The quantitative analysis of interactions takes bioinformatics to the next higher dimension: we go from 1D to 2D with graph theory.
|
|
|
- Robert McDowell
- 5 years ago
- Views:
Transcription
1 1
2 The human protein-protein interaction network of aging-associated genes. A total of 261 aging-associated genes were assembled using the GenAge Human Database. Protein-protein interactions of the human interactome were collected from the 8.0 version of the STRING database using physical contacts only. The network was visualized using the Cytoscape program. The degree (number of neighbors) of nodes is represented by the size of the circle and the font. Note the high number of signaling pathway proteins among hubs (nodes with degrees - and therefore size - much greater than average), exemplified by the MAPK/ERK and PI3K/AKT proteins. Figure and caption from: Simkó G, Gyurkó D, Veres DV, Nánási T, and Peter Csermely P (2009) Network strategies to understand the aging process and help age-related drug design. Genome Med. 2009; 1(9): 90 Plots like these have become ubiquitous in the litearture. They have an undeniable aesthetic quality, but the information about biological processes is limited. The authors mention the high number of signalling pathway proteins among the nodes of large degree, but they have not (i) mentioned how a signalling pathway protein is defined, (ii) described in detail how those aging-related proteins were selected in the first place, (iii) compared this fraction against the fraction of signalling pathway proteins among all genes, (iv) quantified their finding, (v) emphasized this class of proteins by colour in the plot. Plotting networks is in general not a good way to analyze them: the inference process has to go the other way around: analyze the network with computational means, then visualize the results in a plot. This plot may or may not be a graph similar to the one above, in this particular case a simple boxplot of node-degree by GO biological process category would have been more effective. Network plots show interactions, but no hypothesis about those interactions has been suggested that the graph could help us visualize. Let us thus discuss the backgrounds of graph theory and useful measures that we can apply towards biological inference from relationship data. 2
3 The quantitative analysis of interactions takes bioinformatics to the next higher dimension: we go from 1D to 2D with graph theory. 3
4 Definitions. 4
5 5
6 6
7 7
8 8
9 If we have a directed graph, a node may have incoming edges and outgoing edges. In that case, we distinguish between in-degree and out-degree of the node. We can conceptualize that nodes with a high degree lie in the centre of a network, and that nodes with degree 1 constitute the boundary of the network. Therefore the degree of a node is also a topological measure of the graph, i.e. it describes the contribution of nodes to the overall shape of the graph: we call this interpretation degree centrality. 9
10 Degree distributions give us a way to reason about the circumstances that could have produced a network of interactions in the real world. Barabasi and Albert showed scale-free properties for movie-actor networks, pages in the WWW, and the electric power grid. 10
11 It was soon appreciated, that many biological networks also have scale-free properties. This is not trivial, and begs the question what the WWW and metabolic networks could have in common with power-grid layouts and developmental signalling pathways. 11
12 12
13 13
14 Note that the degree-distribution histograms (frequency is on a log-scale) are very characteristic of the generative process. 14
15 The problem here is that it is not obvious why protein-protein interactions should be subject to preferential attachment. Nor is it entirely clear whether actual interaction graphs are indeed scale-free. I do not agree with Barabasi and Albert that the model indicates that growth and preferential attachment play an important role in network development. This would be confusing correlation with causation. They have indeed shown that a model, based on preferential attachment, can reproduce characteristics of many networks. However we must keep an open mind about whether there are other mechanisms that also could give rise to scale-free properties. And indeed there are. Alternative models include: the copy model that creates a scale-free distribution by adding new nodes through copying a fraction of the links of an existing node. Rewiring of random networks towards game-theoretic optimal objective functions also creates scale-free networks. Hierarchical network models are scale-free, as are hyperbolic geometric graphs. All of these have much more straightforward biological analogies than preferential attachment. Besides, the actual networks of biology are not necessarily formed by optimization with a single mechanism towards a common objective function, but are the result of a messy, stochastic, vaguely conserved process of achieving sufficiency of purpose. See also: 15
16 16
17 17
18 18
19 Besides degree distributions, there are several other approaches to the quantitative analysis of biological networks. 19
20 How many diameters does this graph have? I say: six. 20
21 Dijkstra s algortihm finds a shortest path between two nodes. (i) Suppose we want to find the shortest path between the red ( origin ) and the green ( target ) node. (ii) We set the distance of the origin to 0, all other distances to. The origin is our current node. (iii) Next we collect all neighbors of the current node, and set their distance to one-more than the distance of the current node. (iv) We repeat what we did in (iii) by considering all neighbors in turn. However, we never consider a node that we visited previously, i.e. we include only nodes whose distance is. (v) We repeat what we did in (iii) one more time, This is a loop. Lo-and-behold, this time we encounter the target node. (vi) From the target node, we look for a node whose distance is one-less and add it to the shortest path. We repeat this, always going to a node closer to the origin, and adding that to the path. This is called backtracking. Once we have reached the origin, the shortest path is defined. This is generally considered an O( V 2 ) algorithm, but the implementation based on a min-priority queue implemented by a Fibonacci heap runs in O( E + V \log V ) Fredman ML and Tarjan RE. (1984). Fibonacci heaps and their uses in improved network optimization algorithms. 25th Annual Symposium on Foundations of Computer Science. IEEE. pp
22 22
23 Note that in general, betweeness centrality and degree centrality go in the same direction nodes with high degree tend to have high C b values. But the relationship is not absolute e.g. the second highest C b value, 94, is in a node with degree 3, like two other nodes who only have a C b of 56. This node also has a higher C b than the node with C b 73, which has a degree of 4. Stress centrality is a related concept: the stress centrality of a node w is the number of shortest paths passing through w. 23
24 24
25 25
26 26
27 27
V 1 Introduction! Mon, Oct 15, 2012! Bioinformatics 3 Volkhard Helms!
 V 1 Introduction! Mon, Oct 15, 2012! Bioinformatics 3 Volkhard Helms! How Does a Cell Work?! A cell is a crowded environment! => many different proteins,! metabolites, compartments,! On a microscopic level!
V 1 Introduction! Mon, Oct 15, 2012! Bioinformatics 3 Volkhard Helms! How Does a Cell Work?! A cell is a crowded environment! => many different proteins,! metabolites, compartments,! On a microscopic level!
Example for calculation of clustering coefficient Node N 1 has 8 neighbors (red arrows) There are 12 connectivities among neighbors (blue arrows)
 Example for calculation of clustering coefficient Node N 1 has 8 neighbors (red arrows) There are 12 connectivities among neighbors (blue arrows) Average clustering coefficient of a graph Overall measure
Example for calculation of clustering coefficient Node N 1 has 8 neighbors (red arrows) There are 12 connectivities among neighbors (blue arrows) Average clustering coefficient of a graph Overall measure
Advanced Algorithms and Models for Computational Biology -- a machine learning approach
 Advanced Algorithms and Models for Computational Biology -- a machine learning approach Biological Networks & Network Evolution Eric Xing Lecture 22, April 10, 2006 Reading: Molecular Networks Interaction
Advanced Algorithms and Models for Computational Biology -- a machine learning approach Biological Networks & Network Evolution Eric Xing Lecture 22, April 10, 2006 Reading: Molecular Networks Interaction
Network Thinking. Complexity: A Guided Tour, Chapters 15-16
 Network Thinking Complexity: A Guided Tour, Chapters 15-16 Neural Network (C. Elegans) http://gephi.org/wp-content/uploads/2008/12/screenshot-celegans.png Food Web http://1.bp.blogspot.com/_vifbm3t8bou/sbhzqbchiei/aaaaaaaaaxk/rsc-pj45avc/
Network Thinking Complexity: A Guided Tour, Chapters 15-16 Neural Network (C. Elegans) http://gephi.org/wp-content/uploads/2008/12/screenshot-celegans.png Food Web http://1.bp.blogspot.com/_vifbm3t8bou/sbhzqbchiei/aaaaaaaaaxk/rsc-pj45avc/
An Exploratory Journey Into Network Analysis A Gentle Introduction to Network Science and Graph Visualization
 An Exploratory Journey Into Network Analysis A Gentle Introduction to Network Science and Graph Visualization Pedro Ribeiro (DCC/FCUP & CRACS/INESC-TEC) Part 1 Motivation and emergence of Network Science
An Exploratory Journey Into Network Analysis A Gentle Introduction to Network Science and Graph Visualization Pedro Ribeiro (DCC/FCUP & CRACS/INESC-TEC) Part 1 Motivation and emergence of Network Science
V 2 Clusters, Dijkstra, and Graph Layout
 Bioinformatics 3 V 2 Clusters, Dijkstra, and Graph Layout Mon, Oct 31, 2016 Graph Basics A graph G is an ordered pair (V, E) of a set V of vertices and a set E of edges. Degree distribution P(k) Random
Bioinformatics 3 V 2 Clusters, Dijkstra, and Graph Layout Mon, Oct 31, 2016 Graph Basics A graph G is an ordered pair (V, E) of a set V of vertices and a set E of edges. Degree distribution P(k) Random
Network analysis. Martina Kutmon Department of Bioinformatics Maastricht University
 Network analysis Martina Kutmon Department of Bioinformatics Maastricht University What's gonna happen today? Network Analysis Introduction Quiz Hands-on session ConsensusPathDB interaction database Outline
Network analysis Martina Kutmon Department of Bioinformatics Maastricht University What's gonna happen today? Network Analysis Introduction Quiz Hands-on session ConsensusPathDB interaction database Outline
Nick Hamilton Institute for Molecular Bioscience. Essential Graph Theory for Biologists. Image: Matt Moores, The Visible Cell
 Nick Hamilton Institute for Molecular Bioscience Essential Graph Theory for Biologists Image: Matt Moores, The Visible Cell Outline Core definitions Which are the most important bits? What happens when
Nick Hamilton Institute for Molecular Bioscience Essential Graph Theory for Biologists Image: Matt Moores, The Visible Cell Outline Core definitions Which are the most important bits? What happens when
Cytoscape:un approccio sistematico all'analisi dei dati sperimentali
 Cytoscape:un approccio sistematico all'analisi dei dati sperimentali Michele Petterlini michele@petterlini.it Università di Verona Centro di BioMedicina Computazionale The network Network: a graph representation
Cytoscape:un approccio sistematico all'analisi dei dati sperimentali Michele Petterlini michele@petterlini.it Università di Verona Centro di BioMedicina Computazionale The network Network: a graph representation
Case Studies in Complex Networks
 Case Studies in Complex Networks Introduction to Scientific Modeling CS 365 George Bezerra 08/27/2012 The origin of graph theory Königsberg bridge problem Leonard Euler (1707-1783) The Königsberg Bridge
Case Studies in Complex Networks Introduction to Scientific Modeling CS 365 George Bezerra 08/27/2012 The origin of graph theory Königsberg bridge problem Leonard Euler (1707-1783) The Königsberg Bridge
Graph-theoretic Properties of Networks
 Graph-theoretic Properties of Networks Bioinformatics: Sequence Analysis COMP 571 - Spring 2015 Luay Nakhleh, Rice University Graphs A graph is a set of vertices, or nodes, and edges that connect pairs
Graph-theoretic Properties of Networks Bioinformatics: Sequence Analysis COMP 571 - Spring 2015 Luay Nakhleh, Rice University Graphs A graph is a set of vertices, or nodes, and edges that connect pairs
Biological Networks Analysis
 Biological Networks Analysis Introduction and Dijkstra s algorithm Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The clustering problem: partition genes into distinct
Biological Networks Analysis Introduction and Dijkstra s algorithm Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The clustering problem: partition genes into distinct
Properties of Biological Networks
 Properties of Biological Networks presented by: Ola Hamud June 12, 2013 Supervisor: Prof. Ron Pinter Based on: NETWORK BIOLOGY: UNDERSTANDING THE CELL S FUNCTIONAL ORGANIZATION By Albert-László Barabási
Properties of Biological Networks presented by: Ola Hamud June 12, 2013 Supervisor: Prof. Ron Pinter Based on: NETWORK BIOLOGY: UNDERSTANDING THE CELL S FUNCTIONAL ORGANIZATION By Albert-László Barabási
Wednesday, March 8, Complex Networks. Presenter: Jirakhom Ruttanavakul. CS 790R, University of Nevada, Reno
 Wednesday, March 8, 2006 Complex Networks Presenter: Jirakhom Ruttanavakul CS 790R, University of Nevada, Reno Presented Papers Emergence of scaling in random networks, Barabási & Bonabeau (2003) Scale-free
Wednesday, March 8, 2006 Complex Networks Presenter: Jirakhom Ruttanavakul CS 790R, University of Nevada, Reno Presented Papers Emergence of scaling in random networks, Barabási & Bonabeau (2003) Scale-free
Basics of Network Analysis
 Basics of Network Analysis Hiroki Sayama sayama@binghamton.edu Graph = Network G(V, E): graph (network) V: vertices (nodes), E: edges (links) 1 Nodes = 1, 2, 3, 4, 5 2 3 Links = 12, 13, 15, 23,
Basics of Network Analysis Hiroki Sayama sayama@binghamton.edu Graph = Network G(V, E): graph (network) V: vertices (nodes), E: edges (links) 1 Nodes = 1, 2, 3, 4, 5 2 3 Links = 12, 13, 15, 23,
Complex networks Phys 682 / CIS 629: Computational Methods for Nonlinear Systems
 Complex networks Phys 682 / CIS 629: Computational Methods for Nonlinear Systems networks are everywhere (and always have been) - relationships (edges) among entities (nodes) explosion of interest in network
Complex networks Phys 682 / CIS 629: Computational Methods for Nonlinear Systems networks are everywhere (and always have been) - relationships (edges) among entities (nodes) explosion of interest in network
A quick review. The clustering problem: Hierarchical clustering algorithm: Many possible distance metrics K-mean clustering algorithm:
 The clustering problem: partition genes into distinct sets with high homogeneity and high separation Hierarchical clustering algorithm: 1. Assign each object to a separate cluster.. Regroup the pair of
The clustering problem: partition genes into distinct sets with high homogeneity and high separation Hierarchical clustering algorithm: 1. Assign each object to a separate cluster.. Regroup the pair of
TELCOM2125: Network Science and Analysis
 School of Information Sciences University of Pittsburgh TELCOM2125: Network Science and Analysis Konstantinos Pelechrinis Spring 2015 Figures are taken from: M.E.J. Newman, Networks: An Introduction 2
School of Information Sciences University of Pittsburgh TELCOM2125: Network Science and Analysis Konstantinos Pelechrinis Spring 2015 Figures are taken from: M.E.J. Newman, Networks: An Introduction 2
- relationships (edges) among entities (nodes) - technology: Internet, World Wide Web - biology: genomics, gene expression, proteinprotein
 Complex networks Phys 7682: Computational Methods for Nonlinear Systems networks are everywhere (and always have been) - relationships (edges) among entities (nodes) explosion of interest in network structure,
Complex networks Phys 7682: Computational Methods for Nonlinear Systems networks are everywhere (and always have been) - relationships (edges) among entities (nodes) explosion of interest in network structure,
CAIM: Cerca i Anàlisi d Informació Massiva
 1 / 72 CAIM: Cerca i Anàlisi d Informació Massiva FIB, Grau en Enginyeria Informàtica Slides by Marta Arias, José Balcázar, Ricard Gavaldá Department of Computer Science, UPC Fall 2016 http://www.cs.upc.edu/~caim
1 / 72 CAIM: Cerca i Anàlisi d Informació Massiva FIB, Grau en Enginyeria Informàtica Slides by Marta Arias, José Balcázar, Ricard Gavaldá Department of Computer Science, UPC Fall 2016 http://www.cs.upc.edu/~caim
Director. Katherine Icay June 13, 2018
 Director Katherine Icay June 13, 2018 Abstract Director is an R package designed to streamline the visualization of multiple levels of interacting RNA-seq data. It utilizes a modified Sankey plugin of
Director Katherine Icay June 13, 2018 Abstract Director is an R package designed to streamline the visualization of multiple levels of interacting RNA-seq data. It utilizes a modified Sankey plugin of
A quick review. Which molecular processes/functions are involved in a certain phenotype (e.g., disease, stress response, etc.)
 Gene expression profiling A quick review Which molecular processes/functions are involved in a certain phenotype (e.g., disease, stress response, etc.) The Gene Ontology (GO) Project Provides shared vocabulary/annotation
Gene expression profiling A quick review Which molecular processes/functions are involved in a certain phenotype (e.g., disease, stress response, etc.) The Gene Ontology (GO) Project Provides shared vocabulary/annotation
Networks and stability
 Networks and stability Part 1A. Network topology www.weaklink.sote.hu csermelypeter@yahoo.com Peter Csermely 1. network topology 2. network dynamics 3. examples for networks 4. synthesis (complex equilibria,
Networks and stability Part 1A. Network topology www.weaklink.sote.hu csermelypeter@yahoo.com Peter Csermely 1. network topology 2. network dynamics 3. examples for networks 4. synthesis (complex equilibria,
M.E.J. Newman: Models of the Small World
 A Review Adaptive Informatics Research Centre Helsinki University of Technology November 7, 2007 Vocabulary N number of nodes of the graph l average distance between nodes D diameter of the graph d is
A Review Adaptive Informatics Research Centre Helsinki University of Technology November 7, 2007 Vocabulary N number of nodes of the graph l average distance between nodes D diameter of the graph d is
The application of network theory in official statistics
 DGINS Conference 10/10/2018 The application of network theory in official statistics Áron Kincses, HCSO Scheme of presentation 1. Networks 2. The nature of networks 3. Networks in statistics and their
DGINS Conference 10/10/2018 The application of network theory in official statistics Áron Kincses, HCSO Scheme of presentation 1. Networks 2. The nature of networks 3. Networks in statistics and their
Efficient Algorithms for Solving Shortest Paths on Nearly Acyclic Directed Graphs
 Efficient Algorithms for Solving Shortest Paths on Nearly Acyclic Directed Graphs Shane Saunders Tadao Takaoka Department of Computer Science and Software Engineering, University of Canterbury, Christchurch,
Efficient Algorithms for Solving Shortest Paths on Nearly Acyclic Directed Graphs Shane Saunders Tadao Takaoka Department of Computer Science and Software Engineering, University of Canterbury, Christchurch,
Structure of biological networks. Presentation by Atanas Kamburov
 Structure of biological networks Presentation by Atanas Kamburov Seminar Gute Ideen in der theoretischen Biologie / Systembiologie 08.05.2007 Overview Motivation Definitions Large-scale properties of cellular
Structure of biological networks Presentation by Atanas Kamburov Seminar Gute Ideen in der theoretischen Biologie / Systembiologie 08.05.2007 Overview Motivation Definitions Large-scale properties of cellular
(Social) Networks Analysis III. Prof. Dr. Daning Hu Department of Informatics University of Zurich
 (Social) Networks Analysis III Prof. Dr. Daning Hu Department of Informatics University of Zurich Outline Network Topological Analysis Network Models Random Networks Small-World Networks Scale-Free Networks
(Social) Networks Analysis III Prof. Dr. Daning Hu Department of Informatics University of Zurich Outline Network Topological Analysis Network Models Random Networks Small-World Networks Scale-Free Networks
Network Analy Network Anal sis with sis with Cy C toscape
 Network Analysis with Cytoscape Chia-Lang Hsu Ueng-Cheng gyang 2 What is Cytoscape? Cytoscape is a network visualization and analysis tool. An extensible platform for bioinformatics software development.
Network Analysis with Cytoscape Chia-Lang Hsu Ueng-Cheng gyang 2 What is Cytoscape? Cytoscape is a network visualization and analysis tool. An extensible platform for bioinformatics software development.
The Generalized Topological Overlap Matrix in Biological Network Analysis
 The Generalized Topological Overlap Matrix in Biological Network Analysis Andy Yip, Steve Horvath Email: shorvath@mednet.ucla.edu Depts Human Genetics and Biostatistics, University of California, Los Angeles
The Generalized Topological Overlap Matrix in Biological Network Analysis Andy Yip, Steve Horvath Email: shorvath@mednet.ucla.edu Depts Human Genetics and Biostatistics, University of California, Los Angeles
Graph Theory. Graph Theory. COURSE: Introduction to Biological Networks. Euler s Solution LECTURE 1: INTRODUCTION TO NETWORKS.
 Graph Theory COURSE: Introduction to Biological Networks LECTURE 1: INTRODUCTION TO NETWORKS Arun Krishnan Koenigsberg, Russia Is it possible to walk with a route that crosses each bridge exactly once,
Graph Theory COURSE: Introduction to Biological Networks LECTURE 1: INTRODUCTION TO NETWORKS Arun Krishnan Koenigsberg, Russia Is it possible to walk with a route that crosses each bridge exactly once,
Girls Talk Math Summer Camp
 From Brains and Friendships to the Stock Market and the Internet -Sanjukta Krishnagopal 10 July 2018 Girls Talk Math Summer Camp Some real networks Social Networks Networks of acquaintances Collaboration
From Brains and Friendships to the Stock Market and the Internet -Sanjukta Krishnagopal 10 July 2018 Girls Talk Math Summer Camp Some real networks Social Networks Networks of acquaintances Collaboration
V 2 Clusters, Dijkstra, and Graph Layout"
 Bioinformatics 3! V 2 Clusters, Dijkstra, and Graph Layout" Mon, Oct 21, 2013" Graph Basics" A graph G is an ordered pair (V, E) of a set V of vertices and a set E of edges." Degree distribution P(k)!
Bioinformatics 3! V 2 Clusters, Dijkstra, and Graph Layout" Mon, Oct 21, 2013" Graph Basics" A graph G is an ordered pair (V, E) of a set V of vertices and a set E of edges." Degree distribution P(k)!
The Complex Network Phenomena. and Their Origin
 The Complex Network Phenomena and Their Origin An Annotated Bibliography ESL 33C 003180159 Instructor: Gerriet Janssen Match 18, 2004 Introduction A coupled system can be described as a complex network,
The Complex Network Phenomena and Their Origin An Annotated Bibliography ESL 33C 003180159 Instructor: Gerriet Janssen Match 18, 2004 Introduction A coupled system can be described as a complex network,
V 2 Clusters, Dijkstra, and Graph Layout
 Bioinformatics 3 V 2 Clusters, Dijkstra, and Graph Layout Fri, Oct 19, 2012 Graph Basics A graph G is an ordered pair (V, E) of a set V of vertices and a set E of edges. Degree distribution P(k) Random
Bioinformatics 3 V 2 Clusters, Dijkstra, and Graph Layout Fri, Oct 19, 2012 Graph Basics A graph G is an ordered pair (V, E) of a set V of vertices and a set E of edges. Degree distribution P(k) Random
Graph Mining and Social Network Analysis
 Graph Mining and Social Network Analysis Data Mining and Text Mining (UIC 583 @ Politecnico di Milano) References q Jiawei Han and Micheline Kamber, "Data Mining: Concepts and Techniques", The Morgan Kaufmann
Graph Mining and Social Network Analysis Data Mining and Text Mining (UIC 583 @ Politecnico di Milano) References q Jiawei Han and Micheline Kamber, "Data Mining: Concepts and Techniques", The Morgan Kaufmann
Lesson 4. Random graphs. Sergio Barbarossa. UPC - Barcelona - July 2008
 Lesson 4 Random graphs Sergio Barbarossa Graph models 1. Uncorrelated random graph (Erdős, Rényi) N nodes are connected through n edges which are chosen randomly from the possible configurations 2. Binomial
Lesson 4 Random graphs Sergio Barbarossa Graph models 1. Uncorrelated random graph (Erdős, Rényi) N nodes are connected through n edges which are chosen randomly from the possible configurations 2. Binomial
Complex networks: A mixture of power-law and Weibull distributions
 Complex networks: A mixture of power-law and Weibull distributions Ke Xu, Liandong Liu, Xiao Liang State Key Laboratory of Software Development Environment Beihang University, Beijing 100191, China Abstract:
Complex networks: A mixture of power-law and Weibull distributions Ke Xu, Liandong Liu, Xiao Liang State Key Laboratory of Software Development Environment Beihang University, Beijing 100191, China Abstract:
FastA & the chaining problem
 FastA & the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 1 Sources for this lecture: Lectures by Volker Heun, Daniel Huson and Knut Reinert,
FastA & the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 1 Sources for this lecture: Lectures by Volker Heun, Daniel Huson and Knut Reinert,
Overlay (and P2P) Networks
 Overlay (and P2P) Networks Part II Recap (Small World, Erdös Rényi model, Duncan Watts Model) Graph Properties Scale Free Networks Preferential Attachment Evolving Copying Navigation in Small World Samu
Overlay (and P2P) Networks Part II Recap (Small World, Erdös Rényi model, Duncan Watts Model) Graph Properties Scale Free Networks Preferential Attachment Evolving Copying Navigation in Small World Samu
Lectures by Volker Heun, Daniel Huson and Knut Reinert, in particular last years lectures
 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 4.1 Sources for this lecture Lectures by Volker Heun, Daniel Huson and Knut
4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem 4.1 Sources for this lecture Lectures by Volker Heun, Daniel Huson and Knut
Lecture (08, 09) Routing in Switched Networks
 Agenda Lecture (08, 09) Routing in Switched Networks Dr. Ahmed ElShafee Routing protocols Fixed Flooding Random Adaptive ARPANET Routing Strategies ١ Dr. Ahmed ElShafee, ACU Fall 2011, Networks I ٢ Dr.
Agenda Lecture (08, 09) Routing in Switched Networks Dr. Ahmed ElShafee Routing protocols Fixed Flooding Random Adaptive ARPANET Routing Strategies ١ Dr. Ahmed ElShafee, ACU Fall 2011, Networks I ٢ Dr.
Critical Phenomena in Complex Networks
 Critical Phenomena in Complex Networks Term essay for Physics 563: Phase Transitions and the Renormalization Group University of Illinois at Urbana-Champaign Vikyath Deviprasad Rao 11 May 2012 Abstract
Critical Phenomena in Complex Networks Term essay for Physics 563: Phase Transitions and the Renormalization Group University of Illinois at Urbana-Champaign Vikyath Deviprasad Rao 11 May 2012 Abstract
3 Global Properties of Networks
 3 Global Properties of Networks Ralf Steuer a and Gorka Zamora-López b a Humboldt University Berlin, Institute for Theoretical Biology, Invalidenstr. 43, 10115 Berlin, Germany. b University of Potsdam,
3 Global Properties of Networks Ralf Steuer a and Gorka Zamora-López b a Humboldt University Berlin, Institute for Theoretical Biology, Invalidenstr. 43, 10115 Berlin, Germany. b University of Potsdam,
A Comparison of Data Structures for Dijkstra's Single Source Shortest Path Algorithm. Shane Saunders
 A Comparison of Data Structures for Dijkstra's Single Source Shortest Path Algorithm Shane Saunders November 5, 1999 Abstract Dijkstra's algorithm computes the shortest paths between a starting vertex
A Comparison of Data Structures for Dijkstra's Single Source Shortest Path Algorithm Shane Saunders November 5, 1999 Abstract Dijkstra's algorithm computes the shortest paths between a starting vertex
Oh Pott, Oh Pott! or how to detect community structure in complex networks
 Oh Pott, Oh Pott! or how to detect community structure in complex networks Jörg Reichardt Interdisciplinary Centre for Bioinformatics, Leipzig, Germany (Host of the 2012 Olympics) Questions to start from
Oh Pott, Oh Pott! or how to detect community structure in complex networks Jörg Reichardt Interdisciplinary Centre for Bioinformatics, Leipzig, Germany (Host of the 2012 Olympics) Questions to start from
MORO: a Cytoscape App for Relationship Analysis between Modularity and Robustness in Large-Scale Biological Networks
 MORO: a Cytoscape App for Relationship Analysis between Modularity and Robustness in Large-Scale Biological Networks Authors: Cong-Doan Truong, Tien-Dzung Tran and Yung-Keun Kwon October 8, 2016 1 Contents
MORO: a Cytoscape App for Relationship Analysis between Modularity and Robustness in Large-Scale Biological Networks Authors: Cong-Doan Truong, Tien-Dzung Tran and Yung-Keun Kwon October 8, 2016 1 Contents
E6885 Network Science Lecture 5: Network Estimation and Modeling
 E 6885 Topics in Signal Processing -- Network Science E6885 Network Science Lecture 5: Network Estimation and Modeling Ching-Yung Lin, Dept. of Electrical Engineering, Columbia University October 7th,
E 6885 Topics in Signal Processing -- Network Science E6885 Network Science Lecture 5: Network Estimation and Modeling Ching-Yung Lin, Dept. of Electrical Engineering, Columbia University October 7th,
Introduction to the Special Issue on AI & Networks
 Introduction to the Special Issue on AI & Networks Marie desjardins, Matthew E. Gaston, and Dragomir Radev March 16, 2008 As networks have permeated our world, the economy has come to resemble an ecology
Introduction to the Special Issue on AI & Networks Marie desjardins, Matthew E. Gaston, and Dragomir Radev March 16, 2008 As networks have permeated our world, the economy has come to resemble an ecology
Data Structure Lecture#24: Graphs 2 (Chapter 11) U Kang Seoul National University
 Data Structure Lecture#24: Graphs 2 (Chapter 11) U Kang Seoul National University U Kang 1 In This Lecture Shortest path problem Main idea of Dijkstra s algorithm Cost analysis of Dijkstra s algorithm
Data Structure Lecture#24: Graphs 2 (Chapter 11) U Kang Seoul National University U Kang 1 In This Lecture Shortest path problem Main idea of Dijkstra s algorithm Cost analysis of Dijkstra s algorithm
Dijkstra s Single Source Shortest Path Algorithm. Andreas Klappenecker
 Dijkstra s Single Source Shortest Path Algorithm Andreas Klappenecker Single Source Shortest Path Given: a directed or undirected graph G = (V,E) a source node s in V a weight function w: E -> R. Problem:!!
Dijkstra s Single Source Shortest Path Algorithm Andreas Klappenecker Single Source Shortest Path Given: a directed or undirected graph G = (V,E) a source node s in V a weight function w: E -> R. Problem:!!
Overview of Network Theory, I
 Overview of Network Theory, I ECS 253 / MAE 253, Spring 2016, Lecture 1 Prof. Raissa D Souza University of California, Davis Raissa s background: 1999, PhD, Physics, Massachusetts Inst of Tech (MIT): Joint
Overview of Network Theory, I ECS 253 / MAE 253, Spring 2016, Lecture 1 Prof. Raissa D Souza University of California, Davis Raissa s background: 1999, PhD, Physics, Massachusetts Inst of Tech (MIT): Joint
Biological Networks Analysis
 iological Networks nalysis Introduction and ijkstra s algorithm Genome 559: Introduction to Statistical and omputational Genomics Elhanan orenstein The clustering problem: partition genes into distinct
iological Networks nalysis Introduction and ijkstra s algorithm Genome 559: Introduction to Statistical and omputational Genomics Elhanan orenstein The clustering problem: partition genes into distinct
FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:
 FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:56 4001 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem
FastA and the chaining problem, Gunnar Klau, December 1, 2005, 10:56 4001 4 FastA and the chaining problem We will discuss: Heuristics used by the FastA program for sequence alignment Chaining problem
Chapter 1. Social Media and Social Computing. October 2012 Youn-Hee Han
 Chapter 1. Social Media and Social Computing October 2012 Youn-Hee Han http://link.koreatech.ac.kr 1.1 Social Media A rapid development and change of the Web and the Internet Participatory web application
Chapter 1. Social Media and Social Computing October 2012 Youn-Hee Han http://link.koreatech.ac.kr 1.1 Social Media A rapid development and change of the Web and the Internet Participatory web application
To make sense of data, you can start by answering the following questions:
 Taken from the Introductory Biology 1, 181 lab manual, Biological Sciences, Copyright NCSU (with appreciation to Dr. Miriam Ferzli--author of this appendix of the lab manual). Appendix : Understanding
Taken from the Introductory Biology 1, 181 lab manual, Biological Sciences, Copyright NCSU (with appreciation to Dr. Miriam Ferzli--author of this appendix of the lab manual). Appendix : Understanding
Volume 2, Issue 11, November 2014 International Journal of Advance Research in Computer Science and Management Studies
 Volume 2, Issue 11, November 2014 International Journal of Advance Research in Computer Science and Management Studies Research Article / Survey Paper / Case Study Available online at: www.ijarcsms.com
Volume 2, Issue 11, November 2014 International Journal of Advance Research in Computer Science and Management Studies Research Article / Survey Paper / Case Study Available online at: www.ijarcsms.com
Department of Computer Science & Engineering University of Kalyani. Syllabus for Ph.D. Coursework
 Department of Computer Science & Engineering University of Kalyani Syllabus for Ph.D. Coursework Paper 1: A) Literature Review: (Marks - 25) B) Research Methodology: (Marks - 25) Paper 2: Computer Applications:
Department of Computer Science & Engineering University of Kalyani Syllabus for Ph.D. Coursework Paper 1: A) Literature Review: (Marks - 25) B) Research Methodology: (Marks - 25) Paper 2: Computer Applications:
Graph theory and GIS
 Geoinformatics FCE CTU 2009 Workshop 18th september 2009 Contents Graph as such Definition Alternative definition Undirected graphs Graphs as a model of reality Contents Graph as such Definition Alternative
Geoinformatics FCE CTU 2009 Workshop 18th september 2009 Contents Graph as such Definition Alternative definition Undirected graphs Graphs as a model of reality Contents Graph as such Definition Alternative
Constructing a G(N, p) Network
 Random Graph Theory Dr. Natarajan Meghanathan Associate Professor Department of Computer Science Jackson State University, Jackson, MS E-mail: natarajan.meghanathan@jsums.edu Introduction At first inspection,
Random Graph Theory Dr. Natarajan Meghanathan Associate Professor Department of Computer Science Jackson State University, Jackson, MS E-mail: natarajan.meghanathan@jsums.edu Introduction At first inspection,
On the Complexity of the Policy Improvement Algorithm. for Markov Decision Processes
 On the Complexity of the Policy Improvement Algorithm for Markov Decision Processes Mary Melekopoglou Anne Condon Computer Sciences Department University of Wisconsin - Madison 0 West Dayton Street Madison,
On the Complexity of the Policy Improvement Algorithm for Markov Decision Processes Mary Melekopoglou Anne Condon Computer Sciences Department University of Wisconsin - Madison 0 West Dayton Street Madison,
Algorithms and Data Structures 2014 Exercises and Solutions Week 9
 Algorithms and Data Structures 2014 Exercises and Solutions Week 9 November 26, 2014 1 Directed acyclic graphs We are given a sequence (array) of numbers, and we would like to find the longest increasing
Algorithms and Data Structures 2014 Exercises and Solutions Week 9 November 26, 2014 1 Directed acyclic graphs We are given a sequence (array) of numbers, and we would like to find the longest increasing
Why Should We Care? More importantly, it is easy to lie or deceive people with bad plots
 Plots & Graphs Why Should We Care? Everyone uses plots and/or graphs But most people ignore or are unaware of simple principles Default plotting tools (or default settings) are not always the best More
Plots & Graphs Why Should We Care? Everyone uses plots and/or graphs But most people ignore or are unaware of simple principles Default plotting tools (or default settings) are not always the best More
SHAPE, SPACE & MEASURE
 STAGE 1 Know the place value headings up to millions Recall primes to 19 Know the first 12 square numbers Know the Roman numerals I, V, X, L, C, D, M Know the % symbol Know percentage and decimal equivalents
STAGE 1 Know the place value headings up to millions Recall primes to 19 Know the first 12 square numbers Know the Roman numerals I, V, X, L, C, D, M Know the % symbol Know percentage and decimal equivalents
Quake Heaps: A Simple Alternative to Fibonacci Heaps
 Quake Heaps: A Simple Alternative to Fibonacci Heaps Timothy M. Chan Cheriton School of Computer Science, University of Waterloo, Waterloo, Ontario N2L G, Canada, tmchan@uwaterloo.ca Abstract. This note
Quake Heaps: A Simple Alternative to Fibonacci Heaps Timothy M. Chan Cheriton School of Computer Science, University of Waterloo, Waterloo, Ontario N2L G, Canada, tmchan@uwaterloo.ca Abstract. This note
On Reshaping of Clustering Coefficients in Degreebased Topology Generators
 On Reshaping of Clustering Coefficients in Degreebased Topology Generators Xiafeng Li, Derek Leonard, and Dmitri Loguinov Texas A&M University Presented by Derek Leonard Agenda Motivation Statement of
On Reshaping of Clustering Coefficients in Degreebased Topology Generators Xiafeng Li, Derek Leonard, and Dmitri Loguinov Texas A&M University Presented by Derek Leonard Agenda Motivation Statement of
CSE 548: Analysis of Algorithms. Lectures 14 & 15 ( Dijkstra s SSSP & Fibonacci Heaps )
 CSE 548: Analysis of Algorithms Lectures 14 & 15 ( Dijkstra s SSSP & Fibonacci Heaps ) Rezaul A. Chowdhury Department of Computer Science SUNY Stony Brook Fall 2012 Fibonacci Heaps ( Fredman & Tarjan,
CSE 548: Analysis of Algorithms Lectures 14 & 15 ( Dijkstra s SSSP & Fibonacci Heaps ) Rezaul A. Chowdhury Department of Computer Science SUNY Stony Brook Fall 2012 Fibonacci Heaps ( Fredman & Tarjan,
MICROARRAY IMAGE SEGMENTATION USING CLUSTERING METHODS
 Mathematical and Computational Applications, Vol. 5, No. 2, pp. 240-247, 200. Association for Scientific Research MICROARRAY IMAGE SEGMENTATION USING CLUSTERING METHODS Volkan Uslan and Đhsan Ömür Bucak
Mathematical and Computational Applications, Vol. 5, No. 2, pp. 240-247, 200. Association for Scientific Research MICROARRAY IMAGE SEGMENTATION USING CLUSTERING METHODS Volkan Uslan and Đhsan Ömür Bucak
Knowledge Discovery and Data Mining
 Knowledge Discovery and Data Mining Unit # 2 Sajjad Haider Spring 2010 1 Structured vs. Non-Structured Data Most business databases contain structured data consisting of well-defined fields with numeric
Knowledge Discovery and Data Mining Unit # 2 Sajjad Haider Spring 2010 1 Structured vs. Non-Structured Data Most business databases contain structured data consisting of well-defined fields with numeric
BIO 360: Vertebrate Physiology Lab 9: Graphing in Excel. Lab 9: Graphing: how, why, when, and what does it mean? Due 3/26
 Lab 9: Graphing: how, why, when, and what does it mean? Due 3/26 INTRODUCTION Graphs are one of the most important aspects of data analysis and presentation of your of data. They are visual representations
Lab 9: Graphing: how, why, when, and what does it mean? Due 3/26 INTRODUCTION Graphs are one of the most important aspects of data analysis and presentation of your of data. They are visual representations
Introduction to Bioinformatics
 Introduction to Bioinformatics Biological Networks Department of Computing Imperial College London Spring 2010 1. Motivation Large Networks model many real-world phenomena technological: www, internet,
Introduction to Bioinformatics Biological Networks Department of Computing Imperial College London Spring 2010 1. Motivation Large Networks model many real-world phenomena technological: www, internet,
Constructing a G(N, p) Network
 Random Graph Theory Dr. Natarajan Meghanathan Professor Department of Computer Science Jackson State University, Jackson, MS E-mail: natarajan.meghanathan@jsums.edu Introduction At first inspection, most
Random Graph Theory Dr. Natarajan Meghanathan Professor Department of Computer Science Jackson State University, Jackson, MS E-mail: natarajan.meghanathan@jsums.edu Introduction At first inspection, most
Examples of Complex Networks
 Examples of Complex Networks Neural Network (C. Elegans) http://gephi.org/wp-content/uploads/2008/12/screenshot-celegans.png Food Web http://1.bp.blogspot.com/_vifbm3t8bou/sbhzqbchiei/aaaaaaaaaxk/rsc-
Examples of Complex Networks Neural Network (C. Elegans) http://gephi.org/wp-content/uploads/2008/12/screenshot-celegans.png Food Web http://1.bp.blogspot.com/_vifbm3t8bou/sbhzqbchiei/aaaaaaaaaxk/rsc-
1 General description of the projects
 1 General description of the projects The term projects are aimed at providing a test ground for the skills and knowledge you picked up during the course and optionally give you a chance to join ongoing
1 General description of the projects The term projects are aimed at providing a test ground for the skills and knowledge you picked up during the course and optionally give you a chance to join ongoing
Clustering CS 550: Machine Learning
 Clustering CS 550: Machine Learning This slide set mainly uses the slides given in the following links: http://www-users.cs.umn.edu/~kumar/dmbook/ch8.pdf http://www-users.cs.umn.edu/~kumar/dmbook/dmslides/chap8_basic_cluster_analysis.pdf
Clustering CS 550: Machine Learning This slide set mainly uses the slides given in the following links: http://www-users.cs.umn.edu/~kumar/dmbook/ch8.pdf http://www-users.cs.umn.edu/~kumar/dmbook/dmslides/chap8_basic_cluster_analysis.pdf
Response Network Emerging from Simple Perturbation
 Journal of the Korean Physical Society, Vol 44, No 3, March 2004, pp 628 632 Response Network Emerging from Simple Perturbation S-W Son, D-H Kim, Y-Y Ahn and H Jeong Department of Physics, Korea Advanced
Journal of the Korean Physical Society, Vol 44, No 3, March 2004, pp 628 632 Response Network Emerging from Simple Perturbation S-W Son, D-H Kim, Y-Y Ahn and H Jeong Department of Physics, Korea Advanced
Cluster Analysis. Angela Montanari and Laura Anderlucci
 Cluster Analysis Angela Montanari and Laura Anderlucci 1 Introduction Clustering a set of n objects into k groups is usually moved by the aim of identifying internally homogenous groups according to a
Cluster Analysis Angela Montanari and Laura Anderlucci 1 Introduction Clustering a set of n objects into k groups is usually moved by the aim of identifying internally homogenous groups according to a
Topic 1: What is HoTT and why?
 Topic 1: What is HoTT and why? May 5, 2014 Introduction Homotopy type theory (HoTT) is a newly emerging field of mathematics which is currently being developed as a foundation of mathematics which is in
Topic 1: What is HoTT and why? May 5, 2014 Introduction Homotopy type theory (HoTT) is a newly emerging field of mathematics which is currently being developed as a foundation of mathematics which is in
Shortest Path Algorithm
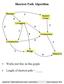 Shortest Path Algorithm C Works just fine on this graph. C Length of shortest path = Copyright 2005 DIMACS BioMath Connect Institute Robert Hochberg Dynamic Programming SP #1 Same Questions, Different
Shortest Path Algorithm C Works just fine on this graph. C Length of shortest path = Copyright 2005 DIMACS BioMath Connect Institute Robert Hochberg Dynamic Programming SP #1 Same Questions, Different
Bidirectional search and Goal-directed Dijkstra
 Computing the shortest path Bidirectional search and Goal-directed Dijkstra Alexander Kozyntsev October 18, 2010 Abstract We study the problem of finding a shortest path between two vertices in a directed
Computing the shortest path Bidirectional search and Goal-directed Dijkstra Alexander Kozyntsev October 18, 2010 Abstract We study the problem of finding a shortest path between two vertices in a directed
Clustering analysis of gene expression data
 Clustering analysis of gene expression data Chapter 11 in Jonathan Pevsner, Bioinformatics and Functional Genomics, 3 rd edition (Chapter 9 in 2 nd edition) Human T cell expression data The matrix contains
Clustering analysis of gene expression data Chapter 11 in Jonathan Pevsner, Bioinformatics and Functional Genomics, 3 rd edition (Chapter 9 in 2 nd edition) Human T cell expression data The matrix contains
GraphLinker. A Visual Comparative Environment of Genomic and Metabolic Networks. Project Update. Niels Hanson
 GraphLinker A Visual Comparative Environment of Genomic and Metabolic Networks Project Update Niels Hanson Monday, November 21 2011 Microbacterial Communities diversity and dynamics largely unexplored
GraphLinker A Visual Comparative Environment of Genomic and Metabolic Networks Project Update Niels Hanson Monday, November 21 2011 Microbacterial Communities diversity and dynamics largely unexplored
Outline. Computer Science 331. Analysis of Prim's Algorithm
 Outline Computer Science 331 Analysis of Prim's Algorithm Mike Jacobson Department of Computer Science University of Calgary Lecture #34 1 Introduction 2, Concluded 3 4 Additional Comments and References
Outline Computer Science 331 Analysis of Prim's Algorithm Mike Jacobson Department of Computer Science University of Calgary Lecture #34 1 Introduction 2, Concluded 3 4 Additional Comments and References
Smallest small-world network
 Smallest small-world network Takashi Nishikawa, 1, * Adilson E. Motter, 1, Ying-Cheng Lai, 1,2 and Frank C. Hoppensteadt 1,2 1 Department of Mathematics, Center for Systems Science and Engineering Research,
Smallest small-world network Takashi Nishikawa, 1, * Adilson E. Motter, 1, Ying-Cheng Lai, 1,2 and Frank C. Hoppensteadt 1,2 1 Department of Mathematics, Center for Systems Science and Engineering Research,
SNA 8: network resilience. Lada Adamic
 SNA 8: network resilience Lada Adamic Outline Node vs. edge percolation Resilience of randomly vs. preferentially grown networks Resilience in real-world networks network resilience Q: If a given fraction
SNA 8: network resilience Lada Adamic Outline Node vs. edge percolation Resilience of randomly vs. preferentially grown networks Resilience in real-world networks network resilience Q: If a given fraction
Scribe from 2014/2015: Jessica Su, Hieu Pham Date: October 6, 2016 Editor: Jimmy Wu
 CS 267 Lecture 3 Shortest paths, graph diameter Scribe from 2014/2015: Jessica Su, Hieu Pham Date: October 6, 2016 Editor: Jimmy Wu Today we will talk about algorithms for finding shortest paths in a graph.
CS 267 Lecture 3 Shortest paths, graph diameter Scribe from 2014/2015: Jessica Su, Hieu Pham Date: October 6, 2016 Editor: Jimmy Wu Today we will talk about algorithms for finding shortest paths in a graph.
Middle School Math Course 2
 Middle School Math Course 2 Correlation of the ALEKS course Middle School Math Course 2 to the Indiana Academic Standards for Mathematics Grade 7 (2014) 1: NUMBER SENSE = ALEKS course topic that addresses
Middle School Math Course 2 Correlation of the ALEKS course Middle School Math Course 2 to the Indiana Academic Standards for Mathematics Grade 7 (2014) 1: NUMBER SENSE = ALEKS course topic that addresses
Summary: What We Have Learned So Far
 Summary: What We Have Learned So Far small-world phenomenon Real-world networks: { Short path lengths High clustering Broad degree distributions, often power laws P (k) k γ Erdös-Renyi model: Short path
Summary: What We Have Learned So Far small-world phenomenon Real-world networks: { Short path lengths High clustering Broad degree distributions, often power laws P (k) k γ Erdös-Renyi model: Short path
ADDENDUM. PRINCIPLES OF DESIGN COURSE Topic YouTube link QR Code
 ADDENDUM PRINCIPLES OF DESIGN COURSE Topic YouTube link QR Code Topic 1 Introduction to Graphic Design https://youtu.be/pacrrojlvui This video discussed on essential skills of a graphic design and its
ADDENDUM PRINCIPLES OF DESIGN COURSE Topic YouTube link QR Code Topic 1 Introduction to Graphic Design https://youtu.be/pacrrojlvui This video discussed on essential skills of a graphic design and its
Algorithm Analysis Graph algorithm. Chung-Ang University, Jaesung Lee
 Algorithm Analysis Graph algorithm Chung-Ang University, Jaesung Lee Basic definitions Graph = (, ) where is a set of vertices and is a set of edges Directed graph = where consists of ordered pairs
Algorithm Analysis Graph algorithm Chung-Ang University, Jaesung Lee Basic definitions Graph = (, ) where is a set of vertices and is a set of edges Directed graph = where consists of ordered pairs
Bramhall High school Year 8 Assessment Descriptors Mathematics
 Grade Description Calculate with negative indices in the context of standard form. 8/9 Multiply (divide) numbers written in standard form. Use inequalities to describe the range of values for a rounded
Grade Description Calculate with negative indices in the context of standard form. 8/9 Multiply (divide) numbers written in standard form. Use inequalities to describe the range of values for a rounded
d. Solution coming soon. 6. Directed Graphs
 Fall 2013 Final Solutions (Beta Edition) 1. TSTs. a. ID, IDD, IDDQD, IDKFA, XA, XX, XEA, XYZZY b. IDAAAA or XDAAAA or XCAAAA, or indeed anything that is six letters long and starts with ID, XD, or XC.
Fall 2013 Final Solutions (Beta Edition) 1. TSTs. a. ID, IDD, IDDQD, IDKFA, XA, XX, XEA, XYZZY b. IDAAAA or XDAAAA or XCAAAA, or indeed anything that is six letters long and starts with ID, XD, or XC.
Complex Networks. Structure and Dynamics
 Complex Networks Structure and Dynamics Ying-Cheng Lai Department of Mathematics and Statistics Department of Electrical Engineering Arizona State University Collaborators! Adilson E. Motter, now at Max-Planck
Complex Networks Structure and Dynamics Ying-Cheng Lai Department of Mathematics and Statistics Department of Electrical Engineering Arizona State University Collaborators! Adilson E. Motter, now at Max-Planck
Centralities for undirected graphs
 Chapter 4 Centralities for undirected graphs The next step in network analysis is to define, using mathematical formalisms, the different features we want to compute. So we present indexes, called centralities,
Chapter 4 Centralities for undirected graphs The next step in network analysis is to define, using mathematical formalisms, the different features we want to compute. So we present indexes, called centralities,
Topology of the Erasmus student mobility network
 Topology of the Erasmus student mobility network Aranka Derzsi, Noemi Derzsy, Erna Káptalan and Zoltán Néda The Erasmus student mobility network Aim: study the network s topology (structure) and its characteristics
Topology of the Erasmus student mobility network Aranka Derzsi, Noemi Derzsy, Erna Káptalan and Zoltán Néda The Erasmus student mobility network Aim: study the network s topology (structure) and its characteristics
The Establishment Game. Motivation
 Motivation Motivation The network models so far neglect the attributes, traits of the nodes. A node can represent anything, people, web pages, computers, etc. Motivation The network models so far neglect
Motivation Motivation The network models so far neglect the attributes, traits of the nodes. A node can represent anything, people, web pages, computers, etc. Motivation The network models so far neglect
The Mathematical Description of Networks
 Modelling Complex Systems University of Manchester, 21 st 23 rd June 2010 Tim Evans Theoretical Physics The Mathematical Description of Networs Page 1 Notation I will focus on Simple Graphs with multiple
Modelling Complex Systems University of Manchester, 21 st 23 rd June 2010 Tim Evans Theoretical Physics The Mathematical Description of Networs Page 1 Notation I will focus on Simple Graphs with multiple
User Guide. v Released June Advaita Corporation 2016
 User Guide v. 0.9 Released June 2016 Copyright Advaita Corporation 2016 Page 2 Table of Contents Table of Contents... 2 Background and Introduction... 4 Variant Calling Pipeline... 4 Annotation Information
User Guide v. 0.9 Released June 2016 Copyright Advaita Corporation 2016 Page 2 Table of Contents Table of Contents... 2 Background and Introduction... 4 Variant Calling Pipeline... 4 Annotation Information
CS-E5740. Complex Networks. Scale-free networks
 CS-E5740 Complex Networks Scale-free networks Course outline 1. Introduction (motivation, definitions, etc. ) 2. Static network models: random and small-world networks 3. Growing network models: scale-free
CS-E5740 Complex Networks Scale-free networks Course outline 1. Introduction (motivation, definitions, etc. ) 2. Static network models: random and small-world networks 3. Growing network models: scale-free
/4 Directions: Graph the functions, then answer the following question.
 1.) Graph y = x. Label the graph. Standard: F-BF.3 Identify the effect on the graph of replacing f(x) by f(x) +k, k f(x), f(kx), and f(x+k), for specific values of k; find the value of k given the graphs.
1.) Graph y = x. Label the graph. Standard: F-BF.3 Identify the effect on the graph of replacing f(x) by f(x) +k, k f(x), f(kx), and f(x+k), for specific values of k; find the value of k given the graphs.
