AutoSOME User Manual
|
|
|
- Herbert Harrington
- 6 years ago
- Views:
Transcription
1 AutoSOME User Manual February 16, 2014 Written by Aaron M. Newman, Ph.D. Institute for Stem Cell Biology and Regenerative Medicine Stanford University
2 Table of Contents Introduction... 1 System Requirements... 1 Installing and Running the AutoSOME GUI... 1 Using AutoSOME... 2 Input Format... 2 Missing Values... 3 AutoSOME GUI... 3 Load (and Filter) Input... 3 Input Adjustment... 6 Basic Fields... 7 Advanced Fields... 8 Fuzzy Cluster Networks... 8 Memory... 8 Algorithm Settings... 8 Self-Organizing Map (SOM)... 8 Cartogram... 9 Running AutoSOME... 9 Output Output Files Cluster tree Heat maps Signal plots View>settings>Image Settings View>settings>scale using entire data set Search Saving data Files written to disk Opening previous clustering results Reset Enhanced GUI vs. Basic GUI AutoSOME Command Line Version Creating Fuzzy Cluster Networks References... 23
3 Introduction AutoSOME is a powerful unsupervised computational method for identifying discrete and fuzzy clusters of diverse geometries from potentially large, multi-dimensional, data sets without prior knowledge of cluster number or structure [Newman and Cooper, 2010]. We have implemented the AutoSOME method as a Graphical User Interface (GUI) and command line tool to facilitate its use by the academic research community. Below, instructions are provided for using both versions of AutoSOME, and a protocol is given for generating AutoSOME fuzzy cluster networks in Cytoscape [Shannon et al., 2005]. Refer to the AutoSOME website for tutorials, software, and FAQ [ System Requirements AutoSOME is coded in Java to promote platform independence. To launch the GUI from the AutoSOME website [ Java Web Start technology needs to be installed on your computer along with Java Standard Edition 1.6, both of which are freely available at The command line version of AutoSOME will run in JAVA 1.5 and perhaps earlier versions. In general, the more memory and processors available, the better the performance of AutoSOME. Without user intervention, both the GUI and command line versions will greedily use all available CPUs. For 32-bit Windows systems, the maximum amount of memory that can be allocated to AutoSOME is about 1.6 GB, while 64-bit operating systems with 64-bit Java can allocate up to ~30 GB of memory. Microarray data sets like the HG U133 Plus 2.0 Affymetrix chipset (>54k probes) with dozens of samples can be run with 1.6 GB RAM. The Java Web Start launch button on the AutoSOME Web Portal homepage will automatically allocate up to 1 GB of RAM. To run AutoSOME with more or less memory, download the.jar file from: Installing and Running the AutoSOME GUI Download AutoSOME from the website, and then unzip all contents of the downloaded archive into the same directory. Using your system console, navigate to the directory where you installed AutoSOME, and type: java Xmx1600m Xms1600m jar autosome_vxxxxxx.jar XXXXXX = the GUI version you downloaded (each version is named by the date of compilation, e.g ). The -Xmx,-Xms arguments allocate additional memory to the 1
4 Java Virtual Machine necessary for running large data sets. Allocate more or less memory as needed for your input data set, parameters, and machine architecture. Also, AutoSOME has not been internationalized. If it is behaving oddly, try adding the following JVM argument -Duser.language=en. For instructions on how to use the AutoSOME command line interface, see AutoSOME Command Line Version below. Using AutoSOME Input Format AutoSOME accepts three kinds of numerical input files. Standard format: The input file is a table of numerical values with optional column labels (row 1) and mandatory row labels (column 1). If column labels are specified, column 1 also needs a label in row 1. All entries must be tab, comma, or space delimited. Input data can be easily formatted using Microsoft Excel. See Table 1 below for an example. Table 1 Probe hesc-1 hesc-2 hesc-3 hesc-4 ipsc-1 ipsc-2 ipsc _at _x_at _at _at _x_at Microarray formats: 1) Gene Expression Omnibus (GEO) Series Matrix. This format contains all chipset expression data as well as user-supplied annotation in a spreadsheet style. AutoSOME can automatically extract expression information content and column names from a raw series matrix file. This allows for rapid analysis of any GEO data set by simply downloading the archive containing the series matrix text file, unzipping it, and loading it into AutoSOME. 2) PCL = Pre CLuster format used for the Cluster software [Eisen et al. (1998) PNAS 95:14863]. For an example, see or the Cluster/TreeView User Manual. The first two columns are reserved for gene annotation (column 1 = row identifiers, column 2 = gene names or annotation). Column 3 is optional, is called GWEIGHT, and specifies how to weigh each gene 2
5 when computing gene-gene similarity. This column is read by AutoSOME, but is ignored since AutoSOME does not cluster genes using a similarity matrix. Row 1 is mandatory and is used to provide column names including names of each array (e.g. ID, NAME, GWEIGHT, array1, array2, ). Row 2 is optional, is called EWEIGHT, and specifies how each array is weighed when computing array-array similarity. In contrast to GWEIGHT, AutoSOME will use EWEIGHT when constructing a distance matrix for transcriptome clustering (fuzzy cluster networks option). Missing Values There are two strategies implemented in AutoSOME for handling missing values (blank space,?, or NA ). 1) Replace missing values with median of columns (or rows). 2) Replace missing values with mean of columns (or rows). Go to File>Settings to launch a window that will allow you to specify which of these two techniques to use. AutoSOME GUI The AutoSOME GUI is launched using Java Web Start from your browser or via the command line, and is used to run AutoSOME as well as to browse and generate publication quality visualizations of the cluster output (Figure 1). Figure 1 Load (and Filter) Input To begin, launch a file browser by either pressing the large INPUT button in the AutoSOME Steps panel on the top left, or by clicking the browse button adjacent to the input file text box in the Input and Output section (Figure 1). A file browser will appear. For standard input (see Input Format above), simply find your file and open it. For 3
6 microarray input, optional data formats are shown at the top of the file browser (see Figure 2). Select the appropriate file format for your microarray data set and load the file. Figure 2 Once loaded, you will be asked whether you want to filter your input data set (Figure 3). Figure 3 If you choose yes you will be redirected to a Filter Data window (Figure 4 below). This window provides two simple data filtration options: i) Remove rows with fold change less than X. The fold change is calculated by the difference between the minimum and maximum values for each row. All rows with fold change <X will be removed. Importantly, if your data are already log2 scaled, make sure you select the corresponding checkbox. ii) Remove rows with mean value below Y. The mean µ of each row is calculated, and the row is removed if µ <Y. After specifying filtration options, press Apply to preview the filtered data, and if you approve, press Accept. 4
7 Figure 4 Figure 5 After your input file has loaded, the number of rows and columns will be displayed in the Input Data panel on the left side of the GUI (see Figure 5). The maximum and minimum values over the entire data set are also provided. When finished clustering, this table will dynamically update based on the contents of selected clusters in the cluster tree (see Output below). 5
8 Input Adjustment AutoSOME implements several input normalization methods in addition to log2 scaling. For an excellent overview of these techniques, see the manual to the Cluster software available at [Eisen et al., 1998]. Press the Show button to the right of the Input Adjustment label to access input scaling and normalization options (see Figure 6 below). All input adjustment operations are performed in the order listed in the GUI, from top to bottom. In brief: i) Log2 Scaling: Logarithmic scaling is routinely used for microarray data sets to amplify small fold changes in gene expression, and is completely reversible. We recommend applying log2 scaling in cases where expression values span several orders of magnitude. All other implemented input adjustment methods irreversibly change the input to make it more suitable for analysis. ii) Unit variance: forces all columns to have zero mean and a standard deviation of one, and is commonly used when there is no a priori reason to treat any column differently from any other. iii) Range [0,x]: Alternatively, data in all columns can be normalized to share lowest and highest values (0,x) by specifying an upper bound x. iv) Median Center Rows/Arrays: For microarray analysis, centering each gene (row) and/or array (column) by subtracting the median value of the row/column eliminates amplitude shifts to cluster distinct expression patterns rather than overall transcript levels. We recommend applying median centering to genes before co-expression clustering (clustering rows). v) Sum of Squares=1 Rows/Arrays: This normalization procedure smoothes the data set by forcing the sum of squares of all expression values to equal 1 for each row/column in the data set. The impact of this method on cluster identification can be significant and tends to result in the detection of larger clusters trailing off into genes with minimal differential expression (after clustering, the signal to noise ratio can be boosted by using the confidence filter; see Output below) As an example, to run AutoSOME co-expression analysis, one may apply log2 scaling, unit variance normalization, and median-centering of genes (and/or arrays). To run AutoSOME transcriptome clustering (fuzzy cluster networks, see tutorial below), it is generally recommended to apply unit variance normalization to your data set. Other normalization settings may also be desirable. Input can also be adjusted using Microsoft Excel or the Cluster software [Eisen et al., 1998] and then imported into AutoSOME. 6
9 Figure 6 Basic Fields Press the Show button to expand the Basic Fields section (see Figure 6). i) Cluster Analysis: Switch among clustering rows (e.g. genes), columns (e.g. experiments), or both rows and columns in a single step. ii) Running Mode: This parameter alters advanced AutoSOME algorithm settings to switch among Precision, Normal, and Speed modes of operation. Precision takes longer, but provides greater training of the SOM node lattice (2X1000 iterations) and greater resolution for density-equalization of the SOM error surface (64X64). For enhanced performance, and especially for first-pass exploratory cluster analysis, choose Speed for less SOM iterations (2X250) and less resolution for density-equalization (16X16). Normal provides a compromise between the other two settings (2X500 SOM iterations and 32X32 cartogram resolution). In our experiments, Normal works quite well, and in fact, often yields comparable results to Precision. Choose Speed for a first-pass exploratory analysis. iii) No. Ensemble Runs: The default of 50 ensemble iterations should be sufficient to begin investigating the cluster structure of most data sets. Although in practice, AutoSOME clustering results can be quite stable with ~50 ensemble iterations (and even as little as 20), for final clustering results, it is recommended to increase this number to at least 100. iv) P-value Threshold: AutoSOME has been extensively benchmarked on a highly diverse array of clustering problems using a P-value cutoff of 0.1. Reduce the p-value threshold to identify tighter clusters. 7
10 v) No. CPUs: To liberate processor resources, decrease No. CPUs. AutoSOME running time will decrease approximately linearly with respect to increasing number of dedicated CPUs. Advanced Fields Fuzzy Cluster Networks. When clustering columns (i.e. Cluster Analysis in Basic Fields is set to columns or both ), the Fuzzy Cluster Networks window will be automatically enabled (see Figure 6). In this window, you can pick one of three distance metrics for calculating the input distance matrix (Euclidean, Pearson s and Uncentered correlation). Euclidean distance is selected by default due to good results obtained from empirical testing. See the review by D haeseleer (2005) for descriptions of Euclidean, Pearson s and Uncentered correlation distance metrics. In brief, Euclidean distance is sensitive to amplitude shifts while Pearson s correlation is not. Experiment with these metrics to get a feel for how they work. If smoothing out the distance matrix is desired, Unit Variance Normalize can be selected to normalize the distance matrix columns to unit variance. When finished clustering, AutoSOME will automatically generate additional output files for building fuzzy cluster networks in Cytoscape (see Fuzzy Cluster Networks below). Memory. Write Intermediate Runs to Disk: If AutoSOME crashes (or seems to hang for a long period of time), then Write Ensemble Runs to Disk should be selected (see Figure 6) as an out of memory error is likely. This may be necessary on systems with insufficient RAM for large data sets and many ensemble runs (e.g. 45k probes, 100 samples, and 2000 ensemble runs). Overall running time will be slower due to reading and writing to disk. Files will be written to a temporary folder in the output directory. Also, try reducing the number of CPUs to free up memory. Algorithm Settings. Press the Show button to expand the Algorithm Settings section (see Figure 6). The GUI permits modification of the most critical algorithm parameters. For access to all algorithm parameters, use the command line version of AutoSOME. Self-Organizing Map (SOM). i) Maximum Grid Length: The maximum number of nodes for the x/y length of the SOM node lattice. ii) Minimum Grid Length: The minimum number of nodes for the x/y length of the SOM node grid. In general increase Maximum Grid Length to provide greater resolution for cluster separation in the SOM, and to increase the number of possible clusters. Increase Minimum Grid Length to force AutoSOME to use an SOM lattice with 8
11 specific minimum dimensions. Set both parameters to the same value to disable automatic adjustment of grid length by the algorithm. The SOM grid length is set, by default, between 5 and 30, providing sufficient cluster resolving power for most clustering problems. If a data set with at least 5,000 rows is imported (e.g. whole-genome microarray), the maximum grid length will be automatically set to 20 for more efficient running time. iii) Training Iterations: The number of iterations for training the SOM node lattice. The value of this parameter dictates the number of iterations for each of two phases, coarse-grained followed by fine-grained training. This value can be toggled between 1000 iterations for precision (used during benchmarking in Newman and Cooper, 2010), 500 iterations for normal, and 250 iterations for speed. iv) Use Square Topology: Select this checkbox to use a square-shaped node lattice rather than a circular node lattice (default). Benchmarking results indicate both topologies yield comparable results. See the Manuscript for more details. Cartogram. Resolution: The density-equalizing cartogram algorithm requires an input array with dimensions that are a power of two. Although all benchmarking tests were conducted using an array size of 64X64, greater resolutions may yield more accurate density-equalization (especially when the SOM dimensions are large). On the other hand, going from 64X64 to 32X32 may increase running time dramatically (up to ~4X), and is sufficient for accurate clustering results in many cases (especially when the SOM dimensions are less than 30X30). Running AutoSOME After the input file and parameters are specified, execute AutoSOME clustering by pressing the green RUN button in the AutoSOME Steps panel (see Figure 1). Progress is shown in the lower left corner of the GUI and elapsed time is displayed (see Figure 7). When finished, the GUI will automatically be redirected to the output panel for browsing the cluster output. The input and output panels can be toggled back and forth using the tabs or the INPUT and OUTPUT buttons in the control panel. Figure 7 9
12 Output Output Files. The Output Files table, located on the center-left side of the GUI, will show links to the AutoSOME output files after clustering is finished (Figure 8). Simply select a file from the table and the file will be automatically displayed. For details of the output files generated by AutoSOME, see files written to disk below. Figure 8 Cluster tree. As shown in Figure 9, all clusters are listed using a dynamic tree structure and are ordered from top to bottom by decreasing size. Figure 9 Once a cluster node is selected using your mouse, the cluster tree can be rapidly traversed by pressing the and keys. To select more than one cluster, hold down the Shift or Control key and select using your mouse. Clusters can be manually modified using the control panel below the cluster tree (Figure 10). Use Split to make a new cluster from a selected group of data points (first expand a cluster node by double-clicking, then select data points with mouse). Press Delete key to erase entire clusters or specific data points. Press Merge to combine the contents of all selected clusters into a single cluster. Press Reset to return to the original clustering output. To filter clustering results by cluster confidence (metric ranging from 1-100, where 100=data point always in cluster x) type in confidence threshold 100 in the Confidence text box and press Update. Figure 10 10
13 By default, the contents of the selected cluster(s) will be displayed as a list of cluster labels. For additional viewing options, see below. Heat maps. Several heat map visualizations are available. Go to View in the menu bar, and select green red, rainbow, gray scale, or blue white from the heat map submenu. Table 2 summarizes the key attributes of the available heat map color modes. Table 2 Heat map High Value Low Value Middle Value(s) Mode (R,G,B) (R,G,B) green red Green (0,255,0) Red (255,0,0) Black rainbow Red (255,0,0) Blue (0,0,255) Blue, Light Blue, Green, Yellow, Red (evenly spaced) gray scale Black (0,0,0) White shades of Gray (255,255,255) blue white Blue (0,0,255) White (255,255,255) shades of Blue The mouse scrollbar is used to zoom in or out. To fit the heat map to the screen, go to view>fit to screen, or right-click your mouse when hovering over the heat map display. By default, cluster confidence is shown as a vertical bar to the left of the heat map with blue=high and red=low confidence (see rainbow-colored heat map in Figure 11 below). For fine-grained control over heat map visualization parameters, select view>settings>image settings (see below). Figure 11 11
14 Signal plots. As indicated in Figure 12, signal plots display all numerical vectors in the selected cluster(s) as a line graph across all column categories (x-axis). Go to View in the menu bar, and select rainbow or red from the signal plot submenu. i) Scale bar: By default, a scale bar is shown on the y-axis. Deselect the scale bar checkbox in View>signal plot to remove. To increase or decrease the scalebar resolution, use the up and down arrow keys in the number pad (8 and 2), respectively. To change the decimal precision of each number, use the left and right arrow keys in the number pad (4 and 6). ii) Mean signal: Select the mean signal checkbox in View>signal plot. Collapses signal plot into one line representing the mean of all vectors in the selected cluster(s). Figure 12 View>settings>Image Settings. The image settings window provides several options for customizing the heat map and signal plot display (see Figure 13 below). i) Dimensions: a. Adjust Height: Determines the vertical resolution of the image (can be used to compress or expand image height). The value of adjust height multiplied by the Zoom Factor (see below) gives the total number of vertical pixels for each heat map cell. b. Zoom Factor: This slider directly controls the number of horizontal pixels in each heat map cell. The overall image size is then adjusted by computing the dimensions (width x height in pixels) of each heat map cell=zoom Factor x (Adjust Height*Zoom Factor). Use this option for saving high-resolution images. ii) Heat Map Contrast: Adjusts the maximum and minimum values used for heat map display. Contrast C increases the original maximum value M and decreases the original minimum value m by (C-1)*(M-m)/2. a. Manually adjust range for contrast: Select this option to manually input maximum and minimum values for contrast adjustment. By default, the maximum and minimum values of the entire input data set are used. To see the maximum and minimum values for a particular cluster (or set of clusters), check the Input Data table on the left panel of the GUI. 12
15 iii) Normalization Tab: By selecting Display Original Data the heat map will render the cluster (s) using your original input data (pre-normalization). You can then select any of the data transformation options (e.g. log2 and median-centering) to re-normalize your data as desired. This option is useful for displaying a more meaningful range of values in the heat map (e.g. down to up-regulated genes). iv) Display Options Tab: Most of these options are self-explanatory. If you select Sort by Decreasing Variance the contents of every cluster will be sorted by decreasing order of row vector variance (e.g. gene probe variance). Although the heat map will update after changing most display settings, you can force the image to refresh by pressing Update. Click Save to launch a file browser for saving your heat map output. All images are saved in the Portable Network Graphics (PNG) format. Importantly, if you want to output a high-resolution image, make sure you select a large Zoom Factor before saving. Figure 13 View>settings>scale using entire dataset. When selected (go to View>settings), both the contrast in the heat map and y- axis for signal plots are scaled using the maximum and minimum values of the data set. When deselected, maximum and minimum values are set based on the selected cluster(s). Deselecting is useful for amplifying cluster-specific signals that are otherwise washed out in the context of the entire input data set. 13
16 Search. To find a particular data item in the cluster output, go to Search>Find. This will invoke a search window. Either find your data point from the Choose Identifier list of manually enter your data item identifier, then click Submit. If it is found, your data item will be highlighted in yellow using the heat map display (you might need to scroll down to find it). Please make sure your data items are uniquely labeled for this feature to work properly. Saving data. i) Images: All images can be saved by selecting File>Export>save image from the menu. ii) Tabular Data: To export tabular data from the selected cluster(s), go to File>Export>save tabular data. This option will save all cluster contents, along with cluster labels and cluster confidence for each data point. You can choose between saving clusters with your original data (pre-normalized) or with data after normalization has been applied ( current normalization ). The current normalization is dependent upon whether you have altered the heat map display using the Display Original Data feature in the image settings window (Figure 13). The output format is identical to output file 3 (see Files written to disk below). Files written to disk. AutoSOME writes the following files to your hard drive: X = No. ensemble iterations (e.g. E100) Y = p-value threshold (e.g. Pval0.1) Z = rows or columns, depending upon the value of Cluster Analysis (see pg. 9) 1) AutoSOME_inputName_EX_PvalY_Summary_Z.html = summary table of file name, parameters used, and cluster output table with size of each cluster, mean cluster confidence, and hyperlinks to data content of each cluster. 2) AutoSOME_inputName_EX_PvalY_Z.html = all clusters and data contents, including individual cluster confidence for each data point. 3) AutoSOME_inputName_EX_PvalY_Z.txt = text file with same data as file (2). First row = column labels (if provided as input), first column CLUST =cluster label, second column CONF =cluster confidence, third column NAME =data point label, all other columns=data vectors. 4) AutoSOME_inputName_rows_columns.txt = text file produced from clustering both rows and columns. Is in same format as (3) with the exception of an additional row (row 2) that stores cluster identifiers for the columns. 14
17 If your data set was normalized using AutoSOME, a code is stored using row 1 of output file (3) in order to properly reopen the file (see Opening old clustering results below). The code legend: #=input data set has column labels, n=log2 scaling, u=unit variance, sx=normalized from [0,X], m=median centering of rows, M=median centering of columns, q=sum of squares normalization of rows, Q=sum of squares normalization of columns. If columns are clustered (see Cluster Analysis, pg. 9), three additional files are written to disk: 5) AutoSOME_inputName_EX_PvalY_Edges.txt = fuzzy edges among data points: first two columns denote connected nodes, third column = pairwise affinity or the fraction of times the two data points were co-clustered (pairwise affinity ranges from -.5, never co-clustered, to 0.5, always co-clustered). 6) AutoSOME_inputName_EX_PvalY_Nodes.txt = All clustered data points: first column= unique data point label, second column=cluster label, third column=original data point label 7) AutoSOME_inputName_EX_PvalY_Matrix.txt = pairwise affinity matrix of all data points compared to all data points; essentially the same information content of output file (4) presented in a form suitable for a heat map display. This file can be immediately read as input into the Cluster 3.0 software [Eisen et al., 1998] to hierarchically reorder the matrix. Results can be visualized with Java TreeView [Saldanha, 2004]. Since Fuzzy Cluster Networks are computed from a distance matrix of all vertical data vectors (e.g. cell samples), output files (1)-(3) and (6) contain data points represented as a distance matrix (not the original input data). Opening previous clustering results. To browse previous clustering results, select Open AutoSOME Results from the File menu. Use the file browser to find the output text file, either file (3) or (4) in files written to disk above, and press Open. The GUI will be immediately redirected to the output window and display the cluster tree. All original data normalization settings are automatically restored for display in the output window. Reset. To reset all AutoSOME settings, go to File>Reset. Enhanced GUI vs. Basic GUI. The enhanced and basic GUIs offer identical functionality. The look and feel of the enhanced GUI makes use of the Substance freeware package ( while the basic GUI uses the default system look and feel. 15
18 AutoSOME Command Line Version >Usage: java -jar autosome_vxxxxxx.jar [Input] [Options] >maximum JVM memory recommended, e.g. java -Xmx1600m -Xms1600m -jar autosome_vxxxxxx.jar [Input] [Options] where XXXXXX=release date of AutoSOME (e.g ) To display all input parameters, run AutoSOME with the parameter o (letter o for options). As indicated below, the command line version of AutoSOME has many options not available in the GUI, including the option to use alternative clustering algorithms (i.e. K- Means, Hierarchical Clustering with four linkage types) and alternative dimensional reduction techniques (i.e. density-equalized SOM, normal SOM, and Sammon s Mapping). For alternative clustering methods, the number of input clusters needs to be specified using the k parameter (see below). In addition, all of these clustering methods, including AutoSOME, can be benchmarked with the b option as long as the data item labels in the input file correspond to numerical cluster labels starting with 1 (1,2,,No. clusters n). Examples using fictitious gene expression data set yeast.txt 1) Perform co-expression clustering using log2 scaling, unit variance and mediancentering of rows normalization, 100 ensemble runs, p-value threshold of 0.05, launch GUI to display results, and write files to C:\output. java -Xmx1600m -Xms1600m -jar autosome_v jar yeast.txt N j1 e100 p.05 v DC:\output 2) Perform transcriptome clustering using unit variance, Uncentered correlation, 100 ensemble runs, launch GUI to display results, and write files to C:\output. java -Xmx1600m -Xms1600m -jar autosome_v jar yeast.txt n1 Q3 e100 v DC:\output 3) Perform transcriptome clustering using unit variance, Pearson s correlation to build the distance matrix, K-means with 5 clusters, 20 ensemble iterations, and launch GUI to display results. java -Xmx1600m -Xms1600m -jar autosome_v jar yeast.txt n1 Q2 K k5 e20 v 16
19 All Command Line Parameters Parameter Description (Default) -t[integer] set number of threads (available CPUs) -e[integer] set number of runs to merge into ensemble (10) -p[0-1] set p-value threshold for minimum spanning tree clustering (0.1) -D[directory] set output directory (same directory as input file) -C read in column headers from first row of input file (auto-detect otherwise) -v launch cluster viewer (false) -v2 display previous clustering results: input=clustering output text file (false) -n[integer] normalize input by unit variance '-n1', to log2 '-n0' or into range [0,X] '-nx' (false) -j[1,2,3] 1=perform median center normalization on all rows;2=columns;3=rows and columns -u[1,2,3] 1=perform sum of squares=1 normalization on all rows;2=columns;3=rows and columns -N normalize input to log2 with unit variance (false) -# apply unit variance normalization to distance matrix (false) -w read in PCL-formatted input file (false) -W read in Gene Expression Omnibus Series Matrix-formatted input file (false) -Q transform columns from input into Euclidean distance matrix (false) -Q[2,3] distance matrix metric; 2 = Pearson's, 3 = Uncentered Correlation (Euclidean) -h[1,2,3,4] fill missing values; 1=means of rows;2=medians of rows;3=means of columns;4=medians of columns (means of rows) -b do benchmarking: 'F-measure, Precision, Recall, NMI, corrected Rand Index',[data items must be labeled: 1,2,3,...,total number of clusters] (false) -c[integer] set number of Monte Carlo simulations for MST clustering (10) -g[integer] set x of SOM grid xy, where y=x (square root of input size*2) -M[integer] set maximum x/y grid of SOM (30) -m[integer] set minimum x/y grid of SOM (5) -P set SOM distance metric to Pearson Correlation (Euclidean) -P2 set SOM distance metric to Uncentered Correlation (Euclidean) -s set SOM topology to square (circle) -i[integer] set number of SOM iterations (1000) -x[integer] set SOM error surface exponent (3) -r[power of 2] set density-equalizing cartogram resolution (64) -E disable Density-Equalizing Cartogram (false) -S invoke Sammon Mapping instead of SOM (false) -S[integer] set number of Sammon Mapping iterations (100) -k[integer] specify number of clusters in data set (false) -K invoke K-Means Clustering; requires option -k (false) -A invoke Agglomerative Clustering; requires option -k (false) -A[method] 1=Single, 2=Complete, 3=Average, 4=Ward's (4) -V print verbose output (false) 17
20 Creating Fuzzy Cluster Networks Fuzzy cluster networks highlight the fuzzy relationships among clustered data points using an intuitive two-dimensional network display. A powerful application of this approach is the visualization of differences between cell lines on the basis of differential gene expression. To display fuzzy cluster networks, the network visualization tool Cytoscape needs to be installed on your computer [Shannon et al., 2005]. Cytoscape is freely available from 1) Import your input file into AutoSOME, set fields and adjust input. 2) Select Fuzzy Cluster Networks and choose distance metric. (Please remember that only vertical data vectors are clustered (e.g. cell samples or time series of a microarray data set) 3) Run AutoSOME 4) Launch Cytoscape 5) In Cytoscape (All screenshots below taken from Cytoscape 2.6.0): a. Go to File>Import>Network from Table (Text/MS Excel) b. Go to Select File(s) and locate AutoSOME output file (4) containing all edges (see Files written to disk in Output above). c. Set Source Interaction to Column 1 and Target Interaction to Column 2. Finally, click on Column 3 in the data Preview window to activate it (it will turn blue). d. Select Import and then Close. A raw network will appear as a grid. 18
21 e. Go to File>Import>Attribute from Table (Text/MS Excel) f. Go to Select File(s) and locate AutoSOME output file (5) containing all nodes and attributes (see Files written to disk in Output above). g. Select Import. h. Change global properties of the network: i. In the Control Panel, select the VizMapper TM tab. ii. Click in the Defaults window (shows a source and target pair with a blue background). A new window will appear. iii. Select the Global tab in the bottom right. iv. Change the background color to white. v. Go back to the Node tab and change the NODE_BORDER_COLOR property to black and increase the NODE_LINE_WIDTH to 2. vi. Select Apply i. Using the VizMapper TM tab: j. Next to Node Label, click ID and select Column 3. All nodes should now be relabeled according to the original data labels. Minimize the Node Label property by selecting the minus icon. k. Under Unused Properties find Edge Color and double-click it. i. Select Column 3 as a value. ii. Then, select Continuous Mapper for Mapping Type. iii. Click on the black-to-white gradient next to Graphical View to launch a Gradient Editor. iv. There are two fixed triangles, one on each end, and two adjustable triangles. Double-click the two leftmost triangles and set their colors to pure red (255,0,0). Double-click the two rightmost triangles and set their colors to pure blue (0,0,255). Drag the leftmost adjustable triangle all the way to the left and likewise drag the rightmost triangle to the right until it stops. 19
22 l. Find Edge Line Width under Unused Properties and double-click it. i. Select Column 3 as a value. ii. Then, select Continuous Mapper for Mapping Type. iii. Click on the graph next to the Graphical View property to launch the Continuous Editor. iv. Adjust the minimum and maximum values denoted by red squares (double-click on squares for precision, otherwise slide squares up or down). For example, set minimum to 0.5 and maximum to 20. Then, exit Continuous Editor. m. Find Edge Opacity under Unused Properties and double-click it. i. Select Column 3 as a value. ii. Then, select Continuous Mapper for Mapping Type. iii. Click on the graph next to the Graphical View property to launch the Continuous Editor. iv. Adjust the minimum and maximum values denoted by red squares (double-click on squares for precision, otherwise slide squares up or down). For example, set minimum to 0.5 and maximum to 60. Then, exit Continuous Editor. 20
23 n. Finally, find Node Color under Unused Properties and double-click it. i. Select Column 2 as a value. ii. Then, select Discrete Mapping for Mapping Type. iii. Right click on Discrete Mapping and go to Generate Discrete Values > Rainbow 1. All nodes are now colored according to cluster labels. Adjust colors as necessary. o. Go to Layout in the main menu and select Settings i. Choose Force-Directed Layout for Layout Algorithm ii. Under Edge Weight Settings, set The minimum edge weight to consider to -0.5, and set The maximum edge weight to consider to 0.5. Further, select Column 3 from The edge attribute that contains the weights. iii. Press Execute Layout to run the layout algorithm. iv. Although the network topology is generally preserved, different runs of the layout algorithm can yield slightly different results in terms of network rotation and local node placement. To increase the repulsion between neighboring nodes (for evenly spaced nodes within a cluster), increase Default Node Mass under Algorithm settings. 21
24 Another layout algorithm that can yield comparable results is the Edge-weighted Spring Embedded algorithm. Before executing the layout, make sure the Edge Weight Settings are adjusted as in (ii) above. This layout algorithm can yield more evenly spaced nodes, but is less stable than Force-Directed Layout. Run a few times. v. Notice that all edges are slightly curved. To straighten edges, save and reopen the Cytoscape file. vi. At this point, make any desirable fine-grained changes to the Edge Color, Edge Line Width and Edge Opacity parameters to emphasize the fuzziness in the network. p. To export the final network, go to File > Export > Network View as Graphics q. Then, select file format and save image! r. To expedite this process for next time, start with your saved network. All visualization parameters will still be specified. Then, simply input your edge and node files and perform the layout. Note that for the Node Color property, it is important to assign colors again using the process given in step (n). 22
25 References 1) D haeseleer P: How does gene expression clustering work? Nature Biotechnology 2005, 23(12): ) Eisen MB, Spellman PT, Brown PO, Botstein D: Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA 1998, 95(25): ) Newman AM and Cooper JB: AutoSOME: a clustering method for identifying gene expression modules without prior knowledge of cluster number. BMC Bioinformatics 2010, 11:117. 4) Saldanha A: Java Treeview extensible visualization of microarray data. Bioinformatics 2004, 20(17): ) Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T: Cytoscape: A Software Environment for Integrated Models of Biomolecular Interaction Networks. Genome Research 2003, 13:
Click Trust to launch TableView.
 Visualizing Expression data using the Co-expression Tool Web service and TableView Introduction. TableView was written by James (Jim) E. Johnson and colleagues at the University of Minnesota Center for
Visualizing Expression data using the Co-expression Tool Web service and TableView Introduction. TableView was written by James (Jim) E. Johnson and colleagues at the University of Minnesota Center for
Numbers Basics Website:
 Website: http://etc.usf.edu/te/ Numbers is Apple's new spreadsheet application. It is installed as part of the iwork suite, which also includes the word processing program Pages and the presentation program
Website: http://etc.usf.edu/te/ Numbers is Apple's new spreadsheet application. It is installed as part of the iwork suite, which also includes the word processing program Pages and the presentation program
The Allen Human Brain Atlas offers three types of searches to allow a user to: (1) obtain gene expression data for specific genes (or probes) of
 Microarray Data MICROARRAY DATA Gene Search Boolean Syntax Differential Search Mouse Differential Search Search Results Gene Classification Correlative Search Download Search Results Data Visualization
Microarray Data MICROARRAY DATA Gene Search Boolean Syntax Differential Search Mouse Differential Search Search Results Gene Classification Correlative Search Download Search Results Data Visualization
Models for Nurses: Quadratic Model ( ) Linear Model Dx ( ) x Models for Doctors:
 The goal of this technology assignment is to graph several formulas in Excel. This assignment assumes that you using Excel 2007. The formula you will graph is a rational function formed from two polynomials,
The goal of this technology assignment is to graph several formulas in Excel. This assignment assumes that you using Excel 2007. The formula you will graph is a rational function formed from two polynomials,
Working with Charts Stratum.Viewer 6
 Working with Charts Stratum.Viewer 6 Getting Started Tasks Additional Information Access to Charts Introduction to Charts Overview of Chart Types Quick Start - Adding a Chart to a View Create a Chart with
Working with Charts Stratum.Viewer 6 Getting Started Tasks Additional Information Access to Charts Introduction to Charts Overview of Chart Types Quick Start - Adding a Chart to a View Create a Chart with
SIVIC GUI Overview. SIVIC GUI Layout Overview
 SIVIC GUI Overview SIVIC GUI Layout Overview At the top of the SIVIC GUI is a row of buttons called the Toolbar. It is a quick interface for loading datasets, controlling how the mouse manipulates the
SIVIC GUI Overview SIVIC GUI Layout Overview At the top of the SIVIC GUI is a row of buttons called the Toolbar. It is a quick interface for loading datasets, controlling how the mouse manipulates the
MetScape User Manual
 MetScape 2.3.2 User Manual A Plugin for Cytoscape National Center for Integrative Biomedical Informatics July 2012 2011 University of Michigan This work is supported by the National Center for Integrative
MetScape 2.3.2 User Manual A Plugin for Cytoscape National Center for Integrative Biomedical Informatics July 2012 2011 University of Michigan This work is supported by the National Center for Integrative
Insight: Measurement Tool. User Guide
 OMERO Beta v2.2: Measurement Tool User Guide - 1 - October 2007 Insight: Measurement Tool User Guide Open Microscopy Environment: http://www.openmicroscopy.org OMERO Beta v2.2: Measurement Tool User Guide
OMERO Beta v2.2: Measurement Tool User Guide - 1 - October 2007 Insight: Measurement Tool User Guide Open Microscopy Environment: http://www.openmicroscopy.org OMERO Beta v2.2: Measurement Tool User Guide
Desktop Studio: Charts. Version: 7.3
 Desktop Studio: Charts Version: 7.3 Copyright 2015 Intellicus Technologies This document and its content is copyrighted material of Intellicus Technologies. The content may not be copied or derived from,
Desktop Studio: Charts Version: 7.3 Copyright 2015 Intellicus Technologies This document and its content is copyrighted material of Intellicus Technologies. The content may not be copied or derived from,
Section 8. 8 Format Editor
 Section 8 8 Format Editor The Format Editor allows the creation and editing of the log presentation or format files. The output of the format editor are files of the type *.prs which are subsequently used
Section 8 8 Format Editor The Format Editor allows the creation and editing of the log presentation or format files. The output of the format editor are files of the type *.prs which are subsequently used
mirnet Tutorial Starting with expression data
 mirnet Tutorial Starting with expression data Computer and Browser Requirements A modern web browser with Java Script enabled Chrome, Safari, Firefox, and Internet Explorer 9+ For best performance and
mirnet Tutorial Starting with expression data Computer and Browser Requirements A modern web browser with Java Script enabled Chrome, Safari, Firefox, and Internet Explorer 9+ For best performance and
Solo 4.6 Release Notes
 June9, 2017 (Updated to include Solo 4.6.4 changes) Solo 4.6 Release Notes This release contains a number of new features, as well as enhancements to the user interface and overall performance. Together
June9, 2017 (Updated to include Solo 4.6.4 changes) Solo 4.6 Release Notes This release contains a number of new features, as well as enhancements to the user interface and overall performance. Together
EXCEL 2003 DISCLAIMER:
 EXCEL 2003 DISCLAIMER: This reference guide is meant for experienced Microsoft Excel users. It provides a list of quick tips and shortcuts for familiar features. This guide does NOT replace training or
EXCEL 2003 DISCLAIMER: This reference guide is meant for experienced Microsoft Excel users. It provides a list of quick tips and shortcuts for familiar features. This guide does NOT replace training or
Microarray Excel Hands-on Workshop Handout
 Microarray Excel Hands-on Workshop Handout Piali Mukherjee (pim2001@med.cornell.edu; http://icb.med.cornell.edu/) Importing Data Excel allows you to import data in tab, comma or space delimited text formats.
Microarray Excel Hands-on Workshop Handout Piali Mukherjee (pim2001@med.cornell.edu; http://icb.med.cornell.edu/) Importing Data Excel allows you to import data in tab, comma or space delimited text formats.
Desktop Studio: Charts
 Desktop Studio: Charts Intellicus Enterprise Reporting and BI Platform Intellicus Technologies info@intellicus.com www.intellicus.com Working with Charts i Copyright 2011 Intellicus Technologies This document
Desktop Studio: Charts Intellicus Enterprise Reporting and BI Platform Intellicus Technologies info@intellicus.com www.intellicus.com Working with Charts i Copyright 2011 Intellicus Technologies This document
Data Should Not be a Four Letter Word Microsoft Excel QUICK TOUR
 Toolbar Tour AutoSum + more functions Chart Wizard Currency, Percent, Comma Style Increase-Decrease Decimal Name Box Chart Wizard QUICK TOUR Name Box AutoSum Numeric Style Chart Wizard Formula Bar Active
Toolbar Tour AutoSum + more functions Chart Wizard Currency, Percent, Comma Style Increase-Decrease Decimal Name Box Chart Wizard QUICK TOUR Name Box AutoSum Numeric Style Chart Wizard Formula Bar Active
Microsoft Office Excel
 Microsoft Office 2007 - Excel Help Click on the Microsoft Office Excel Help button in the top right corner. Type the desired word in the search box and then press the Enter key. Choose the desired topic
Microsoft Office 2007 - Excel Help Click on the Microsoft Office Excel Help button in the top right corner. Type the desired word in the search box and then press the Enter key. Choose the desired topic
User Guide. v7.5. September 4, For the most recent version of this document, visit kcura's Documentation Site.
 User Guide v7.5 September 4, 2013 For the most recent version of this document, visit kcura's Documentation Site. Table of Contents 1 User guide overview 4 2 Relativity objects 4 3 Workspace 6 3.1 Workspaces
User Guide v7.5 September 4, 2013 For the most recent version of this document, visit kcura's Documentation Site. Table of Contents 1 User guide overview 4 2 Relativity objects 4 3 Workspace 6 3.1 Workspaces
IMAGE STUDIO LITE. Tutorial Guide Featuring Image Studio Analysis Software Version 3.1
 IMAGE STUDIO LITE Tutorial Guide Featuring Image Studio Analysis Software Version 3.1 Notice The information contained in this document is subject to change without notice. LI-COR MAKES NO WARRANTY OF
IMAGE STUDIO LITE Tutorial Guide Featuring Image Studio Analysis Software Version 3.1 Notice The information contained in this document is subject to change without notice. LI-COR MAKES NO WARRANTY OF
SUM - This says to add together cells F28 through F35. Notice that it will show your result is
 COUNTA - The COUNTA function will examine a set of cells and tell you how many cells are not empty. In this example, Excel analyzed 19 cells and found that only 18 were not empty. COUNTBLANK - The COUNTBLANK
COUNTA - The COUNTA function will examine a set of cells and tell you how many cells are not empty. In this example, Excel analyzed 19 cells and found that only 18 were not empty. COUNTBLANK - The COUNTBLANK
Technical Documentation Version 7.3 Output
 Technical Documentation Version 7.3 Output These documents are copyrighted by the Regents of the University of Colorado. No part of this document may be reproduced, stored in a retrieval system, or transmitted
Technical Documentation Version 7.3 Output These documents are copyrighted by the Regents of the University of Colorado. No part of this document may be reproduced, stored in a retrieval system, or transmitted
m6aviewer Version Documentation
 m6aviewer Version 1.6.0 Documentation Contents 1. About 2. Requirements 3. Launching m6aviewer 4. Running Time Estimates 5. Basic Peak Calling 6. Running Modes 7. Multiple Samples/Sample Replicates 8.
m6aviewer Version 1.6.0 Documentation Contents 1. About 2. Requirements 3. Launching m6aviewer 4. Running Time Estimates 5. Basic Peak Calling 6. Running Modes 7. Multiple Samples/Sample Replicates 8.
Tree and Data Grid for Micro Charts User Guide
 COMPONENTS FOR XCELSIUS Tree and Data Grid for Micro Charts User Guide Version 1.1 Inovista Copyright 2009 All Rights Reserved Page 1 TABLE OF CONTENTS Components for Xcelsius... 1 Introduction... 4 Data
COMPONENTS FOR XCELSIUS Tree and Data Grid for Micro Charts User Guide Version 1.1 Inovista Copyright 2009 All Rights Reserved Page 1 TABLE OF CONTENTS Components for Xcelsius... 1 Introduction... 4 Data
TraceFinder Analysis Quick Reference Guide
 TraceFinder Analysis Quick Reference Guide This quick reference guide describes the Analysis mode tasks assigned to the Technician role in the Thermo TraceFinder 3.0 analytical software. For detailed descriptions
TraceFinder Analysis Quick Reference Guide This quick reference guide describes the Analysis mode tasks assigned to the Technician role in the Thermo TraceFinder 3.0 analytical software. For detailed descriptions
Creating Web Pages with SeaMonkey Composer
 1 of 26 6/13/2011 11:26 PM Creating Web Pages with SeaMonkey Composer SeaMonkey Composer lets you create your own web pages and publish them on the web. You don't have to know HTML to use Composer; it
1 of 26 6/13/2011 11:26 PM Creating Web Pages with SeaMonkey Composer SeaMonkey Composer lets you create your own web pages and publish them on the web. You don't have to know HTML to use Composer; it
CounselLink Reporting. Designer
 CounselLink Reporting Designer Contents Overview... 1 Introduction to the Document Editor... 2 Create a new document:... 2 Document Templates... 3 Datasets... 3 Document Structure... 3 Layout Area... 4
CounselLink Reporting Designer Contents Overview... 1 Introduction to the Document Editor... 2 Create a new document:... 2 Document Templates... 3 Datasets... 3 Document Structure... 3 Layout Area... 4
ViTraM: VIsualization of TRAnscriptional Modules
 ViTraM: VIsualization of TRAnscriptional Modules Version 1.0 June 1st, 2009 Hong Sun, Karen Lemmens, Tim Van den Bulcke, Kristof Engelen, Bart De Moor and Kathleen Marchal KULeuven, Belgium 1 Contents
ViTraM: VIsualization of TRAnscriptional Modules Version 1.0 June 1st, 2009 Hong Sun, Karen Lemmens, Tim Van den Bulcke, Kristof Engelen, Bart De Moor and Kathleen Marchal KULeuven, Belgium 1 Contents
SAS Visual Analytics 8.2: Working with Report Content
 SAS Visual Analytics 8.2: Working with Report Content About Objects After selecting your data source and data items, add one or more objects to display the results. SAS Visual Analytics provides objects
SAS Visual Analytics 8.2: Working with Report Content About Objects After selecting your data source and data items, add one or more objects to display the results. SAS Visual Analytics provides objects
Tutorial:Basic Expression Analysis in Cytoscape
 Tutorial:Basic Expression Analysis in Cytoscape 1 Tutorial:Basic Expression Analysis in Cytoscape Slideshow Basic Expression Analysis in Cytoscape (30 min) [1] Handout Basic_Expression_Analysis_in_Cytoscape.pdf
Tutorial:Basic Expression Analysis in Cytoscape 1 Tutorial:Basic Expression Analysis in Cytoscape Slideshow Basic Expression Analysis in Cytoscape (30 min) [1] Handout Basic_Expression_Analysis_in_Cytoscape.pdf
EXERCISE: GETTING STARTED WITH SAV
 Sequencing Analysis Viewer (SAV) Overview 1 EXERCISE: GETTING STARTED WITH SAV Purpose This exercise explores the following topics: How to load run data into SAV How to explore run metrics with SAV Getting
Sequencing Analysis Viewer (SAV) Overview 1 EXERCISE: GETTING STARTED WITH SAV Purpose This exercise explores the following topics: How to load run data into SAV How to explore run metrics with SAV Getting
Tutorial: Jump Start on the Human Epigenome Browser at Washington University
 Tutorial: Jump Start on the Human Epigenome Browser at Washington University This brief tutorial aims to introduce some of the basic features of the Human Epigenome Browser, allowing users to navigate
Tutorial: Jump Start on the Human Epigenome Browser at Washington University This brief tutorial aims to introduce some of the basic features of the Human Epigenome Browser, allowing users to navigate
SNPViewer Documentation
 SNPViewer Documentation Module name: Description: Author: SNPViewer Displays SNP data plotting copy numbers and LOH values Jim Robinson (Broad Institute), gp-help@broad.mit.edu Summary: The SNPViewer displays
SNPViewer Documentation Module name: Description: Author: SNPViewer Displays SNP data plotting copy numbers and LOH values Jim Robinson (Broad Institute), gp-help@broad.mit.edu Summary: The SNPViewer displays
Excel 2016 Basics for Mac
 Excel 2016 Basics for Mac Excel 2016 Basics for Mac Training Objective To learn the tools and features to get started using Excel 2016 more efficiently and effectively. What you can expect to learn from
Excel 2016 Basics for Mac Excel 2016 Basics for Mac Training Objective To learn the tools and features to get started using Excel 2016 more efficiently and effectively. What you can expect to learn from
Capstone Appendix. A guide to your lab computer software
 Capstone Appendix A guide to your lab computer software Important Notes Many of the Images will look slightly different from what you will see in lab. This is because each lab setup is different and so
Capstone Appendix A guide to your lab computer software Important Notes Many of the Images will look slightly different from what you will see in lab. This is because each lab setup is different and so
Inserting Information into PowerPoint
 LESSON 6 6.1 Inserting Information into PowerPoint After completing this lesson, you will be able to: Change the layout of a slide. Insert a clip art image. Scale an image. Insert and format a table. Insert
LESSON 6 6.1 Inserting Information into PowerPoint After completing this lesson, you will be able to: Change the layout of a slide. Insert a clip art image. Scale an image. Insert and format a table. Insert
Elixir Ad-hoc Report. Release Elixir Technology Pte Ltd
 Elixir Ad-hoc Report Release 3.5.0 Elixir Technology Pte Ltd Elixir Ad-hoc Report: Release 3.5.0 Elixir Technology Pte Ltd Published 2014 Copyright 2014 Elixir Technology Pte Ltd All rights reserved. Java
Elixir Ad-hoc Report Release 3.5.0 Elixir Technology Pte Ltd Elixir Ad-hoc Report: Release 3.5.0 Elixir Technology Pte Ltd Published 2014 Copyright 2014 Elixir Technology Pte Ltd All rights reserved. Java
All About PlexSet Technology Data Analysis in nsolver Software
 All About PlexSet Technology Data Analysis in nsolver Software PlexSet is a multiplexed gene expression technology which allows pooling of up to 8 samples per ncounter cartridge lane, enabling users to
All About PlexSet Technology Data Analysis in nsolver Software PlexSet is a multiplexed gene expression technology which allows pooling of up to 8 samples per ncounter cartridge lane, enabling users to
Microsoft Excel 2007
 Microsoft Excel 2007 1 Excel is Microsoft s Spreadsheet program. Spreadsheets are often used as a method of displaying and manipulating groups of data in an effective manner. It was originally created
Microsoft Excel 2007 1 Excel is Microsoft s Spreadsheet program. Spreadsheets are often used as a method of displaying and manipulating groups of data in an effective manner. It was originally created
Using the DATAMINE Program
 6 Using the DATAMINE Program 304 Using the DATAMINE Program This chapter serves as a user s manual for the DATAMINE program, which demonstrates the algorithms presented in this book. Each menu selection
6 Using the DATAMINE Program 304 Using the DATAMINE Program This chapter serves as a user s manual for the DATAMINE program, which demonstrates the algorithms presented in this book. Each menu selection
Website Management with the CMS
 Website Management with the CMS In Class Step-by-Step Guidebook Updated 12/22/2010 Quick Reference Links CMS Login http://staging.montgomerycollege.edu/cmslogin.aspx Sample Department Site URLs (staging
Website Management with the CMS In Class Step-by-Step Guidebook Updated 12/22/2010 Quick Reference Links CMS Login http://staging.montgomerycollege.edu/cmslogin.aspx Sample Department Site URLs (staging
Keynote 08 Basics Website:
 Website: http://etc.usf.edu/te/ Keynote is Apple's presentation application. Keynote is installed as part of the iwork suite, which also includes the word processing program Pages and the spreadsheet program
Website: http://etc.usf.edu/te/ Keynote is Apple's presentation application. Keynote is installed as part of the iwork suite, which also includes the word processing program Pages and the spreadsheet program
Introduction to Excel 2007
 Introduction to Excel 2007 These documents are based on and developed from information published in the LTS Online Help Collection (www.uwec.edu/help) developed by the University of Wisconsin Eau Claire
Introduction to Excel 2007 These documents are based on and developed from information published in the LTS Online Help Collection (www.uwec.edu/help) developed by the University of Wisconsin Eau Claire
EXCEL 2010 PROCEDURES
 EXCEL 2010 PROCEDURES Starting Excel 1 Click the Start 2 Click All Programs 3 Click the Microsoft Office folder icon 4 Click Microsoft Excel 2010 Naming and Saving (Ctrl+S) a Workbook 1 Click File 2 Click
EXCEL 2010 PROCEDURES Starting Excel 1 Click the Start 2 Click All Programs 3 Click the Microsoft Office folder icon 4 Click Microsoft Excel 2010 Naming and Saving (Ctrl+S) a Workbook 1 Click File 2 Click
MICROSOFT EXCEL Working with Charts
 MICROSOFT EXCEL 2010 Working with Charts Introduction to charts WORKING WITH CHARTS Charts basically represent your data graphically. The data here refers to numbers. In Excel, you have various types of
MICROSOFT EXCEL 2010 Working with Charts Introduction to charts WORKING WITH CHARTS Charts basically represent your data graphically. The data here refers to numbers. In Excel, you have various types of
Differential Expression Analysis at PATRIC
 Differential Expression Analysis at PATRIC The following step- by- step workflow is intended to help users learn how to upload their differential gene expression data to their private workspace using Expression
Differential Expression Analysis at PATRIC The following step- by- step workflow is intended to help users learn how to upload their differential gene expression data to their private workspace using Expression
2 The Stata user interface
 2 The Stata user interface The windows This chapter introduces the core of Stata s interface: its main windows, its toolbar, its menus, and its dialogs. The five main windows are the Review, Results, Command,
2 The Stata user interface The windows This chapter introduces the core of Stata s interface: its main windows, its toolbar, its menus, and its dialogs. The five main windows are the Review, Results, Command,
Microsoft How to Series
 Microsoft How to Series Getting Started with EXCEL 2007 A B C D E F Tabs Introduction to the Excel 2007 Interface The Excel 2007 Interface is comprised of several elements, with four main parts: Office
Microsoft How to Series Getting Started with EXCEL 2007 A B C D E F Tabs Introduction to the Excel 2007 Interface The Excel 2007 Interface is comprised of several elements, with four main parts: Office
Status Bar: Right click on the Status Bar to add or remove features.
 Excel 2013 Quick Start Guide The Excel Window File Tab: Click to access actions like Print, Save As, etc. Also to set Excel options. Ribbon: Logically organizes actions onto Tabs, Groups, and Buttons to
Excel 2013 Quick Start Guide The Excel Window File Tab: Click to access actions like Print, Save As, etc. Also to set Excel options. Ribbon: Logically organizes actions onto Tabs, Groups, and Buttons to
Excel Select a template category in the Office.com Templates section. 5. Click the Download button.
 Microsoft QUICK Excel 2010 Source Getting Started The Excel Window u v w z Creating a New Blank Workbook 2. Select New in the left pane. 3. Select the Blank workbook template in the Available Templates
Microsoft QUICK Excel 2010 Source Getting Started The Excel Window u v w z Creating a New Blank Workbook 2. Select New in the left pane. 3. Select the Blank workbook template in the Available Templates
40. Sim Module - Common Tools
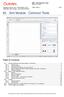 HSC Sim Common Tools 15021-ORC-J 1 (33) 40. Sim Module - Common Tools Table of Contents 40.1. Drawing flowsheets and adding tables to flowsheets... 2 40.1.1. Drawing units... 2 40.1.2. Drawing streams...
HSC Sim Common Tools 15021-ORC-J 1 (33) 40. Sim Module - Common Tools Table of Contents 40.1. Drawing flowsheets and adding tables to flowsheets... 2 40.1.1. Drawing units... 2 40.1.2. Drawing streams...
Excel 2016 Basics for Windows
 Excel 2016 Basics for Windows Excel 2016 Basics for Windows Training Objective To learn the tools and features to get started using Excel 2016 more efficiently and effectively. What you can expect to learn
Excel 2016 Basics for Windows Excel 2016 Basics for Windows Training Objective To learn the tools and features to get started using Excel 2016 more efficiently and effectively. What you can expect to learn
FrontPage 2000 Tutorial -- Advanced
 FrontPage 2000 Tutorial -- Advanced Shared Borders Shared Borders are parts of the web page that share content with the other pages in the web. They are located at the top, bottom, left side, or right
FrontPage 2000 Tutorial -- Advanced Shared Borders Shared Borders are parts of the web page that share content with the other pages in the web. They are located at the top, bottom, left side, or right
Highway Performance Monitoring System
 Highway Performance Monitoring System Version 1.0 June 2011 Quick Start Guide for Version 8.0 Federal Highway Administration Table of Contents Chapter 1 Introduction... 1 Chapter 2 HPMS Workflow... 2 Chapter
Highway Performance Monitoring System Version 1.0 June 2011 Quick Start Guide for Version 8.0 Federal Highway Administration Table of Contents Chapter 1 Introduction... 1 Chapter 2 HPMS Workflow... 2 Chapter
Version 2.4 of Idiogrid
 Version 2.4 of Idiogrid Structural and Visual Modifications 1. Tab delimited grids in Grid Data window. The most immediately obvious change to this newest version of Idiogrid will be the tab sheets that
Version 2.4 of Idiogrid Structural and Visual Modifications 1. Tab delimited grids in Grid Data window. The most immediately obvious change to this newest version of Idiogrid will be the tab sheets that
Gwenview User Manual. Aurélien Gâteau Christopher Martin Henry de Valence
 Aurélien Gâteau Christopher Martin Henry de Valence 2 Contents 1 Introduction 5 1.1 What is Gwenview..................................... 5 2 The Interface 6 2.1 Start Page..........................................
Aurélien Gâteau Christopher Martin Henry de Valence 2 Contents 1 Introduction 5 1.1 What is Gwenview..................................... 5 2 The Interface 6 2.1 Start Page..........................................
Elixir Ad-hoc Report. Release Elixir Technology Pte Ltd
 Elixir Ad-hoc Report Release 4.0.0 Elixir Technology Pte Ltd Elixir Ad-hoc Report: Release 4.0.0 Elixir Technology Pte Ltd Published 2015 Copyright 2015 Elixir Technology Pte Ltd All rights reserved. Java
Elixir Ad-hoc Report Release 4.0.0 Elixir Technology Pte Ltd Elixir Ad-hoc Report: Release 4.0.0 Elixir Technology Pte Ltd Published 2015 Copyright 2015 Elixir Technology Pte Ltd All rights reserved. Java
Blackboard for Faculty: Grade Center (631) In this document:
 1 Blackboard for Faculty: Grade Center (631) 632-2777 Teaching, Learning + Technology Stony Brook University In this document: blackboard@stonybrook.edu http://it.stonybrook.edu 1. What is the Grade Center?..
1 Blackboard for Faculty: Grade Center (631) 632-2777 Teaching, Learning + Technology Stony Brook University In this document: blackboard@stonybrook.edu http://it.stonybrook.edu 1. What is the Grade Center?..
Creating a Pivot Table
 Contents Introduction... 1 Creating a Pivot Table... 1 A One-Dimensional Table... 2 A Two-Dimensional Table... 4 A Three-Dimensional Table... 5 Hiding and Showing Summary Values... 5 Adding New Data and
Contents Introduction... 1 Creating a Pivot Table... 1 A One-Dimensional Table... 2 A Two-Dimensional Table... 4 A Three-Dimensional Table... 5 Hiding and Showing Summary Values... 5 Adding New Data and
In this chapter we explain the structure and use of BestDose, with examples.
 Using BestDose In this chapter we explain the structure and use of BestDose, with examples. General description The image below shows the view you get when you start BestDose. The actual view is called
Using BestDose In this chapter we explain the structure and use of BestDose, with examples. General description The image below shows the view you get when you start BestDose. The actual view is called
WORD Creating Objects: Tables, Charts and More
 WORD 2007 Creating Objects: Tables, Charts and More Microsoft Office 2007 TABLE OF CONTENTS TABLES... 1 TABLE LAYOUT... 1 TABLE DESIGN... 2 CHARTS... 4 PICTURES AND DRAWINGS... 8 USING DRAWINGS... 8 Drawing
WORD 2007 Creating Objects: Tables, Charts and More Microsoft Office 2007 TABLE OF CONTENTS TABLES... 1 TABLE LAYOUT... 1 TABLE DESIGN... 2 CHARTS... 4 PICTURES AND DRAWINGS... 8 USING DRAWINGS... 8 Drawing
INTRODUCTION... 1 UNDERSTANDING CELLS... 2 CELL CONTENT... 4
 Introduction to Microsoft Excel 2016 INTRODUCTION... 1 The Excel 2016 Environment... 1 Worksheet Views... 2 UNDERSTANDING CELLS... 2 Select a Cell Range... 3 CELL CONTENT... 4 Enter and Edit Data... 4
Introduction to Microsoft Excel 2016 INTRODUCTION... 1 The Excel 2016 Environment... 1 Worksheet Views... 2 UNDERSTANDING CELLS... 2 Select a Cell Range... 3 CELL CONTENT... 4 Enter and Edit Data... 4
RITIS Training Module 10 Script. To return to the Florida Analytics main page, select Florida Analytics Tools in the upper left corner of the page.
 RITIS Training Module 10 Script Welcome to the Regional Integrated Transportation Information System or RITIS Module 10 CBT. To begin, select the start button or press Shift+N on your keyboard. To return
RITIS Training Module 10 Script Welcome to the Regional Integrated Transportation Information System or RITIS Module 10 CBT. To begin, select the start button or press Shift+N on your keyboard. To return
Working with PDF s. To open a recent file on the Start screen, double click on the file name.
 Working with PDF s Acrobat DC Start Screen (Home Tab) When Acrobat opens, the Acrobat Start screen (Home Tab) populates displaying a list of recently opened files. The search feature on the top of the
Working with PDF s Acrobat DC Start Screen (Home Tab) When Acrobat opens, the Acrobat Start screen (Home Tab) populates displaying a list of recently opened files. The search feature on the top of the
3DProIN: Protein-Protein Interaction Networks and Structure Visualization
 Columbia International Publishing American Journal of Bioinformatics and Computational Biology doi:10.7726/ajbcb.2014.1003 Research Article 3DProIN: Protein-Protein Interaction Networks and Structure Visualization
Columbia International Publishing American Journal of Bioinformatics and Computational Biology doi:10.7726/ajbcb.2014.1003 Research Article 3DProIN: Protein-Protein Interaction Networks and Structure Visualization
Exploratory data analysis for microarrays
 Exploratory data analysis for microarrays Jörg Rahnenführer Computational Biology and Applied Algorithmics Max Planck Institute for Informatics D-66123 Saarbrücken Germany NGFN - Courses in Practical DNA
Exploratory data analysis for microarrays Jörg Rahnenführer Computational Biology and Applied Algorithmics Max Planck Institute for Informatics D-66123 Saarbrücken Germany NGFN - Courses in Practical DNA
FRAQCEL USER GUIDE
 FRAQCEL 2.7.1 USER GUIDE Author: Kevin Orloske Email Address: orloske.1@osu.edu Overview: This document provides user instructions for Fraqcel (pronounced frack-cell). Fraqcel is an open source fractal
FRAQCEL 2.7.1 USER GUIDE Author: Kevin Orloske Email Address: orloske.1@osu.edu Overview: This document provides user instructions for Fraqcel (pronounced frack-cell). Fraqcel is an open source fractal
PEOPLESOFT TIPS. TABLE OF CONTENTS Background... Error! Bookmark not defined. Navigate PeopleSoft... 3
 PEOPLESOFT TIPS TABLE OF CONTENTS Background... Error! Bookmark not defined. Navigate PeopleSoft... 3 Main Menu in PeopleSoft... 3 Sort the Main Menu... 3 Change the Default Sort Order... 5 Cascading Menu...
PEOPLESOFT TIPS TABLE OF CONTENTS Background... Error! Bookmark not defined. Navigate PeopleSoft... 3 Main Menu in PeopleSoft... 3 Sort the Main Menu... 3 Change the Default Sort Order... 5 Cascading Menu...
CHAPTER 4: MICROSOFT OFFICE: EXCEL 2010
 CHAPTER 4: MICROSOFT OFFICE: EXCEL 2010 Quick Summary A workbook an Excel document that stores data contains one or more pages called a worksheet. A worksheet or spreadsheet is stored in a workbook, and
CHAPTER 4: MICROSOFT OFFICE: EXCEL 2010 Quick Summary A workbook an Excel document that stores data contains one or more pages called a worksheet. A worksheet or spreadsheet is stored in a workbook, and
TUTORIAL - COMMAND CENTER
 FLOTHERM V3.1 Introductory Course TUTORIAL - COMMAND CENTER Introduction This tutorial covers the basic operation of the Command Center Application Window (CC) by walking the user through the main steps
FLOTHERM V3.1 Introductory Course TUTORIAL - COMMAND CENTER Introduction This tutorial covers the basic operation of the Command Center Application Window (CC) by walking the user through the main steps
Report Designer Report Types Table Report Multi-Column Report Label Report Parameterized Report Cross-Tab Report Drill-Down Report Chart with Static
 Table of Contents Report Designer Report Types Table Report Multi-Column Report Label Report Parameterized Report Cross-Tab Report Drill-Down Report Chart with Static Series Chart with Dynamic Series Master-Detail
Table of Contents Report Designer Report Types Table Report Multi-Column Report Label Report Parameterized Report Cross-Tab Report Drill-Down Report Chart with Static Series Chart with Dynamic Series Master-Detail
Advanced Excel. Click Computer if required, then click Browse.
 Advanced Excel 1. Using the Application 1.1. Working with spreadsheets 1.1.1 Open a spreadsheet application. Click the Start button. Select All Programs. Click Microsoft Excel 2013. 1.1.1 Close a spreadsheet
Advanced Excel 1. Using the Application 1.1. Working with spreadsheets 1.1.1 Open a spreadsheet application. Click the Start button. Select All Programs. Click Microsoft Excel 2013. 1.1.1 Close a spreadsheet
Manual. User Reference Guide. Analysis Application (EMG) Electromyography Analysis
 Phone: (888) 765-9735 WWW.MINDWARETECH.COM User Reference Guide Manual Analysis Application Electromyography Analysis (EMG) Copyright 2014 by MindWare Technologies LTD. All Rights Reserved. 1 Phone: (614)
Phone: (888) 765-9735 WWW.MINDWARETECH.COM User Reference Guide Manual Analysis Application Electromyography Analysis (EMG) Copyright 2014 by MindWare Technologies LTD. All Rights Reserved. 1 Phone: (614)
Tutorial: Using the SFLD and Cytoscape to Make Hypotheses About Enzyme Function for an Isoprenoid Synthase Superfamily Sequence
 Tutorial: Using the SFLD and Cytoscape to Make Hypotheses About Enzyme Function for an Isoprenoid Synthase Superfamily Sequence Requirements: 1. A web browser 2. The cytoscape program (available for download
Tutorial: Using the SFLD and Cytoscape to Make Hypotheses About Enzyme Function for an Isoprenoid Synthase Superfamily Sequence Requirements: 1. A web browser 2. The cytoscape program (available for download
About...1. Quick Start...2. Features and options...3. Thematic Map...3. Indicators Panel Graph Panel Options Panel...
 USER GUIDE TABLE OF CONTENTS About...1 Quick Start...2 Features and options...3 Thematic Map...3 Indicators Panel... 5 Graph Panel... 6 Options Panel... 7 Data-table Panel... 8 Selection Panel... 8 Time
USER GUIDE TABLE OF CONTENTS About...1 Quick Start...2 Features and options...3 Thematic Map...3 Indicators Panel... 5 Graph Panel... 6 Options Panel... 7 Data-table Panel... 8 Selection Panel... 8 Time
Genomics - Problem Set 2 Part 1 due Friday, 1/26/2018 by 9:00am Part 2 due Friday, 2/2/2018 by 9:00am
 Genomics - Part 1 due Friday, 1/26/2018 by 9:00am Part 2 due Friday, 2/2/2018 by 9:00am One major aspect of functional genomics is measuring the transcript abundance of all genes simultaneously. This was
Genomics - Part 1 due Friday, 1/26/2018 by 9:00am Part 2 due Friday, 2/2/2018 by 9:00am One major aspect of functional genomics is measuring the transcript abundance of all genes simultaneously. This was
Veco User Guides. Grids, Views, and Grid Reports
 Veco User Guides Grids, Views, and Grid Reports Introduction A Grid is defined as being a list of data records presented to the user. A grid is shown generally when an option is selected from the Tree
Veco User Guides Grids, Views, and Grid Reports Introduction A Grid is defined as being a list of data records presented to the user. A grid is shown generally when an option is selected from the Tree
Xfmea Version 10 First Steps Example
 Xfmea Version 10 First Steps Example This example provides a quick introduction to the Xfmea software by allowing you to experiment with the application s data management, analysis and reporting features.
Xfmea Version 10 First Steps Example This example provides a quick introduction to the Xfmea software by allowing you to experiment with the application s data management, analysis and reporting features.
Guide to WB Annotations
 Guide to WB Annotations 04 May 2016 Annotations are a powerful new feature added to Workbench v1.2.0 (Released May 2016) for placing text and symbols within wb_view tabs and windows. They enable generation
Guide to WB Annotations 04 May 2016 Annotations are a powerful new feature added to Workbench v1.2.0 (Released May 2016) for placing text and symbols within wb_view tabs and windows. They enable generation
Anima-LP. Version 2.1alpha. User's Manual. August 10, 1992
 Anima-LP Version 2.1alpha User's Manual August 10, 1992 Christopher V. Jones Faculty of Business Administration Simon Fraser University Burnaby, BC V5A 1S6 CANADA chris_jones@sfu.ca 1992 Christopher V.
Anima-LP Version 2.1alpha User's Manual August 10, 1992 Christopher V. Jones Faculty of Business Administration Simon Fraser University Burnaby, BC V5A 1S6 CANADA chris_jones@sfu.ca 1992 Christopher V.
BEAWebLogic Server. Using the WebLogic Diagnostic Framework Console Extension
 BEAWebLogic Server Using the WebLogic Diagnostic Framework Console Extension Version 10.0 Revised: March 30, 2007 Contents 1. Introduction and Roadmap What Is the WebLogic Diagnostic Framework Console
BEAWebLogic Server Using the WebLogic Diagnostic Framework Console Extension Version 10.0 Revised: March 30, 2007 Contents 1. Introduction and Roadmap What Is the WebLogic Diagnostic Framework Console
SAS Visual Analytics 8.2: Getting Started with Reports
 SAS Visual Analytics 8.2: Getting Started with Reports Introduction Reporting The SAS Visual Analytics tools give you everything you need to produce and distribute clear and compelling reports. SAS Visual
SAS Visual Analytics 8.2: Getting Started with Reports Introduction Reporting The SAS Visual Analytics tools give you everything you need to produce and distribute clear and compelling reports. SAS Visual
Guide to User Interface 4.3
 Datatel Colleague Guide to User Interface 4.3 Release 18 June 24, 2011 For corrections and clarifications to this manual, see AnswerNet page 1926.37. Guide to User Interface 4.3 All Rights Reserved The
Datatel Colleague Guide to User Interface 4.3 Release 18 June 24, 2011 For corrections and clarifications to this manual, see AnswerNet page 1926.37. Guide to User Interface 4.3 All Rights Reserved The
N2KExtractor. Maretron Data Extraction Software User s Manual
 N2KExtractor Maretron Data Extraction Software User s Manual Revision 3.1.6 Copyright 2017 Maretron, LLP All Rights Reserved Maretron, LLP 9014 N. 23rd Ave #10 Phoenix, AZ 85021-7850 http://www.maretron.com
N2KExtractor Maretron Data Extraction Software User s Manual Revision 3.1.6 Copyright 2017 Maretron, LLP All Rights Reserved Maretron, LLP 9014 N. 23rd Ave #10 Phoenix, AZ 85021-7850 http://www.maretron.com
Skills Funding Agency
 Provider Data Self-Assessment Toolkit (PDSAT) v17 User Guide Contents Introduction... 2 Compatibility and prerequisites... 2 1. Installing PDSAT... 3 2. Opening PDSAT... 6 2.1 Opening Screen... 6 2.2 Updates...
Provider Data Self-Assessment Toolkit (PDSAT) v17 User Guide Contents Introduction... 2 Compatibility and prerequisites... 2 1. Installing PDSAT... 3 2. Opening PDSAT... 6 2.1 Opening Screen... 6 2.2 Updates...
ViTraM: VIsualization of TRAnscriptional Modules
 ViTraM: VIsualization of TRAnscriptional Modules Version 2.0 October 1st, 2009 KULeuven, Belgium 1 Contents 1 INTRODUCTION AND INSTALLATION... 4 1.1 Introduction...4 1.2 Software structure...5 1.3 Requirements...5
ViTraM: VIsualization of TRAnscriptional Modules Version 2.0 October 1st, 2009 KULeuven, Belgium 1 Contents 1 INTRODUCTION AND INSTALLATION... 4 1.1 Introduction...4 1.2 Software structure...5 1.3 Requirements...5
Introduction to Microsoft Office PowerPoint 2010
 Introduction to Microsoft Office PowerPoint 2010 TABLE OF CONTENTS Open PowerPoint 2010... 1 About the Editing Screen... 1 Create a Title Slide... 6 Save Your Presentation... 6 Create a New Slide... 7
Introduction to Microsoft Office PowerPoint 2010 TABLE OF CONTENTS Open PowerPoint 2010... 1 About the Editing Screen... 1 Create a Title Slide... 6 Save Your Presentation... 6 Create a New Slide... 7
v SMS 12.2 Tutorial Observation Prerequisites Requirements Time minutes
 v. 12.2 SMS 12.2 Tutorial Observation Objectives This tutorial will give an overview of using the observation coverage in SMS. Observation points will be created to measure the numerical analysis with
v. 12.2 SMS 12.2 Tutorial Observation Objectives This tutorial will give an overview of using the observation coverage in SMS. Observation points will be created to measure the numerical analysis with
PEOPLESOFT TIPS. TABLE OF CONTENTS Overview... 3 Navigate PeopleSoft... 3
 PEOPLESOFT TIPS TABLE OF CONTENTS Overview... 3 Navigate PeopleSoft... 3 Main Menu in PeopleSoft... 3 Sort the Main Menu... 3 Cascading Menu... 5 Breadcrumb Trail Menu... 6 Search... 7 Searches: Maximum
PEOPLESOFT TIPS TABLE OF CONTENTS Overview... 3 Navigate PeopleSoft... 3 Main Menu in PeopleSoft... 3 Sort the Main Menu... 3 Cascading Menu... 5 Breadcrumb Trail Menu... 6 Search... 7 Searches: Maximum
User Manual. perfectionlearning.com/technical-support
 User Manual perfectionlearning.com/technical-support 1 User Manual Accessing Math X... 3 Login... 3 Forgotten Password... 3 Navigation Menu... 4 Logout... 4 Admin... 5 Creating Classes and Students...
User Manual perfectionlearning.com/technical-support 1 User Manual Accessing Math X... 3 Login... 3 Forgotten Password... 3 Navigation Menu... 4 Logout... 4 Admin... 5 Creating Classes and Students...
SEEK User Manual. Introduction
 SEEK User Manual Introduction SEEK is a computational gene co-expression search engine. It utilizes a vast human gene expression compendium to deliver fast, integrative, cross-platform co-expression analyses.
SEEK User Manual Introduction SEEK is a computational gene co-expression search engine. It utilizes a vast human gene expression compendium to deliver fast, integrative, cross-platform co-expression analyses.
Getting Started. In This Chapter
 Getting Started In This Chapter 2 This chapter introduces concepts and procedures that help you get started with AutoCAD. You learn how to open, close, and manage your drawings. You also learn about the
Getting Started In This Chapter 2 This chapter introduces concepts and procedures that help you get started with AutoCAD. You learn how to open, close, and manage your drawings. You also learn about the
Open Excel by following the directions listed below: Click on Start, select Programs, and the click on Microsoft Excel.
 Candy is Dandy Grading Rubric You have been hired to conduct some market research about M&M's. First, you had your team purchase 4 large bags and the results are given for the contents of those bags. You
Candy is Dandy Grading Rubric You have been hired to conduct some market research about M&M's. First, you had your team purchase 4 large bags and the results are given for the contents of those bags. You
epigenomegateway.wustl.edu
 Everything can be found at epigenomegateway.wustl.edu REFERENCES 1. Zhou X, et al., Nature Methods 8, 989-990 (2011) 2. Zhou X & Wang T, Current Protocols in Bioinformatics Unit 10.10 (2012) 3. Zhou X,
Everything can be found at epigenomegateway.wustl.edu REFERENCES 1. Zhou X, et al., Nature Methods 8, 989-990 (2011) 2. Zhou X & Wang T, Current Protocols in Bioinformatics Unit 10.10 (2012) 3. Zhou X,
User Guide. Version Exago Inc. All rights reserved.
 User Guide Version 2016.2 2016 Exago Inc. All rights reserved. Exago Reporting is a registered trademark of Exago, Inc. Windows is a registered trademark of Microsoft Corporation in the United States and
User Guide Version 2016.2 2016 Exago Inc. All rights reserved. Exago Reporting is a registered trademark of Exago, Inc. Windows is a registered trademark of Microsoft Corporation in the United States and
Workbench Tutorial Flow Over an Airfoil, Page 1 ANSYS Workbench Tutorial Flow Over an Airfoil
 Workbench Tutorial Flow Over an Airfoil, Page 1 ANSYS Workbench Tutorial Flow Over an Airfoil Authors: Scott Richards, Keith Martin, and John M. Cimbala, Penn State University Latest revision: 17 January
Workbench Tutorial Flow Over an Airfoil, Page 1 ANSYS Workbench Tutorial Flow Over an Airfoil Authors: Scott Richards, Keith Martin, and John M. Cimbala, Penn State University Latest revision: 17 January
Excel 2013 Intermediate
 Excel 2013 Intermediate Quick Access Toolbar... 1 Customizing Excel... 2 Keyboard Shortcuts... 2 Navigating the Spreadsheet... 2 Status Bar... 3 Worksheets... 3 Group Column/Row Adjusments... 4 Hiding
Excel 2013 Intermediate Quick Access Toolbar... 1 Customizing Excel... 2 Keyboard Shortcuts... 2 Navigating the Spreadsheet... 2 Status Bar... 3 Worksheets... 3 Group Column/Row Adjusments... 4 Hiding
Book 5. Chapter 1: Slides with SmartArt & Pictures... 1 Working with SmartArt Formatting Pictures Adjust Group Buttons Picture Styles Group Buttons
 Chapter 1: Slides with SmartArt & Pictures... 1 Working with SmartArt Formatting Pictures Adjust Group Buttons Picture Styles Group Buttons Chapter 2: Slides with Charts & Shapes... 12 Working with Charts
Chapter 1: Slides with SmartArt & Pictures... 1 Working with SmartArt Formatting Pictures Adjust Group Buttons Picture Styles Group Buttons Chapter 2: Slides with Charts & Shapes... 12 Working with Charts
FSA Algebra 1 EOC Practice Test Guide
 FSA Algebra 1 EOC Practice Test Guide This guide serves as a walkthrough of the Florida Standards Assessments (FSA) Algebra 1 End-of- Course (EOC) practice test. By reviewing the steps listed below, you
FSA Algebra 1 EOC Practice Test Guide This guide serves as a walkthrough of the Florida Standards Assessments (FSA) Algebra 1 End-of- Course (EOC) practice test. By reviewing the steps listed below, you
Tutorial: RNA-Seq Analysis Part II (Tracks): Non-Specific Matches, Mapping Modes and Expression measures
 : RNA-Seq Analysis Part II (Tracks): Non-Specific Matches, Mapping Modes and February 24, 2014 Sample to Insight : RNA-Seq Analysis Part II (Tracks): Non-Specific Matches, Mapping Modes and : RNA-Seq Analysis
: RNA-Seq Analysis Part II (Tracks): Non-Specific Matches, Mapping Modes and February 24, 2014 Sample to Insight : RNA-Seq Analysis Part II (Tracks): Non-Specific Matches, Mapping Modes and : RNA-Seq Analysis
Adobe Illustrator CS5 Part 2: Vector Graphic Effects
 CALIFORNIA STATE UNIVERSITY, LOS ANGELES INFORMATION TECHNOLOGY SERVICES Adobe Illustrator CS5 Part 2: Vector Graphic Effects Summer 2011, Version 1.0 Table of Contents Introduction...2 Downloading the
CALIFORNIA STATE UNIVERSITY, LOS ANGELES INFORMATION TECHNOLOGY SERVICES Adobe Illustrator CS5 Part 2: Vector Graphic Effects Summer 2011, Version 1.0 Table of Contents Introduction...2 Downloading the
