xmapbridge: Graphically Displaying Numeric Data in the X:Map Genome Browser
|
|
|
- Lindsey Leonard
- 5 years ago
- Views:
Transcription
1 xmapbridge: Graphically Displaying Numeric Data in the X:Map Genome Browser Tim Yates, Crispin J Miller April 30, 2018 Contents 1 Introduction 1 2 Quickstart 2 3 Plotting Graphs, the simple way 4 4 Multiple plots on a graph 6 5 More fine grained control 7 6 Specifying Graph Colours 8 7 Installation and configuration details Installation of the XMapBridge software Configuration of the xmapbridge package Introduction xmapbridge is a package that allows numeric data to be displayed in the web based genome browser, X:Map. X:Map uses the Google Maps API to provide a real-time scrollable view of the genome. It supports a number of species, and can be accessed at To do this, it communicates with X:Map via a small Java application called XMap- Bridge. XMapBridge sits on your local machine, interprets the graph data generated by xmapbridge and uses it to create plots, which are then passed to the genome browser to be displayed via HTTP. Importantly, all graph rendering is performed on the local machine so none of the underlying graph data is uploaded to the X:Map server to generate your graphs. Graph plotting in R is done using calls to the functions xmap.plot and xmap.points, which have parameters that aim to be similar to those used by the standard plot methods in R. These result in data being written to a set of files (in a specific directory 1
2 Figure 1: The XMapBridge in operation structure) that contain the data to be displayed, as well as some additional meta-data describing each of the graphs. When the XMapBridge application is running (figure 1), it monitors the same directory structure used by xmapbridge and uses the information written out from the Bioconductor session to generate its plots. These are then passed via HTTP to the genome browser for display. Note that the XMapBridge must be running on the same machine as your webbrowser. Only one instance of the XMapBridge can be run on a machine at any one time, and the X:Map webpage will try to connect to localhost to load javascript and graph data. If you do not see the Connect button when XMapBridge is running, the most likely reason is this. 2 Quickstart The steps required to get the xmapbridge and XMapBridge programs working are as follows: 1. Install Java 5+ (if it s not already installed). 2. Install XMapBridge (see the section Installation of the XMapBridge software at the end of this Vignette for more details). 3. Install xmapbridge (this package). By default, the XMapBridge java application and the xmapbridge R package use a folder called.xmb cache in your home directory. 2
3 If this folder does not exist when the R functions are first called (or the environment variable isn t set see below), then xmapbridge will offer to create the folder for you. If this folder does not exist when the XMapBridge is first run, (and there is no environment variable set), then the application will run in demo mode, showing three demo graphs for you to see how it works on your system (figure 2). Figure 2: The process to show the graphs in the browser (1a) Click on the connect button (1b) This changes into a drop-down select box (1c) select the graph of choice If you don t want to use this default directory, then you need to create an environment variable XMAP BRIDGE CACHE which contains a path to the folder you want to use as a cache folder instead. The details of how to set this variable on multiple operating systems can be found in the README.TXT file which was included in the XMapBridge download. The function xmap.plot generates a project folder structure, adds a graph to this structure, and adds the plot data in the appropriate directory for XMapBridge to find it. So, for example: > library( xmapbridge ) > set.seed( 100 ) > x <- runif( 100, , ) > y <- 10 * sin( 2 * pi * ( x ) / ) > xmap.plot( x, y, species="homo_sapiens", chr="1", type="line" ) Will first prompt you to ask if you want to create the default directory (if it s not already there) and then generate a line plot between position 1,100,000bp, and 1,200,000bp 3
4 Figure 3: X:Map displaying a simple graph on human chromosome 1. As you can see from the output, it actually starts at location 1,103,015bp due to the runif. (In the following section more complex examples show how to generate graphs for real (Affymetrix exon) array data). If you re-start XMapBridge (assuming it was in demo-mode before), it should now find this directory and the browser (once refreshed or pointed at the X:Map website) should offer you your new graph via the drop-down menu. When you then select it, you should see it displayed, as in figure 3. The next time you use the package, this directory will already exist, so you won t get prompted again, unless you delete or move the cache directory. The XMapBridge java application scans your cache folder every few seconds to see if anything has been updated, however these changes can take some time to bubble up to the javascript in the browser. There should be no need to refresh the X:Map page and reload the bridge connection applet, but it may take up to 30 seconds for the list of graphs to be updated. When the list is updated, the drop-down selection box reverts back to its initial position, removing any currently viewed graph. 3 Plotting Graphs, the simple way As we saw above, the simplest way to plot a graph is simply to call xmap.plot. As well as x,y coordinates, you also need to specify the chromosome and species. Labels can be provided and the type of plot can also be specified. Later we ll see how to set transparency and colours for the plots. Here we explore how to generate some plots for Affymetrix Exon arrays. We first load an example dataset (see?exon.data for more details on the origins of this dataset): > data(xmapbridge) The dataframe exon.data contains information on 1747 probesets targeting 80 different genes, for a triplicate comparison between two cell lines, mcf7 and mcf10a. To generate our plots, we first take our array data and split it into different pieces, one for each gene: 4
5 > l <- split(exon.data,exon.data$"gene") Then, for each gene we need to extract the appropriate x locations (and which chromosome it is on). Initially, we just do it for the first gene in the list: > g <- l[[1]] > x <- g$"seq_region_end" + (g$"seq_region_end" - g$"seq_region_start")/2 > y <- g[,2] > chr <- unique(g$"name") > Now we can plot the data for one of our replicates: > xmap.plot(x,y,chr,species="homo_sapiens",type="area",col="#ff000066") Currently, scatter, line, bar, step, area and steparea plots are supported. These are be pretty self explanatory - the following should generate plots similar to those in figure 4: Figure 4: Different styles of plot: (a) scatter, (b) line, (c) bar, (d) area and (e) steparea > xmap.plot(x,y,chr,species="homo_sapiens",type="scatter",col="#ff000066") 5
6 > xmap.plot(x,y,chr,species="homo_sapiens",type="line",col="#ff000066") > xmap.plot(x,y,chr,species="homo_sapiens",type="bar",col="#ff000066") > xmap.plot(x,y,chr,species="homo_sapiens",type="area",col="#ff000066") > xmap.plot(x,y,chr,species="homo_sapiens",type="steparea",col="#ff000066") 4 Multiple plots on a graph XMapBridge can generate a graph with multiple plots on it. This is done by calling xmap.points after the initial xmap.plot call. Each call to xmap.points will add a new Plot to the current Graph: > #Pick an interesting gene > g <- l[["ensg "]] > x <- g$"seq_region_end" + (g$"seq_region_end" - g$"seq_region_start")/2 > chr <- unique(g$"name") > y <- g[,2] > xmap.plot(x,y,chr,species="homo_sapiens",type="area",col="#ff000033",ylim=c(0,16)) > xmap.points(x,y,chr,type="line",col="#ff0000ff") > #We were plotting the 1st array (2nd column in g), now plot the 4th (5th column in g > y <- g[,5] > xmap.points(x,y,chr,type="area",col="#0000ff33") > xmap.points(x,y,chr,type="line",col="#0000ffff") This should look like figure 6. (For each array we first generate the shaded area, and then add the edge as an additional line plot). Note: As you can see from the output, these calls to xmap.points returns a new Plot object that resides within the same Graph and Project at the plot returned from the initial xmap.plot call. 6
7 Figure 5: Multiple plots on the same graph 5 More fine grained control A quick look at the XMapBridge GUI will show that all of the graphs and plots generated so far have been placed in a hierarchy, beginning with a Project and with a series of graphs and plots underneath that. This is fine if we are happy with all of the plots generated during a single session to be grouped into a single project in this way, but we can express more fine grained control if required. In the next example, we will create a new project and then iterate along our list of genes, l, generating a new graph for each gene. Each of these graphs will be placed in the new project we ve created. First, we set up some colours and names for our individual plots: > cols.bg <- c(rep("#ff000011",3),rep("#0000ff11",3)) > cols.fg <- c(rep("#ff0000ff",3),rep("#0000ffff",3)) > names <- c("7.1","7.2","7.3","10a.1","10a.2","10a.3") (We will explain how colours are defined later on). Then we create a new project and store the resultant id in the variable pid: > pid <- xmap.project.new("mcf7 vs mcf10a") Next, we write a function that can plot a single gene. It takes a matrix of data (i.e. one of the elements of l) and the project id: > plot.a.gene <- function(g,pid) { + x <- g$"seq_region_end" + (g$"seq_region_end" - g$"seq_region_start")/2 + y <- g[,2:7] + chr <- unique(g$"name") + gene <- unique(g$"gene") + xlim <- range(x) + gph <- xmap.graph.new(projectid=pid, name=gene,desc="vignette example", min=0, ma + for(i in 1:6) { + xmap.points(graphid=gph,x=x,y=y[,i],type="area",col=cols.bg[i],xlab="") + xmap.points(graphid=gph,x=x,y=y[,i],type="line",col=cols.fg[i],xlab=names[i]) + } + } 7
8 Note how the function xmap.graph.new is used to create a new graph. It takes a projectid as a parameter, which specifies which project the graph should be placed in, and returns a graphid. This is used by xmap.points to specify which graph to add the plots to. Finally, we can generate a graph for each of our genes using lapply across l: > lapply(l,plot.a.gene,pid=pid) 6 Specifying Graph Colours By default, the XMapBridge allocates unique colours to each plot in the graph, however, note how in the previous example, the colour of the plot is being set as an RGB value with an extra alpha value specifying the opacity. The format for these colours is #RRGGBBAA with each of the Red, Green, Blue and Alpha components being a hexadecimal number from 00 to FF. So, for example, a colour value of # for the first plot gives a (slightly green) blue colour with the opacity set to 33% so that the underlying images from the X:Map website (and the underlying plots on this same graph) can be seen through the current plot (see figure 6). > library( RColorBrewer ) > #A light grey almost transparent step area > xmap.plot( x, y, type="steparea", col="# ", ylab="intensity", + chr="4", species="homo_sapiens" ) > #With an opaque orange edge > xmap.points( x, y, type="step", col="#f7941dff" ) The function xmap.col then provides an easy way to set alpha values: > #Five levels of red, chosen by the RColorBrewer > reds <- brewer.pal( 5, "Reds" ) > reds <- xmap.col( reds, alpha=0x55 ) > for( i in 1:5 ) { + xmap.points( x, y / i, type="area", col=reds[i] ) + } Note that xmap.plot also allows colours to specified as an integer. In this case, the alpha value goes first; the format is 0xAARRGGBB. xmap.col, returns integers in this form. 8
9 Figure 6: Specifying colours and alpha values 7 Installation and configuration details 7.1 Installation of the XMapBridge software The XMapBridge Java application can be downloaded from the X:Map website at http: //xmap.picr.man.ac.uk/download/bridge as a simple ZIP file. When extracted, there are comprehensive installation instructions to be found in the file README.TXT. 7.2 Configuration of the xmapbridge package xmapbridge and XMapBridge can be told to look in another location apart from the default for the directory they share. This is specified by an environment variable with the name XMAP BRIDGE CACHE, which points to a folder you wish to use. Again, comprehensive instructions for doing this on the main operationg systems can be found in the README.TXT file that is part of the XMapBridge download. 9
Mascot Insight is a new application designed to help you to organise and manage your Mascot search and quantitation results. Mascot Insight provides
 1 Mascot Insight is a new application designed to help you to organise and manage your Mascot search and quantitation results. Mascot Insight provides ways to flexibly merge your Mascot search and quantitation
1 Mascot Insight is a new application designed to help you to organise and manage your Mascot search and quantitation results. Mascot Insight provides ways to flexibly merge your Mascot search and quantitation
Ensembl RNASeq Practical. Overview
 Ensembl RNASeq Practical The aim of this practical session is to use BWA to align 2 lanes of Zebrafish paired end Illumina RNASeq reads to chromosome 12 of the zebrafish ZV9 assembly. We have restricted
Ensembl RNASeq Practical The aim of this practical session is to use BWA to align 2 lanes of Zebrafish paired end Illumina RNASeq reads to chromosome 12 of the zebrafish ZV9 assembly. We have restricted
MiTV User Manual Revision 2 July 8, 2015 Prepared by Walter B. Schoustal MicroVideo Learning Systems, Inc.
 MiTV User Manual Revision 2 July 8, 2015 Prepared by Walter B. Schoustal MicroVideo Learning Systems, Inc. http://www.microvideo.com 1 The MiTV Video Scheduling System allows you to schedule and stream
MiTV User Manual Revision 2 July 8, 2015 Prepared by Walter B. Schoustal MicroVideo Learning Systems, Inc. http://www.microvideo.com 1 The MiTV Video Scheduling System allows you to schedule and stream
Cascading style sheets
 Cascading style sheets The best way to create websites is to keep the content separate from the presentation. The best way to create websites is to keep the content separate from the presentation. HTML
Cascading style sheets The best way to create websites is to keep the content separate from the presentation. The best way to create websites is to keep the content separate from the presentation. HTML
Advanced Multivariate Continuous Displays and Diagnostics
 Advanced Multivariate Continuous Displays and Diagnostics 37 This activity explores advanced multivariate plotting. With the advanced diagnostic functions, output graphs are more readable and useful for
Advanced Multivariate Continuous Displays and Diagnostics 37 This activity explores advanced multivariate plotting. With the advanced diagnostic functions, output graphs are more readable and useful for
Akana API Platform: Upgrade Guide
 Akana API Platform: Upgrade Guide Version 8.0 to 8.2 Akana API Platform Upgrade Guide Version 8.0 to 8.2 November, 2016 (update v2) Copyright Copyright 2016 Akana, Inc. All rights reserved. Trademarks
Akana API Platform: Upgrade Guide Version 8.0 to 8.2 Akana API Platform Upgrade Guide Version 8.0 to 8.2 November, 2016 (update v2) Copyright Copyright 2016 Akana, Inc. All rights reserved. Trademarks
Enhancing your Page. Text Effects. Paragraph Effects. Headings
 Enhancing your Page You can make your pages more visually appealing and organize page content by using text effects, paragraph effects, macros, images, tables, etc. To begin, select the "Edit" button for
Enhancing your Page You can make your pages more visually appealing and organize page content by using text effects, paragraph effects, macros, images, tables, etc. To begin, select the "Edit" button for
CS7026 CSS3. CSS3 Graphics Effects
 CS7026 CSS3 CSS3 Graphics Effects What You ll Learn We ll create the appearance of speech bubbles without using any images, just these pieces of pure CSS: The word-wrap property to contain overflowing
CS7026 CSS3 CSS3 Graphics Effects What You ll Learn We ll create the appearance of speech bubbles without using any images, just these pieces of pure CSS: The word-wrap property to contain overflowing
Gale Digital Scholar Lab Getting Started Walkthrough Guide
 Getting Started Logging In Your library or institution will provide you with your login link. You will have the option to sign in with a Google or Microsoft Account, this is so you have a personal account
Getting Started Logging In Your library or institution will provide you with your login link. You will have the option to sign in with a Google or Microsoft Account, this is so you have a personal account
A short Introduction to UCSC Genome Browser
 A short Introduction to UCSC Genome Browser Elodie Girard, Nicolas Servant Institut Curie/INSERM U900 Bioinformatics, Biostatistics, Epidemiology and computational Systems Biology of Cancer 1 Why using
A short Introduction to UCSC Genome Browser Elodie Girard, Nicolas Servant Institut Curie/INSERM U900 Bioinformatics, Biostatistics, Epidemiology and computational Systems Biology of Cancer 1 Why using
ENGL 323: Writing for New Media Repurposing Content for the Web Part Two
 ENGL 323: Writing for New Media Repurposing Content for the Web Part Two Dr. Michael Little michaellittle@kings.edu Hafey-Marian 418 x5917 Using Color to Establish Visual Hierarchies Color is useful in
ENGL 323: Writing for New Media Repurposing Content for the Web Part Two Dr. Michael Little michaellittle@kings.edu Hafey-Marian 418 x5917 Using Color to Establish Visual Hierarchies Color is useful in
The building block of a CSS stylesheet. A rule consists of a selector and a declaration block (one or more declarations).
 WDI Fundamentals Unit 4 CSS Cheat Sheet Rule The building block of a CSS stylesheet. A rule consists of a selector and a declaration block (one or more declarations). Declaration A declaration is made
WDI Fundamentals Unit 4 CSS Cheat Sheet Rule The building block of a CSS stylesheet. A rule consists of a selector and a declaration block (one or more declarations). Declaration A declaration is made
Secure Web Appliance. Basic Usage Guide
 Secure Web Appliance Basic Usage Guide Table of Contents 1. Introduction... 1 1.1. About CYAN Secure Web Appliance... 1 1.2. About this Manual... 1 1.2.1. Document Conventions... 1 2. Description of the
Secure Web Appliance Basic Usage Guide Table of Contents 1. Introduction... 1 1.1. About CYAN Secure Web Appliance... 1 1.2. About this Manual... 1 1.2.1. Document Conventions... 1 2. Description of the
RENDERING TECHNIQUES
 RENDERING TECHNIQUES Colors in Flash In Flash, colors are specified as numbers. A color number can be anything from 0 to 16,777,215 for 24- bit color which is 256 * 256 * 256. Flash uses RGB color, meaning
RENDERING TECHNIQUES Colors in Flash In Flash, colors are specified as numbers. A color number can be anything from 0 to 16,777,215 for 24- bit color which is 256 * 256 * 256. Flash uses RGB color, meaning
Lab 1: Silver Dollar Game 1 CSCI 2101B Fall 2018
 Lab 1: Silver Dollar Game 1 CSCI 2101B Fall 2018 Due: Tuesday, September 18, 11:59 pm Collaboration Policy: Level 1 (review full policy for details) Group Policy: Individual This lab will give you experience
Lab 1: Silver Dollar Game 1 CSCI 2101B Fall 2018 Due: Tuesday, September 18, 11:59 pm Collaboration Policy: Level 1 (review full policy for details) Group Policy: Individual This lab will give you experience
RnBeadsDJ A Quickstart Guide to the RnBeads Data Juggler
 RnBeadsDJ A Quickstart Guide to the RnBeads Data Juggler Fabian Müller, Yassen Assenov, Pavlo Lutsik Contact: rnbeads@mpi-inf.mpg.de Package version: 1.12.2 September 25, 2018 RnBeads is an R package for
RnBeadsDJ A Quickstart Guide to the RnBeads Data Juggler Fabian Müller, Yassen Assenov, Pavlo Lutsik Contact: rnbeads@mpi-inf.mpg.de Package version: 1.12.2 September 25, 2018 RnBeads is an R package for
owncloud Android App Manual
 owncloud Android App Manual Release 2.7.0 The owncloud developers October 30, 2018 CONTENTS 1 Release Notes 1 1.1 Changes in 2.7.0............................................. 1 1.2 Changes in 2.6.0.............................................
owncloud Android App Manual Release 2.7.0 The owncloud developers October 30, 2018 CONTENTS 1 Release Notes 1 1.1 Changes in 2.7.0............................................. 1 1.2 Changes in 2.6.0.............................................
Imagery International website manual
 Imagery International website manual Prepared for: Imagery International Prepared by: Jenn de la Fuente Rosebud Designs http://www.jrosebud.com/designs designs@jrosebud.com 916.538.2133 A brief introduction
Imagery International website manual Prepared for: Imagery International Prepared by: Jenn de la Fuente Rosebud Designs http://www.jrosebud.com/designs designs@jrosebud.com 916.538.2133 A brief introduction
In this project, you ll learn how to create a webpage for your favourite recipe.
 Recipe Introduction In this project, you ll learn how to create a webpage for your favourite recipe. Step 1: Decide on a recipe Before you get coding, you ll need to decide on a recipe. Activity Checklist
Recipe Introduction In this project, you ll learn how to create a webpage for your favourite recipe. Step 1: Decide on a recipe Before you get coding, you ll need to decide on a recipe. Activity Checklist
DATASHEET. Colour Pot Smart Object for Crestron Smart Graphics. Revision: 1.0 Date: 09 February 2015
 Colour Pot Smart Object for Crestron Smart Graphics Revision: 1.0 Date: 09 February 2015 Summary This datasheet relates to Ultamation s Colour Pot Smart Graphics Control for Crestron control systems with
Colour Pot Smart Object for Crestron Smart Graphics Revision: 1.0 Date: 09 February 2015 Summary This datasheet relates to Ultamation s Colour Pot Smart Graphics Control for Crestron control systems with
Things to note: Each week Xampp will need to be installed. Xampp is Windows software, similar software is available for Mac, called Mamp.
 Tutorial 8 Editor Brackets Goals Introduction to PHP and MySql. - Set up and configuration of Xampp - Learning Data flow Things to note: Each week Xampp will need to be installed. Xampp is Windows software,
Tutorial 8 Editor Brackets Goals Introduction to PHP and MySql. - Set up and configuration of Xampp - Learning Data flow Things to note: Each week Xampp will need to be installed. Xampp is Windows software,
User Guide. Remote Support Tool
 Remote Support Tool Remote Support Tool...1 Overview...1 Starting the Support Tool...1 Starting a Remote Support Session...2 Using the Support Tool in an Office...3 Remote Support Tool At a glance...4
Remote Support Tool Remote Support Tool...1 Overview...1 Starting the Support Tool...1 Starting a Remote Support Session...2 Using the Support Tool in an Office...3 Remote Support Tool At a glance...4
CROMWELLSTUDIOS. Content Management System Instruction Manual V1. Content Management System. V1
 Content Management System Instruction Manual V1 www.cromwellstudios.co.uk Cromwell Studios Web Services Content Management System Manual Part 1 Content Management is the system by which you can change
Content Management System Instruction Manual V1 www.cromwellstudios.co.uk Cromwell Studios Web Services Content Management System Manual Part 1 Content Management is the system by which you can change
Tar Heel Chinese Crested Club Instructions for Changing THCCC Website Passwords
 Tar Heel Chinese Crested Club Instructions for Changing THCCC Website Passwords The cpanel is used to maintain the THCCC website. Copy this address: http://www.tarheelchinesecrestedclub.org:2082/ and paste
Tar Heel Chinese Crested Club Instructions for Changing THCCC Website Passwords The cpanel is used to maintain the THCCC website. Copy this address: http://www.tarheelchinesecrestedclub.org:2082/ and paste
Baidu Map Widget Installation & Extension User Guide
 Baidu Map Widget Installation & Extension User Guide Version 1.0b July 2015 Table of Contents 1. Introduction and Installation... 1 2. Baidu Map Extension: Configuration and Use... 5 3. Compatibility...
Baidu Map Widget Installation & Extension User Guide Version 1.0b July 2015 Table of Contents 1. Introduction and Installation... 1 2. Baidu Map Extension: Configuration and Use... 5 3. Compatibility...
File Upload Instructions Customer Access To Transcript Bulletin Publishing s FTP Site
 File Upload Instructions Customer Access To Transcript Bulletin Publishing s FTP Site In order to upload files to our FTP site, you will need a Java-enabled web browser for Microsoft Windows and Mac OS
File Upload Instructions Customer Access To Transcript Bulletin Publishing s FTP Site In order to upload files to our FTP site, you will need a Java-enabled web browser for Microsoft Windows and Mac OS
EKTRON 101: THE BASICS
 EKTRON 101: THE BASICS Table of Contents INTRODUCTION... 2 TERMINOLOGY... 2 WHY DO SOME PAGES LOOK DIFFERENT THAN OTHERS?... 5 LOGGING IN... 8 Choosing an edit mode... 10 Edit in context mode (easy editing)...
EKTRON 101: THE BASICS Table of Contents INTRODUCTION... 2 TERMINOLOGY... 2 WHY DO SOME PAGES LOOK DIFFERENT THAN OTHERS?... 5 LOGGING IN... 8 Choosing an edit mode... 10 Edit in context mode (easy editing)...
Lab: Using R and Bioconductor
 Lab: Using R and Bioconductor Robert Gentleman Florian Hahne Paul Murrell June 19, 2006 Introduction In this lab we will cover some basic uses of R and also begin working with some of the Bioconductor
Lab: Using R and Bioconductor Robert Gentleman Florian Hahne Paul Murrell June 19, 2006 Introduction In this lab we will cover some basic uses of R and also begin working with some of the Bioconductor
CREATING WEBSITES. What you need to build a website Part One The Basics. Chas Large. Welcome one and all
 Slide 1 CREATING WEBSITES What you need to build a website Part One The Basics Chas Large Welcome one and all Short intro about Chas large TV engineer, computer geek, self taught, became IT manager in
Slide 1 CREATING WEBSITES What you need to build a website Part One The Basics Chas Large Welcome one and all Short intro about Chas large TV engineer, computer geek, self taught, became IT manager in
IPM-01 / IPM-01H MODBUS TCP/RTU Bridge User Guide
 VxI Power Ltd. IPM-01 / IPM-01H MODBUS TCP/RTU Bridge User Guide 01/12/2015 Document Number: 14970-020A Issue Number: 2 Contents 1.0 Device Overview... 2 2.0 Getting Started... 3 2.1 Connecting the Device...
VxI Power Ltd. IPM-01 / IPM-01H MODBUS TCP/RTU Bridge User Guide 01/12/2015 Document Number: 14970-020A Issue Number: 2 Contents 1.0 Device Overview... 2 2.0 Getting Started... 3 2.1 Connecting the Device...
Ph3 Mathematica Homework: Week 1
 Ph3 Mathematica Homework: Week 1 Eric D. Black California Institute of Technology v1.1 1 Obtaining, installing, and starting Mathematica Exercise 1: If you don t already have Mathematica, download it and
Ph3 Mathematica Homework: Week 1 Eric D. Black California Institute of Technology v1.1 1 Obtaining, installing, and starting Mathematica Exercise 1: If you don t already have Mathematica, download it and
ChIP-Seq Tutorial on Galaxy
 1 Introduction ChIP-Seq Tutorial on Galaxy 2 December 2010 (modified April 6, 2017) Rory Stark The aim of this practical is to give you some experience handling ChIP-Seq data. We will be working with data
1 Introduction ChIP-Seq Tutorial on Galaxy 2 December 2010 (modified April 6, 2017) Rory Stark The aim of this practical is to give you some experience handling ChIP-Seq data. We will be working with data
DOCMAIL: DATA INTELLIGENCE. Adding Data-driver styles and images
 DOCMAIL: DATA INTELLIGENCE Adding Data-driver styles and images 1 Docmail DATA INTELLIGENCE Docmail is capable of far more than simply mail-merging address data and adding names within content. In this
DOCMAIL: DATA INTELLIGENCE Adding Data-driver styles and images 1 Docmail DATA INTELLIGENCE Docmail is capable of far more than simply mail-merging address data and adding names within content. In this
READSPEAKER BLACKBOARD BUILDING BLOCK
 READSPEAKER BLACKBOARD BUILDING BLOCK System Administrator Guide Version 1.0.4 This guide is intended for Blackboard System Administrators and describes how to install and configure the ReadSpeaker. This
READSPEAKER BLACKBOARD BUILDING BLOCK System Administrator Guide Version 1.0.4 This guide is intended for Blackboard System Administrators and describes how to install and configure the ReadSpeaker. This
Reading How the Web Works
 Reading 1.3 - How the Web Works By Jonathan Lane Introduction Every so often, you get offered a behind-the-scenes look at the cogs and fan belts behind the action. Today is your lucky day. In this article
Reading 1.3 - How the Web Works By Jonathan Lane Introduction Every so often, you get offered a behind-the-scenes look at the cogs and fan belts behind the action. Today is your lucky day. In this article
epigenomegateway.wustl.edu
 Everything can be found at epigenomegateway.wustl.edu REFERENCES 1. Zhou X, et al., Nature Methods 8, 989-990 (2011) 2. Zhou X & Wang T, Current Protocols in Bioinformatics Unit 10.10 (2012) 3. Zhou X,
Everything can be found at epigenomegateway.wustl.edu REFERENCES 1. Zhou X, et al., Nature Methods 8, 989-990 (2011) 2. Zhou X & Wang T, Current Protocols in Bioinformatics Unit 10.10 (2012) 3. Zhou X,
Practical OmicsFusion
 Practical OmicsFusion Introduction In this practical, we will analyse data, from an experiment which aim was to identify the most important metabolites that are related to potato flesh colour, from an
Practical OmicsFusion Introduction In this practical, we will analyse data, from an experiment which aim was to identify the most important metabolites that are related to potato flesh colour, from an
Week 8 Google Maps. This week you ll learn how to embed a Google Map into a web page and add custom markers with custom labels.
 Introduction Hopefully by now you ll have seen the possibilities that jquery provides for rich content on web sites in the form of interaction and media playback. This week we ll be extending this into
Introduction Hopefully by now you ll have seen the possibilities that jquery provides for rich content on web sites in the form of interaction and media playback. This week we ll be extending this into
Using Dreamweaver CC. 6 Styles in Websites. Exercise 1 Linked Styles vs Embedded Styles
 Using Dreamweaver CC 6 So far we have used CSS to arrange the elements on our web page. We have also used CSS for some limited formatting. In this section we will take full advantage of using CSS to format
Using Dreamweaver CC 6 So far we have used CSS to arrange the elements on our web page. We have also used CSS for some limited formatting. In this section we will take full advantage of using CSS to format
Practical Course in Genome Bioinformatics
 Practical Course in Genome Bioinformatics 20/01/2017 Exercises - Day 1 http://ekhidna.biocenter.helsinki.fi/downloads/teaching/spring2017/ Answer questions Q1-Q3 below and include requested Figures 1-5
Practical Course in Genome Bioinformatics 20/01/2017 Exercises - Day 1 http://ekhidna.biocenter.helsinki.fi/downloads/teaching/spring2017/ Answer questions Q1-Q3 below and include requested Figures 1-5
Exploring cdna Data. Achim Tresch, Andreas Buness, Wolfgang Huber, Tim Beißbarth
 Exploring cdna Data Achim Tresch, Andreas Buness, Wolfgang Huber, Tim Beißbarth Practical DNA Microarray Analysis http://compdiag.molgen.mpg.de/ngfn/pma0nov.shtml The following exercise will guide you
Exploring cdna Data Achim Tresch, Andreas Buness, Wolfgang Huber, Tim Beißbarth Practical DNA Microarray Analysis http://compdiag.molgen.mpg.de/ngfn/pma0nov.shtml The following exercise will guide you
KC-SMART Vignette. Jorma de Ronde, Christiaan Klijn April 30, 2018
 KC-SMART Vignette Jorma de Ronde, Christiaan Klijn April 30, 2018 1 Introduction In this Vignette we demonstrate the usage of the KCsmart R package. This R implementation is based on the original implementation
KC-SMART Vignette Jorma de Ronde, Christiaan Klijn April 30, 2018 1 Introduction In this Vignette we demonstrate the usage of the KCsmart R package. This R implementation is based on the original implementation
MAKING MAPS WITH GOOGLE FUSION TABLES. (Data for this tutorial at
 MAKING MAPS WITH GOOGLE FUSION TABLES (Data for this tutorial at www.peteraldhous.com/data) Thanks to Google Fusion Tables, creating maps from data and embedding them on a web page is now easy. We re going
MAKING MAPS WITH GOOGLE FUSION TABLES (Data for this tutorial at www.peteraldhous.com/data) Thanks to Google Fusion Tables, creating maps from data and embedding them on a web page is now easy. We re going
Fish Eye Menu DMXzone.com Fish Eye Menu Manual
 Fish Eye Menu Manual Page 1 of 33 Index Fish Eye Menu Manual... 1 Index... 2 About Fish Eye Menu... 3 Features in Detail... 4 Integrated in Dreamweaver... 6 Before you begin... 7 Installing the extension...
Fish Eye Menu Manual Page 1 of 33 Index Fish Eye Menu Manual... 1 Index... 2 About Fish Eye Menu... 3 Features in Detail... 4 Integrated in Dreamweaver... 6 Before you begin... 7 Installing the extension...
Everything in red on the screenshots has been added for the purpose of this user guide and is the context for the words around it.
 Huddle for Office What is it? Huddle for Office brings the best collaborative parts of Huddle right into your applications. You are able to take the content that you are working on straight from Huddle,
Huddle for Office What is it? Huddle for Office brings the best collaborative parts of Huddle right into your applications. You are able to take the content that you are working on straight from Huddle,
User Guide for The Core
 User Guide for The Core Version 3.1 www.rock7mobile.com General Overview The Back-Office system is designed to provide a single-point of control for Iridium-based tracking devices. The system can support
User Guide for The Core Version 3.1 www.rock7mobile.com General Overview The Back-Office system is designed to provide a single-point of control for Iridium-based tracking devices. The system can support
Gdes2000 Graphic Design and the Internet
 Dreamweaver Rough Photoshop Layouts to DIVs. Using AP DIVs (Absolutely Positioned) Introducing RGB, 72dpi, page sections Last Session: We created a basic DIV Page Layout (Above) We re going to build on
Dreamweaver Rough Photoshop Layouts to DIVs. Using AP DIVs (Absolutely Positioned) Introducing RGB, 72dpi, page sections Last Session: We created a basic DIV Page Layout (Above) We re going to build on
Lab 1 - Introduction to Angular
 Lab 1 - Introduction to Angular In this lab we will build a Hello World style Angular component. The key focus is to learn how to install all the required code and use them from the browser. We wont get
Lab 1 - Introduction to Angular In this lab we will build a Hello World style Angular component. The key focus is to learn how to install all the required code and use them from the browser. We wont get
An Introduction to BigQuery
 An Introduction to BigQuery (in less than 10 minutes) brought to you by The ISB Cancer Genomics Cloud This is what you should see the first time you go to the BigQuery Web UI at bigquery.cloud.google.com
An Introduction to BigQuery (in less than 10 minutes) brought to you by The ISB Cancer Genomics Cloud This is what you should see the first time you go to the BigQuery Web UI at bigquery.cloud.google.com
Lecture 3 - Template and Vectors
 Lecture - Template and Vectors Homework Format and Template: We ll each develop a simple template to use to start any new homework. The idea of a template is to layout the basic structure of what goes
Lecture - Template and Vectors Homework Format and Template: We ll each develop a simple template to use to start any new homework. The idea of a template is to layout the basic structure of what goes
ebook Writing Workshop Practical 1: A First Logfile-Style ebook
 ebook Writing Workshop Practical 1: A First Logfile-Style ebook Introduction In this practical we will aim to get you familiar with both the TREE (Template Reading & Execution Environment) and DEEP (Documents
ebook Writing Workshop Practical 1: A First Logfile-Style ebook Introduction In this practical we will aim to get you familiar with both the TREE (Template Reading & Execution Environment) and DEEP (Documents
CompClustTk Manual & Tutorial
 CompClustTk Manual & Tutorial Brandon King Copyright c California Institute of Technology Version 0.1.10 May 13, 2004 Contents 1 Introduction 1 1.1 Purpose.............................................
CompClustTk Manual & Tutorial Brandon King Copyright c California Institute of Technology Version 0.1.10 May 13, 2004 Contents 1 Introduction 1 1.1 Purpose.............................................
Unifer Documentation. Release V1.0. Matthew S
 Unifer Documentation Release V1.0 Matthew S July 28, 2014 Contents 1 Unifer Tutorial - Notes Web App 3 1.1 Setting up................................................. 3 1.2 Getting the Template...........................................
Unifer Documentation Release V1.0 Matthew S July 28, 2014 Contents 1 Unifer Tutorial - Notes Web App 3 1.1 Setting up................................................. 3 1.2 Getting the Template...........................................
Adaptive Layout Hands-On Challenges
 Adaptive Layout Hands-On Challenges Copyright 2015 Razeware LLC. All rights reserved. No part of this book or corresponding materials (such as text, images, or source code) may be reproduced or distributed
Adaptive Layout Hands-On Challenges Copyright 2015 Razeware LLC. All rights reserved. No part of this book or corresponding materials (such as text, images, or source code) may be reproduced or distributed
WordPress site Import/Export Procedure
 Platform Export single site from WordPress single site (SOURCE SITE) Import to one site on VSB Blog site (DESTINATION SITE) This procedure does not require direct access to the file servers that host the
Platform Export single site from WordPress single site (SOURCE SITE) Import to one site on VSB Blog site (DESTINATION SITE) This procedure does not require direct access to the file servers that host the
This Tutorial is for Word 2007 but 2003 instructions are included in [brackets] after of each step.
![This Tutorial is for Word 2007 but 2003 instructions are included in [brackets] after of each step. This Tutorial is for Word 2007 but 2003 instructions are included in [brackets] after of each step.](/thumbs/72/67280806.jpg) This Tutorial is for Word 2007 but 2003 instructions are included in [brackets] after of each step. Table of Contents Just so you know: Things You Can t Do with Word... 1 Get Organized... 1 Create the
This Tutorial is for Word 2007 but 2003 instructions are included in [brackets] after of each step. Table of Contents Just so you know: Things You Can t Do with Word... 1 Get Organized... 1 Create the
IT153 Midterm Study Guide
 IT153 Midterm Study Guide These are facts about the Adobe Dreamweaver CS4 Application. If you know these facts, you should be able to do well on your midterm. Dreamweaver users work in the Document window
IT153 Midterm Study Guide These are facts about the Adobe Dreamweaver CS4 Application. If you know these facts, you should be able to do well on your midterm. Dreamweaver users work in the Document window
HTML and CSS a further introduction
 HTML and CSS a further introduction By now you should be familiar with HTML and CSS and what they are, HTML dictates the structure of a page, CSS dictates how it looks. This tutorial will teach you a few
HTML and CSS a further introduction By now you should be familiar with HTML and CSS and what they are, HTML dictates the structure of a page, CSS dictates how it looks. This tutorial will teach you a few
Verification and Configuration Instructions. JMP Genomics 3.1 for SAS Required Verification and Configuration Steps.
 Verification and Configuration Instructions JMP Genomics 3.1 for SAS 9.1.3 Required Verification and Configuration Steps Once you have successfully completed the installation of the SAS 9.1.3 Foundation
Verification and Configuration Instructions JMP Genomics 3.1 for SAS 9.1.3 Required Verification and Configuration Steps Once you have successfully completed the installation of the SAS 9.1.3 Foundation
Level 5 Digital technologies Chapter 4
 Level 5 Digital technologies Chapter 4 Digital information database and spreadsheets Technological Practice Strand: Brief Development In this chapter you are going to develop your database and spreadsheet
Level 5 Digital technologies Chapter 4 Digital information database and spreadsheets Technological Practice Strand: Brief Development In this chapter you are going to develop your database and spreadsheet
Joomla 3.X Global Settings Part III Server Settings
 Joomla 3.X Global Settings Part III Server Settings Diagram 1 Path to Temp Folder: This is a text box adjacent to this prompt which holds the path to Joomla s temp folder on the web server. This is the
Joomla 3.X Global Settings Part III Server Settings Diagram 1 Path to Temp Folder: This is a text box adjacent to this prompt which holds the path to Joomla s temp folder on the web server. This is the
How to connect to the University of Exeter VPN service
 How to connect to the University of Exeter VPN service *****Important Part of the process of using the VPN service involves the automatic download and installation of Juniper Network Connect software,
How to connect to the University of Exeter VPN service *****Important Part of the process of using the VPN service involves the automatic download and installation of Juniper Network Connect software,
Hierarchical clustering
 Hierarchical clustering Rebecca C. Steorts, Duke University STA 325, Chapter 10 ISL 1 / 63 Agenda K-means versus Hierarchical clustering Agglomerative vs divisive clustering Dendogram (tree) Hierarchical
Hierarchical clustering Rebecca C. Steorts, Duke University STA 325, Chapter 10 ISL 1 / 63 Agenda K-means versus Hierarchical clustering Agglomerative vs divisive clustering Dendogram (tree) Hierarchical
HTML/CSS Lesson Plans
 HTML/CSS Lesson Plans Course Outline 8 lessons x 1 hour Class size: 15-25 students Age: 10-12 years Requirements Computer for each student (or pair) and a classroom projector Pencil and paper Internet
HTML/CSS Lesson Plans Course Outline 8 lessons x 1 hour Class size: 15-25 students Age: 10-12 years Requirements Computer for each student (or pair) and a classroom projector Pencil and paper Internet
Tutorial: chloroplast genomes
 Tutorial: chloroplast genomes Stacia Wyman Department of Computer Sciences Williams College Williamstown, MA 01267 March 10, 2005 ASSUMPTIONS: You are using Internet Explorer under OS X on the Mac. You
Tutorial: chloroplast genomes Stacia Wyman Department of Computer Sciences Williams College Williamstown, MA 01267 March 10, 2005 ASSUMPTIONS: You are using Internet Explorer under OS X on the Mac. You
Club Site Editing Guide
 Swimming New Zealand Club Site Editing Guide Intelligent Software for Membership All information contained in this document is for the exclusive use of Swimming New Zealand. We are happy that any part
Swimming New Zealand Club Site Editing Guide Intelligent Software for Membership All information contained in this document is for the exclusive use of Swimming New Zealand. We are happy that any part
General Use. Searching for Assets (All users) Browsing for Assets (All users) Viewing and Downloading an Asset (All Users)
 User Guide Rev1.1 Table of Contents General Use... 2 Searching for Assets (All users)... 2 Browsing for Assets (All users)... 2 Viewing and Downloading an Asset (All Users)... 2 Downloading Large Files
User Guide Rev1.1 Table of Contents General Use... 2 Searching for Assets (All users)... 2 Browsing for Assets (All users)... 2 Viewing and Downloading an Asset (All Users)... 2 Downloading Large Files
How To: Run the ENCODE histone ChIP- seq analysis pipeline on DNAnexus
 How To: Run the ENCODE histone ChIP- seq analysis pipeline on DNAnexus Overview: In this exercise, we will run the ENCODE Uniform Processing ChIP- seq Pipeline on a small test dataset containing reads
How To: Run the ENCODE histone ChIP- seq analysis pipeline on DNAnexus Overview: In this exercise, we will run the ENCODE Uniform Processing ChIP- seq Pipeline on a small test dataset containing reads
Yusen Terminal Appointment System. Trucking Company Guide 18 April 2017
 Yusen Terminal Appointment System Trucking Company Guide 18 April 2017 Contents Signing up Creating appointments Reporting Configuring your views Troubleshooting Signing Up Log In / Sign Up Go to the Yusen
Yusen Terminal Appointment System Trucking Company Guide 18 April 2017 Contents Signing up Creating appointments Reporting Configuring your views Troubleshooting Signing Up Log In / Sign Up Go to the Yusen
CS 268 Lab 6 Eclipse Test Server and JSPs
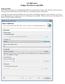 CS 268 Lab 6 Eclipse Test Server and JSPs Setting up Eclipse The first thing you will do is to setup the Eclipse Web Server environment for testing. This will create a local web server running on your
CS 268 Lab 6 Eclipse Test Server and JSPs Setting up Eclipse The first thing you will do is to setup the Eclipse Web Server environment for testing. This will create a local web server running on your
Data Walkthrough: Background
 Data Walkthrough: Background File Types FASTA Files FASTA files are text-based representations of genetic information. They can contain nucleotide or amino acid sequences. For this activity, students will
Data Walkthrough: Background File Types FASTA Files FASTA files are text-based representations of genetic information. They can contain nucleotide or amino acid sequences. For this activity, students will
Blackfin Online Learning & Development
 Presentation Title: Multimedia Starter Kit Presenter Name: George Stephan Chapter 1: Introduction Sub-chapter 1a: Overview Chapter 2: Blackfin Starter Kits Sub-chapter 2a: What is a Starter Kit? Sub-chapter
Presentation Title: Multimedia Starter Kit Presenter Name: George Stephan Chapter 1: Introduction Sub-chapter 1a: Overview Chapter 2: Blackfin Starter Kits Sub-chapter 2a: What is a Starter Kit? Sub-chapter
Beginning HTML. The Nuts and Bolts of building Web pages.
 Beginning HTML The Nuts and Bolts of building Web pages. Overview Today we will cover: 1. what is HTML and what is it not? Building a simple webpage Getting that online. What is HTML? The language of the
Beginning HTML The Nuts and Bolts of building Web pages. Overview Today we will cover: 1. what is HTML and what is it not? Building a simple webpage Getting that online. What is HTML? The language of the
You will always have access to the training area if you want to experiment or repeat this tutorial.
 EasySite Tutorial: Part One Welcome to the EasySite tutorial session. Core Outcomes After this session, you will be able to: Create new pages and edit existing pages on Aston s website. Add different types
EasySite Tutorial: Part One Welcome to the EasySite tutorial session. Core Outcomes After this session, you will be able to: Create new pages and edit existing pages on Aston s website. Add different types
User Guide. Remote Support Tool
 Remote Support Tool Remote Support Tool... 1 User Guide... 1 Overview... 1 Starting the Support Tool... 1 Starting a Remote Support Session... 2 Using TeamViewer... 3 Using the Support Tool in an Office...
Remote Support Tool Remote Support Tool... 1 User Guide... 1 Overview... 1 Starting the Support Tool... 1 Starting a Remote Support Session... 2 Using TeamViewer... 3 Using the Support Tool in an Office...
SISG/SISMID Module 3
 SISG/SISMID Module 3 Introduction to R Ken Rice Tim Thornton University of Washington Seattle, July 2018 Introduction: Course Aims This is a first course in R. We aim to cover; Reading in, summarizing
SISG/SISMID Module 3 Introduction to R Ken Rice Tim Thornton University of Washington Seattle, July 2018 Introduction: Course Aims This is a first course in R. We aim to cover; Reading in, summarizing
Creating an with Constant Contact. A step-by-step guide
 Creating an Email with Constant Contact A step-by-step guide About this Manual Once your Constant Contact account is established, use this manual as a guide to help you create your email campaign Here
Creating an Email with Constant Contact A step-by-step guide About this Manual Once your Constant Contact account is established, use this manual as a guide to help you create your email campaign Here
XML and API Documentation
 ATLauncher XML and API Documentation Version 1.0.22 Last Updated: 18/10/2013 Table of Contents XML Introduction... 2 Basic Layout... 3 ... 4 ... 5 ... 7 ... 8 Mod Types...
ATLauncher XML and API Documentation Version 1.0.22 Last Updated: 18/10/2013 Table of Contents XML Introduction... 2 Basic Layout... 3 ... 4 ... 5 ... 7 ... 8 Mod Types...
In this exercise you will display the Geo-tagged Wikipedia Articles Fusion Table in Google Maps.
 Introduction to the Google Maps API v3 Adding a Fusion Table to Google Maps Fusion Tables, still in the experimental stages of development, is a Google product that allows you to upload and share data
Introduction to the Google Maps API v3 Adding a Fusion Table to Google Maps Fusion Tables, still in the experimental stages of development, is a Google product that allows you to upload and share data
Server Status Dashboard
 The Cisco Prime Network Registrar server status dashboard in the web user interface (web UI) presents a graphical view of the system status, using graphs, charts, and tables, to help in tracking and diagnosis.
The Cisco Prime Network Registrar server status dashboard in the web user interface (web UI) presents a graphical view of the system status, using graphs, charts, and tables, to help in tracking and diagnosis.
ClaNC: The Manual (v1.1)
 ClaNC: The Manual (v1.1) Alan R. Dabney June 23, 2008 Contents 1 Installation 3 1.1 The R programming language............................... 3 1.2 X11 with Mac OS X....................................
ClaNC: The Manual (v1.1) Alan R. Dabney June 23, 2008 Contents 1 Installation 3 1.1 The R programming language............................... 3 1.2 X11 with Mac OS X....................................
Browsing the World Wide Web with Firefox
 Browsing the World Wide Web with Firefox B 660 / 1 Try this Popular and Featurepacked Free Alternative to Internet Explorer Internet Explorer 7 arrived with a bang a few months ago, but it hasn t brought
Browsing the World Wide Web with Firefox B 660 / 1 Try this Popular and Featurepacked Free Alternative to Internet Explorer Internet Explorer 7 arrived with a bang a few months ago, but it hasn t brought
Lecture 11: Recursion (hard)
 Lecture 11: Recursion (hard) Recap/finish: stutter an integer 348 > 334488 We added the base case So how do we deal with larger numbers how can we split them up? We need to split into smaller chunks Since
Lecture 11: Recursion (hard) Recap/finish: stutter an integer 348 > 334488 We added the base case So how do we deal with larger numbers how can we split them up? We need to split into smaller chunks Since
Click on "+" button Select your VCF data files (see #Input Formats->1 above) Remove file from files list:
 CircosVCF: CircosVCF is a web based visualization tool of genome-wide variant data described in VCF files using circos plots. The provided visualization capabilities, gives a broad overview of the genomic
CircosVCF: CircosVCF is a web based visualization tool of genome-wide variant data described in VCF files using circos plots. The provided visualization capabilities, gives a broad overview of the genomic
**Method 3** By attaching a style sheet to your web page, and then placing all your styles in that new style sheet.
 CSS Tutorial Part 1: Introduction: CSS adds style to tags in your html page. With HTML you told the browser what things were (e.g., this is a paragraph). Now you are telling the browser how things look
CSS Tutorial Part 1: Introduction: CSS adds style to tags in your html page. With HTML you told the browser what things were (e.g., this is a paragraph). Now you are telling the browser how things look
ValuePRO Tutorial Internet Explorer 8 Configuration
 ValuePRO Tutorial Internet Explorer 8 Configuration Table of Contents Contents 1. Adding ValuePRO to Trusted Sites... 1 1. Overview... 1 2. Changes Required... 1 2. Enabling Cross Site Scripting... 3 1.
ValuePRO Tutorial Internet Explorer 8 Configuration Table of Contents Contents 1. Adding ValuePRO to Trusted Sites... 1 1. Overview... 1 2. Changes Required... 1 2. Enabling Cross Site Scripting... 3 1.
Website Development Komodo Editor and HTML Intro
 Website Development Komodo Editor and HTML Intro Introduction In this Lecture and Tour we will cover: o Use of the editor that will be used for the Website Development and Javascript Programming sections
Website Development Komodo Editor and HTML Intro Introduction In this Lecture and Tour we will cover: o Use of the editor that will be used for the Website Development and Javascript Programming sections
QUICK REFERENCE GUIDE
 Folders new projects. Organise your folders to find files quickly and easily 1 Look in your yellow storage Folders it can be organised into simple folder structures to help with browsing 2 Click on your
Folders new projects. Organise your folders to find files quickly and easily 1 Look in your yellow storage Folders it can be organised into simple folder structures to help with browsing 2 Click on your
1. Register Computer: Each computer you use to access PaymentNet must now be registered. The first computer you will log on with, will be
 Call or email the PCard Administrators to: 1. Make changes to cardholders profile 2. Deactivate/close a cardholder s account 3. Clarify PCard policies and procedures 4. Resolve declined transactions 5.
Call or email the PCard Administrators to: 1. Make changes to cardholders profile 2. Deactivate/close a cardholder s account 3. Clarify PCard policies and procedures 4. Resolve declined transactions 5.
CS1046 Lab 4. Timing: This lab should take you 85 to 130 minutes. Objectives: By the end of this lab you should be able to:
 CS1046 Lab 4 Timing: This lab should take you 85 to 130 minutes. Objectives: By the end of this lab you should be able to: Define the terms: function, calling and user-defined function and predefined function
CS1046 Lab 4 Timing: This lab should take you 85 to 130 minutes. Objectives: By the end of this lab you should be able to: Define the terms: function, calling and user-defined function and predefined function
Deposit Wizard Panini Installation Guide
 Guide Table of Contents System Requirements... 2 WebScan Overview... 2 Hardware Requirements... 2 Supported Browsers... 2 Driver Installation... 2 Step 1 - Determining Windows Edition & Bit Count... 3
Guide Table of Contents System Requirements... 2 WebScan Overview... 2 Hardware Requirements... 2 Supported Browsers... 2 Driver Installation... 2 Step 1 - Determining Windows Edition & Bit Count... 3
Moose for Java Enterprise Application
 Moose for Java Enterprise Application Perin Fabrizio SCG - Software Composition Group Institut für Informatik und Angewandte Mathematik University of Bern, Switzerland 18/03/10 Revision History Date Ver.
Moose for Java Enterprise Application Perin Fabrizio SCG - Software Composition Group Institut für Informatik und Angewandte Mathematik University of Bern, Switzerland 18/03/10 Revision History Date Ver.
CS Multimedia and Communications REMEMBER TO BRING YOUR MEMORY STICK TO EVERY LAB!
 CS 1033 Multimedia and Communications REMEMBER TO BRING YOUR MEMORY STICK TO EVERY LAB! Lab 06: Introduction to KompoZer (Website Design - Part 3 of 3) Lab 6 Tutorial 1 In this lab we are going to learn
CS 1033 Multimedia and Communications REMEMBER TO BRING YOUR MEMORY STICK TO EVERY LAB! Lab 06: Introduction to KompoZer (Website Design - Part 3 of 3) Lab 6 Tutorial 1 In this lab we are going to learn
1 Lecture 5: Advanced Data Structures
 L5 June 14, 2017 1 Lecture 5: Advanced Data Structures CSCI 1360E: Foundations for Informatics and Analytics 1.1 Overview and Objectives We ve covered list, tuples, sets, and dictionaries. These are the
L5 June 14, 2017 1 Lecture 5: Advanced Data Structures CSCI 1360E: Foundations for Informatics and Analytics 1.1 Overview and Objectives We ve covered list, tuples, sets, and dictionaries. These are the
Dreamweaver MX Overview. Maintaining a Web Site
 Dreamweaver MX Overview Maintaining a Web Site... 1 The Process... 1 Filenames... 1 Starting Dreamweaver... 2 Uploading and Downloading Files... 6 Check In and Check Out Files... 6 Editing Pages in Dreamweaver...
Dreamweaver MX Overview Maintaining a Web Site... 1 The Process... 1 Filenames... 1 Starting Dreamweaver... 2 Uploading and Downloading Files... 6 Check In and Check Out Files... 6 Editing Pages in Dreamweaver...
StudentSignature: Student Number: Your Computer Number. Concepts for Advanced Computer Usage.
 University of Waterloo Place sticker here CS200 Lab Exam Student Name: StudentSignature: Student Number: Handin Code: Your Computer Number Course Abbreviation: Course Title: Course Instructor: CS200 Concepts
University of Waterloo Place sticker here CS200 Lab Exam Student Name: StudentSignature: Student Number: Handin Code: Your Computer Number Course Abbreviation: Course Title: Course Instructor: CS200 Concepts
Top 10 Productivity Tips in Fusion 360
 CP11251 Top 10 Productivity Tips in Fusion 360 Taylor Stein Autodesk Inc. Learning Objectives Learn how to speed up Fusion 360 workflows Learn how to make your selections even easier Learn important best
CP11251 Top 10 Productivity Tips in Fusion 360 Taylor Stein Autodesk Inc. Learning Objectives Learn how to speed up Fusion 360 workflows Learn how to make your selections even easier Learn important best
ecommerce Documentation Powered by
 ecommerce Documentation Powered by Last Updated November 24 th 2015 Contents Introduction... 3 Products... 4 Add/Edit Products... 4 Product Details... 4 Product Categories... 5 Product Tags (optional)...
ecommerce Documentation Powered by Last Updated November 24 th 2015 Contents Introduction... 3 Products... 4 Add/Edit Products... 4 Product Details... 4 Product Categories... 5 Product Tags (optional)...
Day 1: Introduction to MATLAB and Colorizing Images CURIE Academy 2015: Computational Photography Sign-Off Sheet
 Day 1: Introduction to MATLAB and Colorizing Images CURIE Academy 2015: Computational Photography Sign-Off Sheet NAME: NAME: Part 1.1 Part 1.2 Part 1.3 Part 2.1 Part 2.2 Part 3.1 Part 3.2 Sign-Off Milestone
Day 1: Introduction to MATLAB and Colorizing Images CURIE Academy 2015: Computational Photography Sign-Off Sheet NAME: NAME: Part 1.1 Part 1.2 Part 1.3 Part 2.1 Part 2.2 Part 3.1 Part 3.2 Sign-Off Milestone
MAP-BASED WEB EXPORT MANUAL. Publish your own photos to a map-based site.
 MAP-BASED WEB EXPORT MANUAL Publish your own photos to a map-based site. Introduction... 1 Requirements... 1 Contact and Support... 1 Legal Stuff... 1 Features... 2 Demo Restrictions... 2 Installation...
MAP-BASED WEB EXPORT MANUAL Publish your own photos to a map-based site. Introduction... 1 Requirements... 1 Contact and Support... 1 Legal Stuff... 1 Features... 2 Demo Restrictions... 2 Installation...
