GenomeMixer
|
|
|
- Dustin Wells
- 6 years ago
- Views:
Transcription
1 GenomeMixer What is GenomeMixer? GenomeMixer is a cross-platform program that allows you to simulate complex multigenerational breeding schemes for up to several hundred cages (or plots) and several hundred generations. GenomeMixer will handle from 2 to 20,000 parental strains. The number and length of chromosomes is fully editable, and you should be able to model almost any sexually reproducing diploid species. Copyright 2003, Alex Williams, at the University of Tennessee Health Science Center. All rights reserved. Send correspondence to Robert W. Williams <rwilliam@nb.utmem.edu>. This project was funded jointly by the National Institute of Mental Health, National Institute on Drug Abuse, and the National Science Foundation (award P20-MH to KFM and RW). GNU General Public License (GPL): The source code to this program may be used according to the terms of the GNU General Public License (GPL, see for specifics). The source may be freely used as long as the program it is being used in is also released under the GPL. Using figures / images / data from GenomeMixer: Currently, the only visual output directly from GenomeMixer is the graphical representation of chromosome haplotypes available in the "Inspect Progeny" window. You may freely use screen captures, text output, or any other output from this program, as long as it is reasonably clear that the information came from GenomeMixer. Trolltech Qt and Compiling GenomeMixer from source: GenomeMixer is written in C++. It uses the Trolltech Qt toolkit for its window-managing needs, so you will need to have Trolltech Qt installed in order to compile GenomeMixer from the source code. Qt is available for Mac OS X, Linux/Unix, and Windows. Check < for details; we use the commercial version of Qt, but there is also a free GPL-software version that you can download from Trolltech's web site ( Any redistributions of this software in code or binary form must include the complete documentation. Please include all of the documentation and the copyright notices from the source code.
2 2. What do I need to run GenomeMixer? GenomeMixer is available for Mac OS X, Windows, and Unix/Linux, from < If you are running on a Macintosh or Windows system, you should be able to download GenomeMixer as a double-clickable application. On Linux, you can download an x86 binary, but if you are not able to find a binary for your particular system, you can compile GenomeMixer from the source. You can also compile GenomeMixer from the source on a Macintosh or PC. For compiling the source on any platform, however, you will need to download Trolltech Qt ( first. Be aware that Qt is over a hundred megabytes when installed, and takes several hours to compile. 2b. Technical Question: I want to compile GenomeMixer from source. Once I have installed Qt, how do I compile the GenomeMixer source? Once you have installed a copy of Qt, you should then be able to cd to the GenomeMixer directory, type qmake, then type make once that is finished. Aft er that, there should be a GenomeMixer executable program for you to run with./genomemixer. 2c. Technical Question: How do I compile Qt in the first place? Unfortunately, that is beyond the scope of this file. We suggest that you look for help on the Trolltech web site. 3. How to run GenomeMixer: 1. Use a web browser to download GenomeMixer from the links at < 2. Decompress the downloaded file with a utility like StuffIt Expander. 3. On Mac OS X or Windows, you should now be able to double-click the GenomeMixer icon to launch it. On Linux, you can run the program from the command line, but you must have the capability of viewing an X-Windows display on your terminal (text-only will not work). Mac OS X and Linux are also capable of using X11 to run GenomeMixer from a (perhaps faster) remote machine. If you have GenomeMixer installed on one Linux machine, or under the X11 environment on Mac OS X, you can run copies of it remotely on other machines (you will normally have to login with the -X option, indicating that the computer you logged into should export the display over the connection to your machine: for example, ssh -X myname@mixer.computer.xyz.
3 Now that you have GenomeMixer running properly, you should see a window that looks something like one of these: The window to the left is an example of using GenomeMixer over the network; the screenshot was taken on a Mac OS X system using Apple X11 (notice the Aqua buttons at the top), but it is actually running on a Linux machine (hence the appearance of the dialog itself). The version on the right is running natively on OS X. A. The Menu bar Here is a basic overview of the menu options. Note that if you are using the Mac OS X version, the menu bar will be at the top of the screen instead of being attached directly to the GenomeMixer window. i. File Menu Contains options for creating new Settings and Markers. The "Settings" file contains all of the data required for a cross except for the markers on each chromosome, which are stored separately in the Markers file. The Quickload item immediately loads in files from the Saved_Settings folder with the special names quickmarkers.mar and quicksettings.gmx. This saves you the trouble of using the open dialogs to choose your settings each time, if you save your files (or, copies of your files safer, since there is no way to accidentally overwrite the originals) as quickmarkers.mar and quicksettings.gmx inside the Saved_Settings folder. For more information about the markers and settings files, see below. ii. Simulation Menu Contains the steps, in order, for running the simulation. First, you enter the markers and settings (this is the same as the "Set up the Cross..." button). Next, you run the simulation, and finally you inspect the resulting progeny. Page 3 of 22
4 iii. Output Menu The options in this menu correspond to the various output file types. They are disabled (grayed out) until you actually run a simulation and have progeny to export. The three linebreak options at the top allow you to specify what operating system s linebreak format should be followed. More information about the output file formats is available in the output file types section below. iv. Info Menu The info menu contains a limited amount of online help. In general, it is less thorough than this readme file. B. The Output File Types The output menu contains a list of all of the file formats that GenomeMixer can write to. The results of a cross can be exported to various other formats, so that you can take advantage of other programs' analysis features on data exported from GenomeMixer. Most of these programs handle only simple 2- way (A x B) crosses. i. QGene Format: QGene is a Macintosh application (OS 9 only, as of August 2003) by J. C. Nelson. It has excellent support for visualization of marker data. ii. MapManager QTX: MapManager QTX (Manly, KF, Cudmore, Jr, RH, Meer, JM) runs under Mac OS X, OS 9, and Windows, and is used for complex trait analysis. More information on how to import a QTX file is available below. iii. R/qtl: R/qtl by Karl Broman is a mapping program that runs as a module to the R statistical programming environment. R is a free download for nearly all modern computer systems, and R/qtl can run on any computer that supports R. iv. Plain Table format: This export format is just a simple two-dimensional table, with markers at the top of each column (markers from all chromosomes), and each individual as two rows of data (two rows of marker data, rather than one, because the first row lists marker data for the maternal chromosomes of the individual in question, and the second row lists marker data for the paternal chromosomes). Page 4 of 22
5 v. HAPPY: No support is available yet for HAPPY by Richard Mott, but this program can support up to 8-way crosses (8 parental strains). In the future, exporting to this format may be extremely useful for complex GenomeMixer crosses. C. Settings and Running the Simulation The "Open" buttons let you specify an already-existing GenomeMixer settings file or markers file. You can open the "quickload" markers and settings that come with this distribution, if you like, or select "New..." from the File menu and make your own. The three buttons to the right let you edit the current settings, run a simulation using the current settings, and inspect the generated progeny (if any: this button is disabled if there are no progeny to inspect). We discuss the settings in further detail below. Page 5 of 22
6 4. Setting up the simulation: The best way to figure out how to edit the GenomeMixer settings is probably to load in a test file and change the settings until you have a reasonable idea of what is going on. You can load the sample settings that come in the Saved_Settings folder (or you can select "Quickload" from the File menu to automatically load the files in the Saved_Settings folder named quickmarkers.mar and quicksettings.gmx). If you click on the "Set up the Cross" button (or if you select "Edit Markers and Settings" from the Simulation menu), a new window will appear with four tabs at the top: Chromosome, Markers, Purebreds, and Breeding Scheme. Here is an overview of each of the four tabs: Chromosome Tab (Mac interface): In the chromosome tab, you specify the lengths of each chromosome in megabases (millions of base pairs) and centimorgans (cm: a measurement of length based on recombination frequency). The values that come in the default settings files are values for the mouse. A. The sex chromosomes are at the top left. In our model, females have two X chromosomes, and males have an X and a Y chromosome. If the species you want to model works differently from this, you will have to keep that in mind. B. In the table, the values on the left under Len (Mbases) are the lengths in megabases (MBases). This is not really important as far as recombination is
7 concerned, but it is used internally in a limited way, so using approximately correct values is best. Values from 50 to 200 are not unusual, but of course this will vary depending on the species being modeled. C. In the table, the values on the right under Len (cm) are lengths in centimorgans (cm). The greater this value, the more recombinations we expect, on average, in the chromosome. This value is not based directly on the physical length of the chromosome. (It is possible to model recombination hotspots by specifying a large cm interval between two markers, in the markers tab.) D. The Change Interference Model button allows you to decide what kind of effect a recombination has on the probability of another recombination in an adjacent region of the chromosome. You can also turn interference off, so that recombinations become Poisson-distributed events. Currently, only absolute interference (requiring at least X bases between recombination events) is implemented. Recombinations are randomly placed based on centimorgan values between markers (not base locations). Thus, a recombination hotspot can be modeled by a sequence of markers that are close together in bases, but with a relatively large centimorgan distance between them. Each centimorgan has a 1% change of containing the site of a recombination, disregarding interference effects. Markers Tab (Mac interface) In the markers tab, you specify the lengths of each chromosome in megabases (millions of base pairs) and centimorgans (cm: a measurement of length based on recombination frequency). The values that come in the default settings files are (approximately correct) Page 7 of 22
8 values for the mouse. A. The add button adds a new marker after the currently selected one. The delete button deletes the currently selected record. Recombination only occurs between markers in our model, and two adjacent markers may only have one recombination between them per generation at most. It is possible to model recombination hotspots by specifying a large cm interval between two adjacent markers. B. The auto-generate markers button brings up a new dialog box that allows you to automatically fill in the marker table with markers spaced evenly over a certain interval. The only exception to the even spacing is that auto-generate will always create a marker at the beginning and end of each chromosome. C. There is no "Cancel" button, only an "Ok" button and a "Revert" button. The "Ok" this button will apply your changes and return to you the main GenomeMixer window. Since there is no cancel button, you can revert the settings to the way they were when you last selected the current tab by clicking the "Revert tab" button. Note that switching from one tab to another applies your changes. Note that "Ok" does not save your changes to disk. If you want to save changes to disk, you will have to select Save from the menu, after clicking "Ok." Purebreds Tab (Mac interface) In the purebreds tab, you specify the names and symbols associated with the parental purebred strains. Page 8 of 22
9 GenomeMixer Documentation A. The Strain Name field is for a reasonably human-readable name for each purebred strain. You can name your strains "AA" or "BXD32," or anything without a comma, space, or quotation marks. Note that the strain name must be a single word: so instead of using spaces, you should use the underscore character. B. The Symbol field should be either a single character (A, a, B, b, etc ) or a number from 0 to 30,000. If you use a capital letter or a number from 0 to 9, then the Progeny Inspector dialog will be color-coded, so it is recommended that you avoid lower-case or numbers greater than 10 for the symbol if possible. If you need to, you can set different Strain Names to have the same symbol. Normally you probably would not want to do this, but if you want to export GenomeMixer data to an output format that only supports 'A' and 'B' parents, you might have to set some parental strains to have the same symbols. Breeding Scheme Tab (Mac interface) This is the tab in which you set up the actual breeding scheme. You will have to set up at least two parental strains in the purebreds tab before you can do any meaningful crosses in this tab. Page 9 of 22
10 A. Each cell in the breeding scheme table specifies a particular cage and a particular generation. You can use the controls at the bottom to edit the cell, or you can double-click on the cell and edit the text manually. To delete a cell, double-click it and delete all the text in it, and then when you click away from that cell, the data will be gone. Important: Note that when you are editing the parents, any changes you type in will not take effect until you hit return / enter. Selecting a different cell does not automatically save the changes in the parental fields. B. The parental selector controls allow you to choose the parents for the progeny to be generated for the current cell. You can select Purebred from the pulldown menu, and it will bring up a list of valid purebred parents, or you can select From Cell... and then type in the identifier for a particular cell (for example, the top left cell is A1, as specified by the letter in the column header (A) Gen 1 and the number from the row header Cage (1) ). This is the same format that most spreadsheet programs use. Note that you cannot forward reference a cell; if you have selected cell B2 (Generation 2, Cage 2), then you cannot request that an individual from generation 3 (column C) be a parent, because the individuals in generation 3 would not exist at that point. All of the cells in the leftmost column (generation 1 (A)) should have purebred parents. Other cells may have purebred parents, or may specify parents from any of the cells to their left (that is, any earlier generation). C. The symbol represents the female parent and the number of females to be generated. In the Progeny Inspector, it also indicates the chromosome inherited from the individual's mother. The symbol indicates the male / paternal versions of the same things. D. The progeny controls let you decide how many individuals will be generated for this cell, from the mating specified by the parental selector controls. The default generation is 1 male and 1 female, but you can increase (or decrease) these numbers, and add individuals with randomly-assigned sex by incrementing the random counter. Page 10 of 22
11 5. Simulation and Results: Once you have specified all of the data in the four-tabbed Edit Markers and Breeding Scheme window, you should click Ok and return to the main window. There, you will be able to click on the Run Simulation button (or select Run Simulation from the Simulation menu). GenomeMixer will indicate that it has finished running the simulation by updating the general information text to say Simulation finished, running time was X seconds. If an error occurs during the simulation, an error box will pop up to inform you of the error, in which case you can go back and edit the breeding scheme to fix the problem. If the simulation completed without any errors, you can now press the Inspect Progeny button (or select Inspect Progeny from the Simulation menu). A new window like the one below will appear. This window is the Progeny Inspector, where you can look at the genotype at each marker location for each individual. Note that although this view only allows you to inspect chromosomes at marker resolution, recombinations are handled internally at the base pair level. If the marker density is not high enough, there is the possibility of recombinations occurring but remaining undetected. A. The left side of this window is a list of all of the individuals from cells that were tagged as to be exported in the breeding scheme dialog. The list is ordered by ID, which is determined by the order in which the individuals were generated. Purebreds have IDs as well, but they are not exported, which is why the list does not necessarily start at ID #1.
12 GenomeMixer Documentation B. The Parents region in the top-right corner indicates which individuals were mated to produce this offspring. Each individual has a unique ID number, so it can be easily determined if two individuals have the same parents, for example. C. The two resize cells buttons automatically resize all of the columns. Note that you can also manually resize columns and rows (that is why the columns in the sample picture are different sizes). D. The inspection table itself, in the lower-right section of the window, gives you an overview of a selected individual s genotype. The genotype of whichever individual that was selected from the list at the left will appear in this section. Each group of three columns represents a chromosome; one column is labeled with the chromosome number (e.g. Chr 13 ). This column contains the names of all the markers. To the right of each marker is the genotype at that marker, from the chromosome inherited from the mother first (with the female symbol as the column header), and then from the chromosome inherited from the father (with the male symbol as the column header). Here, the Inspect Progeny window is displaying information for individual #829, which is a recombinant inbred male (evident in the homozygous but non-uniform genome). Using a higher-density marker map, we would find that the genome is not 100% homozygous, even though it is extremely close. This individual was created in the 24th generation, and its parents were #798 (female) and #799. Page 12 of 22
13 6. Importing / Exporting: Importing Markers in large numbers (from Excel, or text): GenomeMixer s built-in marker editor is suitable for editing small numbers of markers by hand and for automatically generating markers at evenly-spaced intervals. However, entering large numbers of markers by hand is tedious, and if you already have markers in another format, you will want to import them instead of re-entering them. GenomeMixer marker (.mar) files consist of the following: Regular text comments at the top of the file (warning: if you overwrite this marker file, GenomeMixer will not save the comments). <START MARKERS> (must be alone at the very start of a line) <MARKER> (tab) Name (tab) Chr_Number (tab) Loc_in_bases (tab) Loc_in_cM (newline*) <MARKER> (tab) Name (tab) Chr_Number (tab) Loc_in_bases (tab) Loc_in_cM (newline*) <MARKER> (tab) Name (tab) Chr_Number (tab) Loc_in_bases (tab) Loc_in_cM (newline*) (end of file: nothing special at the end of the file) *Newline: If you are on a Unix system, then newline is the default line ending. However, if you are on a Mac or Windows system, the default line ending contains a carriage return. The newline is represented by \n, and the carriage return is represented by \r. If you are on a Mac or Windows system, you will need to use a utility (such as BBEdit Lite or Josh Aas LineBreak for the Macintosh) to convert your linebreaks to Unix format, or GenomeMixer will not be able to read them. Chr_number should not be the number of the chromosome this marker is on, from 1 onwards. This should always be a number, and never X or Y. The X chromosome is considered the next-to-last chromosome, and Y is considered the last chromosome. So if you have a species with 19 autosomes plus X and Y, X should be chromosome 20, and Y should be chromosome 21. Loc_in_bases should be an integer. It is common for the location of a marker in bases to be up to several hundred million. If a chromosome is 150 megabases long, then the last marker could be located at You cannot use commas in the numbers (although you could type the locations out with commas and then do a search-and-replace to delete them all in the end), and the loc_in_bases cannot be a decimal. Two markers on the same chromosome should not share the same location in bases. Loc_in_cM, on the other hand, will frequently have a decimal point. The length of a chromosome in cm (centimorgans) could be 85.3, , or any other floating-point (or integer) number. This should probably be no greater than 1,000, and in most cases will probably be no greater than 100. If two markers on the same chromosome share the same cm value, then there is no chance of a recombination occurring between them.
14 None of these four parameters can be blank. Each marker must have a name, chromosome number, location in bases, and location in centimorgans. Location in centimorgans is the most important as far as recombination simulation is concerned; if you don t know a location in bases, you can just put in a reasonable estimation. Here is an example of a short, but completely valid, marker file: Some_Random_Marker_Comments_That_Won t_be_saved <START MARKERS> <MARKER> MarkerA <MARKER> TheMarker <MARKER> AloneMarker MarkerA is at the start of Chromosome 1; hence, it s loc_in_bases is zero, as is its loc_in_cm.. In most cases, there should be a marker at the very beginning (0 bases, 0 cm) and very end of every chromosome. Also, notice that in the above example, no recombinations will occur on chromosome 2, because only one marker (the AloneMarker ) is defined on chromosome 2, and in our model, recombinations only occur between markers. If the marker density is too low (< 5 per chromosome), you may get unrealistic recombination effects, because only one recombination can occur between markers (although it can be located anywhere between them unless the chromosomes are < 1 megabase in length, there is virtually no chance of one recombination occurring and then, in a later generation, a subsequent recombination occurring in exactly the same spot). Here is the step-by-step procedure for converting an Excel or text document into a GenomeMixer marker file: 1. Edit marker file in spreadsheet program or text editor. 2. Save in plain text format (not Excel format, for example) 3. Use a program like BBEdit to convert the linebreaks to Unix format. 4. Rename the file so that it has the extension.mar 5. Import the file into GenomeMixer by selecting Open and choosing this marker file. 6. When you go to Edit Markers / Breeding Scheme and select the Markers tab, verify that the imported markers look the way you expected them to. If the markers do not appear in step 6, or if there is only one marker, there are a number of possibilities. Perhaps <START MARKERS> is not at the beginning of its own line, so GenomeMixer just never read any markers in the file. Perhaps <MARKER> (tab) is not at the beginning of every line, or perhaps the file is not Unix-linebreak formatted (check the troubleshooting section for linebreak information). It is also possible that the file was saved in some other format that contains extra data (Excel.xls, for example), instead of plain tab-delimited text. Page 14 of 22
15 Exporting data for MapManager QTX: These are the steps required to make use of exported data in MapManager. After you run a simulation, you can save the exported progeny in MapManager QTX format by selecting the MapManager option from the output menu. A few steps are still required before MapManager can make use of the data. 1. Take note of the (n= ) part of the filename in the MapManager-format file you just saved. If the file is named mm_export_(n=23), then there are 23 individuals in the file. 2. Launch MapManager QTX, and from the File menu, select "Import -> Text..." A new window will appear, with the following data at the top: 3. Set the # Progeny to the same number as the n= in the filename. In our example, it is 23, because the sample filename was mm_export_(n=23). 4. After changing the # Progeny, you must click Apply. 5. After you click the apply button, click "OK," and proceed to a dialog that asks you to Enter a file to import. Select the correct text file, and click open. 6. Now, a dialog named Import Text will appear. You should set the options as they appear in the image below: 7. Once you have made sure the correct options on the left are selected, with 8 checked (Dataset: dataset, progeny names, name. Chr: chr, name. Locus: name, alias, geno) and 11 unchecked, plus tab between items, line feed between records, and none between genotypes, then you should click OK. MapManager should open up a new file with the imported information. Page 15 of 22
16 After you have successfully imported the data into MapManager, you can save it as a native MapManager file, so that you will not need to follow these steps again to open that file. Page 16 of 22
17 X. Troubleshooting After I edited a file with another program, GenomeMixer can no longer read it There several possibilities here: one is that the file is simply using the wrong linebreak format. Check Unix-style linebreaks, below, for a solution to that problem. If you edited the file using a word processor such as Microsoft Word, it probably saved a lot of hidden data in the file (formatting, margins, track changes, etc.), which is invisible within the word processor, but interferes with GenomeMixer s ability to read the file. You can tell if there is hidden data in a file by opening it in a plain text editor, like TextEdit or BBEdit for the Mac, or Notepad for Windows. If there are extra characters besides the bare GenomeMixer data, you will have to re-open the file with the word processor it was edited with, save it as plain text format, and then fix the linebreak format again (again, under Unix-style linebreaks ). If there are no strange characters, and the linebreaks are already Unix-format, then it is likely that an important line or character was inadvertently deleted, or there is an extra space somewhere (especially in the purebred names, which cannot have spaces). For example, in the breeding scheme table, all of the entries must be surrounded in doublequotes, and it is easy to leave these out (also, note that "straight quotes" must be used curly quotes will not be read correctly. If you use Word to edit a GenomeMixer file, you may have to temporarily disable the smart quotes feature). Unix-style Linebreaks (problems with Mac / Win GenomeMixer files) GenomeMixer can only read files that use Unix linebreaks, and although GenomeMixer automatically saves files in this format, if you used another Mac or Windows program to manually edit a file, it is likely that the editor changed the linebreak format. The solution to this problem is to use a utility that can convert linebreaks to Unix format: the freeware Mac OS X drag-and-drop program LineBreak by Josh Aas is one such program. BBEdit and BBEdit Lite also have the ability to convert linebreaks. Simply use one of these applications to change the linebreak format, save your changes, and reload the nowproperly-formatted file. Quickload doesn t work The Quickload menu option depends on two files, named quickmarkers.mar and quicksettings.gmx (case-sensitive), being located inside the Saved_Settings folder (case-sensitive, also note the underscore). The Saved_Settings folder must be located in the same folder as the GenomeMixer application. If Quickload gives an error, check to make sure that valid markers and settings file are in fact in the Saved_Settings folder, that the capitalization of file names is correct, and that you are running the correct copy of GenomeMixer, in case you have more than one copy. If everything seems ok, but the files still do not load, check to make sure that they have Unix-style linebreaks (see Unix-style linebreaks above). Page 17 of 22
18 Page 18 of 22 GenomeMixer Documentation
19 Appendix I: Specification of the GenomeMixer breeding scheme file: The GenomeMixer breeding scheme file (.gmx ) contains all of the information required to specify the type of organism (number of chromosomes, for example) and the breeding scheme used to produce output progeny. (Marker information is stored in a separate.mar marker file.) Several example breeding scheme files are included in the Saved_Settings folder in the default GenomeMixer distribution. Note that under normal circumstances, you will not want to edit the breeding scheme file directly, but if desired, Excel or another spreadsheet program can be used to edit it (provided that the file is saved in plain tab-delimited text format afterward). Here is a sample GenomeMixer beeding scheme (.gmx) file, with annotation: <NAME> This_is_a_configuration_file. The name of the file. This currently has no significance other than to save the name of a particular breeding scheme in a location separate from the filename, in case the file is renamed, for example. <VERSION> 1.01 <CHR> <CHR> <CHR> If there were multiple versions of GenomeMixer breeding scheme input, the version number would allow GenomeMixer to easily distinguish between them for backwards compatibility. Chr MB 107 cm Chr MB 104 cm Chr_My_Name 161 MB 85 cm The autosomes, given with an arbitrary name ( Chr01 ), a length in megabases, and a length in centimorgans. <FEMALE_SEX_CHR> <MALE_SEX_CHR> Chr20X 147 MB 74 cm Chr21Y 30 MB 15 cm The format is the same for the sex chromosomes, except that they are preceded by the tag <... _SEX_CHR> instead of simply <CHR>. <DEFAULT_LITTER_SIZE> 10 When a new cell is made to specify a mating, if the user does not explicitly specify a particular number of progeny from the mating, this default value will be used. <DEFAULT_LITTER_STDEV> 0 Standard deviation is not currently implemented, but it would allow some randomness to be incorporated into litter size determination. Page 19 of 22
20 <ABSOLUTE_INTERFERENCE> Specifies the recombination method used in this breeding scheme. In this case, around every breakpoint there is an absolute threshold of 25,000,000 bases in which no other breakpoint may occur. <PUREBRED> AA A <PUREBRED> BB B <PUREBRED> TwoTails C This is a list of the purebred ancestral strains from which matings can be specified. In this example, there are four distinct ancestral strains. The first parameter ( TwoTails ) is an arbitrary name, but it will appear frequently in the breeding scheme table, so it should be short and contain no spaces or special characters. The second parameter is a letter or number that is used to represent the purebred strain when exporting to other formats ( A and B are good choices for this letter, since some other programs expect A and B parental strains). Note that the letter is case-sensitive: A and a are different. At least two purebred strains will be needed to do any meaningful simulations (if there is only one purebred strain, all progeny will be clones of the purebred strain individuals, no matter how the cross is set up). <GENERATIONS 3> This is the header of the table of GENERATIONS x CAGES (for plant genetics, each cage may be thought of as plot. In the future, it should be possible to specify the terminology here). Note that there is a number inside the <GENERATIONS N> tag, which must be the same as the number of generation numbers (separated by tabs) to the right. So if 8 generations were desired, the line would read <GENERATIONS 8> <CAGE 1> "exp p(aa) x p(bb) r=0 m=1 f=1" "exp A1 x A1 r=0 m=1 f=1" "exp B1 x B3 r=0 m=1 f=1" <CAGE 2> "exp p(bb) x p(aa) r=0 m=1 f=1" "#exp A1 x A2 r=1 m=1 f=1" "" <CAGE 3> "" "A2 x A2 r=2 m=1 f=1" "" <CAGE 4> "" "#A1 x A1 r=5 m=0 f=0" "" Each cage / plot may either contain mating information (surrounded by doublequotes, and separated by tabs) or be blank, which is indicated by either a plain tab or by two consecutive double-quotes ("") with nothing in between. If you are going to edit the files manually, make sure you have plain quotes, as fancy curly quotes / smart quotes ( ) may cause confusion when importing. Cells are numbered down, starting from 1, and lettered across, starting from Page 20 of 22
21 A. So the top-left cell is A1, the cell to its right is B1, and the cell below A1 is A2. The number always indicates the cage, and the letter indications the generation. Z is the 26 th generation, and AA is the 27 th. AB is the 28 th, and so on. The cell immediately to the right of AZ99 would be cell BA99. This is the same format used by nearly every spreadsheet program. The format of the entries is as follows, where items in brackets [ ] are optional: [#] [exp] mother x father r=number m=number f=number A hash (#) indicates a temporarily-disabled cell. No progeny will be generated by any matings specified in a disabled cell. In this example, the second-generation cells for cage 2 and cage 4 are disabled (i.e., cells B2 and B4 are disabled). exp means export the progeny generated in this cell to the output file. If a cell does not have exp in it, the progeny from that mating will still be calculated, but those individuals will not be written to a file. This is useful if you only want the final results of a breeding scheme; the F1 generation, for example, might not contain the exp keyword, if those individuals were genetically uninteresting. r=number: specifies the number of progeny whose sex will be determined randomly. Any non-negative integer is a possible value. Note that r is completely independent of the m and f parameters. r=10 will result in 10 individuals of random (50/50 chance, for now) sex, whereas r=10 m=10 will result in 10 random individuals plus 10 males. m=number: specifies the number of male progeny to generate in this cell, in addition to any progeny of randomly-chosen sex. If this number is 0, then there is no guarantee that any male progeny will be generated in this cell, which could potentially be a problem if a cell in a later generation requests a male individual for a mating. f=number: specifies the number of female progeny to generate in this cell, in addition to any progeny of randomly-chosen sex. If this number is 0, then there is no guarantee that any female progeny will be generated in this cell, which could potentially be a problem if a cell in a later generation requests a female individual for a mating. The mother and father parameters can either be specified by requesting an individual from a previous cell ( A1 will get an individual of the appropriate sex from whatever progeny were produced in cell A1), or by requesting a purebred parental strain. p(name) means get the purebred strain with name NAME. In the example above, p(aa) x p(bb) obtains a female from the A purebred population, and a male from the B population. <END_DATA> This is simply an indication that there is no more breeding scheme information. Comments may be written below this line. Note that if a file is opened in GenomeMixer, edited, and saved again, then the comments will not be saved. Page 21 of 22
22 Page 22 of 22 GenomeMixer Documentation
QTX. Tutorial for. by Kim M.Chmielewicz Kenneth F. Manly. Software for genetic mapping of Mendelian markers and quantitative trait loci.
 Tutorial for QTX by Kim M.Chmielewicz Kenneth F. Manly Software for genetic mapping of Mendelian markers and quantitative trait loci. Available in versions for Mac OS and Microsoft Windows. revised for
Tutorial for QTX by Kim M.Chmielewicz Kenneth F. Manly Software for genetic mapping of Mendelian markers and quantitative trait loci. Available in versions for Mac OS and Microsoft Windows. revised for
RenameMan User Guide. ExtraBit Software
 RenameMan User Guide ExtraBit Software http://www.extrabit.com Version 3.1 August, 2005 Contents Introduction... 5 What does RenameMan do?... 5 Features... 5 Quick tour of RenameMan... 5 Getting started...
RenameMan User Guide ExtraBit Software http://www.extrabit.com Version 3.1 August, 2005 Contents Introduction... 5 What does RenameMan do?... 5 Features... 5 Quick tour of RenameMan... 5 Getting started...
Word: Print Address Labels Using Mail Merge
 Word: Print Address Labels Using Mail Merge No Typing! The Quick and Easy Way to Print Sheets of Address Labels Here at PC Knowledge for Seniors we re often asked how to print sticky address labels in
Word: Print Address Labels Using Mail Merge No Typing! The Quick and Easy Way to Print Sheets of Address Labels Here at PC Knowledge for Seniors we re often asked how to print sticky address labels in
COPYRIGHTED MATERIAL. Making Excel More Efficient
 Making Excel More Efficient If you find yourself spending a major part of your day working with Excel, you can make those chores go faster and so make your overall work life more productive by making Excel
Making Excel More Efficient If you find yourself spending a major part of your day working with Excel, you can make those chores go faster and so make your overall work life more productive by making Excel
Microsoft Excel Level 2
 Microsoft Excel Level 2 Table of Contents Chapter 1 Working with Excel Templates... 5 What is a Template?... 5 I. Opening a Template... 5 II. Using a Template... 5 III. Creating a Template... 6 Chapter
Microsoft Excel Level 2 Table of Contents Chapter 1 Working with Excel Templates... 5 What is a Template?... 5 I. Opening a Template... 5 II. Using a Template... 5 III. Creating a Template... 6 Chapter
How to import text files to Microsoft Excel 2016:
 How to import text files to Microsoft Excel 2016: You would use these directions if you get a delimited text file from a government agency (or some other source). This might be tab-delimited, comma-delimited
How to import text files to Microsoft Excel 2016: You would use these directions if you get a delimited text file from a government agency (or some other source). This might be tab-delimited, comma-delimited
All-In-One E-Sticker Installation and User Guide (Mac Versions)
 r6 All-In-One E-Sticker Installation and User Guide (Mac Versions) Getting Started with Your Mac All-In-One E-Sticker Using the All-In-One E-Sticker is a simple process. Before you begin, however, we highly
r6 All-In-One E-Sticker Installation and User Guide (Mac Versions) Getting Started with Your Mac All-In-One E-Sticker Using the All-In-One E-Sticker is a simple process. Before you begin, however, we highly
Gradekeeper Version 5.7
 Editor Irene Gardner Editorial Project Manager Paul Gardner Editor-in-Chief Sharon Coan, M.S. Ed. Imaging Ralph Olmedo, Jr. Production Manager Phil Garcia Macintosh is a registered trademark of Apple Computer,
Editor Irene Gardner Editorial Project Manager Paul Gardner Editor-in-Chief Sharon Coan, M.S. Ed. Imaging Ralph Olmedo, Jr. Production Manager Phil Garcia Macintosh is a registered trademark of Apple Computer,
Exhibitor Software User s Manual. Exhibitor Software V
 Exhibitor Software User s Manual Exhibitor Software V1.0.1 090908 1 Contents 1. Exhibitor Software 2. Installation 3. Using Exhibitor Program 3.1 Starting the Program 3.2 Logging in to the Program 3.3
Exhibitor Software User s Manual Exhibitor Software V1.0.1 090908 1 Contents 1. Exhibitor Software 2. Installation 3. Using Exhibitor Program 3.1 Starting the Program 3.2 Logging in to the Program 3.3
Genetic Analysis. Page 1
 Genetic Analysis Page 1 Genetic Analysis Objectives: 1) Set up Case-Control Association analysis and the Basic Genetics Workflow 2) Use JMP tools to interact with and explore results 3) Learn advanced
Genetic Analysis Page 1 Genetic Analysis Objectives: 1) Set up Case-Control Association analysis and the Basic Genetics Workflow 2) Use JMP tools to interact with and explore results 3) Learn advanced
Breeding Guide. Customer Services PHENOME-NETWORKS 4Ben Gurion Street, 74032, Nes-Ziona, Israel
 Breeding Guide Customer Services PHENOME-NETWORKS 4Ben Gurion Street, 74032, Nes-Ziona, Israel www.phenome-netwoks.com Contents PHENOME ONE - INTRODUCTION... 3 THE PHENOME ONE LAYOUT... 4 THE JOBS ICON...
Breeding Guide Customer Services PHENOME-NETWORKS 4Ben Gurion Street, 74032, Nes-Ziona, Israel www.phenome-netwoks.com Contents PHENOME ONE - INTRODUCTION... 3 THE PHENOME ONE LAYOUT... 4 THE JOBS ICON...
CROMWELLSTUDIOS. Content Management System Instruction Manual V1. Content Management System. V1
 Content Management System Instruction Manual V1 www.cromwellstudios.co.uk Cromwell Studios Web Services Content Management System Manual Part 1 Content Management is the system by which you can change
Content Management System Instruction Manual V1 www.cromwellstudios.co.uk Cromwell Studios Web Services Content Management System Manual Part 1 Content Management is the system by which you can change
Code::Blocks Student Manual
 Code::Blocks Student Manual Lawrence Goetz, Network Administrator Yedidyah Langsam, Professor and Theodore Raphan, Distinguished Professor Dept. of Computer and Information Science Brooklyn College of
Code::Blocks Student Manual Lawrence Goetz, Network Administrator Yedidyah Langsam, Professor and Theodore Raphan, Distinguished Professor Dept. of Computer and Information Science Brooklyn College of
Population Genetics (52642)
 Population Genetics (52642) Benny Yakir 1 Introduction In this course we will examine several topics that are related to population genetics. In each topic we will discuss briefly the biological background
Population Genetics (52642) Benny Yakir 1 Introduction In this course we will examine several topics that are related to population genetics. In each topic we will discuss briefly the biological background
Chapter 12 Creating Tables of Contents, Indexes and Bibliographies
 Writer Guide Chapter 12 Creating Tables of Contents, Indexes and Bibliographies OpenOffice.org Copyright This document is Copyright 2005 by its contributors as listed in the section titled Authors. You
Writer Guide Chapter 12 Creating Tables of Contents, Indexes and Bibliographies OpenOffice.org Copyright This document is Copyright 2005 by its contributors as listed in the section titled Authors. You
Site Owners: Cascade Basics. May 2017
 Site Owners: Cascade Basics May 2017 Page 2 Logging In & Your Site Logging In Open a browser and enter the following URL (or click this link): http://mordac.itcs.northwestern.edu/ OR http://www.northwestern.edu/cms/
Site Owners: Cascade Basics May 2017 Page 2 Logging In & Your Site Logging In Open a browser and enter the following URL (or click this link): http://mordac.itcs.northwestern.edu/ OR http://www.northwestern.edu/cms/
Function. Description
 Function Check In Get / Checkout Description Checking in a file uploads the file from the user s hard drive into the vault and creates a new file version with any changes to the file that have been saved.
Function Check In Get / Checkout Description Checking in a file uploads the file from the user s hard drive into the vault and creates a new file version with any changes to the file that have been saved.
For many people, learning any new computer software can be an anxietyproducing
 1 Getting to Know Stata 12 For many people, learning any new computer software can be an anxietyproducing task. When that computer program involves statistics, the stress level generally increases exponentially.
1 Getting to Know Stata 12 For many people, learning any new computer software can be an anxietyproducing task. When that computer program involves statistics, the stress level generally increases exponentially.
Convert Dosages to Genotypes Author: Autumn Laughbaum, Golden Helix, Inc.
 Convert Dosages to Genotypes Author: Autumn Laughbaum, Golden Helix, Inc. Overview This script converts allelic dosage values to genotypes based on user-specified thresholds. The dosage data may be in
Convert Dosages to Genotypes Author: Autumn Laughbaum, Golden Helix, Inc. Overview This script converts allelic dosage values to genotypes based on user-specified thresholds. The dosage data may be in
SQL Server. Management Studio. Chapter 3. In This Chapter. Management Studio. c Introduction to SQL Server
 Chapter 3 SQL Server Management Studio In This Chapter c Introduction to SQL Server Management Studio c Using SQL Server Management Studio with the Database Engine c Authoring Activities Using SQL Server
Chapter 3 SQL Server Management Studio In This Chapter c Introduction to SQL Server Management Studio c Using SQL Server Management Studio with the Database Engine c Authoring Activities Using SQL Server
INTViewer Tutorial Cube Tutorial
 INTViewer Tutorial Cube Tutorial This tutorial shows how to use INTViewer to display a seismic cube stored in a Seismic file. Windows created will include INLINE, XLINE, Time Slice and an arbitrary traverse,
INTViewer Tutorial Cube Tutorial This tutorial shows how to use INTViewer to display a seismic cube stored in a Seismic file. Windows created will include INLINE, XLINE, Time Slice and an arbitrary traverse,
Using Microsoft Excel
 Using Microsoft Excel Excel contains numerous tools that are intended to meet a wide range of requirements. Some of the more specialised tools are useful to people in certain situations while others have
Using Microsoft Excel Excel contains numerous tools that are intended to meet a wide range of requirements. Some of the more specialised tools are useful to people in certain situations while others have
Working with Mailbox Manager
 Working with Mailbox Manager A user guide for Mailbox Manager supporting the Message Storage Server component of the Avaya S3400 Message Server Mailbox Manager Version 5.0 February 2003 Copyright 2003
Working with Mailbox Manager A user guide for Mailbox Manager supporting the Message Storage Server component of the Avaya S3400 Message Server Mailbox Manager Version 5.0 February 2003 Copyright 2003
PIXEL SPREADSHEET USER MANUAL April 2011
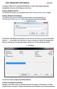 PIXEL SPREADSHEET USER MANUAL April 2011 Greetings! Thank you for using Pixel Spreadsheet. In order to get started using Pixel Spreadsheet, there are a few things you must know: RUNNING PROGRAM FOR MAC
PIXEL SPREADSHEET USER MANUAL April 2011 Greetings! Thank you for using Pixel Spreadsheet. In order to get started using Pixel Spreadsheet, there are a few things you must know: RUNNING PROGRAM FOR MAC
Writer Guide. Chapter 12 Tables of Contents, Indexes, and Bibliographies
 Writer Guide Chapter 12 Tables of Contents, Indexes, and Bibliographies Copyright This document is Copyright 2012-2014 by the LibreOffice Documentation Team. Contributors are listed below. You may distribute
Writer Guide Chapter 12 Tables of Contents, Indexes, and Bibliographies Copyright This document is Copyright 2012-2014 by the LibreOffice Documentation Team. Contributors are listed below. You may distribute
UPGRADE AND CONVERT INSTRUCTION Not available on the ipad. This step must be done on your computer and then transfer your files to ipad.
 UNIFORM MANAGEMENT ASSIGNMENT CENTER II Welcome to Uniform Management Assignment Center. UMAC II is compatible with Windows, Mac OSX, and ipad. Follow the step-by-step instructions to assign your students
UNIFORM MANAGEMENT ASSIGNMENT CENTER II Welcome to Uniform Management Assignment Center. UMAC II is compatible with Windows, Mac OSX, and ipad. Follow the step-by-step instructions to assign your students
Chapter One: Getting Started With IBM SPSS for Windows
 Chapter One: Getting Started With IBM SPSS for Windows Using Windows The Windows start-up screen should look something like Figure 1-1. Several standard desktop icons will always appear on start up. Note
Chapter One: Getting Started With IBM SPSS for Windows Using Windows The Windows start-up screen should look something like Figure 1-1. Several standard desktop icons will always appear on start up. Note
Activity: page 1/10 Introduction to Excel. Getting Started
 Activity: page 1/10 Introduction to Excel Excel is a computer spreadsheet program. Spreadsheets are convenient to use for entering and analyzing data. Although Excel has many capabilities for analyzing
Activity: page 1/10 Introduction to Excel Excel is a computer spreadsheet program. Spreadsheets are convenient to use for entering and analyzing data. Although Excel has many capabilities for analyzing
Importing source database objects from a database
 Importing source database objects from a database We are now at the point where we can finally import our source database objects, source database objects. We ll walk through the process of importing from
Importing source database objects from a database We are now at the point where we can finally import our source database objects, source database objects. We ll walk through the process of importing from
Table Basics. The structure of an table
 TABLE -FRAMESET Table Basics A table is a grid of rows and columns that intersect to form cells. Two different types of cells exist: Table cell that contains data, is created with the A cell that
TABLE -FRAMESET Table Basics A table is a grid of rows and columns that intersect to form cells. Two different types of cells exist: Table cell that contains data, is created with the A cell that
Lab: Supplying Inputs to Programs
 Steven Zeil May 25, 2013 Contents 1 Running the Program 2 2 Supplying Standard Input 4 3 Command Line Parameters 4 1 In this lab, we will look at some of the different ways that basic I/O information can
Steven Zeil May 25, 2013 Contents 1 Running the Program 2 2 Supplying Standard Input 4 3 Command Line Parameters 4 1 In this lab, we will look at some of the different ways that basic I/O information can
ADOBE DREAMWEAVER CS4 BASICS
 ADOBE DREAMWEAVER CS4 BASICS Dreamweaver CS4 2 This tutorial focuses on the basic steps involved in creating an attractive, functional website. In using this tutorial you will learn to design a site layout,
ADOBE DREAMWEAVER CS4 BASICS Dreamweaver CS4 2 This tutorial focuses on the basic steps involved in creating an attractive, functional website. In using this tutorial you will learn to design a site layout,
Chapter 2 The Design Window
 Chapter 2 Objectives Chapter 2 The Design Window Learn about Crystal sections Move objects Use Toolbars, Icons, and Menus Format fields Add Special Fields Change a Group Use the Crystal Field Explorer
Chapter 2 Objectives Chapter 2 The Design Window Learn about Crystal sections Move objects Use Toolbars, Icons, and Menus Format fields Add Special Fields Change a Group Use the Crystal Field Explorer
The first thing we ll need is some numbers. I m going to use the set of times and drug concentration levels in a patient s bloodstream given below.
 Graphing in Excel featuring Excel 2007 1 A spreadsheet can be a powerful tool for analyzing and graphing data, but it works completely differently from the graphing calculator that you re used to. If you
Graphing in Excel featuring Excel 2007 1 A spreadsheet can be a powerful tool for analyzing and graphing data, but it works completely differently from the graphing calculator that you re used to. If you
Introduction to SPSS
 Introduction to SPSS Purpose The purpose of this assignment is to introduce you to SPSS, the most commonly used statistical package in the social sciences. You will create a new data file and calculate
Introduction to SPSS Purpose The purpose of this assignment is to introduce you to SPSS, the most commonly used statistical package in the social sciences. You will create a new data file and calculate
Performer to DP2 Hot Folder Reference Manual Rev There is only one file involved with installing the Performer to DP2 Hot Folder.
 Performer to DP2 Hot Folder Reference Manual Rev. 07.11.05 Install Files: There is only one file involved with installing the Performer to DP2 Hot Folder. The installer file is named PP2DP2_1.x.x.EXE.
Performer to DP2 Hot Folder Reference Manual Rev. 07.11.05 Install Files: There is only one file involved with installing the Performer to DP2 Hot Folder. The installer file is named PP2DP2_1.x.x.EXE.
TUTORIAL FOR IMPORTING OTTAWA FIRE HYDRANT PARKING VIOLATION DATA INTO MYSQL
 TUTORIAL FOR IMPORTING OTTAWA FIRE HYDRANT PARKING VIOLATION DATA INTO MYSQL We have spent the first part of the course learning Excel: importing files, cleaning, sorting, filtering, pivot tables and exporting
TUTORIAL FOR IMPORTING OTTAWA FIRE HYDRANT PARKING VIOLATION DATA INTO MYSQL We have spent the first part of the course learning Excel: importing files, cleaning, sorting, filtering, pivot tables and exporting
Microsoft Excel 2007 Lesson 7: Charts and Comments
 Microsoft Excel 2007 Lesson 7: Charts and Comments Open Example.xlsx if it is not already open. Click on the Example 3 tab to see the worksheet for this lesson. This is essentially the same worksheet that
Microsoft Excel 2007 Lesson 7: Charts and Comments Open Example.xlsx if it is not already open. Click on the Example 3 tab to see the worksheet for this lesson. This is essentially the same worksheet that
Taskbar: Working with Several Windows at Once
 Taskbar: Working with Several Windows at Once Your Best Friend at the Bottom of the Screen How to Make the Most of Your Taskbar The taskbar is the wide bar that stretches across the bottom of your screen,
Taskbar: Working with Several Windows at Once Your Best Friend at the Bottom of the Screen How to Make the Most of Your Taskbar The taskbar is the wide bar that stretches across the bottom of your screen,
SOLOMON: Parentage Analysis 1. Corresponding author: Mark Christie
 SOLOMON: Parentage Analysis 1 Corresponding author: Mark Christie christim@science.oregonstate.edu SOLOMON: Parentage Analysis 2 Table of Contents: Installing SOLOMON on Windows/Linux Pg. 3 Installing
SOLOMON: Parentage Analysis 1 Corresponding author: Mark Christie christim@science.oregonstate.edu SOLOMON: Parentage Analysis 2 Table of Contents: Installing SOLOMON on Windows/Linux Pg. 3 Installing
Individual Animal Records (IAR)
 Page 1 Chapter Contents Introduction 3 Menu Options 4 Getting Started in IAR 4 Customising PAM for IAR 5 Setting up a List of Ear Tags (Visual IDs) 6 Assigning Visual IDs (Tags) to Animals 9 Assigning
Page 1 Chapter Contents Introduction 3 Menu Options 4 Getting Started in IAR 4 Customising PAM for IAR 5 Setting up a List of Ear Tags (Visual IDs) 6 Assigning Visual IDs (Tags) to Animals 9 Assigning
Importing and Merging Data Tutorial
 Importing and Merging Data Tutorial Release 1.0 Golden Helix, Inc. February 17, 2012 Contents 1. Overview 2 2. Import Pedigree Data 4 3. Import Phenotypic Data 6 4. Import Genetic Data 8 5. Import and
Importing and Merging Data Tutorial Release 1.0 Golden Helix, Inc. February 17, 2012 Contents 1. Overview 2 2. Import Pedigree Data 4 3. Import Phenotypic Data 6 4. Import Genetic Data 8 5. Import and
Age & Stage Structure: Elephant Model
 POPULATION MODELS Age & Stage Structure: Elephant Model Terri Donovan recorded: January, 2010 Today we're going to be building an age-structured model for the elephant population. And this will be the
POPULATION MODELS Age & Stage Structure: Elephant Model Terri Donovan recorded: January, 2010 Today we're going to be building an age-structured model for the elephant population. And this will be the
Burning CDs in Windows XP
 B 770 / 1 Make CD Burning a Breeze with Windows XP's Built-in Tools If your PC is equipped with a rewritable CD drive you ve almost certainly got some specialised software for copying files to CDs. If
B 770 / 1 Make CD Burning a Breeze with Windows XP's Built-in Tools If your PC is equipped with a rewritable CD drive you ve almost certainly got some specialised software for copying files to CDs. If
Numbers Basics Website:
 Website: http://etc.usf.edu/te/ Numbers is Apple's new spreadsheet application. It is installed as part of the iwork suite, which also includes the word processing program Pages and the presentation program
Website: http://etc.usf.edu/te/ Numbers is Apple's new spreadsheet application. It is installed as part of the iwork suite, which also includes the word processing program Pages and the presentation program
6. Essential Spreadsheet Operations
 6. Essential Spreadsheet Operations 6.1 Working with Worksheets When you open a new workbook in Excel, the workbook has a designated number of worksheets in it. You can specify how many sheets each new
6. Essential Spreadsheet Operations 6.1 Working with Worksheets When you open a new workbook in Excel, the workbook has a designated number of worksheets in it. You can specify how many sheets each new
How to Edit Your Website
 How to Edit Your Website A guide to using your Content Management System Overview 2 Accessing the CMS 2 Choosing Your Language 2 Resetting Your Password 3 Sites 4 Favorites 4 Pages 5 Creating Pages 5 Managing
How to Edit Your Website A guide to using your Content Management System Overview 2 Accessing the CMS 2 Choosing Your Language 2 Resetting Your Password 3 Sites 4 Favorites 4 Pages 5 Creating Pages 5 Managing
Introduction to Microsoft Excel 2007
 Introduction to Microsoft Excel 2007 Microsoft Excel is a very powerful tool for you to use for numeric computations and analysis. Excel can also function as a simple database but that is another class.
Introduction to Microsoft Excel 2007 Microsoft Excel is a very powerful tool for you to use for numeric computations and analysis. Excel can also function as a simple database but that is another class.
Rescuing Lost Files from CDs and DVDs
 Rescuing Lost Files from CDs and DVDs R 200 / 1 Damaged CD? No Problem Let this Clever Software Recover Your Files! CDs and DVDs are among the most reliable types of computer disk to use for storing your
Rescuing Lost Files from CDs and DVDs R 200 / 1 Damaged CD? No Problem Let this Clever Software Recover Your Files! CDs and DVDs are among the most reliable types of computer disk to use for storing your
DAQFactory U3 Tutorial Getting Started with DAQFactory-Express and your LabJack U3 11/3/06
 DAQFactory U3 Tutorial Getting Started with DAQFactory-Express and your LabJack U3 11/3/06 Congratulations on the purchase of your new LabJack U3. Included on your installation CD is a fully licensed copy
DAQFactory U3 Tutorial Getting Started with DAQFactory-Express and your LabJack U3 11/3/06 Congratulations on the purchase of your new LabJack U3. Included on your installation CD is a fully licensed copy
Word for Research Writing I: Text and Structure
 Word for Research Writing I: Text and Structure Last updated: 10/2017 Shari Hill Sweet dteditor@nd.edu or 631-7545 1. The Graduate School Template...1 1.1 Document structure... 1 1.1.1 Beware of Section
Word for Research Writing I: Text and Structure Last updated: 10/2017 Shari Hill Sweet dteditor@nd.edu or 631-7545 1. The Graduate School Template...1 1.1 Document structure... 1 1.1.1 Beware of Section
My First Cocoa Program
 My First Cocoa Program 1. Tutorial Overview In this tutorial, you re going to create a very simple Cocoa application for the Mac. Unlike a line-command program, a Cocoa program uses a graphical window
My First Cocoa Program 1. Tutorial Overview In this tutorial, you re going to create a very simple Cocoa application for the Mac. Unlike a line-command program, a Cocoa program uses a graphical window
How to Make Graphs with Excel 2007
 Appendix A How to Make Graphs with Excel 2007 A.1 Introduction This is a quick-and-dirty tutorial to teach you the basics of graph creation and formatting in Microsoft Excel. Many of the tasks that you
Appendix A How to Make Graphs with Excel 2007 A.1 Introduction This is a quick-and-dirty tutorial to teach you the basics of graph creation and formatting in Microsoft Excel. Many of the tasks that you
Chapter 10 Linking Calc Data
 Calc Guide Chapter 10 Linking Calc Data Sharing data in and out of Calc This PDF is designed to be read onscreen, two pages at a time. If you want to print a copy, your PDF viewer should have an option
Calc Guide Chapter 10 Linking Calc Data Sharing data in and out of Calc This PDF is designed to be read onscreen, two pages at a time. If you want to print a copy, your PDF viewer should have an option
User Guide. Kronodoc Kronodoc Oy. Intelligent methods for process improvement and project execution
 User Guide Kronodoc 3.0 Intelligent methods for process improvement and project execution 2003 Kronodoc Oy 2 Table of Contents 1 User Guide 5 2 Information Structure in Kronodoc 6 3 Entering and Exiting
User Guide Kronodoc 3.0 Intelligent methods for process improvement and project execution 2003 Kronodoc Oy 2 Table of Contents 1 User Guide 5 2 Information Structure in Kronodoc 6 3 Entering and Exiting
This Tutorial is for Word 2007 but 2003 instructions are included in [brackets] after of each step.
![This Tutorial is for Word 2007 but 2003 instructions are included in [brackets] after of each step. This Tutorial is for Word 2007 but 2003 instructions are included in [brackets] after of each step.](/thumbs/72/67280806.jpg) This Tutorial is for Word 2007 but 2003 instructions are included in [brackets] after of each step. Table of Contents Just so you know: Things You Can t Do with Word... 1 Get Organized... 1 Create the
This Tutorial is for Word 2007 but 2003 instructions are included in [brackets] after of each step. Table of Contents Just so you know: Things You Can t Do with Word... 1 Get Organized... 1 Create the
Export a PROTECT Report to Excel (User s Guide Lesson 21 : Reports) Tips for Using Microsoft Excel and Exported Reports
 Export a PROTECT Report to Excel (User s Guide Lesson 21 : Reports) 1. Run a PROTECT Export report 2. Click the Export button which is at far left on the toolbar in the PROTECT report window. 3. The Export
Export a PROTECT Report to Excel (User s Guide Lesson 21 : Reports) 1. Run a PROTECT Export report 2. Click the Export button which is at far left on the toolbar in the PROTECT report window. 3. The Export
Chapter 6: Creating and Configuring Menus. Using the Menu Manager
 Chapter 6: Creating and Configuring Menus The Menu Manager provides key information about each menu, including: Title. The name of the menu. Type. Its unique name used in programming. Menu Item. A link
Chapter 6: Creating and Configuring Menus The Menu Manager provides key information about each menu, including: Title. The name of the menu. Type. Its unique name used in programming. Menu Item. A link
Linkage analysis with paramlink Session I: Introduction and pedigree drawing
 Linkage analysis with paramlink Session I: Introduction and pedigree drawing In this session we will introduce R, and in particular the package paramlink. This package provides a complete environment for
Linkage analysis with paramlink Session I: Introduction and pedigree drawing In this session we will introduce R, and in particular the package paramlink. This package provides a complete environment for
Statistical Analysis for Genetic Epidemiology (S.A.G.E.) Version 6.4 Graphical User Interface (GUI) Manual
 Statistical Analysis for Genetic Epidemiology (S.A.G.E.) Version 6.4 Graphical User Interface (GUI) Manual Department of Epidemiology and Biostatistics Wolstein Research Building 2103 Cornell Rd Case Western
Statistical Analysis for Genetic Epidemiology (S.A.G.E.) Version 6.4 Graphical User Interface (GUI) Manual Department of Epidemiology and Biostatistics Wolstein Research Building 2103 Cornell Rd Case Western
Writer Guide. Chapter 13 Working with Master Documents
 Writer Guide Chapter 13 Working with Master Documents Copyright This document is Copyright 2011 by its contributors as listed below. You may distribute it and/or modify it under the terms of either the
Writer Guide Chapter 13 Working with Master Documents Copyright This document is Copyright 2011 by its contributors as listed below. You may distribute it and/or modify it under the terms of either the
This Tutorial is for Word 2007 but 2003 instructions are included in [brackets] after of each step.
![This Tutorial is for Word 2007 but 2003 instructions are included in [brackets] after of each step. This Tutorial is for Word 2007 but 2003 instructions are included in [brackets] after of each step.](/thumbs/89/98497436.jpg) This Tutorial is for Word 2007 but 2003 instructions are included in [brackets] after of each step. Table of Contents Get Organized... 1 Create the Home Page... 1 Save the Home Page as a Word Document...
This Tutorial is for Word 2007 but 2003 instructions are included in [brackets] after of each step. Table of Contents Get Organized... 1 Create the Home Page... 1 Save the Home Page as a Word Document...
The main differences with other open source reporting solutions such as JasperReports or mondrian are:
 WYSIWYG Reporting Including Introduction: Content at a glance. Create A New Report: Steps to start the creation of a new report. Manage Data Blocks: Add, edit or remove data blocks in a report. General
WYSIWYG Reporting Including Introduction: Content at a glance. Create A New Report: Steps to start the creation of a new report. Manage Data Blocks: Add, edit or remove data blocks in a report. General
DOING MORE WITH EXCEL: MICROSOFT OFFICE 2010
 DOING MORE WITH EXCEL: MICROSOFT OFFICE 2010 GETTING STARTED PAGE 02 Prerequisites What You Will Learn MORE TASKS IN MICROSOFT EXCEL PAGE 03 Cutting, Copying, and Pasting Data Filling Data Across Columns
DOING MORE WITH EXCEL: MICROSOFT OFFICE 2010 GETTING STARTED PAGE 02 Prerequisites What You Will Learn MORE TASKS IN MICROSOFT EXCEL PAGE 03 Cutting, Copying, and Pasting Data Filling Data Across Columns
Joomla! 2.5.x Training Manual
 Joomla! 2.5.x Training Manual 1 Joomla is an online content management system that keeps track of all content on your website including text, images, links, and documents. This manual includes several
Joomla! 2.5.x Training Manual 1 Joomla is an online content management system that keeps track of all content on your website including text, images, links, and documents. This manual includes several
Barchard Introduction to SPSS Marks
 Barchard Introduction to SPSS 22.0 3 Marks Purpose The purpose of this assignment is to introduce you to SPSS, the most commonly used statistical package in the social sciences. You will create a new data
Barchard Introduction to SPSS 22.0 3 Marks Purpose The purpose of this assignment is to introduce you to SPSS, the most commonly used statistical package in the social sciences. You will create a new data
Writer Guide. Chapter 12 Tables of Contents, Indexes, and Bibliographies
 Writer Guide Chapter 12 Tables of Contents, Indexes, and Bibliographies Copyright This document is Copyright 2011 2012 by its contributors as listed below. You may distribute it and/or modify it under
Writer Guide Chapter 12 Tables of Contents, Indexes, and Bibliographies Copyright This document is Copyright 2011 2012 by its contributors as listed below. You may distribute it and/or modify it under
Here are the steps in downloading the HTML code for signatures:
 I. INTRODUCTION This is a guide on how you download and then install the BBB dynamic seal into your email signature. Note that the code for this type of seal is very modified to work in email and not use
I. INTRODUCTION This is a guide on how you download and then install the BBB dynamic seal into your email signature. Note that the code for this type of seal is very modified to work in email and not use
4D Write. User Reference Mac OS and Windows Versions. 4D Write D SA/4D, Inc. All Rights reserved.
 4D Write User Reference Mac OS and Windows Versions 4D Write 1999-2002 4D SA/4D, Inc. All Rights reserved. 4D Write User Reference Version 6.8 for Mac OS and Windows Copyright 1999 2002 4D SA/4D, Inc.
4D Write User Reference Mac OS and Windows Versions 4D Write 1999-2002 4D SA/4D, Inc. All Rights reserved. 4D Write User Reference Version 6.8 for Mac OS and Windows Copyright 1999 2002 4D SA/4D, Inc.
How to Edit Your Website
 How to Edit Your Website A guide to using your Content Management System Overview 2 Accessing the CMS 2 Choosing Your Language 2 Resetting Your Password 3 Sites 4 Favorites 4 Pages 5 Creating Pages 5 Managing
How to Edit Your Website A guide to using your Content Management System Overview 2 Accessing the CMS 2 Choosing Your Language 2 Resetting Your Password 3 Sites 4 Favorites 4 Pages 5 Creating Pages 5 Managing
Creating and Using Genome Assemblies Tutorial
 Creating and Using Genome Assemblies Tutorial Release 8.1 Golden Helix, Inc. March 18, 2014 Contents 1. Create a Genome Assembly for Danio rerio 2 2. Building Annotation Sources 5 A. Creating a Reference
Creating and Using Genome Assemblies Tutorial Release 8.1 Golden Helix, Inc. March 18, 2014 Contents 1. Create a Genome Assembly for Danio rerio 2 2. Building Annotation Sources 5 A. Creating a Reference
Dreamweaver CS 5.5. University Information Technology Services. Training, Outreach, Learning Technologies, and Video Production
 Dreamweaver CS 5.5 Creating Web Pages with a Template University Information Technology Services Training, Outreach, Learning Technologies, and Video Production Copyright 2012 KSU Department of Information
Dreamweaver CS 5.5 Creating Web Pages with a Template University Information Technology Services Training, Outreach, Learning Technologies, and Video Production Copyright 2012 KSU Department of Information
1.0 Overview For content management, Joomla divides into some basic components: the Article
 Joomla! 3.4.x Training Manual Joomla is an online content management system that keeps track of all content on your website including text, images, links, and documents. This manual includes several tutorials
Joomla! 3.4.x Training Manual Joomla is an online content management system that keeps track of all content on your website including text, images, links, and documents. This manual includes several tutorials
Parish . User Manual
 Parish Email User Manual Table of Contents LOGGING IN TO PARISH EMAIL... 3 GETTING STARTED... 3 GENERAL OVERVIEW OF THE USER INTERFACE... 3 TERMINATE THE SESSION... 4 EMAIL... 4 MESSAGES LIST... 4 Open
Parish Email User Manual Table of Contents LOGGING IN TO PARISH EMAIL... 3 GETTING STARTED... 3 GENERAL OVERVIEW OF THE USER INTERFACE... 3 TERMINATE THE SESSION... 4 EMAIL... 4 MESSAGES LIST... 4 Open
OPENING A LEADS.TXT FILE IN EXCEL 2010
 From the project manager to a team of administrative, programming, and technical specialists, ATS believes in a team approach that brings together all of the necessary elements pre-conference and onsite
From the project manager to a team of administrative, programming, and technical specialists, ATS believes in a team approach that brings together all of the necessary elements pre-conference and onsite
Step-by-Step Guide to Basic Genetic Analysis
 Step-by-Step Guide to Basic Genetic Analysis Page 1 Introduction This document shows you how to clean up your genetic data, assess its statistical properties and perform simple analyses such as case-control
Step-by-Step Guide to Basic Genetic Analysis Page 1 Introduction This document shows you how to clean up your genetic data, assess its statistical properties and perform simple analyses such as case-control
DataPro Quick Start Guide
 DataPro Quick Start Guide Introduction The DataPro application provides the user with the ability to download and analyze data acquired using the ULTRA-LITE PRO range of Auto Meter products. Please see
DataPro Quick Start Guide Introduction The DataPro application provides the user with the ability to download and analyze data acquired using the ULTRA-LITE PRO range of Auto Meter products. Please see
Logger Pro 3. Quick Reference
 Logger Pro 3 Quick Reference Getting Started Logger Pro Requirements To use Logger Pro, you must have the following equipment: Windows 98, 2000, ME, NT, or XP on a Pentium processor or equivalent, 133
Logger Pro 3 Quick Reference Getting Started Logger Pro Requirements To use Logger Pro, you must have the following equipment: Windows 98, 2000, ME, NT, or XP on a Pentium processor or equivalent, 133
1 Introduction to Using Excel Spreadsheets
 Survey of Math: Excel Spreadsheet Guide (for Excel 2007) Page 1 of 6 1 Introduction to Using Excel Spreadsheets This section of the guide is based on the file (a faux grade sheet created for messing with)
Survey of Math: Excel Spreadsheet Guide (for Excel 2007) Page 1 of 6 1 Introduction to Using Excel Spreadsheets This section of the guide is based on the file (a faux grade sheet created for messing with)
MICROSOFT OFFICE. Courseware: Exam: Sample Only EXCEL 2016 CORE. Certification Guide
 MICROSOFT OFFICE Courseware: 3263 2 Exam: 77 727 EXCEL 2016 CORE Certification Guide Microsoft Office Specialist 2016 Series Microsoft Excel 2016 Core Certification Guide Lesson 1: Introducing Excel Lesson
MICROSOFT OFFICE Courseware: 3263 2 Exam: 77 727 EXCEL 2016 CORE Certification Guide Microsoft Office Specialist 2016 Series Microsoft Excel 2016 Core Certification Guide Lesson 1: Introducing Excel Lesson
BICF Nano Course: GWAS GWAS Workflow Development using PLINK. Julia Kozlitina April 28, 2017
 BICF Nano Course: GWAS GWAS Workflow Development using PLINK Julia Kozlitina Julia.Kozlitina@UTSouthwestern.edu April 28, 2017 Getting started Open the Terminal (Search -> Applications -> Terminal), and
BICF Nano Course: GWAS GWAS Workflow Development using PLINK Julia Kozlitina Julia.Kozlitina@UTSouthwestern.edu April 28, 2017 Getting started Open the Terminal (Search -> Applications -> Terminal), and
Exercise 1: Introduction to MapInfo
 Geog 578 Exercise 1: Introduction to MapInfo Page: 1/22 Geog 578: GIS Applications Exercise 1: Introduction to MapInfo Assigned on January 25 th, 2006 Due on February 1 st, 2006 Total Points: 10 0. Convention
Geog 578 Exercise 1: Introduction to MapInfo Page: 1/22 Geog 578: GIS Applications Exercise 1: Introduction to MapInfo Assigned on January 25 th, 2006 Due on February 1 st, 2006 Total Points: 10 0. Convention
Dreamweaver is a full-featured Web application
 Create a Dreamweaver Site Dreamweaver is a full-featured Web application development tool. Dreamweaver s features not only assist you with creating and editing Web pages, but also with managing and maintaining
Create a Dreamweaver Site Dreamweaver is a full-featured Web application development tool. Dreamweaver s features not only assist you with creating and editing Web pages, but also with managing and maintaining
Language Editor User Manual
 Language Editor User Manual June 2010 Contents Introduction... 3 Install the Language Editor... 4 Start using the Language Editor... 6 Editor screen... 8 Section 1: Translating Text... 9 Load Translations...
Language Editor User Manual June 2010 Contents Introduction... 3 Install the Language Editor... 4 Start using the Language Editor... 6 Editor screen... 8 Section 1: Translating Text... 9 Load Translations...
1 Interface Fundamentals
 1 Interface Fundamentals Windows The Media Composer interface is focused on three primary windows: the Composer, the Timeline and the Project. The Composer window contains the source and record monitors
1 Interface Fundamentals Windows The Media Composer interface is focused on three primary windows: the Composer, the Timeline and the Project. The Composer window contains the source and record monitors
Writer 5.4 Guide. Chapter 15 Tables of Contents, Indexes, Bibliographies
 Writer 5.4 Guide Chapter 15 Tables of Contents, Indexes, Bibliographies Copyright This document is Copyright 2017 by the LibreOffice Documentation Team. Contributors are listed below. You may distribute
Writer 5.4 Guide Chapter 15 Tables of Contents, Indexes, Bibliographies Copyright This document is Copyright 2017 by the LibreOffice Documentation Team. Contributors are listed below. You may distribute
My First iphone App. 1. Tutorial Overview
 My First iphone App 1. Tutorial Overview In this tutorial, you re going to create a very simple application on the iphone or ipod Touch. It has a text field, a label, and a button. You can type your name
My First iphone App 1. Tutorial Overview In this tutorial, you re going to create a very simple application on the iphone or ipod Touch. It has a text field, a label, and a button. You can type your name
Content Author's Reference and Cookbook
 Sitecore CMS 7.2 Content Author's Reference and Cookbook Rev. 140225 Sitecore CMS 7.2 Content Author's Reference and Cookbook A Conceptual Overview and Practical Guide to Using Sitecore Table of Contents
Sitecore CMS 7.2 Content Author's Reference and Cookbook Rev. 140225 Sitecore CMS 7.2 Content Author's Reference and Cookbook A Conceptual Overview and Practical Guide to Using Sitecore Table of Contents
Survey of Math: Excel Spreadsheet Guide (for Excel 2016) Page 1 of 9
 Survey of Math: Excel Spreadsheet Guide (for Excel 2016) Page 1 of 9 Contents 1 Introduction to Using Excel Spreadsheets 2 1.1 A Serious Note About Data Security.................................... 2 1.2
Survey of Math: Excel Spreadsheet Guide (for Excel 2016) Page 1 of 9 Contents 1 Introduction to Using Excel Spreadsheets 2 1.1 A Serious Note About Data Security.................................... 2 1.2
Printing Envelopes in Microsoft Word
 Printing Envelopes in Microsoft Word P 730 / 1 Stop Addressing Envelopes by Hand Let Word Print Them for You! One of the most common uses of Microsoft Word is for writing letters. With very little effort
Printing Envelopes in Microsoft Word P 730 / 1 Stop Addressing Envelopes by Hand Let Word Print Them for You! One of the most common uses of Microsoft Word is for writing letters. With very little effort
EXCEL 98 TUTORIAL Chemistry C2407 fall 1998 Andy Eng, Columbia University 1998
 Created on 09/02/98 11:58 PM 1 EXCEL 98 TUTORIAL Chemistry C2407 fall 1998 Andy Eng, Columbia University 1998 Note for Excel 97 users: All features of Excel 98 for Macintosh are available in Excel 97 for
Created on 09/02/98 11:58 PM 1 EXCEL 98 TUTORIAL Chemistry C2407 fall 1998 Andy Eng, Columbia University 1998 Note for Excel 97 users: All features of Excel 98 for Macintosh are available in Excel 97 for
Using Windows 7 Explorer By Len Nasman, Bristol Village Computer Club
 By Len Nasman, Bristol Village Computer Club Understanding Windows 7 Explorer is key to taking control of your computer. If you have ever created a file and later had a hard time finding it, or if you
By Len Nasman, Bristol Village Computer Club Understanding Windows 7 Explorer is key to taking control of your computer. If you have ever created a file and later had a hard time finding it, or if you
MCDOUGAL LITTELL EASYPLANNER USER S GUIDE
 MCDOUGAL LITTELL EASYPLANNER USER S GUIDE 1 Introduction The McDougal Littell EasyPlanner allows you to quickly and easily access Resource materials such as tests, quizzes, books, and other supporting
MCDOUGAL LITTELL EASYPLANNER USER S GUIDE 1 Introduction The McDougal Littell EasyPlanner allows you to quickly and easily access Resource materials such as tests, quizzes, books, and other supporting
Adobe Dreamweaver CS5 Tutorial
 Adobe Dreamweaver CS5 Tutorial GETTING STARTED This tutorial focuses on the basic steps involved in creating an attractive, functional website. In using this tutorial you will learn to design a site layout,
Adobe Dreamweaver CS5 Tutorial GETTING STARTED This tutorial focuses on the basic steps involved in creating an attractive, functional website. In using this tutorial you will learn to design a site layout,
Getting Started With Excel
 Chapter 1 Getting Started With Excel This chapter will familiarize you with various basic features of Excel. Specific features which you need to solve a problem will be introduced as the need arises. When
Chapter 1 Getting Started With Excel This chapter will familiarize you with various basic features of Excel. Specific features which you need to solve a problem will be introduced as the need arises. When
User Manual. Administrator s guide for mass managing VirtueMart products. using. VM Mass Update 1.0
 User Manual Administrator s guide for mass managing VirtueMart products using VM Mass Update 1.0 The ultimate product management solution for VirtueMart! Contents Product Overview... 3 Feature List...
User Manual Administrator s guide for mass managing VirtueMart products using VM Mass Update 1.0 The ultimate product management solution for VirtueMart! Contents Product Overview... 3 Feature List...
Microsoft Access II 1.) Opening a Saved Database Music Click the Options Enable this Content Click OK. *
 Microsoft Access II 1.) Opening a Saved Database Open the Music database saved on your computer s hard drive. *I added more songs and records to the Songs and Artist tables. Click the Options button next
Microsoft Access II 1.) Opening a Saved Database Open the Music database saved on your computer s hard drive. *I added more songs and records to the Songs and Artist tables. Click the Options button next
Polymorphism and Variant Analysis Lab
 Polymorphism and Variant Analysis Lab Arian Avalos PowerPoint by Casey Hanson Polymorphism and Variant Analysis Matt Hudson 2018 1 Exercise In this exercise, we will do the following:. 1. Gain familiarity
Polymorphism and Variant Analysis Lab Arian Avalos PowerPoint by Casey Hanson Polymorphism and Variant Analysis Matt Hudson 2018 1 Exercise In this exercise, we will do the following:. 1. Gain familiarity
Lastly, in case you don t already know this, and don t have Excel on your computers, you can get it for free through IT s website under software.
 Welcome to Basic Excel, presented by STEM Gateway as part of the Essential Academic Skills Enhancement, or EASE, workshop series. Before we begin, I want to make sure we are clear that this is by no means
Welcome to Basic Excel, presented by STEM Gateway as part of the Essential Academic Skills Enhancement, or EASE, workshop series. Before we begin, I want to make sure we are clear that this is by no means
Dreamweaver Domain 4: Adding Content by Using Dreamweaver CS5
 Dreamweaver Domain 4: Adding Content by Using Dreamweaver CS5 Adobe Creative Suite 5 ACA Certification Preparation: Featuring Dreamweaver, Flash, and Photoshop 1 Objectives Define a Dreamweaver site. Create,
Dreamweaver Domain 4: Adding Content by Using Dreamweaver CS5 Adobe Creative Suite 5 ACA Certification Preparation: Featuring Dreamweaver, Flash, and Photoshop 1 Objectives Define a Dreamweaver site. Create,
