MR Diffusion-Based Inference of a Fiber Bundle Model from a Population of Subjects
|
|
|
- Basil Bailey
- 5 years ago
- Views:
Transcription
1 MR Diffusion-Based Inference of a Fiber Bundle Model from a Population of Subjects V. El Kouby 1,3, Y. Cointepas 1,3, C. Poupon 1,3, D. Rivière 1,3, N. Golestani 1,2, J.-B. Poline 4, D. Le Bihan 1,3, and J.-F. Mangin 1,3 1 Service Hospitalier Frédéric Joliot, CEA, Orsay, France elkouby@shfj.cea.fr, 2 INSERM U562 Cognitive Neuro Imaging, Orsay, France 3 Institut Fédératif de Recherche 49 (Imagerie Neurofonctionnelle), Paris Abstract. This paper proposes a method to infer a high level model of the white matter organization from a population of subjects using MR diffusion imaging. This method takes as input for each subject a set of trajectories stemming from any tracking algorithm. Then the inference results from two nested clustering stages. The first clustering converts each individual set of trajectories into a set of bundles supposed to represent large white matter pathways. The second clustering matches these bundles across subjects in order to provide a list of candidates for the bundle model. The method is applied on a population of eleven subjects and leads to the inference of 17 such candidates. 1 Introduction In the recent past, the development of diffusion tensor imaging (DTI) has led to a new field of opportunities for the study of brain white matter [11]. A possible approach for such studies lies in the usual spatial normalization framework. For this purpose, the development of methods for warping tensor fields using full tensor information is a promising undertaking, because it may improve the normalization computed from standard anatomical images [10, 20]. Spatial normalization can be used for instance for voxel-based statistical analysis. Combining normalization with fiber tracking techniques can also lead to statistical parametric maps reflecting variability of bundle location and thickness. Other morphometric approaches can be developped from dedicated parameterizations of the bundles [8, 6]. One of the difficulties for bundle morphometry stems from the poor anatomical knowledge about white matter organization of the human brain. Most of the anatomical techniques used to trace tracts can not be used in the human brain. Therefore, the current knowledge built from Klinger method or observation of Wallerian degeneration after lesion is relatively sparse [7, 13]. There is a need for computational methods aiming at infering a model of the main bundles making up the human brain white matter from MR diffusion imaging. This paper explores this direction of research. A lot of methods have been proposed to compute fiber related pathways from diffusion-weighted MR images. The simplest approaches follow trajectories of maximum diffusion [14, 5, 1]. More sophisticated methods try to overcome partial volume problems induced by fiber crossing using either high angular resolution acquisitions [18, 15], regularization [19, 16], front propagation [12, 9] or Monte Carlo sampling [3, 2]. Most of these techniques convert the raw MR diffusion data into a huge set of trajectories
2 supposed to correspond to putative fascicles. These fascicles can be used to create connectivity matrices related either to functional or anatomical regions of interest (ROIs) [5]. These fascicles can also be organized into larger bundles using clustering algorithms [4], which is addressed in this paper. This last approach is interesting because the potential influence of erroneous definitions of the ROIs is discarded. The proposed method aims at the inference of a model of the bundles that can be detected in most of the subjects. Some of these bundles should be related to the current anatomical knowledge, but the ultimate goal is to enrich the current understanding of white matter organization. The method is made up of two nested clustering algorithms. The first clustering is performed subject by subject in order to reduce the complexity of the full set of fascicles to a small set of large bundles (cf. Fig. 1). The second clustering is performed across subjects in order to match similar bundles that may correspond to a generic anatomical pathway (cf. Fig. 3 and 4). This second stage relies on a simple affine spatial normalization computed from the diffusion-free T2-weighted image included in the dataset. The work described in this paper is exploratory. Therefore, we do not address the current weaknesses of the tracking algorithms that may provide spurious fascicles. Moreover, we use a very simple clustering algorithm in order to get a first insight into the problem before designing more sophisticated dedicated methods. 2 Method 2.1 Clustering the fascicles The first clustering is performed subject by subject according to the following scheme: Global fascicle tracking. The computation of the initial set of fascicles is performed with brainvisa ( an open software implementing Mori s algorithm [14]. Simply speaking, the fascicles are trajectories of highest diffusion reconstructed step by step from the diffusion tensor field. A mask of the brain is computed from the T2-weighted image and one trajectory is obtained for each voxel, leading to a set of about fascicles. In this paper, we have chosen to focus on long bundles which led us to discard the fascicles whose length is less that 5cm. The final set includes about fascicles (see Fig.1.A). ROI map. The clustering is not performed directly on the fascicles but on a set of Regions of Interest (ROIs) defined from Talairach proportional system. The Talairach grid is split into cubic ROIs that are transformed to the subject referential using affine spatial normalization of the T2-weighted image performed via SPM2 ( ion.ucl.ac.uk/spm) (see Fig.1.B). For the study described in this paper, the spatial resolution of the grid of ROIs is 5mm. After masking this grid with a mask of the subject white matter (see Fig.1.D), the final set is made up of about 5000 ROIs. Connectivity matrix. An anatomical connectivity matrix A is computed for the set of ROIs mentioned above. For each pair of ROIs (i, j), A ij is the number of fascicles crossing i and j (see Fig.1.C). In order to focus the clustering on the strongest connectivity information, this matrix is binarized using a threshold on the number of fascicles. For the experiment described further, this threshold has been set to the mean of non null coefficients plus one standard deviation, namely about 40 fascicles depending on the subject.
3 A B C D E Fig. 1. Clustering the fascicles of one subject: A: The set of long fascicles to be clustered into bundles. B: The same set of fascicles embedded into the Talairach grid used to define cubic ROIs of 5mm resolution. C: The connectivity matrix of the ROIs with 2 levels of zoom (about 5000 ROIs). Each point in the matrix is the number of fascicles linking the two related ROIs. D: Result of the clustering for the ROIs. Each color denotes a cluster. Each ROI shape is a cube in Talairach space masked by the subject white matter. E: Result of the clustering at the level of the fascicles. The garbage clusters have been filtered out. Clustering. In the matrix A, each row stands for the connectivity of one ROI with the other ROIs. The ROIs belonging to the same bundle should have similar connectivity patterns. Therefore, we perform a clustering among the ROI set using a distance between ROI corresponding to the Euclidean distance betwwen the rows of A. The clustering is performed using the standard k-means with random initialization of the centers (see Fig.1.D). K-means algorithm assigns each ROI to a cluster, which lead us to assume that the largest cluster in a kind of garbage collector that is discarded before further processing.
4 Fascicle bundles. The final step of the clustering consists in moving back to the fascicle world. Each cluster of ROIs is used to extract a subset of fascicles from the global set. For robustness purpose, a fascicle only needs to be partly included in the ROI cluster to be selected. In practice, a fascicle is selected when at least 60% of its sampling is included (see Fig.1.E). A last processing aims at splitting the resulting set of fascicles into connected components, which is required to deal with some unpredictable behaviour of the K-means. The clustering, indeed, gathers sometimes two bundles symmetric across interhemispheric plane, which calls for the use of better algorithms in the future. This postprocessing can not be performed directly at the level of the ROI clusters because the ROIs including fiber crossing can be assigned to only one cluster, leading sometimes to disconnect a cluster of ROIs corresponding to a reasonable bundle. The analysis of the connectedness of a fascicle set is performed via embedding of the fascicles into a 3D volume. It should be noted that this postprocessing implies that the final number of fascicle bundles depends on the subject. 2.2 Clustering the bundles across subjects The goal of the second stage of clustering is the inference of a model of the bundles that should be found in any human brain. The underlying idea is that a simple clustering performed to match similar bundles could provide a list of candidates for such a model. These candidates would not have to be found in any brain during this inference stage, but should be included in the model of the white matter used to design further pattern recognition systems. What we have in mind is a model of the bundles used to label the fascicles of any new brain in a way similar to what has been developped to deal with cortical sulci [17]. Similarity measure for bundles. In order to match bundles across subjects, a shapebased similarity measure is required. Relying on iconic representations of the bundles allows the definition of discriminant measures stemming from the spatial normalization towards Talairach space. For this purpose, each fascicle bundle is converted first into a binary representation lying into Talairach space. Then, to reduce the influence of the non perfect affine spatial normalization, these binary representations are convoluted with a Gaussian kernel with 3mm standard deviation. Finally, for each pair of bundles (i, j), the similarity C ij is computed as the correlation coefficient between their respective smoothed representations. With a subject-based ordering of the bundles, the similarity matrix C has a block structure (see Fig. 2 and 3). Clustering. The second stage of clustering is also performed with a K-means algorithm. Each bundle is represented by a row of the similarity matrix C. It should be noted that the K-means can merge several bundles of the same subject into the same cluster. It should be noted also that a cluster does not have to include a bundle of each subject. It should be noted finally that the K-means has to assign each bundle to a cluster. These remarks highlight the fact that a postprocessing is required to clean up the clustering before building the bundle models. The idea is first to get rid of the clusters that are not reasonable representation of a bundle, second to get rid of the bundles that look like outliers in the cluster they belong to. Let N be the number of subjects. For the results presented in the following, the clusters gathering less than N/2 bundles were discarded because of a lack of reproductibil-
5 Fig. 2. The block-based structure of the matrix of similarity measures between bundles. Each M ij block corresponds to the one to one correlation coefficients computed between the bundles of the subject i and the bundles of the subjects j. The O ii blocks would be zero-based without smoothing of the binary representations. ity across subjects. The clusters made up of more than 2 N bundles were discarded as garbage collector clusters. Finally, inside each clusters, the statistics of the distance to the cluster center were computed. The outlier bundles were defined as those beyond one mean plus one standard deviation. The remaining clusters are considered as candidates to represent one of the building block of the human brain white matter organization. 3 Results 3.1 Clustering the fascicles Eleven subjects were processed. Their diffusion data have been acquired on a 1.5T with 41 different directions for the diffusion-weighting gradient. The resulting 3D volumes have a 128x128x60 resolution with 1.875x1.875x2 voxel sizes. BrainVISA software was triggered with 42x42x20 5mm cubic ROIs. The K-means was applied with 20 classes for each subject. After the connectivity-based postprocessing, we obtained between 20 and 25 bundles for each subject. An example of result can be found in figure Clustering the bundles across a population The second clustering was applied on the eleven subjects with a 25 classes K-means. After cleaning up the result, we obtained 17 reasonable clusters. The result can be visualized in figures 3 and 4. Some of these clusters seem to fit some well known bundles described in anatomical atlases, while the various bundles resulting from the split of the corpus callosum may have less anatomical meaning. 4 Discussion The inference of a model of the white matter organization will become mandatory to fully exploit the information embedded into MR diffusion-weighted images. While a straightforward approach for this purpose lies in the iconic atlas and spatial normalization principles, this paper advocates an alternative which aims at infering a higher level model of the bundle organization. The work proposed in this paper is still very
6 Axial Sagittal Coronal Fig. 3. Clustering the bundles across subjects: The matrix of correlation coefficient computed for all pairs of bundles and three orthogonal views of the result of the clustering. All subjects bundles have been gathered into Talairach space. Each color denotes a cluster of bundles that may correspond to a generic anatomical entity. Garbage clusters have been filtered out. Among the clusters that are candidates to stand for some known anatomical pathways, we noticed the following: (black) ALIC: Anterior Limb of Internal Capsule; (pink) ATR: Anterior Thalamic Radiation; (red) ILF: Inferior Fronto-Occipital; (red) IFO: Inferior Longitudinal Fasciculus; (braun and dark blue) GCC: Genu of Corpus Callosum; (dark green and violet blue) SCC: Splenium of corpus callosum; (light yellow) FX: Fornix. preliminary. However, it provides an overview of the various difficulties that have to be addressed to tackle the inference of such a model. A lot of the ad hoc choices presented above could be questioned and more sophisticated methods should be derived in the future. The key idea proposed in our preliminary work is the two stage inference strategy: building first a bundle-based representation before trying to match bundles across subjects. This idea allows the matching to deal with reasonable data sizes. The first clustering, however, can not be questioned during the matching stage which is problematic. Therefore, future approaches may lead to build multiscale representations of the in-
7 Fig. 4. Clustering the bundles across subjects: The result of the clustering for ten of the eleven subjects. For each subject two views are presented. Each color denotes a cluster that gathers bundle across most of the subjects. dividual data. For instance, each individual bundle may be split further into smaller bundles using one more non supervised clustering. Some other multiscale representations could stem from selecting other ranges of fascicle length in order to deal with short pathways like U-fiber bundles. As mentioned above, the model inference does not have to find each anatomical entity in every brain. This could be the job of a dedicated pattern recognition system developed to process large databases of diffusion-weighted data. Such a system would face a simpler situation where brains are processed one by one. Hence, the matching with the bundle model could be performed at the fascicle level. Each fascicle would
8 have to be labelled with the name of one of the entities of the model. Such a system would open the door to massive morphometric analysis of the white matter bundles, which would lead to numerous applications in neurosciences. References 1. P. J. Basser, S. Pajevic, C. Pierpaoli, J. Duda, and A. Aldroubi. In vivo fiber tractography using DT-MRI data. MRM, 44(4):625 32, T.E.J Behrens, M.W. Woolrich, M. Jenkinson, H. Johansen-Berg, R.G. Nunes, S. Clare, P.M. Matthews, J.M. Brady, and S.M. Smith. Characterization and propagation of uncertainty in diffusion-weighted MR imaging. Magnetic Resonance in Medicine, 50: , M. Björnemo, A. Brun, R. Kikinis, and C.-F. Westin. Regularized stochastic white matter tractography using diffusion tensor MRI. In MICCAI 02, LNCS 2488, pages , A. Brun, H. Knutsson, H.-J. Park, M. E. Shenton, and C.-F. Westin. Clustering fiber traces using normalized cuts. In MICCAI 2004, LNCS 3216, pages , T. E. Conturo, N. F. Lori, T. S. Cull, E. Akbudak, A. Z. Snyder, J. S. Shimony, R. C. McKinstry, H. Burton, and M. E. Raichle. Tracking neuronal fiber pathways in the living human brain. Proc. Natl. Acad. Sci. USA, 96: , August I. Corouge, S. Gouttard, and G. Gerig. A statistical shape model of individual fiber tracts extracted from diffusion tensor MRI. In MICCAI 2004, LNCS 3217, pages , J. Dejerine. Anatomie des centres nerveux. Paris: Rueff, P. Fillard, J. Gilmore, W. Lin, and G. Gerig. Quantitative analysis of white matter fiber properties along geodesic paths. In MICCAI 03, LNCS 2879, pages 16 23, M. Jackowski, C. Y. Kao, R. T. Constable, and L. H. Staib. Estimation of anatomical connectivity by anisotropic front propagation and diffusion tensor imaging. In MICCAI04, LNCS 3217, pages , D. K. Jones, L. D. Griffin, D. C. Alexander, M. Catani, M. A. Horsfield, R. Howard, and S. C. Williams. Spatial normalization and averaging of diffusion tensor mri data sets. Neuroimage, 17(2): , D Le Bihan, J.-F. Mangin, C. Poupon, C. A. Clark, S. Pappata, N. Molko N, and H. Chabriat. Diffusion tensor imaging: Concepts and applications. Journal of Magnetic Resonance Imaging, 13(4): , C. Lenglet, R. Deriche, and O. Faugeras. Inferring white matter geometry from diffusion tensor mri: Application to connectivity mapping. In 8th ECCV, E. Ludwig and L. Klinger. Atlas cerebri humani. Basel: Karger, S. Mori, B.J. Crain, V. P. Chacko, and P. C. M. van Zijl. Three dimensional tracking of axonal projections in the brain by magnetic resonance imaging. Ann. Neurol., 45: , G. J. M Parker and D. C. Alexander. Probabilistic Monte Carlo based mapping of cerebral connections utilising whole-brain crossing fibre information. In IPMI, pages , C. Poupon, C.A. Clark, V. Frouin, J. Régis, I. Bloch, D. Le Bihan, and J-F. Mangin. Regularization of diffusion-based direction maps for the tracking of brain white matter fascicles. NeuroImage, 12(2): , D. Rivière, J.-F. Mangin, D. Papadopoulos, J.-M. Martinez, V. Frouin, and J. Régis. Automatic recognition of cortical sulci of the human brain using a congregation of neural networks. Medical Image Analysis, 6(2):77 92, D. S. Tuch. Diffusion MRI of complex tissue structure. PhD thesis, MIT, D. Weinstein, G. Kindlmann, and E. Lundberg. Tensorlines: Advection-diffusion based propagation through diffusion tensor fields. In IEEE Visualization, pages , D. Xu, S. Mori, D. Shen, P. C. van Zijl, and C. Davatzikos. Spatial normalization of diffusion tensor fields. Magn Reson Med, 50(1): , 2003.
Fiber Selection from Diffusion Tensor Data based on Boolean Operators
 Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhof 1, G. Greiner 2, M. Buchfelder 3, C. Nimsky 4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhof 1, G. Greiner 2, M. Buchfelder 3, C. Nimsky 4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
Evaluation of Local Filter Approaches for Diffusion Tensor based Fiber Tracking
 Evaluation of Local Filter Approaches for Diffusion Tensor based Fiber Tracking D. Merhof 1, M. Buchfelder 2, C. Nimsky 3 1 Visual Computing, University of Konstanz, Konstanz 2 Department of Neurosurgery,
Evaluation of Local Filter Approaches for Diffusion Tensor based Fiber Tracking D. Merhof 1, M. Buchfelder 2, C. Nimsky 3 1 Visual Computing, University of Konstanz, Konstanz 2 Department of Neurosurgery,
NIH Public Access Author Manuscript Med Image Comput Comput Assist Interv. Author manuscript; available in PMC 2013 May 04.
 NIH Public Access Author Manuscript Published in final edited form as: Med Image Comput Comput Assist Interv. 2005 ; 8(0 1): 180 187. A Hamilton-Jacobi-Bellman approach to high angular resolution diffusion
NIH Public Access Author Manuscript Published in final edited form as: Med Image Comput Comput Assist Interv. 2005 ; 8(0 1): 180 187. A Hamilton-Jacobi-Bellman approach to high angular resolution diffusion
Automatic Quantification of DTI Parameters along Fiber Bundles
 Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein, Simon Hermann, Olaf Konrad, Horst K. Hahn, Heinz-Otto Peitgen MeVis Research, 28359 Bremen Email: klein@mevis.de Abstract. We introduce
Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein, Simon Hermann, Olaf Konrad, Horst K. Hahn, Heinz-Otto Peitgen MeVis Research, 28359 Bremen Email: klein@mevis.de Abstract. We introduce
Automatic Quantification of DTI Parameters along Fiber Bundles
 Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein 1, Simon Hermann 1, Olaf Konrad 1, Horst K. Hahn 1, and Heinz-Otto Peitgen 1 1 MeVis Research, 28359 Bremen Email: klein@mevis.de
Automatic Quantification of DTI Parameters along Fiber Bundles Jan Klein 1, Simon Hermann 1, Olaf Konrad 1, Horst K. Hahn 1, and Heinz-Otto Peitgen 1 1 MeVis Research, 28359 Bremen Email: klein@mevis.de
Fiber Selection from Diffusion Tensor Data based on Boolean Operators
 Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhofl, G. Greiner 2, M. Buchfelder 3, C. Nimsky4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhofl, G. Greiner 2, M. Buchfelder 3, C. Nimsky4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
A Method for Registering Diffusion Weighted Magnetic Resonance Images
 A Method for Registering Diffusion Weighted Magnetic Resonance Images Xiaodong Tao and James V. Miller GE Research, Niskayuna, New York, USA Abstract. Diffusion weighted magnetic resonance (DWMR or DW)
A Method for Registering Diffusion Weighted Magnetic Resonance Images Xiaodong Tao and James V. Miller GE Research, Niskayuna, New York, USA Abstract. Diffusion weighted magnetic resonance (DWMR or DW)
HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Diffusion Tensor Processing and Visualization
 NA-MIC National Alliance for Medical Image Computing http://na-mic.org Diffusion Tensor Processing and Visualization Guido Gerig University of Utah Martin Styner, UNC NAMIC: National Alliance for Medical
NA-MIC National Alliance for Medical Image Computing http://na-mic.org Diffusion Tensor Processing and Visualization Guido Gerig University of Utah Martin Styner, UNC NAMIC: National Alliance for Medical
Reconstruction of Fiber Trajectories via Population-Based Estimation of Local Orientations
 IDEA Reconstruction of Fiber Trajectories via Population-Based Estimation of Local Orientations Pew-Thian Yap, John H. Gilmore, Weili Lin, Dinggang Shen Email: ptyap@med.unc.edu 2011-09-21 Poster: P2-46-
IDEA Reconstruction of Fiber Trajectories via Population-Based Estimation of Local Orientations Pew-Thian Yap, John H. Gilmore, Weili Lin, Dinggang Shen Email: ptyap@med.unc.edu 2011-09-21 Poster: P2-46-
HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu HST.583 Functional Magnetic Resonance Imaging: Data Acquisition and Analysis Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Analysis of White Matter Fiber Tracts via Fiber Clustering and Parametrization
 Analysis of White Matter Fiber Tracts via Fiber Clustering and Parametrization Sylvain Gouttard 2,4, Isabelle Corouge 1,2, Weili Lin 3, John Gilmore 2, and Guido Gerig 1,2 1 Departments of Computer Science,
Analysis of White Matter Fiber Tracts via Fiber Clustering and Parametrization Sylvain Gouttard 2,4, Isabelle Corouge 1,2, Weili Lin 3, John Gilmore 2, and Guido Gerig 1,2 1 Departments of Computer Science,
FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING. Francesca Pizzorni Ferrarese
 FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING Francesca Pizzorni Ferrarese Pipeline overview WM and GM Segmentation Registration Data reconstruction Tractography
FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING Francesca Pizzorni Ferrarese Pipeline overview WM and GM Segmentation Registration Data reconstruction Tractography
Quantitative Analysis of White Matter Fiber Properties along Geodesic Paths
 Quantitative Analysis of White Matter Fiber Properties along Geodesic Paths Pierre Fillard 1,4, John Gilmore 2, Joseph Piven 2, Weili Lin 3, and Guido Gerig 1,2 1 Department of Computer Science, 2 Department
Quantitative Analysis of White Matter Fiber Properties along Geodesic Paths Pierre Fillard 1,4, John Gilmore 2, Joseph Piven 2, Weili Lin 3, and Guido Gerig 1,2 1 Department of Computer Science, 2 Department
NEURO M203 & BIOMED M263 WINTER 2014
 NEURO M203 & BIOMED M263 WINTER 2014 MRI Lab 2: Neuroimaging Connectivity Lab In today s lab we will work with sample diffusion imaging data and the group averaged fmri data collected during your scanning
NEURO M203 & BIOMED M263 WINTER 2014 MRI Lab 2: Neuroimaging Connectivity Lab In today s lab we will work with sample diffusion imaging data and the group averaged fmri data collected during your scanning
Neural Network-Assisted Fiber Tracking of Synthetic and White Matter DT-MR Images
 Neural Network-Assisted Fiber Tracking of Synthetic and White Matter DT-MR Images L.M. San-José-Revuelta, M. Martín-Fernández and C. Alberola-López Abstract In this paper, a recently developed fiber tracking
Neural Network-Assisted Fiber Tracking of Synthetic and White Matter DT-MR Images L.M. San-José-Revuelta, M. Martín-Fernández and C. Alberola-López Abstract In this paper, a recently developed fiber tracking
White Matter Tractography Using Diffusion Tensor Deflection
 Human Brain Mapping 18:306 321(2003) White Matter Tractography Using Diffusion Tensor Deflection Mariana Lazar, 1 David M. Weinstein, 2 Jay S. Tsuruda, 3 Khader M. Hasan, 4,8 Konstantinos Arfanakis, 4
Human Brain Mapping 18:306 321(2003) White Matter Tractography Using Diffusion Tensor Deflection Mariana Lazar, 1 David M. Weinstein, 2 Jay S. Tsuruda, 3 Khader M. Hasan, 4,8 Konstantinos Arfanakis, 4
Optimized diffusion gradient orientation schemes for corrupted clinical DTI data sets
 Magn Reson Mater Phy (2006) DOI 10.1007/s10334-006-0036-0 RESEARCH ARTICLE J. Dubois C. Poupon F. Lethimonnier D. Le Bihan Optimized diffusion gradient orientation schemes for corrupted clinical DTI data
Magn Reson Mater Phy (2006) DOI 10.1007/s10334-006-0036-0 RESEARCH ARTICLE J. Dubois C. Poupon F. Lethimonnier D. Le Bihan Optimized diffusion gradient orientation schemes for corrupted clinical DTI data
Introduction of a Quantitative Evaluation Method for White Matter Tractography using a HARDI-based Reference
 Introduction of a Quantitative Evaluation Method for White Matter Tractography using a HARDI-based Reference Peter F. Neher 1, Bram Stieltjes 2, Hans-Peter Meinzer 1, and Klaus H. Fritzsche 1,2, 1 German
Introduction of a Quantitative Evaluation Method for White Matter Tractography using a HARDI-based Reference Peter F. Neher 1, Bram Stieltjes 2, Hans-Peter Meinzer 1, and Klaus H. Fritzsche 1,2, 1 German
On Classifying Disease-Induced Patterns in the Brain Using Diffusion Tensor Images
 On Classifying Disease-Induced Patterns in the Brain Using Diffusion Tensor Images Peng Wang 1,2 and Ragini Verma 1 1 Section of Biomedical Image Analysis, Department of Radiology, University of Pennsylvania,
On Classifying Disease-Induced Patterns in the Brain Using Diffusion Tensor Images Peng Wang 1,2 and Ragini Verma 1 1 Section of Biomedical Image Analysis, Department of Radiology, University of Pennsylvania,
Probabilistic Monte Carlo Based Mapping of Cerebral Connections Utilising Whole-Brain Crossing Fibre Information
 Probabilistic Monte Carlo Based Mapping of Cerebral Connections Utilising Whole-Brain Crossing Fibre Information Geoff J.M. Parker 1 and Daniel C. Alexander 2 1 Imaging Science & Biomedical Engineering,
Probabilistic Monte Carlo Based Mapping of Cerebral Connections Utilising Whole-Brain Crossing Fibre Information Geoff J.M. Parker 1 and Daniel C. Alexander 2 1 Imaging Science & Biomedical Engineering,
Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge
 Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge Christian Wasserthal 1, Karin Engel 1, Karsten Rink 1 und André Brechmann
Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge Christian Wasserthal 1, Karin Engel 1, Karsten Rink 1 und André Brechmann
ARTICLE IN PRESS YNIMG-03232; No. of pages: 12; 4C: 6, 7, 8, 9, 10
 YNIMG-03232; No. of pages: 12; 4C: 6, 7, 8, 9, 10 DTD 5 www.elsevier.com/locate/ynimg NeuroImage xx (2005) xxx xxx Flow-based fiber tracking with diffusion tensor and q-ball data: Validation and comparison
YNIMG-03232; No. of pages: 12; 4C: 6, 7, 8, 9, 10 DTD 5 www.elsevier.com/locate/ynimg NeuroImage xx (2005) xxx xxx Flow-based fiber tracking with diffusion tensor and q-ball data: Validation and comparison
Uncertainty in White Matter Fiber Tractography
 Uncertainty in White Matter Fiber Tractography Ola Friman and Carl-Fredrik Westin Laboratory of Mathematics in Imaging, Department of Radiology Brigham and Women s Hospital, Harvard Medical School Abstract.
Uncertainty in White Matter Fiber Tractography Ola Friman and Carl-Fredrik Westin Laboratory of Mathematics in Imaging, Department of Radiology Brigham and Women s Hospital, Harvard Medical School Abstract.
Group Statistics of DTI Fiber Bundles Using Spatial Functions of Tensor Measures
 Group Statistics of DTI Fiber Bundles Using Spatial Functions of Tensor Measures Casey B. Goodlett 1,2, P. Thomas Fletcher 1,2, John H. Gilmore 3, and Guido Gerig 1,2 1 School of Computing, University
Group Statistics of DTI Fiber Bundles Using Spatial Functions of Tensor Measures Casey B. Goodlett 1,2, P. Thomas Fletcher 1,2, John H. Gilmore 3, and Guido Gerig 1,2 1 School of Computing, University
The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy
 The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy Sokratis K. Makrogiannis, PhD From post-doctoral research at SBIA lab, Department of Radiology,
The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy Sokratis K. Makrogiannis, PhD From post-doctoral research at SBIA lab, Department of Radiology,
Tractography via the Ensemble Average Propagator in diffusion MRI
 Tractography via the Ensemble Average Propagator in diffusion MRI Sylvain Merlet, Anne-Charlotte Philippe, Rachid Deriche, Maxime Descoteaux To cite this version: Sylvain Merlet, Anne-Charlotte Philippe,
Tractography via the Ensemble Average Propagator in diffusion MRI Sylvain Merlet, Anne-Charlotte Philippe, Rachid Deriche, Maxime Descoteaux To cite this version: Sylvain Merlet, Anne-Charlotte Philippe,
Coloring of DT-MRI Fiber Traces using Laplacian Eigenmaps
 Coloring of DT-MRI Fiber Traces using Laplacian Eigenmaps Anders Brun 1, Hae-Jeong Park 2, Hans Knutsson 3, and Carl-Fredrik Westin 1 1 Laboratory of Mathematics in Imaging, Brigham and Women s Hospital,
Coloring of DT-MRI Fiber Traces using Laplacian Eigenmaps Anders Brun 1, Hae-Jeong Park 2, Hans Knutsson 3, and Carl-Fredrik Westin 1 1 Laboratory of Mathematics in Imaging, Brigham and Women s Hospital,
MITK Global Tractography
 MITK Global Tractography Peter F. Neher a, Bram Stieltjes b, Marco Reisert c, Ignaz Reicht a, Hans-Peter Meinzer a, Klaus H. Fritzsche a,b a German Cancer Research Center, Medical and Biological Informatics,
MITK Global Tractography Peter F. Neher a, Bram Stieltjes b, Marco Reisert c, Ignaz Reicht a, Hans-Peter Meinzer a, Klaus H. Fritzsche a,b a German Cancer Research Center, Medical and Biological Informatics,
3D Stochastic Completion Fields for Fiber Tractography
 3D Stochastic Completion Fields for Fiber Tractography Parya Momayyez School of Computer Science Centre for Intelligent Machines McGill University, Montréal, QC, Canada pmamay@cim.mcgill.ca Kaleem Siddiqi
3D Stochastic Completion Fields for Fiber Tractography Parya Momayyez School of Computer Science Centre for Intelligent Machines McGill University, Montréal, QC, Canada pmamay@cim.mcgill.ca Kaleem Siddiqi
Direct Registration of White Matter Tractographies with Application to Atlas Construction
 Direct Registration of White Matter Tractographies with Application to Atlas Construction Arnaldo Mayer, Hayit Greenspan Medical image processing lab, Tel-Aviv University, Ramat-Aviv, Israel arnakdom@post.tau.ac.il,
Direct Registration of White Matter Tractographies with Application to Atlas Construction Arnaldo Mayer, Hayit Greenspan Medical image processing lab, Tel-Aviv University, Ramat-Aviv, Israel arnakdom@post.tau.ac.il,
Tractography via the Ensemble Average Propagator in Diffusion MRI
 Tractography via the Ensemble Average Propagator in Diffusion MRI Sylvain Merlet 1, Anne-Charlotte Philippe 1, Rachid Deriche 1, and Maxime Descoteaux 2 1 Athena Project-Team, INRIA Sophia Antipolis -
Tractography via the Ensemble Average Propagator in Diffusion MRI Sylvain Merlet 1, Anne-Charlotte Philippe 1, Rachid Deriche 1, and Maxime Descoteaux 2 1 Athena Project-Team, INRIA Sophia Antipolis -
syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets
 syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets Julien Gervais; Lisa Chuah Siemens Healthcare, Magnetic
syngo.mr Neuro 3D: Your All-In-One Post Processing, Visualization and Reporting Engine for BOLD Functional and Diffusion Tensor MR Imaging Datasets Julien Gervais; Lisa Chuah Siemens Healthcare, Magnetic
Acceleration of Probabilistic Tractography Using Multi-GPU Parallel Processing. Jungsoo Lee, Sun Mi Park, Dae-Shik Kim
 Acceleration of Probabilistic Tractography Using Multi-GPU Parallel Processing Jungsoo Lee, Sun Mi Park, Dae-Shik Kim Introduction In particular, probabilistic tractography requires relatively long computation
Acceleration of Probabilistic Tractography Using Multi-GPU Parallel Processing Jungsoo Lee, Sun Mi Park, Dae-Shik Kim Introduction In particular, probabilistic tractography requires relatively long computation
Labeling of ambiguous subvoxel fibre bundle configurations in high angular resolution diffusion MRI
 www.elsevier.com/locate/ynimg NeuroImage 41 (2008) 58 68 Labeling of ambiguous subvoxel fibre bundle configurations in high angular resolution diffusion MRI Peter Savadjiev, a, Jennifer S.W. Campbell,
www.elsevier.com/locate/ynimg NeuroImage 41 (2008) 58 68 Labeling of ambiguous subvoxel fibre bundle configurations in high angular resolution diffusion MRI Peter Savadjiev, a, Jennifer S.W. Campbell,
NIH Public Access Author Manuscript Proc SPIE. Author manuscript; available in PMC 2012 October 10.
 NIH Public Access Author Manuscript Published in final edited form as: Proc SPIE. 2011 ; 7962: 79620S. doi:10.1117/12.878291. Efficient, Graph-based White Matter Connectivity from Orientation Distribution
NIH Public Access Author Manuscript Published in final edited form as: Proc SPIE. 2011 ; 7962: 79620S. doi:10.1117/12.878291. Efficient, Graph-based White Matter Connectivity from Orientation Distribution
A Ray-based Approach for Boundary Estimation of Fiber Bundles Derived from Diffusion Tensor Imaging
 A Ray-based Approach for Boundary Estimation of Fiber Bundles Derived from Diffusion Tensor Imaging M. H. A. Bauer 1,3, S. Barbieri 2, J. Klein 2, J. Egger 1,3, D. Kuhnt 1, B. Freisleben 3, H.-K. Hahn
A Ray-based Approach for Boundary Estimation of Fiber Bundles Derived from Diffusion Tensor Imaging M. H. A. Bauer 1,3, S. Barbieri 2, J. Klein 2, J. Egger 1,3, D. Kuhnt 1, B. Freisleben 3, H.-K. Hahn
Fiber Tract Clustering on Manifolds With Dual Rooted-Graphs
 Fiber Tract Clustering on Manifolds With Dual Rooted-Graphs Andy Tsai 1,2 Carl-Fredrik Westin 1 Alfred O. Hero III 3 Alan S. Willsky 2 1 Dept. of Radiology 2 Dept. of Electrical Eng. 3 Dept. of Electrical
Fiber Tract Clustering on Manifolds With Dual Rooted-Graphs Andy Tsai 1,2 Carl-Fredrik Westin 1 Alfred O. Hero III 3 Alan S. Willsky 2 1 Dept. of Radiology 2 Dept. of Electrical Eng. 3 Dept. of Electrical
Appendix E1. Supplementary Methods. MR Image Acquisition. MR Image Analysis
 RSNA, 2015 10.1148/radiol.2015150532 Appendix E1 Supplementary Methods MR Image Acquisition By using a 1.5-T system (Avanto, Siemens Medical, Erlangen, Germany) under a program of regular maintenance (no
RSNA, 2015 10.1148/radiol.2015150532 Appendix E1 Supplementary Methods MR Image Acquisition By using a 1.5-T system (Avanto, Siemens Medical, Erlangen, Germany) under a program of regular maintenance (no
Diffusion Tractography of the Corticospinal Tract with Multi-fiber Orientation Filtering
 Diffusion Tractography of the Corticospinal Tract with Multi-fiber Orientation Filtering Ryan P. Cabeen, David H. Laidlaw Computer Science Department Brown University Providence, RI, USA {cabeen,dhl}@cs.brown.edu
Diffusion Tractography of the Corticospinal Tract with Multi-fiber Orientation Filtering Ryan P. Cabeen, David H. Laidlaw Computer Science Department Brown University Providence, RI, USA {cabeen,dhl}@cs.brown.edu
Corpus Callosum Subdivision based on a Probabilistic Model of Inter-Hemispheric Connectivity
 Corpus Callosum Subdivision based on a Probabilistic Model of Inter-Hemispheric Connectivity Martin A. Styner 1,2, Ipek Oguz 1, Rachel Gimpel Smith 2, Carissa Cascio 2, and Matthieu Jomier 1 1 Dept. of
Corpus Callosum Subdivision based on a Probabilistic Model of Inter-Hemispheric Connectivity Martin A. Styner 1,2, Ipek Oguz 1, Rachel Gimpel Smith 2, Carissa Cascio 2, and Matthieu Jomier 1 1 Dept. of
Generation of Hulls Encompassing Neuronal Pathways Based on Tetrahedralization and 3D Alpha Shapes
 Generation of Hulls Encompassing Neuronal Pathways Based on Tetrahedralization and 3D Alpha Shapes Dorit Merhof 1,2, Martin Meister 1, Ezgi Bingöl 1, Peter Hastreiter 1,2, Christopher Nimsky 2,3, Günther
Generation of Hulls Encompassing Neuronal Pathways Based on Tetrahedralization and 3D Alpha Shapes Dorit Merhof 1,2, Martin Meister 1, Ezgi Bingöl 1, Peter Hastreiter 1,2, Christopher Nimsky 2,3, Günther
Feasibility and Advantages of Diffusion Weighted Imaging Atlas Construction in Q-Space
 Feasibility and Advantages of Diffusion Weighted Imaging Atlas Construction in Q-Space Thijs Dhollander 1,2, Jelle Veraart 3,WimVanHecke 1,4,5, Frederik Maes 1,2, Stefan Sunaert 1,4,JanSijbers 3, and Paul
Feasibility and Advantages of Diffusion Weighted Imaging Atlas Construction in Q-Space Thijs Dhollander 1,2, Jelle Veraart 3,WimVanHecke 1,4,5, Frederik Maes 1,2, Stefan Sunaert 1,4,JanSijbers 3, and Paul
DIFFUSION TENSOR IMAGING ANALYSIS. Using Analyze
 DIFFUSION TENSOR IMAGING ANALYSIS Using Analyze 2 Table of Contents 1. Introduction page 3 2. Loading DTI Data page 4 3. Computing DTI Maps page 5 4. Defining ROIs for Fiber Tracking page 6 5. Visualizing
DIFFUSION TENSOR IMAGING ANALYSIS Using Analyze 2 Table of Contents 1. Introduction page 3 2. Loading DTI Data page 4 3. Computing DTI Maps page 5 4. Defining ROIs for Fiber Tracking page 6 5. Visualizing
The Application of GPU Particle Tracing to Diffusion Tensor Field Visualization
 The Application of GPU Particle Tracing to Diffusion Tensor Field Visualization Polina Kondratieva, Jens Krüger, Rüdiger Westermann Computer Graphics and Visualization Group Technische Universität München
The Application of GPU Particle Tracing to Diffusion Tensor Field Visualization Polina Kondratieva, Jens Krüger, Rüdiger Westermann Computer Graphics and Visualization Group Technische Universität München
THE brain is a massively interconnected organ. Individual
 IEEE TRANSACTIONS ON VISUALIZATION AND COMPUTER GRAPHICS 1 Exploring Connectivity of the Brain s White Matter with Dynamic Queries Anthony Sherbondy, David Akers, Rachel Mackenzie, Robert Dougherty, and
IEEE TRANSACTIONS ON VISUALIZATION AND COMPUTER GRAPHICS 1 Exploring Connectivity of the Brain s White Matter with Dynamic Queries Anthony Sherbondy, David Akers, Rachel Mackenzie, Robert Dougherty, and
Diffusion Imaging Models 1: from DTI to HARDI models
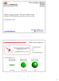 Diffusion Imaging Models 1: from DTI to HARDI models Flavio Dell Acqua, PhD. www.natbrainlab.com flavio.dellacqua@kcl.ac.uk @flaviodellacqua Diffusion Tensor Imaging (DTI) z λ 1 λ 2 The profile of the
Diffusion Imaging Models 1: from DTI to HARDI models Flavio Dell Acqua, PhD. www.natbrainlab.com flavio.dellacqua@kcl.ac.uk @flaviodellacqua Diffusion Tensor Imaging (DTI) z λ 1 λ 2 The profile of the
A Tract-Specific Framework for White Matter Morphometry Combining Macroscopic and Microscopic Tract Features
 A Tract-Specific Framework for White Matter Morphometry Combining Macroscopic and Microscopic Tract Features Hui Zhang, Suyash P. Awate, Sandhitsu R. Das, John H. Woo, Elias R. Melhem, James C. Gee, and
A Tract-Specific Framework for White Matter Morphometry Combining Macroscopic and Microscopic Tract Features Hui Zhang, Suyash P. Awate, Sandhitsu R. Das, John H. Woo, Elias R. Melhem, James C. Gee, and
Advanced Visual Medicine: Techniques for Visual Exploration & Analysis
 Advanced Visual Medicine: Techniques for Visual Exploration & Analysis Interactive Visualization of Multimodal Volume Data for Neurosurgical Planning Felix Ritter, MeVis Research Bremen Multimodal Neurosurgical
Advanced Visual Medicine: Techniques for Visual Exploration & Analysis Interactive Visualization of Multimodal Volume Data for Neurosurgical Planning Felix Ritter, MeVis Research Bremen Multimodal Neurosurgical
Diffusion-MRI processing for group analysis
 Diffusion-MRI processing for group analysis Felix Renard IRMaGe: Inserm US 17 / CNRS UMS 3552 University Hospital of Grenoble - France 25/09/2015 felixrenard@gmail.com 1 Diffusion-MRI processing for group
Diffusion-MRI processing for group analysis Felix Renard IRMaGe: Inserm US 17 / CNRS UMS 3552 University Hospital of Grenoble - France 25/09/2015 felixrenard@gmail.com 1 Diffusion-MRI processing for group
Tech Report Computer Science Department, UNC Chapel Hill
 Tech Report Computer Science Department, UNC Chapel Hill Brain morphometry by distance measurement in a non-euclidean, curvilinear space Martin Styner 1, Thomas Coradi 2 and Guido Gerig 1 1 Dept. of Computer
Tech Report Computer Science Department, UNC Chapel Hill Brain morphometry by distance measurement in a non-euclidean, curvilinear space Martin Styner 1, Thomas Coradi 2 and Guido Gerig 1 1 Dept. of Computer
Diffusion Tensor Imaging and Reading Development
 Diffusion Tensor Imaging and Reading Development Bob Dougherty Stanford Institute for Reading and Learning Reading and Anatomy Every brain is different... Not all brains optimized for highly proficient
Diffusion Tensor Imaging and Reading Development Bob Dougherty Stanford Institute for Reading and Learning Reading and Anatomy Every brain is different... Not all brains optimized for highly proficient
Unveiling the White Matter Microstructure in 22q11.2 Deletion Syndrome. Julio Villalón
 Unveiling the White Matter Microstructure in 22q11.2 Deletion Syndrome Julio Villalón What is 22q11.2 Deletion Syndrome? Yes, there is a mouse model!! Modified from Meechan et al. 2015 How are neurons
Unveiling the White Matter Microstructure in 22q11.2 Deletion Syndrome Julio Villalón What is 22q11.2 Deletion Syndrome? Yes, there is a mouse model!! Modified from Meechan et al. 2015 How are neurons
A Novel Contrast for DTI Visualization for Thalamus Delineation
 A Novel Contrast for DTI Visualization for Thalamus Delineation Xian Fan a, Meredith Thompson a,b, John A. Bogovic a, Pierre-Louis Bazin c, Jerry L. Prince a,c a Johns Hopkins University, Baltimore, MD,
A Novel Contrast for DTI Visualization for Thalamus Delineation Xian Fan a, Meredith Thompson a,b, John A. Bogovic a, Pierre-Louis Bazin c, Jerry L. Prince a,c a Johns Hopkins University, Baltimore, MD,
ISMI: A classification index for High Angular Resolution Diffusion Imaging
 ISMI: A classification index for High Angular Resolution Diffusion Imaging D. Röttger, D. Dudai D. Merhof and S. Müller Institute for Computational Visualistics, University of Koblenz-Landau, Germany Visual
ISMI: A classification index for High Angular Resolution Diffusion Imaging D. Röttger, D. Dudai D. Merhof and S. Müller Institute for Computational Visualistics, University of Koblenz-Landau, Germany Visual
CONTRACTING ORGANIZATION: The Cleveland Clinic Foundation, Cleveland, OH 44195
 AD Award Number: W81XWH-11-1-0362 TITLE: Whole Brain Networks for Treatment for Epilepsy PRINCIPAL INVESTIGATOR: Ken Sakaie, Ph.D. CONTRACTING ORGANIZATION: The Cleveland Clinic Foundation, Cleveland,
AD Award Number: W81XWH-11-1-0362 TITLE: Whole Brain Networks for Treatment for Epilepsy PRINCIPAL INVESTIGATOR: Ken Sakaie, Ph.D. CONTRACTING ORGANIZATION: The Cleveland Clinic Foundation, Cleveland,
NIH Public Access Author Manuscript Med Image Comput Comput Assist Interv. Author manuscript; available in PMC 2009 December 4.
 NIH Public Access Author Manuscript Med Image Comput Comput Assist Interv. Author manuscript; available in PMC 2009 December 4. Published in final edited form as: Med Image Comput Comput Assist Interv.
NIH Public Access Author Manuscript Med Image Comput Comput Assist Interv. Author manuscript; available in PMC 2009 December 4. Published in final edited form as: Med Image Comput Comput Assist Interv.
Clustering Fiber Traces Using Normalized Cuts
 Clustering Fiber Traces Using Normalized Cuts Anders Brun 1,4, Hans Knutsson 1, Hae-Jeong Park 2,3,4, Martha E. Shenton 2, and Carl-Fredrik Westin 4 1 Department. of Biomedical Engineering, Linköping University,
Clustering Fiber Traces Using Normalized Cuts Anders Brun 1,4, Hans Knutsson 1, Hae-Jeong Park 2,3,4, Martha E. Shenton 2, and Carl-Fredrik Westin 4 1 Department. of Biomedical Engineering, Linköping University,
Identifying white-matter fiber bundles in DTI data using an automated proximity-based fiber-clustering method
 1 Identifying white-matter fiber bundles in DTI data using an automated proximity-based fiber-clustering method Song Zhang 1, Stephen Correia 2, and David H. Laidlaw 3 1 Department of Computer Science
1 Identifying white-matter fiber bundles in DTI data using an automated proximity-based fiber-clustering method Song Zhang 1, Stephen Correia 2, and David H. Laidlaw 3 1 Department of Computer Science
MITK-DI. A new Diffusion Imaging Component for MITK. Klaus Fritzsche, Hans-Peter Meinzer
 MITK-DI A new Diffusion Imaging Component for MITK Klaus Fritzsche, Hans-Peter Meinzer Division of Medical and Biological Informatics, DKFZ Heidelberg k.fritzsche@dkfz-heidelberg.de Abstract. Diffusion-MRI
MITK-DI A new Diffusion Imaging Component for MITK Klaus Fritzsche, Hans-Peter Meinzer Division of Medical and Biological Informatics, DKFZ Heidelberg k.fritzsche@dkfz-heidelberg.de Abstract. Diffusion-MRI
Supplementary Online Content
 Supplementary Online Content Sun H, Lui S, Yao L, et al. Two patterns of white matter abnormalities in medication-naive patients with first-episode schizophrenia revealed by diffusion tensor imaging and
Supplementary Online Content Sun H, Lui S, Yao L, et al. Two patterns of white matter abnormalities in medication-naive patients with first-episode schizophrenia revealed by diffusion tensor imaging and
Generating Fiber Crossing Phantoms Out of Experimental DWIs
 Generating Fiber Crossing Phantoms Out of Experimental DWIs Matthan Caan 1,2, Anne Willem de Vries 2, Ganesh Khedoe 2,ErikAkkerman 1, Lucas van Vliet 2, Kees Grimbergen 1, and Frans Vos 1,2 1 Department
Generating Fiber Crossing Phantoms Out of Experimental DWIs Matthan Caan 1,2, Anne Willem de Vries 2, Ganesh Khedoe 2,ErikAkkerman 1, Lucas van Vliet 2, Kees Grimbergen 1, and Frans Vos 1,2 1 Department
NIH Public Access Author Manuscript Med Image Comput Comput Assist Interv. Author manuscript; available in PMC 2012 February 27.
 NIH Public Access Author Manuscript Med Image Comput Comput Assist Interv. Author manuscript; available in PMC 2012 February 27. Published in final edited form as: Med Image Comput Comput Assist Interv.
NIH Public Access Author Manuscript Med Image Comput Comput Assist Interv. Author manuscript; available in PMC 2012 February 27. Published in final edited form as: Med Image Comput Comput Assist Interv.
Diffusion Tensor Imaging Analysis of Regional Whiter Matter Changes. along the Cingulum in Mild Cognitive Impairment
 1 Diffusion Tensor Imaging Analysis of Regional Whiter Matter Changes along the Cingulum in Mild Cognitive Impairment Xuwei Liang, Ning Kang, Stephen E. Rose, Jonathan B. Chalk, Jun Zhang Laboratory for
1 Diffusion Tensor Imaging Analysis of Regional Whiter Matter Changes along the Cingulum in Mild Cognitive Impairment Xuwei Liang, Ning Kang, Stephen E. Rose, Jonathan B. Chalk, Jun Zhang Laboratory for
Estimation of the Underlying Fiber Orientation Using Spherical k-means Method from the Diffusion ODF in HARDI Data
 Estimation of the Underlying Fiber Orientation Using Spherical k-means Method from the Diffusion ODF in HARDI Data Huaizhong Zhang, Martin McGinnity, Sonya Coleman and Min Jing Intelligent Systems Research
Estimation of the Underlying Fiber Orientation Using Spherical k-means Method from the Diffusion ODF in HARDI Data Huaizhong Zhang, Martin McGinnity, Sonya Coleman and Min Jing Intelligent Systems Research
The organization of the human cerebral cortex estimated by intrinsic functional connectivity
 1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
Atelier 2 : Calcul Haute Performance et Sciences du Vivant Forum er juillet, Paris, France
 From Diffusion MR Image Analysis to Whole Brain Connectivity Simulation Jean-Philippe Thiran EPFL Lausanne, Switzerland EPFL - Lausanne HPC in life sciences at EPFL The Blue Brain project: create a biologically
From Diffusion MR Image Analysis to Whole Brain Connectivity Simulation Jean-Philippe Thiran EPFL Lausanne, Switzerland EPFL - Lausanne HPC in life sciences at EPFL The Blue Brain project: create a biologically
Introduction to Neuroimaging Janaina Mourao-Miranda
 Introduction to Neuroimaging Janaina Mourao-Miranda Neuroimaging techniques have changed the way neuroscientists address questions about functional anatomy, especially in relation to behavior and clinical
Introduction to Neuroimaging Janaina Mourao-Miranda Neuroimaging techniques have changed the way neuroscientists address questions about functional anatomy, especially in relation to behavior and clinical
Measuring longitudinal brain changes in humans and small animal models. Christos Davatzikos
 Measuring longitudinal brain changes in humans and small animal models Christos Davatzikos Section of Biomedical Image Analysis University of Pennsylvania (Radiology) http://www.rad.upenn.edu/sbia Computational
Measuring longitudinal brain changes in humans and small animal models Christos Davatzikos Section of Biomedical Image Analysis University of Pennsylvania (Radiology) http://www.rad.upenn.edu/sbia Computational
The Effects of Inter-slice Gap Thickness on Parametric Methods for Diffusion Tensor Imaging Abstract
 The Effects of Inter-slice Gap Thickness on Parametric Methods for Diffusion Tensor Imaging Beth Hutchinson; Neuroscience Training program, Statistics 592 Abstract Although high-resolution, whole-brain
The Effects of Inter-slice Gap Thickness on Parametric Methods for Diffusion Tensor Imaging Beth Hutchinson; Neuroscience Training program, Statistics 592 Abstract Although high-resolution, whole-brain
Complex Fiber Visualization
 Annales Mathematicae et Informaticae 34 (2007) pp. 103 109 http://www.ektf.hu/tanszek/matematika/ami Complex Fiber Visualization Henrietta Tomán a, Róbert Tornai b, Marianna Zichar c a Department of Computer
Annales Mathematicae et Informaticae 34 (2007) pp. 103 109 http://www.ektf.hu/tanszek/matematika/ami Complex Fiber Visualization Henrietta Tomán a, Róbert Tornai b, Marianna Zichar c a Department of Computer
Super-Resolution Reconstruction of Diffusion-Weighted Images from Distortion Compensated Orthogonal Anisotropic Acquisitions.
 Super-Resolution Reconstruction of Diffusion-Weighted Images from Distortion Compensated Orthogonal Anisotropic Acquisitions. Benoit Scherrer Ali Gholipour Simon K. Warfield Children s Hospital Boston,
Super-Resolution Reconstruction of Diffusion-Weighted Images from Distortion Compensated Orthogonal Anisotropic Acquisitions. Benoit Scherrer Ali Gholipour Simon K. Warfield Children s Hospital Boston,
Dual Tensor Atlas Generation Based on a Cohort of Coregistered non-hardi Datasets
 Dual Tensor Atlas Generation Based on a Cohort of Coregistered non-hardi Datasets Matthan Caan 1,2, Caroline Sage 3, Maaike van der Graaf 1, Cornelis Grimbergen 1, Stefan Sunaert 3, Lucas van Vliet 2,
Dual Tensor Atlas Generation Based on a Cohort of Coregistered non-hardi Datasets Matthan Caan 1,2, Caroline Sage 3, Maaike van der Graaf 1, Cornelis Grimbergen 1, Stefan Sunaert 3, Lucas van Vliet 2,
Surface-Based Structural Group Analysis of fmri Data
 Surface-Based Structural Group Analysis of fmri Data Grégory Operto 1,Cédric Clouchoux 1,Rémy Bulot 1,Jean-LucAnton 2, and Olivier Coulon 1 1 Laboratoire LSIS, UMR CNRS 6168, Marseille, France gregory.operto@univmed.fr
Surface-Based Structural Group Analysis of fmri Data Grégory Operto 1,Cédric Clouchoux 1,Rémy Bulot 1,Jean-LucAnton 2, and Olivier Coulon 1 1 Laboratoire LSIS, UMR CNRS 6168, Marseille, France gregory.operto@univmed.fr
Diffusion Imaging Visualization
 Diffusion Imaging Visualization Thomas Schultz URL: http://cg.cs.uni-bonn.de/schultz/ E-Mail: schultz@cs.uni-bonn.de 1 Outline Introduction to Diffusion Imaging Basic techniques Glyph-based Visualization
Diffusion Imaging Visualization Thomas Schultz URL: http://cg.cs.uni-bonn.de/schultz/ E-Mail: schultz@cs.uni-bonn.de 1 Outline Introduction to Diffusion Imaging Basic techniques Glyph-based Visualization
Automatic Generation of Training Data for Brain Tissue Classification from MRI
 Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. COCOSCO, Alex P. ZIJDENBOS, and Alan C. EVANS http://www.bic.mni.mcgill.ca/users/crisco/ McConnell Brain Imaging
Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. COCOSCO, Alex P. ZIJDENBOS, and Alan C. EVANS http://www.bic.mni.mcgill.ca/users/crisco/ McConnell Brain Imaging
A GA-based Approach for Parameter Estimation in DT-MRI Tracking Algorithms
 A GA-based Approach for Parameter Estimation in DT-MRI Tracking Algorithms L.M. San-José-Revuelta, M. Martín-Fernández and C. Alberola-López Abstract This paper expands upon previous work of the authors
A GA-based Approach for Parameter Estimation in DT-MRI Tracking Algorithms L.M. San-José-Revuelta, M. Martín-Fernández and C. Alberola-López Abstract This paper expands upon previous work of the authors
NeuroImage. Tract-based morphometry for white matter group analysis. Lauren J. O'Donnell a,b,, Carl-Fredrik Westin b, Alexandra J.
 NeuroImage 45 (2009) 832 844 Contents lists available at ScienceDirect NeuroImage journal homepage: www.elsevier.com/locate/ynimg Tract-based morphometry for white matter group analysis Lauren J. O'Donnell
NeuroImage 45 (2009) 832 844 Contents lists available at ScienceDirect NeuroImage journal homepage: www.elsevier.com/locate/ynimg Tract-based morphometry for white matter group analysis Lauren J. O'Donnell
Supplementary Online Content
 Supplementary Online Content Palacios EM, Sala-Llonch R, Junque C, Roig T, Tormos JM, Bargallo N, Vendrella P. Restingstate functional magnetic resonance imaging activity and connectivity and cognitive
Supplementary Online Content Palacios EM, Sala-Llonch R, Junque C, Roig T, Tormos JM, Bargallo N, Vendrella P. Restingstate functional magnetic resonance imaging activity and connectivity and cognitive
Local Atlas-based Adaptive Fiber Tracking
 Local Atlas-based Adaptive Fiber Tracking Jan Klein 1, Monique Meuschke 1, Benjamin Geisler 1, and Horst K. Hahn 1 Fraunhofer MEVIS Institute for Medical Image Computing, Bremen, Germany, {jan.klein,monique.meuschke,benjamin.geisler,horst.hahn}@mevis.fraunhofer.de
Local Atlas-based Adaptive Fiber Tracking Jan Klein 1, Monique Meuschke 1, Benjamin Geisler 1, and Horst K. Hahn 1 Fraunhofer MEVIS Institute for Medical Image Computing, Bremen, Germany, {jan.klein,monique.meuschke,benjamin.geisler,horst.hahn}@mevis.fraunhofer.de
Image Deformation using Velocity Fields: An Exact Solution
 Image Deformation using Velocity Fields: An Exact Solution Jeff Orchard University of Waterloo, Waterloo Ontario N2L 3G1, Canada, jorchard@cs.uwaterloo.ca Abstract. In image deformation, one of the challenges
Image Deformation using Velocity Fields: An Exact Solution Jeff Orchard University of Waterloo, Waterloo Ontario N2L 3G1, Canada, jorchard@cs.uwaterloo.ca Abstract. In image deformation, one of the challenges
Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm
 Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm Lin Chen and Gudrun Wagenknecht Central Institute for Electronics, Research Center Jülich, Jülich, Germany Email: l.chen@fz-juelich.de
Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm Lin Chen and Gudrun Wagenknecht Central Institute for Electronics, Research Center Jülich, Jülich, Germany Email: l.chen@fz-juelich.de
Network connectivity via inference over curvature-regularizing line graphs
 Network connectivity via inference over curvature-regularizing line graphs Asian Conference on Computer Vision Maxwell D. Collins 1,2, Vikas Singh 2,1, Andrew L. Alexander 3 1 Department of Computer Sciences
Network connectivity via inference over curvature-regularizing line graphs Asian Conference on Computer Vision Maxwell D. Collins 1,2, Vikas Singh 2,1, Andrew L. Alexander 3 1 Department of Computer Sciences
Streamline Flows for White Matter Fibre Pathway Segmentation in Diffusion MRI
 Streamline Flows for White Matter Fibre Pathway Segmentation in Diffusion MRI Peter Savadjiev 1, Jennifer S.W. Campbell 1,2,G.BrucePike 2, and Kaleem Siddiqi 1 McGill University, Montréal, QC, Canada 1
Streamline Flows for White Matter Fibre Pathway Segmentation in Diffusion MRI Peter Savadjiev 1, Jennifer S.W. Campbell 1,2,G.BrucePike 2, and Kaleem Siddiqi 1 McGill University, Montréal, QC, Canada 1
NeuroQLab A Software Assistant for Neurosurgical Planning and Quantitative Image Analysis
 NeuroQLab A Software Assistant for Neurosurgical Planning and Quantitative Image Analysis Florian Weiler 1, Jan Rexilius 2, Jan Klein 1, Horst K. Hahn 1 1 Fraunhofer MEVIS, Universitätsallee 29, 28359
NeuroQLab A Software Assistant for Neurosurgical Planning and Quantitative Image Analysis Florian Weiler 1, Jan Rexilius 2, Jan Klein 1, Horst K. Hahn 1 1 Fraunhofer MEVIS, Universitätsallee 29, 28359
Multi-Diffusion-Tensor Fitting via Spherical Deconvolution: A Unifying Framework
 Multi-Diffusion-Tensor Fitting via Spherical Deconvolution: A Unifying Framework Thomas Schultz 1, Carl-Fredrik Westin 2, and Gordon Kindlmann 1 1 Computer Science Department and Computation Institute,
Multi-Diffusion-Tensor Fitting via Spherical Deconvolution: A Unifying Framework Thomas Schultz 1, Carl-Fredrik Westin 2, and Gordon Kindlmann 1 1 Computer Science Department and Computation Institute,
Spatial Normalization of Diffusion Tensor Fields
 Magnetic Resonance in Medicine 50:175 182 (2003) Spatial Normalization of Diffusion Tensor Fields Dongrong Xu, 1 * Susumu Mori, 2 Dinggang Shen, 1 Peter C.M. van Zijl, 2 and Christos Davatzikos 1 A method
Magnetic Resonance in Medicine 50:175 182 (2003) Spatial Normalization of Diffusion Tensor Fields Dongrong Xu, 1 * Susumu Mori, 2 Dinggang Shen, 1 Peter C.M. van Zijl, 2 and Christos Davatzikos 1 A method
Department of Biomedical Engineering, University of Florida, Gainesville, FL 32611, USA
 Int. J. Bioinformatics and Research Applications, Vol. x, No. x, 2012 1 Perpendicular Fiber Tracking for Neural Fiber Bundle Analysis using Diffusion MRI S. Ray Department of Biomedical Engineering, University
Int. J. Bioinformatics and Research Applications, Vol. x, No. x, 2012 1 Perpendicular Fiber Tracking for Neural Fiber Bundle Analysis using Diffusion MRI S. Ray Department of Biomedical Engineering, University
Where are we now? Structural MRI processing and analysis
 Where are we now? Structural MRI processing and analysis Pierre-Louis Bazin bazin@cbs.mpg.de Leipzig, Germany Structural MRI processing: why bother? Just use the standards? SPM FreeSurfer FSL However:
Where are we now? Structural MRI processing and analysis Pierre-Louis Bazin bazin@cbs.mpg.de Leipzig, Germany Structural MRI processing: why bother? Just use the standards? SPM FreeSurfer FSL However:
Detecting Changes In Non-Isotropic Images
 Detecting Changes In Non-Isotropic Images K.J. Worsley 1, M. Andermann 1, T. Koulis 1, D. MacDonald, 2 and A.C. Evans 2 August 4, 1999 1 Department of Mathematics and Statistics, 2 Montreal Neurological
Detecting Changes In Non-Isotropic Images K.J. Worsley 1, M. Andermann 1, T. Koulis 1, D. MacDonald, 2 and A.C. Evans 2 August 4, 1999 1 Department of Mathematics and Statistics, 2 Montreal Neurological
High-Order Diffusion Tensor Connectivity Mapping on the GPU
 High-Order Diffusion Tensor Connectivity Mapping on the GPU Tim McGraw and Donald Herring Purdue University Abstract. We present an efficient approach to computing white matter fiber connectivity on the
High-Order Diffusion Tensor Connectivity Mapping on the GPU Tim McGraw and Donald Herring Purdue University Abstract. We present an efficient approach to computing white matter fiber connectivity on the
Building an Average Population HARDI Atlas
 Building an Average Population HARDI Atlas Sylvain Bouix 1, Yogesh Rathi 1, and Mert Sabuncu 2 1 Psychiatry Neuroimaging Laboratory, Brigham and Women s Hospital, Harvard Medical School, Boston, MA, USA.
Building an Average Population HARDI Atlas Sylvain Bouix 1, Yogesh Rathi 1, and Mert Sabuncu 2 1 Psychiatry Neuroimaging Laboratory, Brigham and Women s Hospital, Harvard Medical School, Boston, MA, USA.
1192 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 29, NO. 5, MAY 2010
 1192 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 29, NO. 5, MAY 2010 F-TIMER: Fast Tensor Image Morphing for Elastic Registration Pew-Thian Yap, Guorong Wu, Hongtu Zhu, Weili Lin, and Dinggang Shen*, Senior
1192 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 29, NO. 5, MAY 2010 F-TIMER: Fast Tensor Image Morphing for Elastic Registration Pew-Thian Yap, Guorong Wu, Hongtu Zhu, Weili Lin, and Dinggang Shen*, Senior
MARS: Multiple Atlases Robust Segmentation
 Software Release (1.0.1) Last updated: April 30, 2014. MARS: Multiple Atlases Robust Segmentation Guorong Wu, Minjeong Kim, Gerard Sanroma, and Dinggang Shen {grwu, mjkim, gerard_sanroma, dgshen}@med.unc.edu
Software Release (1.0.1) Last updated: April 30, 2014. MARS: Multiple Atlases Robust Segmentation Guorong Wu, Minjeong Kim, Gerard Sanroma, and Dinggang Shen {grwu, mjkim, gerard_sanroma, dgshen}@med.unc.edu
Using a Statistical Shape Model to Extract Sulcal Curves on the Outer Cortex of the Human Brain
 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 21, NO. 5, MAY 2002 513 Using a Statistical Shape Model to Extract Sulcal Curves on the Outer Cortex of the Human Brain Xiaodong Tao, Jerry L. Prince, and Christos
IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 21, NO. 5, MAY 2002 513 Using a Statistical Shape Model to Extract Sulcal Curves on the Outer Cortex of the Human Brain Xiaodong Tao, Jerry L. Prince, and Christos
Structural connectivity of the brain measured by diffusion tensor imaging [DTI]
![Structural connectivity of the brain measured by diffusion tensor imaging [DTI] Structural connectivity of the brain measured by diffusion tensor imaging [DTI]](/thumbs/74/71065400.jpg) Structural connectivity of the brain measured by diffusion tensor imaging [DTI] Lars T. Westlye CSHC / Centre for Advanced Study PSY4320 06.05.2012 l.t.westlye@psykologi.uio.no ~ 40 % of the brain is white
Structural connectivity of the brain measured by diffusion tensor imaging [DTI] Lars T. Westlye CSHC / Centre for Advanced Study PSY4320 06.05.2012 l.t.westlye@psykologi.uio.no ~ 40 % of the brain is white
White Pixel Artifact. Caused by a noise spike during acquisition Spike in K-space <--> sinusoid in image space
 White Pixel Artifact Caused by a noise spike during acquisition Spike in K-space sinusoid in image space Susceptibility Artifacts Off-resonance artifacts caused by adjacent regions with different
White Pixel Artifact Caused by a noise spike during acquisition Spike in K-space sinusoid in image space Susceptibility Artifacts Off-resonance artifacts caused by adjacent regions with different
H. Mirzaalian, L. Ning, P. Savadjiev, O. Pasternak, S. Bouix, O. Michailovich, M. Kubicki, C-F Westin, M.E.Shenton, Y. Rathi
 Harmonizing diffusion MRI data from multiple scanners H. Mirzaalian, L. Ning, P. Savadjiev, O. Pasternak, S. Bouix, O. Michailovich, M. Kubicki, C-F Westin, M.E.Shenton, Y. Rathi Synopsis Diffusion MRI
Harmonizing diffusion MRI data from multiple scanners H. Mirzaalian, L. Ning, P. Savadjiev, O. Pasternak, S. Bouix, O. Michailovich, M. Kubicki, C-F Westin, M.E.Shenton, Y. Rathi Synopsis Diffusion MRI
Technische Universiteit Eindhoven Department of Mathematics and Computer Science. Master s Thesis HIERARCHICAL VISUALIZATION USING FIBER CLUSTERING
 Technische Universiteit Eindhoven Department of Mathematics and Computer Science Master s Thesis HIERARCHICAL VISUALIZATION USING FIBER CLUSTERING by Ing. B. Moberts Supervisors: Dr. A. Vilanova Prof.dr.ir.
Technische Universiteit Eindhoven Department of Mathematics and Computer Science Master s Thesis HIERARCHICAL VISUALIZATION USING FIBER CLUSTERING by Ing. B. Moberts Supervisors: Dr. A. Vilanova Prof.dr.ir.
Algebraic Methods for Direct and Feature Based Registration of Diffusion Tensor Images
 Algebraic Methods for Direct and Feature Based Registration of Diffusion Tensor Images Alvina Goh and René Vidal Center for Imaging Science, Department of BME, Johns Hopkins University 308B Clark Hall,
Algebraic Methods for Direct and Feature Based Registration of Diffusion Tensor Images Alvina Goh and René Vidal Center for Imaging Science, Department of BME, Johns Hopkins University 308B Clark Hall,
