Slicer3 Training Compendium. Slicer3 Training Tutorial ARCTIC (v1.2)
|
|
|
- Melissa Manning
- 5 years ago
- Views:
Transcription
1 Slicer3 Training Compendium Slicer3 Training Tutorial ARCTIC (v1.2) ( ThICkness (Automatic Regional Cortical University of North Carolina, Chapel Hill: Neuro Image Research and Analysis Lab Neurodevelopmental Disorders Research Center Cedric Mathieu, Clement Vachet, Martin Styner, Heather Cody Hazlett Contact : ced.mathieu@gmail.com / cvachet@ .unc.edu -1-
2 Learning Objective Following this tutorial, you will be able to perform an individual analysis of regional cortical thickness. You will learn how to load input volumes, run the end-to-end module ARCTIC to generate cortical thickness information and display output volumes. Cortical thickness on WM surface Cortical thickness on GM surface -2-
3 Prerequisites This tutorial assumes that you have already completed the tutorial Data Loading and Visualization. Tutorials for Slicer3 are available at the following location: Slicer3 tutorials -3-
4 Materials This tutorial requires the installation of Slicer3, BatchMake wrappers, the tutorial dataset and the external modules. They are available at the following locations: ( build Slicer3 download page (Slicer nightly ( ARCTIC_BatchMake_Wrapper_1.2 )pagedownloadwrapperbatchmake ( page(arctic_tutorial_example_1.0 downloaddatasettutorial ( ARCTIC_Executables_1.2 )pagedownloadmodulesexternal ( page(unc_pediatric_brain_atlas Atlas download Disclaimer: It is the responsibility of the user of Slicer to comply with both the terms of the license and with the applicable laws, regulations, and rules. -4-
5 Materials: Tutorial dataset The tutorial dataset (ARCTIC_Tutorial_example_1.0) is a ZIP file. Unzip this file somewhere in your computer. An ARCTIC_Tutorial_example_1.0 folder will be created, containing: A pediatric case: T1-weighted and T2-weighted images. An ARTIC-Results/ directory, in which results of the tutorial example will be saved. -5-
6 Materials: External modules The executables are in a ZIP file : ARCTIC_Executables_1.2_linux32/64/Mac Unzip this file somewhere in your computer. An ARCTIC_Executables_1.2 folder will be created, containing executables needed to perform the cortical thickness analysis. To add the executables as Slicer3 external modules: - Open Slicer3 - Go to View Application Settings Module Settings - Click on the add a preset button - Select the ARCTIC_Executables_1.2 folder and confirm - Close Slicer3-6-
7 Materials: Atlas The atlas and its related files are in a ZIP file (UNC_Pediatric_Brain_Atlas). Create a pediatric-atlas-4years-sym-t1-rai folder somewhere in your computer. Unzip the ZIP file in this new folder. The pediatric-atlas-4years-sym-t1-rai folder will thus contain the atlas and its related files. You can then unzip all the images (gunzip command). -7-
8 Prerequisites Add the executables in the PATH. -tcsh usage : setenv PATH ARCTIC-Executables-Directory:Slicer3-Plugins-Directory:Slicer3-Bin-Directory:${PATH} -bash usage : export PATH=ARCTIC-Executables-Directory:Slicer3-Plugins-Directory:Slicer3-Bin-Directory:${PATH} Notice : To execute ARCTIC within Slicer3D, it is not necessary to add "Slicer3D-Bin-Directory" and "Slicer3-Bin- Directory" in the PATH. Set ARCTIC environment variable -tcsh usage : setenv BatchmakeWrapper_Dir Batchmake-Wrapper-Directory -bash usage : export BatchmakeWrapper_Dir=Batchmake-Wrapper-Directory WITH: ( ARTIC_Executables_1.2 ) ARCTIC-Executables-Directory/ : Downloaded folder Slicer3-Plugins-Directory/ : Directory containing Slicer3 plugins Nightly build version : Slicer3Dir /lib/slicer3/plugins Slicer3-Bin-Directory/ : Directory containing Slicer3 binary files Nightly build version : Slicer3Dir /lib/slicer3/bin ( ARTIC_Batchmake_Wrapper_1.2 ) Batchmake-Wrapper-Directory/ : Downloaded folder -8-
9 Set Slicer libraries variable (only for command line) Slicer-nightly-build /lib SLICERLIBPATHsetenv -tcsh usage : setenv LD_LIBRARY_PATH ${LD_LIBRARY_PATH}:${SLICERLIBPATH}/BatchMake:${SLICERLIBPATH}/bmModuleDescriptionParser:${SLICERLIB PATH}/FreeSurfer:${SLICERLIBPATH}/GenerateCLP:${SLICERLIBPATH}/GenerateLM:${SLICERLIBPATH}/IGT:${SLIC ERLIBPATH}/igtl:${SLICERLIBPATH}/InsightToolkit:${SLICERLIBPATH}/ITKCommandIO:${SLICERLIBPATH}/KWWidget s:${slicerlibpath}/loadablemodule:${slicerlibpath}/mghimageio:${slicerlibpath}/moduledescriptionparse r:${slicerlibpath}/mrml:${slicerlibpath}/mrmlidimageio:${slicerlibpath}/openigtlink:${slicerlibpat H}/Python/lib:${SLICERLIBPATH}/Qdec:${SLICERLIBPATH}/RemoteIO:${SLICERLIBPATH}/Slicer3:${SLICERLIBPATH}/ SlicerIO:${SLICERLIBPATH}/tclap:${SLICERLIBPATH}/TclTk/lib:${SLICERLIBPATH}/Teem :${SLICERLIBPATH}/vtk-5.2:${SLICERLIBPATH}/vtkITK:${SLICERLIBPATH}/vtkTeem SLICERLIBPATH= Slicer-nightly-build /libexport -bash usage : export LD_LIBRARY_PATH=${LD_LIBRARY_PATH}:${SLICERLIBPATH}/BatchMake:${SLICERLIBPATH}/bmModuleDescriptio nparser:${slicerlibpath}/freesurfer:${slicerlibpath}/generateclp:${slicerlibpath}/generatelm:${slicer LIBPATH}/IGT:${SLICERLIBPATH}/igtl:${SLICERLIBPATH}/InsightToolkit:${SLICERLIBPATH}/ITKCommandIO:${SLICE RLIBPATH}/KWWidgets:${SLICERLIBPATH}/LoadableModule:${SLICERLIBPATH}/MGHImageIO:${SLICERLIBPATH}/M oduledescriptionparser:${slicerlibpath}/mrml:${slicerlibpath}/mrmlidimageio:${slicerlibpath}/openig TLink:${SLICERLIBPATH}/Python/lib:${SLICERLIBPATH}/Qdec:${SLICERLIBPATH}/RemoteIO:${SLICERLIBPATH}/Slic er3:${slicerlibpath}/slicerio:${slicerlibpath}/tclap:${slicerlibpath}/tcltk/lib:${slicerlibpath}/teem :${SLICERLIBPATH}/vtk-5.2:${SLICERLIBPATH}/vtkITK:${SLICERLIBPATH}/vtkTee WITH: Slicer-nightly-build : path of Slicer nightly build in your computer Prerequisites -9-
10 Overview 1- Pipeline overview 2- Input images 3- Pipeline description 4- Ouput images and organisation 5- Execution within Slicer 6- Example with tutorial dataset 7- Command line execution -10-
11 Overview 1- Pipeline overview 2- Input images 3- Pipeline description 4- Ouput images and organisation 5- Execution within Slicer 6- Example with tutorial dataset 7- Command line execution -11-
12 Pipeline Overview All the tools used in the current pipeline are Slicer3 modules, some of them being UNC external modules. The user can thus perform a regional cortical thickness analysis on an individual subject within Slicer3. Two different modes can be used, depending on the input images: ( PD Raw images (T1-weighted, T2-weighted, Tissue segmentation label image -12-
13 Overview 1- Pipeline overview 2- Input images 3- Pipeline description 4- Ouput images and organisation 5- Execution within Slicer 6- Example with tutorial dataset 7- Command line execution -13-
14 Input images What you need Raw images Segmented image T1-weighted image Tissue segmentation atlas directory Optional T2-weighted image PD-weighted image Atlas raw image + its parcellation Case parcellation image Raw image Tissue segmentation label image Optional Atlas raw image + its parcellation Case parcellation image -14-
15 Overview 1- Pipeline overview 2- Input images 3- Pipeline description 4- Ouput images and organisation 5- Execution within Slicer 6- Example with tutorial dataset 7- Command line execution -15-
16 Pipeline Description If T1-weighted Image Tissue Segmentation itkems Input Volumes If Tissue Segmentation Label Image If no Parcellation If Atlas Parcellation Sparse Cortical Thickness CortThick If Case Parcellation Sparse Cortical Thickness ( option CortThick (w. Parcellation Deformable Registration SegPostProcess RegisterImages ResampleVolume2 Sparse Cortical Thickness ( option CortThick (w. Parcellation Statistics ImageMath/ImageStat Mesh creation ModelMaker Mrml scene creation (for quality control) -16-
17 1. Tissue segmentation Module : itkems (UNC Slicer3 external module) 2. Regional atlas deformable registration 3.1. Skull stripping ( module Module : SegPostProcess (UNC Slicer3 external 3.2. Deformable registration of T1-weighted atlas Module : RegisterImages (Slicer3 module) 3.3. Applying transformation to its parcellation map Module : ResampleVolume2 (Slicer3 module) 3. Sparse and asymmetric Cortical Thickness Module : CortThick (UNC Slicer3 module) 4. Statistics Modules : ImageMath, ImageStat (UNC Slicer3 external modules) 5. Mesh Creation Module : ModelMaker (Slicer3 module) 6. Mrml scene Creation Pipeline Description -17-
18 Pipeline Description ( module Tissue segmentation (itkems external Probabilistic atlas-based automatic tissue segmentation via an Expectation-Maximization scheme. ItkEMS also performs an intensity inhomogeneity correction of the input image that removes gradual variations in the image intensities mainly due to RF coil imperfection Input_T1-Image.nrrd Image_corrected_EMS.nrr d Image_labels_EMS.nrrd -18-
19 Pipeline Description ( module Skull Stripping (SegPostProcess external This step is performed using the previously computed tissue segmentation label image. Image_corrected_EMS.nrr d Image_corrected_EMS_stripped.nrrd -19-
20 Pipeline Description Deformable registration of T1-weighted atlas ( module (RegisterImages B-spline pipeline registration. A transformation file is created and will be used by the next step. Module link TransformFile.txt Atlas.nrr d AtlasRegistered_Image_corrected_EMS_stripped.nrrd -20-
21 Pipeline Description mapparcellation atlasthetotransformationapplying ( module (ResampleVolume2 Module link : TransformFile.txt Parcellation.nrrd ParcellationRegistered_Image_corrected_EMS_stripped.nrrd -21-
22 Pipeline Description ( module Cortical Thickness (CortThick external Sparse and asymmetric local cortical thickness Optional Outputs WM_AvgBoundary.nrrd Regional Cortical Thickness information Image_labels_EMS.nrrd GM_AvgBoundary.nrrd -22-
23 Pipeline Description ( modules Statistics (ImageStat and ImageMath Both modules are used to generate volume information in the following files : - TissueSegmentationVolumes.csv White matter, gray matter and CSF volumes. ( provided - ParcellationMapVolumes.csv (if the parcellation image is White matter, gray matter and CSF volumes per lobe. Notice : values are in mm -23-
24 Pipeline Description ( module Mesh creation (ModelMaker Module link : Two meshes are created : - WM_Surface.vtk White matter mesh. - GM_Surface.vtk Gray matter mesh. -24-
25 Pipeline Description Mrml scene creation A Mrml scene is created to display all the steps of the pipeline. There is one snapshot per step. This Mrml scene is created to make a quality control. -25-
26 Overview 1- Pipeline overview 2- Input images 3- Pipeline description 4- Output images and organization 5- Execution within Slicer 6- Example with tutorial dataset 7- Command line execution -26-
27 Output and Organisation Input Files Directory/ Input images ARCTIC/ 2 BatchMake Scripts TissueSegmentation/ InputVolumes_corrected_EMS.nrrd InputVolume_labels_EMS.nrrd WM/GM/CSF_Map.nrrd Registration/ Image_corrected_EMS_stripped.nrrd AtlasRegistered_Image_corrected_EMS_stripped.nrrd AtlasTransform_Image_corrected_EMS_stripped.txt ParcellationRegistered_Image_corrected_EMS_stripped.nrrd CorticalThickness/ If the parcellation image is not provided: Image_labels_EMS_WhiteMatDistanceMap.csv If the parcellation image is provided: Image_labels_EMS_WhiteMatDistanceMap_par.csv Image_labels_EMS_WhiteMatDistanceMap_par_array.csv TissueSegmentationVolumes.csv ParcellationMapVolumes.csv Stat/ Mesh/ WM_Surface.vtk GM_Surface.vtk -27-
28 Overview 1- Pipeline overview 2- Input images 3- Pipeline description 4- Ouput images and organisation 5- Execution within Slicer 6- Example with tutorial dataset 7- Command line execution -28-
29 Execution within Slicer Load input images Demonstration with «Raw Images» Demonstration with «Segmented Image» -29-
30 Demonstration : Load the input images 1 How to load raw images (case and atlas)? Select the image in the browser 2- Set the image origin as «centered» 3- Click on «Apply» to load How to load parcellation and label images? 1 1- Select the image in the browser 2- Set the image origin as «centered» 3- Check the «label map» button 4- Click on «Apply» to load
31 Demonstration in Slicer Load input images Demonstration with «Raw Images» Demonstration with «Segmented Image» Parcellation option Advanced parameters -31-
32 Demonstration : Raw Images Input 1 1 ( Modules 1- Select the «ARCTIC» module (in All 2- Add the T1-weighted image 3- Set the Tissue Segmentation Atlas Directory for the tissue segmentation 4- Set the output directory 5- Set a prefix which will be added to all the outputs 6- Click on the «Apply» button to process the data
33 Demonstration : Raw Images Input 1 Verifications / Options If available, set the T2 and/or PD-weighted images to improve the tissue segmentation 2- Check the tissue segmentation atlas type (T1- weighted or T2-weighted image) 3 3- Set the output images to be displayed in Slicer («(«Create a new volume» instead of «None -33-
34 Demonstration in Slicer Load input images Demonstration with «Raw Images» Demonstration with «Segmented Image» Parcellation option Advanced parameters -34-
35 Demonstration : Segmented Image Input 2 1 ( Modules 1- Select the «ARCTIC» module (in All 2- Set the tissue segmentation label image and check the related tissue labels 2 3- Set its raw image (T1-weighted, T2-weighted, PDweighted)and change the orientation if necessary 3 4- Set the output directory 5- Set a prefix which will be added to all the outputs Click on the «Apply» button to process the data 6-35-
36 Options Demonstration : Segmented Image Input 2 1- Set the output images to be displayed in Slicer («(«Create a new volume» instead of «None 1 Cortical Thickness on WM Boundary Cortical Thickness on GM Boundary -36-
37 Demonstration in Slicer Load input images Demonstration with «Raw Images» Demonstration with «Segmented Image» Parcellation option Advanced parameters -37-
38 Parcellation options Parcellation options If you want to perform a lobar cortical thickness analysis, choose between the two possibilities a- Add a parcellation image which is defined in the input coordinate space («Case («Parcellation Image b- Add the atlas raw image and its parcellation, defined in the atlas coordinate space («(«Atlas Parcellation Image a b -38-
39 Demonstration in Slicer Load input images Demonstration with «Raw Images» Demonstration with «Segmented Image» Parcellation option Advanced parameters -39-
40 Advanced parameters Tissue segmentation parameters a- Filter options: specifies smoothing parameters prior to the segmentation b- Priors weighting the tissue classes for the segmentation c- Atlas warping options: - No atlas warping: - Unchecked by default: atlas to subject B-Spline registration is performed - Checked: atlas to subject affine registration is performed warping instead of the - Grid size X,Y,Z: grid controls points for atlas warping a b Skull stripping parameters a- Check to apply a dilation of the mask (necessary if the ( quality tissue segmentation has a low c Cortical Thickness parameters a a- To remove the interpolation of the cortical thickness or set the threshold used to match the cortical thickness map with the parcellation a Atlas registration parameters a- Different initialization methods a -40-
41 Overview 1- Pipeline overview 2- Input images 3- Pipeline description 4- Ouput images and organisation 5- Execution within Slicer 6- Example with tutorial dataset 7- Command line execution -41-
42 Example with tutorial dataset Load input images Run ARCTIC -42-
43 Load input images In Slicer, select the module «Volumes» to load the input images. Then click on the «Select Volume File» button to load the images. -43-
44 Load input images A new window Open Volume File is now open. Select the «ARCTIC_Tutorial_example» directory. Select the «pediatric_t1_rai.nrrd» file in the Data directory and click on «Open». -44-
45 Load input images Now, select the Image Origin as «Centered». And click on «Apply». -45-
46 Load input images The first image is now loaded. You can check it in the «Active Volume» widget. -46-
47 Load input images Apply the same steps to load the T2-weighted and atlas images. One can find the T2-weighted image in the same directory than the T1- weighted one. The atlas image, named «template-stripped.nrrd» is in the «pediatricatlas-4years-sym-t1-rai» directory. -47-
48 Load input images Now we will load the parcellation image. Click on the «Select Volume File» button to load the parcellation. -48-
49 Load input images»theselect. opennowis FileVolumeOpen windownewa pediatric-atlas- 4years-sym-T1-RAI» directory. Select the «Parcellation.nrrd» file and click on «Open». -49-
50 Load input images Now, select the Image Origin as «Centered». Then, check the «Label Map» case to load the parcellation as a label image. And click on «Apply». -50-
51 Load input images The dataset is now loaded. You can check it in the «Active Volume» widget while displaying the 4 images. -51-
52 Example with tutorial dataset Input images loading ARCTIC execution -52-
53 Module execution 1 1- Select the «ARCTIC» module (in All ( Modules 2- Set the T1-weighted images ( pediatric_t1_rai.gipl ) 3- Click on the «Tissue Segmentation Atlas Directory» button
54 Module execution A new window is now open to select the tissue segmentation atlas. Search and select the «pediatric-atlas-4years-sym- T1-RAI/» folder. Click on the «OK» button to confirm. -54-
55 Module execution 1- Select «Create a new volume» to display output images 2- Click on the «Cortical Thickness Results Directory» button
56 Module execution 1- Set the prefix for the outputs
57 Module execution Select the «ARCTIC-Results» folder in the ARCTIC_Tutorial_Example directory Click on the «Save» button to confirm your choice. -57-
58 Module execution 1- Add the «Parcellation.gipl» as Atlas Parcellation Image, and the «template-stripped.gipl» as Atlas Image 2- Click on the «Apply» button to start the process
59 Module execution Once the execution is finished, several images are displayed within Slicer. You can compare your images with the following ones to perform a quick quality control. Cortical Thickness on WM Boundary Cortical Thickness on GM Boundary -59-
60 Overview 1- Pipeline overview 2- Input images 3- Pipeline description 4- Ouput images and organisation 5- Execution within Slicer 6- Example with tutorial dataset 7- Command line execution -60-
61 Use the command line «Raw Images» Mode Global analysis ARCTIC --T1 Image_T1.nrrd --segatlasdir TissueSegmentationAtlasDirectory/ Lobar cortical thickness analysis If the atlas raw image and its parcellation are provided: ARCTIC --T1 Image_T1.nrrd --segatlasdir TissueSegmentationAtlasDirectory/ --atlas Atlas.nrrd --atlasparcellation Parcellation.nrrd If the case parcellation image is provided: ARCTIC --T1 Image_T1.nrrd --segatlasdir TissueSegmentationAtlasDirectory/ --caseparcellation CaseRegisteredParcellation.nrrd -61-
62 Use the command line «Raw Images» Mode Complementary flags --T2 Image_T2.gipl / --pd Image_PD.gipl : T2 and/or Pd-weighted image(s) can be added to improve tissue segmentation --atlasorientation RAI : if the orientation of your atlas/parcellation is different than the default value (RAI), add this flag to set the right orientation --atlastype T1 : if the type of your tissue segmentation atlas is different than T1 (default value) --outputdir output_directory/ : if you want to select the output directory, add this flag and indicate the path an existing folder --IDNumber prefix : if you want to add a prefix before all the outputs generated by the pipeline --SaveWM WMCorticalThicknessMap.gipl / --SaveGM GMCorticalThicknessMap.gipl : those flags are used to save a volume with information of the average cortical thickness on WM/GM boundary(ies), the filename needed is a path with the name of the output volume -62-
63 Use the command line «Segmented Image» Mode Global analysis ARCTIC --label TissueSegmentationImage.nrrd --rawimage Image_T1.nrrd Lobar cortical thickness analysis If the atlas raw image and its parcellation are provided: ARCTIC --label TissueSegmentationImage.nrrd --rawimage Image_T1.nrrd --atlas Atlas.nrrd --atlasparcellation Parcellation.nrrd If the case parcellation image is provided: ARCTIC --label TissueSegmentationImage.nrrd --rawimage Image_T1.nrrd --caseparcellation CaseRegisteredParcellation.nrrd -63-
64 Use the command line «Segmented Image» Mode Complementary flags --WMLabel 1 / --GMLabel 2 / --CSFLabel 3 : if your label are different than the default value --outputdir output_directory/ : if you want to select the output directory, add this flag and indicate the path an existing folder --IDNumber prefix : if you want to add a prefix before all the outputs generated by the pipeline --SaveWM WMCorticalThicknessMap.gipl / --SaveGM GMCorticalThicknessMap.gipl : those flags are used to save a volume with information of the average cortical thickness on WM/GM boundary(ies), the filename needed is a path with the name of the output volum -64-
65 Conclusion Slicer3 toolkit provides an accessible and versatile platform to conduct image processing of MRI data, in this case, regional cortical thickness analysis using ARCTIC. Thanks to this tutorial you are now ready to perform a regional cortical thickness analysis on your own dataset. -65-
66 Acknowledgements NIH U54EB UNC Chapel Hill Neurodevelopmental Disorders Research Center Neuro Image Research Analysis Laboratories -66-
67 Acknowledgements ( ModelMaker ) Nicole Aucoin, Surgical Planning Lab, Boston ( RegisterImages ) Steven Aylward, Kitware Inc. François Budin, Psychiatry Neuroimaging Laboratory, Boston (ResampleVolume2) ( Batchmake ) Julien Jomier, Kitware Inc. Steve Pieper, Isomics Inc. Joseph Piven, Neurodevelopmental Disorders Research Center, UNC Chapel Hill ( itkems ) Marcel Prastawa, Scientific Computing and Imaging Institute, Utah Delphine Ribes, Sylvain Gouttard, Cassian Marc, NIRAL, UNC (CortThick) -67-
Automatic SPHARM Shape Analysis in 3D Slicer
 NA-MIC http://na-mic.org Automatic SPHARM Shape Analysis in 3D Slicer Corentin Hamel, Clement Vachet, Beatriz Paniagua, Nicolas Augier, Martin Styner University of North Carolina, Chapel Hill: Neuro Image
NA-MIC http://na-mic.org Automatic SPHARM Shape Analysis in 3D Slicer Corentin Hamel, Clement Vachet, Beatriz Paniagua, Nicolas Augier, Martin Styner University of North Carolina, Chapel Hill: Neuro Image
Slicer3 Training Tutorial Using EM Segmenter with Non- Human Primate Images
 Slicer3 Training Compendium Slicer3 Training Tutorial Using EM Segmenter with Non- Human Primate Images Vidya Rajagopalan Christopher Wyatt BioImaging Systems Lab Dept. of Electrical Engineering Virginia
Slicer3 Training Compendium Slicer3 Training Tutorial Using EM Segmenter with Non- Human Primate Images Vidya Rajagopalan Christopher Wyatt BioImaging Systems Lab Dept. of Electrical Engineering Virginia
Subcortical Structure Segmentation using Probabilistic Atlas Priors
 Subcortical Structure Segmentation using Probabilistic Atlas Priors Sylvain Gouttard 1, Martin Styner 1,2, Sarang Joshi 3, Brad Davis 2, Rachel G. Smith 1, Heather Cody Hazlett 1, Guido Gerig 1,2 1 Department
Subcortical Structure Segmentation using Probabilistic Atlas Priors Sylvain Gouttard 1, Martin Styner 1,2, Sarang Joshi 3, Brad Davis 2, Rachel G. Smith 1, Heather Cody Hazlett 1, Guido Gerig 1,2 1 Department
3D Visualization of FreeSurfer Data
 3D Visualization of FreeSurfer Data Sonia Pujol, Ph.D. Silas Mann, B.Sc. Randy Gollub, MD., Ph.D. Surgical Planning Laboratory Athinoula A. Martinos Center Harvard University Acknowledgements NIH U54EB005149
3D Visualization of FreeSurfer Data Sonia Pujol, Ph.D. Silas Mann, B.Sc. Randy Gollub, MD., Ph.D. Surgical Planning Laboratory Athinoula A. Martinos Center Harvard University Acknowledgements NIH U54EB005149
Automatic Brain Segmentation in Rhesus Monkeys
 Automatic Brain Segmentation in Rhesus Monkeys Martin Styner a,b, Rebecca Knickmeyer b, Sarang Joshi c, Christopher Coe d, Sarah J Short d, John Gilmore b a Department of Psychiatry, University of North
Automatic Brain Segmentation in Rhesus Monkeys Martin Styner a,b, Rebecca Knickmeyer b, Sarang Joshi c, Christopher Coe d, Sarah J Short d, John Gilmore b a Department of Psychiatry, University of North
EMSegmenter Tutorial (Advanced Mode)
 EMSegmenter Tutorial (Advanced Mode) Dominique Belhachemi Section of Biomedical Image Analysis Department of Radiology University of Pennsylvania 1/65 Overview The goal of this tutorial is to apply the
EMSegmenter Tutorial (Advanced Mode) Dominique Belhachemi Section of Biomedical Image Analysis Department of Radiology University of Pennsylvania 1/65 Overview The goal of this tutorial is to apply the
NIH Public Access Author Manuscript Proc Soc Photo Opt Instrum Eng. Author manuscript; available in PMC 2011 September 7.
 NIH Public Access Author Manuscript Published in final edited form as: Proc Soc Photo Opt Instrum Eng. 2011 March ; 7962: 796225-1 796225-7. doi:10.1117/12.878405. Automatic Skull-stripping of Rat MRI/DTI
NIH Public Access Author Manuscript Published in final edited form as: Proc Soc Photo Opt Instrum Eng. 2011 March ; 7962: 796225-1 796225-7. doi:10.1117/12.878405. Automatic Skull-stripping of Rat MRI/DTI
Detecting White Matter Lesions in Lupus
 Slicer3 Training Compendium Detecting White Matter Lesions in Lupus Version 2.0 1/6/2009 H. Jeremy Bockholt Mark Scully -1- Learning objective Following this tutorial, you ll be able to load scans into
Slicer3 Training Compendium Detecting White Matter Lesions in Lupus Version 2.0 1/6/2009 H. Jeremy Bockholt Mark Scully -1- Learning objective Following this tutorial, you ll be able to load scans into
Automatic corpus callosum segmentation using a deformable active Fourier contour model
 Automatic corpus callosum segmentation using a deformable active Fourier contour model Clement Vachet a, Benjamin Yvernault a, Kshamta Bhatt a, Rachel G. Smith a,b, Guido Gerig c, Heather Cody Hazlett
Automatic corpus callosum segmentation using a deformable active Fourier contour model Clement Vachet a, Benjamin Yvernault a, Kshamta Bhatt a, Rachel G. Smith a,b, Guido Gerig c, Heather Cody Hazlett
Diffusion Imaging Quality Control with DTIPrep
 NA-MIC National Alliance for Medical Image Computing http://na-mic.org Diffusion Imaging Quality Control with DTIPrep Martin Styner Francois Budin University of North Carolina Neuro Image Research and
NA-MIC National Alliance for Medical Image Computing http://na-mic.org Diffusion Imaging Quality Control with DTIPrep Martin Styner Francois Budin University of North Carolina Neuro Image Research and
Subcortical Structure Segmentation using Probabilistic Atlas Priors
 Subcortical Structure Segmentation using Probabilistic Atlas Priors Sylvain Gouttard a, Martin Styner a,b, Sarang Joshi c, Rachel G. Smith a, Heather Cody a, Guido Gerig a,b a Department of Psychiatry,
Subcortical Structure Segmentation using Probabilistic Atlas Priors Sylvain Gouttard a, Martin Styner a,b, Sarang Joshi c, Rachel G. Smith a, Heather Cody a, Guido Gerig a,b a Department of Psychiatry,
Automated MR Image Analysis Pipelines
 Automated MR Image Analysis Pipelines Andy Simmons Centre for Neuroimaging Sciences, Kings College London Institute of Psychiatry. NIHR Biomedical Research Centre for Mental Health at IoP & SLAM. Neuroimaging
Automated MR Image Analysis Pipelines Andy Simmons Centre for Neuroimaging Sciences, Kings College London Institute of Psychiatry. NIHR Biomedical Research Centre for Mental Health at IoP & SLAM. Neuroimaging
GLIRT: Groupwise and Longitudinal Image Registration Toolbox
 Software Release (1.0.1) Last updated: March. 30, 2011. GLIRT: Groupwise and Longitudinal Image Registration Toolbox Guorong Wu 1, Qian Wang 1,2, Hongjun Jia 1, and Dinggang Shen 1 1 Image Display, Enhancement,
Software Release (1.0.1) Last updated: March. 30, 2011. GLIRT: Groupwise and Longitudinal Image Registration Toolbox Guorong Wu 1, Qian Wang 1,2, Hongjun Jia 1, and Dinggang Shen 1 1 Image Display, Enhancement,
Detecting White Matter Lesions in Lupus
 Slicer3 Training Compendium Detecting White Matter Lesions in Lupus Version 2.1 6/25/2009 H. Jeremy Bockholt Mark Scully -1- Learning objective This tutorial demonstrates an automated, multi-level method
Slicer3 Training Compendium Detecting White Matter Lesions in Lupus Version 2.1 6/25/2009 H. Jeremy Bockholt Mark Scully -1- Learning objective This tutorial demonstrates an automated, multi-level method
Characterizing growth patterns in longitudinal MRI using image contrast
 Characterizing growth patterns in longitudinal MRI using image contrast Avantika Vardhan a,b, Marcel Prastawa a,c, Clement Vachet a, Joseph Piven d for IBIS *, Guido Gerig a,b,c a Scientific Computing
Characterizing growth patterns in longitudinal MRI using image contrast Avantika Vardhan a,b, Marcel Prastawa a,c, Clement Vachet a, Joseph Piven d for IBIS *, Guido Gerig a,b,c a Scientific Computing
Slicer3 Training Tutorial Manual segmentation of orbit structures from isotropic MRI data within 3D Slicer for windows
 Slicer3 Training Compendium Slicer3 Training Tutorial Manual segmentation of orbit structures from isotropic MRI data within 3D Slicer for windows Dr Raphael Olszewski Surgical Planning Laboratory Brigham
Slicer3 Training Compendium Slicer3 Training Tutorial Manual segmentation of orbit structures from isotropic MRI data within 3D Slicer for windows Dr Raphael Olszewski Surgical Planning Laboratory Brigham
MARS: Multiple Atlases Robust Segmentation
 Software Release (1.0.1) Last updated: April 30, 2014. MARS: Multiple Atlases Robust Segmentation Guorong Wu, Minjeong Kim, Gerard Sanroma, and Dinggang Shen {grwu, mjkim, gerard_sanroma, dgshen}@med.unc.edu
Software Release (1.0.1) Last updated: April 30, 2014. MARS: Multiple Atlases Robust Segmentation Guorong Wu, Minjeong Kim, Gerard Sanroma, and Dinggang Shen {grwu, mjkim, gerard_sanroma, dgshen}@med.unc.edu
Subvoxel Segmentation and Representation of Brain Cortex Using Fuzzy Clustering and Gradient Vector Diffusion
 Subvoxel Segmentation and Representation of Brain Cortex Using Fuzzy Clustering and Gradient Vector Diffusion Ming-Ching Chang Xiaodong Tao GE Global Research Center {changm, taox} @ research.ge.com SPIE
Subvoxel Segmentation and Representation of Brain Cortex Using Fuzzy Clustering and Gradient Vector Diffusion Ming-Ching Chang Xiaodong Tao GE Global Research Center {changm, taox} @ research.ge.com SPIE
NIH Public Access Author Manuscript Proc SPIE. Author manuscript; available in PMC 2013 December 31.
 NIH Public Access Author Manuscript Published in final edited form as: Proc SPIE. 2013 March 13; 8669:. doi:10.1117/12.2007093. UNC-Utah NA-MIC DTI framework: Atlas Based Fiber Tract Analysis with Application
NIH Public Access Author Manuscript Published in final edited form as: Proc SPIE. 2013 March 13; 8669:. doi:10.1117/12.2007093. UNC-Utah NA-MIC DTI framework: Atlas Based Fiber Tract Analysis with Application
Slicer3 Minute Tutorial
 Slicer3 Minute Tutorial Surgical Planning Laboratory Harvard Medical School Sonia Pujol, PhD Slicer3 Minute Tutorial This tutorial is a short introduction to the advanced 3D visualization capabilities
Slicer3 Minute Tutorial Surgical Planning Laboratory Harvard Medical School Sonia Pujol, PhD Slicer3 Minute Tutorial This tutorial is a short introduction to the advanced 3D visualization capabilities
Assessment of Reliability of Multi-site Neuroimaging Via Traveling Phantom Study
 Assessment of Reliability of Multi-site Neuroimaging Via Traveling Phantom Study Sylvain Gouttard 1, Martin Styner 2,3, Marcel Prastawa 1, Joseph Piven 3, and Guido Gerig 1 1 Scientific Computing and Imaging
Assessment of Reliability of Multi-site Neuroimaging Via Traveling Phantom Study Sylvain Gouttard 1, Martin Styner 2,3, Marcel Prastawa 1, Joseph Piven 3, and Guido Gerig 1 1 Scientific Computing and Imaging
MRI Segmentation MIDAS, 2007, 2010
 MRI Segmentation MIDAS, 2007, 2010 Lawrence O. Hall, Dmitry Goldgof, Yuhua Gu, Prodip Hore Dept. of Computer Science & Engineering University of South Florida CONTENTS: 1. Introduction... 1 2. Installing
MRI Segmentation MIDAS, 2007, 2010 Lawrence O. Hall, Dmitry Goldgof, Yuhua Gu, Prodip Hore Dept. of Computer Science & Engineering University of South Florida CONTENTS: 1. Introduction... 1 2. Installing
Slicer3 minute tutorial
 Slicer3 minute tutorial Sonia Pujol, Ph.D. Surgical Planning Laboratory Harvard Medical School -1- Slicer3 minute tutorial This tutorial is a short introduction to the advanced 3D visualization capabilities
Slicer3 minute tutorial Sonia Pujol, Ph.D. Surgical Planning Laboratory Harvard Medical School -1- Slicer3 minute tutorial This tutorial is a short introduction to the advanced 3D visualization capabilities
Slicer4Minute Tutorial. Sonia Pujol, Ph.D. Surgical Planning Laboratory Harvard Medical School
 Slicer4Minute Tutorial Sonia Pujol, Ph.D. Surgical Planning Laboratory Harvard Medical School 1 Slicer4 minute tutorial This tutorial is a 4-minute introduction to the 3D visualization capabilities of
Slicer4Minute Tutorial Sonia Pujol, Ph.D. Surgical Planning Laboratory Harvard Medical School 1 Slicer4 minute tutorial This tutorial is a 4-minute introduction to the 3D visualization capabilities of
Neuroimaging and mathematical modelling Lesson 2: Voxel Based Morphometry
 Neuroimaging and mathematical modelling Lesson 2: Voxel Based Morphometry Nivedita Agarwal, MD Nivedita.agarwal@apss.tn.it Nivedita.agarwal@unitn.it Volume and surface morphometry Brain volume White matter
Neuroimaging and mathematical modelling Lesson 2: Voxel Based Morphometry Nivedita Agarwal, MD Nivedita.agarwal@apss.tn.it Nivedita.agarwal@unitn.it Volume and surface morphometry Brain volume White matter
Assessment of Reliability of Multi-site Neuroimaging via Traveling Phantom Study
 Assessment of Reliability of Multi-site Neuroimaging via Traveling Phantom Study Sylvain Gouttard 1, Martin Styner 2,3, Marcel Prastawa 1, Joseph Piven 3, and Guido Gerig 1 1 Scientific Computing and Imaging
Assessment of Reliability of Multi-site Neuroimaging via Traveling Phantom Study Sylvain Gouttard 1, Martin Styner 2,3, Marcel Prastawa 1, Joseph Piven 3, and Guido Gerig 1 1 Scientific Computing and Imaging
MARS: Multiple Atlases Robust Segmentation
 Software Release (1.0.1) Last updated: April 30, 2014. MARS: Multiple Atlases Robust Segmentation Guorong Wu, Minjeong Kim, Gerard Sanroma, and Dinggang Shen {grwu, mjkim, gerard_sanroma, dgshen}@med.unc.edu
Software Release (1.0.1) Last updated: April 30, 2014. MARS: Multiple Atlases Robust Segmentation Guorong Wu, Minjeong Kim, Gerard Sanroma, and Dinggang Shen {grwu, mjkim, gerard_sanroma, dgshen}@med.unc.edu
Slicer3 Tutorial: Registration Library Case 14. Intra-subject Brain PET-MRI fusion
 NA-MIC Slicer3 Tutorial: Registration Library Case 14 Intra-subject Brain PET-MRI fusion Dominik Meier, Ron Kikinis March 2010 Overview 1. Introduction 2. Prerequisites 3. Modules Used takes how long to
NA-MIC Slicer3 Tutorial: Registration Library Case 14 Intra-subject Brain PET-MRI fusion Dominik Meier, Ron Kikinis March 2010 Overview 1. Introduction 2. Prerequisites 3. Modules Used takes how long to
EMSegment Tutorial. How to Define and Fine-Tune Automatic Brain Compartment Segmentation and the Detection of White Matter Hyperintensities
 EMSegment Tutorial How to Define and Fine-Tune Automatic Brain Compartment Segmentation and the Detection of White Matter Hyperintensities This documentation serves as a tutorial to learn to customize
EMSegment Tutorial How to Define and Fine-Tune Automatic Brain Compartment Segmentation and the Detection of White Matter Hyperintensities This documentation serves as a tutorial to learn to customize
Automatic Registration-Based Segmentation for Neonatal Brains Using ANTs and Atropos
 Automatic Registration-Based Segmentation for Neonatal Brains Using ANTs and Atropos Jue Wu and Brian Avants Penn Image Computing and Science Lab, University of Pennsylvania, Philadelphia, USA Abstract.
Automatic Registration-Based Segmentation for Neonatal Brains Using ANTs and Atropos Jue Wu and Brian Avants Penn Image Computing and Science Lab, University of Pennsylvania, Philadelphia, USA Abstract.
CIVET 2.0 (marching cubes), An Automated Pipeline For Structural Human MRI
 CIVET 2.0 (marching cubes), An Automated Pipeline For Structural Human MRI Lindsay B. Lewis, Ph.D. OMM March 17, 2014 SurfStat Outline Who is behind CIVET? What is CIVET? What s new in CIVET 2.0? Preparing
CIVET 2.0 (marching cubes), An Automated Pipeline For Structural Human MRI Lindsay B. Lewis, Ph.D. OMM March 17, 2014 SurfStat Outline Who is behind CIVET? What is CIVET? What s new in CIVET 2.0? Preparing
Image Registration Driven by Combined Probabilistic and Geometric Descriptors
 Image Registration Driven by Combined Probabilistic and Geometric Descriptors Linh Ha 1, Marcel Prastawa 1, Guido Gerig 1, John H. Gilmore 2, Cláudio T. Silva 1, Sarang Joshi 1 1 Scientific Computing and
Image Registration Driven by Combined Probabilistic and Geometric Descriptors Linh Ha 1, Marcel Prastawa 1, Guido Gerig 1, John H. Gilmore 2, Cláudio T. Silva 1, Sarang Joshi 1 1 Scientific Computing and
KWMeshVisu: A Mesh Visualization Tool for Shape Analysis
 KWMeshVisu: A Mesh Visualization Tool for Shape Analysis Release 1.00 Ipek Oguz 1, Guido Gerig 2, Sebastien Barre 3, and Martin Styner 4 July 13, 2006 1 University of North Carolina at Chapel Hill, ipek@cs.unc.edu
KWMeshVisu: A Mesh Visualization Tool for Shape Analysis Release 1.00 Ipek Oguz 1, Guido Gerig 2, Sebastien Barre 3, and Martin Styner 4 July 13, 2006 1 University of North Carolina at Chapel Hill, ipek@cs.unc.edu
ASAP_2.0 (Automatic Software for ASL Processing) USER S MANUAL
 ASAP_2.0 (Automatic Software for ASL Processing) USER S MANUAL ASAP was developed as part of the COST Action "Arterial Spin Labelling Initiative in Dementia (AID)" by: Department of Neuroimaging, Institute
ASAP_2.0 (Automatic Software for ASL Processing) USER S MANUAL ASAP was developed as part of the COST Action "Arterial Spin Labelling Initiative in Dementia (AID)" by: Department of Neuroimaging, Institute
ShapePopulationViewer
 ShapePopulationViewer User Tutorial V1.3.2 Alexis Girault, Francois Budin, Beatriz Paniagua, Martin Styner Neuro Image Research and Analysis Laboratories University of North Carolina at Chapel Hill 1 ShapePopulationViewer
ShapePopulationViewer User Tutorial V1.3.2 Alexis Girault, Francois Budin, Beatriz Paniagua, Martin Styner Neuro Image Research and Analysis Laboratories University of North Carolina at Chapel Hill 1 ShapePopulationViewer
UNC 4D Infant Cortical Surface Atlases, from Neonate to 6 Years of Age
 UNC 4D Infant Cortical Surface Atlases, from Neonate to 6 Years of Age Version 1.0 UNC 4D infant cortical surface atlases from neonate to 6 years of age contain 11 time points, including 1, 3, 6, 9, 12,
UNC 4D Infant Cortical Surface Atlases, from Neonate to 6 Years of Age Version 1.0 UNC 4D infant cortical surface atlases from neonate to 6 years of age contain 11 time points, including 1, 3, 6, 9, 12,
Software for ABSORB: An algorithm for effective groupwise registration
 Software Release (1.0.5) Last updated: Sep. 01, 2010. Software for ABSORB: An algorithm for effective groupwise registration Hongjun Jia 1, Guorong Wu 1, Qian Wang 1,2 and Dinggang Shen 1 1 Image Display,
Software Release (1.0.5) Last updated: Sep. 01, 2010. Software for ABSORB: An algorithm for effective groupwise registration Hongjun Jia 1, Guorong Wu 1, Qian Wang 1,2 and Dinggang Shen 1 1 Image Display,
Automatic Segmentation of Neonatal Brain MRI
 Automatic Segmentation of Neonatal Brain MRI 1 Marcel Prastawa, 2 John Gilmore, 3 Weili Lin, and 1,2 Guido Gerig Department of 1 Computer Science, 2 Psychiatry, and 3 Radiology University of North Carolina,
Automatic Segmentation of Neonatal Brain MRI 1 Marcel Prastawa, 2 John Gilmore, 3 Weili Lin, and 1,2 Guido Gerig Department of 1 Computer Science, 2 Psychiatry, and 3 Radiology University of North Carolina,
Data Loading & 3D Visualization
 Neuroimage Analysis Center Data Loading & 3D Visualization Sonia Pujol, Ph.D. Surgical Planning Laboratory Harvard Medical School Leonardo da Vinci (1452-1519), Virgin and Child Alte Pinakothek, München
Neuroimage Analysis Center Data Loading & 3D Visualization Sonia Pujol, Ph.D. Surgical Planning Laboratory Harvard Medical School Leonardo da Vinci (1452-1519), Virgin and Child Alte Pinakothek, München
Corpus Callosum Subdivision based on a Probabilistic Model of Inter-Hemispheric Connectivity
 Corpus Callosum Subdivision based on a Probabilistic Model of Inter-Hemispheric Connectivity Martin A. Styner 1,2, Ipek Oguz 1, Rachel Gimpel Smith 2, Carissa Cascio 2, and Matthieu Jomier 1 1 Dept. of
Corpus Callosum Subdivision based on a Probabilistic Model of Inter-Hemispheric Connectivity Martin A. Styner 1,2, Ipek Oguz 1, Rachel Gimpel Smith 2, Carissa Cascio 2, and Matthieu Jomier 1 1 Dept. of
GLM for fmri data analysis Lab Exercise 1
 GLM for fmri data analysis Lab Exercise 1 March 15, 2013 Medical Image Processing Lab Medical Image Processing Lab GLM for fmri data analysis Outline 1 Getting Started 2 AUDIO 1 st level Preprocessing
GLM for fmri data analysis Lab Exercise 1 March 15, 2013 Medical Image Processing Lab Medical Image Processing Lab GLM for fmri data analysis Outline 1 Getting Started 2 AUDIO 1 st level Preprocessing
Lilla Zöllei A.A. Martinos Center, MGH; Boston, MA
 Lilla Zöllei lzollei@nmr.mgh.harvard.edu A.A. Martinos Center, MGH; Boston, MA Bruce Fischl Gheorghe Postelnicu Jean Augustinack Anastasia Yendiki Allison Stevens Kristen Huber Sita Kakonoori + the FreeSurfer
Lilla Zöllei lzollei@nmr.mgh.harvard.edu A.A. Martinos Center, MGH; Boston, MA Bruce Fischl Gheorghe Postelnicu Jean Augustinack Anastasia Yendiki Allison Stevens Kristen Huber Sita Kakonoori + the FreeSurfer
Structural Segmentation
 Structural Segmentation FAST tissue-type segmentation FIRST sub-cortical structure segmentation FSL-VBM voxelwise grey-matter density analysis SIENA atrophy analysis FAST FMRIB s Automated Segmentation
Structural Segmentation FAST tissue-type segmentation FIRST sub-cortical structure segmentation FSL-VBM voxelwise grey-matter density analysis SIENA atrophy analysis FAST FMRIB s Automated Segmentation
A midas plugin to enable construction of reproducible web-based image processing pipelines
 NEUROINFORMATICS TECHNOLOGY REPORT ARTICLE published: 30 December 2013 doi: 10.3389/fninf.2013.00046 A midas plugin to enable construction of reproducible web-based image processing pipelines Michael Grauer
NEUROINFORMATICS TECHNOLOGY REPORT ARTICLE published: 30 December 2013 doi: 10.3389/fninf.2013.00046 A midas plugin to enable construction of reproducible web-based image processing pipelines Michael Grauer
NIH Public Access Author Manuscript Proc IEEE Int Symp Biomed Imaging. Author manuscript; available in PMC 2014 January 15.
 NIH Public Access Author Manuscript Published in final edited form as: Proc IEEE Int Symp Biomed Imaging. 2013 December 31; 2013: 1396 1399. doi:10.1109/isbi. 2013.6556794. MODELING LONGITUDINAL MRI CHANGES
NIH Public Access Author Manuscript Published in final edited form as: Proc IEEE Int Symp Biomed Imaging. 2013 December 31; 2013: 1396 1399. doi:10.1109/isbi. 2013.6556794. MODELING LONGITUDINAL MRI CHANGES
The organization of the human cerebral cortex estimated by intrinsic functional connectivity
 1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
Structural Segmentation
 Structural Segmentation FAST tissue-type segmentation FIRST sub-cortical structure segmentation FSL-VBM voxelwise grey-matter density analysis SIENA atrophy analysis FAST FMRIB s Automated Segmentation
Structural Segmentation FAST tissue-type segmentation FIRST sub-cortical structure segmentation FSL-VBM voxelwise grey-matter density analysis SIENA atrophy analysis FAST FMRIB s Automated Segmentation
How to create a head model
 How to create a head model This document describes the command line tools: mri2mesh: Central tool to reconstruct a head model from T1w and T2w data dwi2cond: Reconstruct conductivity tensors for brain
How to create a head model This document describes the command line tools: mri2mesh: Central tool to reconstruct a head model from T1w and T2w data dwi2cond: Reconstruct conductivity tensors for brain
Automatic Generation of Training Data for Brain Tissue Classification from MRI
 Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. COCOSCO, Alex P. ZIJDENBOS, and Alan C. EVANS http://www.bic.mni.mcgill.ca/users/crisco/ McConnell Brain Imaging
Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. COCOSCO, Alex P. ZIJDENBOS, and Alan C. EVANS http://www.bic.mni.mcgill.ca/users/crisco/ McConnell Brain Imaging
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit
 Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
Atlas Registration & Label Merging
 NA-MIC Slicer3 Tutorial Atlas Registration & Label Merging Dominik Meier, Ron Kikinis February 2010 Overview 1. Introduction 2. Prerequisites 3. Modules Used takes how long to do? 4. Loading Example Dataset
NA-MIC Slicer3 Tutorial Atlas Registration & Label Merging Dominik Meier, Ron Kikinis February 2010 Overview 1. Introduction 2. Prerequisites 3. Modules Used takes how long to do? 4. Loading Example Dataset
Slicer3 Tutorial: Registration Library Case 08
 NA-MIC Slicer3 Tutorial: Registration Library Case 08 Serial PET-CT Dominik Meier, Ron Kikinis March 2010 Introduction / Scenario We have two sets of PET-CT scans, a baseline and follow-up scan. baseline
NA-MIC Slicer3 Tutorial: Registration Library Case 08 Serial PET-CT Dominik Meier, Ron Kikinis March 2010 Introduction / Scenario We have two sets of PET-CT scans, a baseline and follow-up scan. baseline
Joint longitudinal modeling of brain appearance in multimodal MRI for the characterization of early brain developmental processes
 Joint longitudinal modeling of brain appearance in multimodal MRI for the characterization of early brain developmental processes Avantika Vardhan 1, Marcel Prastawa, Neda Sadeghi 1, Clement Vachet 1,
Joint longitudinal modeling of brain appearance in multimodal MRI for the characterization of early brain developmental processes Avantika Vardhan 1, Marcel Prastawa, Neda Sadeghi 1, Clement Vachet 1,
Low-Rank to the Rescue Atlas-based Analyses in the Presence of Pathologies
 Low-Rank to the Rescue Atlas-based Analyses in the Presence of Pathologies Xiaoxiao Liu 1, Marc Niethammer 2, Roland Kwitt 3, Matthew McCormick 1, and Stephen Aylward 1 1 Kitware Inc., USA 2 University
Low-Rank to the Rescue Atlas-based Analyses in the Presence of Pathologies Xiaoxiao Liu 1, Marc Niethammer 2, Roland Kwitt 3, Matthew McCormick 1, and Stephen Aylward 1 1 Kitware Inc., USA 2 University
Berger Jean-Baptiste
 Berger Jean-Baptiste jean-baptiste.berger@cpe.fr 1 Index Introduction - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - 3 Step1: Select your Data - - - - - - - - - -
Berger Jean-Baptiste jean-baptiste.berger@cpe.fr 1 Index Introduction - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - - 3 Step1: Select your Data - - - - - - - - - -
Preprocessing II: Between Subjects John Ashburner
 Preprocessing II: Between Subjects John Ashburner Pre-processing Overview Statistics or whatever fmri time-series Anatomical MRI Template Smoothed Estimate Spatial Norm Motion Correct Smooth Coregister
Preprocessing II: Between Subjects John Ashburner Pre-processing Overview Statistics or whatever fmri time-series Anatomical MRI Template Smoothed Estimate Spatial Norm Motion Correct Smooth Coregister
Programming in Slicer4
 Programming in Slicer4 Sonia Pujol, Ph.D. Surgical Planning Laboratory, Harvard Medical School Paul Cézanne, Moulin sur la Couleuvre à Pontoise, 1881, Staatliche Museen zu Berlin, Na:onalgalerie Steve
Programming in Slicer4 Sonia Pujol, Ph.D. Surgical Planning Laboratory, Harvard Medical School Paul Cézanne, Moulin sur la Couleuvre à Pontoise, 1881, Staatliche Museen zu Berlin, Na:onalgalerie Steve
Programming in Slicer4
 Programming in Slicer4 Sonia Pujol, Ph.D. Surgical Planning Laboratory, Harvard Medical School Paul Cézanne, Moulin sur la Couleuvre à Pontoise, 1881, Staatliche Museen zu Berlin, Na:onalgalerie Steve
Programming in Slicer4 Sonia Pujol, Ph.D. Surgical Planning Laboratory, Harvard Medical School Paul Cézanne, Moulin sur la Couleuvre à Pontoise, 1881, Staatliche Museen zu Berlin, Na:onalgalerie Steve
Multiple Sclerosis Brain MRI Segmentation Workflow deployment on the EGEE grid
 Multiple Sclerosis Brain MRI Segmentation Workflow deployment on the EGEE grid Erik Pernod 1, Jean-Christophe Souplet 1, Javier Rojas Balderrama 2, Diane Lingrand 2, Xavier Pennec 1 Speaker: Grégoire Malandain
Multiple Sclerosis Brain MRI Segmentation Workflow deployment on the EGEE grid Erik Pernod 1, Jean-Christophe Souplet 1, Javier Rojas Balderrama 2, Diane Lingrand 2, Xavier Pennec 1 Speaker: Grégoire Malandain
Comparison Study of Clinical 3D MRI Brain Segmentation Evaluation
 Comparison Study of Clinical 3D MRI Brain Segmentation Evaluation Ting Song 1, Elsa D. Angelini 2, Brett D. Mensh 3, Andrew Laine 1 1 Heffner Biomedical Imaging Laboratory Department of Biomedical Engineering,
Comparison Study of Clinical 3D MRI Brain Segmentation Evaluation Ting Song 1, Elsa D. Angelini 2, Brett D. Mensh 3, Andrew Laine 1 1 Heffner Biomedical Imaging Laboratory Department of Biomedical Engineering,
BrainSuite Lab Exercises. presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014
 BrainSuite Lab Exercises presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014 1. Opening and Displaying an MRI Start BrainSuite Drag and drop the T1 image from the native space
BrainSuite Lab Exercises presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014 1. Opening and Displaying an MRI Start BrainSuite Drag and drop the T1 image from the native space
ZipStudy. Utility for Packaging Study Files. A. Maudsley, 6/2015
 ZipStudy Utility for Packaging Study Files A. Maudsley, 6/2015 CONTENTS: 1. Introduction... 1 2. Starting the Program... 1 3. Copying and Compressing Subject Data... 2 4. Restoring MIDAS Data from a Zipped
ZipStudy Utility for Packaging Study Files A. Maudsley, 6/2015 CONTENTS: 1. Introduction... 1 2. Starting the Program... 1 3. Copying and Compressing Subject Data... 2 4. Restoring MIDAS Data from a Zipped
Processing math: 100% Intensity Normalization
 Intensity Normalization Overall Pipeline 2/21 Intensity normalization Conventional MRI intensites (T1-w, T2-w, PD, FLAIR) are acquired in arbitrary units Images are not comparable across scanners, subjects,
Intensity Normalization Overall Pipeline 2/21 Intensity normalization Conventional MRI intensites (T1-w, T2-w, PD, FLAIR) are acquired in arbitrary units Images are not comparable across scanners, subjects,
NA-MIC National Alliance for Medical Image Computing fmri Data Analysis
 NA-MIC fmri Data Analysis Sonia Pujol, Ph.D. Wendy Plesniak, Ph.D. Randy Gollub, M.D., Ph.D. Acknowledgments NIH U54EB005149 Neuroimage Analysis Center NIH P41RR013218 FIRST Biomedical Informatics Research
NA-MIC fmri Data Analysis Sonia Pujol, Ph.D. Wendy Plesniak, Ph.D. Randy Gollub, M.D., Ph.D. Acknowledgments NIH U54EB005149 Neuroimage Analysis Center NIH P41RR013218 FIRST Biomedical Informatics Research
PANDA Manual Zaixu Cui & Suyu Zhong & Gaolang Gong
 PANDA Manual Zaixu Cui & Suyu Zhong & Gaolang Gong National Key Laboratory of Cognitive Neuroscience and Learning Beijing Normal University, China Contents Overview Setup Files/Directories selection Preparing
PANDA Manual Zaixu Cui & Suyu Zhong & Gaolang Gong National Key Laboratory of Cognitive Neuroscience and Learning Beijing Normal University, China Contents Overview Setup Files/Directories selection Preparing
Automatic MS Lesion Segmentation by Outlier Detection and Information Theoretic Region Partitioning Release 0.00
 Automatic MS Lesion Segmentation by Outlier Detection and Information Theoretic Region Partitioning Release 0.00 Marcel Prastawa 1 and Guido Gerig 1 Abstract July 17, 2008 1 Scientific Computing and Imaging
Automatic MS Lesion Segmentation by Outlier Detection and Information Theoretic Region Partitioning Release 0.00 Marcel Prastawa 1 and Guido Gerig 1 Abstract July 17, 2008 1 Scientific Computing and Imaging
The Insight Toolkit. Image Registration Algorithms & Frameworks
 The Insight Toolkit Image Registration Algorithms & Frameworks Registration in ITK Image Registration Framework Multi Resolution Registration Framework Components PDE Based Registration FEM Based Registration
The Insight Toolkit Image Registration Algorithms & Frameworks Registration in ITK Image Registration Framework Multi Resolution Registration Framework Components PDE Based Registration FEM Based Registration
Where are we now? Structural MRI processing and analysis
 Where are we now? Structural MRI processing and analysis Pierre-Louis Bazin bazin@cbs.mpg.de Leipzig, Germany Structural MRI processing: why bother? Just use the standards? SPM FreeSurfer FSL However:
Where are we now? Structural MRI processing and analysis Pierre-Louis Bazin bazin@cbs.mpg.de Leipzig, Germany Structural MRI processing: why bother? Just use the standards? SPM FreeSurfer FSL However:
Input tab. Insert the path to the DTI Image that you want to tract.
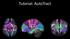 Tutorial: AutoTract Input tab Insert the path to the DTI Image that you want to tract. Input tab (optional)if you already have a WM/CSF mask you want to use to process the tracts, insert the path here.
Tutorial: AutoTract Input tab Insert the path to the DTI Image that you want to tract. Input tab (optional)if you already have a WM/CSF mask you want to use to process the tracts, insert the path here.
PRANA. Project Review and Analysis. March (Pre-release and still under development!)
 PRANA Project Review and Analysis March 2010- (Pre-release and still under development!) A.A. Maudsley Contents: PRANA... 1 1. Introduction... 2 1.1. Examples... 2 2. Subject/Study Selection and Filter...
PRANA Project Review and Analysis March 2010- (Pre-release and still under development!) A.A. Maudsley Contents: PRANA... 1 1. Introduction... 2 1.1. Examples... 2 2. Subject/Study Selection and Filter...
An Introduction To Automatic Tissue Classification Of Brain MRI. Colm Elliott Mar 2014
 An Introduction To Automatic Tissue Classification Of Brain MRI Colm Elliott Mar 2014 Tissue Classification Tissue classification is part of many processing pipelines. We often want to classify each voxel
An Introduction To Automatic Tissue Classification Of Brain MRI Colm Elliott Mar 2014 Tissue Classification Tissue classification is part of many processing pipelines. We often want to classify each voxel
Combined SPHARM-PDM and entropy-based particle systems shape analysis framework
 Combined SPHARM-PDM and entropy-based particle systems shape analysis framework Beatriz Paniagua a, Lucile Bompard a, Josh Cates b, Ross Whitaker b, Manasi Datar b, Clement Vachet a, Martin Styner a a
Combined SPHARM-PDM and entropy-based particle systems shape analysis framework Beatriz Paniagua a, Lucile Bompard a, Josh Cates b, Ross Whitaker b, Manasi Datar b, Clement Vachet a, Martin Styner a a
White Matter Lesion Segmentation (WMLS) Manual
 White Matter Lesion Segmentation (WMLS) Manual 1. Introduction White matter lesions (WMLs) are brain abnormalities that appear in different brain diseases, such as multiple sclerosis (MS), head injury,
White Matter Lesion Segmentation (WMLS) Manual 1. Introduction White matter lesions (WMLs) are brain abnormalities that appear in different brain diseases, such as multiple sclerosis (MS), head injury,
3D Slicer. NA-MIC National Alliance for Medical Image Computing 4 February 2011
 NA-MIC http://na-mic.org 3D Slicer 4 February 2011 Andrey Fedorov, PhD Steve Pieper, PhD Ron Kikinis, MD Surgical Planning Lab Brigham and Women's Hospital Acknowledgments Picture courtesy Kapur, Jakab,
NA-MIC http://na-mic.org 3D Slicer 4 February 2011 Andrey Fedorov, PhD Steve Pieper, PhD Ron Kikinis, MD Surgical Planning Lab Brigham and Women's Hospital Acknowledgments Picture courtesy Kapur, Jakab,
An ITK Filter for Bayesian Segmentation: itkbayesianclassifierimagefilter
 An ITK Filter for Bayesian Segmentation: itkbayesianclassifierimagefilter John Melonakos 1, Karthik Krishnan 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332, USA {jmelonak,
An ITK Filter for Bayesian Segmentation: itkbayesianclassifierimagefilter John Melonakos 1, Karthik Krishnan 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332, USA {jmelonak,
Automatic Vascular Tree Formation Using the Mahalanobis Distance
 Automatic Vascular Tree Formation Using the Mahalanobis Distance Julien Jomier, Vincent LeDigarcher, and Stephen R. Aylward Computer-Aided Diagnosis and Display Lab, Department of Radiology The University
Automatic Vascular Tree Formation Using the Mahalanobis Distance Julien Jomier, Vincent LeDigarcher, and Stephen R. Aylward Computer-Aided Diagnosis and Display Lab, Department of Radiology The University
NIH Public Access Author Manuscript Proc SPIE. Author manuscript; available in PMC 2012 October 10.
 NIH Public Access Author Manuscript Published in final edited form as: Proc SPIE. 2011 ; 7962: 79620S. doi:10.1117/12.878291. Efficient, Graph-based White Matter Connectivity from Orientation Distribution
NIH Public Access Author Manuscript Published in final edited form as: Proc SPIE. 2011 ; 7962: 79620S. doi:10.1117/12.878291. Efficient, Graph-based White Matter Connectivity from Orientation Distribution
Brain Portion Peeling from T2 Axial MRI Head Scans using Clustering and Morphological Operation
 159 Brain Portion Peeling from T2 Axial MRI Head Scans using Clustering and Morphological Operation K. Somasundaram Image Processing Lab Dept. of Computer Science and Applications Gandhigram Rural Institute
159 Brain Portion Peeling from T2 Axial MRI Head Scans using Clustering and Morphological Operation K. Somasundaram Image Processing Lab Dept. of Computer Science and Applications Gandhigram Rural Institute
Scientific Computing and Imaging Institute, University of Utah, Salt Lake City
 UNC-Utah NA-MIC DTI framework: Atlas Based Fiber Tract Analysis with Application to a Study of Nicotine Smoking Addiction Audrey R Verde1, Jean-Baptiste Berger1, Aditya Gupta1,7, Mahshid Farzinfar1, Adrien
UNC-Utah NA-MIC DTI framework: Atlas Based Fiber Tract Analysis with Application to a Study of Nicotine Smoking Addiction Audrey R Verde1, Jean-Baptiste Berger1, Aditya Gupta1,7, Mahshid Farzinfar1, Adrien
Joint Segmentation and Registration for Infant Brain Images
 Joint Segmentation and Registration for Infant Brain Images Guorong Wu 1, Li Wang 1, John Gilmore 2, Weili Lin 1, and Dinggang Shen 1(&) 1 Department of Radiology and BRIC, University of North Carolina,
Joint Segmentation and Registration for Infant Brain Images Guorong Wu 1, Li Wang 1, John Gilmore 2, Weili Lin 1, and Dinggang Shen 1(&) 1 Department of Radiology and BRIC, University of North Carolina,
Deformetrica: a software for statistical analysis of anatomical shapes
 Deformetrica: a software for statistical analysis of anatomical shapes Alexandre Routier, Marcel Prastawa, Benjamin Charlier, Cédric Doucet, Joan Alexis Glaunès, Stanley Durrleman To cite this version:
Deformetrica: a software for statistical analysis of anatomical shapes Alexandre Routier, Marcel Prastawa, Benjamin Charlier, Cédric Doucet, Joan Alexis Glaunès, Stanley Durrleman To cite this version:
NIH Public Access Author Manuscript Proc SPIE. Author manuscript; available in PMC 2014 July 31.
 NIH Public Access Author Manuscript Published in final edited form as: Proc SPIE. 2013 March 13; 8669:. doi:10.1117/12.2006651. Consistent 4D Brain Extraction of Serial Brain MR Images Yaping Wang a,b,
NIH Public Access Author Manuscript Published in final edited form as: Proc SPIE. 2013 March 13; 8669:. doi:10.1117/12.2006651. Consistent 4D Brain Extraction of Serial Brain MR Images Yaping Wang a,b,
An integrated software solution for improving neuroimaging data archival, management, and processing - The experience from the Queen Square MS Centre
 An integrated software solution for improving neuroimaging data archival, management, and processing - The experience from the Queen Square MS Centre Dr Baris Kanber, Translational Imaging Group, Centre
An integrated software solution for improving neuroimaging data archival, management, and processing - The experience from the Queen Square MS Centre Dr Baris Kanber, Translational Imaging Group, Centre
Measuring longitudinal brain changes in humans and small animal models. Christos Davatzikos
 Measuring longitudinal brain changes in humans and small animal models Christos Davatzikos Section of Biomedical Image Analysis University of Pennsylvania (Radiology) http://www.rad.upenn.edu/sbia Computational
Measuring longitudinal brain changes in humans and small animal models Christos Davatzikos Section of Biomedical Image Analysis University of Pennsylvania (Radiology) http://www.rad.upenn.edu/sbia Computational
Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge
 Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge Christian Wasserthal 1, Karin Engel 1, Karsten Rink 1 und André Brechmann
Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge Christian Wasserthal 1, Karin Engel 1, Karsten Rink 1 und André Brechmann
MIDAS Processing of Segmentation Data for Brain Lesions
 MIDAS Processing of Segmentation Data for Brain Lesions A. A. Maudsley 4/6/2012 For studies that contain lesions, the MIDAS processing has two steps that benefit from a modified processing pipeline to
MIDAS Processing of Segmentation Data for Brain Lesions A. A. Maudsley 4/6/2012 For studies that contain lesions, the MIDAS processing has two steps that benefit from a modified processing pipeline to
Methods for data preprocessing
 Methods for data preprocessing John Ashburner Wellcome Trust Centre for Neuroimaging, 12 Queen Square, London, UK. Overview Voxel-Based Morphometry Morphometry in general Volumetrics VBM preprocessing
Methods for data preprocessing John Ashburner Wellcome Trust Centre for Neuroimaging, 12 Queen Square, London, UK. Overview Voxel-Based Morphometry Morphometry in general Volumetrics VBM preprocessing
Objectives Learn how GMS uses rasters to support all kinds of digital elevation models and how rasters can be used for interpolation in GMS.
 v. 9.1 GMS 9.1 Tutorial Using rasters for interpolation and visualization in GMS Objectives Learn how GMS uses rasters to support all kinds of digital elevation models and how rasters can be used for interpolation
v. 9.1 GMS 9.1 Tutorial Using rasters for interpolation and visualization in GMS Objectives Learn how GMS uses rasters to support all kinds of digital elevation models and how rasters can be used for interpolation
NIH Public Access Author Manuscript Proc IEEE Comput Soc Conf Comput Vis Pattern Recognit. Author manuscript; available in PMC 2010 September 20.
 NIH Public Access Author Manuscript Proc IEEE Comput Soc Conf Comput Vis Pattern Recognit. Author manuscript; available in PMC 2010 September 20. Published in final edited form as: Proc IEEE Comput Soc
NIH Public Access Author Manuscript Proc IEEE Comput Soc Conf Comput Vis Pattern Recognit. Author manuscript; available in PMC 2010 September 20. Published in final edited form as: Proc IEEE Comput Soc
SPHARM-PDM. User Tutorial. Jonathan Perdomo, Beatriz Paniagua, Martin Styner July 2015
 SPHARM-PDM User Tutorial Jonathan Perdomo, Beatriz Paniagua, Martin Styner July 2015 3D Slicer Installation Go to www.slicer.org to download Slicer for your respective operating system. SPHARM-PDM Installation
SPHARM-PDM User Tutorial Jonathan Perdomo, Beatriz Paniagua, Martin Styner July 2015 3D Slicer Installation Go to www.slicer.org to download Slicer for your respective operating system. SPHARM-PDM Installation
Neuroimage Processing
 Neuroimage Processing Instructor: Moo K. Chung mkchung@wisc.edu Lecture 2-3. General Linear Models (GLM) Voxel-based Morphometry (VBM) September 11, 2009 What is GLM The general linear model (GLM) is a
Neuroimage Processing Instructor: Moo K. Chung mkchung@wisc.edu Lecture 2-3. General Linear Models (GLM) Voxel-based Morphometry (VBM) September 11, 2009 What is GLM The general linear model (GLM) is a
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit
 Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
LST: A lesion segmentation tool for SPM
 LST: A lesion segmentation tool for SPM Manual/Documentation for version 2.0.15 June 2017 Paul Schmidt Lucie Wink Contents 1 Getting started 3 1.1 License................................. 3 1.2 Installation...............................
LST: A lesion segmentation tool for SPM Manual/Documentation for version 2.0.15 June 2017 Paul Schmidt Lucie Wink Contents 1 Getting started 3 1.1 License................................. 3 1.2 Installation...............................
Free-Form B-spline Deformation Model for Groupwise Registration
 Free-Form B-spline Deformation Model for Groupwise Registration The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Balci
Free-Form B-spline Deformation Model for Groupwise Registration The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Balci
BDP: BrainSuite Diffusion Pipeline. Chitresh Bhushan
 BDP: BrainSuite Diffusion Pipeline Chitresh Bhushan Why diffusion MRI? T 2 weighted MPRAGE FA map Fiber track Quantify microstructural tissue characteristics Structural connectivity Connectome Clinical
BDP: BrainSuite Diffusion Pipeline Chitresh Bhushan Why diffusion MRI? T 2 weighted MPRAGE FA map Fiber track Quantify microstructural tissue characteristics Structural connectivity Connectome Clinical
Cortical Enhanced Tissue Segmentation of Neonatal Brain MR Images Acquired by a Dedicated Phased Array Coil
 Cortical Enhanced Tissue Segmentation of Neonatal Brain MR Images Acquired by a Dedicated Phased Array Coil Feng Shi 1, Pew-Thian Yap 1, Yong Fan 1, Jie-Zhi Cheng 1, Lawrence L. Wald 2,3, Guido Gerig 4,
Cortical Enhanced Tissue Segmentation of Neonatal Brain MR Images Acquired by a Dedicated Phased Array Coil Feng Shi 1, Pew-Thian Yap 1, Yong Fan 1, Jie-Zhi Cheng 1, Lawrence L. Wald 2,3, Guido Gerig 4,
Using rasters for interpolation and visualization in GMS
 v. 10.3 GMS 10.3 Tutorial Using rasters for interpolation and visualization in GMS Objectives This tutorial teaches how GMS uses rasters to support all kinds of digital elevation models and how rasters
v. 10.3 GMS 10.3 Tutorial Using rasters for interpolation and visualization in GMS Objectives This tutorial teaches how GMS uses rasters to support all kinds of digital elevation models and how rasters
3D Slicer Overview. Andras Lasso, PhD PerkLab, Queen s University
 3D Slicer Overview Andras Lasso, PhD PerkLab, Queen s University Right tool for the job Technological prototype Research tool Clinical tool Can it be done? Jalopnik.com Innovative, not robust, usually
3D Slicer Overview Andras Lasso, PhD PerkLab, Queen s University Right tool for the job Technological prototype Research tool Clinical tool Can it be done? Jalopnik.com Innovative, not robust, usually
Structural MRI analysis
 Structural MRI analysis volumetry and voxel-based morphometry cortical thickness measurements structural covariance network mapping Boris Bernhardt, PhD Department of Social Neuroscience, MPI-CBS bernhardt@cbs.mpg.de
Structural MRI analysis volumetry and voxel-based morphometry cortical thickness measurements structural covariance network mapping Boris Bernhardt, PhD Department of Social Neuroscience, MPI-CBS bernhardt@cbs.mpg.de
Fmri Spatial Processing
 Educational Course: Fmri Spatial Processing Ray Razlighi Jun. 8, 2014 Spatial Processing Spatial Re-alignment Geometric distortion correction Spatial Normalization Smoothing Why, When, How, Which Why is
Educational Course: Fmri Spatial Processing Ray Razlighi Jun. 8, 2014 Spatial Processing Spatial Re-alignment Geometric distortion correction Spatial Normalization Smoothing Why, When, How, Which Why is
