Quantitative Biology II!
|
|
|
- Darrell Berry
- 5 years ago
- Views:
Transcription
1 Quantitative Biology II! Lecture 3: Markov Chain Monte Carlo! March 9, 2015!
2 2! Plan for Today!! Introduction to Sampling!! Introduction to MCMC!! Metropolis Algorithm!! Metropolis-Hastings Algorithm!! Gibbs Sampling!! Monitoring Convergence!! Examples!
3 3! Sampling Motivation!! So far we have focused on models for which exact inference is possible!! In general, this will not be true e.g., models with non-gaussian continuous distributions or large clique sizes!! There are two main options available in such cases:!! Approximate inference!! Sampling methods!! Today: sampling (Monte Carlo) methods, Mickey: approximate inference!
4 4! Sampling!! Suppose exact inference is impossible for a pdf p (x), but samples x (1), x (2),..., x (N) can be drawn!! Many properties of interest can be estimated if N is sufficiently large, e.g.,!! Note that the samples need not be independent, but if not then N must be larger!
5 5! Toy Example! Radius: r = 1! Side: s = 2! Number of darts: N! Number in circle: k! E[k/N] = π/4! π 4k/N!
6 6! Monte Carlo History!! We have estimated the area of the circle by Monte Carlo integration!! Monte Carlo methods were pioneered by mathematicians and statistical physicists during and after the Manhattan Project (esp. Stan Ulam, Nicholas Metropolis, John von Neumann)!! Interest in sampling theory dates back to the early days of probability theory, but putting it to work required electronic computers!
7 7! Simple Sampling!! Uniform distribution: generate a pseudorandom number between 0 and large M (e.g., RAND_MAX), then divide by M!! More complex distributions:!! Inversion method!! Rejection sampling!! Importance sampling!! Sampling-importance-resampling!
8 8! Inversion Method! Example:! If h(y) is a CDF, and y is a random variate from the desired distribution, then x = h(y) is uniformly distributed: x ~ U(0, 1)!! Thus, a uniform random variate xʹ can be converted to a random variate yʹ by inverting h: yʹ = h 1 (xʹ )
9 9! Rejection Sampling! Algorithm:!! Sample x0 from q(x)!! Sample u ~ U(0, 1)!! Accept x0 if:!! Otherwise reject x0 and continue sampling!!
10 Adaptive Rejection 10! Sampling!! Suppose x univariate, p (x) concave!! Given a set of points, {x1,, xn}, define piecewise linear envelope function for ln p(x)!! Drawing from the envelope function is straightforward piecewise exponential form!! Initialize with a grid of points. As new points are drawn, they can be added to the set, improving the envelope function!
11 11! Importance Sampling!! Suppose we seek f = E[f(X)]! We canʼt sample from p(x), but we can evaluate the density!! Suppose, in addition, we can sample from a simpler q(x)! Importance sampling follows from:! More generally, for unnormalized distributions,! where!
12 Sampling-Importance- 12! Resampling!! The same idea can be incorporated into a sampling scheme!! Start by drawing N points from q(x) and computing weights, similar to those above!! Now draw M points with probabilities given by these weights!! As N approaches infinity, the resampling distribution approaches p(x)
13 MCMC! 13!! The basic idea of MCMC is to sample variables (or subsets of variables) conditional on previous samples!! Typically, these conditional distributions are easier to work with than the full joint distribution!! Successive samples will be correlated. The samples form a Markov chain whose state space equals the support of the joint distribution!! MCMC is designed such that the long-term average (stationary) distribution of the chain equals the desired distribution!! Basic approach: collect many samples, try to show convergence!
14 Notation! 14!! As with EM, assume some variables are observed and denote them x!! Assume other variables are latent and denote them z! The observed variables will be held fixed throughout the procedure, while the latent variables will be sampled!! The state space of the Markov chain therefore equals the space of possible values of z, and its stationary distribution is p (z x)! Key problem: what should p(z (t+1) z (t), x) be?!
15 15! Illustration of MCMC! The transition probabilities must be designed so that the stationary distribution is p(z x).! After a suitable burn-in period, samples drawn from each p(z t z t 1, x) will be representative of p (z x).! However, they will not be independent samples.!
16 16! Bivariate Normal Example!! Suppose x is a set of n points on the two-dimensional plane!! These points are assumed to be drawn independently from a bivariate normal distribution with unknown mean μ! The goal is to infer the distribution of μ given x (the posterior)!! A (diffuse) normal prior is assumed:!
17 17! Bivariate Normal, cont.!! In this case, we can derive an exact closed form solution for the posterior distribution, but suppose we wish to use MCMC instead!! Here z is the mean μ, and the state space of the Markov chain is points on the twodimensional plane. The observed variable x is fixed at the given set of points!! Transitions can be thought of as moves from one point on the plane to another, and a sequence of samples will trace a 2d trajectory!! Over the long term, points from this trajectory will represent the posterior p(μ x)!
18 Illustration! 18!
19 19! How Does MCMC Work?! How can we set the transition probabilities such that the equilibrium distribution is the posterior, without knowing what the posterior is?!
20 20! Marginals for a Markov Chain!! Let z = (z (1), z (2),..., z (N) ) be a (first-order) Markov chain, with z (t) S for t {1,..., N}. For simplicity, assume S is a finite set.!! Let π (t) be the marginal distribution of z (t) :!! Thus,! or, in matrix notation,!! Given an initial distribution π (0), π (t) is given by:!
21 21! Stationary Distribution!! We say the chain is invariant, or stationary, when π (t) = π (t+1) = π *, i.e.,!! A Markov chain may have more than one stationary distribution. For example, every distribution is invariant when A = I!!! If the Markov chain is ergodic, however, then it will always converge to a single stationary distribution:!! This distribution is given by the eigenvector corresponding to the largest eigenvalue of A
22 Ergodicity! 22!! To be ergodic, the chain must be:!! Irreducible must be positive probability of reaching any state from any other!! Aperiodic must not cycle through states deterministically!! Non-transient must always be able to return to a state after visiting it!! In designing transition distributions for MCMC, irreducibility is typically the critical property!! Ergodicity is automatic if the transitions to all states have nonzero probability!
23 23! Reversibility!! A Markov chain is said to be reversible if:!! Reversibility with respect to a distribution π * is sufficient to make π * invariant:!! Thus, if a Markov chain is constructed to be ergodic and reversible with respect to some π *, then it will converge to π *
24 Metropolis Algorithm! 24!! Suppose transitions are proposed from a symmetric distribution q(z (t) z (t 1) ) i.e., such that q(z (t) =a z (t 1) =b) = q(z (t) =b z (t 1) =a)!! Now suppose proposals are accepted with probability (implicitly conditioning on x):!! Thus:!
25 25! Implications!! This simple procedure guarantees reversibility of the Markov chain with respect to the posterior p (z) simply by evaluating ratios of densities!! Furthermore, ratios of posterior densities can be computed as ratios of complete data densities:!! As discussed, reversibility with respect to p(z) implies that p(z) is a stationary distribution of the Markov chain!! If the Markov chain is also ergodic, then p(z) is a unique stationary distribution of the Markov chain!
26 Logistics! 26!! The proposal distribution has to be designed to guarantee ergodicity!! The chain will not reach stationarity immediately; a burn-in period is required. Suppose it consists of B steps!! Suppose S samples are collected following the B burn-in steps!! A sample can be collected on each iteration, but successive samples may be highly correlated, resulting in an effective sample size << S. It may be more efficient to retain every kth sample!
27 27! Metropolis Algorithm! initialize with z (0) s.t. p(z (0) x) > 0! t 1! repeat! sample z (t) from q(z (t) z (t 1), x)! compute:! draw u from U(0,1)! if (u > a(z (t 1), z (t) )) z (t) z (t 1) /* reject proposal */! if (t > B and t mod k = 0) retain sample z (t)! t t + 1! until enough samples (t = B + Sk)!
28 28! Recall: Bivariate Normal!! Suppose x is a set of n points on the two-dimensional plane!! These points are assumed to be drawn independently from a bivariate normal distribution with unknown mean μ! The goal is to infer the distribution of μ given x (the posterior) [assume fixed var. I]!! A (diffuse) normal prior is assumed:!
29 29! Bivariate Normal, cont.!! As a symmetric proposal distribution for moves on the 2d plane, assume a simple Gaussian random walk:!! The acceptance probabilities will be:!! The variance σ 2 determines the average step size, and can be used as a tuning parameter!
30 30! Illustration! Small σ 2 : small steps, high acceptance rate! Large σ 2 : big steps, low acceptance rate! Minimizing the correlation between successive samples, hence minimizing the number of samples needed, requires a tradeoff!
31 Remarks! 31!! Notice that probabilities (densities) are always computed from fully observed variables; no integration is necessary!! Furthermore, only ratios of densities are needed. As a result, unnormalized distributions can be used.!! The key design parameter is the proposal distribution. It must ensure that the chain is ergodic, keep the acceptance rate high, and facilitate mixing (low correlation of successive samples)!! There is tradeoff between bold and cautious proposals in optimizing mixing!
32 Asymmetric Proposals! 32!! The requirement of a symmetric proposal distr. is easily circumvented!! An additional term in the acceptance probability corrects for any asymmetry:!! Now:!
33 33! Metropolis-Hastings! initialize with z (0) s.t. p(z (0) x) > 0! t 1! repeat! sample z (t) from q(z (t) z (t 1), x)! compute:! draw u from U(0,1)! if (u > a(z (t 1), z (t) )) z (t) z (t 1) /* reject proposal */! if (t > B and t mod k = 0) retain sample z (t)! t t + 1! until enough samples (t = B + Sk)!
34 More Remarks! 34!! MCMC is enormously versatile: a sampler can easily be constructed for almost any model!! It is also flexible: not only can the posterior be approximated, but so can any function of the posterior!! The critical issue is convergence. How long does the chain have to run? How can we be sure it has converged? Even if it has, have enough samples been drawn?!! Bottom line: hard problems are still hard, but MCMC with clever proposal distributions can help!
35 35! Proposing Subsets!! If z has high dimension, it may be hard to find a proposal distribution that will result in a sufficiently high acceptance rate!! A possible solution is to partition the variables into W subsets, and to sample individual subsets conditional on the others!! On each step t consider a subset zi (randomly or by round robin) and propose a new value from:!
36 Illustration! 36!
37 37! Gibbs Sampling!! Gibbs sampling is the special case in which the proposal distribution is defined by the exact conditional distribution:!! This proposal distribution guarantees a perfect acceptance rate!!
38 38! Simple Example!! Suppose three latent variables, z1, z2, z3!! Gibbs sampling will sample each in turn conditional on the other two (and on x), using the exact conditionals:!! z1 (t) ~ p(z1 z2 (t 1), z3 (t 1), x)! z2 (t+1) ~ p(z2 z1 (t), z3 (t), x)! z3 (t+2) ~ p(z3 z1 (t+1), z2 (t+1), x)! It can either cycle through them in order, or visit them randomly (provided each is visited with sufficiently high probability)!
39 39! Gibbs Sampling Algorithm! initialize with z (0) s.t. p(z (0) x) > 0! t 1! repeat! for i 1 to W! sample zi (t) from p(zi (t) z i (t 1), x) z-i (t) z-i (t 1)! if (t > B and t mod k = 0) retain sample z (t)! t t + 1! end for! until enough samples (t = B + Sk)!
40 40! Another Way to See It!! It can be shown more directly that Gibbs sampling must produce the desired stationary distribution!! Suppose the Markov chain has reached a point at which z (t) ~ p(z x). Note that p(z x) = p(z i x) p(zi z i, x)!! Each Gibbs step holds z i (t) fixed and draws zi (t+1) from the exact conditional; thus z (t+1) ~ p(z x)!! It is also easy to show directly that the chain is reversible wrt p(z x)
41 41! Ergodicity!! For the posterior to be a unique equilibrium distribution, the chain must also be ergodic (as usual)!! If all conditional distributions are nonzero everywhere, then ergodicity must hold!! Otherwise, it must be proven explicitly!
42 Bivariate Normal Gibbs! 42!
43 43! Gaussian Mixtures!! Gibbs sampling allows the Gaussian mixtures problem to be addressed in a fully Bayesian way:!! Assign cluster means a (Gaussian) prior!! Mean sampling: For each cluster, sample new mean based on prior and currently assigned data points!! Assignment sampling: Sample new cluster assignment for each data point given current cluster means!! Upon termination, summarize groupings from samples of joint posterior!
44 44!
45 45! Comparison with EM!! Both EM and Gibbs alternate between setting variables and setting parameters!! EM avoids hard assignments, instead using expectations!! Gibbs makes hard assignments but does so stochastically!! EM maximizes parameters based on expectations of rvʼs; Gibbs does not distinguish between parameters and rvʼs!! Gibbs can be seen as a stochastic hill climbing algorithm. It may do better than EM at avoiding local maxima!
46 46! Assessing Convergence!! Simplest approach: plot complete log likelihood, visually assess stationarity!! Using this method can usually make a good guess at appropriate burn-in length B!! Can apply to logl or estimated scalars!! Good idea to start multiple chains and see whether they end up behaving the same!! More rigorously, can run multiple chains and compare within chain and between chain variances!
47 Visual Inspection! 47!
48 Another Example! 48!
49 49! Monitoring Scalar Estimands!! Run J parallel chains, initializing from an overdispersed distribution. Collect n samples from each.!! Compute within-chain (W) and betweenchain (B) variances for scalar samples!! Monitor convergence via scale reduction,! Gelman et al. Bayesian Data Analysis, 1995!
50 50! Sampling Motifs! initialize! extract counts,! sample from Dirichlet! compute posteriors,! sample positions!
51 51! Sampling Alignments! V! L! S! P! A! D! K! H! L! A! E! S! K!
52 52! Sampling Alignments! VLSPAD-K! HL--AESK! H! L! V! L! S! P! A! D! K! A! E! S! K!
53 53! Sampling Alignments! VLSPAD-K! HL--AESK! VL--SPADK! HLAES---K! H! L! A! E! V! L! S! P! A! D! K! S! K!
54 54! Sampling Alignments! VLSPAD-K! HL--AESK! VL--SPADK! HLAES---K! -VLSPADK! H-LAES-K! H! L! A! E! S! K! V! L! S! P! A! D! K!
55 55! Measuring Confidence! Lunter et al., Genome Res, 2008!
56 56! Thatʼs All!! Bishop has good introduction to sampling and MCMC!! Sampling alignments is covered in Durbin et al.!! Gelman et al. good reference on applied Bayesian analysis!! Thanks for listening!!
Overview. Monte Carlo Methods. Statistics & Bayesian Inference Lecture 3. Situation At End Of Last Week
 Statistics & Bayesian Inference Lecture 3 Joe Zuntz Overview Overview & Motivation Metropolis Hastings Monte Carlo Methods Importance sampling Direct sampling Gibbs sampling Monte-Carlo Markov Chains Emcee
Statistics & Bayesian Inference Lecture 3 Joe Zuntz Overview Overview & Motivation Metropolis Hastings Monte Carlo Methods Importance sampling Direct sampling Gibbs sampling Monte-Carlo Markov Chains Emcee
Computer vision: models, learning and inference. Chapter 10 Graphical Models
 Computer vision: models, learning and inference Chapter 10 Graphical Models Independence Two variables x 1 and x 2 are independent if their joint probability distribution factorizes as Pr(x 1, x 2 )=Pr(x
Computer vision: models, learning and inference Chapter 10 Graphical Models Independence Two variables x 1 and x 2 are independent if their joint probability distribution factorizes as Pr(x 1, x 2 )=Pr(x
MCMC Methods for data modeling
 MCMC Methods for data modeling Kenneth Scerri Department of Automatic Control and Systems Engineering Introduction 1. Symposium on Data Modelling 2. Outline: a. Definition and uses of MCMC b. MCMC algorithms
MCMC Methods for data modeling Kenneth Scerri Department of Automatic Control and Systems Engineering Introduction 1. Symposium on Data Modelling 2. Outline: a. Definition and uses of MCMC b. MCMC algorithms
Issues in MCMC use for Bayesian model fitting. Practical Considerations for WinBUGS Users
 Practical Considerations for WinBUGS Users Kate Cowles, Ph.D. Department of Statistics and Actuarial Science University of Iowa 22S:138 Lecture 12 Oct. 3, 2003 Issues in MCMC use for Bayesian model fitting
Practical Considerations for WinBUGS Users Kate Cowles, Ph.D. Department of Statistics and Actuarial Science University of Iowa 22S:138 Lecture 12 Oct. 3, 2003 Issues in MCMC use for Bayesian model fitting
Approximate Bayesian Computation. Alireza Shafaei - April 2016
 Approximate Bayesian Computation Alireza Shafaei - April 2016 The Problem Given a dataset, we are interested in. The Problem Given a dataset, we are interested in. The Problem Given a dataset, we are interested
Approximate Bayesian Computation Alireza Shafaei - April 2016 The Problem Given a dataset, we are interested in. The Problem Given a dataset, we are interested in. The Problem Given a dataset, we are interested
1 Methods for Posterior Simulation
 1 Methods for Posterior Simulation Let p(θ y) be the posterior. simulation. Koop presents four methods for (posterior) 1. Monte Carlo integration: draw from p(θ y). 2. Gibbs sampler: sequentially drawing
1 Methods for Posterior Simulation Let p(θ y) be the posterior. simulation. Koop presents four methods for (posterior) 1. Monte Carlo integration: draw from p(θ y). 2. Gibbs sampler: sequentially drawing
MCMC Diagnostics. Yingbo Li MATH Clemson University. Yingbo Li (Clemson) MCMC Diagnostics MATH / 24
 MCMC Diagnostics Yingbo Li Clemson University MATH 9810 Yingbo Li (Clemson) MCMC Diagnostics MATH 9810 1 / 24 Convergence to Posterior Distribution Theory proves that if a Gibbs sampler iterates enough,
MCMC Diagnostics Yingbo Li Clemson University MATH 9810 Yingbo Li (Clemson) MCMC Diagnostics MATH 9810 1 / 24 Convergence to Posterior Distribution Theory proves that if a Gibbs sampler iterates enough,
Markov Chain Monte Carlo (part 1)
 Markov Chain Monte Carlo (part 1) Edps 590BAY Carolyn J. Anderson Department of Educational Psychology c Board of Trustees, University of Illinois Spring 2018 Depending on the book that you select for
Markov Chain Monte Carlo (part 1) Edps 590BAY Carolyn J. Anderson Department of Educational Psychology c Board of Trustees, University of Illinois Spring 2018 Depending on the book that you select for
Markov chain Monte Carlo methods
 Markov chain Monte Carlo methods (supplementary material) see also the applet http://www.lbreyer.com/classic.html February 9 6 Independent Hastings Metropolis Sampler Outline Independent Hastings Metropolis
Markov chain Monte Carlo methods (supplementary material) see also the applet http://www.lbreyer.com/classic.html February 9 6 Independent Hastings Metropolis Sampler Outline Independent Hastings Metropolis
Clustering Relational Data using the Infinite Relational Model
 Clustering Relational Data using the Infinite Relational Model Ana Daglis Supervised by: Matthew Ludkin September 4, 2015 Ana Daglis Clustering Data using the Infinite Relational Model September 4, 2015
Clustering Relational Data using the Infinite Relational Model Ana Daglis Supervised by: Matthew Ludkin September 4, 2015 Ana Daglis Clustering Data using the Infinite Relational Model September 4, 2015
Chapter 1. Introduction
 Chapter 1 Introduction A Monte Carlo method is a compuational method that uses random numbers to compute (estimate) some quantity of interest. Very often the quantity we want to compute is the mean of
Chapter 1 Introduction A Monte Carlo method is a compuational method that uses random numbers to compute (estimate) some quantity of interest. Very often the quantity we want to compute is the mean of
Bayesian Estimation for Skew Normal Distributions Using Data Augmentation
 The Korean Communications in Statistics Vol. 12 No. 2, 2005 pp. 323-333 Bayesian Estimation for Skew Normal Distributions Using Data Augmentation Hea-Jung Kim 1) Abstract In this paper, we develop a MCMC
The Korean Communications in Statistics Vol. 12 No. 2, 2005 pp. 323-333 Bayesian Estimation for Skew Normal Distributions Using Data Augmentation Hea-Jung Kim 1) Abstract In this paper, we develop a MCMC
Probabilistic Graphical Models
 Probabilistic Graphical Models Lecture 17 EM CS/CNS/EE 155 Andreas Krause Announcements Project poster session on Thursday Dec 3, 4-6pm in Annenberg 2 nd floor atrium! Easels, poster boards and cookies
Probabilistic Graphical Models Lecture 17 EM CS/CNS/EE 155 Andreas Krause Announcements Project poster session on Thursday Dec 3, 4-6pm in Annenberg 2 nd floor atrium! Easels, poster boards and cookies
Short-Cut MCMC: An Alternative to Adaptation
 Short-Cut MCMC: An Alternative to Adaptation Radford M. Neal Dept. of Statistics and Dept. of Computer Science University of Toronto http://www.cs.utoronto.ca/ radford/ Third Workshop on Monte Carlo Methods,
Short-Cut MCMC: An Alternative to Adaptation Radford M. Neal Dept. of Statistics and Dept. of Computer Science University of Toronto http://www.cs.utoronto.ca/ radford/ Third Workshop on Monte Carlo Methods,
CS281 Section 9: Graph Models and Practical MCMC
 CS281 Section 9: Graph Models and Practical MCMC Scott Linderman November 11, 213 Now that we have a few MCMC inference algorithms in our toolbox, let s try them out on some random graph models. Graphs
CS281 Section 9: Graph Models and Practical MCMC Scott Linderman November 11, 213 Now that we have a few MCMC inference algorithms in our toolbox, let s try them out on some random graph models. Graphs
Sampling informative/complex a priori probability distributions using Gibbs sampling assisted by sequential simulation
 Sampling informative/complex a priori probability distributions using Gibbs sampling assisted by sequential simulation Thomas Mejer Hansen, Klaus Mosegaard, and Knud Skou Cordua 1 1 Center for Energy Resources
Sampling informative/complex a priori probability distributions using Gibbs sampling assisted by sequential simulation Thomas Mejer Hansen, Klaus Mosegaard, and Knud Skou Cordua 1 1 Center for Energy Resources
10-701/15-781, Fall 2006, Final
 -7/-78, Fall 6, Final Dec, :pm-8:pm There are 9 questions in this exam ( pages including this cover sheet). If you need more room to work out your answer to a question, use the back of the page and clearly
-7/-78, Fall 6, Final Dec, :pm-8:pm There are 9 questions in this exam ( pages including this cover sheet). If you need more room to work out your answer to a question, use the back of the page and clearly
An Introduction to Markov Chain Monte Carlo
 An Introduction to Markov Chain Monte Carlo Markov Chain Monte Carlo (MCMC) refers to a suite of processes for simulating a posterior distribution based on a random (ie. monte carlo) process. In other
An Introduction to Markov Chain Monte Carlo Markov Chain Monte Carlo (MCMC) refers to a suite of processes for simulating a posterior distribution based on a random (ie. monte carlo) process. In other
ADAPTIVE METROPOLIS-HASTINGS SAMPLING, OR MONTE CARLO KERNEL ESTIMATION
 ADAPTIVE METROPOLIS-HASTINGS SAMPLING, OR MONTE CARLO KERNEL ESTIMATION CHRISTOPHER A. SIMS Abstract. A new algorithm for sampling from an arbitrary pdf. 1. Introduction Consider the standard problem of
ADAPTIVE METROPOLIS-HASTINGS SAMPLING, OR MONTE CARLO KERNEL ESTIMATION CHRISTOPHER A. SIMS Abstract. A new algorithm for sampling from an arbitrary pdf. 1. Introduction Consider the standard problem of
Convexization in Markov Chain Monte Carlo
 in Markov Chain Monte Carlo 1 IBM T. J. Watson Yorktown Heights, NY 2 Department of Aerospace Engineering Technion, Israel August 23, 2011 Problem Statement MCMC processes in general are governed by non
in Markov Chain Monte Carlo 1 IBM T. J. Watson Yorktown Heights, NY 2 Department of Aerospace Engineering Technion, Israel August 23, 2011 Problem Statement MCMC processes in general are governed by non
Clustering web search results
 Clustering K-means Machine Learning CSE546 Emily Fox University of Washington November 4, 2013 1 Clustering images Set of Images [Goldberger et al.] 2 1 Clustering web search results 3 Some Data 4 2 K-means
Clustering K-means Machine Learning CSE546 Emily Fox University of Washington November 4, 2013 1 Clustering images Set of Images [Goldberger et al.] 2 1 Clustering web search results 3 Some Data 4 2 K-means
Clustering K-means. Machine Learning CSEP546 Carlos Guestrin University of Washington February 18, Carlos Guestrin
 Clustering K-means Machine Learning CSEP546 Carlos Guestrin University of Washington February 18, 2014 Carlos Guestrin 2005-2014 1 Clustering images Set of Images [Goldberger et al.] Carlos Guestrin 2005-2014
Clustering K-means Machine Learning CSEP546 Carlos Guestrin University of Washington February 18, 2014 Carlos Guestrin 2005-2014 1 Clustering images Set of Images [Goldberger et al.] Carlos Guestrin 2005-2014
GAMES Webinar: Rendering Tutorial 2. Monte Carlo Methods. Shuang Zhao
 GAMES Webinar: Rendering Tutorial 2 Monte Carlo Methods Shuang Zhao Assistant Professor Computer Science Department University of California, Irvine GAMES Webinar Shuang Zhao 1 Outline 1. Monte Carlo integration
GAMES Webinar: Rendering Tutorial 2 Monte Carlo Methods Shuang Zhao Assistant Professor Computer Science Department University of California, Irvine GAMES Webinar Shuang Zhao 1 Outline 1. Monte Carlo integration
MSA101/MVE Lecture 5
 MSA101/MVE187 2017 Lecture 5 Petter Mostad Chalmers University September 12, 2017 1 / 15 Importance sampling MC integration computes h(x)f (x) dx where f (x) is a probability density function, by simulating
MSA101/MVE187 2017 Lecture 5 Petter Mostad Chalmers University September 12, 2017 1 / 15 Importance sampling MC integration computes h(x)f (x) dx where f (x) is a probability density function, by simulating
ECE521: Week 11, Lecture March 2017: HMM learning/inference. With thanks to Russ Salakhutdinov
 ECE521: Week 11, Lecture 20 27 March 2017: HMM learning/inference With thanks to Russ Salakhutdinov Examples of other perspectives Murphy 17.4 End of Russell & Norvig 15.2 (Artificial Intelligence: A Modern
ECE521: Week 11, Lecture 20 27 March 2017: HMM learning/inference With thanks to Russ Salakhutdinov Examples of other perspectives Murphy 17.4 End of Russell & Norvig 15.2 (Artificial Intelligence: A Modern
Monte Carlo for Spatial Models
 Monte Carlo for Spatial Models Murali Haran Department of Statistics Penn State University Penn State Computational Science Lectures April 2007 Spatial Models Lots of scientific questions involve analyzing
Monte Carlo for Spatial Models Murali Haran Department of Statistics Penn State University Penn State Computational Science Lectures April 2007 Spatial Models Lots of scientific questions involve analyzing
Probabilistic Graphical Models
 Overview of Part Two Probabilistic Graphical Models Part Two: Inference and Learning Christopher M. Bishop Exact inference and the junction tree MCMC Variational methods and EM Example General variational
Overview of Part Two Probabilistic Graphical Models Part Two: Inference and Learning Christopher M. Bishop Exact inference and the junction tree MCMC Variational methods and EM Example General variational
Stochastic Simulation: Algorithms and Analysis
 Soren Asmussen Peter W. Glynn Stochastic Simulation: Algorithms and Analysis et Springer Contents Preface Notation v xii I What This Book Is About 1 1 An Illustrative Example: The Single-Server Queue 1
Soren Asmussen Peter W. Glynn Stochastic Simulation: Algorithms and Analysis et Springer Contents Preface Notation v xii I What This Book Is About 1 1 An Illustrative Example: The Single-Server Queue 1
10.4 Linear interpolation method Newton s method
 10.4 Linear interpolation method The next best thing one can do is the linear interpolation method, also known as the double false position method. This method works similarly to the bisection method by
10.4 Linear interpolation method The next best thing one can do is the linear interpolation method, also known as the double false position method. This method works similarly to the bisection method by
Monte Carlo Methods and Statistical Computing: My Personal E
 Monte Carlo Methods and Statistical Computing: My Personal Experience Department of Mathematics & Statistics Indian Institute of Technology Kanpur November 29, 2014 Outline Preface 1 Preface 2 3 4 5 6
Monte Carlo Methods and Statistical Computing: My Personal Experience Department of Mathematics & Statistics Indian Institute of Technology Kanpur November 29, 2014 Outline Preface 1 Preface 2 3 4 5 6
Level-set MCMC Curve Sampling and Geometric Conditional Simulation
 Level-set MCMC Curve Sampling and Geometric Conditional Simulation Ayres Fan John W. Fisher III Alan S. Willsky February 16, 2007 Outline 1. Overview 2. Curve evolution 3. Markov chain Monte Carlo 4. Curve
Level-set MCMC Curve Sampling and Geometric Conditional Simulation Ayres Fan John W. Fisher III Alan S. Willsky February 16, 2007 Outline 1. Overview 2. Curve evolution 3. Markov chain Monte Carlo 4. Curve
The Multi Stage Gibbs Sampling: Data Augmentation Dutch Example
 The Multi Stage Gibbs Sampling: Data Augmentation Dutch Example Rebecca C. Steorts Bayesian Methods and Modern Statistics: STA 360/601 Module 8 1 Example: Data augmentation / Auxiliary variables A commonly-used
The Multi Stage Gibbs Sampling: Data Augmentation Dutch Example Rebecca C. Steorts Bayesian Methods and Modern Statistics: STA 360/601 Module 8 1 Example: Data augmentation / Auxiliary variables A commonly-used
Inference and Representation
 Inference and Representation Rachel Hodos New York University Lecture 5, October 6, 2015 Rachel Hodos Lecture 5: Inference and Representation Today: Learning with hidden variables Outline: Unsupervised
Inference and Representation Rachel Hodos New York University Lecture 5, October 6, 2015 Rachel Hodos Lecture 5: Inference and Representation Today: Learning with hidden variables Outline: Unsupervised
Clustering K-means. Machine Learning CSEP546 Carlos Guestrin University of Washington February 18, Carlos Guestrin
 Clustering K-means Machine Learning CSEP546 Carlos Guestrin University of Washington February 18, 2014 Carlos Guestrin 2005-2014 1 Clustering images Set of Images [Goldberger et al.] Carlos Guestrin 2005-2014
Clustering K-means Machine Learning CSEP546 Carlos Guestrin University of Washington February 18, 2014 Carlos Guestrin 2005-2014 1 Clustering images Set of Images [Goldberger et al.] Carlos Guestrin 2005-2014
STA 4273H: Statistical Machine Learning
 STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 5 Inference
STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 5 Inference
A Short History of Markov Chain Monte Carlo
 A Short History of Markov Chain Monte Carlo Christian Robert and George Casella 2010 Introduction Lack of computing machinery, or background on Markov chains, or hesitation to trust in the practicality
A Short History of Markov Chain Monte Carlo Christian Robert and George Casella 2010 Introduction Lack of computing machinery, or background on Markov chains, or hesitation to trust in the practicality
An Efficient Model Selection for Gaussian Mixture Model in a Bayesian Framework
 IEEE SIGNAL PROCESSING LETTERS, VOL. XX, NO. XX, XXX 23 An Efficient Model Selection for Gaussian Mixture Model in a Bayesian Framework Ji Won Yoon arxiv:37.99v [cs.lg] 3 Jul 23 Abstract In order to cluster
IEEE SIGNAL PROCESSING LETTERS, VOL. XX, NO. XX, XXX 23 An Efficient Model Selection for Gaussian Mixture Model in a Bayesian Framework Ji Won Yoon arxiv:37.99v [cs.lg] 3 Jul 23 Abstract In order to cluster
Recap: The E-M algorithm. Biostatistics 615/815 Lecture 22: Gibbs Sampling. Recap - Local minimization methods
 Recap: The E-M algorithm Biostatistics 615/815 Lecture 22: Gibbs Sampling Expectation step (E-step) Given the current estimates of parameters λ (t), calculate the conditional distribution of latent variable
Recap: The E-M algorithm Biostatistics 615/815 Lecture 22: Gibbs Sampling Expectation step (E-step) Given the current estimates of parameters λ (t), calculate the conditional distribution of latent variable
Math 494: Mathematical Statistics
 Math 494: Mathematical Statistics Instructor: Jimin Ding jmding@wustl.edu Department of Mathematics Washington University in St. Louis Class materials are available on course website (www.math.wustl.edu/
Math 494: Mathematical Statistics Instructor: Jimin Ding jmding@wustl.edu Department of Mathematics Washington University in St. Louis Class materials are available on course website (www.math.wustl.edu/
Metropolis Light Transport
 Metropolis Light Transport CS295, Spring 2017 Shuang Zhao Computer Science Department University of California, Irvine CS295, Spring 2017 Shuang Zhao 1 Announcements Final presentation June 13 (Tuesday)
Metropolis Light Transport CS295, Spring 2017 Shuang Zhao Computer Science Department University of California, Irvine CS295, Spring 2017 Shuang Zhao 1 Announcements Final presentation June 13 (Tuesday)
AN ADAPTIVE POPULATION IMPORTANCE SAMPLER. Luca Martino*, Victor Elvira\ David Luengcfi, Jukka Corander*
 AN ADAPTIVE POPULATION IMPORTANCE SAMPLER Luca Martino*, Victor Elvira\ David Luengcfi, Jukka Corander* * Dep. of Mathematics and Statistics, University of Helsinki, 00014 Helsinki (Finland). t Dep. of
AN ADAPTIVE POPULATION IMPORTANCE SAMPLER Luca Martino*, Victor Elvira\ David Luengcfi, Jukka Corander* * Dep. of Mathematics and Statistics, University of Helsinki, 00014 Helsinki (Finland). t Dep. of
Modified Metropolis-Hastings algorithm with delayed rejection
 Modified Metropolis-Hastings algorithm with delayed reection K.M. Zuev & L.S. Katafygiotis Department of Civil Engineering, Hong Kong University of Science and Technology, Hong Kong, China ABSTRACT: The
Modified Metropolis-Hastings algorithm with delayed reection K.M. Zuev & L.S. Katafygiotis Department of Civil Engineering, Hong Kong University of Science and Technology, Hong Kong, China ABSTRACT: The
CS839: Probabilistic Graphical Models. Lecture 10: Learning with Partially Observed Data. Theo Rekatsinas
 CS839: Probabilistic Graphical Models Lecture 10: Learning with Partially Observed Data Theo Rekatsinas 1 Partially Observed GMs Speech recognition 2 Partially Observed GMs Evolution 3 Partially Observed
CS839: Probabilistic Graphical Models Lecture 10: Learning with Partially Observed Data Theo Rekatsinas 1 Partially Observed GMs Speech recognition 2 Partially Observed GMs Evolution 3 Partially Observed
Nested Sampling: Introduction and Implementation
 UNIVERSITY OF TEXAS AT SAN ANTONIO Nested Sampling: Introduction and Implementation Liang Jing May 2009 1 1 ABSTRACT Nested Sampling is a new technique to calculate the evidence, Z = P(D M) = p(d θ, M)p(θ
UNIVERSITY OF TEXAS AT SAN ANTONIO Nested Sampling: Introduction and Implementation Liang Jing May 2009 1 1 ABSTRACT Nested Sampling is a new technique to calculate the evidence, Z = P(D M) = p(d θ, M)p(θ
CIS 520, Machine Learning, Fall 2015: Assignment 7 Due: Mon, Nov 16, :59pm, PDF to Canvas [100 points]
![CIS 520, Machine Learning, Fall 2015: Assignment 7 Due: Mon, Nov 16, :59pm, PDF to Canvas [100 points] CIS 520, Machine Learning, Fall 2015: Assignment 7 Due: Mon, Nov 16, :59pm, PDF to Canvas [100 points]](/thumbs/89/100746783.jpg) CIS 520, Machine Learning, Fall 2015: Assignment 7 Due: Mon, Nov 16, 2015. 11:59pm, PDF to Canvas [100 points] Instructions. Please write up your responses to the following problems clearly and concisely.
CIS 520, Machine Learning, Fall 2015: Assignment 7 Due: Mon, Nov 16, 2015. 11:59pm, PDF to Canvas [100 points] Instructions. Please write up your responses to the following problems clearly and concisely.
Convergence and Efficiency of Adaptive MCMC. Jinyoung Yang
 Convergence and Efficiency of Adaptive MCMC by Jinyoung Yang A thesis submitted in conformity with the requirements for the degree of Doctor of Philosophy Graduate Department of Statistical Sciences University
Convergence and Efficiency of Adaptive MCMC by Jinyoung Yang A thesis submitted in conformity with the requirements for the degree of Doctor of Philosophy Graduate Department of Statistical Sciences University
A noninformative Bayesian approach to small area estimation
 A noninformative Bayesian approach to small area estimation Glen Meeden School of Statistics University of Minnesota Minneapolis, MN 55455 glen@stat.umn.edu September 2001 Revised May 2002 Research supported
A noninformative Bayesian approach to small area estimation Glen Meeden School of Statistics University of Minnesota Minneapolis, MN 55455 glen@stat.umn.edu September 2001 Revised May 2002 Research supported
Tutorial using BEAST v2.4.1 Troubleshooting David A. Rasmussen
 Tutorial using BEAST v2.4.1 Troubleshooting David A. Rasmussen 1 Background The primary goal of most phylogenetic analyses in BEAST is to infer the posterior distribution of trees and associated model
Tutorial using BEAST v2.4.1 Troubleshooting David A. Rasmussen 1 Background The primary goal of most phylogenetic analyses in BEAST is to infer the posterior distribution of trees and associated model
The Plan: Basic statistics: Random and pseudorandom numbers and their generation: Chapter 16.
 Scientific Computing with Case Studies SIAM Press, 29 http://www.cs.umd.edu/users/oleary/sccswebpage Lecture Notes for Unit IV Monte Carlo Computations Dianne P. O Leary c 28 What is a Monte-Carlo method?
Scientific Computing with Case Studies SIAM Press, 29 http://www.cs.umd.edu/users/oleary/sccswebpage Lecture Notes for Unit IV Monte Carlo Computations Dianne P. O Leary c 28 What is a Monte-Carlo method?
K-Means and Gaussian Mixture Models
 K-Means and Gaussian Mixture Models David Rosenberg New York University June 15, 2015 David Rosenberg (New York University) DS-GA 1003 June 15, 2015 1 / 43 K-Means Clustering Example: Old Faithful Geyser
K-Means and Gaussian Mixture Models David Rosenberg New York University June 15, 2015 David Rosenberg (New York University) DS-GA 1003 June 15, 2015 1 / 43 K-Means Clustering Example: Old Faithful Geyser
Machine Learning
 Machine Learning 10-601 Tom M. Mitchell Machine Learning Department Carnegie Mellon University April 1, 2019 Today: Inference in graphical models Learning graphical models Readings: Bishop chapter 8 Bayesian
Machine Learning 10-601 Tom M. Mitchell Machine Learning Department Carnegie Mellon University April 1, 2019 Today: Inference in graphical models Learning graphical models Readings: Bishop chapter 8 Bayesian
Image analysis. Computer Vision and Classification Image Segmentation. 7 Image analysis
 7 Computer Vision and Classification 413 / 458 Computer Vision and Classification The k-nearest-neighbor method The k-nearest-neighbor (knn) procedure has been used in data analysis and machine learning
7 Computer Vision and Classification 413 / 458 Computer Vision and Classification The k-nearest-neighbor method The k-nearest-neighbor (knn) procedure has been used in data analysis and machine learning
Bayesian Statistics Group 8th March Slice samplers. (A very brief introduction) The basic idea
 Bayesian Statistics Group 8th March 2000 Slice samplers (A very brief introduction) The basic idea lacements To sample from a distribution, simply sample uniformly from the region under the density function
Bayesian Statistics Group 8th March 2000 Slice samplers (A very brief introduction) The basic idea lacements To sample from a distribution, simply sample uniformly from the region under the density function
Reddit Recommendation System Daniel Poon, Yu Wu, David (Qifan) Zhang CS229, Stanford University December 11 th, 2011
 Reddit Recommendation System Daniel Poon, Yu Wu, David (Qifan) Zhang CS229, Stanford University December 11 th, 2011 1. Introduction Reddit is one of the most popular online social news websites with millions
Reddit Recommendation System Daniel Poon, Yu Wu, David (Qifan) Zhang CS229, Stanford University December 11 th, 2011 1. Introduction Reddit is one of the most popular online social news websites with millions
Stat 547 Assignment 3
 Stat 547 Assignment 3 Release Date: Saturday April 16, 2011 Due Date: Wednesday, April 27, 2011 at 4:30 PST Note that the deadline for this assignment is one day before the final project deadline, and
Stat 547 Assignment 3 Release Date: Saturday April 16, 2011 Due Date: Wednesday, April 27, 2011 at 4:30 PST Note that the deadline for this assignment is one day before the final project deadline, and
CSCI 599 Class Presenta/on. Zach Levine. Markov Chain Monte Carlo (MCMC) HMM Parameter Es/mates
 CSCI 599 Class Presenta/on Zach Levine Markov Chain Monte Carlo (MCMC) HMM Parameter Es/mates April 26 th, 2012 Topics Covered in this Presenta2on A (Brief) Review of HMMs HMM Parameter Learning Expecta2on-
CSCI 599 Class Presenta/on Zach Levine Markov Chain Monte Carlo (MCMC) HMM Parameter Es/mates April 26 th, 2012 Topics Covered in this Presenta2on A (Brief) Review of HMMs HMM Parameter Learning Expecta2on-
Discussion on Bayesian Model Selection and Parameter Estimation in Extragalactic Astronomy by Martin Weinberg
 Discussion on Bayesian Model Selection and Parameter Estimation in Extragalactic Astronomy by Martin Weinberg Phil Gregory Physics and Astronomy Univ. of British Columbia Introduction Martin Weinberg reported
Discussion on Bayesian Model Selection and Parameter Estimation in Extragalactic Astronomy by Martin Weinberg Phil Gregory Physics and Astronomy Univ. of British Columbia Introduction Martin Weinberg reported
Grundlagen der Künstlichen Intelligenz
 Grundlagen der Künstlichen Intelligenz Unsupervised learning Daniel Hennes 29.01.2018 (WS 2017/18) University Stuttgart - IPVS - Machine Learning & Robotics 1 Today Supervised learning Regression (linear
Grundlagen der Künstlichen Intelligenz Unsupervised learning Daniel Hennes 29.01.2018 (WS 2017/18) University Stuttgart - IPVS - Machine Learning & Robotics 1 Today Supervised learning Regression (linear
Machine Learning
 Machine Learning 10-601 Tom M. Mitchell Machine Learning Department Carnegie Mellon University October 9, 2012 Today: Graphical models Bayes Nets: Inference Learning Readings: Required: Bishop chapter
Machine Learning 10-601 Tom M. Mitchell Machine Learning Department Carnegie Mellon University October 9, 2012 Today: Graphical models Bayes Nets: Inference Learning Readings: Required: Bishop chapter
Clustering Lecture 5: Mixture Model
 Clustering Lecture 5: Mixture Model Jing Gao SUNY Buffalo 1 Outline Basics Motivation, definition, evaluation Methods Partitional Hierarchical Density-based Mixture model Spectral methods Advanced topics
Clustering Lecture 5: Mixture Model Jing Gao SUNY Buffalo 1 Outline Basics Motivation, definition, evaluation Methods Partitional Hierarchical Density-based Mixture model Spectral methods Advanced topics
Approximate (Monte Carlo) Inference in Bayes Nets. Monte Carlo (continued)
 Approximate (Monte Carlo) Inference in Bayes Nets Basic idea: Let s repeatedly sample according to the distribution represented by the Bayes Net. If in 400/1000 draws, the variable X is true, then we estimate
Approximate (Monte Carlo) Inference in Bayes Nets Basic idea: Let s repeatedly sample according to the distribution represented by the Bayes Net. If in 400/1000 draws, the variable X is true, then we estimate
Introduction to Machine Learning CMU-10701
 Introduction to Machine Learning CMU-10701 Clustering and EM Barnabás Póczos & Aarti Singh Contents Clustering K-means Mixture of Gaussians Expectation Maximization Variational Methods 2 Clustering 3 K-
Introduction to Machine Learning CMU-10701 Clustering and EM Barnabás Póczos & Aarti Singh Contents Clustering K-means Mixture of Gaussians Expectation Maximization Variational Methods 2 Clustering 3 K-
10703 Deep Reinforcement Learning and Control
 10703 Deep Reinforcement Learning and Control Russ Salakhutdinov Machine Learning Department rsalakhu@cs.cmu.edu Policy Gradient I Used Materials Disclaimer: Much of the material and slides for this lecture
10703 Deep Reinforcement Learning and Control Russ Salakhutdinov Machine Learning Department rsalakhu@cs.cmu.edu Policy Gradient I Used Materials Disclaimer: Much of the material and slides for this lecture
Scientific Computing with Case Studies SIAM Press, Lecture Notes for Unit IV Monte Carlo
 Scientific Computing with Case Studies SIAM Press, 2009 http://www.cs.umd.edu/users/oleary/sccswebpage Lecture Notes for Unit IV Monte Carlo Computations Dianne P. O Leary c 2008 1 What is a Monte-Carlo
Scientific Computing with Case Studies SIAM Press, 2009 http://www.cs.umd.edu/users/oleary/sccswebpage Lecture Notes for Unit IV Monte Carlo Computations Dianne P. O Leary c 2008 1 What is a Monte-Carlo
Machine Learning A W 1sst KU. b) [1 P] Give an example for a probability distributions P (A, B, C) that disproves
![Machine Learning A W 1sst KU. b) [1 P] Give an example for a probability distributions P (A, B, C) that disproves Machine Learning A W 1sst KU. b) [1 P] Give an example for a probability distributions P (A, B, C) that disproves](/thumbs/93/111796408.jpg) Machine Learning A 708.064 11W 1sst KU Exercises Problems marked with * are optional. 1 Conditional Independence I [2 P] a) [1 P] Give an example for a probability distribution P (A, B, C) that disproves
Machine Learning A 708.064 11W 1sst KU Exercises Problems marked with * are optional. 1 Conditional Independence I [2 P] a) [1 P] Give an example for a probability distribution P (A, B, C) that disproves
Variational Methods for Graphical Models
 Chapter 2 Variational Methods for Graphical Models 2.1 Introduction The problem of probabb1istic inference in graphical models is the problem of computing a conditional probability distribution over the
Chapter 2 Variational Methods for Graphical Models 2.1 Introduction The problem of probabb1istic inference in graphical models is the problem of computing a conditional probability distribution over the
Analysis of Incomplete Multivariate Data
 Analysis of Incomplete Multivariate Data J. L. Schafer Department of Statistics The Pennsylvania State University USA CHAPMAN & HALL/CRC A CR.C Press Company Boca Raton London New York Washington, D.C.
Analysis of Incomplete Multivariate Data J. L. Schafer Department of Statistics The Pennsylvania State University USA CHAPMAN & HALL/CRC A CR.C Press Company Boca Raton London New York Washington, D.C.
RJaCGH, a package for analysis of
 RJaCGH, a package for analysis of CGH arrays with Reversible Jump MCMC 1. CGH Arrays: Biological problem: Changes in number of DNA copies are associated to cancer activity. Microarray technology: Oscar
RJaCGH, a package for analysis of CGH arrays with Reversible Jump MCMC 1. CGH Arrays: Biological problem: Changes in number of DNA copies are associated to cancer activity. Microarray technology: Oscar
Mesh segmentation. Florent Lafarge Inria Sophia Antipolis - Mediterranee
 Mesh segmentation Florent Lafarge Inria Sophia Antipolis - Mediterranee Outline What is mesh segmentation? M = {V,E,F} is a mesh S is either V, E or F (usually F) A Segmentation is a set of sub-meshes
Mesh segmentation Florent Lafarge Inria Sophia Antipolis - Mediterranee Outline What is mesh segmentation? M = {V,E,F} is a mesh S is either V, E or F (usually F) A Segmentation is a set of sub-meshes
Mixture Models and the EM Algorithm
 Mixture Models and the EM Algorithm Padhraic Smyth, Department of Computer Science University of California, Irvine c 2017 1 Finite Mixture Models Say we have a data set D = {x 1,..., x N } where x i is
Mixture Models and the EM Algorithm Padhraic Smyth, Department of Computer Science University of California, Irvine c 2017 1 Finite Mixture Models Say we have a data set D = {x 1,..., x N } where x i is
L10. PARTICLE FILTERING CONTINUED. NA568 Mobile Robotics: Methods & Algorithms
 L10. PARTICLE FILTERING CONTINUED NA568 Mobile Robotics: Methods & Algorithms Gaussian Filters The Kalman filter and its variants can only model (unimodal) Gaussian distributions Courtesy: K. Arras Motivation
L10. PARTICLE FILTERING CONTINUED NA568 Mobile Robotics: Methods & Algorithms Gaussian Filters The Kalman filter and its variants can only model (unimodal) Gaussian distributions Courtesy: K. Arras Motivation
GiRaF: a toolbox for Gibbs Random Fields analysis
 GiRaF: a toolbox for Gibbs Random Fields analysis Julien Stoehr *1, Pierre Pudlo 2, and Nial Friel 1 1 University College Dublin 2 Aix-Marseille Université February 24, 2016 Abstract GiRaF package offers
GiRaF: a toolbox for Gibbs Random Fields analysis Julien Stoehr *1, Pierre Pudlo 2, and Nial Friel 1 1 University College Dublin 2 Aix-Marseille Université February 24, 2016 Abstract GiRaF package offers
Machine Learning
 Machine Learning 10-601 Tom M. Mitchell Machine Learning Department Carnegie Mellon University March 4, 2015 Today: Graphical models Bayes Nets: EM Mixture of Gaussian clustering Learning Bayes Net structure
Machine Learning 10-601 Tom M. Mitchell Machine Learning Department Carnegie Mellon University March 4, 2015 Today: Graphical models Bayes Nets: EM Mixture of Gaussian clustering Learning Bayes Net structure
CSC412: Stochastic Variational Inference. David Duvenaud
 CSC412: Stochastic Variational Inference David Duvenaud Admin A3 will be released this week and will be shorter Motivation for REINFORCE Class projects Class Project ideas Develop a generative model for
CSC412: Stochastic Variational Inference David Duvenaud Admin A3 will be released this week and will be shorter Motivation for REINFORCE Class projects Class Project ideas Develop a generative model for
arxiv: v2 [stat.co] 19 Feb 2016
![arxiv: v2 [stat.co] 19 Feb 2016 arxiv: v2 [stat.co] 19 Feb 2016](/thumbs/73/68585270.jpg) Noname manuscript No. (will be inserted by the editor) Issues in the Multiple Try Metropolis mixing L. Martino F. Louzada Received: date / Accepted: date arxiv:158.4253v2 [stat.co] 19 Feb 216 Abstract
Noname manuscript No. (will be inserted by the editor) Issues in the Multiple Try Metropolis mixing L. Martino F. Louzada Received: date / Accepted: date arxiv:158.4253v2 [stat.co] 19 Feb 216 Abstract
CS 229 Midterm Review
 CS 229 Midterm Review Course Staff Fall 2018 11/2/2018 Outline Today: SVMs Kernels Tree Ensembles EM Algorithm / Mixture Models [ Focus on building intuition, less so on solving specific problems. Ask
CS 229 Midterm Review Course Staff Fall 2018 11/2/2018 Outline Today: SVMs Kernels Tree Ensembles EM Algorithm / Mixture Models [ Focus on building intuition, less so on solving specific problems. Ask
Lecture 6: Spectral Graph Theory I
 A Theorist s Toolkit (CMU 18-859T, Fall 013) Lecture 6: Spectral Graph Theory I September 5, 013 Lecturer: Ryan O Donnell Scribe: Jennifer Iglesias 1 Graph Theory For this course we will be working on
A Theorist s Toolkit (CMU 18-859T, Fall 013) Lecture 6: Spectral Graph Theory I September 5, 013 Lecturer: Ryan O Donnell Scribe: Jennifer Iglesias 1 Graph Theory For this course we will be working on
arxiv: v3 [stat.co] 27 Apr 2012
![arxiv: v3 [stat.co] 27 Apr 2012 arxiv: v3 [stat.co] 27 Apr 2012](/thumbs/86/93793267.jpg) A multi-point Metropolis scheme with generic weight functions arxiv:1112.4048v3 stat.co 27 Apr 2012 Abstract Luca Martino, Victor Pascual Del Olmo, Jesse Read Department of Signal Theory and Communications,
A multi-point Metropolis scheme with generic weight functions arxiv:1112.4048v3 stat.co 27 Apr 2012 Abstract Luca Martino, Victor Pascual Del Olmo, Jesse Read Department of Signal Theory and Communications,
Hierarchical Bayesian Modeling with Ensemble MCMC. Eric B. Ford (Penn State) Bayesian Computing for Astronomical Data Analysis June 12, 2014
 Hierarchical Bayesian Modeling with Ensemble MCMC Eric B. Ford (Penn State) Bayesian Computing for Astronomical Data Analysis June 12, 2014 Simple Markov Chain Monte Carlo Initialise chain with θ 0 (initial
Hierarchical Bayesian Modeling with Ensemble MCMC Eric B. Ford (Penn State) Bayesian Computing for Astronomical Data Analysis June 12, 2014 Simple Markov Chain Monte Carlo Initialise chain with θ 0 (initial
Slice sampler algorithm for generalized Pareto distribution
 Slice sampler algorithm for generalized Pareto distribution Mohammad Rostami, Mohd Bakri Adam Yahya, Mohamed Hisham Yahya, Noor Akma Ibrahim Abstract In this paper, we developed the slice sampler algorithm
Slice sampler algorithm for generalized Pareto distribution Mohammad Rostami, Mohd Bakri Adam Yahya, Mohamed Hisham Yahya, Noor Akma Ibrahim Abstract In this paper, we developed the slice sampler algorithm
Particle Filters for Visual Tracking
 Particle Filters for Visual Tracking T. Chateau, Pascal Institute, Clermont-Ferrand 1 Content Particle filtering: a probabilistic framework SIR particle filter MCMC particle filter RJMCMC particle filter
Particle Filters for Visual Tracking T. Chateau, Pascal Institute, Clermont-Ferrand 1 Content Particle filtering: a probabilistic framework SIR particle filter MCMC particle filter RJMCMC particle filter
This chapter explains two techniques which are frequently used throughout
 Chapter 2 Basic Techniques This chapter explains two techniques which are frequently used throughout this thesis. First, we will introduce the concept of particle filters. A particle filter is a recursive
Chapter 2 Basic Techniques This chapter explains two techniques which are frequently used throughout this thesis. First, we will introduce the concept of particle filters. A particle filter is a recursive
Introduction to Pattern Recognition Part II. Selim Aksoy Bilkent University Department of Computer Engineering
 Introduction to Pattern Recognition Part II Selim Aksoy Bilkent University Department of Computer Engineering saksoy@cs.bilkent.edu.tr RETINA Pattern Recognition Tutorial, Summer 2005 Overview Statistical
Introduction to Pattern Recognition Part II Selim Aksoy Bilkent University Department of Computer Engineering saksoy@cs.bilkent.edu.tr RETINA Pattern Recognition Tutorial, Summer 2005 Overview Statistical
Bayesian Modelling with JAGS and R
 Bayesian Modelling with JAGS and R Martyn Plummer International Agency for Research on Cancer Rencontres R, 3 July 2012 CRAN Task View Bayesian Inference The CRAN Task View Bayesian Inference is maintained
Bayesian Modelling with JAGS and R Martyn Plummer International Agency for Research on Cancer Rencontres R, 3 July 2012 CRAN Task View Bayesian Inference The CRAN Task View Bayesian Inference is maintained
Parallel Gibbs Sampling From Colored Fields to Thin Junction Trees
 Parallel Gibbs Sampling From Colored Fields to Thin Junction Trees Joseph Gonzalez Yucheng Low Arthur Gretton Carlos Guestrin Draw Samples Sampling as an Inference Procedure Suppose we wanted to know the
Parallel Gibbs Sampling From Colored Fields to Thin Junction Trees Joseph Gonzalez Yucheng Low Arthur Gretton Carlos Guestrin Draw Samples Sampling as an Inference Procedure Suppose we wanted to know the
Expectation-Maximization Methods in Population Analysis. Robert J. Bauer, Ph.D. ICON plc.
 Expectation-Maximization Methods in Population Analysis Robert J. Bauer, Ph.D. ICON plc. 1 Objective The objective of this tutorial is to briefly describe the statistical basis of Expectation-Maximization
Expectation-Maximization Methods in Population Analysis Robert J. Bauer, Ph.D. ICON plc. 1 Objective The objective of this tutorial is to briefly describe the statistical basis of Expectation-Maximization
Clustering: Classic Methods and Modern Views
 Clustering: Classic Methods and Modern Views Marina Meilă University of Washington mmp@stat.washington.edu June 22, 2015 Lorentz Center Workshop on Clusters, Games and Axioms Outline Paradigms for clustering
Clustering: Classic Methods and Modern Views Marina Meilă University of Washington mmp@stat.washington.edu June 22, 2015 Lorentz Center Workshop on Clusters, Games and Axioms Outline Paradigms for clustering
Warped Mixture Models
 Warped Mixture Models Tomoharu Iwata, David Duvenaud, Zoubin Ghahramani Cambridge University Computational and Biological Learning Lab March 11, 2013 OUTLINE Motivation Gaussian Process Latent Variable
Warped Mixture Models Tomoharu Iwata, David Duvenaud, Zoubin Ghahramani Cambridge University Computational and Biological Learning Lab March 11, 2013 OUTLINE Motivation Gaussian Process Latent Variable
Improved Adaptive Rejection Metropolis Sampling Algorithms
 Improved Adaptive Rejection Metropolis Sampling Algorithms 1 Luca Martino, Jesse Read, David Luengo Department of Signal Theory and Communications, Universidad Carlos III de Madrid. arxiv:1205.5494v4 [stat.co]
Improved Adaptive Rejection Metropolis Sampling Algorithms 1 Luca Martino, Jesse Read, David Luengo Department of Signal Theory and Communications, Universidad Carlos III de Madrid. arxiv:1205.5494v4 [stat.co]
Expectation Maximization (EM) and Gaussian Mixture Models
 Expectation Maximization (EM) and Gaussian Mixture Models Reference: The Elements of Statistical Learning, by T. Hastie, R. Tibshirani, J. Friedman, Springer 1 2 3 4 5 6 7 8 Unsupervised Learning Motivation
Expectation Maximization (EM) and Gaussian Mixture Models Reference: The Elements of Statistical Learning, by T. Hastie, R. Tibshirani, J. Friedman, Springer 1 2 3 4 5 6 7 8 Unsupervised Learning Motivation
COMPUTATIONAL STATISTICS UNSUPERVISED LEARNING
 COMPUTATIONAL STATISTICS UNSUPERVISED LEARNING Luca Bortolussi Department of Mathematics and Geosciences University of Trieste Office 238, third floor, H2bis luca@dmi.units.it Trieste, Winter Semester
COMPUTATIONAL STATISTICS UNSUPERVISED LEARNING Luca Bortolussi Department of Mathematics and Geosciences University of Trieste Office 238, third floor, H2bis luca@dmi.units.it Trieste, Winter Semester
Statistical techniques for data analysis in Cosmology
 Statistical techniques for data analysis in Cosmology arxiv:0712.3028; arxiv:0911.3105 Numerical recipes (the bible ) Licia Verde ICREA & ICC UB-IEEC http://icc.ub.edu/~liciaverde outline Lecture 1: Introduction
Statistical techniques for data analysis in Cosmology arxiv:0712.3028; arxiv:0911.3105 Numerical recipes (the bible ) Licia Verde ICREA & ICC UB-IEEC http://icc.ub.edu/~liciaverde outline Lecture 1: Introduction
Probabilistic Graphical Models
 Overview of Part One Probabilistic Graphical Models Part One: Graphs and Markov Properties Christopher M. Bishop Graphs and probabilities Directed graphs Markov properties Undirected graphs Examples Microsoft
Overview of Part One Probabilistic Graphical Models Part One: Graphs and Markov Properties Christopher M. Bishop Graphs and probabilities Directed graphs Markov properties Undirected graphs Examples Microsoft
( ) =cov X Y = W PRINCIPAL COMPONENT ANALYSIS. Eigenvectors of the covariance matrix are the principal components
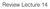 Review Lecture 14 ! PRINCIPAL COMPONENT ANALYSIS Eigenvectors of the covariance matrix are the principal components 1. =cov X Top K principal components are the eigenvectors with K largest eigenvalues
Review Lecture 14 ! PRINCIPAL COMPONENT ANALYSIS Eigenvectors of the covariance matrix are the principal components 1. =cov X Top K principal components are the eigenvectors with K largest eigenvalues
Probabilistic Robotics
 Probabilistic Robotics Discrete Filters and Particle Filters Models Some slides adopted from: Wolfram Burgard, Cyrill Stachniss, Maren Bennewitz, Kai Arras and Probabilistic Robotics Book SA-1 Probabilistic
Probabilistic Robotics Discrete Filters and Particle Filters Models Some slides adopted from: Wolfram Burgard, Cyrill Stachniss, Maren Bennewitz, Kai Arras and Probabilistic Robotics Book SA-1 Probabilistic
Missing Data Analysis for the Employee Dataset
 Missing Data Analysis for the Employee Dataset 67% of the observations have missing values! Modeling Setup Random Variables: Y i =(Y i1,...,y ip ) 0 =(Y i,obs, Y i,miss ) 0 R i =(R i1,...,r ip ) 0 ( 1
Missing Data Analysis for the Employee Dataset 67% of the observations have missing values! Modeling Setup Random Variables: Y i =(Y i1,...,y ip ) 0 =(Y i,obs, Y i,miss ) 0 R i =(R i1,...,r ip ) 0 ( 1
Physics 736. Experimental Methods in Nuclear-, Particle-, and Astrophysics. - Statistical Methods -
 Physics 736 Experimental Methods in Nuclear-, Particle-, and Astrophysics - Statistical Methods - Karsten Heeger heeger@wisc.edu Course Schedule and Reading course website http://neutrino.physics.wisc.edu/teaching/phys736/
Physics 736 Experimental Methods in Nuclear-, Particle-, and Astrophysics - Statistical Methods - Karsten Heeger heeger@wisc.edu Course Schedule and Reading course website http://neutrino.physics.wisc.edu/teaching/phys736/
Statistical Matching using Fractional Imputation
 Statistical Matching using Fractional Imputation Jae-Kwang Kim 1 Iowa State University 1 Joint work with Emily Berg and Taesung Park 1 Introduction 2 Classical Approaches 3 Proposed method 4 Application:
Statistical Matching using Fractional Imputation Jae-Kwang Kim 1 Iowa State University 1 Joint work with Emily Berg and Taesung Park 1 Introduction 2 Classical Approaches 3 Proposed method 4 Application:
SGN (4 cr) Chapter 11
 SGN-41006 (4 cr) Chapter 11 Clustering Jussi Tohka & Jari Niemi Department of Signal Processing Tampere University of Technology February 25, 2014 J. Tohka & J. Niemi (TUT-SGN) SGN-41006 (4 cr) Chapter
SGN-41006 (4 cr) Chapter 11 Clustering Jussi Tohka & Jari Niemi Department of Signal Processing Tampere University of Technology February 25, 2014 J. Tohka & J. Niemi (TUT-SGN) SGN-41006 (4 cr) Chapter
Linear Modeling with Bayesian Statistics
 Linear Modeling with Bayesian Statistics Bayesian Approach I I I I I Estimate probability of a parameter State degree of believe in specific parameter values Evaluate probability of hypothesis given the
Linear Modeling with Bayesian Statistics Bayesian Approach I I I I I Estimate probability of a parameter State degree of believe in specific parameter values Evaluate probability of hypothesis given the
