NAMD: Biomolecular Simulation on Thousands of Processors
|
|
|
- Adela Barker
- 5 years ago
- Views:
Transcription
1 NAMD: Biomolecular Simulation on Thousands of Processors James Phillips Gengbin Zheng Laxmikant Kalé Abstract NAMD is a parallel, object-oriented molecular dynamics program designed for high performance simulation of large biomolecular systems. NAMD employs the prioritized message-driven execution capabilities of the Charm++/Converse parallel runtime system, allowing excellent parallel scaling on both massively parallel supercomputers and commodity workstation clusters. This paper discusses the techniques which have allowed NAMD to effectively employ over one thousand processors in production simulations of biomedical relevance. 1 Introduction NAMD is a parallel, object-oriented molecular dynamics program designed for high performance simulation of large biomolecular systems [7]. NAMD employs the prioritized message-driven execution capabilities of the Charm++/Converse parallel runtime system, 1 allowing excellent parallel scaling on both massively parallel supercomputers and commodity workstation clusters. NAMD is distributed free of charge via the web 2 to over 3000 registered users as both source code and convenient precompiled binaries. NAMD development and support is a service of the National Institutes of Health Resource for Macromolecular Modeling and Bioinformatics, located at the University of Illinois at Urbana-Champaign. 3 Molecular dynamics simulation. In a molecular dynamics (MD) simulation, full atomic coordinates of the proteins, nucleic acids, and/or lipids of interest, as well as explicit water and ions, are obtained from known crystallographic or other structures. An empirical energy function, which consists of approximations of covalent interactions in addition to long-range Lennard-Jones and electrostatic terms, is applied. The resulting Newtonian equations of motion are typically integrated by symplectic and reversible methods using a timestep of 1 fs. Modifications are made to the equations of motion to control temperature and pressure during the simulation. The basic protocol for MD simulations consists of minimization (to eliminate initial contacts which would destabilize the integrator), equilibration to a temperature of 300 K and a pressure of 1 atm, and simulation in an isobaric (NPT) ensemble for 1-10 ns. This is sufficient to test the stability of a biomolecular aggregate or to observe the relaxation of the system into a more favorable This work was supported by the National Institutes of Health (NIH PHS 5 P41 RR ) and the National Science Foundation (NSF/GCAG BIR and NSF BIR EQ). Beckman Institute, University of Illinois, Urbana, IL. Department of Computer Science and Beckman Institute, University of Illinois, Urbana, IL. Department of Computer Science and Beckman Institute, University of Illinois, Urbana, IL
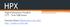
2 Figure 1: Simulations have increased exponentially in size, from BPTI (upper left, about 3K atoms), through the estrogen receptor (lower left, 36K atoms, 1996), to F 1 -ATPase (right, 327K atoms, 2001). (Atom counts include solvent.) conformation. However, if the goal is to study events which would not spontaneously occur during the timespan of the simulation, the system may be forced to undergo a transition via the application of forces in a steered molecular dynamics protocol. Important observations may be made during such a simulation of non-equilibrium events, even though the simulated time scale is much shorter than the natural one. Importance of molecular dynamics simulation software. The application of molecular dynamics simulation methods in biomedicine is directly dependent upon the capability, performance, usability and availability of the required simulation and analysis software. Simulations often require substantial computing resources, sometimes available only at supercomputing centers. By developing the program NAMD, the Resource provides the biomedical research community with a freely available tool for performing high quality MD simulations of large biomolecular systems using a variety of available and cost-effective hardware [7, 10]. Increases in system size and simulation length. With continuing increases in high performance computing technology, the domain of biomolecular simulation has rapidly expanded from isolated proteins in solvent to include complex aggregates, often in a lipid environment. Such simulations can easily exceed 100,000 atoms (see Fig. 1). Similarly, studying the function of even the simplest of biomolecular machines requires simulations of 10 ns or longer, even when techniques for accelerating processes of interest are employed. The goal of interactive simulation of smaller molecules places even greater demands on the performance of MD software, as the sensitivity of 2
3 the haptic interface increases quadratically with simulation speed [14]. Importance of parallel computing to simulations. Despite the seemingly unending progress in microprocessor performance indicated by Moore s law, the urgent nature and computational needs of biomedical research demand that we pursue the additional factors of tens, hundreds, or thousands in total performance which may be had by harnessing a multitude of processors for a single calculation. While the MD algorithm is blessed with a large ratio of calculation to data, its parallelization to large numbers of processors is not straightforward [4]. The Resource has been particularly prescient in recognizing and pursuing the benefits of parallel computing for biomedical research. Increasing availability of large parallel resources. The Accelerated Strategic Computing Initiative, 4 a U.S. Department of Energy program, has provided an unprecedented impetus for the application of massively parallel teraflop computing to the problems of the physical sciences and engineering. The National Science Foundation has followed this lead, funding terascale facilities at the national centers with the intent of enabling research that can employ many or all of the processors on these new machines. 5 The concept of grid computing promises to make this computational power readily available from any desktop. Huge computing resources will soon be available to the biomedical researcher who is able to harness it using software such as NAMD. Other available simulation programs. The biomolecular modeling community sustains a variety of software packages with overlapping core functionality but varying strengths and motivations. For comparison, we select AMBER, CHARMM, GROMACS, NWChem, and TINKER. AMBER [15] and CHARMM [3] are often considered the standard community codes of structural biology, having been developed over many years by a wide variety of researchers. Both AMBER and CHARMM support their own force field development efforts, although the form of the energy functions themselves is quite similar. Both codes are implemented in FORTRAN 77, although AMBER takes the form of a large package of specialized programs while CHARMM is a single binary. The parallelization of these codes is limited and not uniform across features, e.g., the GIBBS module of AMBER is limited to costly shared memory machines. Neither program is freely available, although the academic versions are highly discounted in comparison to commercial licenses. GROMACS 3.0 [9], the version recently released, claims the title of fastest MD. This can be attributed largely to the GROMOS force field, which neglects most hydrogen atoms and eliminates van der Waals interactions for those that remain. In contrast, the AMBER and CHARMM force fields represent all atoms and new development has centered on increasing accuracy via additional terms. Additional performance on Intel x86 processors comes from the implementation of inner loops in assembly code. GROMACS is implemented in C as a large package of programs and is newly released under the GNU General Public License (GPL). Distribution takes the form of source code and Linux binaries. Plans for future support and development are unclear for this program. NWChem [6] is a comprehensive molecular simulation system developed by a large group of researchers at the PNNL EMSL, primarily to meet internal requirements. The code centers on quantum mechanical methods but includes an MD component. Implemented in C and FORTRAN,
4 NWChem is parallelized using MPI and a Global Arrays library 6 which automatically redistributes data on distributed memory machines. Parallel scaling is respectable given sufficient workload, although published benchmarks tend to use abnormally large cutoffs rather than the 12 Å (or PME) typically used in biomolecular simulations. Access to NWChem source code is available with the submission of a signed license agreement, although support is explicitly unavailable outside of PNNL. TINKER [12] is a small FORTRAN code developed primarily for the testing of new methods. It incorporates a variety of force fields, in addition to its own, and includes many experimental methods. The code is freely available, but is not parallelized, and is therefore inappropriate for traditional large-scale biomolecular simulations. It does, however, provide the community with a simple code for experiments in method development. Motivation for NAMD development. NAMD builds upon other available programs in the field by incorporating popular force fields and reading file formats from other codes. NAMD complements existing capabilities by providing a higher performance alternative for simulations on the full range of available parallel platforms. Through NAMD, the Resource expands the MD user community by providing free, ready-to-run, easy-to-use software for most available platforms, complete with web-based educational materials and a safety net of support via . NAMD also provides easily modified object-oriented C++ source code to support method development both within and beyond the Resource. 2 NAMD Parallelization Strategy We have approached the scalability challenge by adopting message-driven approaches and reducing the complexity associated with these methods by combining multithreading and an object-oriented implementation in C++. Charm++ parallel object-based programming model. The dynamic components of NAMD are implemented in the Charm++[8] parallel language. Charm++ implements an object-based message-driven execution model. In Charm++ applications, there are collections of C++ objects, which communicate by remotely invoking methods on other objects by messages. Compared with conventional programming models such as message passing, shared memory or data parallel programming, Charm++ has several advantages in improving locality, parallelism and load balance. The flexibility provided by Charm++ is a key to the high performance achieved by NAMD on thousands of processors. In Charm++ applications, users decompose the problem into objects, and since they decide the granularity of the objects, it is easier for them to generate parallelism. As described in the paragraph below, NAMD uses a novel way of decomposition that easily generates the large amount of parallelism needed to occupy thousands of processors. Charm++ s object-based decomposition also help users to improve data locality. Objects encapsulate states, Charm++ objects are only allowed to directly access their own local memory. Access to other data is only possible via asynchronous method invocation to other objects. Charm++ s parallel objects and data-driven execution adaptively overlaps communication and computation and hide communication latency: when an object is waiting for some incoming data, entry functions of other objects with all data ready are free to execute
5 Bonded Force Objects Patch A Patch B Proxy C Proxy D Non-bonded Non-bonded Non-bonded Non-bonded Pair Compute Self Compute Self Compute Pair Compute Objects Objects Objects Objects PROCESSOR 1 Figure 2: NAMD 2 hybrid force/spatial decomposition. In Charm++, objects may even migrate from processor to processor at runtime. Object migration is controlled by Charm++ load balancer. Charm++ implements a measurement based load balancing framework which automatically instruments all Charm++ objects, collectes computation load and communication pattern during execution and stores them into a load balancing database. Charm++ then provides a collection of load balancing strategies whose job is to decide on a new mapping of objects to processors based on information from the database. Load balancing strategies are implemented in Charm++ as libraries. Programmers can easily experiment with different existing strategies by linking different strategy modules and specify which strategy to use at runtime via command line options. This involves very little efforts from programmers while achieving significant improvements in performance in adaptive applications. Application specific load balancing stategies can also be developed by users and plugged in easily. In the following paragraphs, we will describe the load balancing strategies optimized for NAMD2 in details. Hybrid force/spatial decomposition. NAMD 1 is parallelized via a form of spatial decomposition using cubes whose dimensions are slightly larger than the cutoff radius. Thus, atoms in one cube need to interact only with their 26 neighboring cubes. However, one problem with this spatial decomposition is that the number of cubes is limited by the simulation space. Even on a relatively large molecular system, such as the 92,442 atom ApoA-I benchmark, we only have 245 (7 7 5) cubes. Further, as density of the system varies across space, one may encounter strong load imbalances. NAMD 2 addresses this problem with a novel combination of force [11] and spatial decomposition. For each pair of neighboring cubes, we assign a non-bonded force computation object, which can be independently mapped to any processor. The number of such objects is therefore 14 times (26/2 + 1 self-interaction) the number of cubes. The cubes described above are represented in NAMD 2 by objects called home patches. Each home patch is responsible for distributing coordinate data, retrieving forces, and integrating the 5
6 equations of motion for all of the atoms in the cube of space owned by the patch. The forces used by the patches are computed by a variety of compute objects. There are several varieties of compute objects, responsible for computing the different types of forces (bond, electrostatic, constraint, etc.). Some compute objects require data from one patch, and only calculate interactions between atoms within that single patch. Other compute objects are responsible for interactions between atoms distributed among neighboring patches. Relationships among objects are illustrated in Fig. 2. Dynamic communication and execution. When running in parallel, some compute objects require data from patches not on the compute object s processor. In this case, a proxy patch takes the place of the home patch on the compute object s processor. During each time step, the home patch requests new forces from local compute objects, and sends its atom positions to all its proxy patches. Each proxy patch informs the compute objects on the proxy patch s processor that new forces must be calculated. When the compute objects provide the forces to the proxy, the proxy returns the data to the home patch, which combines all incoming forces before integrating. Thus, all computation and communication is scheduled based on priority and the availability of required data. Measurement-based load balancing. Some compute objects are permanently placed on processors at the start of the simulation, but others are moved during periodic load balancing phases. Ideally, all compute objects would be able to be moved around at any time. However, where calculations must be performed for atoms in several patches, it is more efficient to assume that some compute objects will not move during the course of the simulation. In general, the bulk of the computational load is represented by the non-bonded (electrostatic and van der Waals) interactions, and certain types of bonds. These objects are designed to be able to migrate during the simulation to optimize parallel efficiency. The non-migratable objects, including computations for bonds spanning multiple patches, represent only a small fraction of the work, so good load balance can be achieved without making them migratable. NAMD uses a measurement-based load balancer, employing the Charm++ load balancing framework. When a simulation begins, patches are distributed according to a recursive coordinate bisection scheme [2], so that each processor receives a number of neighboring patches. All compute objects are then distributed to a processor owning at least one home patch, insuring that each patch has at most seven proxies. The dynamic load balancer uses the load measurement capabilities of Converse to refine the initial distribution. The framework measures the execution time of each compute object (the object loads), and records other (non-migratable) patch work as background load. After the simulation runs for several timesteps (typically several seconds to several minutes), the program suspends the simulation to trigger the initial load balancing. NAMD retrieves the object times and background load from the framework, computes an improved load distribution, and redistributes the migratable compute objects. Success on large supercomputers. NAMD s combination of force and spatial decomposition allows large simulations to be efficiently decomposed onto hundreds of processors. Our success in employing this technique is demonstrated by a parallel speedup of 1252 on 2048 CPUs of the Sandia ASCI Red for a 206,617 atom cutoff (non-pme) simulation [4] and by our selection as a finalist for the 2000 Gordon Bell prize. When combined with ongoing work on improving the parallelization of PME simulations, NAMD is uniquely positioned as the tool of choice for simulations employing up to a thousand processors on the new generation of terascale platforms, i.e., the 3,000 6
7 128 NAMD 2.1 NAMD 2.2 NAMD 2.3 NAMD 2.4 NAMD ApoA1 (PME) Benchmark on PSC Cray T3E Processors x Time per Step (seconds) 64 Runtime for 1 ns Simulation 1 month 2 weeks 1 week 4 days 2 days Number of Processors Figure 3: Total resources consumed per step for 92K atom benchmark with and without particle mesh Ewald by NAMD versions on varying numbers of processors of the PSC T3E. Perfect linear scaling is a horizontal line. Diagonal scale shows runtime per ns. Note that performance increases for both PME and cutoff simulations with each release. processor AlphaServer SC and the 1,000 processor Itanium cluster at the Pittsburgh and Illinois supercomputing centers. 3 Particle Mesh Ewald in NAMD Incorporating, tuning, and parallelizing particle mesh Ewald (PME) [5] full electrostatics calculations has been a recurring theme in NAMD 2 development and this one feature has primarily determined the performance and scalability observed by users. As seen in Fig. 3, significant progress has been made. Development started with the incorporation of the external DPME 7 package into NAMD 2.0, providing a stable base functionality. The PME reciprocal sum was serialized in this version because the target workstation clusters for which DPME was developed obtained sufficient scaling by only distributing the direct interactions. In NAMD, the direct interactions were incorporated into the existing parallel cutoff non-bonded calculation. The reciprocal sum was reimplemented more efficiently in NAMD 2.1, providing a base for later parallelization. An effective parallel implementation of the PME reciprocal sum was finally achieved in NAMD 2.2. The final design was elegant but far from obvious, as numerous false starts were attempted along the way. In particular, the parallel implementation of FFTW 8 was found to be inappropriate for our purposes so a new one was written for NAMD. The reciprocal sum is currently parallelized only to the size of the PME grid, which is typically between 50 and 128. However, this is sufficient to allow 7 Distributed Particle Mesh Ewald, 8 Fastest Fourier Transform in the West, 7
8 NAMD simulations to scale to several hundred processors as the bottleneck has been significantly reduced. In addition, the message-driven design of NAMD interleaves the PME communication with other work, giving good performance even on high latency networks. Improvements to the PME direct sum in NAMD 2.3 were obtained by eliminating expensive calls to erfc(). 9 This was accomplished by incorporating the entire short-range non-bonded potential into an interpolation table. By interpolating based on the square of the interaction distance, the calculation of 1/ r 2 was eliminated as well. The interpolation table is carefully constructed to avoid cancellation errors and is iteratively refined during program startup to eliminate discontinuities in the calculated forces. Simulations performed with the new code are up to 50% faster than before and are of equivalent accuracy. Shared memory optimizations. Although NAMD 2 was initially designed to employ multiple threads on multiprocessor shared-memory machines, particularly in combination with messagepassing on a cluster of SMP nodes, this was never implemented. Even on modern clusters with SMP nodes, standard MPI-based message passing is normally the encouraged method of programming. However, it is certainly possible to exploit superior communication speeds between processors on the same node through algorithm design in a single-threaded implementation. Modifications to the NAMD load balancing system and proxy communication algorithms will reduce transmission of coordinates and forces between nodes. Distributing PME calculations to at most one processor per node for large machines in NAMD 2.4 has reduced contention for available interconnect bandwidth and other switch or network interface resources. This has increase the scalability of NAMD 2 simulations using PME to 1024 processors on the existing machines at NCSA, PSC, and SDSC with 2, 4, and 8 processors per node. 4 Scalability Results Figure 4 illustrates the portable scalability of NAMD on a variety of recent platforms employed for production simulations by researchers at the Resource. Figure 5 shows the greatest scalability attained by NAMD on the 3000 processor LeMieux cluster at PSC. NAMD scales particularly well for larger simulations, which are those for which improved performance is most greatly desired by researchers. 5 Remaining Challenges Continuation after external failures. The increasing size and complexity of parallel computing platforms will soon reduce the mean time between failures for hardware and system software to the point that large jobs will be more likely to fail due to an external error than to complete normally. Application software must address this problem internally, as general methods which may be developed are likely to be incomplete or inflexible. While NAMD 2 periodically saves atomic coordinates and velocities, the user is required to alter input files and restart the calculation manually. NAMD 3 will provide a single comprehensive restart file which will save all relevant information, including the state of the command interpreter. This will allow a simulation which has ended prematurely to be continued automatically. In a second, more aggressive approach, we 9 The erfc(x) function computes the complement of the error function of x. 8
9 64 NCSA Platinum 2xP3/1000 Myri SDSC IBM SP 8xPWR3/375 Resource Linux K7/ bT NCSA Titan 2xIA64/800 Myri PSC LeMieux 4xev6/1000 NAMD 2.4 ApoA1 (PME) Benchmark 1 month 2 weeks 1 week Processors x Time per Step (seconds) Runtime for 1 ns Simulation 4 days 2 days 1 day 12 hours Number of Processors Figure 4: Total resources consumed per step for 92K atom benchmark with particle mesh Ewald by NAMD 2.4 on varying numbers of processors for recent parallel platforms. Perfect linear scaling is a horizontal line. Diagonal scale shows runtime per ns. will develop methods to allow a running simulation to survive and rapidly recover from the loss of a small number of nodes, dynamically reallocating calculations to the remaining processors. Repeatable and deterministic parallel simulations. In NAMD 2, the order of operations, particularly the summation of forces, is determined dynamically by the load balancer and the order of message arrival. Combined with the non-associativity of floating-point arithmetic and the chaotic nature of MD simulations, this causes parallel simulations starting from identical initial conditions to diverge after a few hundred steps, even when run on the same machine and number of processors. In order to obtain the benefits of exact repeatability for both the developer (needed to perform long running regression tests) and for the user (repeating a simulation in order to extract additional details), NAMD 3 will optionally employ associative simulated fixed-point addition for force summation. This can be done at almost no additional cost [13]. Accelerating interactive simulation. Interactive simulation will become truly useful when a researcher can obtain 1 ps of simulation time per second of wall clock time. At this rate 1 ns is simulated in 20 minutes and 1 µs in under two weeks. Obtaining this level of performance, which is at least a factor of ten beyond current performance on any platform, will require the distribution of a simulation of 10,000 atoms efficiently to 256 processors of a high performance, tightly coupled machine. Latency-tolerant minimally coupled algorithms will be essential in this environment, as any synchronization would harm performance. Force computations must be partitioned into even smaller, fine-grained, objects than currently attained. In addition to the parallelization challenges, careful compromises between performance and accuracy will need to be made in selecting algorithms 9
10 64 ATPase (327K atoms) ApoA1 (92K atoms) NAMD 2.4 (PME) on PSC LeMieux Processors x Time per Step (seconds) Runtime for 1 ns Simulation 2 weeks 1 week 4 days 2 days 1 day 12 hours 6 hours Number of Processors Figure 5: Total resources consumed per step for 92K and 327K atom benchmarks with particle mesh Ewald by NAMD 2.4 on varying numbers of processors for PSC LeMieux. Perfect linear scaling is a horizontal line. Diagonal scale shows runtime per ns. for both force calculation and integration. Large capability clusters. All recent terascale systems have combined standard multiprocessor workstations with proprietary low latency interconnects. Common examples are the IBM SP (San Diego), 10 AlphaServer SC (Pittsburgh), 11 and Myrinet Linux clusters (Illinois). 12 In order to scale to thousands of processors on these platforms, NAMD will have to take advantage (through Converse) of the low level interfaces to the interconnect hardware (e.g., IBM s LAPI [1]) which translate to a message driven design much better than the common MPI interface. Multiprocessor nodes are used on these machines (up to 32 CPUs per node on ASCI Q, the AlphaServer SC being installed at Los Alamos 13 ), so NAMD will be adapted to this multilevel communication hierarchy. This will be in the form of multiple threads on some platforms while merely modifying communication patterns on others. In either case, load balancing will need to consider the disparity of communication speeds between processors. Massively parallel platforms. Supercomputers with over 1000 processors are available at the national centers (San Diego, Pittsburgh, and Illinois) and machines with over 10,000 processors are in use at DOE labs. We expect computational facilities with 10,000 to 100,000 processors to be available for simulations by the middle of the proposed funding period, e.g., IBM is expected to build
11 parallel machines in this range by To scale NAMD to such large machines, we will develop and implement new parallel algorithms. Fine-grained decomposition techniques will be used to allow each time step to be completed in less than a millisecond for simulations involving over 100,000 atoms. We will experiment with variations of our hybrid spatial-and-force decomposition strategy that lead to smaller computational pieces. This fine grained decomposition will be supported by a new low-overhead runtime system, which can schedule each computational object with 1 2 microseconds overhead. Total communication on such large machines is typically limited because of physical considerations. We will develop new load balancers that place communicating objects in such a way as to minimize the average number of hops traveled by messages, while simultaneously minimizing load imbalance and communication volume. Processor-in-memory architectures. During years 3 5 of the proposed funding period, machines with processors-in-memory chips are expected to become available. Such machines will provide a much higher aggregate bandwidth between processor and memory modules, by integrating multiple processors and memory within one chip. We will harness the high performance potential of such machines by developing a version of NAMD which supports a low memory-to-processor ratio (by eliminating replicated molecule data, for example). New algorithms that can deal with on-demand movement of data among memory modules will be implemented, to support prefetching of data. One of the potential machines in this category will be a Teraflop Board consisting of multiple multiprocessor chips designed for the original IBM Blue Gene machine. 14 Accommodating large biomolecular aggregates. Larger simulations generally scale better on multi-processor machines than smaller simulations simply because there is more computation to be distributed. However, the memory usage of the master node in NAMD 2 is proportional to the number of atoms in the simulation, and does not decrease as additional processors are added. Because of this, systems larger than 100,000 atoms cannot run on a Cray T3E with only 128 MB of memory per node and 300,000 atoms can require 512 MB on the master node to perform load balancing. To address this problem, a parallel load balancing algorithm will be implemented that will not collect all load data on the master node. Additionally, the replicated molecular structure will be compressed by a general method which locates and eliminates repeated patterns such as water molecules, lipids, and amino acids. 6 Parallel Architecture Design It is immensely ingenious, immensely complicated, and extremely effective, but somehow at the same time crude, wasteful, and inelegant, and one feels that there must be a better way of doing things. C. Strachey (1962, concerning the IBM Stretch computer) When developing complex software, the first forays into a new application domain or approach to software design often lead to a collection of code which, although correct, is more indicative of the learning process of its creator than the final understanding which was gained. Successive implementations do not avoid this fate, but merely encounter it in the pursuit of new and more complex functionality
12 NAMD 2 demonstrated that object oriented design and cooperative multithreading make it possible to write and maintain a complex message-driven program. However, a cleansing of the source code with the implementation of NAMD 3 will prevent its degeneration into the prevalent software architecture recently described as a big ball of mud. 15 Insights to be applied in the design of NAMD 3 are summarized below. Threadless control structure. NAMD 2 uses cooperative user-space threads to provide individual control loops for each fixed-size spatial domain (or patch) and for global aspects of the algorithm such as pressure control. These threads are scheduled by Charm++/Converse in the same queue as other tasks. In order to improve portability and allow for simplified debugging, control loops will be implemented using a simple state machine and preprocessor. Using object member variables in place of a stack, and providing additional control logic to handle dependencies, will allow for the clear expression of algorithms in this restrictive environment. Exploiting multiprocessor nodes. NAMD 3 will run on clusters with multiprocessor nodes either with threads or as independent processes. Tasks will be permanently assigned to a given processor based on periodic load balancing. The alternative method of using a single work queue for all processors on a node would require a complex protocol to ensure that all processors completed at the same time and would also depend heavily on expensive synchronization and mutual exclusion functions. The primary concern for multiprocessor nodes will be optimizing data layout and dependencies to limit inter-node communication, a development which is not dependent on shared memory protocols. Patterns for parallel operations. The parallel efficiency of NAMD 2 is due to a limited number of parallel operations, some of which are necessarily complex. The remainder of the code, however, only affects startup but is implemented via a variety of methods which were selected based on immediate convenience. The implementation and maintenance of NAMD 3 will be greatly aided by selecting a small number of useful patterns and encapsulating them as templates for repeated use. Distributed simulation components. The general molecular dynamics algorithms and other utility code employed in NAMD 3 will be separated from the parallel control structures in order to provide a basis of components which can be independently tested and reused in other applications such as VMD. The imposition of clearly defined interfaces between these components will also aid the development of test harnesses for NAMD. References [1] M. Banikazemi, R. K. Govindaraju, R. Blackmore, and D. K. Panda. MPI-LAPI: An efficient implementation of MPI for IBM RS/6000 SP systems. IEEE Trans. Parallel and Distributed Systems, In press. [2] M. Berger and S. Bokhari. A partitioning strategy for nonuniform problems on multiprocessors. IEEE Trans. Computers, C-36: ,
13 [3] B. R. Brooks, R. E. Bruccoleri, B. D. Olafson, D. J. States, S. Swaminathan, and M. Karplus. CHARMM: A program for macromolecular energy, minimization, and dynamics calculations. J. Comp. Chem., 4: , [4] R. K. Brunner, J. C. Phillips, and L. V. Kalé. Scalable molecular dynamics for large biomolecular systems. In Proceedings of the 2000 ACM/IEEE SC2000 Conference. ACM, [5] T. Darden, D. York, and L. Pedersen. Particle mesh Ewald. An N log(n) method for Ewald sums in large systems. J. Chem. Phys., 98: , [6] High Performance Computational Chemistry Group. NWChem, a computational chemistry package for parallel computers, version [7] L. Kalé, R. Skeel, M. Bhandarkar, R. Brunner, A. Gursoy, N. Krawetz, J. Phillips, A. Shinozaki, K. Varadarajan, and K. Schulten. NAMD2: Greater scalability for parallel molecular dynamics. J. Comp. Phys., 151: , [8] L. V. Kalé and S. Krishnan. Charm++: Parallel programming with message-driven objects. In G. V. Wilson and P. Lu, editors, Parallel Programming using C++, pages MIT Press, [9] E. Lindahl, B. Hess, and D. van der Spoel. GROMACS 3.0: a package for molecular simulation and trajectory analysis. J. Mol. Mod., [10] M. Nelson, W. Humphrey, A. Gursoy, A. Dalke, L. Kalé, R. D. Skeel, and K. Schulten. NAMD A parallel, object-oriented molecular dynamics program. Int. J. Supercomp. Appl. High Perform. Comp., 10: , [11] S. J. Plimpton and B. A. Hendrickson. A new parallel method for molecular dynamics simulation of macromolecular systems. J. Comp. Chem., 17(3): , [12] J. W. Ponder and F. M. Richards. An efficient Newton-like method for molecular mechanics energy minimization of large molecules. J. Comp. Chem., 8: , [13] R. D. Skeel. Symplectic integration with floating-point arithmetic and other approximations. Appl. Numer. Math., 29:3 18, [14] J. Stone, J. Gullingsrud, P. Grayson, and K. Schulten. A system for interactive molecular dynamics simulation. In J. F. Huges and C. H. Séquin, editors, 2001 ACM Symposium on Interactive 3D Graphics, pages , New York, ACM SIGGRAPH. [15] P. K. Weiner and P. A. Kollman. AMBER: Assisted model building with energy refinement. A general program for modeling molecules and their interactions. J. Comp. Chem., 2(3): ,
NAMD: Biomolecular Simulation on Thousands of Processors
 NAMD: Biomolecular Simulation on Thousands of Processors James C. Phillips Gengbin Zheng Sameer Kumar Laxmikant V. Kalé Abstract NAMD is a fully featured, production molecular dynamics program for high
NAMD: Biomolecular Simulation on Thousands of Processors James C. Phillips Gengbin Zheng Sameer Kumar Laxmikant V. Kalé Abstract NAMD is a fully featured, production molecular dynamics program for high
NAMD Serial and Parallel Performance
 NAMD Serial and Parallel Performance Jim Phillips Theoretical Biophysics Group Serial performance basics Main factors affecting serial performance: Molecular system size and composition. Cutoff distance
NAMD Serial and Parallel Performance Jim Phillips Theoretical Biophysics Group Serial performance basics Main factors affecting serial performance: Molecular system size and composition. Cutoff distance
Using GPUs to compute the multilevel summation of electrostatic forces
 Using GPUs to compute the multilevel summation of electrostatic forces David J. Hardy Theoretical and Computational Biophysics Group Beckman Institute for Advanced Science and Technology University of
Using GPUs to compute the multilevel summation of electrostatic forces David J. Hardy Theoretical and Computational Biophysics Group Beckman Institute for Advanced Science and Technology University of
A Parallel-Object Programming Model for PetaFLOPS Machines and BlueGene/Cyclops Gengbin Zheng, Arun Singla, Joshua Unger, Laxmikant Kalé
 A Parallel-Object Programming Model for PetaFLOPS Machines and BlueGene/Cyclops Gengbin Zheng, Arun Singla, Joshua Unger, Laxmikant Kalé Parallel Programming Laboratory Department of Computer Science University
A Parallel-Object Programming Model for PetaFLOPS Machines and BlueGene/Cyclops Gengbin Zheng, Arun Singla, Joshua Unger, Laxmikant Kalé Parallel Programming Laboratory Department of Computer Science University
Abhinav Bhatele, Laxmikant V. Kale University of Illinois at Urbana Champaign Sameer Kumar IBM T. J. Watson Research Center
 Abhinav Bhatele, Laxmikant V. Kale University of Illinois at Urbana Champaign Sameer Kumar IBM T. J. Watson Research Center Motivation: Contention Experiments Bhatele, A., Kale, L. V. 2008 An Evaluation
Abhinav Bhatele, Laxmikant V. Kale University of Illinois at Urbana Champaign Sameer Kumar IBM T. J. Watson Research Center Motivation: Contention Experiments Bhatele, A., Kale, L. V. 2008 An Evaluation
NAMD at Extreme Scale. Presented by: Eric Bohm Team: Eric Bohm, Chao Mei, Osman Sarood, David Kunzman, Yanhua, Sun, Jim Phillips, John Stone, LV Kale
 NAMD at Extreme Scale Presented by: Eric Bohm Team: Eric Bohm, Chao Mei, Osman Sarood, David Kunzman, Yanhua, Sun, Jim Phillips, John Stone, LV Kale Overview NAMD description Power7 Tuning Support for
NAMD at Extreme Scale Presented by: Eric Bohm Team: Eric Bohm, Chao Mei, Osman Sarood, David Kunzman, Yanhua, Sun, Jim Phillips, John Stone, LV Kale Overview NAMD description Power7 Tuning Support for
Accelerating Molecular Modeling Applications with Graphics Processors
 Accelerating Molecular Modeling Applications with Graphics Processors John Stone Theoretical and Computational Biophysics Group University of Illinois at Urbana-Champaign Research/gpu/ SIAM Conference
Accelerating Molecular Modeling Applications with Graphics Processors John Stone Theoretical and Computational Biophysics Group University of Illinois at Urbana-Champaign Research/gpu/ SIAM Conference
NAMD: A Portable and Highly Scalable Program for Biomolecular Simulations
 NAMD: A Portable and Highly Scalable Program for Biomolecular Simulations Abhinav Bhatele 1, Sameer Kumar 3, Chao Mei 1, James C. Phillips 2, Gengbin Zheng 1, Laxmikant V. Kalé 1 1 Department of Computer
NAMD: A Portable and Highly Scalable Program for Biomolecular Simulations Abhinav Bhatele 1, Sameer Kumar 3, Chao Mei 1, James C. Phillips 2, Gengbin Zheng 1, Laxmikant V. Kalé 1 1 Department of Computer
Improving NAMD Performance on Volta GPUs
 Improving NAMD Performance on Volta GPUs David Hardy - Research Programmer, University of Illinois at Urbana-Champaign Ke Li - HPC Developer Technology Engineer, NVIDIA John Stone - Senior Research Programmer,
Improving NAMD Performance on Volta GPUs David Hardy - Research Programmer, University of Illinois at Urbana-Champaign Ke Li - HPC Developer Technology Engineer, NVIDIA John Stone - Senior Research Programmer,
Installation and Test of Molecular Dynamics Simulation Packages on SGI Altix and Hydra-Cluster at JKU Linz
 Installation and Test of Molecular Dynamics Simulation Packages on SGI Altix and Hydra-Cluster at JKU Linz Rene Kobler May 25, 25 Contents 1 Abstract 2 2 Status 2 3 Changelog 2 4 Installation Notes 3 4.1
Installation and Test of Molecular Dynamics Simulation Packages on SGI Altix and Hydra-Cluster at JKU Linz Rene Kobler May 25, 25 Contents 1 Abstract 2 2 Status 2 3 Changelog 2 4 Installation Notes 3 4.1
Simulating Life at the Atomic Scale
 Simulating Life at the Atomic Scale James Phillips Beckman Institute, University of Illinois Research/namd/ Beckman Institute University of Illinois at Urbana-Champaign Theoretical and Computational Biophysics
Simulating Life at the Atomic Scale James Phillips Beckman Institute, University of Illinois Research/namd/ Beckman Institute University of Illinois at Urbana-Champaign Theoretical and Computational Biophysics
Molecular Dynamics and Quantum Mechanics Applications
 Understanding the Performance of Molecular Dynamics and Quantum Mechanics Applications on Dell HPC Clusters High-performance computing (HPC) clusters are proving to be suitable environments for running
Understanding the Performance of Molecular Dynamics and Quantum Mechanics Applications on Dell HPC Clusters High-performance computing (HPC) clusters are proving to be suitable environments for running
Multiprocessing and Scalability. A.R. Hurson Computer Science and Engineering The Pennsylvania State University
 A.R. Hurson Computer Science and Engineering The Pennsylvania State University 1 Large-scale multiprocessor systems have long held the promise of substantially higher performance than traditional uniprocessor
A.R. Hurson Computer Science and Engineering The Pennsylvania State University 1 Large-scale multiprocessor systems have long held the promise of substantially higher performance than traditional uniprocessor
Gromacs 4 tips & tricks Or: Speeding up your simulations
 Gromacs 4 tips & tricks Or: Speeding up your simulations Erik Lindahl lindahl@cbr.su.se Center for Biomembrane Research Stockholm University, Sweden CBR Topics Algorithms behind the options Tips & tricks
Gromacs 4 tips & tricks Or: Speeding up your simulations Erik Lindahl lindahl@cbr.su.se Center for Biomembrane Research Stockholm University, Sweden CBR Topics Algorithms behind the options Tips & tricks
Charm++ for Productivity and Performance
 Charm++ for Productivity and Performance A Submission to the 2011 HPC Class II Challenge Laxmikant V. Kale Anshu Arya Abhinav Bhatele Abhishek Gupta Nikhil Jain Pritish Jetley Jonathan Lifflander Phil
Charm++ for Productivity and Performance A Submission to the 2011 HPC Class II Challenge Laxmikant V. Kale Anshu Arya Abhinav Bhatele Abhishek Gupta Nikhil Jain Pritish Jetley Jonathan Lifflander Phil
Multilevel Summation of Electrostatic Potentials Using GPUs
 Multilevel Summation of Electrostatic Potentials Using GPUs David J. Hardy Theoretical and Computational Biophysics Group Beckman Institute for Advanced Science and Technology University of Illinois at
Multilevel Summation of Electrostatic Potentials Using GPUs David J. Hardy Theoretical and Computational Biophysics Group Beckman Institute for Advanced Science and Technology University of Illinois at
Resource allocation and utilization in the Blue Gene/L supercomputer
 Resource allocation and utilization in the Blue Gene/L supercomputer Tamar Domany, Y Aridor, O Goldshmidt, Y Kliteynik, EShmueli, U Silbershtein IBM Labs in Haifa Agenda Blue Gene/L Background Blue Gene/L
Resource allocation and utilization in the Blue Gene/L supercomputer Tamar Domany, Y Aridor, O Goldshmidt, Y Kliteynik, EShmueli, U Silbershtein IBM Labs in Haifa Agenda Blue Gene/L Background Blue Gene/L
The midpoint method for parallelization of particle simulations
 THE JOURNAL OF CHEMICAL PHYSICS 124, 184109 2006 The midpoint method for parallelization of particle simulations Kevin J. Bowers and Ron O. Dror D. E. Shaw Research, LLC, 39th Floor, Tower 45, 120 West
THE JOURNAL OF CHEMICAL PHYSICS 124, 184109 2006 The midpoint method for parallelization of particle simulations Kevin J. Bowers and Ron O. Dror D. E. Shaw Research, LLC, 39th Floor, Tower 45, 120 West
Graphics Processor Acceleration and YOU
 Graphics Processor Acceleration and YOU James Phillips Research/gpu/ Goals of Lecture After this talk the audience will: Understand how GPUs differ from CPUs Understand the limits of GPU acceleration Have
Graphics Processor Acceleration and YOU James Phillips Research/gpu/ Goals of Lecture After this talk the audience will: Understand how GPUs differ from CPUs Understand the limits of GPU acceleration Have
Accelerating NAMD with Graphics Processors
 Accelerating NAMD with Graphics Processors James Phillips John Stone Klaus Schulten Research/namd/ NAMD: Practical Supercomputing 24,000 users can t all be computer experts. 18% are NIH-funded; many in
Accelerating NAMD with Graphics Processors James Phillips John Stone Klaus Schulten Research/namd/ NAMD: Practical Supercomputing 24,000 users can t all be computer experts. 18% are NIH-funded; many in
HANDLING LOAD IMBALANCE IN DISTRIBUTED & SHARED MEMORY
 HANDLING LOAD IMBALANCE IN DISTRIBUTED & SHARED MEMORY Presenters: Harshitha Menon, Seonmyeong Bak PPL Group Phil Miller, Sam White, Nitin Bhat, Tom Quinn, Jim Phillips, Laxmikant Kale MOTIVATION INTEGRATED
HANDLING LOAD IMBALANCE IN DISTRIBUTED & SHARED MEMORY Presenters: Harshitha Menon, Seonmyeong Bak PPL Group Phil Miller, Sam White, Nitin Bhat, Tom Quinn, Jim Phillips, Laxmikant Kale MOTIVATION INTEGRATED
NAMD, CUDA, and Clusters: Taking GPU Molecular Dynamics Beyond the Deskop
 NAMD, CUDA, and Clusters: Taking GPU Molecular Dynamics Beyond the Deskop James Phillips Research/gpu/ Beckman Institute University of Illinois at Urbana-Champaign Theoretical and Computational Biophysics
NAMD, CUDA, and Clusters: Taking GPU Molecular Dynamics Beyond the Deskop James Phillips Research/gpu/ Beckman Institute University of Illinois at Urbana-Champaign Theoretical and Computational Biophysics
Overcoming Scaling Challenges in Biomolecular Simulations across Multiple Platforms
 Overcoming Scaling Challenges in Biomolecular Simulations across Multiple Platforms Abhinav Bhatelé, Sameer Kumar, Chao Mei, James C. Phillips, Gengbin Zheng, Laxmikant V. Kalé Department of Computer Science
Overcoming Scaling Challenges in Biomolecular Simulations across Multiple Platforms Abhinav Bhatelé, Sameer Kumar, Chao Mei, James C. Phillips, Gengbin Zheng, Laxmikant V. Kalé Department of Computer Science
In-Situ Visualization and Analysis of Petascale Molecular Dynamics Simulations with VMD
 In-Situ Visualization and Analysis of Petascale Molecular Dynamics Simulations with VMD John Stone Theoretical and Computational Biophysics Group Beckman Institute for Advanced Science and Technology University
In-Situ Visualization and Analysis of Petascale Molecular Dynamics Simulations with VMD John Stone Theoretical and Computational Biophysics Group Beckman Institute for Advanced Science and Technology University
Performance Study of Popular Computational Chemistry Software Packages on Cray HPC Systems
 Performance Study of Popular Computational Chemistry Software Packages on Cray HPC Systems Junjie Li (lijunj@iu.edu) Shijie Sheng (shengs@iu.edu) Raymond Sheppard (rsheppar@iu.edu) Pervasive Technology
Performance Study of Popular Computational Chemistry Software Packages on Cray HPC Systems Junjie Li (lijunj@iu.edu) Shijie Sheng (shengs@iu.edu) Raymond Sheppard (rsheppar@iu.edu) Pervasive Technology
Lecture 9: MIMD Architectures
 Lecture 9: MIMD Architectures Introduction and classification Symmetric multiprocessors NUMA architecture Clusters Zebo Peng, IDA, LiTH 1 Introduction MIMD: a set of general purpose processors is connected
Lecture 9: MIMD Architectures Introduction and classification Symmetric multiprocessors NUMA architecture Clusters Zebo Peng, IDA, LiTH 1 Introduction MIMD: a set of general purpose processors is connected
Computing architectures Part 2 TMA4280 Introduction to Supercomputing
 Computing architectures Part 2 TMA4280 Introduction to Supercomputing NTNU, IMF January 16. 2017 1 Supercomputing What is the motivation for Supercomputing? Solve complex problems fast and accurately:
Computing architectures Part 2 TMA4280 Introduction to Supercomputing NTNU, IMF January 16. 2017 1 Supercomputing What is the motivation for Supercomputing? Solve complex problems fast and accurately:
ENABLING NEW SCIENCE GPU SOLUTIONS
 ENABLING NEW SCIENCE TESLA BIO Workbench The NVIDIA Tesla Bio Workbench enables biophysicists and computational chemists to push the boundaries of life sciences research. It turns a standard PC into a
ENABLING NEW SCIENCE TESLA BIO Workbench The NVIDIA Tesla Bio Workbench enables biophysicists and computational chemists to push the boundaries of life sciences research. It turns a standard PC into a
Enzo-P / Cello. Formation of the First Galaxies. San Diego Supercomputer Center. Department of Physics and Astronomy
 Enzo-P / Cello Formation of the First Galaxies James Bordner 1 Michael L. Norman 1 Brian O Shea 2 1 University of California, San Diego San Diego Supercomputer Center 2 Michigan State University Department
Enzo-P / Cello Formation of the First Galaxies James Bordner 1 Michael L. Norman 1 Brian O Shea 2 1 University of California, San Diego San Diego Supercomputer Center 2 Michigan State University Department
COMPILED HARDWARE ACCELERATION OF MOLECULAR DYNAMICS CODE. Jason Villarreal and Walid A. Najjar
 COMPILED HARDWARE ACCELERATION OF MOLECULAR DYNAMICS CODE Jason Villarreal and Walid A. Najjar Department of Computer Science and Engineering University of California, Riverside villarre, najjar@cs.ucr.edu
COMPILED HARDWARE ACCELERATION OF MOLECULAR DYNAMICS CODE Jason Villarreal and Walid A. Najjar Department of Computer Science and Engineering University of California, Riverside villarre, najjar@cs.ucr.edu
Scalable Fault Tolerance Schemes using Adaptive Runtime Support
 Scalable Fault Tolerance Schemes using Adaptive Runtime Support Laxmikant (Sanjay) Kale http://charm.cs.uiuc.edu Parallel Programming Laboratory Department of Computer Science University of Illinois at
Scalable Fault Tolerance Schemes using Adaptive Runtime Support Laxmikant (Sanjay) Kale http://charm.cs.uiuc.edu Parallel Programming Laboratory Department of Computer Science University of Illinois at
ECE519 Advanced Operating Systems
 IT 540 Operating Systems ECE519 Advanced Operating Systems Prof. Dr. Hasan Hüseyin BALIK (10 th Week) (Advanced) Operating Systems 10. Multiprocessor, Multicore and Real-Time Scheduling 10. Outline Multiprocessor
IT 540 Operating Systems ECE519 Advanced Operating Systems Prof. Dr. Hasan Hüseyin BALIK (10 th Week) (Advanced) Operating Systems 10. Multiprocessor, Multicore and Real-Time Scheduling 10. Outline Multiprocessor
Multiprocessors and Thread Level Parallelism Chapter 4, Appendix H CS448. The Greed for Speed
 Multiprocessors and Thread Level Parallelism Chapter 4, Appendix H CS448 1 The Greed for Speed Two general approaches to making computers faster Faster uniprocessor All the techniques we ve been looking
Multiprocessors and Thread Level Parallelism Chapter 4, Appendix H CS448 1 The Greed for Speed Two general approaches to making computers faster Faster uniprocessor All the techniques we ve been looking
The Use of Cloud Computing Resources in an HPC Environment
 The Use of Cloud Computing Resources in an HPC Environment Bill, Labate, UCLA Office of Information Technology Prakashan Korambath, UCLA Institute for Digital Research & Education Cloud computing becomes
The Use of Cloud Computing Resources in an HPC Environment Bill, Labate, UCLA Office of Information Technology Prakashan Korambath, UCLA Institute for Digital Research & Education Cloud computing becomes
Algorithm Engineering with PRAM Algorithms
 Algorithm Engineering with PRAM Algorithms Bernard M.E. Moret moret@cs.unm.edu Department of Computer Science University of New Mexico Albuquerque, NM 87131 Rome School on Alg. Eng. p.1/29 Measuring and
Algorithm Engineering with PRAM Algorithms Bernard M.E. Moret moret@cs.unm.edu Department of Computer Science University of New Mexico Albuquerque, NM 87131 Rome School on Alg. Eng. p.1/29 Measuring and
Analysis and Visualization Algorithms in VMD
 1 Analysis and Visualization Algorithms in VMD David Hardy Research/~dhardy/ NAIS: State-of-the-Art Algorithms for Molecular Dynamics (Presenting the work of John Stone.) VMD Visual Molecular Dynamics
1 Analysis and Visualization Algorithms in VMD David Hardy Research/~dhardy/ NAIS: State-of-the-Art Algorithms for Molecular Dynamics (Presenting the work of John Stone.) VMD Visual Molecular Dynamics
An Introduction to Parallel Programming
 An Introduction to Parallel Programming Ing. Andrea Marongiu (a.marongiu@unibo.it) Includes slides from Multicore Programming Primer course at Massachusetts Institute of Technology (MIT) by Prof. SamanAmarasinghe
An Introduction to Parallel Programming Ing. Andrea Marongiu (a.marongiu@unibo.it) Includes slides from Multicore Programming Primer course at Massachusetts Institute of Technology (MIT) by Prof. SamanAmarasinghe
NAMD-QM/MM TUTORIAL. Unix/MacOSX Version. NAMD-QM/MM Suite Developers: Marcelo C. R. Melo, Rafael C. Bernardi
 University of Illinois at Urbana-Champaign NIH Resource for Macromolecular Modelling and Bioinformatics Beckman Institute Computational Biophysics Workshop NAMD-QM/MM TUTORIAL Unix/MacOSX Version NAMD-QM/MM
University of Illinois at Urbana-Champaign NIH Resource for Macromolecular Modelling and Bioinformatics Beckman Institute Computational Biophysics Workshop NAMD-QM/MM TUTORIAL Unix/MacOSX Version NAMD-QM/MM
Partitioning Effects on MPI LS-DYNA Performance
 Partitioning Effects on MPI LS-DYNA Performance Jeffrey G. Zais IBM 138 Third Street Hudson, WI 5416-1225 zais@us.ibm.com Abbreviations: MPI message-passing interface RISC - reduced instruction set computing
Partitioning Effects on MPI LS-DYNA Performance Jeffrey G. Zais IBM 138 Third Street Hudson, WI 5416-1225 zais@us.ibm.com Abbreviations: MPI message-passing interface RISC - reduced instruction set computing
Strong Scaling for Molecular Dynamics Applications
 Strong Scaling for Molecular Dynamics Applications Scaling and Molecular Dynamics Strong Scaling How does solution time vary with the number of processors for a fixed problem size Classical Molecular Dynamics
Strong Scaling for Molecular Dynamics Applications Scaling and Molecular Dynamics Strong Scaling How does solution time vary with the number of processors for a fixed problem size Classical Molecular Dynamics
Lecture 9: MIMD Architecture
 Lecture 9: MIMD Architecture Introduction and classification Symmetric multiprocessors NUMA architecture Cluster machines Zebo Peng, IDA, LiTH 1 Introduction MIMD: a set of general purpose processors is
Lecture 9: MIMD Architecture Introduction and classification Symmetric multiprocessors NUMA architecture Cluster machines Zebo Peng, IDA, LiTH 1 Introduction MIMD: a set of general purpose processors is
GROMACS Performance Benchmark and Profiling. August 2011
 GROMACS Performance Benchmark and Profiling August 2011 Note The following research was performed under the HPC Advisory Council activities Participating vendors: Intel, Dell, Mellanox Compute resource
GROMACS Performance Benchmark and Profiling August 2011 Note The following research was performed under the HPC Advisory Council activities Participating vendors: Intel, Dell, Mellanox Compute resource
Toward Runtime Power Management of Exascale Networks by On/Off Control of Links
 Toward Runtime Power Management of Exascale Networks by On/Off Control of Links, Nikhil Jain, Laxmikant Kale University of Illinois at Urbana-Champaign HPPAC May 20, 2013 1 Power challenge Power is a major
Toward Runtime Power Management of Exascale Networks by On/Off Control of Links, Nikhil Jain, Laxmikant Kale University of Illinois at Urbana-Champaign HPPAC May 20, 2013 1 Power challenge Power is a major
Load Balancing Techniques for Asynchronous Spacetime Discontinuous Galerkin Methods
 Load Balancing Techniques for Asynchronous Spacetime Discontinuous Galerkin Methods Aaron K. Becker (abecker3@illinois.edu) Robert B. Haber Laxmikant V. Kalé University of Illinois, Urbana-Champaign Parallel
Load Balancing Techniques for Asynchronous Spacetime Discontinuous Galerkin Methods Aaron K. Becker (abecker3@illinois.edu) Robert B. Haber Laxmikant V. Kalé University of Illinois, Urbana-Champaign Parallel
Data I/O Optimization in GROMACS Using the Global Arrays Toolkit
 Available online at www.prace-ri.eu Partnership for Advanced Computing in Europe Data I/O Optimization in GROMACS Using the Global Arrays Toolkit Valentin Pavlov*, Peicho Petkov NCSA, Acad. G. Bonchev
Available online at www.prace-ri.eu Partnership for Advanced Computing in Europe Data I/O Optimization in GROMACS Using the Global Arrays Toolkit Valentin Pavlov*, Peicho Petkov NCSA, Acad. G. Bonchev
CS 475: Parallel Programming Introduction
 CS 475: Parallel Programming Introduction Wim Bohm, Sanjay Rajopadhye Colorado State University Fall 2014 Course Organization n Let s make a tour of the course website. n Main pages Home, front page. Syllabus.
CS 475: Parallel Programming Introduction Wim Bohm, Sanjay Rajopadhye Colorado State University Fall 2014 Course Organization n Let s make a tour of the course website. n Main pages Home, front page. Syllabus.
What are Clusters? Why Clusters? - a Short History
 What are Clusters? Our definition : A parallel machine built of commodity components and running commodity software Cluster consists of nodes with one or more processors (CPUs), memory that is shared by
What are Clusters? Our definition : A parallel machine built of commodity components and running commodity software Cluster consists of nodes with one or more processors (CPUs), memory that is shared by
Designing a Cluster for a Small Research Group
 Designing a Cluster for a Small Research Group Jim Phillips, John Stone, Tim Skirvin Low-cost Linux Clusters for Biomolecular Simulations Using NAMD Outline Why and why not clusters? Consider your Users
Designing a Cluster for a Small Research Group Jim Phillips, John Stone, Tim Skirvin Low-cost Linux Clusters for Biomolecular Simulations Using NAMD Outline Why and why not clusters? Consider your Users
Optimizing Fine-grained Communication in a Biomolecular Simulation Application on Cray XK6
 Optimizing Fine-grained Communication in a Biomolecular Simulation Application on Cray XK6 Yanhua Sun, Gengbin Zheng, Chao Mei, Eric J. Bohm, James C. Phillips, Laxmikant V. Kalé University of Illinois
Optimizing Fine-grained Communication in a Biomolecular Simulation Application on Cray XK6 Yanhua Sun, Gengbin Zheng, Chao Mei, Eric J. Bohm, James C. Phillips, Laxmikant V. Kalé University of Illinois
BlueGene/L. Computer Science, University of Warwick. Source: IBM
 BlueGene/L Source: IBM 1 BlueGene/L networking BlueGene system employs various network types. Central is the torus interconnection network: 3D torus with wrap-around. Each node connects to six neighbours
BlueGene/L Source: IBM 1 BlueGene/L networking BlueGene system employs various network types. Central is the torus interconnection network: 3D torus with wrap-around. Each node connects to six neighbours
A Study of High Performance Computing and the Cray SV1 Supercomputer. Michael Sullivan TJHSST Class of 2004
 A Study of High Performance Computing and the Cray SV1 Supercomputer Michael Sullivan TJHSST Class of 2004 June 2004 0.1 Introduction A supercomputer is a device for turning compute-bound problems into
A Study of High Performance Computing and the Cray SV1 Supercomputer Michael Sullivan TJHSST Class of 2004 June 2004 0.1 Introduction A supercomputer is a device for turning compute-bound problems into
Scalable In-memory Checkpoint with Automatic Restart on Failures
 Scalable In-memory Checkpoint with Automatic Restart on Failures Xiang Ni, Esteban Meneses, Laxmikant V. Kalé Parallel Programming Laboratory University of Illinois at Urbana-Champaign November, 2012 8th
Scalable In-memory Checkpoint with Automatic Restart on Failures Xiang Ni, Esteban Meneses, Laxmikant V. Kalé Parallel Programming Laboratory University of Illinois at Urbana-Champaign November, 2012 8th
Operating Systems: Internals and Design Principles. Chapter 2 Operating System Overview Seventh Edition By William Stallings
 Operating Systems: Internals and Design Principles Chapter 2 Operating System Overview Seventh Edition By William Stallings Operating Systems: Internals and Design Principles Operating systems are those
Operating Systems: Internals and Design Principles Chapter 2 Operating System Overview Seventh Edition By William Stallings Operating Systems: Internals and Design Principles Operating systems are those
Introduction to Parallel Computing
 Portland State University ECE 588/688 Introduction to Parallel Computing Reference: Lawrence Livermore National Lab Tutorial https://computing.llnl.gov/tutorials/parallel_comp/ Copyright by Alaa Alameldeen
Portland State University ECE 588/688 Introduction to Parallel Computing Reference: Lawrence Livermore National Lab Tutorial https://computing.llnl.gov/tutorials/parallel_comp/ Copyright by Alaa Alameldeen
Parallel and High Performance Computing CSE 745
 Parallel and High Performance Computing CSE 745 1 Outline Introduction to HPC computing Overview Parallel Computer Memory Architectures Parallel Programming Models Designing Parallel Programs Parallel
Parallel and High Performance Computing CSE 745 1 Outline Introduction to HPC computing Overview Parallel Computer Memory Architectures Parallel Programming Models Designing Parallel Programs Parallel
A Framework for Collective Personalized Communication
 A Framework for Collective Personalized Communication Laxmikant V. Kalé, Sameer Kumar Department of Computer Science University of Illinois at Urbana-Champaign {kale,skumar2}@cs.uiuc.edu Krishnan Varadarajan
A Framework for Collective Personalized Communication Laxmikant V. Kalé, Sameer Kumar Department of Computer Science University of Illinois at Urbana-Champaign {kale,skumar2}@cs.uiuc.edu Krishnan Varadarajan
Motivation for Parallelism. Motivation for Parallelism. ILP Example: Loop Unrolling. Types of Parallelism
 Motivation for Parallelism Motivation for Parallelism The speed of an application is determined by more than just processor speed. speed Disk speed Network speed... Multiprocessors typically improve the
Motivation for Parallelism Motivation for Parallelism The speed of an application is determined by more than just processor speed. speed Disk speed Network speed... Multiprocessors typically improve the
Multiprocessor scheduling
 Chapter 10 Multiprocessor scheduling When a computer system contains multiple processors, a few new issues arise. Multiprocessor systems can be categorized into the following: Loosely coupled or distributed.
Chapter 10 Multiprocessor scheduling When a computer system contains multiple processors, a few new issues arise. Multiprocessor systems can be categorized into the following: Loosely coupled or distributed.
Scalable Computing: Practice and Experience Volume 8, Number 4, pp
 Scalable Computing: Practice and Experience Volume 8, Number 4, pp. 395 410. http://www.scpe.org ISSN 1895-1767 2007 SWPS THROUGHPUT IMPROVEMENT OF MOLECULAR DYNAMICS SIMULATIONS USING RECONFIGURABLE COMPUTING
Scalable Computing: Practice and Experience Volume 8, Number 4, pp. 395 410. http://www.scpe.org ISSN 1895-1767 2007 SWPS THROUGHPUT IMPROVEMENT OF MOLECULAR DYNAMICS SIMULATIONS USING RECONFIGURABLE COMPUTING
Dr. Gengbin Zheng and Ehsan Totoni. Parallel Programming Laboratory University of Illinois at Urbana-Champaign
 Dr. Gengbin Zheng and Ehsan Totoni Parallel Programming Laboratory University of Illinois at Urbana-Champaign April 18, 2011 A function level simulator for parallel applications on peta scale machine An
Dr. Gengbin Zheng and Ehsan Totoni Parallel Programming Laboratory University of Illinois at Urbana-Champaign April 18, 2011 A function level simulator for parallel applications on peta scale machine An
Lecture 9: MIMD Architectures
 Lecture 9: MIMD Architectures Introduction and classification Symmetric multiprocessors NUMA architecture Clusters Zebo Peng, IDA, LiTH 1 Introduction A set of general purpose processors is connected together.
Lecture 9: MIMD Architectures Introduction and classification Symmetric multiprocessors NUMA architecture Clusters Zebo Peng, IDA, LiTH 1 Introduction A set of general purpose processors is connected together.
HPX. High Performance ParalleX CCT Tech Talk Series. Hartmut Kaiser
 HPX High Performance CCT Tech Talk Hartmut Kaiser (hkaiser@cct.lsu.edu) 2 What s HPX? Exemplar runtime system implementation Targeting conventional architectures (Linux based SMPs and clusters) Currently,
HPX High Performance CCT Tech Talk Hartmut Kaiser (hkaiser@cct.lsu.edu) 2 What s HPX? Exemplar runtime system implementation Targeting conventional architectures (Linux based SMPs and clusters) Currently,
High Performance Computing
 The Need for Parallelism High Performance Computing David McCaughan, HPC Analyst SHARCNET, University of Guelph dbm@sharcnet.ca Scientific investigation traditionally takes two forms theoretical empirical
The Need for Parallelism High Performance Computing David McCaughan, HPC Analyst SHARCNET, University of Guelph dbm@sharcnet.ca Scientific investigation traditionally takes two forms theoretical empirical
Chapter 20: Database System Architectures
 Chapter 20: Database System Architectures Chapter 20: Database System Architectures Centralized and Client-Server Systems Server System Architectures Parallel Systems Distributed Systems Network Types
Chapter 20: Database System Architectures Chapter 20: Database System Architectures Centralized and Client-Server Systems Server System Architectures Parallel Systems Distributed Systems Network Types
Introduction to Parallel Performance Engineering
 Introduction to Parallel Performance Engineering Markus Geimer, Brian Wylie Jülich Supercomputing Centre (with content used with permission from tutorials by Bernd Mohr/JSC and Luiz DeRose/Cray) Performance:
Introduction to Parallel Performance Engineering Markus Geimer, Brian Wylie Jülich Supercomputing Centre (with content used with permission from tutorials by Bernd Mohr/JSC and Luiz DeRose/Cray) Performance:
A CASE STUDY OF COMMUNICATION OPTIMIZATIONS ON 3D MESH INTERCONNECTS
 A CASE STUDY OF COMMUNICATION OPTIMIZATIONS ON 3D MESH INTERCONNECTS Abhinav Bhatele, Eric Bohm, Laxmikant V. Kale Parallel Programming Laboratory Euro-Par 2009 University of Illinois at Urbana-Champaign
A CASE STUDY OF COMMUNICATION OPTIMIZATIONS ON 3D MESH INTERCONNECTS Abhinav Bhatele, Eric Bohm, Laxmikant V. Kale Parallel Programming Laboratory Euro-Par 2009 University of Illinois at Urbana-Champaign
Chapter 18: Database System Architectures.! Centralized Systems! Client--Server Systems! Parallel Systems! Distributed Systems!
 Chapter 18: Database System Architectures! Centralized Systems! Client--Server Systems! Parallel Systems! Distributed Systems! Network Types 18.1 Centralized Systems! Run on a single computer system and
Chapter 18: Database System Architectures! Centralized Systems! Client--Server Systems! Parallel Systems! Distributed Systems! Network Types 18.1 Centralized Systems! Run on a single computer system and
NAMD Performance Benchmark and Profiling. February 2012
 NAMD Performance Benchmark and Profiling February 2012 Note The following research was performed under the HPC Advisory Council activities Participating vendors: AMD, Dell, Mellanox Compute resource -
NAMD Performance Benchmark and Profiling February 2012 Note The following research was performed under the HPC Advisory Council activities Participating vendors: AMD, Dell, Mellanox Compute resource -
Dynamic Load Partitioning Strategies for Managing Data of Space and Time Heterogeneity in Parallel SAMR Applications
 Dynamic Load Partitioning Strategies for Managing Data of Space and Time Heterogeneity in Parallel SAMR Applications Xiaolin Li and Manish Parashar The Applied Software Systems Laboratory Department of
Dynamic Load Partitioning Strategies for Managing Data of Space and Time Heterogeneity in Parallel SAMR Applications Xiaolin Li and Manish Parashar The Applied Software Systems Laboratory Department of
ParalleX. A Cure for Scaling Impaired Parallel Applications. Hartmut Kaiser
 ParalleX A Cure for Scaling Impaired Parallel Applications Hartmut Kaiser (hkaiser@cct.lsu.edu) 2 Tianhe-1A 2.566 Petaflops Rmax Heterogeneous Architecture: 14,336 Intel Xeon CPUs 7,168 Nvidia Tesla M2050
ParalleX A Cure for Scaling Impaired Parallel Applications Hartmut Kaiser (hkaiser@cct.lsu.edu) 2 Tianhe-1A 2.566 Petaflops Rmax Heterogeneous Architecture: 14,336 Intel Xeon CPUs 7,168 Nvidia Tesla M2050
Designing Parallel Programs. This review was developed from Introduction to Parallel Computing
 Designing Parallel Programs This review was developed from Introduction to Parallel Computing Author: Blaise Barney, Lawrence Livermore National Laboratory references: https://computing.llnl.gov/tutorials/parallel_comp/#whatis
Designing Parallel Programs This review was developed from Introduction to Parallel Computing Author: Blaise Barney, Lawrence Livermore National Laboratory references: https://computing.llnl.gov/tutorials/parallel_comp/#whatis
Multicore Computing and Scientific Discovery
 scientific infrastructure Multicore Computing and Scientific Discovery James Larus Dennis Gannon Microsoft Research In the past half century, parallel computers, parallel computation, and scientific research
scientific infrastructure Multicore Computing and Scientific Discovery James Larus Dennis Gannon Microsoft Research In the past half century, parallel computers, parallel computation, and scientific research
Scalable Replay with Partial-Order Dependencies for Message-Logging Fault Tolerance
 Scalable Replay with Partial-Order Dependencies for Message-Logging Fault Tolerance Jonathan Lifflander*, Esteban Meneses, Harshitha Menon*, Phil Miller*, Sriram Krishnamoorthy, Laxmikant V. Kale* jliffl2@illinois.edu,
Scalable Replay with Partial-Order Dependencies for Message-Logging Fault Tolerance Jonathan Lifflander*, Esteban Meneses, Harshitha Menon*, Phil Miller*, Sriram Krishnamoorthy, Laxmikant V. Kale* jliffl2@illinois.edu,
Scalable Interaction with Parallel Applications
 Scalable Interaction with Parallel Applications Filippo Gioachin Chee Wai Lee Laxmikant V. Kalé Department of Computer Science University of Illinois at Urbana-Champaign Outline Overview Case Studies Charm++
Scalable Interaction with Parallel Applications Filippo Gioachin Chee Wai Lee Laxmikant V. Kalé Department of Computer Science University of Illinois at Urbana-Champaign Outline Overview Case Studies Charm++
Multi-Domain Pattern. I. Problem. II. Driving Forces. III. Solution
 Multi-Domain Pattern I. Problem The problem represents computations characterized by an underlying system of mathematical equations, often simulating behaviors of physical objects through discrete time
Multi-Domain Pattern I. Problem The problem represents computations characterized by an underlying system of mathematical equations, often simulating behaviors of physical objects through discrete time
Performance of relational database management
 Building a 3-D DRAM Architecture for Optimum Cost/Performance By Gene Bowles and Duke Lambert As systems increase in performance and power, magnetic disk storage speeds have lagged behind. But using solidstate
Building a 3-D DRAM Architecture for Optimum Cost/Performance By Gene Bowles and Duke Lambert As systems increase in performance and power, magnetic disk storage speeds have lagged behind. But using solidstate
Objective. We will study software systems that permit applications programs to exploit the power of modern high-performance computers.
 CS 612 Software Design for High-performance Architectures 1 computers. CS 412 is desirable but not high-performance essential. Course Organization Lecturer:Paul Stodghill, stodghil@cs.cornell.edu, Rhodes
CS 612 Software Design for High-performance Architectures 1 computers. CS 412 is desirable but not high-performance essential. Course Organization Lecturer:Paul Stodghill, stodghil@cs.cornell.edu, Rhodes
Accelerating Scientific Applications with GPUs
 Accelerating Scientific Applications with GPUs John Stone Theoretical and Computational Biophysics Group University of Illinois at Urbana-Champaign Research/gpu/ Workshop on Programming Massively Parallel
Accelerating Scientific Applications with GPUs John Stone Theoretical and Computational Biophysics Group University of Illinois at Urbana-Champaign Research/gpu/ Workshop on Programming Massively Parallel
Parallel Programming Patterns Overview and Concepts
 Parallel Programming Patterns Overview and Concepts Partners Funding Reusing this material This work is licensed under a Creative Commons Attribution- NonCommercial-ShareAlike 4.0 International License.
Parallel Programming Patterns Overview and Concepts Partners Funding Reusing this material This work is licensed under a Creative Commons Attribution- NonCommercial-ShareAlike 4.0 International License.
Analyzing the Performance of IWAVE on a Cluster using HPCToolkit
 Analyzing the Performance of IWAVE on a Cluster using HPCToolkit John Mellor-Crummey and Laksono Adhianto Department of Computer Science Rice University {johnmc,laksono}@rice.edu TRIP Meeting March 30,
Analyzing the Performance of IWAVE on a Cluster using HPCToolkit John Mellor-Crummey and Laksono Adhianto Department of Computer Science Rice University {johnmc,laksono}@rice.edu TRIP Meeting March 30,
Performance Analysis and Petascaling Enabling of GROMACS
 Fabio Affinito (a), Andrew Emerson (a) Leandar Litov (b), Peicho Petkov, (b) Rossen Apostolov (c) (d), Lilit Axner (c) Berk Hess (d) and Erik Lindahl (d) Maria Francesca Iozzi (e) a) CINECA Supercomputing,
Fabio Affinito (a), Andrew Emerson (a) Leandar Litov (b), Peicho Petkov, (b) Rossen Apostolov (c) (d), Lilit Axner (c) Berk Hess (d) and Erik Lindahl (d) Maria Francesca Iozzi (e) a) CINECA Supercomputing,
3/24/2014 BIT 325 PARALLEL PROCESSING ASSESSMENT. Lecture Notes:
 BIT 325 PARALLEL PROCESSING ASSESSMENT CA 40% TESTS 30% PRESENTATIONS 10% EXAM 60% CLASS TIME TABLE SYLLUBUS & RECOMMENDED BOOKS Parallel processing Overview Clarification of parallel machines Some General
BIT 325 PARALLEL PROCESSING ASSESSMENT CA 40% TESTS 30% PRESENTATIONS 10% EXAM 60% CLASS TIME TABLE SYLLUBUS & RECOMMENDED BOOKS Parallel processing Overview Clarification of parallel machines Some General
Achieving Strong Scaling with NAMD on Blue Gene/L
 Achieving Strong Scaling with NAMD on Blue Gene/L Sameer Kumar 1, Chao Huang 2, Gheorghe Almasi 1, Laxmikant V. Kalé 2 1 IBM T.J. Watson Research Center 2 University of Illinois at Urbana-Champaign Yorktown
Achieving Strong Scaling with NAMD on Blue Gene/L Sameer Kumar 1, Chao Huang 2, Gheorghe Almasi 1, Laxmikant V. Kalé 2 1 IBM T.J. Watson Research Center 2 University of Illinois at Urbana-Champaign Yorktown
Review: Creating a Parallel Program. Programming for Performance
 Review: Creating a Parallel Program Can be done by programmer, compiler, run-time system or OS Steps for creating parallel program Decomposition Assignment of tasks to processes Orchestration Mapping (C)
Review: Creating a Parallel Program Can be done by programmer, compiler, run-time system or OS Steps for creating parallel program Decomposition Assignment of tasks to processes Orchestration Mapping (C)
CHRONO::HPC DISTRIBUTED MEMORY FLUID-SOLID INTERACTION SIMULATIONS. Felipe Gutierrez, Arman Pazouki, and Dan Negrut University of Wisconsin Madison
 CHRONO::HPC DISTRIBUTED MEMORY FLUID-SOLID INTERACTION SIMULATIONS Felipe Gutierrez, Arman Pazouki, and Dan Negrut University of Wisconsin Madison Support: Rapid Innovation Fund, U.S. Army TARDEC ASME
CHRONO::HPC DISTRIBUTED MEMORY FLUID-SOLID INTERACTION SIMULATIONS Felipe Gutierrez, Arman Pazouki, and Dan Negrut University of Wisconsin Madison Support: Rapid Innovation Fund, U.S. Army TARDEC ASME
VMD: Immersive Molecular Visualization and Interactive Ray Tracing for Domes, Panoramic Theaters, and Head Mounted Displays
 VMD: Immersive Molecular Visualization and Interactive Ray Tracing for Domes, Panoramic Theaters, and Head Mounted Displays John E. Stone Theoretical and Computational Biophysics Group Beckman Institute
VMD: Immersive Molecular Visualization and Interactive Ray Tracing for Domes, Panoramic Theaters, and Head Mounted Displays John E. Stone Theoretical and Computational Biophysics Group Beckman Institute
Center Extreme Scale CS Research
 Center Extreme Scale CS Research Center for Compressible Multiphase Turbulence University of Florida Sanjay Ranka Herman Lam Outline 10 6 10 7 10 8 10 9 cores Parallelization and UQ of Rocfun and CMT-Nek
Center Extreme Scale CS Research Center for Compressible Multiphase Turbulence University of Florida Sanjay Ranka Herman Lam Outline 10 6 10 7 10 8 10 9 cores Parallelization and UQ of Rocfun and CMT-Nek
Petascale Multiscale Simulations of Biomolecular Systems. John Grime Voth Group Argonne National Laboratory / University of Chicago
 Petascale Multiscale Simulations of Biomolecular Systems John Grime Voth Group Argonne National Laboratory / University of Chicago About me Background: experimental guy in grad school (LSCM, drug delivery)
Petascale Multiscale Simulations of Biomolecular Systems John Grime Voth Group Argonne National Laboratory / University of Chicago About me Background: experimental guy in grad school (LSCM, drug delivery)
Client Server & Distributed System. A Basic Introduction
 Client Server & Distributed System A Basic Introduction 1 Client Server Architecture A network architecture in which each computer or process on the network is either a client or a server. Source: http://webopedia.lycos.com
Client Server & Distributed System A Basic Introduction 1 Client Server Architecture A network architecture in which each computer or process on the network is either a client or a server. Source: http://webopedia.lycos.com
Multiprocessors and Thread-Level Parallelism. Department of Electrical & Electronics Engineering, Amrita School of Engineering
 Multiprocessors and Thread-Level Parallelism Multithreading Increasing performance by ILP has the great advantage that it is reasonable transparent to the programmer, ILP can be quite limited or hard to
Multiprocessors and Thread-Level Parallelism Multithreading Increasing performance by ILP has the great advantage that it is reasonable transparent to the programmer, ILP can be quite limited or hard to
ARCHER ecse Technical Report. Algorithmic Enhancements to the Solvaware Package for the Analysis of Hydration. 4 Technical Report (publishable)
 4 Technical Report (publishable) ARCHER ecse Technical Report Algorithmic Enhancements to the Solvaware Package for the Analysis of Hydration Technical staff: Arno Proeme (EPCC, University of Edinburgh)
4 Technical Report (publishable) ARCHER ecse Technical Report Algorithmic Enhancements to the Solvaware Package for the Analysis of Hydration Technical staff: Arno Proeme (EPCC, University of Edinburgh)
CMSC 714 Lecture 6 MPI vs. OpenMP and OpenACC. Guest Lecturer: Sukhyun Song (original slides by Alan Sussman)
 CMSC 714 Lecture 6 MPI vs. OpenMP and OpenACC Guest Lecturer: Sukhyun Song (original slides by Alan Sussman) Parallel Programming with Message Passing and Directives 2 MPI + OpenMP Some applications can
CMSC 714 Lecture 6 MPI vs. OpenMP and OpenACC Guest Lecturer: Sukhyun Song (original slides by Alan Sussman) Parallel Programming with Message Passing and Directives 2 MPI + OpenMP Some applications can
HPC Considerations for Scalable Multidiscipline CAE Applications on Conventional Linux Platforms. Author: Correspondence: ABSTRACT:
 HPC Considerations for Scalable Multidiscipline CAE Applications on Conventional Linux Platforms Author: Stan Posey Panasas, Inc. Correspondence: Stan Posey Panasas, Inc. Phone +510 608 4383 Email sposey@panasas.com
HPC Considerations for Scalable Multidiscipline CAE Applications on Conventional Linux Platforms Author: Stan Posey Panasas, Inc. Correspondence: Stan Posey Panasas, Inc. Phone +510 608 4383 Email sposey@panasas.com
High Performance Computing: Architecture, Applications, and SE Issues. Peter Strazdins
 High Performance Computing: Architecture, Applications, and SE Issues Peter Strazdins Department of Computer Science, Australian National University e-mail: peter@cs.anu.edu.au May 17, 2004 COMP1800 Seminar2-1
High Performance Computing: Architecture, Applications, and SE Issues Peter Strazdins Department of Computer Science, Australian National University e-mail: peter@cs.anu.edu.au May 17, 2004 COMP1800 Seminar2-1
Let s say I give you a homework assignment today with 100 problems. Each problem takes 2 hours to solve. The homework is due tomorrow.
 Let s say I give you a homework assignment today with 100 problems. Each problem takes 2 hours to solve. The homework is due tomorrow. Big problems and Very Big problems in Science How do we live Protein
Let s say I give you a homework assignment today with 100 problems. Each problem takes 2 hours to solve. The homework is due tomorrow. Big problems and Very Big problems in Science How do we live Protein
MPI+X on The Way to Exascale. William Gropp
 MPI+X on The Way to Exascale William Gropp http://wgropp.cs.illinois.edu Some Likely Exascale Architectures Figure 1: Core Group for Node (Low Capacity, High Bandwidth) 3D Stacked Memory (High Capacity,
MPI+X on The Way to Exascale William Gropp http://wgropp.cs.illinois.edu Some Likely Exascale Architectures Figure 1: Core Group for Node (Low Capacity, High Bandwidth) 3D Stacked Memory (High Capacity,
A Scalable Adaptive Mesh Refinement Framework For Parallel Astrophysics Applications
 A Scalable Adaptive Mesh Refinement Framework For Parallel Astrophysics Applications James Bordner, Michael L. Norman San Diego Supercomputer Center University of California, San Diego 15th SIAM Conference
A Scalable Adaptive Mesh Refinement Framework For Parallel Astrophysics Applications James Bordner, Michael L. Norman San Diego Supercomputer Center University of California, San Diego 15th SIAM Conference
Chapter 1: Introduction to Parallel Computing
 Parallel and Distributed Computing Chapter 1: Introduction to Parallel Computing Jun Zhang Laboratory for High Performance Computing & Computer Simulation Department of Computer Science University of Kentucky
Parallel and Distributed Computing Chapter 1: Introduction to Parallel Computing Jun Zhang Laboratory for High Performance Computing & Computer Simulation Department of Computer Science University of Kentucky
UCLA UCLA Previously Published Works
 UCLA UCLA Previously Published Works Title Parallel Markov chain Monte Carlo simulations Permalink https://escholarship.org/uc/item/4vh518kv Authors Ren, Ruichao Orkoulas, G. Publication Date 2007-06-01
UCLA UCLA Previously Published Works Title Parallel Markov chain Monte Carlo simulations Permalink https://escholarship.org/uc/item/4vh518kv Authors Ren, Ruichao Orkoulas, G. Publication Date 2007-06-01
Game-changing Extreme GPU computing with The Dell PowerEdge C4130
 Game-changing Extreme GPU computing with The Dell PowerEdge C4130 A Dell Technical White Paper This white paper describes the system architecture and performance characterization of the PowerEdge C4130.
Game-changing Extreme GPU computing with The Dell PowerEdge C4130 A Dell Technical White Paper This white paper describes the system architecture and performance characterization of the PowerEdge C4130.
