The TargetSearch Package
|
|
|
- Lee Lesley Chambers
- 6 years ago
- Views:
Transcription
1 The TargetSearch Package Alvaro Cuadros-Inostroza, Jan Lisec, Henning Redestig and Matthew A Hannah Max Planck Institute for Molecular Plant Physiology Potsdam, Germany October 30, 2017 This document describes how to use TargetSearch to preprocess GC-MS data. 1 Supplied Files This section describes the files that have to be prepared before running TargetSearch. These comprise a sample definition file, a reference library file and a retention marker definition file. Throughout this manual, we will use the example files provided by the package TargetSearchData. 1.1 NetCDF Files and Sample File TargetSearch can currently read only NetCDF files. Many GC-MS software packages are able to convert raw chromatograms to NetCDF. Although some baseline correction functionality is included in TargetSearch, it is recommended to baseline correct your chromatograms before exporting to NetCDF to increase processing speed. Please refer to your software documentation for further details. Export the NetCDF files into a convenient location. Then prepare a tab-delimited text file describing your samples. It must be contain (at least) the two columns entitled: CDF FILE and MEASUREMENT DAY. Other columns such as sample name, sample group, treatment, etc. may be additionally included to aid sample sub-setting and downstream analyses. An example is shown in table 1. To import the sample list into R, use the function ImportSamples() specifying the options CDFpath (directory containing the NetCDF files) and path (directory where the transformed cdf files, containing the retention index corrected apex data, also called -files, will be saved). 1
2 CDF FILE MEASUREMENT DAY TIME POINT 7235eg08.cdf eg11.cdf eg26.cdf eg04.cdf eg30.cdf eg32.cdf Table 1: Sample file example, samples.txt > library(targetsearchdata) > library(targetsearch) > cdf.path <- system.file("gc-ms-data", package = "TargetSearchData") > sample.file <- file.path(cdf.path, "samples.txt") > samples <- ImportSamples(sample.file, CDFpath = cdf.path, path = ".") Alternatively, you could create a tssample object by using the sample class methods. > cdffiles <- dir(cdf.path, pattern="cdf$") > # define the file names > rifiles <- paste("_", sub("cdf","txt", cdffiles), sep = "") > # take the measurement day info from the cdf file names. > days <- sub("^([[:digit:]]+).*$","\\1",cdffiles) > # sample names > smp_names <- sub("\\.cdf", "", cdffiles) > # add some sample info > smp_data <- data.frame(cdf_file =cdffiles, GROUP = gl(5,3)) > # create the sample object > samples <- new("tssample", Names = smp_names, CDFfiles = cdffiles, + CDFpath = cdf.path, path = ".", days = days, + files = rifiles, data = smp_data) 1.2 Retention Time () Markers To align different chromatograms using markers, a tab-delimited definition file for these markers has to be provided. At a minimum it should contain three columns specifying the search window (lower and upper threshod) and the fixed standard value for each marker (table 2). Further, a characteristic ion mass (m/z) has to be provided either in an additional column or as an argument mass to function ImportFameSettings(). > rim.file <- file.path(cdf.path, "rimlimits.txt") > rimlimits <- ImportFameSettings(rim.file, mass = 87) 2
3 LowerLimit UpperLimit standard Table 2: Retention time definition file example, rimlimits.txt This will import the limits in rimlimits.txt file and set the marker mass to 87 for all markers. If you do not use markers, you can skip this part by setting the parameter rim- Limits to NULL in correct function. Please note that in this case, no retention time correction will be performed. 2 Baseline correction, peak identification and correction Initially, TargetSearch identifies the local apex intensities in all chromatograms, finds the retention time (RT) of the markers and converts RT to using linear interpolation (Van den Dool and Kratz, 1963). This is done by the function correct(). Parameter to this function are sample and retention time limits objects, the mass range to extract ( MassRange = c(85,500)), the intensity threshhold (IntThreshold = 10 ), the peak picking method (pp.method) and a Window parameter that will be used by said method. The function will return a matrix with the retention times of all markers and create a tab-delimited file ( file) for each chromatogram in the specified directory. These files contain the extracted peak list of the respective NetCDF file. > matrix <- correct(samples, rimlimits, massrange = c(85,500), + IntThreshold = 10, pp.method = "smoothing", Window = 15) There are two peak picking methods available: smoothing implements the algorithm used by Tagfinder Luedemann et al. (2008), while the ppc algorithm is a adapted based on the function ppc.peaks from the package ppc. ppc.peaks searches for local maxima within a given time window. Therefore the Window parameter sets the window size of the chosen peak picking method. In case the chromatograms were not baseline corrected by the platform specific GC- MS software, it is possible to achieve that in TargetSearch by setting baseline to TRUE in function correct() which calls the function baselinecorrection(). This algorithm is based on the work of Chang et al. (2007) and a description is found in the baselinecorrection() documentation. Some options can be passed to baselinecorrection() via baseline.opts. After that, it is possible to check for outliers in the samples by using the function FAMEoutliers(). It creates a PDF report of the markers including information 3
4 about possible outliers that the user can remove or not. Alternatively, markers can be checked manually using the function plotfame (Figure 1). > outliers <- FAMEoutliers(samples, matrix, threshold = 3) Outliers Report: =============== No outliers were found. Here, threshold sets the number of standard deviations a value has to be away from the mean before it will be considered an outlier. In this example, however, no outliers were detected. > plotfame(samples, matrix, 1) Marker 1 R.T Samples Figure 1: Retention Index Marker 1. Note that TargetSearch assumes that the markers are injected together with the biological samples. A different approach is to inject them separately. If this is the case, please look at the script RetentionIndexCorrection.R for a workaround. 4
5 3 Library Search 3.1 Reference Library File The reference library file contains the information of the metabolites or mass spectral tags (MSTs) that will be searched for in the chromatograms. A public spectra database could be found here at The Golm Metabolome Database (Kopka et al., 2005; Hummel et al., 2007, 2013). Required information is the metabolite name ( Name ), expected retention time index ( ), selective masses ( SEL MASS ), most abundant masses ( TOP MASS ), spectrum ( SPECTRUM ) and deviations ( Win 1, Win 2, Win 3 ). See example in table 3. The columns Name and are mandatory and you have at least to include one of the columns SEL MASS, TOP MASS or SPECTRUM in the file (see below). The deviation columns are optional. Name Win 1 SEL MASS SPECTRUM Pyruvic acid ;115;158;174;189 85:7 86:14 87:7 88: Glycine (2TMS) ;102;147;176;204 86:26 87:19 88:8 89:4... Valine ;144;156;218;246 85:8 86:14 87:6 88: Glycerol (3TMS) ;117;205;293 85:14 86:2 87:16 88:13... Leucine ;158;232; : : :45... Isoleucine ;103;158;163;218 90:11 91:2 92:1 93: Glycine ;100;174;248;276 85:6 86:245 87:24 88:12... Table 3: Reference Library example, library.txt In this file, masses and intensities must be positive integers. s and deviations can be any positive real number. The selective and most abundant masses list must be delimited by semicolon (;). The spectrum is described by a list of mass and intensity pair. Every mass-intensity pair is separated by colon (:) and different pairs are separated by spaces. The function ImportLibrary() imports the reference file. > lib.file <- file.path(cdf.path, "library.txt") > lib <- ImportLibrary(lib.file, _dev = c(2000,1000,200), + TopMasses = 15, ExcludeMasses = c(147, 148, 149)) Here we set the window deviations to 2000, 1000 and 200 units. Since Win 1 column is already in the file, the first value (2000) is ignored. Also, the 15th most abundant masses are taken but excluding the masses 147, 148 and 149 (common confounding masses) 3.2 Library Search Algorithm The library search is performed in three steps. First, for every metabolite, selective masses are searched in a given time window around the expected. This is done by the 5
6 function medianlib(). This function calculates the median of the selective masses and return new library object with the updated. The time deviation is given either in the library file (column Win 1 ) or when the library is imported (see ImportLibrary()). > lib <- medianlib(samples, lib) It is also posible to examinate visually the deviation of the metabolites by setting the parameter makereport=true, which creates a pdf report like the one shown in figure 2. This may help to set or update the expected deviation. Pyruvic acid Glycine (2TMS) Valine Glycerol (3TMS) Leucine Isoleucine Glycine Benzoic acid Serine (major) Figure 2: deviation of first 9 metabolites in the library. In the second step, the function sample() searches the selective masses again, but using the updated and the deviation defined in the library object ( Win 2 ). After that, the intensities of the selected masses are normalised to the median of the day, and then used to extract other masses with correlated apex profiles. The masses for which the Pearson correlation coefficient is above r thres are taken as metabolite markers and 6
7 their s are averaged on a per sample basis. This average represents the exact position where the metabolite elutes in the respective sample, which is returned in a matrix form. > cor_ <- sample(samples, lib, r_thres = 0.95, + method = "daynorm") The third step will look up for all the masses (selective and most abundant masses) in all the samples. This is done by the function peakfind(). It returns a tsmsdata object with the intensities and of every mass (rows) and every sample (columns) that were search for. > peakdata <- peakfind(samples, lib, cor_) The intensity and slots can be accessed by using the Intensity and retindex methods. Each slot is a list, where every component of the list is a matrix, which is related to a metabolite in the library. There should be as many components as metabolites in the library. In every matrix, columns are samples and rows are masses. > met. <- retindex(peakdata) > met.intensity <- Intensity(peakData) > # show the intensity values of the first metabolite. > met.intensity[[1]] 7235eg eg eg eg eg eg NA NA NA NA NA NA NA eg eg eg eg eg eg NA NA
8 NA NA NA 1193 NA NA eg eg eg NA 325 NA NA NA NA NA NA NA Metabolite Profile The function Profile makes a profile of the MS data by averaging all the normalised mass intensities whose Pearson coefficient is greater that r thresh. > MetabProfile <- Profile(samples, lib, peakdata, r_thres = 0.95, + method = "daynorm") A msprofile object is returned. The averaged intensities and matrices that can be obtained by Intensity and retindex methods. The profile information is represented by a data.frame in the info slot (accessible by profileinfo method). The columns are: Name The metabolite/analyte name. Lib The expected (library ). Mass count The number of correlating masses. 8
9 Non consecutive Mass count Same as above, but not counting the consecutive masses. Sample Count per Mass The number of samples in which a correlating mass was found. The numbers are separated by semi-colon (;) and each number corresponds to one correlating masses (same order). If all the numbers are the same, only that unique number is shown. Masses The correlating masses. The average. Score all masses The similarity score calculated using the average intensity of all the masses that were searched for, regardless of whether they are correlating masses. Score cor masses Same as above, but only correlating masses are considered. As metabolites with similar selective masses and s can be present in metabolite libraries, it is necessary to reduce redundancy. This is performed by the function ProfileCleanUp which selects peaks for which the gap is smaller than timesplit and computes the Pearson correlation between them. When two metabolites within such a time-group are tightly correlated (given by r thres) only the one with more correlated masses is retained. > finalprofile <- ProfileCleanUp(MetabProfile, timesplit = 500, + r_thres = 0.95) The function returns a msprofile object. The info slot is similar as described above, but extra columns with a Cor preffix (e.g., Cor Name ) are included. They provide information about metabolite redundancy. 5 Peaks and Spectra Visualisation Finally, it may be of interest to check the chromatographic peak of selected metabolites and compare the median spectra of the metabolites, i.e., the median intensities of the selected masses across all the samples, with the reference spectra of the library. There are two functions to do so: plotpeak and plotspectra. For example, we can compare the median spectrum of Valine against its spectrum reference. Here we look for the library index of Valine and plot the spectra comparison in a head-tail plot (figure 3). To look at the chromatographic peak of Valine in a given sample, we use the functions peakcdfextraction to extract the raw chromatogram and plotpeak to plot the peak (figure 4). Refer to the documentation of the functions plotpeak and plotspectra for further options not covered here. 9
10 > grep("valine", libname(lib)) [1] 3 > plotspectra(lib, peakdata, libid = 3, type = "ht") Valine Intensity median spectrum reference spectrum Figure 3: Spectra comparison of Valine 10
11 > # we select the first sample > sample.id <- 1 > cdf.file <- file.path(cdf.path, cdffiles[sample.id]) > rawpeaks <- peakcdfextraction(cdf.file, massrange = c(85,500)) > # which.met=3 (id of Valine) > plotpeak(samples, lib, MetabProfile, rawpeaks, which.smp=sample.id, + which.met=3, massrange=c(85,500), cormass=false) Sample: 7235eg04 Metab: Valine Intensity Retention Index () Figure 4: Chromatographic peak of Valine. 11
12 6 TargetSearch GUI We provide a grahical user interface intended to facilitate the use of TargetSearch for users unfamiliar with R. Many parameters that would be set calling the individual TargetSearch functions as described in this document can be set here in one go before running the complete analysis. A screenshot of the GUI is shown in figure 5. Figure 5: The TargetSearch GUI. This is a descript of all the GUI options. Working Directory : Use the Browse-button to select the folder on your hard drive 12
13 containing all your GC-MS data files. The output of TargetSearch will be written to this folder too. File Import : Clicking NetCDF Data or Apex Data radio buttons will open a file select dialog. Choose the files you would like to be processed. You may check your selection pressing the Show-button. Baseline Correction : Clicking on/off button will perform baseline correction before peak detection. If selected, the threshold parameter is a numeric value between 0 and 1. A value of one returns a baseline above the noise, 0.5 in the middle of the noise and 0 below the noise. See baselinecorrection documentation for further details. Peak Detection : Intensity Counts threshold defines the minimum apex intensity incorporated in the analysis. A value of 1 would include all peaks. Mass Range allows to limit the mass values (m/z) to be included in the analysis. Smoothing averages raw data to eliminate some inherent noise leading to multiple peaks otherwise. Retention Index Correction : Retention Index Correction is neccessary and applied only if you supply NetCDF Data (Apex Data contain already Retention Indices). You may Load or Create the search windows for your -Markers here. Library : A Library (to detect metabolites) usable by TargetSearch contains at least information about the metabolite Name, its expected and the selective masses in its spectrum SEL MASS. You may Load or Create one yourself using the respective buttons. A more detailed description of the file formats can be found in ImportLibrary. Search Windows refers to the allowed deviation of your metabolites which are narrowed in 3 consecutive searches. No. of top masses is the number of most abundant masses that will be selected from the spectra, besides the selected masses. ExcludeMasses is only used when masses are obtained from the spectra. For example, 126,147:149,160 means that the masses 126, 147, 148, 149, and 160 will be excluded from the analysis (unless they are used as selected masses). Normalization : This selects how the data will be normalized during the metabolite search. Options are daynorm, a day based median normalization, mediannorm, normalization using the median of all the intensities of a given mass, and none, no normalization at all. Final Profiles : Here you may set the parameters used by the functions Profile and ProfileCleanUp. timesplit sets an window that will be used to look for metabolites that could have been redundanly identified. correl. thr. is the correlation threshold and min. number of correlation samples is a threshold used to make sure that correlations are computed with at least said number of observations. 13
14 prioritization is used to resolv ambiguities by taking the metabolite with the most correlating masses (corr. mass) or the highest similarity score. Metabolites with corr. mass and score higher than the thresholds will be suggested above the one that do not pass the thresholds, which are marked as unidentified. Quantification Mass : This section is used to generate an intensity matrix based on one quantification mass. The quantification mass can be specified in the library file (column QUANT MASS ) or selected automatically by choosing max intensity or max observations. max intensity selects the mass with the highest intensity across the samples, while max observation takes the mass with less missing values. Parameters : You may Save the current parameters as an *.RData file or Load previously saved parameters to compare the outcome of different settings or just repeat the analysis. Program : Run starts to process all currently selected files using the current parameters and saving output to Working Directory. Quit closes the GUI. 7 Untargeted search Although TargetSearch was designed to be targeted oriented, it is possible to perform untargeted searches. The basic idea is to create a library that contains evenly distributed metabolites in time and every metabolite uses the whole range of possible masses as selective masses. An example: > met <- seq(200000, , by = 5000) > metmz <- 85:250 > metnames <- paste("metab",format(met,digits=6), sep = "_") Here we define a set of metabolites located every 5000 units from each other in the range , with selective masses in the range of , and assign them a name. After that, we create an tslib object. > metlib <- new("tslib", Name = metnames, = met, + selmass = rep(list(metmz), length(met)), dev = c(3000, 1500, 500)) Now we can use this library object to perform a targetive search with this library on the E. coli samples as we did before. > metlib <- medianlib(samples, metlib) > metcor <- sample(samples, metlib) > metpeakdata <- peakfind(samples, metlib, metcor) > metprofile <- Profile(samples, metlib, metpeakdata) > metfinprof <- ProfileCleanUp(metProfile, timesplit = 500) 14
15 The metfinprof object can be used to create a new library by taking only the metabolites that have, for example, more than 5 correlating masses and using the correlating masses as selective ones. > sum( profileinfo(metfinprof)$mass_count > 5) [1] 13 > tmp <- profileinfo(metfinprof)$mass_count > 5 > met <- profileinfo(metfinprof)$[tmp] > metnames <- as.character( profileinfo(metfinprof)$name[tmp] ) > metmz <- sapply(profileinfo(metfinprof)$masses[tmp], + function(x) as.numeric(unlist(strsplit(x,";"))) ) > metlib <- new("tslib", Name = metnames, = met, + selmass = metmz, dev = c(1500, 750, 250)) After the new library object is created, the process can be repeated as shown above. Finally, by using the function writemsp, it is possible to export the spectrum of the unknown metabolites to MSP format used by NIST mass spectra search ( so the unknown spectra can be search against known metabolite spectra databases. References D. Chang, C. D. Banack, and S. L. Shah. Robust baseline correction algorithm for signal dense NMR spectra. J. Magn. Reson., 187(2): , J. Hummel, J. Selbig, D. Walther, and J. Kopka. The golm metabolome database: a database for gc-ms based metabolite profiling. In J. Nielsen and M.C. Jewett, editors, Metabolomics, volume 18, pages Springer-Verlag, Berlin, Heidelberg, New York, ISBN doi: / J. Hummel, N. Strehmel, C. Bölling, S. Schmidt, D. Walther, and J. Kopka. Mass spectral search and analysis using the golm metabolome database. In The Handbook of Plant Metabolomics, pages Wiley-VCH Verlag GmbH & Co. KGaA, doi: / ch18. J. Kopka, N. Schauer, S. Krueger, C. Birkemeyer, B. Usadel, E. Bergmuller, P. Dormann, W. Weckwerth, Y. Gibon, M. Stitt, L. Willmitzer, A. R. Fernie, and D. Steinhauser. Gmd@csb.db: the Golm Metabolome database. Bioinformatics, 21(8): , A. Luedemann, K. Strassburg, A. Erban, and J. Kopka. Tagfinder for the quantitative analysis of gas chromatography - mass spectrometry (GC-MS)-based metabolite profiling experiments. Bioinformatics, 24(5): ,
16 H. Van den Dool and P. D. Kratz. A generalization of retention index system including linear temperature programmed gas-liquid partition chromatography. J. Chromatography, 11(4):463 &,
An introduction to Baitmet package
 An introduction to Baitmet package Version 1.0.1 Xavier Domingo-Almenara (Maintainer) xdomingo@scripps.edu May 17, 2017 This vignette presents Baitmet, an R package for library driven compound profiling
An introduction to Baitmet package Version 1.0.1 Xavier Domingo-Almenara (Maintainer) xdomingo@scripps.edu May 17, 2017 This vignette presents Baitmet, an R package for library driven compound profiling
Package Metab. September 18, 2018
 Version 1.14.0 Date 2013-10-11 Package Metab September 18, 2018 Title Metab: An R Package for a High-Throughput Analysis of Metabolomics Data Generated by GC-MS. Author Raphael Aggio
Version 1.14.0 Date 2013-10-11 Package Metab September 18, 2018 Title Metab: An R Package for a High-Throughput Analysis of Metabolomics Data Generated by GC-MS. Author Raphael Aggio
Progenesis CoMet User Guide
 Progenesis CoMet User Guide Analysis workflow guidelines for version 2.0 Contents Introduction... 3 How to use this document... 3 How can I analyse my own runs using CoMet?... 3 LC-MS Data used in this
Progenesis CoMet User Guide Analysis workflow guidelines for version 2.0 Contents Introduction... 3 How to use this document... 3 How can I analyse my own runs using CoMet?... 3 LC-MS Data used in this
Tutorial 2: Analysis of DIA/SWATH data in Skyline
 Tutorial 2: Analysis of DIA/SWATH data in Skyline In this tutorial we will learn how to use Skyline to perform targeted post-acquisition analysis for peptide and inferred protein detection and quantification.
Tutorial 2: Analysis of DIA/SWATH data in Skyline In this tutorial we will learn how to use Skyline to perform targeted post-acquisition analysis for peptide and inferred protein detection and quantification.
Data processing. Filters and normalisation. Mélanie Pétéra W4M Core Team 31/05/2017 v 1.0.0
 Data processing Filters and normalisation Mélanie Pétéra W4M Core Team 31/05/2017 v 1.0.0 Presentation map 1) Processing the data W4M table format for Galaxy 2) A generic tool to filter in Galaxy a) Generic
Data processing Filters and normalisation Mélanie Pétéra W4M Core Team 31/05/2017 v 1.0.0 Presentation map 1) Processing the data W4M table format for Galaxy 2) A generic tool to filter in Galaxy a) Generic
MS data processing. Filtering and correcting data. W4M Core Team. 22/09/2015 v 1.0.0
 MS data processing Filtering and correcting data W4M Core Team 22/09/2015 v 1.0.0 Presentation map 1) Processing the data W4M table format for Galaxy 2) Filters for mass spectrometry extracted data a)
MS data processing Filtering and correcting data W4M Core Team 22/09/2015 v 1.0.0 Presentation map 1) Processing the data W4M table format for Galaxy 2) Filters for mass spectrometry extracted data a)
Package HiResTEC. August 7, 2018
 Type Package Package HiResTEC August 7, 2018 Title Non-Targeted Fluxomics on High-Resolution Mass-Spectrometry Data Version 0.54 Date 2018-08-07 Maintainer Jan Lisec Identifying labeled
Type Package Package HiResTEC August 7, 2018 Title Non-Targeted Fluxomics on High-Resolution Mass-Spectrometry Data Version 0.54 Date 2018-08-07 Maintainer Jan Lisec Identifying labeled
SIMAT: GC-SIM-MS Analayis Tool
 SIMAT: GC-SIM-MS Analayis Tool Mo R. Nezami Ranjbar September 10, 2018 1 Selected Ion Monitoring Gas chromatography coupled with mass spectrometry (GC-MS) is one of the promising technologies for qualitative
SIMAT: GC-SIM-MS Analayis Tool Mo R. Nezami Ranjbar September 10, 2018 1 Selected Ion Monitoring Gas chromatography coupled with mass spectrometry (GC-MS) is one of the promising technologies for qualitative
Thermo Xcalibur Getting Started (Quantitative Analysis)
 Thermo Xcalibur Getting Started (Quantitative Analysis) XCALI-97207 Revision B September 2010 2010 Thermo Fisher Scientific Inc. All rights reserved. Xcalibur, Surveyor, and Accela are registered trademarks
Thermo Xcalibur Getting Started (Quantitative Analysis) XCALI-97207 Revision B September 2010 2010 Thermo Fisher Scientific Inc. All rights reserved. Xcalibur, Surveyor, and Accela are registered trademarks
Progenesis CoMet User Guide
 Progenesis CoMet User Guide Analysis workflow guidelines for version 1.0 Contents Introduction... 3 How to use this document... 3 How can I analyse my own runs using CoMet?... 3 LC-MS Data used in this
Progenesis CoMet User Guide Analysis workflow guidelines for version 1.0 Contents Introduction... 3 How to use this document... 3 How can I analyse my own runs using CoMet?... 3 LC-MS Data used in this
High-throughput Processing and Analysis of LC-MS Spectra
 High-throughput Processing and Analysis of LC-MS Spectra By Jianguo Xia (jianguox@ualberta.ca) Last update : 02/05/2012 This tutorial shows how to process and analyze LC-MS spectra using methods provided
High-throughput Processing and Analysis of LC-MS Spectra By Jianguo Xia (jianguox@ualberta.ca) Last update : 02/05/2012 This tutorial shows how to process and analyze LC-MS spectra using methods provided
Tutorial 7: Automated Peak Picking in Skyline
 Tutorial 7: Automated Peak Picking in Skyline Skyline now supports the ability to create custom advanced peak picking and scoring models for both selected reaction monitoring (SRM) and data-independent
Tutorial 7: Automated Peak Picking in Skyline Skyline now supports the ability to create custom advanced peak picking and scoring models for both selected reaction monitoring (SRM) and data-independent
Protein Deconvolution Quick Start Guide
 Protein Deconvolution Quick Start Guide The electrospray ionization (ESI) of intact peptides and proteins produces mass spectra containing a series of multiply charged ions with associated mass-to-charge
Protein Deconvolution Quick Start Guide The electrospray ionization (ESI) of intact peptides and proteins produces mass spectra containing a series of multiply charged ions with associated mass-to-charge
Package cosmiq. April 11, 2018
 Type Package Package cosmiq April 11, 2018 Title cosmiq - COmbining Single Masses Into Quantities Version 1.12.0 Author David Fischer , Christian Panse , Endre
Type Package Package cosmiq April 11, 2018 Title cosmiq - COmbining Single Masses Into Quantities Version 1.12.0 Author David Fischer , Christian Panse , Endre
TraceFinder Analysis Quick Reference Guide
 TraceFinder Analysis Quick Reference Guide This quick reference guide describes the Analysis mode tasks assigned to the Technician role in the Thermo TraceFinder 3.0 analytical software. For detailed descriptions
TraceFinder Analysis Quick Reference Guide This quick reference guide describes the Analysis mode tasks assigned to the Technician role in the Thermo TraceFinder 3.0 analytical software. For detailed descriptions
Package CorrectOverloadedPeaks
 Type Package Package CorrectOverloadedPeaks July 10, 2018 Title Correct Overloaded Peaks from GC-APCI-MS Data Version 1.2.15 Date 2018-07-10 Author Jan Lisec [aut, cre] Analyzes and modifies metabolomics
Type Package Package CorrectOverloadedPeaks July 10, 2018 Title Correct Overloaded Peaks from GC-APCI-MS Data Version 1.2.15 Date 2018-07-10 Author Jan Lisec [aut, cre] Analyzes and modifies metabolomics
Analysis of GC-MS metabolomics data with metams
 Analysis of GC-MS metabolomics data with metams Ron Wehrens October 30, 2017 1 Introduction Many packages are available for the analysis of data from GC-MS and LC-MS experiments typically, hardware vendors
Analysis of GC-MS metabolomics data with metams Ron Wehrens October 30, 2017 1 Introduction Many packages are available for the analysis of data from GC-MS and LC-MS experiments typically, hardware vendors
Building Agilent GC/MSD Deconvolution Reporting Libraries for Any Application Technical Overview
 Building Agilent GC/MSD Deconvolution Reporting Libraries for Any Application Technical Overview Authors Xiaofei Ping Agilent Technologies (Shanghai) Co Ltd 412 Yinglun Road, Shanghai, 200131 P.R. China
Building Agilent GC/MSD Deconvolution Reporting Libraries for Any Application Technical Overview Authors Xiaofei Ping Agilent Technologies (Shanghai) Co Ltd 412 Yinglun Road, Shanghai, 200131 P.R. China
Quick Guide to the Star Bar
 Saturn View Saturn Writer Quick Guide to the Star Bar Last active Chromatogram in SaturnView Last active method in the Method Editor Click with the right mouse button on the Star Bar to get this menu Sample,
Saturn View Saturn Writer Quick Guide to the Star Bar Last active Chromatogram in SaturnView Last active method in the Method Editor Click with the right mouse button on the Star Bar to get this menu Sample,
MRMPROBS tutorial. Hiroshi Tsugawa RIKEN Center for Sustainable Resource Science
 MRMPROBS tutorial Edited in 2014/10/6 Introduction MRMPROBS is a tool for the analysis of data from multiple reaction monitoring (MRM)- or selected reaction monitoring (SRM)-based metabolomics studies.
MRMPROBS tutorial Edited in 2014/10/6 Introduction MRMPROBS is a tool for the analysis of data from multiple reaction monitoring (MRM)- or selected reaction monitoring (SRM)-based metabolomics studies.
Adaptive Processing of LC/MS Metabolomics data
 Adaptive Processing of LC/MS Metabolomics data Tianwei Yu August 1, 2012 1. Introduction The aplcms package is designed for the processing of high resolution LC/MS data. The main characteristics include:
Adaptive Processing of LC/MS Metabolomics data Tianwei Yu August 1, 2012 1. Introduction The aplcms package is designed for the processing of high resolution LC/MS data. The main characteristics include:
Xcalibur Library Browser
 Thermo Xcalibur Library Browser User Guide Creating and Searching Spectral Libraries Software Version 3.0 XCALI-97552 Revision A June 2013 2013 Thermo Fisher Scientific Inc. All rights reserved. Xcalibur
Thermo Xcalibur Library Browser User Guide Creating and Searching Spectral Libraries Software Version 3.0 XCALI-97552 Revision A June 2013 2013 Thermo Fisher Scientific Inc. All rights reserved. Xcalibur
Skyline MS1 Full Scan Filtering
 Skyline MS1 Full Scan Filtering The Skyline Targeted Proteomics Environment provides informative visual displays of the raw mass spectrometer data you import into your Skyline project. These displays allow
Skyline MS1 Full Scan Filtering The Skyline Targeted Proteomics Environment provides informative visual displays of the raw mass spectrometer data you import into your Skyline project. These displays allow
Reference Manual MarkerView Software Reference Manual Revision: February, 2010
 Reference Manual MarkerView 1.2.1 Software Reference Manual - 1 - Revision: February, 2010 This document is provided to customers who have purchased AB SCIEX equipment to use in the operation of such AB
Reference Manual MarkerView 1.2.1 Software Reference Manual - 1 - Revision: February, 2010 This document is provided to customers who have purchased AB SCIEX equipment to use in the operation of such AB
Metabolomic Data Analysis with MetaboAnalyst
 Metabolomic Data Analysis with MetaboAnalyst User ID: guest6522519400069885256 April 14, 2009 1 Data Processing and Normalization 1.1 Reading and Processing the Raw Data MetaboAnalyst accepts a variety
Metabolomic Data Analysis with MetaboAnalyst User ID: guest6522519400069885256 April 14, 2009 1 Data Processing and Normalization 1.1 Reading and Processing the Raw Data MetaboAnalyst accepts a variety
MS-FINDER tutorial. Last edited in Aug 10, 2018
 MS-FINDER tutorial Last edited in Aug 10, 2018 Abstract The purpose of metabolomics is to perform the comprehensive analysis for small biomolecules of living organisms. Gas chromatography coupled with
MS-FINDER tutorial Last edited in Aug 10, 2018 Abstract The purpose of metabolomics is to perform the comprehensive analysis for small biomolecules of living organisms. Gas chromatography coupled with
MRMPROBS tutorial. Hiroshi Tsugawa RIKEN Center for Sustainable Resource Science MRMPROBS screenshot
 MRMPROBS tutorial Edited in 2016/11/16 Introduction MRMPROBS is launched as a universal program for targeted metabolomics using not only multiple reaction monitoring (MRM)- or selected reaction monitoring
MRMPROBS tutorial Edited in 2016/11/16 Introduction MRMPROBS is launched as a universal program for targeted metabolomics using not only multiple reaction monitoring (MRM)- or selected reaction monitoring
Chromatography Software Training Materials. Contents
 Chromatography Software Training Materials This document contains information on how to build a method, start the instrument to acquire data, and then process the data using the Galaxie Program. You will
Chromatography Software Training Materials This document contains information on how to build a method, start the instrument to acquire data, and then process the data using the Galaxie Program. You will
TraceFinder Analysis Quick Reference Guide
 TraceFinder Analysis Quick Reference Guide This quick reference guide describes the Analysis mode tasks assigned to the Technician role in Thermo TraceFinder analytical software. For detailed descriptions
TraceFinder Analysis Quick Reference Guide This quick reference guide describes the Analysis mode tasks assigned to the Technician role in Thermo TraceFinder analytical software. For detailed descriptions
REVIEW. MassFinder 3. Navigation of the Chromatogram
 REVIEW MassFinder 3 Dr. Detlev H. Hochmuth, Author Hochmuth Scientific Consulting Hamburg, Germany http://www.massfinder.com Euro 1850, which includes latest version of MassFinder 3, a terpene spectral
REVIEW MassFinder 3 Dr. Detlev H. Hochmuth, Author Hochmuth Scientific Consulting Hamburg, Germany http://www.massfinder.com Euro 1850, which includes latest version of MassFinder 3, a terpene spectral
Statistical Process Control in Proteomics SProCoP
 Statistical Process Control in Proteomics SProCoP This tutorial will guide you through the installation of SProCoP and using it to perform statistical analysis on a sample Skyline file. Getting Started
Statistical Process Control in Proteomics SProCoP This tutorial will guide you through the installation of SProCoP and using it to perform statistical analysis on a sample Skyline file. Getting Started
Retention Time Locking with the MSD Productivity ChemStation. Technical Overview. Introduction. When Should I Lock My Methods?
 Retention Time Locking with the MSD Productivity ChemStation Technical Overview Introduction A retention time is the fundamental qualitative measurement of chromatography. Most peak identification is performed
Retention Time Locking with the MSD Productivity ChemStation Technical Overview Introduction A retention time is the fundamental qualitative measurement of chromatography. Most peak identification is performed
Agilent 6400 Series Triple Quadrupole LC/MS System
 Agilent 6400 Series Triple Quadrupole LC/MS System Quick Start Guide Where to find information 4 Getting Started 6 Step 1. Start the Data Acquisition software 7 Step 2. Prepare the LC modules 13 Step 3.
Agilent 6400 Series Triple Quadrupole LC/MS System Quick Start Guide Where to find information 4 Getting Started 6 Step 1. Start the Data Acquisition software 7 Step 2. Prepare the LC modules 13 Step 3.
Galaxie Photodiode Array Software
 Varian, Inc. 2700 Mitchell Drive Walnut Creek, CA 94598-1675/USA Galaxie Photodiode Array Software User s Guide Varian, Inc. 2002-2006 03-914950-00:6 Table of Contents Introduction... 3 General... 3 PDA
Varian, Inc. 2700 Mitchell Drive Walnut Creek, CA 94598-1675/USA Galaxie Photodiode Array Software User s Guide Varian, Inc. 2002-2006 03-914950-00:6 Table of Contents Introduction... 3 General... 3 PDA
ChromQuest 5.0 Quick Reference Guide
 ChromQuest 5.0 Quick Reference Guide This guide contains an overview of the ChromQuest chromatography data system, with topics organized by workflow. For more information, refer to the ChromQuest User
ChromQuest 5.0 Quick Reference Guide This guide contains an overview of the ChromQuest chromatography data system, with topics organized by workflow. For more information, refer to the ChromQuest User
Skyline High Resolution Metabolomics (Draft)
 Skyline High Resolution Metabolomics (Draft) The Skyline Targeted Proteomics Environment provides informative visual displays of the raw mass spectrometer data you import into your Skyline documents. Originally
Skyline High Resolution Metabolomics (Draft) The Skyline Targeted Proteomics Environment provides informative visual displays of the raw mass spectrometer data you import into your Skyline documents. Originally
Progenesis QI for proteomics User Guide. Analysis workflow guidelines for DDA data
 Progenesis QI for proteomics User Guide Analysis workflow guidelines for DDA data Contents Introduction... 3 How to use this document... 3 How can I analyse my own runs using Progenesis QI for proteomics?...
Progenesis QI for proteomics User Guide Analysis workflow guidelines for DDA data Contents Introduction... 3 How to use this document... 3 How can I analyse my own runs using Progenesis QI for proteomics?...
Package envipick. June 6, 2016
 Type Package Package envipick June 6, 2016 Title Peak Picking for High Resolution Mass Spectrometry Data Version 1.5 Date 2016-06-03 Author Maintainer Sequential partitioning, clustering
Type Package Package envipick June 6, 2016 Title Peak Picking for High Resolution Mass Spectrometry Data Version 1.5 Date 2016-06-03 Author Maintainer Sequential partitioning, clustering
NMRProcFlow Macro-command Reference Guide
 NMRProcFlow Macro-command Reference Guide This document is the reference guide of the macro-commands Daniel Jacob UMR 1332 BFP, Metabolomics Facility CGFB Bordeaux, MetaboHUB - 2018 1 NMRProcFlow - Macro-command
NMRProcFlow Macro-command Reference Guide This document is the reference guide of the macro-commands Daniel Jacob UMR 1332 BFP, Metabolomics Facility CGFB Bordeaux, MetaboHUB - 2018 1 NMRProcFlow - Macro-command
Using OPUS to Process Evolved Gas Data (8/12/15 edits highlighted)
 Using OPUS to Process Evolved Gas Data (8/12/15 edits highlighted) Once FTIR data has been acquired for the gases evolved during your DSC/TGA run, you will process using the OPUS software package. Select
Using OPUS to Process Evolved Gas Data (8/12/15 edits highlighted) Once FTIR data has been acquired for the gases evolved during your DSC/TGA run, you will process using the OPUS software package. Select
MassHunter File Reader
 MassHunter File Reader vers 1.0.0 2015 Quadtech Associates, Inc. All Rights Reserved Release date: November 18, 2015 www.quadtechassociates.com MassHunter File Reader Welcome to MassHunter File Reader.
MassHunter File Reader vers 1.0.0 2015 Quadtech Associates, Inc. All Rights Reserved Release date: November 18, 2015 www.quadtechassociates.com MassHunter File Reader Welcome to MassHunter File Reader.
MS-FINDER tutorial. Last edited in Sep. 10, 2018
 MS-FINDER tutorial Last edited in Sep. 10, 2018 Abstract The purpose of metabolomics is to perform the comprehensive analysis for small biomolecules of living organisms. Gas chromatography coupled with
MS-FINDER tutorial Last edited in Sep. 10, 2018 Abstract The purpose of metabolomics is to perform the comprehensive analysis for small biomolecules of living organisms. Gas chromatography coupled with
MassHunter Personal Compound Database and Library Manager for Forensic Toxicology
 MassHunter Personal Compound Database and Library Manager for Forensic Toxicology Quick Start Guide What is MassHunter Personal Compound Database and Library Manager? 2 Installation 3 Main Window 4 Getting
MassHunter Personal Compound Database and Library Manager for Forensic Toxicology Quick Start Guide What is MassHunter Personal Compound Database and Library Manager? 2 Installation 3 Main Window 4 Getting
Skyline irt Retention Time Prediction
 Skyline irt Retention Time Prediction Predicting peptide retention time has long been of interest in targeted proteomics. As early as version 0.2, Skyline integrated the SSRCalc hydrophobicity calculator
Skyline irt Retention Time Prediction Predicting peptide retention time has long been of interest in targeted proteomics. As early as version 0.2, Skyline integrated the SSRCalc hydrophobicity calculator
Chromeleon software orientation
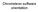 Chromeleon software orientation Upon opening of Chromeleon shortcut, a blue screen should appear (called control panel). If this does not occur, the green circled shortcut will open this screen. To ensure
Chromeleon software orientation Upon opening of Chromeleon shortcut, a blue screen should appear (called control panel). If this does not occur, the green circled shortcut will open this screen. To ensure
Agilent G6854 MassHunter Personal Pesticide Database
 Agilent G6854 MassHunter Personal Pesticide Database Quick Start Guide What is MassHunter Personal Pesticide Database? 2 Installation 3 Main Window 4 Getting Started 11 Database operations 12 Searching
Agilent G6854 MassHunter Personal Pesticide Database Quick Start Guide What is MassHunter Personal Pesticide Database? 2 Installation 3 Main Window 4 Getting Started 11 Database operations 12 Searching
Annotation of LC-MS metabolomics datasets by the metams package
 Annotation of LC-MS metabolomics datasets by the metams package Pietro Franceschi April 30, 2018 1 Introduction metams is designed to perform the analysis of LC-MS and GC-MSbased metabolomics assays. For
Annotation of LC-MS metabolomics datasets by the metams package Pietro Franceschi April 30, 2018 1 Introduction metams is designed to perform the analysis of LC-MS and GC-MSbased metabolomics assays. For
Skyline Targeted Method Refinement
 Skyline Targeted Method Refinement This tutorial will introduce the features available in the Skyline Targeted Proteomics Environment for refining instrument methods for Selected Reaction Monitoring (SRM,
Skyline Targeted Method Refinement This tutorial will introduce the features available in the Skyline Targeted Proteomics Environment for refining instrument methods for Selected Reaction Monitoring (SRM,
Project Report on. De novo Peptide Sequencing. Course: Math 574 Gaurav Kulkarni Washington State University
 Project Report on De novo Peptide Sequencing Course: Math 574 Gaurav Kulkarni Washington State University Introduction Protein is the fundamental building block of one s body. Many biological processes
Project Report on De novo Peptide Sequencing Course: Math 574 Gaurav Kulkarni Washington State University Introduction Protein is the fundamental building block of one s body. Many biological processes
Advance XCMS Data Processing. H. Paul Benton
 Advance XCMS Data Processing H. Paul Benton Reminder of what we re trying to do Peak Detection Grouping Groups similar Peaks across replicates Retention Time Alignment Statistical Analysis of Classes Total
Advance XCMS Data Processing H. Paul Benton Reminder of what we re trying to do Peak Detection Grouping Groups similar Peaks across replicates Retention Time Alignment Statistical Analysis of Classes Total
cief Data Analysis Chapter Overview Chapter 12:
 page 285 Chapter 12: cief Data Analysis Chapter Overview Analysis Screen Overview Opening Run Files How Run Data is Displayed Viewing Run Data Data Notifications and Warnings Checking Your Results Group
page 285 Chapter 12: cief Data Analysis Chapter Overview Analysis Screen Overview Opening Run Files How Run Data is Displayed Viewing Run Data Data Notifications and Warnings Checking Your Results Group
SUPPLEMENTARY INFORMATION
 Electronic Supplementary Material (ESI) for Analytical Methods. This journal is The Royal Society of Chemistry 2015 1 SUPPLEMENTARY INFORMATION 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23
Electronic Supplementary Material (ESI) for Analytical Methods. This journal is The Royal Society of Chemistry 2015 1 SUPPLEMENTARY INFORMATION 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23
Huqun Liu, Varian Inc., Lake Forest CA
 Rapid development of an LC method for separating high molecular weight degradants from a biopharmaceutical product using an automated Design of Experiments (DOE) approach. Introduction Huqun Liu, Varian
Rapid development of an LC method for separating high molecular weight degradants from a biopharmaceutical product using an automated Design of Experiments (DOE) approach. Introduction Huqun Liu, Varian
Progenesis LC-MS Tutorial Including Data File Import, Alignment, Filtering, Progenesis Stats and Protein ID
 Progenesis LC-MS Tutorial Including Data File Import, Alignment, Filtering, Progenesis Stats and Protein ID 1 Introduction This tutorial takes you through a complete analysis of 9 LC-MS runs (3 replicate
Progenesis LC-MS Tutorial Including Data File Import, Alignment, Filtering, Progenesis Stats and Protein ID 1 Introduction This tutorial takes you through a complete analysis of 9 LC-MS runs (3 replicate
SIVIC GUI Overview. SIVIC GUI Layout Overview
 SIVIC GUI Overview SIVIC GUI Layout Overview At the top of the SIVIC GUI is a row of buttons called the Toolbar. It is a quick interface for loading datasets, controlling how the mouse manipulates the
SIVIC GUI Overview SIVIC GUI Layout Overview At the top of the SIVIC GUI is a row of buttons called the Toolbar. It is a quick interface for loading datasets, controlling how the mouse manipulates the
TraceFinder Shortcut Menus Quick Reference Guide
 TraceFinder Shortcut Menus Quick Reference Guide This quick reference guide describes the right-click shortcut menus available in the Thermo TraceFinder application. Contents Acquisition Mode Analysis
TraceFinder Shortcut Menus Quick Reference Guide This quick reference guide describes the right-click shortcut menus available in the Thermo TraceFinder application. Contents Acquisition Mode Analysis
Tutorial script for whole-cell MALDI-TOF analysis
 Tutorial script for whole-cell MALDI-TOF analysis Julien Textoris June 19, 2013 Contents 1 Required libraries 2 2 Data loading 2 3 Spectrum visualization and pre-processing 4 4 Analysis and comparison
Tutorial script for whole-cell MALDI-TOF analysis Julien Textoris June 19, 2013 Contents 1 Required libraries 2 2 Data loading 2 3 Spectrum visualization and pre-processing 4 4 Analysis and comparison
Agilent MassHunter Workstation Software 7200 Accurate-Mass Quadrupole Time of Flight GC/MS
 Agilent MassHunter Workstation Software 7200 Accurate-Mass Quadrupole Time of Flight GC/MS Familiarization Guide Before you begin 3 Prepare your system 3 Prepare the samples required for data acquisition
Agilent MassHunter Workstation Software 7200 Accurate-Mass Quadrupole Time of Flight GC/MS Familiarization Guide Before you begin 3 Prepare your system 3 Prepare the samples required for data acquisition
Package batman. Installation and Testing
 Package batman Installation and Testing Table of Contents 1. INSTALLATION INSTRUCTIONS... 1 2. TESTING... 3 Test 1: Single spectrum from designed mixture data... 3 Test 2: Multiple spectra from designed
Package batman Installation and Testing Table of Contents 1. INSTALLATION INSTRUCTIONS... 1 2. TESTING... 3 Test 1: Single spectrum from designed mixture data... 3 Test 2: Multiple spectra from designed
Corra v2.0 User s Guide
 Corra v2.0 User s Guide Corra is an open source software Licensed under the Apache License, Version 2.0 and it s source code, demo data and this guide can be downloaded at the http://tools.proteomecenter.org/corra/corra.html.
Corra v2.0 User s Guide Corra is an open source software Licensed under the Apache License, Version 2.0 and it s source code, demo data and this guide can be downloaded at the http://tools.proteomecenter.org/corra/corra.html.
Skyline Targeted Method Refinement
 Skyline Targeted Method Refinement This tutorial will introduce the features available in the Skyline Targeted Proteomics Environment for refining instrument methods for Selected Reaction Monitoring (SRM,
Skyline Targeted Method Refinement This tutorial will introduce the features available in the Skyline Targeted Proteomics Environment for refining instrument methods for Selected Reaction Monitoring (SRM,
An R package to process LC/MS metabolomic data: MAIT (Metabolite Automatic Identification Toolkit)
 An R package to process LC/MS metabolomic data: MAIT (Metabolite Automatic Identification Toolkit) Francesc Fernández-Albert, Rafael Llorach, Cristina Andrés-Lacueva, Alexandre Perera June 13, 2018 1 Abstract
An R package to process LC/MS metabolomic data: MAIT (Metabolite Automatic Identification Toolkit) Francesc Fernández-Albert, Rafael Llorach, Cristina Andrés-Lacueva, Alexandre Perera June 13, 2018 1 Abstract
This manual describes step-by-step instructions to perform basic operations for data analysis.
 HDXanalyzer User Manual The program HDXanalyzer is available for the analysis of the deuterium exchange mass spectrometry data obtained on high-resolution mass spectrometers. Currently, the program is
HDXanalyzer User Manual The program HDXanalyzer is available for the analysis of the deuterium exchange mass spectrometry data obtained on high-resolution mass spectrometers. Currently, the program is
Package ALS. August 3, 2015
 Type Package Package ALS August 3, 2015 Title Multivariate Curve Resolution Alternating Least Squares (MCR-ALS) Version 0.0.6 Author Katharine M. Mullen Maintainer Katharine Mullen
Type Package Package ALS August 3, 2015 Title Multivariate Curve Resolution Alternating Least Squares (MCR-ALS) Version 0.0.6 Author Katharine M. Mullen Maintainer Katharine Mullen
SpectroDive 8 - Coelacanth. User Manual
 SpectroDive 8 - Coelacanth User Manual Table of Contents 1 System Requirements... 4 2 General Information... 4 2.1 Supported Instruments... 4 3 Getting Started... 5 3.1 Getting SpectroDive... 5 3.2 SpectroDive
SpectroDive 8 - Coelacanth User Manual Table of Contents 1 System Requirements... 4 2 General Information... 4 2.1 Supported Instruments... 4 3 Getting Started... 5 3.1 Getting SpectroDive... 5 3.2 SpectroDive
OpenLynx User's Guide
 OpenLynx User s Guide OpenLynx User's Guide Version 4.0 Waters part No 715000405 Micromass Part No - 6666670 5 February, 2002 i OpenLynx User s Guide OpenLynx User s Guide The software described in this
OpenLynx User s Guide OpenLynx User's Guide Version 4.0 Waters part No 715000405 Micromass Part No - 6666670 5 February, 2002 i OpenLynx User s Guide OpenLynx User s Guide The software described in this
[ WHITE PAPER ] System Policies and Privileges for Processing Data in Empower Software with Data Integrity in Mind INTRODUCTION
![[ WHITE PAPER ] System Policies and Privileges for Processing Data in Empower Software with Data Integrity in Mind INTRODUCTION [ WHITE PAPER ] System Policies and Privileges for Processing Data in Empower Software with Data Integrity in Mind INTRODUCTION](/thumbs/87/96630216.jpg) System Policies and Privileges for Processing Data in Empower Software with Data Integrity in Mind Explore how Empower Chromatography Data Software System Policies and privileges can be used to control
System Policies and Privileges for Processing Data in Empower Software with Data Integrity in Mind Explore how Empower Chromatography Data Software System Policies and privileges can be used to control
User Manual STATISTICS
 Version 6 User Manual STATISTICS 2006 BRUKER OPTIK GmbH, Rudolf-Plank-Straße 27, D-76275 Ettlingen, www.brukeroptics.com All rights reserved. No part of this manual may be reproduced or transmitted in
Version 6 User Manual STATISTICS 2006 BRUKER OPTIK GmbH, Rudolf-Plank-Straße 27, D-76275 Ettlingen, www.brukeroptics.com All rights reserved. No part of this manual may be reproduced or transmitted in
What s New in Empower 3
 What s New in Empower 3 Revision A Copyright Waters Corporation 2010 All rights reserved Copyright notice 2010 WATERS CORPORATION. PRINTED IN THE UNITED STATES OF AMERICA AND IN IRELAND. ALL RIGHTS RESERVED.
What s New in Empower 3 Revision A Copyright Waters Corporation 2010 All rights reserved Copyright notice 2010 WATERS CORPORATION. PRINTED IN THE UNITED STATES OF AMERICA AND IN IRELAND. ALL RIGHTS RESERVED.
Multivariate Calibration Quick Guide
 Last Updated: 06.06.2007 Table Of Contents 1. HOW TO CREATE CALIBRATION MODELS...1 1.1. Introduction into Multivariate Calibration Modelling... 1 1.1.1. Preparing Data... 1 1.2. Step 1: Calibration Wizard
Last Updated: 06.06.2007 Table Of Contents 1. HOW TO CREATE CALIBRATION MODELS...1 1.1. Introduction into Multivariate Calibration Modelling... 1 1.1.1. Preparing Data... 1 1.2. Step 1: Calibration Wizard
Exploring Data. This guide describes the facilities in SPM to gain initial insights about a dataset by viewing and generating descriptive statistics.
 This guide describes the facilities in SPM to gain initial insights about a dataset by viewing and generating descriptive statistics. 2018 by Minitab Inc. All rights reserved. Minitab, SPM, SPM Salford
This guide describes the facilities in SPM to gain initial insights about a dataset by viewing and generating descriptive statistics. 2018 by Minitab Inc. All rights reserved. Minitab, SPM, SPM Salford
Fatty Acid Methyl Ester (FAME) RTL Databases for GC and GC/MS
 Fatty Acid Methyl Ester (FAME) RTL Databases for GC and GC/MS User contributed by: Frank David and Pat Sandra Research Institute for Chromatography Pres. Kennedypark 20, B-8500 Kortrijk, Belgium and by:
Fatty Acid Methyl Ester (FAME) RTL Databases for GC and GC/MS User contributed by: Frank David and Pat Sandra Research Institute for Chromatography Pres. Kennedypark 20, B-8500 Kortrijk, Belgium and by:
Automated Bioinformatics Analysis System on Chip ABASOC. version 1.1
 Automated Bioinformatics Analysis System on Chip ABASOC version 1.1 Phillip Winston Miller, Priyam Patel, Daniel L. Johnson, PhD. University of Tennessee Health Science Center Office of Research Molecular
Automated Bioinformatics Analysis System on Chip ABASOC version 1.1 Phillip Winston Miller, Priyam Patel, Daniel L. Johnson, PhD. University of Tennessee Health Science Center Office of Research Molecular
PSS718 - Data Mining
 Lecture 5 - Hacettepe University October 23, 2016 Data Issues Improving the performance of a model To improve the performance of a model, we mostly improve the data Source additional data Clean up the
Lecture 5 - Hacettepe University October 23, 2016 Data Issues Improving the performance of a model To improve the performance of a model, we mostly improve the data Source additional data Clean up the
For additional information, see these documents on the Breeze 2 software DVD:
 R E L E A S E N O T E S Waters Breeze 2 Read these release notes before you install and operate Breeze 2. They contain information necessary to ensure successful liquid chromatography results, including:
R E L E A S E N O T E S Waters Breeze 2 Read these release notes before you install and operate Breeze 2. They contain information necessary to ensure successful liquid chromatography results, including:
Statistical Analysis of Metabolomics Data. Xiuxia Du Department of Bioinformatics & Genomics University of North Carolina at Charlotte
 Statistical Analysis of Metabolomics Data Xiuxia Du Department of Bioinformatics & Genomics University of North Carolina at Charlotte Outline Introduction Data pre-treatment 1. Normalization 2. Centering,
Statistical Analysis of Metabolomics Data Xiuxia Du Department of Bioinformatics & Genomics University of North Carolina at Charlotte Outline Introduction Data pre-treatment 1. Normalization 2. Centering,
Using SIMDIS Expert 9 (Separation Systems) with Clarity
 Using SIMDIS Expert 9 (Separation Systems) with Clarity The SIMDIS Expert 9 may be used with Clarity or Clarity Lite software to process the acquired chromatograms according to Simulated Distillation.
Using SIMDIS Expert 9 (Separation Systems) with Clarity The SIMDIS Expert 9 may be used with Clarity or Clarity Lite software to process the acquired chromatograms according to Simulated Distillation.
BIOINF 4399B Computational Proteomics and Metabolomics
 BIOINF 4399B Computational Proteomics and Metabolomics Sven Nahnsen WS 13/14 6. Quantification Part II Overview Label-free quantification Definition of features Feature finding on centroided data Absolute
BIOINF 4399B Computational Proteomics and Metabolomics Sven Nahnsen WS 13/14 6. Quantification Part II Overview Label-free quantification Definition of features Feature finding on centroided data Absolute
Package biosigner. March 6, 2019
 Type Package Title Signature discovery from omics data Version 1.10.0 Date 2018-04-15 Package biosigner March 6, 2019 Author Philippe Rinaudo , Etienne Thevenot
Type Package Title Signature discovery from omics data Version 1.10.0 Date 2018-04-15 Package biosigner March 6, 2019 Author Philippe Rinaudo , Etienne Thevenot
Importing data in a database with levels
 BioNumerics Tutorial: Importing data in a database with levels 1 Aim In this tutorial you will learn how to import data in a BioNumerics database with levels and how to replicate and summarize level-specific
BioNumerics Tutorial: Importing data in a database with levels 1 Aim In this tutorial you will learn how to import data in a BioNumerics database with levels and how to replicate and summarize level-specific
PDA EXTENSION. Clarity Extension. Code/Rev.: M054/80A Date: 6/8/2018. Fax: Petrzilkova 2583/ Prague 5
 PDA EXTENSION Clarity Extension ENG Code/Rev.: M054/80A Date: 6/8/2018 Phone: +420 251 013 400 DataApex Ltd. Fax: +420 251 013 401 Petrzilkova 2583/13 clarity@dataapex.com 158 00 Prague 5 www.dataapex.com
PDA EXTENSION Clarity Extension ENG Code/Rev.: M054/80A Date: 6/8/2018 Phone: +420 251 013 400 DataApex Ltd. Fax: +420 251 013 401 Petrzilkova 2583/13 clarity@dataapex.com 158 00 Prague 5 www.dataapex.com
Introduction to Time-of-Flight Mass
 Introduction to Time-of-Flight Mass Spectrometry and Deconvolution Nicholas Hall Speaker Mark Libardoni, Pete Stevens, Joe Binkley Life Science & Chemical Analysis Centre St. Joseph, Michigan, USA e-seminar
Introduction to Time-of-Flight Mass Spectrometry and Deconvolution Nicholas Hall Speaker Mark Libardoni, Pete Stevens, Joe Binkley Life Science & Chemical Analysis Centre St. Joseph, Michigan, USA e-seminar
LC-MS Data Pre-Processing. Xiuxia Du, Ph.D. Department of Bioinformatics and Genomics University of North Carolina at Charlotte
 LC-MS Data Pre-Processing Xiuxia Du, Ph.D. Department of Bioinformatics and Genomics University of North Carolina at Charlotte Outline Raw LC-MS data - Profile and centroid data - Mass vs. retention time
LC-MS Data Pre-Processing Xiuxia Du, Ph.D. Department of Bioinformatics and Genomics University of North Carolina at Charlotte Outline Raw LC-MS data - Profile and centroid data - Mass vs. retention time
Rat 2D EPSI Dual Band Variable Flip Angle 13 C Dynamic Spectroscopy
 Rat 2D EPSI Dual Band Variable Flip Angle 13 C Dynamic Spectroscopy In this example you will load a dynamic MRS animal data set acquired on a GE 3T scanner. This data was acquired with an EPSI sequence
Rat 2D EPSI Dual Band Variable Flip Angle 13 C Dynamic Spectroscopy In this example you will load a dynamic MRS animal data set acquired on a GE 3T scanner. This data was acquired with an EPSI sequence
Agilent G2721AA Spectrum Mill MS Proteomics Workbench Quick Start Guide
 Agilent G2721AA Spectrum Mill MS Proteomics Workbench Quick Start Guide A guide to the Spectrum Mill workbench Use this reference for your first steps with the Spectrum Mill workbench. What is the Spectrum
Agilent G2721AA Spectrum Mill MS Proteomics Workbench Quick Start Guide A guide to the Spectrum Mill workbench Use this reference for your first steps with the Spectrum Mill workbench. What is the Spectrum
Direct Infusion Mass Spectrometry Processing (DIMaSP) Instructions for use
 Direct Infusion Mass Spectrometry Processing (DIMaSP) Instructions for use The following instructions assume the user has a batch of paired sample and blank spectra that have been exported to a comma-separated
Direct Infusion Mass Spectrometry Processing (DIMaSP) Instructions for use The following instructions assume the user has a batch of paired sample and blank spectra that have been exported to a comma-separated
Galaxie Report Editor
 Varian, Inc. 2700 Mitchell Drive Walnut Creek, CA 94598-1675/USA Galaxie Report Editor User s Guide Varian, Inc. 2008 Printed in U.S.A. 03-914949-00: Rev 6 Galaxie Report Editor i Table of Contents Introduction...
Varian, Inc. 2700 Mitchell Drive Walnut Creek, CA 94598-1675/USA Galaxie Report Editor User s Guide Varian, Inc. 2008 Printed in U.S.A. 03-914949-00: Rev 6 Galaxie Report Editor i Table of Contents Introduction...
Impact Analysis for Software changes in OpenLAB CDS A.01.04
 Impact Analysis for Software changes in OpenLAB CDS A.01.04 Document Information: Filename Product Identifier Product Revision Project Identifier Document Revision Impact_Analysis_OpenLAB_CDS_A0104 OpenLAB
Impact Analysis for Software changes in OpenLAB CDS A.01.04 Document Information: Filename Product Identifier Product Revision Project Identifier Document Revision Impact_Analysis_OpenLAB_CDS_A0104 OpenLAB
Chromeleon / MSQ Plus Operator s Guide Document No Revision 03 October 2009
 MSQ Plus Chromeleon / MSQ Plus Operator s Guide Document No. 065322 Revision 03 October 2009 Chromeleon / MSQ Plus Operator s Guide 2009 by Dionex Corporation All rights reserved worldwide. Printed in
MSQ Plus Chromeleon / MSQ Plus Operator s Guide Document No. 065322 Revision 03 October 2009 Chromeleon / MSQ Plus Operator s Guide 2009 by Dionex Corporation All rights reserved worldwide. Printed in
Package GCalignR. January 16, 2018
 Package GCalignR January 16, 2018 Title Simple Peak Alignment for Gas-Chromatography Data Version 1.0.1 Date 2018-01-16 Aligns peak based on peak retention times and matches homologous peaks across samples.
Package GCalignR January 16, 2018 Title Simple Peak Alignment for Gas-Chromatography Data Version 1.0.1 Date 2018-01-16 Aligns peak based on peak retention times and matches homologous peaks across samples.
Data Handling and Reports
 Varian Analytical Instruments 2700 Mitchell Drive Walnut Creek, CA 94598-1675/USA Star Chromatography Workstation Version 6 Data Handling and Reports Tutorials Varian, Inc. 2002 03-914736-00:Rev. 4 Trademark
Varian Analytical Instruments 2700 Mitchell Drive Walnut Creek, CA 94598-1675/USA Star Chromatography Workstation Version 6 Data Handling and Reports Tutorials Varian, Inc. 2002 03-914736-00:Rev. 4 Trademark
R-software multims-toolbox. (User Guide)
 R-software multims-toolbox (User Guide) Pavel Cejnar 1, Štěpánka Kučková 2 1 Department of Computing and Control Engineering, Institute of Chemical Technology, Technická 3, 166 28 Prague 6, Czech Republic,
R-software multims-toolbox (User Guide) Pavel Cejnar 1, Štěpánka Kučková 2 1 Department of Computing and Control Engineering, Institute of Chemical Technology, Technická 3, 166 28 Prague 6, Czech Republic,
NIST MS AUTOIMP feature looks for and finds secondary locator file.
 INTRODUCTION This document describes how to start the NIST MS (Mass Spectral) Search Program Version 2.0 and higher or the AMDIS (Automated Mass spectral Deconvolution and Identification) program from
INTRODUCTION This document describes how to start the NIST MS (Mass Spectral) Search Program Version 2.0 and higher or the AMDIS (Automated Mass spectral Deconvolution and Identification) program from
PROTEOMIC COMMAND LINE SOLUTION. Linux User Guide December, B i. Bioinformatics Solutions Inc.
 >_ PROTEOMIC COMMAND LINE SOLUTION Linux User Guide December, 2015 B i Bioinformatics Solutions Inc. www.bioinfor.com 1. Introduction Liquid chromatography-tandem mass spectrometry (LC-MS/MS) based proteomics
>_ PROTEOMIC COMMAND LINE SOLUTION Linux User Guide December, 2015 B i Bioinformatics Solutions Inc. www.bioinfor.com 1. Introduction Liquid chromatography-tandem mass spectrometry (LC-MS/MS) based proteomics
Agilent Triple Quadrupole LC/MS Peptide Quantitation with Skyline
 Agilent Triple Quadrupole LC/MS Peptide Quantitation with Skyline Workflow Guide A Peptide Optimization B Review Results in Skyline Create QQQ method in Skyline Edit Skyline settings Import peptides into
Agilent Triple Quadrupole LC/MS Peptide Quantitation with Skyline Workflow Guide A Peptide Optimization B Review Results in Skyline Create QQQ method in Skyline Edit Skyline settings Import peptides into
UPDATING PESTICIDE RETENTION TIME LIBRARIES FOR THE AGILENT INTUVO 9000 GC
 UPDATING PESTICIDE RETENTION TIME LIBRARIES FOR THE AGILENT INTUVO 9000 GC How to Update Retention Time Libraries on Intuvo Introduction When applying methods to a new gas chromatographic (GC) system,
UPDATING PESTICIDE RETENTION TIME LIBRARIES FOR THE AGILENT INTUVO 9000 GC How to Update Retention Time Libraries on Intuvo Introduction When applying methods to a new gas chromatographic (GC) system,
To get started download the dataset, unzip files, start Maven, and follow steps below.
 Getting Started. This document provides basic overview of Maven functionality. For the purpose of demonstration we will use an example CMV Viral Infection time course dataset. (Dataset Download). There
Getting Started. This document provides basic overview of Maven functionality. For the purpose of demonstration we will use an example CMV Viral Infection time course dataset. (Dataset Download). There
Agilent G2727AA LC/MS Data Browser Quick Start
 What is Data Browser? Agilent G2727AA LC/MS Data Browser Quick Start A guide to get started with Data Browser Use this quick reference guide as a road map for your first steps with the Data Browser software,
What is Data Browser? Agilent G2727AA LC/MS Data Browser Quick Start A guide to get started with Data Browser Use this quick reference guide as a road map for your first steps with the Data Browser software,
CHROMuLAN. Basic instruction
 PiKRON s.r.o Apr 25 2002 CHROMuLAN 1. Introduction This freeware is designed for the control system of various sets of apparatuses and subsequent assessment of results. The project is initialised and sponsored
PiKRON s.r.o Apr 25 2002 CHROMuLAN 1. Introduction This freeware is designed for the control system of various sets of apparatuses and subsequent assessment of results. The project is initialised and sponsored
Real-time Data Compression for Mass Spectrometry Jose de Corral 2015 GPU Technology Conference
 Real-time Data Compression for Mass Spectrometry Jose de Corral 2015 GPU Technology Conference 2015 Waters Corporation 1 Introduction to LC/IMS/MS LC/IMS/MS is the combination of three analytical techniques
Real-time Data Compression for Mass Spectrometry Jose de Corral 2015 GPU Technology Conference 2015 Waters Corporation 1 Introduction to LC/IMS/MS LC/IMS/MS is the combination of three analytical techniques
