Text The 1 st concentration, if used. Conc2 Text The 2 nd concentration ConcN Text The N th concentration NumberOfFactors Integer The number of user factors defined. Factor1 Float 1 St User Factor if required. MassLynx currently only supports 1 User Factor. Factor2 Float The 2 nd User Factor. Not currently used. FactorN Float N th User Factor. Not currently used. SampleID Text Sample Identifier SampleDescription Text Sample Description Conditions Test Sample conditions MSDataName Text The name of the MS data file to be processed and/or created. MSInjectionVolume Float The amount of sample to inject, for an MS analysis. NumberOfUserFields Integer The number of user fields defined. User1 Text 1 St User Field, if required. User2 Text The 2 nd User Field. UserN Text N th User Field. Page 22
25 OpenLynx Browser Sections The file will hold section blocks of data. The start of a section block will always start with the name of the section surrounded by square parentheses on a line by itself. i.e. [SectionName]. The main body of the data will be enclosed by a pair of braces, the opening and closing braces will be the only characters on their respective lines. i.e. [SectionName] {...Data... } Section blocks can have section blocks embedded in them, in some cases multiple instances of sections with the same name can be allowed, depending on the type of data. e.g. [SectionName1] {...Data... [SectionName2] {...Data... } } Fields Sections will have a group of fields, each field will be on one line. The field name will be the first item on the line, followed by a list of TAB separated data items for that field. The type of data will depend on the field type. i.e. [SectionName] { FieldName1 FieldName2... } Data1 Data2_1Data2_2 Comments A comment line is denoted by the first character being a semi-colon. A comment line can be any where in a section. Multiple line comments have a semi-colon preceding each line. Page 23
26 Tabulated Sections Some sections can be considered to be a table, or list, of fields of the same type. Tabulated sections do not have any field names to describe the data but will normally have a comment line describing the names of the fields. e.g. [TableSection] { ;Field1 Field2 Data1_1Data1_2 Data2_1Data2_2... } Free Format Text Sections The information in these sections contains free format text which may stretch over several lines. A line will be terminated with a <carriage return><linefeed> pair. Page 24
27 File Section Descriptions SAMPLE Field/Section Name Data Type Description Sample Integer The number of the sample in the batch. Well Text The description of the well position, as a pair of co-ordinates for plate position in rack and well position in plate, or an absolute position. There are three formats for Well. 1. platex,platey:wellx,welly 2. platex,platey:wellawell1 (or well1wella ) 3. Absolute rack position. Where X and Y are integers and A is an alphabetical character. FileName Text The MassLynx NT MS data file name. SampleType Integer Sample type: Unknown = 0, Standard = 1, QC = 2 or Blank = 3. SampleID Text MassLynx Sample List input field. SampleDescription Text MassLynx Sample List input field. Date Text Date that MS data file was acquired. Time Text Time that MS data file was acquired. JobCode Text MassLynx Sample List input field. TaskCode Text MassLynx Sample List input field. UserName Text Current MassLynx user name. LabName Text Laboratory name. Instrument Text Instrument name. Conditions Text MassLynx Sample List input field. Submitter Text MassLynx Sample List input field. Plate Text Description of micro-titre plate, if applicable. See below for detailed description. INLET PARAMETERS Section Description of compound for testing. COMPOUND Section Description of compound for testing. FRACTION Section Results of any FractionLynx fraction collection. FUNCTION Section MS function data and test results. There will be one FUNCTION section per MassLynx MS and DAD data functions. ANALOG Section Analogue chromatographic data. Page 25
28 PLATE FIELD For samples acquired using a Gilson or Waters 2700 autosampler the following information will be written to the file, otherwise this field will be blank. Version Number ( two digits ) Origin Location ( TL - top left, TR - top right, BL - bottom left, BR - bottom right ) Priority ( XY - x coordinate before y coordinate, YX - y coordinate before x coordinate ) Reference scheme ( LN - letter number, NL - number letter, NN - number number, LL - letter letter, SC - sequential continuous, SD sequential discontinuous ) 1:Number Of Rows 2:Number Of Columns 3:Row Spacing 4:Column Spacing 5:Row Offset 6:Column Offset 7:Which Rows Offset ( O or E ) 8:Which Columns Offset ( O or E ) 9:Plate Width 10:Plate Height 11:Vial/Well Depth 12:Vial/Well Diameter 13:Vial One X 14:Vial One Y Note that all spacings, etc are given to 0.1 of a mm. Page 26
29 e.g. 01,TL,XY,LN,1:8,2:12,3:120.0,4:120.0,5:0.0,6:0.0,7:O,8:O,9:1250.0,10:870.0,11:4 00.0,12:90.0,13:80.0,14:130.0 Which means: Version = 01. Origin Location = top left. Priority = x coordinate before y coordinate. Reference scheme = Letter Number. 1:Number Of Rows = 8. 2:Number Of Columns = 12. 3:Row Spacing = 12mm. 4:Column Spacing = 12mm. 5:Row Offset = 0.0mm. 6:Column Offset = 0.0mm. 7:Which Rows Offset ( O or E ) = Odd numbers, but because Row Offset = 0, this has no effect. 8:Which Columns Offset ( O or E ) = Odd numbers, but because Column Offset = 0, this has no effect. 9:Plate Width = 125mm. 10:Plate Height = 87mm. 11:Vial/Well Depth = 40mm. 12:Vial/Well Diameter = 9mm. 13:Vial One X = 8.0mm. 14:Vial One Y = 13.0mm. Page 27
30 COMPOUND Field Name Data Type Description Mono Mass Float The mass used for targeting. This could be the mono-isotopic mass or the mass with the greatest intensity, derived from the formula s isotope cluster. Formula Text The chemical composition, if entered. Name Text The name of the compound. Unused at present. FRACTION Field Name Data Type Description Mass Float The mass collected. Org. Target Float The original mass minus any adducts. Ion Mode Text The ionisation mode used to collect the fraction. Start Time Float The time at which the fraction collection started. End Time Float The time at which the fraction collection ended. Start Site Text The name of the site at which the fraction collection started. No. Of Tubes Integer The number of tubes the fraction was collected in. INLET PARAMETERS FUNCTION This is a free format text section. Field/Section Name Data Type Description Function Integer The MassLynx MS data function number. IonMode Text The type of ionisation mode e.g. AP+. Type Text The type of MS scan function e.g. MS for full scan. Description Text A brief description of the MS scan function. SPECTRUM Section MS data and results of testing. There can be more than one SPECTRUM section in a function. CHROMATOGRAM Section Chromatographic data. There can be more than one CHROMATOGRAM section in a function. Page 28
31 SPECTRUM Field/Section Name Data Type Description ProcDesc Text Description of processing performed on spectrum. E.g. Combine (54:56-(38:39+74:75)) Process Integer The number of the MassLynx saved mass spectrum. Only present if the processed spectrum was saved with data file. State Text The result of testing the quality of the mass spectral data. Peak ID Integer Spectrum Peak Identification number. Peak Ref Integer Peak Reference Number. Time Float The retention time of the mass spectrum. TIC Float Total Ion Current, the sum of all the peak intensities in the spectrum. BPI Float The intensity of the base peak in the mass spectrum. BPM Float The mass of the base peak in the mass spectrum. Continuum Text TRUE if the spectrum is stored in continuum format. Otherwise FALSE. Saved Text TRUE if the processed spectrum was saved with the data file. Otherwise FALSE. MASSES Section Section describing the results of testing, and finding significant masses. This includes all adducts. RESULTS Section Section describing the results of the targeting of the expected compound. Masses from the same compound are combined into one item. MS Section Section that holds all the mass/intensity pairs, for this mass spectrum. SEARCH Section Section holding the results of a library search on this spectrum. This section will not be created if no search exists. Page 29
32 MASSES Field Name Data Type Description Mass Float The input mass. Exp. Mass Float The expected mass due to adduction. Obs. Mass Float The mass that was actually observed. % BPI Float The intensity of the mass peak as a percentage of the base peak intensity. Int. Abs Float The absolute intensity of the MS peak. RESULTS Field Name Data Type Description Mass Float The input mass. Found Boolean 1 if any of the possible adduct ions have been found for this compound, otherwise 0 % BPI Float The sum of the percentage intensities for the compound. % Purity Float A measure of the contribution of this compound to the spectrum as percentage of the TIC. This includes all MS peaks associated with this compound. Confirmed Boolean False if any of the confirmation tests made failed. If not specified regarded as being True. Status Text Description of confirmation failure. Not present if confirmed = TRUE MS Field Name Data Type Description Mass Float Observed mass of peak. % BPI Float Intensity of peak as a percentage of the base peak. Page 30
33 ELEMENTAL Contains results of elemental calculations if defined, otherwise this section does not appear. Field Name Data Type Description Mass Float Actual observed mass for which elemental composition was searched. Calc. Mass Float Calculated mass of reported formula mda Float Difference between observed mass and calculated mass in milli-daltons PPM Float Difference between observed mass and calculated mass in parts per million DBE Float Double Bond Equivalence. Formula Text Representation of calculated chemical formula. E.g. C19 H14 O2 SEARCH Contains Spectrum library search results if library search was defined, otherwise this section does not appear. Field Name Data Type Description Library Text Name of library in which the search was made. HIT Section Contains information about a library hit. There can be several or no hits for each search HIT Field Name Data Type Description Entry Integer Library entry number hit was made against. For Integer Forward search match value. Value between 0 and Rev Integer Reverse search match value. Value between 0 and Name Text Library entry name F1 Float Library entry general purpose filter 1 value. F2 Float Library entry general purpose filter 2 value. Mass Float Library entry mass value. Page 31
34 CHROMATOGRAM Field/Section Name Data Type Description TraceNumber Integer Number of chromatogram trace. Description Text The description of the chromatogram type. ProcDesc Text Description of processing performed on chromatogram. E.g. Smooth (Mn, 2x2) MaxIntensity Float The absolute intensity of the point with maximum intensity in the chromatographic data. TRACE Section A section that holds the chromatographic data. PEAK Section Section(s) that hold information about peak(s) detected in the chromatogram. TRACE Field Name Data Type Description Time Float The time of the chromatographic point. Int. %Max Float Intensity of the chromatographic point as a percentage of the maximum intensity. Page 32
35 PEAK Field Name Data Type Description Peak ID Integer Peak Identification number. Peak Ref Integer Peak Reference Number. Time Float Peak retention time in decimal minutes. Peak Float + Float Peak start and end retention times in decimal minutes. Two floats separated by a TAB character. Intensity Float + Float Peak baseline start and end intensity. Two floats separated by a TAB character. Height Float Detected peak height. AreaAbs Float Detected peak area. Area %BP Float Detected peak area as percentage of the largest peak in the chromatogram. Area %Total Float Detected peak area as percentage of the sum of all the peak areas in the chromatogram. Width Float Peak width in decimal minutes. RT Index Float Retention time index for peak. Field only output if RT Index has been calculated. RT LogP Float Retention time LogP for peak. Field only output if RT LogP has been calculated. Calc Conc Float Calculated concentration for peak. Field only output if calculated. Calc Amount Float Calculated amount for peak. Field only output if calculated. ANALOG Field/Section Name Data Type Description Number Integer The number of the MassLynx analog channel. Description Text The description of the analog data source. MaxIntensity Float The absolute intensity of the point with maximum intensity in the analog channel data. TRACE Section A section that holds the chromatographic data. See description above. PEAK Section Section(s) that hold information about peak(s) detected in the chromatogram. See description above. Page 33
36 MetaboLynx Browser Sections The file will hold section blocks of data. The start of a section block will always start with the name of the section surrounded by square parentheses on a line by itself. i.e. [SectionName]. The main body of the data will be enclosed by a pair of braces, the opening and closing braces will be the only characters on their respective lines. i.e. [SectionName] {...Data... } Section blocks can have section blocks embedded in them, in some cases multiple instances of sections with the same name can be allowed, depending on the type of data. e.g. [SectionName1] {...Data... [SectionName2] {...Data... } } Fields Sections will have a group of fields, each field will be on one line. The field name will be the first item on the line, followed by a list of TAB separated data items for that field. The type of data will depend on the field type. i.e. [SectionName] { FieldName1 FieldName2... } Data1 Data2_1Data2_2 Comments A comment line is denoted by the first character being a semi-colon. A comment line can be any where in a section. Multiple line comments have a semi-colon preceding each line. Page 34
37 Tabulated Sections Some sections can be considered to be a table, or list, of fields of the same type. Tabulated sections do not have any field names to describe the data but will normally have a comment line describing the names of the fields. e.g. [TableSection] { ;Field1 Field2 Data1_1Data1_2 Data2_1Data2_2... } Free Format Text Sections The information in these sections contains free format text which may stretch over several lines. A line will be terminated with a <carriage return><linefeed> pair. Page 35
38 File Section Descriptions SAMPLE Field/Section Name Data Type Description Sample Integer The number of the sample in the batch. ProcessParameters Text MassLynx Sample List input field. Name of the MEP file used to process the data. InjectionVolume Float MassLynx Sample List input field. Volume of sample injected, in micro-litres. InletFileName Text MassLynx Sample List input field. Name of the Inlet Parameters file used to acquire the data. MSMethodFileName Text MassLynx Sample List input field. Name of the Method File used to acquire the data. MSTuneFileName Text MassLynx Sample List input field. Name of the Instrument Tune File used for the data acquisition. Well Text The description of the well position, as a pair of co-ordinates for plate position in rack and well position in plate, or an absolute position. There are three formats for Well. 1. platex,platey:wellx,welly 2. platex,platey:wellawell1 (or well1wella ) 3. Absolute rack position. Where X and Y are integers and A is an alphabetical character. FileName Text The MassLynx NT MS data file name. SampleType Text QC = Control Sample, Analyte = metabolised sample. SampleID Text MassLynx Sample List input field. SampleDescription Text MassLynx Sample List input field. Date Text Date that MS data file was acquired. Time Text Time that MS data file was acquired. JobCode Text MassLynx Sample List input field. TaskCode Text MassLynx Sample List input field. UserName Text Current MassLynx user name. LabName Text Laboratory name. Page 36
39 Field/Section Name Data Type Description Instrument Text Instrument name. Conditions Text MassLynx Sample List input field. Submitter Text MassLynx Sample List input field. Plate Text Description of micro-titre plate, if applicable. See below for detailed description. COMPOUND Section Description of parent drug(s); the masses taken from the MassLynx Sample List. FUNCTION Section MS function data and test results. There will be one FUNCTION section per MassLynx MS and DAD data functions. ANALOG Section Analogue chromatographic data. EXPECTED METABOLITES UNEXPECTED METABOLITES Section Section Description of metabolites searched for. Description of potential metabolites found as a result of comparing the sample with a control. PLATE FIELD For samples acquired using a Gilson or Waters 2700 autosampler the following information will be written to the file, otherwise this field will be blank. Version Number ( two digits ) Origin Location ( TL - top left, TR - top right, BL - bottom left, BR - bottom right ) Priority ( XY - x coordinate before y coordinate, YX - y coordinate before x coordinate ) Reference scheme ( LN - letter number, NL - number letter, NN - number number, LL - letter letter, SC - sequential continuous, SD sequential discontinuous ) 1:Number Of Rows 2:Number Of Columns 3:Row Spacing 4:Column Spacing 5:Row Offset 6:Column Offset 7:Which Rows Offset ( O or E ) 8:Which Columns Offset ( O or E ) 9:Plate Width 10:Plate Height Page 37
40 11:Vial/Well Depth 12:Vial/Well Diameter 13:Vial One X 14:Vial One Y Note that all spacings, etc are given to 0.1 of a mm. e.g. 01,TL,XY,LN,1:8,2:12,3:120.0,4:120.0,5:0.0,6:0.0,7:O,8:O,9:1250.0,10:870.0,11:4 00.0,12:90.0,13:80.0,14:130.0 Which means: Version = 01. Origin Location = top left. Priority = x coordinate before y coordinate. Reference scheme = Letter Number. 1:Number Of Rows = 8. 2:Number Of Columns = 12. 3:Row Spacing = 12mm. 4:Column Spacing = 12mm. 5:Row Offset = 0.0mm. 6:Column Offset = 0.0mm. 7:Which Rows Offset ( O or E ) = Odd numbers, but because Row Offset = 0, this has no effect. 8:Which Columns Offset ( O or E ) = Odd numbers, but because Column Offset = 0, this has no effect. 9:Plate Width = 125mm. 10:Plate Height = 87mm. 11:Vial/Well Depth = 40mm. 12:Vial/Well Diameter = 9mm. 13:Vial One X = 8.0mm. 14:Vial One Y = 13.0mm. Page 38
41 COMPOUND Field Name Data Type Description Mono Mass Float The parent drug used for targeting. This could be the monoisotopic mass or the mass with the greatest intensity, derived from the formula s isotope cluster. Formula Text The chemical composition, if entered. FUNCTION Field/Section Name Data Type Description Function Integer The MassLynx MS data function number. IonMode Text The type of ionisation mode e.g. AP+. Type Text The type of MS scan function e.g. MS for full scan. Description Text A brief description of the MS scan function. ScanCycleTime Float The scan cycle time in seconds i.e. the scan duration + the inter-scan delay time. InterScanDelay Float The inter-scan delay in seconds. SPECTRUM Section MS data and results of testing. There can be more than one SPECTRUM section in a function. CHROMATOGRAM Section Chromatographic data. There can be more than one CHROMATOGRAM section in a function. Page 39
42 SPECTRUM Field/Section Name Data Type Description ProcDesc Text Description of processing performed on spectrum. E.g. Combine (54:56-(38:39+74:75)) Process Integer The number of the MassLynx saved mass spectrum. Only present if the processed spectrum was saved with data file. State Text The result of testing the quality of the mass spectral data. Peak ID Integer Spectrum Peak Identification number. Peak Ref Integer Peak Reference Number. Peak Cluster ID Integer If this peak is considered to be the same as another peak (according to Setup parameter Min Peak Separation) then this is the number of the cluster containing this and all other similar peaks. A value of 1 indicates that this peak is considered separate from all others. Control Peak ID Integer If the sample is an analyte and this spectrum matches one in the control, then the Peak ID of the control spectrum is written to this field. Only present if this spectrum has a matching control spectrum. Time Float The retention time of the mass spectrum. TIC Float Total Ion Current, the sum of all the peak intensities in the spectrum. BPI Float The intensity of the base peak in the mass spectrum. BPM Float The mass of the base peak in the mass spectrum. Continuum Text TRUE if the spectrum is stored in continuum format. Otherwise FALSE. Saved Text TRUE if the processed spectrum was saved with the data file. Otherwise FALSE. MASSES Section Section describing the results of testing, and finding significant masses. This includes all adducts. RESULTS Section Section describing the results of the targeting of the expected compound. Masses from the same compound are combined into one item. MS Section Section that holds all the mass/intensity pairs, for this mass spectrum. Page 40
43 MASSES Field Name Data Type Description Mass Float The input mass. Exp. Mass Float The expected mass due to adduction. Obs. Mass Float The mass that was actually observed. % BPI Float The intensity of the mass peak as a percentage of the base peak intensity. Int. Abs Float The absolute intensity of the MS peak. RESULTS Field Name Data Type Description Mass Float The input mass. Found Boolean 1 if any of the possible adduct ions have been found for this compound, otherwise 0. % BPI Float The sum of the percentage intensities for the compound. % Purity Float A measure of the contribution of this compound to the spectrum as percentage of the TIC. This includes all MS peaks associated with this compound. Confirmed Boolean False if any of the confirmation tests made failed. If not specified regarded as being True. Status Text Description of confirmation failure. Not present if confirmed = TRUE MS Field Name Data Type Description Mass Float Observed mass of peak. % BPI Float Intensity of peak as a percentage of the base peak. Page 41
44 ELEMENTAL Contains results of elemental calculations if defined, otherwise this section does not appear. Field Name Data Type Description Mass Float Actual observed mass for which elemental composition was searched. Calc. Mass Float Calculated mass of reported formula Mda Float Difference between observed mass and calculated mass in milli-daltons PPM Float Difference between observed mass and calculated mass in parts per million DBE Float Double Bond Equivalence. Formula Text Representation of calculated chemical formula. E.g. C19 H14 O2 CHROMATOGRAM Field/Section Name Data Type Description TraceNumber Integer Number of chromatogram trace. Description Text The description of the chromatogram type. ProcDesc Text Description of processing performed on chromatogram. E.g. Smooth (Mn, 2x2) MaxIntensity Float The absolute intensity of the point with maximum intensity in the chromatographic data. TraceNumberIn Function Integer The number of the chromatogram trace relative to the start of the function. TRACE Section A section that holds the chromatographic data. PEAK Section Section(s) that hold information about peak(s) detected in the chromatogram. Page 42
45 TRACE Field Name Data Type Description Time Float The time of the chromatographic point. Int. %Max Float Intensity of the chromatographic point as a percentage of the maximum intensity. PEAK Field Name Data Type Description Peak ID Integer Peak Identification number. Peak Ref Integer Peak Reference Number. Peak Cluster ID Integer If this peak is considered to be the same as another peak (according to Setup parameter Min Peak Separation) then this is the number of the cluster containing this and all other similar peaks. A value of 1 indicates that this peak is considered separate from all others. Time Float Peak retention time in decimal minutes. Peak Float + Float Peak start and end retention times in decimal minutes. Two floats separated by a TAB character. Intensity Float + Float Peak baseline start and end intensity. Two floats separated by a TAB character. Height Float Detected peak height. AreaAbs Float Detected peak area. Area %BP Float The peak area as a percentage of the largest peak in the sample in which a metabolite was detected. If no metabolite found = 0. Area %Total Float The peak area as a percentage of the total area of all peaks in the sample in which a metabolite was detected. If no metabolite found = 0. Width Float Peak width in decimal minutes. Page 43
46 ANALOG Field/Section Name Data Type Description Number Integer The number of the MassLynx analog channel. Description Text The description of the analog data source. MaxIntensity Float The absolute intensity of the point with maximum intensity in the analog channel data. TRACE Section A section that holds the chromatographic data. See above. PEAK Section Section(s) that hold information about peak(s) detected in the chromatogram. See above. EXPECTED METABOLITES This is a tabulated section. Field Name Data Type Description Compound Float The number of the compound that this metabolite is associated with, starting from zero. Precursor Float Mass to be searched for, the sum of the compound and metabolite. Metabolite Float Mass of the metabolite, as specified on the MetaboLynx Setup Metabolites page. It is the mass difference between the Compound Mass and the Precursor. Found Boolean 1 indicates mass was found, 0 not found. Formula Text The formula of the metabolite if known. Absolute formula if the parent mass was specified as a formula (e.g. C18H24O2) or relative formula if parent mass was specified as a mass (e.g. +CO2). If the metabolite is not recognised the formula is 0. ChroTrace Integer list This field is a comma-separated list of chromatogram traces searched when looking for this mass; the trace numbers start from 1. Peak Text list If the mass was found, this field is a comma-separated list of chromatogram peak numbers in which the mass was found. Each peak is identified by a string of text containing four forward slash separated numbers in the following format: peak-number/function-number/trace-number/mass-found where the first 3 numbers are integers, the fourth a float. If the mass was not found, this field is 0. MetName Text Descriptive name of the metabolite Page 44
47 UNEXPECTED METABOLITES This is a tabulated section. Field Name Data Type Description Compound Float The number of the compound that this metabolite is associated with, starting from zero. Precursor Float Mass to be searched for, the sum of the compound and metabolite. Metabolite Float Mass of the metabolite, as specified on the MetaboLynx Setup Metabolites page. It is the mass difference between the Compound Mass and the Precursor. Found Boolean 1 indicates mass was found, 0 not found. Formula Text The formula of the metabolite if known. Absolute formula if the parent mass was specified as a formula (e.g. C18H24O2) or relative formula if parent mass was specified as a mass (e.g. +CO2). If the metabolite is not recognised the formula is 0. ChroTrace Integer list This field is a comma-separated list of chromatogram traces searched when looking for this mass; the trace numbers start from 1. Peak Text list If the mass was found, this field is a comma-separated list of chromatogram peak numbers in which the mass was found. Each peak is identified by a string of text containing four forward slash separated numbers in the following format: peak-number/function-number/trace-number/mass-found where the first 3 numbers are integers, the fourth a float. If the mass was not found, this field is 0. MetName Text Descriptive name of the metabolite Page 45
48 ProteinLynx Browser Sections The file will hold section blocks of data. The start of a section block will always start with the name of the section surrounded by square parentheses on a line by itself. i.e. [SectionName]. The main body of the data will be enclosed by a pair of braces, the opening and closing braces will be the only characters on their respective lines. e.g. [SectionName] {...Data... } Section blocks can have section blocks embedded in them, in some cases multiple instances of sections with the same name can be allowed, depending on the type of data. e.g. [SectionName1] {...Data... [SectionName2] {...Data... } } Fields Sections will have a group of fields, each field will be on one line. The field name will be the first item on the line, followed by a list of TAB separated data items for that field. The type of data will depend on the field type. i.e. [SectionName] { FieldName1 FieldName2... } Data1 Data2_1Data2_2 Tabulated Sections Some sections can be considered to be a table, or list, of fields of the same type. Tabulated sections do not require any field names to describe the data. e.g. [TableSection] { ;Field1 Field2 Data1_1Data1_2 Data2_1Data2_2... } Page 46
49 File Section Descriptions SAMPLE Note: The fields may not appear in the same order as the table below and some empty fields will not appear at all. Field/Section Name Data Type Description Sample Integer The number of the sample in the batch. Well Text The description of the well position, as a pair of co-ordinates for plate position in rack and well position in plate, or an absolute position. There are three formats for Well. 1. wellx,welly 2. wellawell1 (or well1wella ) 3. Absolute rack position. Where X and Y are integers and A is an alphabetical character. Data_File_Name Text The MassLynx NT MS data file name. Sample_ID Text MassLynx Sample ID. Sample_Description Text MassLynx Sample description. Spare1 Text User definable spare. Spare2 Text User definable spare. Spare3 Text User definable spare. Spare4 Text User definable spare. Spare5 Text User definable spare. Date Text Date that MS data file was acquired. Time Text Time that MS data file was acquired. Job_Code Text MassLynx Sample List Jobcode field. Task_Code Text MassLynx Sample List Taskcode field. User_Name Text Current MassLynx user name. Instrument Text Instrument name. Plate Text Description of micro-titre plate, if applicable. MS_File Text The MassLynx MS data file name. Process_Parameters Text The MassLynx process parameter file name. Page 47
50 Field/Section Name Data Type Description FUNCTION Section MS function data and results. There will be one FUNCTION section per MassLynx MS data functions. FUNCTION Field/Section Name Data Type Description Function Integer The MassLynx MS data function number. IonMode Text The type of ionisation mode e.g. AP+. Type Text The type of MS scan function e.g. MS for full scan. Description Text A brief description of the MS scan function. PROCESS_PARAMETERS Section MS process parameter data and results. There will only be one PROCESS_PARAMETERS section in a FUNCTION. SEARCH_PARAMETERS Section MS search parameter data. There will only be one SEARCH_PARAMETERS section in a FUNCTION. SEARCH_RESULTS Section MS search results. There will only be one SEARCH_RESULTS section in a FUNCTION. SPECTRUM Section MS raw spectrum data. There will only be one SPECTRUM in a function. Note: Only present for Maldi data. PEAKLIST Section MSMS raw spectrum data. There will only be one PEAKLIST in a function. Note: Only present for MSMS data. Page 48
51 PROCESS_PARAMETERS Field Name Data Type Description Do_Process Boolean 1 if processing has been performed, otherwise 0. Combine_Mode Text Spectrum combine mode. DoBackgroundSubtract Boolean 1 if background subtraction has been performed, otherwise 0. PolynomialOrder Int Number indicating the polynomial order used in background subtraction. BelowCurve Int Number indicating the below curve % used in background subtraction. DoSmooth Boolean 1 if smoothing has been performed, otherwise 0. SmoothChannels Float Number of channels smoothed. NumberOfSmooths Int Number of smooths performed. Smooth_Mode Text The smoothing method used. E.g. Savitzky-Golay Min_Peak_Width Int Minimum spectral peak width. Centroid_Percent Float Intensity of peak as a percentage of the base peak. Centroid_Mode Text Spectrum centroid mode used. PreCharge Int Pre charge state. Min_Mass_To_Start Float Minimum start search mass. Isotope_Factor Float Minimum isotopic factor value. Relative_Deiso_Intensity Int The relative deiso intensity. LockMassPeak_1 Int First lock mass value. LockMassPeak_2 Int Second lock mass value.. LockMassPeak_3 Int Third lock mass value. Page 49
52 SEARCH_PARAMETERS Field Name Data Type Description Do_Search Boolean 1 if search performed, otherwise 0. Database_Type Text Type of database searched. Index_Path Text Path of database index. Sequence_File Text Database sequence file name. Trembl_Path Text Path of Trembl database SearchServer Text Name of search server Output_Type Text Type of output file. PEAKLIST_location Text Location of Sequest output file. MASS Text Location of mascot output file. PepSea_location Text Location of PepSea output file. MOWSE_location Text Location of mass intensity output file. Organism Text Search organism. Freetext Text Search free text. Keyword Text Search key word. Author Text Search author. Accession Text The database accession code. Subsequence Text Search sub sequence. BileEqual Boolean 1 if treat I/L and Q/K as equivalent box checked, otherwise 0 Do_Monoiso Boolean 1 if search monoisotopic peaks box checked, otherwise 0. MAX_HITS Integer Maximum number of hits to return from the search. 0 if All selected. SORT_TYPE Text BIO_SORT_PROB, BIO_SORT_LIKELIHOOD, BIO_SORT_ENTRY, BIO_SORT_COVERAGE or BIO_SORT_MATCH depending on the sort method. Exclude_Lockmass Boolean 1 if lockmass excluded, otherwise 0. Tolerance Float Peptide tolerance value. Tolerance_Units Text Peptide tolerance units. Page 50
53 Field Name Data Type Description Charge Float Peptide charge state. MinToMatch Int Number of required matches. Do_Digest Boolean 1 if digest simulation was performed, otherwise 0. Fast_Digest_Reagent Text Name of digest reagent when using and index. Digest_Index Text Digest index s file path and name. Simulated_Digest_Reagent Text Name of primary digest reagent. Secondary_Digest_Reagent Text Name of secondary digest reagent. pfactor Float The pfactor value entered. Missed_cleavages Integer Number of missed cleavage sites. Do_Incomplete Boolean 1 if incomplete cleavage processing performed, otherwise 0. Do_MW Boolean 1 if molecular weight restriction was specified, otherwise 0. MW_from Float Molecular weight range minimum. MW_to Float Molecular weight range maximum. Do_PI Boolean 1 if isoelectric point restriction was specified, otherwise 0. PI_from Float Isoelectric point range minimum. PI_to Float Isoelectric point range maximum. Do_Modifiers Boolean 1 if modification was specified, otherwise 0. Reduced_cysteine Boolean 1 if reduced cysteine was specified, otherwise 0. MODIFIERS Section Description of the modifiers used during processing. MODIFIERS Field Name Data Type Description Name of modifier Boolean 1 if always use this modifier box checked, 0 if use when needed to match peptide box checked. Page 51
54 SEARCH_RESULTS Field Name Data Type Description File_Name Text Search result file name with its full path PRP_File_Name Text The hit section does not appear in this file but in the file named above. One HIT section defines one database Hit. There can be more than one HIT sections. SPECTRUM Field/Section Name Data Type Description LockMassUsed Float The lock mass value used MONOISOMASSES Section MS monoisomasses data. There will only be one MONOISOMASSES in a SPECTRUM. CORRECTION_FACTOR Section MS raw spectrum data. There will only be one CORRECTION_FACTOR in a SPECTRUM. MONOISOMASSES Field Name Data Type Description Float One monoisomass value per line. There can be more than one line CORRECTION_FACTOR Field Name Data Type Description Correction_Factor Float Correction factor applied to the whole spectrum. Calculated from the lock mass. Offset_Factor Float Correction factor applied to the whole spectrum. Calculated from the lock mass. ProcessHistory Text Process history information text string Page 52
55 PEAK_LIST Field Name Data Type Description Output_Filename Text The name of the Mascot or Sequest compatible peak information file. e.g. Mascot file C:\TEMP\hilo pkl where 004 = the scan number 2 = the function number 1 = the charge state Sequest file C:\MASSLYNX\hilo dta where 0001 = the scan number 01 = the function number. Bio-Rad File Descriptions If the Bio-Rad box was checked then a SampleID.txt file is produced with the following format. Field Name Data Type Description VIEW_ONE Section Database protein entry information. There will only be one VIEW_ONE section per HIT. VIEW_TWO Section Database match information There will only be one VIEW_TWO section per HIT. VIEW_THREE Section Database protein sequence There will only be one VIEW_THREE section per HIT. Page 53
56 VIEW_ONE Some entries do not contain data on all the line types, and some line types occur many times in a single entry. Field Name ID AC DT DE GN OS OG OC RN Description Identification line showing the entry name of the sequence. The accession number, a unique number used to identify entries. Date of entry or last modification. Description line containing general descriptive information. Gene name. The organism species line specifies the organism which was the source of the sequence. In most cases this is the Latin name followed by the English name (in parentheses). For viruses only the common English name is given. If the sequence is identical for more than one species the OS lines will list the names of all those species. The Organelle lines indicate if the gene coding for a protein originates from the mitochondria, the chloroplast, a cyanelle or a plasmid. The organism classification lines contain the taxonomic classification of the source organism. Reference number RP Reference position RC Reference comment RX Reference cross-reference RA Reference authors RL Reference location (Journal information) CC DR KW FT Comments or notes used to convey useful information. The database cross-reference lines are used as pointers to related information in other data collections. E.g. Protein Data Bank (PDB). Keyword lines provide information which can be used to generate cross-references to other entries. Feature table data lists regions of interest in the sequence - post-transitional modifications, binding sites, enzyme active sites etc. SQ Sequence header followed by sequence data. Page 54
57 VIEW_TWO Some entries do not contain data on all the line types, and some line types occur many times in a single entry. Field Name ID Accession number Description Species Number of matched peptides Probability score Percent coverage Description Identification line showing the entry name of the sequence. The accession number, a unique number used to identify entries. Description line containing general descriptive information. The organism species line specifies the organism which was the source of the sequence. The number of matched peptides MOWSE type score. % sequence coverage. Isoelectric point The isoelectric point of the entry. Molecular weight The molecular weight of the entry. Matching peptides Unmatched masses A list of matching fragment masses with their molecular weight (MW), mass error (Delta), the position of the peptide in the sequence (start and end) and the sequence. If a * appears to the left of one of these entries then it is a single partial fragment (i.e. one missed cleavage site), if ** appears it is a double partial fragment (i.e. two missed cleavage sites). A list of unmatched masses showing the Searched mass, the Query mass and the Charge State. VIEW_THREE This view displays the sequence specific to the entry as a text string. Page 55
58 File Formats Sample List The Sample Lists are held in Microsoft Access which mean their contents are readily available to numerous other windows programs. Sample Lists created by the MassLynx system have the.spl extension. Label Format Description VERSION Double Database version number. Index Double Used internally to keep samples sequenced when editing. FILE_NAME MS_FILE MS_TUNE_FILE INLET_FILE INLET_PRERUN INLET_POSTRUN INLET_SWITCH AUTO_FILE Char (255) Raw data file name for this sample. Can be name or full path. Char (255) MS method parameter file to use when acquiring this sample. Can be name or full path. Char (255) MS tuning parameter file to use when acquiring this sample. Can be name or full path. If empty the current tune settings will be used. Char (255) Inlet method parameter file to use when acquiring this sample. Can be name or full path. Only required if a programmable inlet acquisition system is being used to acquire sample. Char (255) Inlet pre-run method parameter file to use prior to acquiring this sample. Can be name or full path. If empty no pre-run is performed. Only required if a programmable inlet acquisition system is being used to acquire sample. Char (255) Inlet postrun method parameter file to use subsequent to acquiring this sample. Can be name or full path. If empty no post-run is performed. Only required if a programmable inlet acquisition system is being used to acquire sample. Char (255) Inlet pre-run method parameter file to use prior to acquiring this sample if inlet system has had to make a column switch, this will be used in preference to INLET_PRERUN. Can be name or full path. If empty no column switch pre-run is performed. Only required if a programmable inlet acquisition system is being used to acquire sample. Char (255) Autosampler method parameter file to use when acquiring this sample. Can be name or full path. Only required if a programmable autosampler system is being used to acquire sample. SAMPLE_LOCATION Char (255) Defines location of sample to be acquired on autosampler bed. Format will depend on type of autosampler being used. INJ_VOL Double Volume of sample to inject when acquiring with programmable autosampler Page 56
59 Label Format Description PROCESS PROCESS_PARAMS PROCESS_OPTIONS FILE_TEXT Char (255) External process to run to perform processing of sample. Can be name or full path. Only required if external processes are being run, if empty no process will be run for this sample. Char (255) Name of external process parameter file to use when processing samples. Available to external process via MLCURSMP.TXT file. Char (255) Options to specify on command line of external process when it is executed Char (255) Sample text description to be used for sample. Will be recorded in data file header. JOB Char(255) Job description for sample. Will be recorded in data file header. TASK Char(255) Task description for sample. Will be recorded in data file header. USER Char(255) User name for sample. Will be recorded in data file header. SUBMITTER Char(255) Submitter for sample. Will be recorded in data file header. CONDITIONS Char(255) Condition information for sample. Will be recorded in data file header. TYPE Char (255) Sample type to use during Quantify. Currently can be one of ANALYTE, BLANK, QC or STANDARD ID Char(255) The sample ID from OpenLynx. CONC_A CONC_T Char (255) 20 fields used to specify the concentration levels of compounds within this sample. Used during the quantify process. WAVELENGTH_A WAVELENGTH_J Double 10 fields used to specify wavelengths to analyse for this sample. Used during OpenLynx processing. MASS_A MASS_T Char (255) 20 fields used to specify masses to analyse for this sample. Used during the OpenLynx process. Masses can be real numbers or chemical formulae STOCK_DIL Double Not currently used reserved for future use USER_DIVISOR_1 Double Divisor used during concentration calculation stage of Quantify. Defaults to 1 if not specified. USER_FACTOR_1 USER_FACTOR_3 Double Multipliers used during concentration calculation stage of Quantify. Defaults to 1 if not specified. HPLC_FILE Char(255) The additional LC information for this sample (used by OpenLynx) SPARE_1 SPARE_5 ACQU_PROCESS Char (255) General purpose fields available to the User to store extra information about the sample. Char (255) External process to run when acquiring data. Can be name or full path. Only required if external processes are being run, if empty no process will be run for this sample. Page 57
60 Label Format Description ACQU_PROCESS_ PARAMS ACQU_PROCESS_ OPTIONS SAMPLE_GROUP Char (255) Name of external process parameter file to use when acquiring samples. Available to external process via MLCURSMP.TXT file. Char (255) Options to specify on command line of external process when it is executed Char (255) Used too inform QuanLynx that the optimisation file for an Analysis sample should combine the optimisations for a number of compounds sample groups are used. Each sample group item is a unique one or two letter code in upper case. Thus A:B:C, AA BB CC, A.ZZ.QP are all valid three item groups. If the sample group field is empty then all the compounds will be optimised; one method file will be created and used for all the analysis samples. If a sample group in the analysis sample list does not have a corresponding entry in the compound list, then that group will not be run. FRACTION_FILE FRACTION_1 to FRACTION_4 QUAN_REF AUTO_ADDITION METH_DB CURVE_DB DUMP_FILE PROCESS_ACTION Char (255) Raw data file name for this sample. Can be name or full path. Char (255) References to the masses (MASS_A to MASS_T) to be monitored when performing fraction collection acquisition. Char (255) This field specifies which sample (if any) is to be used to adjust the retention time window of the quantification method created for the analysis. The first sample in the current group that has this field set to anything other than blank will be used. Char (255) The Waters 2790 autosampler can take samples from up to 15 vials and inject them sequentially. This field contains a list of the sample locations separated by a semi-colon (;) e.g. A:1;A:2;A:3. The volume specified in the INJ_VOL field is drawn from each vial. Char (255) The full path to the quantify method file used for this sample group in QuanLynx. Char (255) The full path to the calibration curve file used for this sample group in QuanLynx. Char (255) The full path to the text export file used for this sample group in QuanLynx. Char (255) The Action that the process manager is to perform should the external application return an error Page 58
61 An Example Generic Sample List File Below is an example of a small single sample generic sample list file. [Batch] MSMethod=DEFAULT.MDB MSTune=DEFAULT.DBF LCMethod=DEFAULT ProcessParameters=C:\MASSLYNX\LC1.olp Process=C:\MASSLYNX\OPENLYNX.EXE UserName=john doe BatchID=13 NumberOfPlates=1 [Plate:1] JobCode=13 UserName=john doe Rows=0 Columns=0 Track=R NumberOfWells=1 [Sample:1:1] Well=25 SampleDescription=testsample SampleID=tst11 MSDataName=test1 SampleType=0 MSInjectionVolume= NumberOfMasses=1 Mass1= NumberOfWaveLengths=1 Wavelength1=254.0 NumberOfConcs=2 Conc1= Conc2= NumberOfFormulae=1 Formula1=CH3CH2Cl Page 59
62 AutoLynx AutoLynx is an application that enables batches to be submitted to the MassLynx queue for automatic acquisition, processing and report generation. Applications can be written (e.g. in Visual Basic) to: Take information from other software packages (e.g. LIMS systems) and automatically create batch files. Add these batch files to the AutoLynx queue. (AutoLynx will then add them to the MassLynx queue and perform the processing defined on the AutoLynx settings dialog). Monitor the progress of the batch file. Automatically read the acquired data, or processed results, for further processing and/or exporting into other software packages. Page 60
63 To produce the required Browser file the Process column in the Sample List must be defined as: OpenLynx to produce an OpenLynx Browser file. PeptideAuto to produce an ProteinLynx Browser file. MetaboLynx to produce an MetaboLynx Browser file. For more information on AutoLynx see the MassLynx NT Users Guide. Page 61
64 Status.ini The Status.ini file contains details of the current status of the Mass Spectrometer, LC system and the MassLynx Queue. To create a Status.ini file select Options from the MassLynx Tools menu to display the options dialog. Figure 3 The Options dialog On the Status page check the Update Status box to write details to a status file. By default the details in this file are updated every 60 seconds, to change this enter a new time in the Refresh rate box. The default filename and location is C:\MASSLYNX\status.ini. To change this press the File Name button, select a directory using the browser displayed, enter a new file name and press Open. Note: do not use the name of an existing *.ini file in the current directory as this will cause problems with other software. If any of these fields are changed MassLynx must be closed down and restarted for the changes to take effect. Page 62
OpenLynx User's Guide
 OpenLynx User s Guide OpenLynx User's Guide Version 4.0 Waters part No 715000405 Micromass Part No - 6666670 5 February, 2002 i OpenLynx User s Guide OpenLynx User s Guide The software described in this
OpenLynx User s Guide OpenLynx User's Guide Version 4.0 Waters part No 715000405 Micromass Part No - 6666670 5 February, 2002 i OpenLynx User s Guide OpenLynx User s Guide The software described in this
TraceFinder Analysis Quick Reference Guide
 TraceFinder Analysis Quick Reference Guide This quick reference guide describes the Analysis mode tasks assigned to the Technician role in the Thermo TraceFinder 3.0 analytical software. For detailed descriptions
TraceFinder Analysis Quick Reference Guide This quick reference guide describes the Analysis mode tasks assigned to the Technician role in the Thermo TraceFinder 3.0 analytical software. For detailed descriptions
TraceFinder Analysis Quick Reference Guide
 TraceFinder Analysis Quick Reference Guide This quick reference guide describes the Analysis mode tasks assigned to the Technician role in Thermo TraceFinder analytical software. For detailed descriptions
TraceFinder Analysis Quick Reference Guide This quick reference guide describes the Analysis mode tasks assigned to the Technician role in Thermo TraceFinder analytical software. For detailed descriptions
ChromQuest 5.0 Quick Reference Guide
 ChromQuest 5.0 Quick Reference Guide This guide contains an overview of the ChromQuest chromatography data system, with topics organized by workflow. For more information, refer to the ChromQuest User
ChromQuest 5.0 Quick Reference Guide This guide contains an overview of the ChromQuest chromatography data system, with topics organized by workflow. For more information, refer to the ChromQuest User
Thermo Xcalibur Getting Started (Quantitative Analysis)
 Thermo Xcalibur Getting Started (Quantitative Analysis) XCALI-97207 Revision B September 2010 2010 Thermo Fisher Scientific Inc. All rights reserved. Xcalibur, Surveyor, and Accela are registered trademarks
Thermo Xcalibur Getting Started (Quantitative Analysis) XCALI-97207 Revision B September 2010 2010 Thermo Fisher Scientific Inc. All rights reserved. Xcalibur, Surveyor, and Accela are registered trademarks
FractionLynx Software
 FractionLynx Software User s Guide 34 Maple Street Milford, MA 01757 71500054302, Revision A NOTICE The information in this document is subject to change without notice and should not be construed as a
FractionLynx Software User s Guide 34 Maple Street Milford, MA 01757 71500054302, Revision A NOTICE The information in this document is subject to change without notice and should not be construed as a
TraceFinder Administrator Quick Reference Guide
 TraceFinder Administrator Quick Reference Guide This quick reference guide discusses the application configuration tasks assigned to the ITAdmin and LabDirector roles when user security is activated in
TraceFinder Administrator Quick Reference Guide This quick reference guide discusses the application configuration tasks assigned to the ITAdmin and LabDirector roles when user security is activated in
GPS Explorer Software For Protein Identification Using the Applied Biosystems 4700 Proteomics Analyzer
 GPS Explorer Software For Protein Identification Using the Applied Biosystems 4700 Proteomics Analyzer Getting Started Guide GPS Explorer Software For Protein Identification Using the Applied Biosystems
GPS Explorer Software For Protein Identification Using the Applied Biosystems 4700 Proteomics Analyzer Getting Started Guide GPS Explorer Software For Protein Identification Using the Applied Biosystems
Data Acquisition with CP-2002/2003 Micro-GC Control
 Varian Analytical Instruments 2700 Mitchell Drive Walnut Creek, CA 94598 Star Chromatography Workstation Version 6 Data Acquisition with CP-2002/2003 Micro-GC Control Operation Manual Varian, Inc. 2002
Varian Analytical Instruments 2700 Mitchell Drive Walnut Creek, CA 94598 Star Chromatography Workstation Version 6 Data Acquisition with CP-2002/2003 Micro-GC Control Operation Manual Varian, Inc. 2002
Agilent G2727AA LC/MS Data Browser Quick Start
 What is Data Browser? Agilent G2727AA LC/MS Data Browser Quick Start A guide to get started with Data Browser Use this quick reference guide as a road map for your first steps with the Data Browser software,
What is Data Browser? Agilent G2727AA LC/MS Data Browser Quick Start A guide to get started with Data Browser Use this quick reference guide as a road map for your first steps with the Data Browser software,
Protein Deconvolution Quick Start Guide
 Protein Deconvolution Quick Start Guide The electrospray ionization (ESI) of intact peptides and proteins produces mass spectra containing a series of multiply charged ions with associated mass-to-charge
Protein Deconvolution Quick Start Guide The electrospray ionization (ESI) of intact peptides and proteins produces mass spectra containing a series of multiply charged ions with associated mass-to-charge
TraceFinder Shortcut Menus Quick Reference Guide
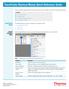 TraceFinder Shortcut Menus Quick Reference Guide This quick reference guide describes the right-click shortcut menus available in the Thermo TraceFinder application. Contents Acquisition Mode Analysis
TraceFinder Shortcut Menus Quick Reference Guide This quick reference guide describes the right-click shortcut menus available in the Thermo TraceFinder application. Contents Acquisition Mode Analysis
MRMPROBS tutorial. Hiroshi Tsugawa RIKEN Center for Sustainable Resource Science
 MRMPROBS tutorial Edited in 2014/10/6 Introduction MRMPROBS is a tool for the analysis of data from multiple reaction monitoring (MRM)- or selected reaction monitoring (SRM)-based metabolomics studies.
MRMPROBS tutorial Edited in 2014/10/6 Introduction MRMPROBS is a tool for the analysis of data from multiple reaction monitoring (MRM)- or selected reaction monitoring (SRM)-based metabolomics studies.
Getting Started. Environmental Analysis Software
 Getting Started Environmental Analysis Software Environmental Software Overview The Environmental software is a data acquisition and data analysis package designed to assist you in complying with United
Getting Started Environmental Analysis Software Environmental Software Overview The Environmental software is a data acquisition and data analysis package designed to assist you in complying with United
Xcalibur. QuickQuan. User Guide. XCALI Revision C July For Research Use Only Not for use in Diagnostic Procedures
 Xcalibur QuickQuan User Guide XCALI-97195 Revision C July 2007 For Research Use Only Not for use in Diagnostic Procedures 2007 Thermo Fisher Scientific Inc. All rights reserved. Microsoft, Access, Excel,
Xcalibur QuickQuan User Guide XCALI-97195 Revision C July 2007 For Research Use Only Not for use in Diagnostic Procedures 2007 Thermo Fisher Scientific Inc. All rights reserved. Microsoft, Access, Excel,
Agilent 6400 Series Triple Quadrupole LC/MS System
 Agilent 6400 Series Triple Quadrupole LC/MS System Quick Start Guide Where to find information 4 Getting Started 6 Step 1. Start the Data Acquisition software 7 Step 2. Prepare the LC modules 13 Step 3.
Agilent 6400 Series Triple Quadrupole LC/MS System Quick Start Guide Where to find information 4 Getting Started 6 Step 1. Start the Data Acquisition software 7 Step 2. Prepare the LC modules 13 Step 3.
Tutorial 7: Automated Peak Picking in Skyline
 Tutorial 7: Automated Peak Picking in Skyline Skyline now supports the ability to create custom advanced peak picking and scoring models for both selected reaction monitoring (SRM) and data-independent
Tutorial 7: Automated Peak Picking in Skyline Skyline now supports the ability to create custom advanced peak picking and scoring models for both selected reaction monitoring (SRM) and data-independent
For additional information, see these documents on the Breeze 2 software DVD:
 R E L E A S E N O T E S Waters Breeze 2 Read these release notes before you install and operate Breeze 2. They contain information necessary to ensure successful liquid chromatography results, including:
R E L E A S E N O T E S Waters Breeze 2 Read these release notes before you install and operate Breeze 2. They contain information necessary to ensure successful liquid chromatography results, including:
Using OPUS to Process Evolved Gas Data (8/12/15 edits highlighted)
 Using OPUS to Process Evolved Gas Data (8/12/15 edits highlighted) Once FTIR data has been acquired for the gases evolved during your DSC/TGA run, you will process using the OPUS software package. Select
Using OPUS to Process Evolved Gas Data (8/12/15 edits highlighted) Once FTIR data has been acquired for the gases evolved during your DSC/TGA run, you will process using the OPUS software package. Select
Quick Guide to the Star Bar
 Saturn View Saturn Writer Quick Guide to the Star Bar Last active Chromatogram in SaturnView Last active method in the Method Editor Click with the right mouse button on the Star Bar to get this menu Sample,
Saturn View Saturn Writer Quick Guide to the Star Bar Last active Chromatogram in SaturnView Last active method in the Method Editor Click with the right mouse button on the Star Bar to get this menu Sample,
Reference Manual MarkerView Software Reference Manual Revision: February, 2010
 Reference Manual MarkerView 1.2.1 Software Reference Manual - 1 - Revision: February, 2010 This document is provided to customers who have purchased AB SCIEX equipment to use in the operation of such AB
Reference Manual MarkerView 1.2.1 Software Reference Manual - 1 - Revision: February, 2010 This document is provided to customers who have purchased AB SCIEX equipment to use in the operation of such AB
What s New in Empower 3
 What s New in Empower 3 Revision A Copyright Waters Corporation 2010 All rights reserved Copyright notice 2010 WATERS CORPORATION. PRINTED IN THE UNITED STATES OF AMERICA AND IN IRELAND. ALL RIGHTS RESERVED.
What s New in Empower 3 Revision A Copyright Waters Corporation 2010 All rights reserved Copyright notice 2010 WATERS CORPORATION. PRINTED IN THE UNITED STATES OF AMERICA AND IN IRELAND. ALL RIGHTS RESERVED.
The Detection Flags and Primary Flags Fields in QuanLynx and TargetLynx Version 4.1
 TECN1851799 New Doc. No. Rev. 01 Page 1 of 5 The Detection Flags and Primary Flags Fields in QuanLynx and TargetLynx Version 4.1 The Detection Flags field in QuanLynx and the Primary Flags field in TargetLynx
TECN1851799 New Doc. No. Rev. 01 Page 1 of 5 The Detection Flags and Primary Flags Fields in QuanLynx and TargetLynx Version 4.1 The Detection Flags field in QuanLynx and the Primary Flags field in TargetLynx
Agilent MassHunter Qualitative Data Analysis
 Agilent MassHunter Qualitative Data Analysis Qualitative Navigator B.08.00 Presenters: Kevin Costalunga Stephen Harnos With Matt Leyden & Kevin Costalunga 1 MassHunter Webinar Series MassHunter Qualitative
Agilent MassHunter Qualitative Data Analysis Qualitative Navigator B.08.00 Presenters: Kevin Costalunga Stephen Harnos With Matt Leyden & Kevin Costalunga 1 MassHunter Webinar Series MassHunter Qualitative
MassLynx 4.1 SCN 804. Release Notes. SQ Detector 2
 R E L E A S E N O T E S MassLynx 4.1 SCN 804 Release Notes SQ Detector 2 43464 Rev 01 Copyright Waters Corporation 2011. All rights reserved. Copyright 2011 Waters Corporation 43464_1. Waters is a registered
R E L E A S E N O T E S MassLynx 4.1 SCN 804 Release Notes SQ Detector 2 43464 Rev 01 Copyright Waters Corporation 2011. All rights reserved. Copyright 2011 Waters Corporation 43464_1. Waters is a registered
Xcalibur Library Browser
 Thermo Xcalibur Library Browser User Guide Creating and Searching Spectral Libraries Software Version 3.0 XCALI-97552 Revision A June 2013 2013 Thermo Fisher Scientific Inc. All rights reserved. Xcalibur
Thermo Xcalibur Library Browser User Guide Creating and Searching Spectral Libraries Software Version 3.0 XCALI-97552 Revision A June 2013 2013 Thermo Fisher Scientific Inc. All rights reserved. Xcalibur
EXCEL 2007 TIP SHEET. Dialog Box Launcher these allow you to access additional features associated with a specific Group of buttons within a Ribbon.
 EXCEL 2007 TIP SHEET GLOSSARY AutoSum a function in Excel that adds the contents of a specified range of Cells; the AutoSum button appears on the Home ribbon as a. Dialog Box Launcher these allow you to
EXCEL 2007 TIP SHEET GLOSSARY AutoSum a function in Excel that adds the contents of a specified range of Cells; the AutoSum button appears on the Home ribbon as a. Dialog Box Launcher these allow you to
Galaxie Report Editor
 Varian, Inc. 2700 Mitchell Drive Walnut Creek, CA 94598-1675/USA Galaxie Report Editor User s Guide Varian, Inc. 2008 Printed in U.S.A. 03-914949-00: Rev 6 Galaxie Report Editor i Table of Contents Introduction...
Varian, Inc. 2700 Mitchell Drive Walnut Creek, CA 94598-1675/USA Galaxie Report Editor User s Guide Varian, Inc. 2008 Printed in U.S.A. 03-914949-00: Rev 6 Galaxie Report Editor i Table of Contents Introduction...
Chromatography Software Training Materials. Contents
 Chromatography Software Training Materials This document contains information on how to build a method, start the instrument to acquire data, and then process the data using the Galaxie Program. You will
Chromatography Software Training Materials This document contains information on how to build a method, start the instrument to acquire data, and then process the data using the Galaxie Program. You will
Agilent G6854 MassHunter Personal Pesticide Database
 Agilent G6854 MassHunter Personal Pesticide Database Quick Start Guide What is MassHunter Personal Pesticide Database? 2 Installation 3 Main Window 4 Getting Started 11 Database operations 12 Searching
Agilent G6854 MassHunter Personal Pesticide Database Quick Start Guide What is MassHunter Personal Pesticide Database? 2 Installation 3 Main Window 4 Getting Started 11 Database operations 12 Searching
NiceForm User Guide. English Edition. Rev Euro Plus d.o.o. & Niceware International LLC All rights reserved.
 www.nicelabel.com, info@nicelabel.com English Edition Rev-0910 2009 Euro Plus d.o.o. & Niceware International LLC All rights reserved. www.nicelabel.com Head Office Euro Plus d.o.o. Ulica Lojzeta Hrovata
www.nicelabel.com, info@nicelabel.com English Edition Rev-0910 2009 Euro Plus d.o.o. & Niceware International LLC All rights reserved. www.nicelabel.com Head Office Euro Plus d.o.o. Ulica Lojzeta Hrovata
Tutorial 2: Analysis of DIA/SWATH data in Skyline
 Tutorial 2: Analysis of DIA/SWATH data in Skyline In this tutorial we will learn how to use Skyline to perform targeted post-acquisition analysis for peptide and inferred protein detection and quantification.
Tutorial 2: Analysis of DIA/SWATH data in Skyline In this tutorial we will learn how to use Skyline to perform targeted post-acquisition analysis for peptide and inferred protein detection and quantification.
Agilent G6854AA MassHunter Personal Compound Database
 Agilent G6854AA MassHunter Personal Compound Database Quick Start Guide This guide describes how to install and use MassHunter Personal Compound Database. Where to find more information What is Personal
Agilent G6854AA MassHunter Personal Compound Database Quick Start Guide This guide describes how to install and use MassHunter Personal Compound Database. Where to find more information What is Personal
Agilent EZChrom Elite. PDA Analysis
 Agilent EZChrom Elite PDA Analysis Notices Copyright Scientific Software, Inc 1997-2003 Agilent Technologies, Inc. 2006. No part of this manual may be reproduced in any form or by any means (including
Agilent EZChrom Elite PDA Analysis Notices Copyright Scientific Software, Inc 1997-2003 Agilent Technologies, Inc. 2006. No part of this manual may be reproduced in any form or by any means (including
Agilent G6825AA METLIN Personal Metabolite Database for MassHunter Workstation
 Agilent G6825AA METLIN Personal Metabolite Database for MassHunter Workstation Quick Start Guide This guide describes how to install and use METLIN Personal Metabolite Database for MassHunter Workstation.
Agilent G6825AA METLIN Personal Metabolite Database for MassHunter Workstation Quick Start Guide This guide describes how to install and use METLIN Personal Metabolite Database for MassHunter Workstation.
Q Exactive HF-X Software Manual
 Q Exactive HF-X Software Manual Exactive Series 2.9 BRE0012262 Revision A May 2017 Q Exactive HF-X Software Manual Exactive Series 2.9 BRE0012262 Revision A May 2017 Legal Notices 2017 Thermo Fisher Scientific
Q Exactive HF-X Software Manual Exactive Series 2.9 BRE0012262 Revision A May 2017 Q Exactive HF-X Software Manual Exactive Series 2.9 BRE0012262 Revision A May 2017 Legal Notices 2017 Thermo Fisher Scientific
Data Handling and Reports
 Varian Analytical Instruments 2700 Mitchell Drive Walnut Creek, CA 94598-1675/USA Star Chromatography Workstation Version 6 Data Handling and Reports Tutorials Varian, Inc. 2002 03-914736-00:Rev. 4 Trademark
Varian Analytical Instruments 2700 Mitchell Drive Walnut Creek, CA 94598-1675/USA Star Chromatography Workstation Version 6 Data Handling and Reports Tutorials Varian, Inc. 2002 03-914736-00:Rev. 4 Trademark
Clarity. Clarity Demo. Code/Rev.: M003/60A Date: Fax: Petrzilkova 2583/ Prague 5
 Clarity Clarity Demo Clarity ENG Code/Rev.: M003/60A Date: 21.4.2015 Phone: +420 251 013 400 DataApex Ltd. Fax: +420 251 013 401 Petrzilkova 2583/13 clarity@dataapex.com 158 00 Prague 5 www.dataapex.com
Clarity Clarity Demo Clarity ENG Code/Rev.: M003/60A Date: 21.4.2015 Phone: +420 251 013 400 DataApex Ltd. Fax: +420 251 013 401 Petrzilkova 2583/13 clarity@dataapex.com 158 00 Prague 5 www.dataapex.com
UniPoint System Software User s Guide
 UniPoint System Software User s Guide LT2137 1998 Gilson, Inc. All rights reserved Exercise 3-Creating operations list... 4-22 Create and name operations list... 4-22 Set up steps... 4-23 Identify sample
UniPoint System Software User s Guide LT2137 1998 Gilson, Inc. All rights reserved Exercise 3-Creating operations list... 4-22 Create and name operations list... 4-22 Set up steps... 4-23 Identify sample
Skyline High Resolution Metabolomics (Draft)
 Skyline High Resolution Metabolomics (Draft) The Skyline Targeted Proteomics Environment provides informative visual displays of the raw mass spectrometer data you import into your Skyline documents. Originally
Skyline High Resolution Metabolomics (Draft) The Skyline Targeted Proteomics Environment provides informative visual displays of the raw mass spectrometer data you import into your Skyline documents. Originally
Formulas and Functions
 Conventions used in this document: Keyboard keys that must be pressed will be shown as Enter or Ctrl. Controls to be activated with the mouse will be shown as Start button > Settings > System > About.
Conventions used in this document: Keyboard keys that must be pressed will be shown as Enter or Ctrl. Controls to be activated with the mouse will be shown as Start button > Settings > System > About.
Introduction to Excel 2007
 Introduction to Excel 2007 These documents are based on and developed from information published in the LTS Online Help Collection (www.uwec.edu/help) developed by the University of Wisconsin Eau Claire
Introduction to Excel 2007 These documents are based on and developed from information published in the LTS Online Help Collection (www.uwec.edu/help) developed by the University of Wisconsin Eau Claire
Basic tasks in Excel 2013
 Basic tasks in Excel 2013 Excel is an incredibly powerful tool for getting meaning out of vast amounts of data. But it also works really well for simple calculations and tracking almost any kind of information.
Basic tasks in Excel 2013 Excel is an incredibly powerful tool for getting meaning out of vast amounts of data. But it also works really well for simple calculations and tracking almost any kind of information.
MassLynx 4.2 SCN 989 Release Notes
 MassLynx 4.2 SCN 989 Release Notes Page 1 of 20 Waters, THE SCIENCE OF WHAT S POSSIBLE., ACQUITY, Alliance, MassLynx, nanoacquity, and Xevo are registered trademarks and OpenLynx, Quanpedia, and TargetLynx
MassLynx 4.2 SCN 989 Release Notes Page 1 of 20 Waters, THE SCIENCE OF WHAT S POSSIBLE., ACQUITY, Alliance, MassLynx, nanoacquity, and Xevo are registered trademarks and OpenLynx, Quanpedia, and TargetLynx
Agilent Triple Quadrupole LC/MS Peptide Quantitation with Skyline
 Agilent Triple Quadrupole LC/MS Peptide Quantitation with Skyline Workflow Guide A Peptide Optimization B Review Results in Skyline Create QQQ method in Skyline Edit Skyline settings Import peptides into
Agilent Triple Quadrupole LC/MS Peptide Quantitation with Skyline Workflow Guide A Peptide Optimization B Review Results in Skyline Create QQQ method in Skyline Edit Skyline settings Import peptides into
SequencePro Data Analysis Application. User Guide
 SequencePro Data Analysis Application User Guide SequencePro Data Analysis Application User Guide DRAFT October 31, 2001 12:52 pm, Title_page.fm Copyright 2001, Applied Biosystems. All rights reserved.
SequencePro Data Analysis Application User Guide SequencePro Data Analysis Application User Guide DRAFT October 31, 2001 12:52 pm, Title_page.fm Copyright 2001, Applied Biosystems. All rights reserved.
ECDL Module 4 REFERENCE MANUAL
 ECDL Module 4 REFERENCE MANUAL Spreadsheets Microsoft Excel XP Edition for ECDL Syllabus Four PAGE 2 - ECDL MODULE 4 (USING MICROSOFT EXCEL XP) - MANUAL 4.1 USING THE APPLICATION... 4 4.1.1 FIRST STEPS
ECDL Module 4 REFERENCE MANUAL Spreadsheets Microsoft Excel XP Edition for ECDL Syllabus Four PAGE 2 - ECDL MODULE 4 (USING MICROSOFT EXCEL XP) - MANUAL 4.1 USING THE APPLICATION... 4 4.1.1 FIRST STEPS
DESCRIPTION. DataApex Clarity 32bit multi instrument chromatography station for Windows
 wwwdataapexcom e-mail: clarity@dataapexcom DESCRIPTION DataApex Clarity 32bit multi instrument chromatography station for Windows Multi-detector instruments Easy to install and easy to use Graphical user
wwwdataapexcom e-mail: clarity@dataapexcom DESCRIPTION DataApex Clarity 32bit multi instrument chromatography station for Windows Multi-detector instruments Easy to install and easy to use Graphical user
Galaxie Photodiode Array Software
 Varian, Inc. 2700 Mitchell Drive Walnut Creek, CA 94598-1675/USA Galaxie Photodiode Array Software User s Guide Varian, Inc. 2002-2006 03-914950-00:6 Table of Contents Introduction... 3 General... 3 PDA
Varian, Inc. 2700 Mitchell Drive Walnut Creek, CA 94598-1675/USA Galaxie Photodiode Array Software User s Guide Varian, Inc. 2002-2006 03-914950-00:6 Table of Contents Introduction... 3 General... 3 PDA
MRMPROBS tutorial. Hiroshi Tsugawa RIKEN Center for Sustainable Resource Science MRMPROBS screenshot
 MRMPROBS tutorial Edited in 2016/11/16 Introduction MRMPROBS is launched as a universal program for targeted metabolomics using not only multiple reaction monitoring (MRM)- or selected reaction monitoring
MRMPROBS tutorial Edited in 2016/11/16 Introduction MRMPROBS is launched as a universal program for targeted metabolomics using not only multiple reaction monitoring (MRM)- or selected reaction monitoring
cief Data Analysis Chapter Overview Chapter 12:
 page 285 Chapter 12: cief Data Analysis Chapter Overview Analysis Screen Overview Opening Run Files How Run Data is Displayed Viewing Run Data Data Notifications and Warnings Checking Your Results Group
page 285 Chapter 12: cief Data Analysis Chapter Overview Analysis Screen Overview Opening Run Files How Run Data is Displayed Viewing Run Data Data Notifications and Warnings Checking Your Results Group
Excel Main Screen. Fundamental Concepts. General Keyboard Shortcuts Open a workbook Create New Save Preview and Print Close a Workbook
 Excel 2016 Main Screen Fundamental Concepts General Keyboard Shortcuts Open a workbook Create New Save Preview and Print Close a Ctrl + O Ctrl + N Ctrl + S Ctrl + P Ctrl + W Help Run Spell Check Calculate
Excel 2016 Main Screen Fundamental Concepts General Keyboard Shortcuts Open a workbook Create New Save Preview and Print Close a Ctrl + O Ctrl + N Ctrl + S Ctrl + P Ctrl + W Help Run Spell Check Calculate
Batch Processing (CasaXPS version and above)
 Copyright 2004, Casa Software Ltd. All Rights Reserved. 1 of 8 Batch Processing (CasaXPS version 2.2.50 and above) Batch processing is aimed at situations where a set of essentially equivalent samples
Copyright 2004, Casa Software Ltd. All Rights Reserved. 1 of 8 Batch Processing (CasaXPS version 2.2.50 and above) Batch processing is aimed at situations where a set of essentially equivalent samples
EXCEL BASICS: MICROSOFT OFFICE 2007
 EXCEL BASICS: MICROSOFT OFFICE 2007 GETTING STARTED PAGE 02 Prerequisites What You Will Learn USING MICROSOFT EXCEL PAGE 03 Opening Microsoft Excel Microsoft Excel Features Keyboard Review Pointer Shapes
EXCEL BASICS: MICROSOFT OFFICE 2007 GETTING STARTED PAGE 02 Prerequisites What You Will Learn USING MICROSOFT EXCEL PAGE 03 Opening Microsoft Excel Microsoft Excel Features Keyboard Review Pointer Shapes
Quant Browser for the Agilent Technologies GCMS Chemstation
 http://www.waleson.eu Quant Browser for the Agilent Technologies GCMS Chemstation Main purposes of Quant Browser Improve productivity of the quantitation process. Collect quantitation results of a complete
http://www.waleson.eu Quant Browser for the Agilent Technologies GCMS Chemstation Main purposes of Quant Browser Improve productivity of the quantitation process. Collect quantitation results of a complete
EZ IQ. Chromatography Software PN P1A
 O P E R A T I N G M A N U A L EZ IQ Chromatography Software Trademarks The trademarks of the products mentioned in this manual are held by the companies that produce them. Windows is a registered trademark
O P E R A T I N G M A N U A L EZ IQ Chromatography Software Trademarks The trademarks of the products mentioned in this manual are held by the companies that produce them. Windows is a registered trademark
GCSE CCEA GCSE EXCEL 2010 USER GUIDE. Business and Communication Systems
 GCSE CCEA GCSE EXCEL 2010 USER GUIDE Business and Communication Systems For first teaching from September 2017 Contents Page Define the purpose and uses of a spreadsheet... 3 Define a column, row, and
GCSE CCEA GCSE EXCEL 2010 USER GUIDE Business and Communication Systems For first teaching from September 2017 Contents Page Define the purpose and uses of a spreadsheet... 3 Define a column, row, and
MassHunter File Reader
 MassHunter File Reader vers 1.0.0 2015 Quadtech Associates, Inc. All Rights Reserved Release date: November 18, 2015 www.quadtechassociates.com MassHunter File Reader Welcome to MassHunter File Reader.
MassHunter File Reader vers 1.0.0 2015 Quadtech Associates, Inc. All Rights Reserved Release date: November 18, 2015 www.quadtechassociates.com MassHunter File Reader Welcome to MassHunter File Reader.
Installation Instructions
 Analyst QS Service Pack 5 (SP5) Release Notes Final Version Analyst QS Service Pack 5 includes all previous improvements released in Analyst QS SP1 through Analyst QS SP4. It addresses a number of issues
Analyst QS Service Pack 5 (SP5) Release Notes Final Version Analyst QS Service Pack 5 includes all previous improvements released in Analyst QS SP1 through Analyst QS SP4. It addresses a number of issues
Operating Manual. High Performance Liquid Chromatograph. Scientific Equipment Center, Prince Of Songkla University
 Operating Manual High Performance Liquid Chromatograph Agilent 1100 ; VWD, DAD, FLD and RID Scientific Equipment Center, Prince Of Songkla University Operating Manual High Performance Liquid Chromatograph
Operating Manual High Performance Liquid Chromatograph Agilent 1100 ; VWD, DAD, FLD and RID Scientific Equipment Center, Prince Of Songkla University Operating Manual High Performance Liquid Chromatograph
Building Agilent GC/MSD Deconvolution Reporting Libraries for Any Application Technical Overview
 Building Agilent GC/MSD Deconvolution Reporting Libraries for Any Application Technical Overview Authors Xiaofei Ping Agilent Technologies (Shanghai) Co Ltd 412 Yinglun Road, Shanghai, 200131 P.R. China
Building Agilent GC/MSD Deconvolution Reporting Libraries for Any Application Technical Overview Authors Xiaofei Ping Agilent Technologies (Shanghai) Co Ltd 412 Yinglun Road, Shanghai, 200131 P.R. China
StarFinder. Operation Manual. Star Chromatography Workstation Version 6
 Varian Analytical Instruments 2700 Mitchell Drive Walnut Creek, CA 94598-1675/USA Star Chromatography Workstation Version 6 StarFinder Operation Manual Varian, Inc. 2002 03-914723-00:Rev. 2 Trademark Acknowledgment
Varian Analytical Instruments 2700 Mitchell Drive Walnut Creek, CA 94598-1675/USA Star Chromatography Workstation Version 6 StarFinder Operation Manual Varian, Inc. 2002 03-914723-00:Rev. 2 Trademark Acknowledgment
Agilent ChemStation Plus
 Agilent ChemStation Plus Getting Started Guide Agilent Technologies Notices Agilent Technologies, Inc. 2004, 2006-2008 No part of this manual may be reproduced in any form or by any means (including electronic
Agilent ChemStation Plus Getting Started Guide Agilent Technologies Notices Agilent Technologies, Inc. 2004, 2006-2008 No part of this manual may be reproduced in any form or by any means (including electronic
Introduction to the workbook and spreadsheet
 Excel Tutorial To make the most of this tutorial I suggest you follow through it while sitting in front of a computer with Microsoft Excel running. This will allow you to try things out as you follow along.
Excel Tutorial To make the most of this tutorial I suggest you follow through it while sitting in front of a computer with Microsoft Excel running. This will allow you to try things out as you follow along.
Quick Reference Guide 8 Excel 2013 for Windows Keyboard Shortcut Keys
 Quick Reference Guide 8 Excel 2013 for Windows Keyboard Shortcut Keys Control Shortcut s Ctrl + PgDn Ctrl + PgUp Ctrl + Shift + & Ctrl + Shift_ Ctrl + Shift + ~ Ctrl + Shift + $ Ctrl + Shift + % Ctrl +
Quick Reference Guide 8 Excel 2013 for Windows Keyboard Shortcut Keys Control Shortcut s Ctrl + PgDn Ctrl + PgUp Ctrl + Shift + & Ctrl + Shift_ Ctrl + Shift + ~ Ctrl + Shift + $ Ctrl + Shift + % Ctrl +
-Table of Contents- 1. Overview Installation and removal Operation Main menu Trend graph... 13
 Thank you for buying Data Analysis Software. In order to use this software correctly and safely and to prevent trouble, please read this manual carefully. Notice 1. No part of this manual can be reproduced
Thank you for buying Data Analysis Software. In order to use this software correctly and safely and to prevent trouble, please read this manual carefully. Notice 1. No part of this manual can be reproduced
EXCEL 2003 DISCLAIMER:
 EXCEL 2003 DISCLAIMER: This reference guide is meant for experienced Microsoft Excel users. It provides a list of quick tips and shortcuts for familiar features. This guide does NOT replace training or
EXCEL 2003 DISCLAIMER: This reference guide is meant for experienced Microsoft Excel users. It provides a list of quick tips and shortcuts for familiar features. This guide does NOT replace training or
Skyline Targeted MS/MS
 Skyline Targeted MS/MS Skyline now supports several methods of extracting chromatography-based quantitative measurements from the raw data files of full-scan mass spectrometers, such as ion trap and Q-TOF
Skyline Targeted MS/MS Skyline now supports several methods of extracting chromatography-based quantitative measurements from the raw data files of full-scan mass spectrometers, such as ion trap and Q-TOF
Agilent MassHunter Metabolite ID Software. Installation and Getting Started Guide
 Agilent MassHunter Metabolite ID Software Installation and Getting Started Guide Notices Agilent Technologies, Inc. 2011 No part of this manual may be reproduced in any form or by any means (including
Agilent MassHunter Metabolite ID Software Installation and Getting Started Guide Notices Agilent Technologies, Inc. 2011 No part of this manual may be reproduced in any form or by any means (including
Cell to Cell mouse arrow Type Tab Enter Scroll Bars Page Up Page Down Crtl + Home Crtl + End Value Label Formula Note:
 1 of 1 NOTE: IT IS RECOMMENDED THAT YOU READ THE ACCOMPANYING DOCUMENT CALLED INTRO TO EXCEL LAYOUT 2007 TO FULLY GRASP THE BASICS OF EXCEL Introduction A spreadsheet application allows you to enter data
1 of 1 NOTE: IT IS RECOMMENDED THAT YOU READ THE ACCOMPANYING DOCUMENT CALLED INTRO TO EXCEL LAYOUT 2007 TO FULLY GRASP THE BASICS OF EXCEL Introduction A spreadsheet application allows you to enter data
Multivariate Calibration Quick Guide
 Last Updated: 06.06.2007 Table Of Contents 1. HOW TO CREATE CALIBRATION MODELS...1 1.1. Introduction into Multivariate Calibration Modelling... 1 1.1.1. Preparing Data... 1 1.2. Step 1: Calibration Wizard
Last Updated: 06.06.2007 Table Of Contents 1. HOW TO CREATE CALIBRATION MODELS...1 1.1. Introduction into Multivariate Calibration Modelling... 1 1.1.1. Preparing Data... 1 1.2. Step 1: Calibration Wizard
download instant at
 CHAPTER 1 - LAB SESSION INTRODUCTION TO EXCEL INTRODUCTION: This lab session is designed to introduce you to the statistical aspects of Microsoft Excel. During this session you will learn how to enter
CHAPTER 1 - LAB SESSION INTRODUCTION TO EXCEL INTRODUCTION: This lab session is designed to introduce you to the statistical aspects of Microsoft Excel. During this session you will learn how to enter
INTRODUCTION... 1 UNDERSTANDING CELLS... 2 CELL CONTENT... 4
 Introduction to Microsoft Excel 2016 INTRODUCTION... 1 The Excel 2016 Environment... 1 Worksheet Views... 2 UNDERSTANDING CELLS... 2 Select a Cell Range... 3 CELL CONTENT... 4 Enter and Edit Data... 4
Introduction to Microsoft Excel 2016 INTRODUCTION... 1 The Excel 2016 Environment... 1 Worksheet Views... 2 UNDERSTANDING CELLS... 2 Select a Cell Range... 3 CELL CONTENT... 4 Enter and Edit Data... 4
1. Math symbols Operation Symbol Example Order
 Excel 2 Microsoft Excel 2013 Mercer County Library System Brian M. Hughes, County Executive Excel s Order of Calculation 1. Math symbols Operation Symbol Example Order Parentheses ( ) =(4+2)*8 1st Exponents
Excel 2 Microsoft Excel 2013 Mercer County Library System Brian M. Hughes, County Executive Excel s Order of Calculation 1. Math symbols Operation Symbol Example Order Parentheses ( ) =(4+2)*8 1st Exponents
Excel Select a template category in the Office.com Templates section. 5. Click the Download button.
 Microsoft QUICK Excel 2010 Source Getting Started The Excel Window u v w z Creating a New Blank Workbook 2. Select New in the left pane. 3. Select the Blank workbook template in the Available Templates
Microsoft QUICK Excel 2010 Source Getting Started The Excel Window u v w z Creating a New Blank Workbook 2. Select New in the left pane. 3. Select the Blank workbook template in the Available Templates
Intro to Excel. To start a new workbook, click on the Blank workbook icon in the middle of the screen.
 Excel is a spreadsheet application that allows for the storing, organizing and manipulation of data that is entered into it. Excel has variety of built in tools that allow users to perform both simple
Excel is a spreadsheet application that allows for the storing, organizing and manipulation of data that is entered into it. Excel has variety of built in tools that allow users to perform both simple
Agilent MassHunter Workstation Software 7200 Accurate-Mass Quadrupole Time of Flight GC/MS
 Agilent MassHunter Workstation Software 7200 Accurate-Mass Quadrupole Time of Flight GC/MS Familiarization Guide Before you begin 3 Prepare your system 3 Prepare the samples required for data acquisition
Agilent MassHunter Workstation Software 7200 Accurate-Mass Quadrupole Time of Flight GC/MS Familiarization Guide Before you begin 3 Prepare your system 3 Prepare the samples required for data acquisition
"Excel"-erate Your Worksheets! Shortcuts and Power Tips NDSU Information Technology Services December 18, 2006
 "Excel"-erate Your Worksheets! Shortcuts and Power Tips NDSU Information Technology Services December 18, 2006 1. Check Which Version of Excel You're Using a. Click Help, About Microsoft Office Excel 2.
"Excel"-erate Your Worksheets! Shortcuts and Power Tips NDSU Information Technology Services December 18, 2006 1. Check Which Version of Excel You're Using a. Click Help, About Microsoft Office Excel 2.
Working with Data and Charts
 PART 9 Working with Data and Charts In Excel, a formula calculates a value based on the values in other cells of the workbook. Excel displays the result of a formula in a cell as a numeric value. A function
PART 9 Working with Data and Charts In Excel, a formula calculates a value based on the values in other cells of the workbook. Excel displays the result of a formula in a cell as a numeric value. A function
MS Excel Henrico County Public Library. I. Tour of the Excel Window
 MS Excel 2013 I. Tour of the Excel Window Start Excel by double-clicking on the Excel icon on the desktop. Excel may also be opened by clicking on the Start button>all Programs>Microsoft Office>Excel.
MS Excel 2013 I. Tour of the Excel Window Start Excel by double-clicking on the Excel icon on the desktop. Excel may also be opened by clicking on the Start button>all Programs>Microsoft Office>Excel.
TraceFinder Acquisition Quick Reference Guide
 TraceFinder Acquisition Quick Reference Guide This quick reference guide describes the Thermo TraceFinder acquisition tasks assigned to the Technician role. Contents Acquisition Mode Selecting a Batch
TraceFinder Acquisition Quick Reference Guide This quick reference guide describes the Thermo TraceFinder acquisition tasks assigned to the Technician role. Contents Acquisition Mode Selecting a Batch
Retention Time Locking with the MSD Productivity ChemStation. Technical Overview. Introduction. When Should I Lock My Methods?
 Retention Time Locking with the MSD Productivity ChemStation Technical Overview Introduction A retention time is the fundamental qualitative measurement of chromatography. Most peak identification is performed
Retention Time Locking with the MSD Productivity ChemStation Technical Overview Introduction A retention time is the fundamental qualitative measurement of chromatography. Most peak identification is performed
MALDI Software Users Guide (simplified version)
 MALDI Software Users Guide (simplified version) Central Analytical Lab Ekeley E266 The Acquisition program is called Voyager Control Panel. Once you shoot your sample and determine it looks good, you transfer
MALDI Software Users Guide (simplified version) Central Analytical Lab Ekeley E266 The Acquisition program is called Voyager Control Panel. Once you shoot your sample and determine it looks good, you transfer
Basic Microsoft Excel 2007
 Basic Microsoft Excel 2007 Contents Starting Excel... 2 Excel Window Properties... 2 The Ribbon... 3 Tabs... 3 Contextual Tabs... 3 Dialog Box Launchers... 4 Galleries... 5 Minimizing the Ribbon... 5 The
Basic Microsoft Excel 2007 Contents Starting Excel... 2 Excel Window Properties... 2 The Ribbon... 3 Tabs... 3 Contextual Tabs... 3 Dialog Box Launchers... 4 Galleries... 5 Minimizing the Ribbon... 5 The
Tutorials. Saturn GC/MS Workstation Version 5.4. Varian Analytical Instruments 2700 Mitchell Drive Walnut Creek, CA /usa
 Varian Analytical Instruments 2700 Mitchell Drive Walnut Creek, CA 94598-1675/usa Saturn GC/MS Workstation Version 5.4 Tutorials Varian, Inc. 1999 03-914765-00:Rev. 1 1 Contents How to Exercises...1 Overview...1
Varian Analytical Instruments 2700 Mitchell Drive Walnut Creek, CA 94598-1675/usa Saturn GC/MS Workstation Version 5.4 Tutorials Varian, Inc. 1999 03-914765-00:Rev. 1 1 Contents How to Exercises...1 Overview...1
Agilent MassHunter Workstation Software Report Designer Add-in
 Agilent MassHunter Workstation Software Report Designer Add-in Quick Start Guide What is the Agilent MassHunter Workstation Software Report Designer Add-in? 2 Report Designer UI elements 3 Getting Started
Agilent MassHunter Workstation Software Report Designer Add-in Quick Start Guide What is the Agilent MassHunter Workstation Software Report Designer Add-in? 2 Report Designer UI elements 3 Getting Started
Analyst QS. Administrator s Guide. Part Number: B July 2004
 Analyst QS Administrator s Guide Part Number: 1001930 B July 2004 http://www.appliedbiosystems.com This document is provided to customers who have purchased MDS Sciex equipment to use in the operation
Analyst QS Administrator s Guide Part Number: 1001930 B July 2004 http://www.appliedbiosystems.com This document is provided to customers who have purchased MDS Sciex equipment to use in the operation
MassHunter Personal Compound Database and Library Manager for Forensic Toxicology
 MassHunter Personal Compound Database and Library Manager for Forensic Toxicology Quick Start Guide What is MassHunter Personal Compound Database and Library Manager? 2 Installation 3 Main Window 4 Getting
MassHunter Personal Compound Database and Library Manager for Forensic Toxicology Quick Start Guide What is MassHunter Personal Compound Database and Library Manager? 2 Installation 3 Main Window 4 Getting
Getting low-resolution GC-MS data on the JEOL GCMate Introduction
 Getting low-resolution GC-MS data on the JEOL GCMate Introduction GC-MS is a technique that combines gas phase separation and mass-based detection/characterization of analytes. It s a very sensitive method
Getting low-resolution GC-MS data on the JEOL GCMate Introduction GC-MS is a technique that combines gas phase separation and mass-based detection/characterization of analytes. It s a very sensitive method
Microsoft Excel 2010 Basics
 Microsoft Excel 2010 Basics Starting Word 2010 with XP: Click the Start Button, All Programs, Microsoft Office, Microsoft Excel 2010 Starting Word 2010 with 07: Click the Microsoft Office Button with the
Microsoft Excel 2010 Basics Starting Word 2010 with XP: Click the Start Button, All Programs, Microsoft Office, Microsoft Excel 2010 Starting Word 2010 with 07: Click the Microsoft Office Button with the
Galaxie Workstation Configuration Manager
 Varian, Inc. 2700 Mitchell Drive Walnut Creek, CA 94598-1675/USA Galaxie Workstation Configuration Manager User s Guide Varian, Inc. 2002-2005 Printed in U.S.A. 03-914973-00:Rev. 3 Table of Contents Using
Varian, Inc. 2700 Mitchell Drive Walnut Creek, CA 94598-1675/USA Galaxie Workstation Configuration Manager User s Guide Varian, Inc. 2002-2005 Printed in U.S.A. 03-914973-00:Rev. 3 Table of Contents Using
Microsoft Excel 2010
 Microsoft Excel 2010 omar 2013-2014 First Semester 1. Exploring and Setting Up Your Excel Environment Microsoft Excel 2010 2013-2014 The Ribbon contains multiple tabs, each with several groups of commands.
Microsoft Excel 2010 omar 2013-2014 First Semester 1. Exploring and Setting Up Your Excel Environment Microsoft Excel 2010 2013-2014 The Ribbon contains multiple tabs, each with several groups of commands.
EQuIS Data Processor (EDP) User Manual
 EQuIS Data Processor (EDP) User Manual Introduction EQuIS Data Processor (EDP) Introduction The EQuIS Data Processor, or EDP, is today s answer to the many data quality issues that plague data managers.
EQuIS Data Processor (EDP) User Manual Introduction EQuIS Data Processor (EDP) Introduction The EQuIS Data Processor, or EDP, is today s answer to the many data quality issues that plague data managers.
P6 Professional Reporting Guide Version 18
 P6 Professional Reporting Guide Version 18 August 2018 Contents About the P6 Professional Reporting Guide... 7 Producing Reports and Graphics... 9 Report Basics... 9 Reporting features... 9 Report Wizard...
P6 Professional Reporting Guide Version 18 August 2018 Contents About the P6 Professional Reporting Guide... 7 Producing Reports and Graphics... 9 Report Basics... 9 Reporting features... 9 Report Wizard...
Lecture- 5. Introduction to Microsoft Excel
 Lecture- 5 Introduction to Microsoft Excel The Microsoft Excel Window Microsoft Excel is an electronic spreadsheet. You can use it to organize your data into rows and columns. You can also use it to perform
Lecture- 5 Introduction to Microsoft Excel The Microsoft Excel Window Microsoft Excel is an electronic spreadsheet. You can use it to organize your data into rows and columns. You can also use it to perform
Procedures for Group Quantitation
 Procedures for Group Quantitation Global Application Development Center Analytical & Measuring Instruments Division Shimadzu Corporation Group Quantitation In anionic surfactant analyses based on inspection
Procedures for Group Quantitation Global Application Development Center Analytical & Measuring Instruments Division Shimadzu Corporation Group Quantitation In anionic surfactant analyses based on inspection
Business Online TM. Positive Pay - Adding Issued Items. Quick Reference Guide
 Business Online TM Positive Pay - Adding Issued Items Quick Reference Guide Positive Pay Adding Issued Items Manually or Using Templates Positive Pay is a risk management solution that provides the ability
Business Online TM Positive Pay - Adding Issued Items Quick Reference Guide Positive Pay Adding Issued Items Manually or Using Templates Positive Pay is a risk management solution that provides the ability
Excel for Gen Chem General Chemistry Laboratory September 15, 2014
 Excel for Gen Chem General Chemistry Laboratory September 15, 2014 Excel is a ubiquitous data analysis software. Mastery of Excel can help you succeed in a first job and in your further studies with expertise
Excel for Gen Chem General Chemistry Laboratory September 15, 2014 Excel is a ubiquitous data analysis software. Mastery of Excel can help you succeed in a first job and in your further studies with expertise
Agilent MassHunter LC/SQ ChemStation Integration Software
 Agilent MassHunter LC/SQ ChemStation Integration Software Quick Start Guide What is the Agilent MassHunter LC/SQ ChemStation Integration Software? 2 Installation 4 Getting Started 5 LC/SQ Chemstation To
Agilent MassHunter LC/SQ ChemStation Integration Software Quick Start Guide What is the Agilent MassHunter LC/SQ ChemStation Integration Software? 2 Installation 4 Getting Started 5 LC/SQ Chemstation To
Skyline irt Retention Time Prediction
 Skyline irt Retention Time Prediction Predicting peptide retention time has long been of interest in targeted proteomics. As early as version 0.2, Skyline integrated the SSRCalc hydrophobicity calculator
Skyline irt Retention Time Prediction Predicting peptide retention time has long been of interest in targeted proteomics. As early as version 0.2, Skyline integrated the SSRCalc hydrophobicity calculator
