Plaid models, biclustering, clustering on subsets of attributes, feature selection in clustering, et al.
|
|
|
- Milo Gardner
- 6 years ago
- Views:
Transcription
1 Plaid models, biclustering, clustering on subsets of attributes, feature selection in clustering, et al. Ramón Díaz-Uriarte rdiaz Unidad de Bioinformática Centro Nacional de Investigaciones Oncológicas (CNIO) (Spanish National Cancer Center) Systems Biology Seminar at CNIO, Plaid models et al. p. 1
2 Introduction Limitations of usual clustering: Uses all genes to cluster the samples (and/or uses all samples to cluster all genes). Disjoint clusters. However, might want to allow clusters of genes to be defined only with respect to a subset of samples (and vice-versa); some genes to be in more than one cluster; Plaid models et al. p. 2
3 Of particular interest because: Often suspect groups can be heterogeneous even in supervised settings. We are concerned about possible interactions: non-additive effects of genes. Plaid models, biclusters, et al., can be of potential use in these exploratory pursuits (of course, we are no longer in the more clearly defined setting of hypothesis testing). Plaid models et al. p. 3
4 Objectives Review several (most) plaid-like and biclustering techniques, with emphasis on those that are already implemented (see side note ). Show an example with real data. Discussion, suggestions for future research, and extensions. Plaid models et al. p. 4
5 Side note: software should be available If a promising technique is to live up to its promises, software to use the technique ought to be freely available. Applied statisticians and biologists do not have the time to implement any and every idea that is published, nor to deal with the complications of patented algorithms. It is not OK to get answers such as the code is not available but my method is straightforward to implement from the explanations in my paper ;... we can think of a collaboration, and I ll analyze the data for you.... the method will soon be available (exclusively) as part of program XYZ (which is proprietary). Plaid models et al. p. 5
6 reference implementation (Ripley, 2002): will allow users to run the procedure in moderately sized problems and repeat results. Ideally: Source code available in a widely used language (C/C++, FORTRAN, R); allows: fixing bugs; understanding exactly what is done; modifying the code for exploration/extensions; Compiled or ready to run version for a variety of platforms. This talk will be biased in favor of methods with code: in the end, a pragmatic decision. Plaid models et al. p. 6
7 Methods we ll cover Plaid-like models (including Plaid, Cheng & Church, FLOC, xmotif, PRN). SAMBA. Optimal variable weighting in clustering. Clustering on subsets of attributes. Plaid models et al. p. 7
8 Biclustering: definition The intuitive notion of a bicluster is a subset of genes that exhibit similar expression patterns over a subset of conditions. Following this intuition we define a bicluster as a subset of genes that jointly respond across a subset of conditions, where a gene is termed responding in some condition if its expression level changes significantly at that condition w.r.t. its normal level. (Analysis of Gene Expression Data, Lecture 8, Spring 2002, by R. Shamir and R. Sharan, p. 6.) Plaid models et al. p. 8
9 What does similar expression patterns mean, exactly? Plaid models et al. p. 9
10 What does similar expression patterns mean, exactly? What does jointly respond mean, exactly? Should all genes change by exactly the same amount, or are proportional changes allowed (e.g., gene 1: 2x; gene 2: 4x; etc)? Are subject (or array) effects allowed? (genes 1 to 5 increase, but in subject 1 they increase 2x, and in subject 2 they increase 3x). Plaid models et al. p. 9
11 What does similar expression patterns mean, exactly? What does jointly respond mean, exactly? Should all genes change by exactly the same amount, or are proportional changes allowed (e.g., gene 1: 2x; gene 2: 4x; etc)? Are subject (or array) effects allowed? (genes 1 to 5 increase, but in subject 1 they increase 2x, and in subject 2 they increase 3x). (... ) expression level changes significantly : are we discretizing changes (up, down no change)? how? why not model the original, continuous, data? Plaid models et al. p. 9
12 Methods we ll cover Plaid-like models (including Plaid, Cheng & Church, FLOC, xmotif, PRN). SAMBA. Optimal variable weighting in clustering. Clustering on subsets of attributes. Plaid models et al. p. 10
13 Plaid model Plaid model: Lazzeroni & Owen, 2002, Statistica Sinica, (also Y : matrix of gene expression data: Y ij expression of gene i from subject (or array) j. i from 1,.., n, j from 1,..., p. Approximate Y ij as a sum of layers: each Y ij is modeled as the sum of several layers to which the entry ij belongs. Not all genes of a subject (not all i of a j) are part of the same layers (and vice-versa for the j of a i). Plaid models et al. p. 11
14 Plaid (II) Each layer k an Analysis of Variance Model. Four types: θ ijk = µ k θ ijk = µ k + α ik θ ijk = µ k + βjk θ ijk = µ k + α ik + βjk Not all layers need to be of the same type (e.g., first can have α and β, and second only µ). As we said, we approximate each Y ij as the sum of contributions from different layers: Y ij. K = θ ijk ρ ik κ jk k=0 Plaid models et al. p. 12
15 Plaid (III) We approximate: Y ij θ ij0 is the background layer.. K = θ ijk ρ ik κ jk k=0 ρ ik and κ jk are indicators: ρ ik = 1 if gene i is in the k th gene-block (0 otherwise) and κ jk = 1 if subject k is in the k th sample-block (0 otherwise). We allow genes and samples to belong to more than one layer ( k ρ ik 2 for some i, and similar for subjects), and some genes/samples not to belong to any layer ( k ρ ik = 0 for some i, and similar for subjects). (Constraints i ρ ikα ik = 0 and similarly for β k to avoid overparameterization). Plaid models et al. p. 13
16 Plaid (IV) We are trying to minimize n p (Y ij K θ ijk ρ ik κ jk ) 2 i=1 j=1 k=0 Like minimizing residual sum of squares: i.e., do the best possible job predicting Y ij as a sum of layers of (two-way) ANOVAs. (We call Z ij = Y ij K 1 k=0 θ ijkρ ik κ jk the residual from the first K 1 layers.). Plaid models et al. p. 14
17 Plaid (V) Values of α ik and β jk : orderings of the effects of layer k upon genes and samples. Combine gene clustering with variable selection on the samples, and sample clustering with variable selection on the genes. Can set constraints so that, for a given layer every µ + α i and every µ + β j have the same sign (i.e., all genes and samples in a layer are either over or underexpressed). Can constraint so that we exclude from a layer genes and/or samples that are not well explained by that layer (i.e., not large enough decrease in residual variance). Like setting a minimum R 2. Plaid models et al. p. 15
18 Plaid: algorithm See details in paper of how search for a layer carried out. A greedy algorithm that adds one layer at a time. When to stop? Importance or size of layer σk 2 = i θ ijk 2 ρ ik κ jk j After finding a layer, permute (residual) elements of Y ij (i.e., the Z ij ) by row and by column. Compute importance of permuted layers, and see if larger or smaller than importance of layer k. Plaid models et al. p. 16
19 Geometry of plaids (Really neat; easy to see if you draw 2-D case). For a layer with only µ, if we represent subjects as points in the space with genes as axes, all are clustered around µ (and similarly if we represent genes). For a layer with µ + β: Represent genes in space with subjects as axes: all genes clustered around point (µ + β 1, µ + β 2,..., µ + β p ). Represent subjects in space with genes as axes: all subjects cluster along a line segment (of slope 1) through the center µ; for each subject k its position is µ + β j. Also induces a positive correlation among genes: if good fitting model, values spread in an ellipsoid with major axis along the above segment. Analogous for µ + α. Plaid models et al. p. 17
20 Geometry of plaids (II) For a layer with µ + α + β: Genes in space with subjects as axes: clustered along a line segment with center (µ + β 1, µ + β 2,..., µ + β p ); for each gene i, its position is center + α i. Analogous for subjects in space with genes as axes. Plaid models et al. p. 18
21 Plaid: software Windows executable available from Source code might become available in the near future. Plaid models et al. p. 19
22 Other Plaid-like (I) Other approaches, developed independently. But they are really special cases of Plaid. Cheng & Church, Biclustering of Expression Data, Proc. ISMB 00, and the later improvement in FLOC (Yang, Wang, Wang, Yu, Enhanced Biclustering on Expression Data, BIBE 2003, or are like Plaid but: Do not allow setting any α or β equal to 0. Including a background layer does not seem supported. Formulated in a different way. Original algorithm problematic (substitution of layers by random numbers). FLOC solves the later problem, but same basic model. Is software available? Plaid models et al. p. 20
23 Other Plaid-like (II) xmotif, by Murali & Kasif ( Extracting conserved gene expression motifs from gene expression data, Like Plaid, but restricting β = 0. Software available at the above page (though I haven t been able to compile it). Plaid models et al. p. 21
24 Other Plaid-like (III) Probabilistic Relational Networks (PRN) by Segal, Battle & Koller ( Decomposing gene expression into cellular processes., PSB 2003, see also Segal et al., Bioinformatics, 2001, 1 (1): 1 9). Like Plaid, but sets α = 0, β 0. Since a richer probabilistic framework, allows incorporation of additional information, and heterogeneous data sets. A bayesian network: computational issues. All the model estimated in a single go (no greedy algorithm). Is this really an advantage? Need to decide number of layers before analyses. No help on how to do it (AIC-like approaches not straightforward). Software not available (nor trivial to implement). Plaid models et al. p. 22
25 Methods we ll cover Plaid-like models (including Plaid, Cheng & Church, FLOC, xmotif, PRN). SAMBA. Optimal variable weighting in clustering. Clustering on subsets of attributes. Plaid models et al. p. 23
26 SAMBA (I) By Tanay, Sharan, Shamir. ( Discovering statistically significant biclusters in gene expression data. Bioinformatics, 2002, 18: S136 S144. Also Given the gene expression data, form a bipartite graph. Connect conditions (subjects, arrays) with genes. Only connect (i.e., draw an edge from) a sample to a gene if the gene is differentially expressed in that sample. (And how is this determined?) Search for heavy subgraphs: subgraphs with a lot of connections. These are the layers. It does allow to incorporate additional information (GO terms, transcription factors) for layer formation. Plaid models et al. p. 24
27 SAMBA (II) Issues: How is a gene determined to be differentially expressed? What if we wanted to use a richer model (e.g., Parmigiani et al., in POE)? I am not sure to understand the underlying statistical model (might be my limitations), but it seems intrinsically more limited than Plaid. However, it has a lot more parameters (related to the optimization?). Software available as Java executable but: Not yet completely implemented nor documented. There are many parameters that can be changed (settings file) and are undocumented. The intrinsic validation of clusters (like a p-value) not available. Plaid models et al. p. 25
28 Methods we ll cover Plaid-like models (including Plaid, Cheng & Church, FLOC, xmotif, PRN). SAMBA. Optimal variable weighting in clustering. Clustering on subsets of attributes. Plaid models et al. p. 26
29 Optimal variable weighting (OVW) A large literature; main contributions by De Soete. We will follow Makarenkov & Legendre (J. Classification, 2001). See also For a typical problem of clustering p objects (e.g., arrays, subjects) on n variables (e.g., genes). Determine the optimal weights for each variable so that the dissimilarities (Euclidean distances) among objects satisfy certain optimality criteria (the dissimilarity between objects r and s is [ n i=1 w i(y ri y si ) 2 ] 1/2 ). (Note: notation changed to agree with Plaid). Can be applied to either k-means partitions or hierarchical clustering (different optimality criteria). Very important: the weights are the same for all objects. Plaid models et al. p. 27
30 Optimal variable weighting (II) Is it appropriate for microarray data? Might be, if we have already defined sets of groups of genes. But not really a biclustering problem as usually defined: a gene has a weight that is the same over all subjects. This is not really a model. Might lead to easier interpretation than other approaches. Anyway, software available as source code (and Win32 executable) from (the OVW program of Makarenkov & Legendre). [Note:compiles out of the box in GNU/Linux]. Plaid models et al. p. 28
31 Methods we ll cover Plaid-like models (including Plaid, Cheng & Church, FLOC, xmotif, PRN). SAMBA. Optimal variable weighting in clustering. Clustering on subsets of attributes. Plaid models et al. p. 29
32 Clustering on subsets of attributes Looks like OVW but Clustering on Subsets of Attributes, COSA, weights for a gene are different in different groups of subjects. By Friedman & Meulman ( Clusterin objects on subsets of attributes, Other technical differences in how optimization is carried out. For a set of groups or clusters C, with n variables (genes), we want to find the encoder function (the function which assigns each subject to a cluster) and the gene weights so that the within cluster distance is the smallest possible. This within cluster distance is a weighted average of distances over each variable (gene). This is the same as OVW for k-means. Now, consider the possibility that each gene has different weights for different groups or clusters. Plaid models et al. p. 30
33 COSA (II) We want to find the optimal cluster membership AND the optimal weights for each variable in each cluster, so that the within cluster distance is smallest. For all the genes n, and for a cluster c, we can write the distance between two subjects, r and s as: D rs = n i=1 w icd rsi (notation changed to follow Plaid). And we want to minimize the overall within cluster distance, or the sum over all clusters, of D rs, for all the r, s, that belong to the same cluster. Note that the above makes explicit that the weights are the weights of a gene in a cluster (w ic ). The above criterion yields groups that cluster only one one attribute (gene). Add an incentive (negative penalty) for multiple attribute solutions (i.e., several w ic > 0). Plaid models et al. p. 31
34 COSA (III) Some modifications are needed to make the above work in the hierarchical setting. Adds knear parameter, that controls the size of neighborhoods. Targeted clustering: search, explicitly, for value near a target (single target), such as very large values, or for values near two possible targets (dual target), such as either very large or very small. Once the procedure is run, evaluate if the variable importances are larger than expected from randomly formed clusters of the given size. Software available as an R package for Windows (uses a Windows executable). GNU/Linux version available soon? Plaid models et al. p. 32
35 COSA vs. Plaid Different objectives. Plaid: a model of gene expression data. COSA: a clustering algorithm. Plaid yields predictions of values (fitted values) based on a, hopefully, small number of parameters. COSA does not yield predictions. Plaid has overlapping clusters of subjects, COSA doesn t. Plaid s gene membership to a layer can be understood as gene weights, but they are crisp : either 0 or 1, in contrast to COSA. COSA might be easier to interpret (a small number of results). But COSA has many, many parameters and choices, with non-intuitive interpretations. COSA might encourage running hundreds of different parameter combinations. Plaid models et al. p. 33
36 Some results with real data All the papers show examples with real data. We will use a data set from a group at CNIO. For confidentiality reasons, we mask most details. Suffice to know we have 41 subjects and 6519 genes. Data normalized using print-tip loess, then a reference value for each gene subtracted. Researchers think it is likely that there are several subgroups of subjects defined w.r.t. subsets of genes. Plaid models et al. p. 34
37 Plaid examples We run two Plaid models (both include a background layer with µ k + α ik + βjk). Larger plaid : Use θ ijk = µ k + α ik + βjk for all layers. Eliminate rows and columns with R 2 < 0.5. These are the defaults used in Lazzeroni & Owen s gene expression example. Simpler plaid : Fit the background layer. Set minimum R 2 = 0.7. Fit layers with only µ (until no further layers found). Fit layers without β (until no further layers found). Fit layers without α (until no further layers found). Fit full two-way models (θ ijk = µ k + α ik + βjk) until no further rows or columns retained (total of 43 layers). Plaid models et al. p. 35
38 Larger plaid (I) Will show results with 53 layers (many more could be found; at least up to 98). The typical look of some of the output (summary description): Layer Rows Cols HasA HasB DF +/- SST SSM SSA SSB Y Y Y Y Y Y Y Y Y Y Y Y Y Y Y Y Y Y Y Y Y Y Plaid models et al. p. 36
39 Larger plaid (II) A model with parameters (about 8% of the data). Most layers a large number of genes: Min. 1st Qu. Median Mean 3rd Qu. Max Most models, few subjects: Number of subjects: Number of layers: Mean squares for genes are often very small (75% of them < 0.4) but mean squares for arrays are larger (50% > 1.7; 25% > 4.9). Plaid models et al. p. 37
40 Larger plaid (III) And how does it work (recall number of parameters 8% of data) Observed Plaid, 53 layers Fitted Plaid models et al. p. 38
41 Simpler plaid (I) Will show results with 43 layers; no more layers found with both rows and columns. First 13 layers: Layer Rows Cols HasA HasB DF +/- SST SSM SSA SSB Y Y Y Y Y Y Y Y Y Y Y Y Y Y Y Y Plaid models et al. p. 39
42 Simpler plaid (II) A model with 9013 parameters (about 3% of the data). Smaller number of genes per layer (recall release if R 2 < 0.7). Min. 1st Qu. Median Mean 3rd Qu. Max Most models, fewer subjects per layer, but not such a large difference: Number of subjects: Number of layers: Mean squares for genes are larger (50% of them > 0.4) and mean squares for arrays are smaller (50% > 0.8; 25% > 1.86). Layers are more intense w.r.t. genes. Plaid models et al. p. 40
43 Simpler plaid (III) And how does it work (recall number of parameters 3% of data) Observed Plaid, 43 layers Fitted Plaid models et al. p. 41
44 Simpler vs. Larger plaid Simpler plaid Simpler plaid Fitted values 6 Residuals from fit Larger plaid Larger plaid Plaid models et al. p. 42
45 Interpreting these Plaid models Of course, what remains to be done is that someone with subject matter knowledge take a look at the results... Plaid models et al. p. 43
46 COSA example I tried more than 100 different possible combinations of parameters and settings: type of target value, incentive for multiple clustering, size of near-neighborhoods, type of distance calculation. For every case, I first examined the dendrogram looking for decent dendrograms : initial branches much longer than final ones. If a half-decent dendrogram was found, I examined the existence of attribute importances larger than achievable by random grouping of subjects. Not difficult (setting the incentive for multiple clustering very close to 0) to find at least one cluster where just one attribute has a high importance; but, is this credible with 6500 genes? Plaid models et al. p. 44
47 COSA example (II) Plaid models et al. p. 45
48 COSA example (III) Plaid models et al. p. 46
49 COSA example (IV) Plaid models et al. p. 47
50 COSA example (V) Plaid models et al. p. 48
51 COSA example (VI) Plaid models et al. p. 49
52 Discussion Potential issues with COSA: How are parameters chosen? Can encourage running hundreds of models until something is found. Requires using the black art of clustering interpretation. A large enough cluster size is needed (to obtain good estimates of a scale term). Why do we want to use hierarchical clustering? Clusters of subjects are not often interpreted as if they belonged to a hierarchy (in contrast to what is often done in taxonomy). Easy to find clusters where just a single attribute is relevant. How stable are results to sampling variation? How dependent are results on sample size? Plaid models et al. p. 50
53 Discussion (II) Potential issues with Plaid: Not as many parameters as with COSA, but some decisions need to be made, and not easy to grasp the consequences down the analyses. Often a huge number of layers is found. Is this reasonable? Yes it is, if we think of how few parameters (in relative terms) this adds to the model. Order of analyses matters (e.g., first only α, till exhaustion, then α + β, and then we can fit only α again.) Sure, we know this from linear models, but annoying anyway. Difficult to know, ahead of time, what might be a reasonable strategy? How stable are results to sampling variation? How dependent are results on sample size and on size of layers? Plaid models et al. p. 51
54 Discussion (III) Some further research: How stable are results to sampling variation? How dependent are results on sample size and on size of layers? Integration of results from COSA and Plaid. Use of Plaid as a backbone for further developments on molecular signature problem. Plaid offers a nice, understandable, and extensible model. Plaid models et al. p. 52
55 Discussion (IV) A few caveats: these are not simple problems. These cannot be simple problems. We should not expect simple answers. We seem to be searching for homogeneous groups w.r.t. variables, and we don t know the groups or the variables. A discovery problem. Criteria to be optimized are often vague and there are many different choices. Sample size is still an issue. Much easier to get stable, convincing results if we have 500 subjects rather than 50 (assuming both sampled from same population). Thus, there are both conceptual and optimization difficulties. Plaid models et al. p. 53
56 Acknowledgements A. Pérez, M.J. Artiga, and M.A. Piris for the data. Laura Lazzeroni and Art Owen for clarifications about Plaid. Plaid models et al. p. 54
DNA chips and other techniques measure the expression level of a large number of genes, perhaps all
 INESC-ID TECHNICAL REPORT 1/2004, JANUARY 2004 1 Biclustering Algorithms for Biological Data Analysis: A Survey* Sara C. Madeira and Arlindo L. Oliveira Abstract A large number of clustering approaches
INESC-ID TECHNICAL REPORT 1/2004, JANUARY 2004 1 Biclustering Algorithms for Biological Data Analysis: A Survey* Sara C. Madeira and Arlindo L. Oliveira Abstract A large number of clustering approaches
2. Background. 2.1 Clustering
 2. Background 2.1 Clustering Clustering involves the unsupervised classification of data items into different groups or clusters. Unsupervised classificaiton is basically a learning task in which learning
2. Background 2.1 Clustering Clustering involves the unsupervised classification of data items into different groups or clusters. Unsupervised classificaiton is basically a learning task in which learning
Biclustering Bioinformatics Data Sets. A Possibilistic Approach
 Possibilistic algorithm Bioinformatics Data Sets: A Possibilistic Approach Dept Computer and Information Sciences, University of Genova ITALY EMFCSC Erice 20/4/2007 Bioinformatics Data Sets Outline Introduction
Possibilistic algorithm Bioinformatics Data Sets: A Possibilistic Approach Dept Computer and Information Sciences, University of Genova ITALY EMFCSC Erice 20/4/2007 Bioinformatics Data Sets Outline Introduction
Clustering CS 550: Machine Learning
 Clustering CS 550: Machine Learning This slide set mainly uses the slides given in the following links: http://www-users.cs.umn.edu/~kumar/dmbook/ch8.pdf http://www-users.cs.umn.edu/~kumar/dmbook/dmslides/chap8_basic_cluster_analysis.pdf
Clustering CS 550: Machine Learning This slide set mainly uses the slides given in the following links: http://www-users.cs.umn.edu/~kumar/dmbook/ch8.pdf http://www-users.cs.umn.edu/~kumar/dmbook/dmslides/chap8_basic_cluster_analysis.pdf
Biclustering Algorithms for Gene Expression Analysis
 Biclustering Algorithms for Gene Expression Analysis T. M. Murali August 19, 2008 Problems with Hierarchical Clustering It is a global clustering algorithm. Considers all genes to be equally important
Biclustering Algorithms for Gene Expression Analysis T. M. Murali August 19, 2008 Problems with Hierarchical Clustering It is a global clustering algorithm. Considers all genes to be equally important
Biclustering for Microarray Data: A Short and Comprehensive Tutorial
 Biclustering for Microarray Data: A Short and Comprehensive Tutorial 1 Arabinda Panda, 2 Satchidananda Dehuri 1 Department of Computer Science, Modern Engineering & Management Studies, Balasore 2 Department
Biclustering for Microarray Data: A Short and Comprehensive Tutorial 1 Arabinda Panda, 2 Satchidananda Dehuri 1 Department of Computer Science, Modern Engineering & Management Studies, Balasore 2 Department
Exploratory data analysis for microarrays
 Exploratory data analysis for microarrays Jörg Rahnenführer Computational Biology and Applied Algorithmics Max Planck Institute for Informatics D-66123 Saarbrücken Germany NGFN - Courses in Practical DNA
Exploratory data analysis for microarrays Jörg Rahnenführer Computational Biology and Applied Algorithmics Max Planck Institute for Informatics D-66123 Saarbrücken Germany NGFN - Courses in Practical DNA
Biclustering with δ-pcluster John Tantalo. 1. Introduction
 Biclustering with δ-pcluster John Tantalo 1. Introduction The subject of biclustering is chiefly concerned with locating submatrices of gene expression data that exhibit shared trends between genes. That
Biclustering with δ-pcluster John Tantalo 1. Introduction The subject of biclustering is chiefly concerned with locating submatrices of gene expression data that exhibit shared trends between genes. That
Unsupervised Learning and Clustering
 Unsupervised Learning and Clustering Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Spring 2008 CS 551, Spring 2008 c 2008, Selim Aksoy (Bilkent University)
Unsupervised Learning and Clustering Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Spring 2008 CS 551, Spring 2008 c 2008, Selim Aksoy (Bilkent University)
Unsupervised Learning and Clustering
 Unsupervised Learning and Clustering Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Spring 2009 CS 551, Spring 2009 c 2009, Selim Aksoy (Bilkent University)
Unsupervised Learning and Clustering Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Spring 2009 CS 551, Spring 2009 c 2009, Selim Aksoy (Bilkent University)
General Factorial Models
 In Chapter 8 in Oehlert STAT:5201 Week 9 - Lecture 2 1 / 34 It is possible to have many factors in a factorial experiment. In DDD we saw an example of a 3-factor study with ball size, height, and surface
In Chapter 8 in Oehlert STAT:5201 Week 9 - Lecture 2 1 / 34 It is possible to have many factors in a factorial experiment. In DDD we saw an example of a 3-factor study with ball size, height, and surface
Mining Deterministic Biclusters in Gene Expression Data
 Mining Deterministic Biclusters in Gene Expression Data Zonghong Zhang 1 Alvin Teo 1 BengChinOoi 1,2 Kian-Lee Tan 1,2 1 Department of Computer Science National University of Singapore 2 Singapore-MIT-Alliance
Mining Deterministic Biclusters in Gene Expression Data Zonghong Zhang 1 Alvin Teo 1 BengChinOoi 1,2 Kian-Lee Tan 1,2 1 Department of Computer Science National University of Singapore 2 Singapore-MIT-Alliance
DNA chips and other techniques measure the expression
 24 IEEE TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS, VOL. 1, NO. 1, JANUARY-MARCH 2004 Biclustering Algorithms for Biological Data Analysis: A Survey Sara C. Madeira and Arlindo L. Oliveira
24 IEEE TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS, VOL. 1, NO. 1, JANUARY-MARCH 2004 Biclustering Algorithms for Biological Data Analysis: A Survey Sara C. Madeira and Arlindo L. Oliveira
Mean Square Residue Biclustering with Missing Data and Row Inversions
 Mean Square Residue Biclustering with Missing Data and Row Inversions Stefan Gremalschi a, Gulsah Altun b, Irina Astrovskaya a, and Alexander Zelikovsky a a Department of Computer Science, Georgia State
Mean Square Residue Biclustering with Missing Data and Row Inversions Stefan Gremalschi a, Gulsah Altun b, Irina Astrovskaya a, and Alexander Zelikovsky a a Department of Computer Science, Georgia State
Gene expression & Clustering (Chapter 10)
 Gene expression & Clustering (Chapter 10) Determining gene function Sequence comparison tells us if a gene is similar to another gene, e.g., in a new species Dynamic programming Approximate pattern matching
Gene expression & Clustering (Chapter 10) Determining gene function Sequence comparison tells us if a gene is similar to another gene, e.g., in a new species Dynamic programming Approximate pattern matching
General Factorial Models
 In Chapter 8 in Oehlert STAT:5201 Week 9 - Lecture 1 1 / 31 It is possible to have many factors in a factorial experiment. We saw some three-way factorials earlier in the DDD book (HW 1 with 3 factors:
In Chapter 8 in Oehlert STAT:5201 Week 9 - Lecture 1 1 / 31 It is possible to have many factors in a factorial experiment. We saw some three-way factorials earlier in the DDD book (HW 1 with 3 factors:
Application of Hierarchical Clustering to Find Expression Modules in Cancer
 Application of Hierarchical Clustering to Find Expression Modules in Cancer T. M. Murali August 18, 2008 Innovative Application of Hierarchical Clustering A module map showing conditional activity of expression
Application of Hierarchical Clustering to Find Expression Modules in Cancer T. M. Murali August 18, 2008 Innovative Application of Hierarchical Clustering A module map showing conditional activity of expression
e-ccc-biclustering: Related work on biclustering algorithms for time series gene expression data
 : Related work on biclustering algorithms for time series gene expression data Sara C. Madeira 1,2,3, Arlindo L. Oliveira 1,2 1 Knowledge Discovery and Bioinformatics (KDBIO) group, INESC-ID, Lisbon, Portugal
: Related work on biclustering algorithms for time series gene expression data Sara C. Madeira 1,2,3, Arlindo L. Oliveira 1,2 1 Knowledge Discovery and Bioinformatics (KDBIO) group, INESC-ID, Lisbon, Portugal
9/29/13. Outline Data mining tasks. Clustering algorithms. Applications of clustering in biology
 9/9/ I9 Introduction to Bioinformatics, Clustering algorithms Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Outline Data mining tasks Predictive tasks vs descriptive tasks Example
9/9/ I9 Introduction to Bioinformatics, Clustering algorithms Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Outline Data mining tasks Predictive tasks vs descriptive tasks Example
Gene Clustering & Classification
 BINF, Introduction to Computational Biology Gene Clustering & Classification Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Introduction to Gene Clustering
BINF, Introduction to Computational Biology Gene Clustering & Classification Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Introduction to Gene Clustering
Unsupervised Learning : Clustering
 Unsupervised Learning : Clustering Things to be Addressed Traditional Learning Models. Cluster Analysis K-means Clustering Algorithm Drawbacks of traditional clustering algorithms. Clustering as a complex
Unsupervised Learning : Clustering Things to be Addressed Traditional Learning Models. Cluster Analysis K-means Clustering Algorithm Drawbacks of traditional clustering algorithms. Clustering as a complex
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 15: Microarray clustering http://compbio.pbworks.com/f/wood2.gif Some slides were adapted from Dr. Shaojie Zhang (University of Central Florida) Microarray
EECS730: Introduction to Bioinformatics Lecture 15: Microarray clustering http://compbio.pbworks.com/f/wood2.gif Some slides were adapted from Dr. Shaojie Zhang (University of Central Florida) Microarray
Statistical Pattern Recognition
 Statistical Pattern Recognition Features and Feature Selection Hamid R. Rabiee Jafar Muhammadi Spring 2012 http://ce.sharif.edu/courses/90-91/2/ce725-1/ Agenda Features and Patterns The Curse of Size and
Statistical Pattern Recognition Features and Feature Selection Hamid R. Rabiee Jafar Muhammadi Spring 2012 http://ce.sharif.edu/courses/90-91/2/ce725-1/ Agenda Features and Patterns The Curse of Size and
What to come. There will be a few more topics we will cover on supervised learning
 Summary so far Supervised learning learn to predict Continuous target regression; Categorical target classification Linear Regression Classification Discriminative models Perceptron (linear) Logistic regression
Summary so far Supervised learning learn to predict Continuous target regression; Categorical target classification Linear Regression Classification Discriminative models Perceptron (linear) Logistic regression
3. Cluster analysis Overview
 Université Laval Multivariate analysis - February 2006 1 3.1. Overview 3. Cluster analysis Clustering requires the recognition of discontinuous subsets in an environment that is sometimes discrete (as
Université Laval Multivariate analysis - February 2006 1 3.1. Overview 3. Cluster analysis Clustering requires the recognition of discontinuous subsets in an environment that is sometimes discrete (as
Recall the expression for the minimum significant difference (w) used in the Tukey fixed-range method for means separation:
 Topic 11. Unbalanced Designs [ST&D section 9.6, page 219; chapter 18] 11.1 Definition of missing data Accidents often result in loss of data. Crops are destroyed in some plots, plants and animals die,
Topic 11. Unbalanced Designs [ST&D section 9.6, page 219; chapter 18] 11.1 Definition of missing data Accidents often result in loss of data. Crops are destroyed in some plots, plants and animals die,
Based on Raymond J. Mooney s slides
 Instance Based Learning Based on Raymond J. Mooney s slides University of Texas at Austin 1 Example 2 Instance-Based Learning Unlike other learning algorithms, does not involve construction of an explicit
Instance Based Learning Based on Raymond J. Mooney s slides University of Texas at Austin 1 Example 2 Instance-Based Learning Unlike other learning algorithms, does not involve construction of an explicit
Using Machine Learning to Optimize Storage Systems
 Using Machine Learning to Optimize Storage Systems Dr. Kiran Gunnam 1 Outline 1. Overview 2. Building Flash Models using Logistic Regression. 3. Storage Object classification 4. Storage Allocation recommendation
Using Machine Learning to Optimize Storage Systems Dr. Kiran Gunnam 1 Outline 1. Overview 2. Building Flash Models using Logistic Regression. 3. Storage Object classification 4. Storage Allocation recommendation
Clustering and Visualisation of Data
 Clustering and Visualisation of Data Hiroshi Shimodaira January-March 28 Cluster analysis aims to partition a data set into meaningful or useful groups, based on distances between data points. In some
Clustering and Visualisation of Data Hiroshi Shimodaira January-March 28 Cluster analysis aims to partition a data set into meaningful or useful groups, based on distances between data points. In some
CSE 6242 A / CS 4803 DVA. Feb 12, Dimension Reduction. Guest Lecturer: Jaegul Choo
 CSE 6242 A / CS 4803 DVA Feb 12, 2013 Dimension Reduction Guest Lecturer: Jaegul Choo CSE 6242 A / CS 4803 DVA Feb 12, 2013 Dimension Reduction Guest Lecturer: Jaegul Choo Data is Too Big To Do Something..
CSE 6242 A / CS 4803 DVA Feb 12, 2013 Dimension Reduction Guest Lecturer: Jaegul Choo CSE 6242 A / CS 4803 DVA Feb 12, 2013 Dimension Reduction Guest Lecturer: Jaegul Choo Data is Too Big To Do Something..
CSE 255 Lecture 6. Data Mining and Predictive Analytics. Community Detection
 CSE 255 Lecture 6 Data Mining and Predictive Analytics Community Detection Dimensionality reduction Goal: take high-dimensional data, and describe it compactly using a small number of dimensions Assumption:
CSE 255 Lecture 6 Data Mining and Predictive Analytics Community Detection Dimensionality reduction Goal: take high-dimensional data, and describe it compactly using a small number of dimensions Assumption:
CS Introduction to Data Mining Instructor: Abdullah Mueen
 CS 591.03 Introduction to Data Mining Instructor: Abdullah Mueen LECTURE 8: ADVANCED CLUSTERING (FUZZY AND CO -CLUSTERING) Review: Basic Cluster Analysis Methods (Chap. 10) Cluster Analysis: Basic Concepts
CS 591.03 Introduction to Data Mining Instructor: Abdullah Mueen LECTURE 8: ADVANCED CLUSTERING (FUZZY AND CO -CLUSTERING) Review: Basic Cluster Analysis Methods (Chap. 10) Cluster Analysis: Basic Concepts
Lecture 5 Finding meaningful clusters in data. 5.1 Kleinberg s axiomatic framework for clustering
 CSE 291: Unsupervised learning Spring 2008 Lecture 5 Finding meaningful clusters in data So far we ve been in the vector quantization mindset, where we want to approximate a data set by a small number
CSE 291: Unsupervised learning Spring 2008 Lecture 5 Finding meaningful clusters in data So far we ve been in the vector quantization mindset, where we want to approximate a data set by a small number
Cluster Analysis. Mu-Chun Su. Department of Computer Science and Information Engineering National Central University 2003/3/11 1
 Cluster Analysis Mu-Chun Su Department of Computer Science and Information Engineering National Central University 2003/3/11 1 Introduction Cluster analysis is the formal study of algorithms and methods
Cluster Analysis Mu-Chun Su Department of Computer Science and Information Engineering National Central University 2003/3/11 1 Introduction Cluster analysis is the formal study of algorithms and methods
CSE 6242 A / CX 4242 DVA. March 6, Dimension Reduction. Guest Lecturer: Jaegul Choo
 CSE 6242 A / CX 4242 DVA March 6, 2014 Dimension Reduction Guest Lecturer: Jaegul Choo Data is Too Big To Analyze! Limited memory size! Data may not be fitted to the memory of your machine! Slow computation!
CSE 6242 A / CX 4242 DVA March 6, 2014 Dimension Reduction Guest Lecturer: Jaegul Choo Data is Too Big To Analyze! Limited memory size! Data may not be fitted to the memory of your machine! Slow computation!
Information Retrieval and Web Search Engines
 Information Retrieval and Web Search Engines Lecture 7: Document Clustering December 4th, 2014 Wolf-Tilo Balke and José Pinto Institut für Informationssysteme Technische Universität Braunschweig The Cluster
Information Retrieval and Web Search Engines Lecture 7: Document Clustering December 4th, 2014 Wolf-Tilo Balke and José Pinto Institut für Informationssysteme Technische Universität Braunschweig The Cluster
Chapter 2 Basic Structure of High-Dimensional Spaces
 Chapter 2 Basic Structure of High-Dimensional Spaces Data is naturally represented geometrically by associating each record with a point in the space spanned by the attributes. This idea, although simple,
Chapter 2 Basic Structure of High-Dimensional Spaces Data is naturally represented geometrically by associating each record with a point in the space spanned by the attributes. This idea, although simple,
Machine Learning and Data Mining. Clustering (1): Basics. Kalev Kask
 Machine Learning and Data Mining Clustering (1): Basics Kalev Kask Unsupervised learning Supervised learning Predict target value ( y ) given features ( x ) Unsupervised learning Understand patterns of
Machine Learning and Data Mining Clustering (1): Basics Kalev Kask Unsupervised learning Supervised learning Predict target value ( y ) given features ( x ) Unsupervised learning Understand patterns of
Clustering. CE-717: Machine Learning Sharif University of Technology Spring Soleymani
 Clustering CE-717: Machine Learning Sharif University of Technology Spring 2016 Soleymani Outline Clustering Definition Clustering main approaches Partitional (flat) Hierarchical Clustering validation
Clustering CE-717: Machine Learning Sharif University of Technology Spring 2016 Soleymani Outline Clustering Definition Clustering main approaches Partitional (flat) Hierarchical Clustering validation
CLUSTERING IN BIOINFORMATICS
 CLUSTERING IN BIOINFORMATICS CSE/BIMM/BENG 8 MAY 4, 0 OVERVIEW Define the clustering problem Motivation: gene expression and microarrays Types of clustering Clustering algorithms Other applications of
CLUSTERING IN BIOINFORMATICS CSE/BIMM/BENG 8 MAY 4, 0 OVERVIEW Define the clustering problem Motivation: gene expression and microarrays Types of clustering Clustering algorithms Other applications of
1. Mesh Coloring a.) Assign unique color to each polygon based on the polygon id.
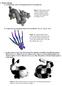 1. Mesh Coloring a.) Assign unique color to each polygon based on the polygon id. Figure 1: The dragon model is shown rendered using a coloring scheme based on coloring each triangle face according to
1. Mesh Coloring a.) Assign unique color to each polygon based on the polygon id. Figure 1: The dragon model is shown rendered using a coloring scheme based on coloring each triangle face according to
3. Cluster analysis Overview
 Université Laval Analyse multivariable - mars-avril 2008 1 3.1. Overview 3. Cluster analysis Clustering requires the recognition of discontinuous subsets in an environment that is sometimes discrete (as
Université Laval Analyse multivariable - mars-avril 2008 1 3.1. Overview 3. Cluster analysis Clustering requires the recognition of discontinuous subsets in an environment that is sometimes discrete (as
10. Clustering. Introduction to Bioinformatics Jarkko Salojärvi. Based on lecture slides by Samuel Kaski
 10. Clustering Introduction to Bioinformatics 30.9.2008 Jarkko Salojärvi Based on lecture slides by Samuel Kaski Definition of a cluster Typically either 1. A group of mutually similar samples, or 2. A
10. Clustering Introduction to Bioinformatics 30.9.2008 Jarkko Salojärvi Based on lecture slides by Samuel Kaski Definition of a cluster Typically either 1. A group of mutually similar samples, or 2. A
Part I. Hierarchical clustering. Hierarchical Clustering. Hierarchical clustering. Produces a set of nested clusters organized as a
 Week 9 Based in part on slides from textbook, slides of Susan Holmes Part I December 2, 2012 Hierarchical Clustering 1 / 1 Produces a set of nested clusters organized as a Hierarchical hierarchical clustering
Week 9 Based in part on slides from textbook, slides of Susan Holmes Part I December 2, 2012 Hierarchical Clustering 1 / 1 Produces a set of nested clusters organized as a Hierarchical hierarchical clustering
Variable Selection 6.783, Biomedical Decision Support
 6.783, Biomedical Decision Support (lrosasco@mit.edu) Department of Brain and Cognitive Science- MIT November 2, 2009 About this class Why selecting variables Approaches to variable selection Sparsity-based
6.783, Biomedical Decision Support (lrosasco@mit.edu) Department of Brain and Cognitive Science- MIT November 2, 2009 About this class Why selecting variables Approaches to variable selection Sparsity-based
2D/3D Geometric Transformations and Scene Graphs
 2D/3D Geometric Transformations and Scene Graphs Week 4 Acknowledgement: The course slides are adapted from the slides prepared by Steve Marschner of Cornell University 1 A little quick math background
2D/3D Geometric Transformations and Scene Graphs Week 4 Acknowledgement: The course slides are adapted from the slides prepared by Steve Marschner of Cornell University 1 A little quick math background
Hierarchical Clustering
 What is clustering Partitioning of a data set into subsets. A cluster is a group of relatively homogeneous cases or observations Hierarchical Clustering Mikhail Dozmorov Fall 2016 2/61 What is clustering
What is clustering Partitioning of a data set into subsets. A cluster is a group of relatively homogeneous cases or observations Hierarchical Clustering Mikhail Dozmorov Fall 2016 2/61 What is clustering
Statistical Pattern Recognition
 Statistical Pattern Recognition Features and Feature Selection Hamid R. Rabiee Jafar Muhammadi Spring 2013 http://ce.sharif.edu/courses/91-92/2/ce725-1/ Agenda Features and Patterns The Curse of Size and
Statistical Pattern Recognition Features and Feature Selection Hamid R. Rabiee Jafar Muhammadi Spring 2013 http://ce.sharif.edu/courses/91-92/2/ce725-1/ Agenda Features and Patterns The Curse of Size and
Clustering. Chapter 10 in Introduction to statistical learning
 Clustering Chapter 10 in Introduction to statistical learning 16 14 12 10 8 6 4 2 0 2 4 6 8 10 12 14 1 Clustering ² Clustering is the art of finding groups in data (Kaufman and Rousseeuw, 1990). ² What
Clustering Chapter 10 in Introduction to statistical learning 16 14 12 10 8 6 4 2 0 2 4 6 8 10 12 14 1 Clustering ² Clustering is the art of finding groups in data (Kaufman and Rousseeuw, 1990). ² What
10701 Machine Learning. Clustering
 171 Machine Learning Clustering What is Clustering? Organizing data into clusters such that there is high intra-cluster similarity low inter-cluster similarity Informally, finding natural groupings among
171 Machine Learning Clustering What is Clustering? Organizing data into clusters such that there is high intra-cluster similarity low inter-cluster similarity Informally, finding natural groupings among
Today. Lecture 4: Last time. The EM algorithm. We examine clustering in a little more detail; we went over it a somewhat quickly last time
 Today Lecture 4: We examine clustering in a little more detail; we went over it a somewhat quickly last time The CAD data will return and give us an opportunity to work with curves (!) We then examine
Today Lecture 4: We examine clustering in a little more detail; we went over it a somewhat quickly last time The CAD data will return and give us an opportunity to work with curves (!) We then examine
Statistical Pattern Recognition
 Statistical Pattern Recognition Features and Feature Selection Hamid R. Rabiee Jafar Muhammadi Spring 2014 http://ce.sharif.edu/courses/92-93/2/ce725-2/ Agenda Features and Patterns The Curse of Size and
Statistical Pattern Recognition Features and Feature Selection Hamid R. Rabiee Jafar Muhammadi Spring 2014 http://ce.sharif.edu/courses/92-93/2/ce725-2/ Agenda Features and Patterns The Curse of Size and
MSA220 - Statistical Learning for Big Data
 MSA220 - Statistical Learning for Big Data Lecture 13 Rebecka Jörnsten Mathematical Sciences University of Gothenburg and Chalmers University of Technology Clustering Explorative analysis - finding groups
MSA220 - Statistical Learning for Big Data Lecture 13 Rebecka Jörnsten Mathematical Sciences University of Gothenburg and Chalmers University of Technology Clustering Explorative analysis - finding groups
Unsupervised Learning
 Unsupervised Learning Unsupervised learning Until now, we have assumed our training samples are labeled by their category membership. Methods that use labeled samples are said to be supervised. However,
Unsupervised Learning Unsupervised learning Until now, we have assumed our training samples are labeled by their category membership. Methods that use labeled samples are said to be supervised. However,
Lecture 3: Linear Classification
 Lecture 3: Linear Classification Roger Grosse 1 Introduction Last week, we saw an example of a learning task called regression. There, the goal was to predict a scalar-valued target from a set of features.
Lecture 3: Linear Classification Roger Grosse 1 Introduction Last week, we saw an example of a learning task called regression. There, the goal was to predict a scalar-valued target from a set of features.
1 Homophily and assortative mixing
 1 Homophily and assortative mixing Networks, and particularly social networks, often exhibit a property called homophily or assortative mixing, which simply means that the attributes of vertices correlate
1 Homophily and assortative mixing Networks, and particularly social networks, often exhibit a property called homophily or assortative mixing, which simply means that the attributes of vertices correlate
TELCOM2125: Network Science and Analysis
 School of Information Sciences University of Pittsburgh TELCOM2125: Network Science and Analysis Konstantinos Pelechrinis Spring 2015 2 Part 4: Dividing Networks into Clusters The problem l Graph partitioning
School of Information Sciences University of Pittsburgh TELCOM2125: Network Science and Analysis Konstantinos Pelechrinis Spring 2015 2 Part 4: Dividing Networks into Clusters The problem l Graph partitioning
Clustering. Informal goal. General types of clustering. Applications: Clustering in information search and analysis. Example applications in search
 Informal goal Clustering Given set of objects and measure of similarity between them, group similar objects together What mean by similar? What is good grouping? Computation time / quality tradeoff 1 2
Informal goal Clustering Given set of objects and measure of similarity between them, group similar objects together What mean by similar? What is good grouping? Computation time / quality tradeoff 1 2
Enhancing Clustering Results In Hierarchical Approach By Mvs Measures
 International Journal of Engineering Research and Development e-issn: 2278-067X, p-issn: 2278-800X, www.ijerd.com Volume 10, Issue 6 (June 2014), PP.25-30 Enhancing Clustering Results In Hierarchical Approach
International Journal of Engineering Research and Development e-issn: 2278-067X, p-issn: 2278-800X, www.ijerd.com Volume 10, Issue 6 (June 2014), PP.25-30 Enhancing Clustering Results In Hierarchical Approach
Information Retrieval and Web Search Engines
 Information Retrieval and Web Search Engines Lecture 7: Document Clustering May 25, 2011 Wolf-Tilo Balke and Joachim Selke Institut für Informationssysteme Technische Universität Braunschweig Homework
Information Retrieval and Web Search Engines Lecture 7: Document Clustering May 25, 2011 Wolf-Tilo Balke and Joachim Selke Institut für Informationssysteme Technische Universität Braunschweig Homework
Clustering. Supervised vs. Unsupervised Learning
 Clustering Supervised vs. Unsupervised Learning So far we have assumed that the training samples used to design the classifier were labeled by their class membership (supervised learning) We assume now
Clustering Supervised vs. Unsupervised Learning So far we have assumed that the training samples used to design the classifier were labeled by their class membership (supervised learning) We assume now
Classification and Regression Trees
 Classification and Regression Trees David S. Rosenberg New York University April 3, 2018 David S. Rosenberg (New York University) DS-GA 1003 / CSCI-GA 2567 April 3, 2018 1 / 51 Contents 1 Trees 2 Regression
Classification and Regression Trees David S. Rosenberg New York University April 3, 2018 David S. Rosenberg (New York University) DS-GA 1003 / CSCI-GA 2567 April 3, 2018 1 / 51 Contents 1 Trees 2 Regression
Clustering Techniques
 Clustering Techniques Bioinformatics: Issues and Algorithms CSE 308-408 Fall 2007 Lecture 16 Lopresti Fall 2007 Lecture 16-1 - Administrative notes Your final project / paper proposal is due on Friday,
Clustering Techniques Bioinformatics: Issues and Algorithms CSE 308-408 Fall 2007 Lecture 16 Lopresti Fall 2007 Lecture 16-1 - Administrative notes Your final project / paper proposal is due on Friday,
(Refer Slide Time 6:48)
 Digital Circuits and Systems Prof. S. Srinivasan Department of Electrical Engineering Indian Institute of Technology Madras Lecture - 8 Karnaugh Map Minimization using Maxterms We have been taking about
Digital Circuits and Systems Prof. S. Srinivasan Department of Electrical Engineering Indian Institute of Technology Madras Lecture - 8 Karnaugh Map Minimization using Maxterms We have been taking about
Genotype x Environmental Analysis with R for Windows
 Genotype x Environmental Analysis with R for Windows Biometrics and Statistics Unit Angela Pacheco CIMMYT,Int. 23-24 Junio 2015 About GEI In agricultural experimentation, a large number of genotypes are
Genotype x Environmental Analysis with R for Windows Biometrics and Statistics Unit Angela Pacheco CIMMYT,Int. 23-24 Junio 2015 About GEI In agricultural experimentation, a large number of genotypes are
Mixture Models and the EM Algorithm
 Mixture Models and the EM Algorithm Padhraic Smyth, Department of Computer Science University of California, Irvine c 2017 1 Finite Mixture Models Say we have a data set D = {x 1,..., x N } where x i is
Mixture Models and the EM Algorithm Padhraic Smyth, Department of Computer Science University of California, Irvine c 2017 1 Finite Mixture Models Say we have a data set D = {x 1,..., x N } where x i is
Variables and Data Representation
 You will recall that a computer program is a set of instructions that tell a computer how to transform a given set of input into a specific output. Any program, procedural, event driven or object oriented
You will recall that a computer program is a set of instructions that tell a computer how to transform a given set of input into a specific output. Any program, procedural, event driven or object oriented
Comparisons and validation of statistical clustering techniques for microarray gene expression data. Outline. Microarrays.
 Comparisons and validation of statistical clustering techniques for microarray gene expression data Susmita Datta and Somnath Datta Presented by: Jenni Dietrich Assisted by: Jeffrey Kidd and Kristin Wheeler
Comparisons and validation of statistical clustering techniques for microarray gene expression data Susmita Datta and Somnath Datta Presented by: Jenni Dietrich Assisted by: Jeffrey Kidd and Kristin Wheeler
Small Survey on Perfect Graphs
 Small Survey on Perfect Graphs Michele Alberti ENS Lyon December 8, 2010 Abstract This is a small survey on the exciting world of Perfect Graphs. We will see when a graph is perfect and which are families
Small Survey on Perfect Graphs Michele Alberti ENS Lyon December 8, 2010 Abstract This is a small survey on the exciting world of Perfect Graphs. We will see when a graph is perfect and which are families
Supplementary text S6 Comparison studies on simulated data
 Supplementary text S Comparison studies on simulated data Peter Langfelder, Rui Luo, Michael C. Oldham, and Steve Horvath Corresponding author: shorvath@mednet.ucla.edu Overview In this document we illustrate
Supplementary text S Comparison studies on simulated data Peter Langfelder, Rui Luo, Michael C. Oldham, and Steve Horvath Corresponding author: shorvath@mednet.ucla.edu Overview In this document we illustrate
Cluster Analysis. Angela Montanari and Laura Anderlucci
 Cluster Analysis Angela Montanari and Laura Anderlucci 1 Introduction Clustering a set of n objects into k groups is usually moved by the aim of identifying internally homogenous groups according to a
Cluster Analysis Angela Montanari and Laura Anderlucci 1 Introduction Clustering a set of n objects into k groups is usually moved by the aim of identifying internally homogenous groups according to a
Machine Learning (BSMC-GA 4439) Wenke Liu
 Machine Learning (BSMC-GA 4439) Wenke Liu 01-25-2018 Outline Background Defining proximity Clustering methods Determining number of clusters Other approaches Cluster analysis as unsupervised Learning Unsupervised
Machine Learning (BSMC-GA 4439) Wenke Liu 01-25-2018 Outline Background Defining proximity Clustering methods Determining number of clusters Other approaches Cluster analysis as unsupervised Learning Unsupervised
CPSC 340: Machine Learning and Data Mining. Deep Learning Fall 2018
 CPSC 340: Machine Learning and Data Mining Deep Learning Fall 2018 Last Time: Multi-Dimensional Scaling Multi-dimensional scaling (MDS): Non-parametric visualization: directly optimize the z i locations.
CPSC 340: Machine Learning and Data Mining Deep Learning Fall 2018 Last Time: Multi-Dimensional Scaling Multi-dimensional scaling (MDS): Non-parametric visualization: directly optimize the z i locations.
Fly wing length data Sokal and Rohlf Box 10.1 Ch13.xls. on chalk board
 Model Based Statistics in Biology. Part IV. The General Linear Model. Multiple Explanatory Variables. Chapter 13.6 Nested Factors (Hierarchical ANOVA ReCap. Part I (Chapters 1,2,3,4), Part II (Ch 5, 6,
Model Based Statistics in Biology. Part IV. The General Linear Model. Multiple Explanatory Variables. Chapter 13.6 Nested Factors (Hierarchical ANOVA ReCap. Part I (Chapters 1,2,3,4), Part II (Ch 5, 6,
Parametric curves. Brian Curless CSE 457 Spring 2016
 Parametric curves Brian Curless CSE 457 Spring 2016 1 Reading Required: Angel 10.1-10.3, 10.5.2, 10.6-10.7, 10.9 Optional Bartels, Beatty, and Barsky. An Introduction to Splines for use in Computer Graphics
Parametric curves Brian Curless CSE 457 Spring 2016 1 Reading Required: Angel 10.1-10.3, 10.5.2, 10.6-10.7, 10.9 Optional Bartels, Beatty, and Barsky. An Introduction to Splines for use in Computer Graphics
CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS
 CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS CHAPTER 4 CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS 4.1 Introduction Optical character recognition is one of
CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS CHAPTER 4 CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS 4.1 Introduction Optical character recognition is one of
Lecture 19: Generative Adversarial Networks
 Lecture 19: Generative Adversarial Networks Roger Grosse 1 Introduction Generative modeling is a type of machine learning where the aim is to model the distribution that a given set of data (e.g. images,
Lecture 19: Generative Adversarial Networks Roger Grosse 1 Introduction Generative modeling is a type of machine learning where the aim is to model the distribution that a given set of data (e.g. images,
Cluster Analysis. Prof. Thomas B. Fomby Department of Economics Southern Methodist University Dallas, TX April 2008 April 2010
 Cluster Analysis Prof. Thomas B. Fomby Department of Economics Southern Methodist University Dallas, TX 7575 April 008 April 010 Cluster Analysis, sometimes called data segmentation or customer segmentation,
Cluster Analysis Prof. Thomas B. Fomby Department of Economics Southern Methodist University Dallas, TX 7575 April 008 April 010 Cluster Analysis, sometimes called data segmentation or customer segmentation,
Parametric curves. Reading. Curves before computers. Mathematical curve representation. CSE 457 Winter Required:
 Reading Required: Angel 10.1-10.3, 10.5.2, 10.6-10.7, 10.9 Parametric curves CSE 457 Winter 2014 Optional Bartels, Beatty, and Barsky. An Introduction to Splines for use in Computer Graphics and Geometric
Reading Required: Angel 10.1-10.3, 10.5.2, 10.6-10.7, 10.9 Parametric curves CSE 457 Winter 2014 Optional Bartels, Beatty, and Barsky. An Introduction to Splines for use in Computer Graphics and Geometric
Regularization and model selection
 CS229 Lecture notes Andrew Ng Part VI Regularization and model selection Suppose we are trying select among several different models for a learning problem. For instance, we might be using a polynomial
CS229 Lecture notes Andrew Ng Part VI Regularization and model selection Suppose we are trying select among several different models for a learning problem. For instance, we might be using a polynomial
Slides for Data Mining by I. H. Witten and E. Frank
 Slides for Data Mining by I. H. Witten and E. Frank 7 Engineering the input and output Attribute selection Scheme-independent, scheme-specific Attribute discretization Unsupervised, supervised, error-
Slides for Data Mining by I. H. Witten and E. Frank 7 Engineering the input and output Attribute selection Scheme-independent, scheme-specific Attribute discretization Unsupervised, supervised, error-
Lecture 25: Review I
 Lecture 25: Review I Reading: Up to chapter 5 in ISLR. STATS 202: Data mining and analysis Jonathan Taylor 1 / 18 Unsupervised learning In unsupervised learning, all the variables are on equal standing,
Lecture 25: Review I Reading: Up to chapter 5 in ISLR. STATS 202: Data mining and analysis Jonathan Taylor 1 / 18 Unsupervised learning In unsupervised learning, all the variables are on equal standing,
Project and Production Management Prof. Arun Kanda Department of Mechanical Engineering Indian Institute of Technology, Delhi
 Project and Production Management Prof. Arun Kanda Department of Mechanical Engineering Indian Institute of Technology, Delhi Lecture - 8 Consistency and Redundancy in Project networks In today s lecture
Project and Production Management Prof. Arun Kanda Department of Mechanical Engineering Indian Institute of Technology, Delhi Lecture - 8 Consistency and Redundancy in Project networks In today s lecture
Clustering: Classic Methods and Modern Views
 Clustering: Classic Methods and Modern Views Marina Meilă University of Washington mmp@stat.washington.edu June 22, 2015 Lorentz Center Workshop on Clusters, Games and Axioms Outline Paradigms for clustering
Clustering: Classic Methods and Modern Views Marina Meilă University of Washington mmp@stat.washington.edu June 22, 2015 Lorentz Center Workshop on Clusters, Games and Axioms Outline Paradigms for clustering
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT MD Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org 19
GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT MD Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org 19
Network Clustering. Balabhaskar Balasundaram, Sergiy Butenko
 Network Clustering Balabhaskar Balasundaram, Sergiy Butenko Department of Industrial & Systems Engineering Texas A&M University College Station, Texas 77843, USA. Introduction Clustering can be loosely
Network Clustering Balabhaskar Balasundaram, Sergiy Butenko Department of Industrial & Systems Engineering Texas A&M University College Station, Texas 77843, USA. Introduction Clustering can be loosely
Lecture 3: February Local Alignment: The Smith-Waterman Algorithm
 CSCI1820: Sequence Alignment Spring 2017 Lecture 3: February 7 Lecturer: Sorin Istrail Scribe: Pranavan Chanthrakumar Note: LaTeX template courtesy of UC Berkeley EECS dept. Notes are also adapted from
CSCI1820: Sequence Alignment Spring 2017 Lecture 3: February 7 Lecturer: Sorin Istrail Scribe: Pranavan Chanthrakumar Note: LaTeX template courtesy of UC Berkeley EECS dept. Notes are also adapted from
Classification. 1 o Semestre 2007/2008
 Classification Departamento de Engenharia Informática Instituto Superior Técnico 1 o Semestre 2007/2008 Slides baseados nos slides oficiais do livro Mining the Web c Soumen Chakrabarti. Outline 1 2 3 Single-Class
Classification Departamento de Engenharia Informática Instituto Superior Técnico 1 o Semestre 2007/2008 Slides baseados nos slides oficiais do livro Mining the Web c Soumen Chakrabarti. Outline 1 2 3 Single-Class
Multivariate Analysis
 Multivariate Analysis Cluster Analysis Prof. Dr. Anselmo E de Oliveira anselmo.quimica.ufg.br anselmo.disciplinas@gmail.com Unsupervised Learning Cluster Analysis Natural grouping Patterns in the data
Multivariate Analysis Cluster Analysis Prof. Dr. Anselmo E de Oliveira anselmo.quimica.ufg.br anselmo.disciplinas@gmail.com Unsupervised Learning Cluster Analysis Natural grouping Patterns in the data
BIMAX. Lecture 11: December 31, Introduction Model An Incremental Algorithm.
 Analysis of DNA Chips and Gene Networks Spring Semester, 2009 Lecturer: Ron Shamir BIMAX 11.1 Introduction. Lecture 11: December 31, 2009 Scribe: Boris Kostenko In the course we have already seen different
Analysis of DNA Chips and Gene Networks Spring Semester, 2009 Lecturer: Ron Shamir BIMAX 11.1 Introduction. Lecture 11: December 31, 2009 Scribe: Boris Kostenko In the course we have already seen different
Machine Learning Techniques for Data Mining
 Machine Learning Techniques for Data Mining Eibe Frank University of Waikato New Zealand 10/25/2000 1 PART V Credibility: Evaluating what s been learned 10/25/2000 2 Evaluation: the key to success How
Machine Learning Techniques for Data Mining Eibe Frank University of Waikato New Zealand 10/25/2000 1 PART V Credibility: Evaluating what s been learned 10/25/2000 2 Evaluation: the key to success How
Computational Genomics and Molecular Biology, Fall
 Computational Genomics and Molecular Biology, Fall 2015 1 Sequence Alignment Dannie Durand Pairwise Sequence Alignment The goal of pairwise sequence alignment is to establish a correspondence between the
Computational Genomics and Molecular Biology, Fall 2015 1 Sequence Alignment Dannie Durand Pairwise Sequence Alignment The goal of pairwise sequence alignment is to establish a correspondence between the
Application of fuzzy set theory in image analysis. Nataša Sladoje Centre for Image Analysis
 Application of fuzzy set theory in image analysis Nataša Sladoje Centre for Image Analysis Our topics for today Crisp vs fuzzy Fuzzy sets and fuzzy membership functions Fuzzy set operators Approximate
Application of fuzzy set theory in image analysis Nataša Sladoje Centre for Image Analysis Our topics for today Crisp vs fuzzy Fuzzy sets and fuzzy membership functions Fuzzy set operators Approximate
Clustering. Robert M. Haralick. Computer Science, Graduate Center City University of New York
 Clustering Robert M. Haralick Computer Science, Graduate Center City University of New York Outline K-means 1 K-means 2 3 4 5 Clustering K-means The purpose of clustering is to determine the similarity
Clustering Robert M. Haralick Computer Science, Graduate Center City University of New York Outline K-means 1 K-means 2 3 4 5 Clustering K-means The purpose of clustering is to determine the similarity
Exploratory Analysis: Clustering
 Exploratory Analysis: Clustering (some material taken or adapted from slides by Hinrich Schutze) Heejun Kim June 26, 2018 Clustering objective Grouping documents or instances into subsets or clusters Documents
Exploratory Analysis: Clustering (some material taken or adapted from slides by Hinrich Schutze) Heejun Kim June 26, 2018 Clustering objective Grouping documents or instances into subsets or clusters Documents
One way ANOVA when the data are not normally distributed (The Kruskal-Wallis test).
 One way ANOVA when the data are not normally distributed (The Kruskal-Wallis test). Suppose you have a one way design, and want to do an ANOVA, but discover that your data are seriously not normal? Just
One way ANOVA when the data are not normally distributed (The Kruskal-Wallis test). Suppose you have a one way design, and want to do an ANOVA, but discover that your data are seriously not normal? Just
Computer Graphics Prof. Sukhendu Das Dept. of Computer Science and Engineering Indian Institute of Technology, Madras Lecture - 14
 Computer Graphics Prof. Sukhendu Das Dept. of Computer Science and Engineering Indian Institute of Technology, Madras Lecture - 14 Scan Converting Lines, Circles and Ellipses Hello everybody, welcome again
Computer Graphics Prof. Sukhendu Das Dept. of Computer Science and Engineering Indian Institute of Technology, Madras Lecture - 14 Scan Converting Lines, Circles and Ellipses Hello everybody, welcome again
The Generalized Topological Overlap Matrix in Biological Network Analysis
 The Generalized Topological Overlap Matrix in Biological Network Analysis Andy Yip, Steve Horvath Email: shorvath@mednet.ucla.edu Depts Human Genetics and Biostatistics, University of California, Los Angeles
The Generalized Topological Overlap Matrix in Biological Network Analysis Andy Yip, Steve Horvath Email: shorvath@mednet.ucla.edu Depts Human Genetics and Biostatistics, University of California, Los Angeles
CPSC 340: Machine Learning and Data Mining. Probabilistic Classification Fall 2017
 CPSC 340: Machine Learning and Data Mining Probabilistic Classification Fall 2017 Admin Assignment 0 is due tonight: you should be almost done. 1 late day to hand it in Monday, 2 late days for Wednesday.
CPSC 340: Machine Learning and Data Mining Probabilistic Classification Fall 2017 Admin Assignment 0 is due tonight: you should be almost done. 1 late day to hand it in Monday, 2 late days for Wednesday.
Math background. 2D Geometric Transformations. Implicit representations. Explicit representations. Read: CS 4620 Lecture 6
 Math background 2D Geometric Transformations CS 4620 Lecture 6 Read: Chapter 2: Miscellaneous Math Chapter 5: Linear Algebra Notation for sets, functions, mappings Linear transformations Matrices Matrix-vector
Math background 2D Geometric Transformations CS 4620 Lecture 6 Read: Chapter 2: Miscellaneous Math Chapter 5: Linear Algebra Notation for sets, functions, mappings Linear transformations Matrices Matrix-vector
