seam Documentation Release 0.0 Scott Burns
|
|
|
- Catherine Howard
- 6 years ago
- Views:
Transcription
1 seam Documentation Release 0.0 Scott Burns April 08, 2014
2
3 Contents 1 Contents Installation Philosophy Contributing Tools Support 9 3 License 11 Python Module Index 13 i
4 ii
5 Seam is a simple layer between neuroimaging tools and your data. It makes no decisions about how to organize your data or execute data analyses. It simply provides commands to intelligently call standard neuroimaging tools. Contents 1
6 2 Contents
7 CHAPTER 1 Contents 1.1 Installation Install the latest version with pip: $ pip install seam Seam is also distributed in the wheel format, so you can install it as a wheel: $ pip install --use-wheel seam Note, this requires the wheel package along with an up-to-date pip. To install the bleeding edge from the github repo, use the following: $ pip install -e git+ Either way, it has no dependencies. 1.2 Philosophy Software is best written in layers. Each layer should encapsulate knowledge about how to best use the next lower layer. Its functionality should be exposed through as simple an API as possible. Seam extends this rationale to neuroimaging data analysis. It is up to you to organize your data and run analyses, but seam will generate commands for you to apply your data against standard neuroimaging tools. Simply put, it builds commands around common neuroimaging tools. You re free to do with these commands whatever you wish. Importantly, seam is oblivious to these two otherwise-important factors: Data organization Command execution Seam has no dependencies and requires minimal effort to use it. Seam can easily integrate into any application ranging from a single script to something much more complicated Recipes vs Functions Where possible, functionality provided by seam is organized in two layers: 3
8 Functions: low-level, highly configurable functions to produce commands or scripts. Recipes: high-level, less configurable functions to produce complete workflows around a tool. Recipes will be exposed as both binaries that can be executed in a shell and as importable functions in python code. Users should first try to use recipes to accomplish their goals as these expose best practices around the tool. If more customization is needed, the low-level functions are also available. Understandably, more effort is required to piece together the low-level functions into a meaningful processing stream Backwards Compatiblity Seam makes every effort to ensure backward compatibility for the commands it generates. Because neuroimaging projects can last for very long periods, the commands generated by seam will be versioned so you can upgrade seam and still produce the same commands for a given project. Newer projects, though, can use newer versions of commands using the same seam. 1.3 Contributing If you would like to contribute commands or find errors and/or better ways to build commands, please consider contributing to seam. To do so, please use the following workflow: Fork the repository to your own account. Checkout an aptly-named branch and commit your changes. Please add tests (and documentation) and make sure they pass. You can use $ make test to run the suite. Push your commits to your fork and submit a pull-request. 1.4 Tools DTI_QA This interface assists in executing Bennett Landman s DTI_QA toolkit. TBD: More documentation here. Because DTI_QA is built upon matlab, the generated commands are m-code. Functions seam.dti_qa.dtiqa_mcode(images, basedir, dtiqa_path, n_b0=1) Returns m-code that can be executed in matlab to run DTI_QA Usage: Parameters images (str,list) path to single raw DTI image or list of multiple basedir (str) base directory to write results dtiqa_path (str) path to DTI_QA installation n_b0 (int) # of b0 averages (default is 1) 4 Chapter 1. Contents
9 >>> from seam.dti_qa.v1 import dtiqa_mcode >>> images = [ /path/to/first.nii, /path/to/second.nii ] >>> basedir, n_b0, dtiqa_path = /path/to/output, 6, /path/to/dtiqa >>> f = open( dti_qa.m, w ) >>> f.write(dtiqa_mcode(images, basedir, dtiqa_path, n_b0)) >>> f.close() Versioning: V1 defines the following: seam.dti_qa.v1.dtiqa_mcode() that generates m-code to run DTI_QA Freesurfer Freesurfer is a set of software tools for the study of cortical and subcortical anatomy. In the cortical surface stream, the tools construct models of the boundary between white matter and cortical gray matter as well as the pial surface. Once these surfaces are known, an array of anatomical measures becomes possible, including: cortical thickness, surface area, curvature, and surface normal at each point on the cortex. The surfaces can be inflated and/or flattened for improved visualization. The surfaces can also be used to constrain the solutions to inverse optical, EEG and MEG problems. In addition, a cortical surface-based atlas has been defined based on average folding patterns mapped to a sphere. Surfaces from individuals can be aligned with this atlas with a high-dimensional nonlinear registration algorithm. The registration is based on aligning the cortical folding patterns and so directly aligns the anatomy instead of image intensities. The spherical atlas naturally forms a coordinate system in which point-to-point correspondence between subjects can be achieved. This coordinate system can then be used to create group maps (similar to how Talairach space is used for volumetric measurements). Most of the FreeSurfer pipeline is automated, which makes it ideal for use on large data sets. Recipes seam.freesurfer.v1.recipe.build_recipe(subject_id, input_data, script_dir, use_xvfb=false, recon_flags=none) This function builds a complete pipeline around Freesufer. It does the following: Imports data using recon-all -i Runs the main recon-all command with the following flags: -qcache -measure thickness -measure curv -measure sulc -measure area -measure jacobian_white Takes volumetric snapshots using tkmedit Per hemisphere: Converts the aparc.a2009s.annot file to labels in the subject s label directory Takes screenshots of the inflated surface with and without the advanced labels Tools 5
10 Parameters subject_id (str) subject identifier input_data (str,list) list of paths or string to subject s T1 images script_dir (str) directory to write scripts & screenshots use_xvfb (boolean) Wrap tksurfer & tkmedit commands in xvfb-run, useful if running in a non-graphical (ie cluster) environment. recon_flags (list) other flags to pass to recon-all Return type tuple Returns paths to recon script, tkmedit script and lh & rh tksurfer scripts Note the main script is set as executable Note This function is exposed on the command line through build-recon-v1 Functions These are lower-level functions to be used when more customization is needed than provided by the above recipes. seam.freesurfer.v1.core.recon_input(subject_id, data) The function supplies the recon-all -i command. This command initializes the Freesurfer directory for a subject and converts the raw data into an internal format for use by the rest of the recon-all pipeline. Usage: Parameters subject_id (str) Subject identifier data (str,list) path(s) to input data Returns command that will execute recon-all -i command Return type str >>> from seam.freesurfer import recon_input >>> recon_input( sub0001, /full/path/to/data.nii ) recon-all -s sub0001 -i /full/path/to/data.nii >>> # or with multiple inputs... >>> recon_input( sub0001, [ /path/first.nii, /path/second.nii ]) recon-all -s sub0001 -i /path/first.nii -i /path/second.nii seam.freesurfer.v1.core.recon_all(subject_id, flags=none) This function supplies the recon-all -all command. This command will run the entire anatomical analysis suite of Freesurfer. Usage: Note Use seam.freesurfer.recon_input() to setup this subject Parameters subject_id (str) Subject identifier on which to run recon-all flags (list) command-line flags to pass to recon-all Returns command that will execute recon-all -all Return type str 6 Chapter 1. Contents
11 >>> from seam.freesurfer import recon_all >>> recon_all( sub0001, flags=[ -use-gpu ]) recon-all -s sub0001 -all -qcache -measure thickness -measure curv -measure sulc -measure area seam.freesurfer.v1.core.tkmedit_screenshot_tcl(basepath, beg=5, end=256, step=10) Supplies a tcl string that can be used to take screenshots of a volume using tkmedit Images are written to *basepath*/tkmedit-$i.tiff where beg <= i <= end in increments of step Usage: Parameters basepath base directory to write images in beg (int) Beginning slice where screenshots begin end (int) End slice where screenshots end step (int) Value to increment successive screenshots >>> from seam.freesurfer import tkmedit_screenshot_tcl >>> f = open( tmedit_screenshots.tcl, w ) >>> f.write(tkmedit_screenshot_tcl( /path/to/image_dir )) >>> f.close() $ tkmedit sub0001 brain.finalsurfs.mgz -aseg -surfs -tcl tkmedit_screenshots.tcl seam.freesurfer.v1.core.tkmedit_screenshot_cmd(subject_id, volume, tcl_path, flags=none) Supplies a command to execute a tcl script in tkmedit for subject_id s volume Usage: Parameters subject_id (str) subject identifier volume (str) Volume for tkmedit to load tcl_path (str) Path to tcl script flags (list) Flags to pass to tkmedit >>> from seam.freesurfer import tkmedit_screenshot_cmd >>> tkmedit_screenshot_cmd( sub0001, brain.finalsurfs.mgz, /path/tkmedit.tcl, [ -aseg, -s tkmedit sub0001 brain.finalsurfs.mgz -aseg -surfs -tcl /path/tkmedit.tcl an- seam.freesurfer.v1.core.tksurfer_screenshot_tcl(basepath, not= aparc.a2009s.annot ) Supplies a tcl command to take screenshots of a surface using tksurfer Four screenshots are taken: lateral view (default view when tksurfer opens) saved to basepath-lateral.tiff medial view saved to basepath-medial.tiff Lateral view with an annotation loaded (given by annot) saved to basepath-annot-lateral.tiff Medial view with an annotation loaded (given by annot) saved to basepath-annot-medial.tiff Parameters basepath (str) prefix for images to be saved annot (str) annotation file to load for overlay on the surface 1.4. Tools 7
12 Usage: >>> from seam.freesurfer import tksurfer_screenshot_tcl >>> f = open( tksurfer.lh.tcl, w ) >>> f.write(tksurfer_screenshot_tcl( /path/to/screenshots/lh )) >>> f.close() seam.freesurfer.v1.core.tksurfer_screenshot_cmd(subject_id, hemi, surface, tcl_path, flags=none) Supply a command that will run tksurfer using the surface from subject_id s hemi hemisphere and execute a tcl script. Usage: Parameters subject_id (str) subject identifier hemi (str) lh or rh, hemisphere to open in tksurfer surface (str) surface to view tcl_path (str) path to tcl script to execute flags (list) flags to pass into tksurfer >>> from seam.freesurfer import tksurfer_screenshot_cmd >>> tksurfer_screenshot_cmd( sub0001, lh, inflated, /path/tksurfer.lh.tcl, [ -gray ]) tksurfer sub0001 lh inflated -gray -tcl /path/tksurfer.lh.tcl seam.freesurfer.v1.core.annot2label_cmd(subject_id, hemi, annot_path, outdir, surface= white ) Build the mri_annotation2label commandline string. Parameters subject_id (str) subject identifier hemi (str) lh or rh, hemisphere to use annot_path (str) path to annotation file outdir (str) output directory to place labels surface (str) surface to use when generating coords in labels Versions: V1 defines the following recipes: seam.freesurfer.v1.build_recipe() for building a complete script for executing the recon-all pipeline. V1 defines the following functions: recon-all -all exposed through seam.freesurfer.v1.recon_all() recon-all -i exposed through seam.freesurfer.v1.recon_input() seam.freesurfer.v1.tkmedit_screenshot_tcl() for generating tcl to take screenshots of a volume loaded in tkmedit. seam.freesurfer.v1.tkmedit_screenshot_cmd() for supplying a command to execute tkmedit with a tcl script. seam.freesurfer.v1.tksurfer_screenshot_tcl() for generating a tcl script to take screenshots of a hemisphere using tksurfer 8 Chapter 1. Contents
13 seam.freesurfer.v1.tksurfer_screenshot_cmd() for supplying a command to run tksurfer and generate screenshots. seam.freesufer.v1.annot2label_cmd() for building a mri_annotation2label command Tools 9
14 10 Chapter 1. Contents
15 CHAPTER 2 Support If you are having problems, please raise an issue on Github. I can t guarantee support but promise to help where I can. Issue Tracker: Source Code: 11
16 12 Chapter 2. Support
17 CHAPTER 3 License The project is licensed under the MIT license. 13
18 14 Chapter 3. License
19 Python Module Index s seam.dti_qa, 4 seam.dti_qa.v1, 4 seam.freesurfer.v1, 7 15
1. FreeSurfer Tutorial
 1. FreeSurfer Tutorial FreeSurfer is brought to you by the Martinos Center for Biomedical Imaging (supported by NCRR/P41RR14075), Massachusetts General Hospital, Boston, MA USA 2012-03-27 15:02 top previous
1. FreeSurfer Tutorial FreeSurfer is brought to you by the Martinos Center for Biomedical Imaging (supported by NCRR/P41RR14075), Massachusetts General Hospital, Boston, MA USA 2012-03-27 15:02 top previous
Surface-based Analysis: Inter-subject Registration and Smoothing
 Surface-based Analysis: Inter-subject Registration and Smoothing Outline Exploratory Spatial Analysis Coordinate Systems 3D (Volumetric) 2D (Surface-based) Inter-subject registration Volume-based Surface-based
Surface-based Analysis: Inter-subject Registration and Smoothing Outline Exploratory Spatial Analysis Coordinate Systems 3D (Volumetric) 2D (Surface-based) Inter-subject registration Volume-based Surface-based
FreeSurfer Tutorial :02
 FreeSurfer Tutorial FreeSurfer is brought to you by the Martinos Center for Biomedical Imaging (supported by NCRR/P41RR14075), Massachusetts General Hospital, Boston, MA USA 2009-06-01 12:02 1 FSL/FreeSurfer
FreeSurfer Tutorial FreeSurfer is brought to you by the Martinos Center for Biomedical Imaging (supported by NCRR/P41RR14075), Massachusetts General Hospital, Boston, MA USA 2009-06-01 12:02 1 FSL/FreeSurfer
Introduc)on to FreeSurfer h0p://surfer.nmr.mgh.harvard.edu. Jenni Pacheco.
 Introduc)on to FreeSurfer h0p://surfer.nmr.mgh.harvard.edu Jenni Pacheco jpacheco@mail.utexas.edu Analyzing the individual subject What happens? How do I do that? Now what? Tools you will use recon all
Introduc)on to FreeSurfer h0p://surfer.nmr.mgh.harvard.edu Jenni Pacheco jpacheco@mail.utexas.edu Analyzing the individual subject What happens? How do I do that? Now what? Tools you will use recon all
FreeSurfer Tutorial :42
 FreeSurfer Tutorial FreeSurfer is brought to you by the Martinos Center for Biomedical Imaging (supported by NCRR/P41RR14075), Massachusetts General Hospital, Boston, MA USA 2010-05-12 13:42 1 FSL/FreeSurfer
FreeSurfer Tutorial FreeSurfer is brought to you by the Martinos Center for Biomedical Imaging (supported by NCRR/P41RR14075), Massachusetts General Hospital, Boston, MA USA 2010-05-12 13:42 1 FSL/FreeSurfer
3D Visualization of FreeSurfer Data
 3D Visualization of FreeSurfer Data Sonia Pujol, Ph.D. Silas Mann, B.Sc. Randy Gollub, MD., Ph.D. Surgical Planning Laboratory Athinoula A. Martinos Center Harvard University Acknowledgements NIH U54EB005149
3D Visualization of FreeSurfer Data Sonia Pujol, Ph.D. Silas Mann, B.Sc. Randy Gollub, MD., Ph.D. Surgical Planning Laboratory Athinoula A. Martinos Center Harvard University Acknowledgements NIH U54EB005149
UNC 4D Infant Cortical Surface Atlases, from Neonate to 6 Years of Age
 UNC 4D Infant Cortical Surface Atlases, from Neonate to 6 Years of Age Version 1.0 UNC 4D infant cortical surface atlases from neonate to 6 years of age contain 11 time points, including 1, 3, 6, 9, 12,
UNC 4D Infant Cortical Surface Atlases, from Neonate to 6 Years of Age Version 1.0 UNC 4D infant cortical surface atlases from neonate to 6 years of age contain 11 time points, including 1, 3, 6, 9, 12,
Introduc)on to FreeSurfer h0p://surfer.nmr.mgh.harvard.edu. Jenni Pacheco.
 Introduc)on to FreeSurfer h0p://surfer.nmr.mgh.harvard.edu Jenni Pacheco jpacheco@mail.utexas.edu fmri Integra)on Visualize individual fmri results on surface volume ROI Volume Study: Count number of voxels
Introduc)on to FreeSurfer h0p://surfer.nmr.mgh.harvard.edu Jenni Pacheco jpacheco@mail.utexas.edu fmri Integra)on Visualize individual fmri results on surface volume ROI Volume Study: Count number of voxels
The organization of the human cerebral cortex estimated by intrinsic functional connectivity
 1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
FreeSurfer Register Download Platforms: Linux, Mac, Windows.
 FreeSurfer http://surfer.nmr.mgh.harvard.edu Register Download Platforms: Linux, Mac, Windows freesurfer@nmr.mgh.harvard.edu Big Picture Input: T1-weighted (MPRAGE,SPGR) 1mm 3 resolution (.dcm) Output:
FreeSurfer http://surfer.nmr.mgh.harvard.edu Register Download Platforms: Linux, Mac, Windows freesurfer@nmr.mgh.harvard.edu Big Picture Input: T1-weighted (MPRAGE,SPGR) 1mm 3 resolution (.dcm) Output:
FreeSurfer Register Download Platforms: Linux, Mac, Windows.
 FreeSurfer http://surfer.nmr.mgh.harvard.edu Register Download Platforms: Linux, Mac, Windows freesurfer@nmr.mgh.harvard.edu Analyzing the Individual Subject in FreeSurfer What happens? How do I do that?
FreeSurfer http://surfer.nmr.mgh.harvard.edu Register Download Platforms: Linux, Mac, Windows freesurfer@nmr.mgh.harvard.edu Analyzing the Individual Subject in FreeSurfer What happens? How do I do that?
BrainSuite Lab Exercises. presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014
 BrainSuite Lab Exercises presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014 1. Opening and Displaying an MRI Start BrainSuite Drag and drop the T1 image from the native space
BrainSuite Lab Exercises presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014 1. Opening and Displaying an MRI Start BrainSuite Drag and drop the T1 image from the native space
FreeSurfer: Troubleshooting surfer.nmr.mgh.harvard.edu
 FreeSurfer: Troubleshooting surfer.nmr.mgh.harvard.edu 1 Hard and Soft Failures Categories of errors: Hard & Soft Failures Hard = recon-all quits before it finishes Soft = recon-all finishes but results
FreeSurfer: Troubleshooting surfer.nmr.mgh.harvard.edu 1 Hard and Soft Failures Categories of errors: Hard & Soft Failures Hard = recon-all quits before it finishes Soft = recon-all finishes but results
Anatomical Analysis with FreeSurfer surfer.nmr.mgh.harvard.edu
 Anatomical Analysis with FreeSurfer surfer.nmr.mgh.harvard.edu 1 Processing Stream Overview T1 Weighted Input Skull Stripping Volumetric Labeling Intensity Normalization Gyral Labeling Surface Extraction
Anatomical Analysis with FreeSurfer surfer.nmr.mgh.harvard.edu 1 Processing Stream Overview T1 Weighted Input Skull Stripping Volumetric Labeling Intensity Normalization Gyral Labeling Surface Extraction
Automated MR Image Analysis Pipelines
 Automated MR Image Analysis Pipelines Andy Simmons Centre for Neuroimaging Sciences, Kings College London Institute of Psychiatry. NIHR Biomedical Research Centre for Mental Health at IoP & SLAM. Neuroimaging
Automated MR Image Analysis Pipelines Andy Simmons Centre for Neuroimaging Sciences, Kings College London Institute of Psychiatry. NIHR Biomedical Research Centre for Mental Health at IoP & SLAM. Neuroimaging
Lilla Zöllei A.A. Martinos Center, MGH; Boston, MA
 Lilla Zöllei lzollei@nmr.mgh.harvard.edu A.A. Martinos Center, MGH; Boston, MA Bruce Fischl Gheorghe Postelnicu Jean Augustinack Anastasia Yendiki Allison Stevens Kristen Huber Sita Kakonoori + the FreeSurfer
Lilla Zöllei lzollei@nmr.mgh.harvard.edu A.A. Martinos Center, MGH; Boston, MA Bruce Fischl Gheorghe Postelnicu Jean Augustinack Anastasia Yendiki Allison Stevens Kristen Huber Sita Kakonoori + the FreeSurfer
Caret5 Tutorial Segmentation, Flattening, and Registration
 Caret5 Tutorial Segmentation, Flattening, and Registration John Harwell, Donna Dierker, Heather A. Drury, and David C. Van Essen Version 5.5 September 2006 Copyright 1995-2006 Washington University Washington
Caret5 Tutorial Segmentation, Flattening, and Registration John Harwell, Donna Dierker, Heather A. Drury, and David C. Van Essen Version 5.5 September 2006 Copyright 1995-2006 Washington University Washington
Caret Tutorial Contours and Sections April 17, 2007
 Caret Tutorial Contours and Sections April 17, 2007 John Harwell, Donna Dierker, and David C. Van Essen Washington University School of Medicine Department of Anatomy and Neurobiology Saint Louis, Missouri,
Caret Tutorial Contours and Sections April 17, 2007 John Harwell, Donna Dierker, and David C. Van Essen Washington University School of Medicine Department of Anatomy and Neurobiology Saint Louis, Missouri,
FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING. Francesca Pizzorni Ferrarese
 FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING Francesca Pizzorni Ferrarese Pipeline overview WM and GM Segmentation Registration Data reconstruction Tractography
FROM IMAGE RECONSTRUCTION TO CONNECTIVITY ANALYSIS: A JOURNEY THROUGH THE BRAIN'S WIRING Francesca Pizzorni Ferrarese Pipeline overview WM and GM Segmentation Registration Data reconstruction Tractography
Electrical Engineering, Vanderbilt University, Nashville, TN, USA b. Computer Science, Vanderbilt University, Nashville, TN, USA c
 Improved Stability of Whole Brain Surface Parcellation with Multi-Atlas Segmentation Yuankai Huo* a, Shunxing Bao b, Prasanna Parvathaneni a, Bennett A. Landman a,b,c a Electrical Engineering, Vanderbilt
Improved Stability of Whole Brain Surface Parcellation with Multi-Atlas Segmentation Yuankai Huo* a, Shunxing Bao b, Prasanna Parvathaneni a, Bennett A. Landman a,b,c a Electrical Engineering, Vanderbilt
MNE software version 2.6. Matti Hämäläinen. MGH/MIT/HMS Athinoula A. Martinos Center for Biomedical Imaging Charlestown, MA USA
 MNE software version 2.6 Matti Hämäläinen MGH/MIT/HMS Athinoula A. Martinos Center for Biomedical Imaging Charlestown, MA USA Overall changes Manual: description of the MEG/EEG and MRI coordinate systems
MNE software version 2.6 Matti Hämäläinen MGH/MIT/HMS Athinoula A. Martinos Center for Biomedical Imaging Charlestown, MA USA Overall changes Manual: description of the MEG/EEG and MRI coordinate systems
Github/Git Primer. Tyler Hague
 Github/Git Primer Tyler Hague Why Use Github? Github keeps all of our code up to date in one place Github tracks changes so we can see what is being worked on Github has issue tracking for keeping up with
Github/Git Primer Tyler Hague Why Use Github? Github keeps all of our code up to date in one place Github tracks changes so we can see what is being worked on Github has issue tracking for keeping up with
Multimodal Imaging Brain Connectivity Analysis (MIBCA)
 Multimodal Imaging Brain Connectivity Analysis (MIBCA) Andre Santos Ribeiro, Luis Miguel Lacerda, Hugo Ferreira April 23, 2015 Abstract In recent years, connectivity studies using neuroimaging data have
Multimodal Imaging Brain Connectivity Analysis (MIBCA) Andre Santos Ribeiro, Luis Miguel Lacerda, Hugo Ferreira April 23, 2015 Abstract In recent years, connectivity studies using neuroimaging data have
Machine learning improves automated cortical surface reconstruction in human MRI studies
 University of Iowa Iowa Research Online Theses and Dissertations Spring 2017 Machine learning improves automated cortical surface reconstruction in human MRI studies David G. Ellis University of Iowa Copyright
University of Iowa Iowa Research Online Theses and Dissertations Spring 2017 Machine learning improves automated cortical surface reconstruction in human MRI studies David G. Ellis University of Iowa Copyright
Commits and Commit Messages
 Commits and Commit Messages What is a commit? Small set of modifications to a code base Each commit should contain one (atomic) change Commits should be standalone (independent of other commits) Open Source
Commits and Commit Messages What is a commit? Small set of modifications to a code base Each commit should contain one (atomic) change Commits should be standalone (independent of other commits) Open Source
Redis Timeseries Documentation
 Redis Timeseries Documentation Release 0.1.8 Ryan Anguiano Jul 26, 2017 Contents 1 Redis Timeseries 3 1.1 Install................................................... 3 1.2 Usage...................................................
Redis Timeseries Documentation Release 0.1.8 Ryan Anguiano Jul 26, 2017 Contents 1 Redis Timeseries 3 1.1 Install................................................... 3 1.2 Usage...................................................
BanzaiDB Documentation
 BanzaiDB Documentation Release 0.3.0 Mitchell Stanton-Cook Jul 19, 2017 Contents 1 BanzaiDB documentation contents 3 2 Indices and tables 11 i ii BanzaiDB is a tool for pairing Microbial Genomics Next
BanzaiDB Documentation Release 0.3.0 Mitchell Stanton-Cook Jul 19, 2017 Contents 1 BanzaiDB documentation contents 3 2 Indices and tables 11 i ii BanzaiDB is a tool for pairing Microbial Genomics Next
Getting the files for the first time...2. Making Changes, Commiting them and Pull Requests:...5. Update your repository from the upstream master...
 Table of Contents Getting the files for the first time...2 Making Changes, Commiting them and Pull Requests:...5 Update your repository from the upstream master...8 Making a new branch (for leads, do this
Table of Contents Getting the files for the first time...2 Making Changes, Commiting them and Pull Requests:...5 Update your repository from the upstream master...8 Making a new branch (for leads, do this
Git. Presenter: Haotao (Eric) Lai Contact:
 Git Presenter: Haotao (Eric) Lai Contact: haotao.lai@gmail.com 1 Acknowledge images with white background is from the following link: http://marklodato.github.io/visual-git-guide/index-en.html images with
Git Presenter: Haotao (Eric) Lai Contact: haotao.lai@gmail.com 1 Acknowledge images with white background is from the following link: http://marklodato.github.io/visual-git-guide/index-en.html images with
CPSC 491. Lecture 19 & 20: Source Code Version Control. VCS = Version Control Software SCM = Source Code Management
 CPSC 491 Lecture 19 & 20: Source Code Version Control VCS = Version Control Software SCM = Source Code Management Exercise: Source Code (Version) Control 1. Pretend like you don t have a version control
CPSC 491 Lecture 19 & 20: Source Code Version Control VCS = Version Control Software SCM = Source Code Management Exercise: Source Code (Version) Control 1. Pretend like you don t have a version control
Python Project Example Documentation
 Python Project Example Documentation Release 0.1.0 Neil Stoddard Mar 22, 2017 Contents 1 Neilvana Example 3 1.1 Features.................................................. 3 1.2 Credits..................................................
Python Project Example Documentation Release 0.1.0 Neil Stoddard Mar 22, 2017 Contents 1 Neilvana Example 3 1.1 Features.................................................. 3 1.2 Credits..................................................
git-pr Release dev2+ng5b0396a
 git-pr Release 0.2.1.dev2+ng5b0396a Mar 20, 2017 Contents 1 Table Of Contents 3 1.1 Installation................................................ 3 1.2 Usage...................................................
git-pr Release 0.2.1.dev2+ng5b0396a Mar 20, 2017 Contents 1 Table Of Contents 3 1.1 Installation................................................ 3 1.2 Usage...................................................
Lecture 3: Processing Language Data, Git/GitHub. LING 1340/2340: Data Science for Linguists Na-Rae Han
 Lecture 3: Processing Language Data, Git/GitHub LING 1340/2340: Data Science for Linguists Na-Rae Han Objectives What do linguistic data look like? Homework 1: What did you process? How does collaborating
Lecture 3: Processing Language Data, Git/GitHub LING 1340/2340: Data Science for Linguists Na-Rae Han Objectives What do linguistic data look like? Homework 1: What did you process? How does collaborating
TPS Documentation. Release Thomas Roten
 TPS Documentation Release 0.1.0 Thomas Roten Sep 27, 2017 Contents 1 TPS: TargetProcess in Python! 3 2 Installation 5 3 Contributing 7 3.1 Types of Contributions..........................................
TPS Documentation Release 0.1.0 Thomas Roten Sep 27, 2017 Contents 1 TPS: TargetProcess in Python! 3 2 Installation 5 3 Contributing 7 3.1 Types of Contributions..........................................
flask-dynamo Documentation
 flask-dynamo Documentation Release 0.1.2 Randall Degges January 22, 2018 Contents 1 User s Guide 3 1.1 Quickstart................................................ 3 1.2 Getting Help...............................................
flask-dynamo Documentation Release 0.1.2 Randall Degges January 22, 2018 Contents 1 User s Guide 3 1.1 Quickstart................................................ 3 1.2 Getting Help...............................................
Roman Numeral Converter Documentation
 Roman Numeral Converter Documentation Release 0.1.0 Adrian Cruz October 07, 2014 Contents 1 Roman Numeral Converter 3 1.1 Features.................................................. 3 2 Installation 5
Roman Numeral Converter Documentation Release 0.1.0 Adrian Cruz October 07, 2014 Contents 1 Roman Numeral Converter 3 1.1 Features.................................................. 3 2 Installation 5
Managing Network Configurations with Git and GitLab
 Managing Network Configurations with Git and GitLab Matthew DeNapoli Developer Advocate, DevNet Twitter: @thedenap Season 1, Workshop 3 https://developer.cisco.com/netdevops/live What are we going to talk
Managing Network Configurations with Git and GitLab Matthew DeNapoli Developer Advocate, DevNet Twitter: @thedenap Season 1, Workshop 3 https://developer.cisco.com/netdevops/live What are we going to talk
ACCURATE NONLINEAR MAPPING BETWEEN MNI152/COLIN27 VOLUMETRIC AND FREESURFER SURFACE COORDINATE SYSTEMS
 ACCURATE NONLINEAR MAPPING BETWEEN MNI152/COLIN27 VOLUMETRIC AND FREESURFER SURFACE COORDINATE SYSTEMS WU JIANXIAO (B.Eng. (Hons.), NUS) A THESIS SUBMITTED FOR THE DEGREE OF MASTER OF ENGINEERING DEPARTMENT
ACCURATE NONLINEAR MAPPING BETWEEN MNI152/COLIN27 VOLUMETRIC AND FREESURFER SURFACE COORDINATE SYSTEMS WU JIANXIAO (B.Eng. (Hons.), NUS) A THESIS SUBMITTED FOR THE DEGREE OF MASTER OF ENGINEERING DEPARTMENT
FAQ Q: Where/in which branch do I create new code/modify existing code? A: Q: How do I commit new changes? A:
 FAQ Q: Where/in which branch do I create new code/modify existing code? A: We strongly recommend only modifying the source code within the local master branch: Git Repository View Woped repository Branches
FAQ Q: Where/in which branch do I create new code/modify existing code? A: We strongly recommend only modifying the source code within the local master branch: Git Repository View Woped repository Branches
TangeloHub Documentation
 TangeloHub Documentation Release None Kitware, Inc. September 21, 2015 Contents 1 User s Guide 3 1.1 Managing Data.............................................. 3 1.2 Running an Analysis...........................................
TangeloHub Documentation Release None Kitware, Inc. September 21, 2015 Contents 1 User s Guide 3 1.1 Managing Data.............................................. 3 1.2 Running an Analysis...........................................
chatterbot-weather Documentation
 chatterbot-weather Documentation Release 0.1.1 Gunther Cox Nov 23, 2018 Contents 1 chatterbot-weather 3 1.1 Installation................................................ 3 1.2 Example.................................................
chatterbot-weather Documentation Release 0.1.1 Gunther Cox Nov 23, 2018 Contents 1 chatterbot-weather 3 1.1 Installation................................................ 3 1.2 Example.................................................
nacelle Documentation
 nacelle Documentation Release 0.4.1 Patrick Carey August 16, 2014 Contents 1 Standing on the shoulders of giants 3 2 Contents 5 2.1 Getting Started.............................................. 5 2.2
nacelle Documentation Release 0.4.1 Patrick Carey August 16, 2014 Contents 1 Standing on the shoulders of giants 3 2 Contents 5 2.1 Getting Started.............................................. 5 2.2
A Design Toolbox to Generate Complex Phantoms for the Evaluation of Medical Image Processing Algorithms
 A Design Toolbox to Generate Complex Phantoms for the Evaluation of Medical Image Processing Algorithms Omar Hamo, Georg Nelles, Gudrun Wagenknecht Central Institute for Electronics, Research Center Juelich,
A Design Toolbox to Generate Complex Phantoms for the Evaluation of Medical Image Processing Algorithms Omar Hamo, Georg Nelles, Gudrun Wagenknecht Central Institute for Electronics, Research Center Juelich,
Git for Subversion users
 Git for Subversion users Zend webinar, 23-02-2012 Stefan who? Stefan who? Freelancer: Ingewikkeld Stefan who? Freelancer: Ingewikkeld Symfony Community Manager Stefan who? Freelancer: Ingewikkeld Symfony
Git for Subversion users Zend webinar, 23-02-2012 Stefan who? Stefan who? Freelancer: Ingewikkeld Stefan who? Freelancer: Ingewikkeld Symfony Community Manager Stefan who? Freelancer: Ingewikkeld Symfony
dicompyler-core Documentation
 dicompyler-core Documentation Release 0.5.3 Aditya Panchal Nov 08, 2017 Contents 1 dicompyler-core 3 1.1 Other information............................................ 3 1.2 Dependencies...............................................
dicompyler-core Documentation Release 0.5.3 Aditya Panchal Nov 08, 2017 Contents 1 dicompyler-core 3 1.1 Other information............................................ 3 1.2 Dependencies...............................................
Programming for Data Science Syllabus
 Programming for Data Science Syllabus Learn to use Python and SQL to solve problems with data Before You Start Prerequisites: There are no prerequisites for this program, aside from basic computer skills.
Programming for Data Science Syllabus Learn to use Python and SQL to solve problems with data Before You Start Prerequisites: There are no prerequisites for this program, aside from basic computer skills.
e24paymentpipe Documentation
 e24paymentpipe Documentation Release 1.2.0 Burhan Khalid Oct 30, 2017 Contents 1 e24paymentpipe 3 1.1 Features.................................................. 3 1.2 Todo...................................................
e24paymentpipe Documentation Release 1.2.0 Burhan Khalid Oct 30, 2017 Contents 1 e24paymentpipe 3 1.1 Features.................................................. 3 1.2 Todo...................................................
API RI. Application Programming Interface Reference Implementation. Policies and Procedures Discussion
 API Working Group Meeting, Harris County, TX March 22-23, 2016 Policies and Procedures Discussion Developing a Mission Statement What do we do? How do we do it? Whom do we do it for? What value are we
API Working Group Meeting, Harris County, TX March 22-23, 2016 Policies and Procedures Discussion Developing a Mission Statement What do we do? How do we do it? Whom do we do it for? What value are we
Python wrapper for Viscosity.app Documentation
 Python wrapper for Viscosity.app Documentation Release Paul Kremer March 08, 2014 Contents 1 Python wrapper for Viscosity.app 3 1.1 Features.................................................. 3 2 Installation
Python wrapper for Viscosity.app Documentation Release Paul Kremer March 08, 2014 Contents 1 Python wrapper for Viscosity.app 3 1.1 Features.................................................. 3 2 Installation
Merge Conflicts p. 92 More GitHub Workflows: Forking and Pull Requests p. 97 Using Git to Make Life Easier: Working with Past Commits p.
 Preface p. xiii Ideology: Data Skills for Robust and Reproducible Bioinformatics How to Learn Bioinformatics p. 1 Why Bioinformatics? Biology's Growing Data p. 1 Learning Data Skills to Learn Bioinformatics
Preface p. xiii Ideology: Data Skills for Robust and Reproducible Bioinformatics How to Learn Bioinformatics p. 1 Why Bioinformatics? Biology's Growing Data p. 1 Learning Data Skills to Learn Bioinformatics
Function-Structure Integration in FreeSurfer
 Function-Structure Integration in FreeSurfer Outline Function-Structure Integration Function-Structure Registration in FreeSurfer fmri Analysis Preprocessing First-Level Analysis Higher-Level (Group) Analysis
Function-Structure Integration in FreeSurfer Outline Function-Structure Integration Function-Structure Registration in FreeSurfer fmri Analysis Preprocessing First-Level Analysis Higher-Level (Group) Analysis
Configuration Management
 Configuration Management VIMIMA11 Design and integration of embedded systems Budapest University of Technology and Economics Department of Measurement and Information Systems BME-MIT 2017 Configuration
Configuration Management VIMIMA11 Design and integration of embedded systems Budapest University of Technology and Economics Department of Measurement and Information Systems BME-MIT 2017 Configuration
smsghussd Documentation
 smsghussd Documentation Release 0.1.0 Mawuli Adzaku July 11, 2015 Contents 1 How to use 3 2 Author 7 3 LICENSE 9 3.1 Contents:................................................. 9 3.2 Feedback.................................................
smsghussd Documentation Release 0.1.0 Mawuli Adzaku July 11, 2015 Contents 1 How to use 3 2 Author 7 3 LICENSE 9 3.1 Contents:................................................. 9 3.2 Feedback.................................................
Where are we now? Structural MRI processing and analysis
 Where are we now? Structural MRI processing and analysis Pierre-Louis Bazin bazin@cbs.mpg.de Leipzig, Germany Structural MRI processing: why bother? Just use the standards? SPM FreeSurfer FSL However:
Where are we now? Structural MRI processing and analysis Pierre-Louis Bazin bazin@cbs.mpg.de Leipzig, Germany Structural MRI processing: why bother? Just use the standards? SPM FreeSurfer FSL However:
gunny Documentation Release David Blewett
 gunny Documentation Release 0.1.0 David Blewett December 29, 2013 Contents 1 gunny 3 1.1 Features.................................................. 3 2 Installation 5 2.1 Dependencies...............................................
gunny Documentation Release 0.1.0 David Blewett December 29, 2013 Contents 1 gunny 3 1.1 Features.................................................. 3 2 Installation 5 2.1 Dependencies...............................................
First tutorial session
 First tutorial session Vincent Dumoulin January 15, 2015 Outline 1 Solution to the numpy + MNIST + MLP assignment 2 Git primer 3 Theano primer 4 Porting numpy + MNIST + MLP to theano + MNIST + MLP Solution
First tutorial session Vincent Dumoulin January 15, 2015 Outline 1 Solution to the numpy + MNIST + MLP assignment 2 Git primer 3 Theano primer 4 Porting numpy + MNIST + MLP to theano + MNIST + MLP Solution
About the Tutorial. Audience. Prerequisites. Copyright & Disclaimer. Gerrit
 Gerrit About the Tutorial Gerrit is a web-based code review tool, which is integrated with Git and built on top of Git version control system (helps developers to work together and maintain the history
Gerrit About the Tutorial Gerrit is a web-based code review tool, which is integrated with Git and built on top of Git version control system (helps developers to work together and maintain the history
Version Control for the 2- Pizza Team: Merge Conflicts (ELLS 9.5) Armando Fox
 Version Control for the 2- Pizza Team: Merge Conflicts (ELLS 9.5) Armando Fox 2012 Armando Fox & David Patterson Licensed under Creative Commons Attribution- NonCommercial-ShareAlike 3.0 Unported License
Version Control for the 2- Pizza Team: Merge Conflicts (ELLS 9.5) Armando Fox 2012 Armando Fox & David Patterson Licensed under Creative Commons Attribution- NonCommercial-ShareAlike 3.0 Unported License
Introduction to Git and Github
 Introduction to Git and Github Computing in Optimization and Statistics: Lecture 1 Jackie Baek MIT January 10, 2017 What is git and GitHub? git is a version control system. Other version control systems
Introduction to Git and Github Computing in Optimization and Statistics: Lecture 1 Jackie Baek MIT January 10, 2017 What is git and GitHub? git is a version control system. Other version control systems
Functional MRI data preprocessing. Cyril Pernet, PhD
 Functional MRI data preprocessing Cyril Pernet, PhD Data have been acquired, what s s next? time No matter the design, multiple volumes (made from multiple slices) have been acquired in time. Before getting
Functional MRI data preprocessing Cyril Pernet, PhD Data have been acquired, what s s next? time No matter the design, multiple volumes (made from multiple slices) have been acquired in time. Before getting
Lab 08. Command Line and Git
 Lab 08 Command Line and Git Agenda Final Project Information All Things Git! Make sure to come to lab next week for Python! Final Projects Connect 4 Arduino ios Creative AI Being on a Team - How To Maximize
Lab 08 Command Line and Git Agenda Final Project Information All Things Git! Make sure to come to lab next week for Python! Final Projects Connect 4 Arduino ios Creative AI Being on a Team - How To Maximize
Introduc)on to FreeSurfer h0p://surfer.nmr.mgh.harvard.edu. Jenni Pacheco.
 Introduc)on to FreeSurfer h0p://surfer.nmr.mgh.harvard.edu Jenni Pacheco jpacheco@mail.utexas.edu Outline Processing Stages Command line Stream Assemble Data (mris_preproc, mri_surf2surf) Design/Contrast
Introduc)on to FreeSurfer h0p://surfer.nmr.mgh.harvard.edu Jenni Pacheco jpacheco@mail.utexas.edu Outline Processing Stages Command line Stream Assemble Data (mris_preproc, mri_surf2surf) Design/Contrast
Django MFA Documentation
 Django MFA Documentation Release 1.0 Micro Pyramid Sep 20, 2018 Contents 1 Getting started 3 1.1 Requirements............................................... 3 1.2 Installation................................................
Django MFA Documentation Release 1.0 Micro Pyramid Sep 20, 2018 Contents 1 Getting started 3 1.1 Requirements............................................... 3 1.2 Installation................................................
Simple libtorrent streaming module Documentation
 Simple libtorrent streaming module Documentation Release 0.1.0 David Francos August 31, 2015 Contents 1 Simple libtorrent streaming module 3 1.1 Dependences...............................................
Simple libtorrent streaming module Documentation Release 0.1.0 David Francos August 31, 2015 Contents 1 Simple libtorrent streaming module 3 1.1 Dependences...............................................
withenv Documentation
 withenv Documentation Release 0.7.0 Eric Larson Aug 02, 2017 Contents 1 withenv 3 2 Installation 5 3 Usage 7 3.1 YAML Format.............................................. 7 3.2 Command Substitutions.........................................
withenv Documentation Release 0.7.0 Eric Larson Aug 02, 2017 Contents 1 withenv 3 2 Installation 5 3 Usage 7 3.1 YAML Format.............................................. 7 3.2 Command Substitutions.........................................
contribution-guide.org Release
 contribution-guide.org Release August 06, 2018 Contents 1 About 1 1.1 Sources.................................................. 1 2 Submitting bugs 3 2.1 Due diligence...............................................
contribution-guide.org Release August 06, 2018 Contents 1 About 1 1.1 Sources.................................................. 1 2 Submitting bugs 3 2.1 Due diligence...............................................
Instructions part 1 (rev.) and 2 ENIGMA-MDD hippocampal subfields project
 Instructions part 1 (rev.) and 2 ENIGMA-MDD hippocampal subfields project 12 th of December 2016 Philipp Sämann, Neda Janashad, Laura van Velzen, Derrek Hibar, Theo van Erp, Christopher Whelan, Lianne
Instructions part 1 (rev.) and 2 ENIGMA-MDD hippocampal subfields project 12 th of December 2016 Philipp Sämann, Neda Janashad, Laura van Velzen, Derrek Hibar, Theo van Erp, Christopher Whelan, Lianne
The Allen Human Brain Atlas offers three types of searches to allow a user to: (1) obtain gene expression data for specific genes (or probes) of
 Microarray Data MICROARRAY DATA Gene Search Boolean Syntax Differential Search Mouse Differential Search Search Results Gene Classification Correlative Search Download Search Results Data Visualization
Microarray Data MICROARRAY DATA Gene Search Boolean Syntax Differential Search Mouse Differential Search Search Results Gene Classification Correlative Search Download Search Results Data Visualization
Using GitHub and SourceTree to work with DITA TC repositories
 Using GitHub and SourceTree to work with DITA TC repositories Kristen James Eberlein Eberlein Consulting LLC Agenda 1. Before you begin 2. Getting set up: 1. Fork the DITA TC repository 2. Clone your fork
Using GitHub and SourceTree to work with DITA TC repositories Kristen James Eberlein Eberlein Consulting LLC Agenda 1. Before you begin 2. Getting set up: 1. Fork the DITA TC repository 2. Clone your fork
699DR git/github Tutorial
 699DR git/github Tutorial Sep 20 2017 This tutorial gives a high-level introduction into basic usage of the version control software git in combination with the online platform Github. The git commands
699DR git/github Tutorial Sep 20 2017 This tutorial gives a high-level introduction into basic usage of the version control software git in combination with the online platform Github. The git commands
Python simple arp table reader Documentation
 Python simple arp table reader Documentation Release 0.0.1 David Francos Nov 17, 2017 Contents 1 Python simple arp table reader 3 1.1 Features.................................................. 3 1.2 Usage...................................................
Python simple arp table reader Documentation Release 0.0.1 David Francos Nov 17, 2017 Contents 1 Python simple arp table reader 3 1.1 Features.................................................. 3 1.2 Usage...................................................
Version Control Systems
 Nothing to see here. Everything is under control! September 16, 2015 Change tracking File moving Teamwork Undo! Undo! UNDO!!! What strategies do you use for tracking changes to files? Change tracking File
Nothing to see here. Everything is under control! September 16, 2015 Change tracking File moving Teamwork Undo! Undo! UNDO!!! What strategies do you use for tracking changes to files? Change tracking File
ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION
 ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION Abstract: MIP Project Report Spring 2013 Gaurav Mittal 201232644 This is a detailed report about the course project, which was to implement
ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION Abstract: MIP Project Report Spring 2013 Gaurav Mittal 201232644 This is a detailed report about the course project, which was to implement
FPLLL. Contributing. Martin R. Albrecht 2017/07/06
 FPLLL Contributing Martin R. Albrecht 2017/07/06 Outline Communication Setup Reporting Bugs Topic Branches and Pull Requests How to Get your Pull Request Accepted Documentation Overview All contributions
FPLLL Contributing Martin R. Albrecht 2017/07/06 Outline Communication Setup Reporting Bugs Topic Branches and Pull Requests How to Get your Pull Request Accepted Documentation Overview All contributions
Version control CSE 403
 Version control CSE 403 Goals of a version control system Keep a history of your work Explain the purpose of each change Checkpoint specific versions (known good state) Recover specific state (fix bugs,
Version control CSE 403 Goals of a version control system Keep a history of your work Explain the purpose of each change Checkpoint specific versions (known good state) Recover specific state (fix bugs,
sainsmart Documentation
 sainsmart Documentation Release 0.3.1 Victor Yap Jun 21, 2017 Contents 1 sainsmart 3 1.1 Install................................................... 3 1.2 Usage...................................................
sainsmart Documentation Release 0.3.1 Victor Yap Jun 21, 2017 Contents 1 sainsmart 3 1.1 Install................................................... 3 1.2 Usage...................................................
Review Version Control Concepts
 Review Version Control Concepts SWEN-261 Introduction to Software Engineering Department of Software Engineering Rochester Institute of Technology Managing change is a constant aspect of software development.
Review Version Control Concepts SWEN-261 Introduction to Software Engineering Department of Software Engineering Rochester Institute of Technology Managing change is a constant aspect of software development.
Poulpe Documentation. Release Edouard Klein
 Poulpe Documentation Release 0.0.5 Edouard Klein Jul 18, 2017 Contents 1 Poulpe 1 1.1 Features.................................................. 1 2 Usage 3 3 Installation 5 4 Contributing 7 4.1 Types
Poulpe Documentation Release 0.0.5 Edouard Klein Jul 18, 2017 Contents 1 Poulpe 1 1.1 Features.................................................. 1 2 Usage 3 3 Installation 5 4 Contributing 7 4.1 Types
Python Finite State Machine. Release 0.1.5
 Python Finite State Machine Release 0.1.5 Sep 15, 2017 Contents 1 Overview 1 1.1 Installation................................................ 1 1.2 Documentation..............................................
Python Finite State Machine Release 0.1.5 Sep 15, 2017 Contents 1 Overview 1 1.1 Installation................................................ 1 1.2 Documentation..............................................
TractoR and Other Software
 TractoR and Other Software Jon Clayden DIBS Teaching Seminar, 11 Dec 2015 Photo by José Martín Ramírez Carrasco https://www.behance.net/martini_rc TractoR A set of R packages Additional
TractoR and Other Software Jon Clayden DIBS Teaching Seminar, 11 Dec 2015 Photo by José Martín Ramírez Carrasco https://www.behance.net/martini_rc TractoR A set of R packages Additional
Release Nicholas A. Del Grosso
 wavefront r eaderdocumentation Release 0.1.0 Nicholas A. Del Grosso Apr 12, 2017 Contents 1 wavefront_reader 3 1.1 Features.................................................. 3 1.2 Credits..................................................
wavefront r eaderdocumentation Release 0.1.0 Nicholas A. Del Grosso Apr 12, 2017 Contents 1 wavefront_reader 3 1.1 Features.................................................. 3 1.2 Credits..................................................
The Old World. Have you ever had to collaborate on a project by
 What the Git? The Old World Have you ever had to collaborate on a project by Shuttling a USB drive back and forth Using Dropbox E-mailing your document around Have you ever accidentally deleted someone
What the Git? The Old World Have you ever had to collaborate on a project by Shuttling a USB drive back and forth Using Dropbox E-mailing your document around Have you ever accidentally deleted someone
timegate Documentation
 timegate Documentation Release 0.5.0.dev20160000 LANL Jul 16, 2018 Contents 1 About 3 2 User s Guide 5 2.1 Introduction............................................... 5 2.2 Installation................................................
timegate Documentation Release 0.5.0.dev20160000 LANL Jul 16, 2018 Contents 1 About 3 2 User s Guide 5 2.1 Introduction............................................... 5 2.2 Installation................................................
Software Development I
 6.148 Software Development I Two things How to write code for web apps. How to collaborate and keep track of your work. A text editor A text editor A text editor Anything that you re used to using Even
6.148 Software Development I Two things How to write code for web apps. How to collaborate and keep track of your work. A text editor A text editor A text editor Anything that you re used to using Even
CS 520: VCS and Git. Intermediate Topics Ben Kushigian
 CS 520: VCS and Git Intermediate Topics Ben Kushigian https://people.cs.umass.edu/~rjust/courses/2017fall/cs520/2017_09_19.zip Our Goal Our Goal (Overture) Overview the basics of Git w/ an eye towards
CS 520: VCS and Git Intermediate Topics Ben Kushigian https://people.cs.umass.edu/~rjust/courses/2017fall/cs520/2017_09_19.zip Our Goal Our Goal (Overture) Overview the basics of Git w/ an eye towards
Distributed Version Control
 Distributed Version Control David Grellscheid 2014-03-17 Workshop on Advanced Techniques for Scientific Programming and Management of Open Source Software Packages 10 21 March 2014 Version management In
Distributed Version Control David Grellscheid 2014-03-17 Workshop on Advanced Techniques for Scientific Programming and Management of Open Source Software Packages 10 21 March 2014 Version management In
Multiple Cortical Surface Correspondence Using Pairwise Shape Similarity
 Multiple Cortical Surface Correspondence Using Pairwise Shape Similarity Pahal Dalal 1, Feng Shi 2, Dinggang Shen 2, and Song Wang 1 1 Department of Computer Science and Engineering, University of South
Multiple Cortical Surface Correspondence Using Pairwise Shape Similarity Pahal Dalal 1, Feng Shi 2, Dinggang Shen 2, and Song Wang 1 1 Department of Computer Science and Engineering, University of South
Tips on how to set up a GitHub account:
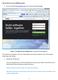 Tips on how to set up a GitHub account: 1. Go to the website https://github.com/, you will see the following page: Figure 1: The GitHub main webpage (before you create an account and sign in) Then choose
Tips on how to set up a GitHub account: 1. Go to the website https://github.com/, you will see the following page: Figure 1: The GitHub main webpage (before you create an account and sign in) Then choose
Slicer3 Training Compendium. Slicer3 Training Tutorial ARCTIC (v1.2)
 Slicer3 Training Compendium Slicer3 Training Tutorial ARCTIC (v1.2) ( ThICkness (Automatic Regional Cortical University of North Carolina, Chapel Hill: Neuro Image Research and Analysis Lab Neurodevelopmental
Slicer3 Training Compendium Slicer3 Training Tutorial ARCTIC (v1.2) ( ThICkness (Automatic Regional Cortical University of North Carolina, Chapel Hill: Neuro Image Research and Analysis Lab Neurodevelopmental
Job Submitter Documentation
 Job Submitter Documentation Release 0+untagged.133.g5a1e521.dirty Juan Eiros February 27, 2017 Contents 1 Job Submitter 3 1.1 Before you start............................................. 3 1.2 Features..................................................
Job Submitter Documentation Release 0+untagged.133.g5a1e521.dirty Juan Eiros February 27, 2017 Contents 1 Job Submitter 3 1.1 Before you start............................................. 3 1.2 Features..................................................
Caret 5 User s Guide to Analysis Procedures John Harwell, Heather A. Drury, Donna Dierker, and David C. Van Essen
 Caret 5 User s Guide to Analysis Procedures John Harwell, Heather A. Drury, Donna Dierker, and David C. Van Essen Caret Version 5.5 22 September, 2006 Copyright 1995-2006 Washington University Washington
Caret 5 User s Guide to Analysis Procedures John Harwell, Heather A. Drury, Donna Dierker, and David C. Van Essen Caret Version 5.5 22 September, 2006 Copyright 1995-2006 Washington University Washington
G E T T I N G S TA R T E D W I T H G I T
 G E T T I N G S TA R T E D W I T H G I T A A R O N H O O V E R & B R A D M I N C H J A N U A R Y 2 2, 2 0 1 8 1 Why use a version control system? Much of this document was blatantly cribbed from Allen
G E T T I N G S TA R T E D W I T H G I T A A R O N H O O V E R & B R A D M I N C H J A N U A R Y 2 2, 2 0 1 8 1 Why use a version control system? Much of this document was blatantly cribbed from Allen
SFO17-315: OpenDataPlane Testing in Travis. Dmitry Eremin-Solenikov, Cavium Maxim Uvarov, Linaro
 SFO17-315: OpenDataPlane Testing in Travis Dmitry Eremin-Solenikov, Cavium Maxim Uvarov, Linaro What is ODP (OpenDataPlane) The ODP project is an open-source, cross-platform set of APIs for the networking
SFO17-315: OpenDataPlane Testing in Travis Dmitry Eremin-Solenikov, Cavium Maxim Uvarov, Linaro What is ODP (OpenDataPlane) The ODP project is an open-source, cross-platform set of APIs for the networking
Python data pipelines similar to R Documentation
 Python data pipelines similar to R Documentation Release 0.1.0 Jan Schulz October 23, 2016 Contents 1 Python data pipelines 3 1.1 Features.................................................. 3 1.2 Documentation..............................................
Python data pipelines similar to R Documentation Release 0.1.0 Jan Schulz October 23, 2016 Contents 1 Python data pipelines 3 1.1 Features.................................................. 3 1.2 Documentation..............................................
BrainSuite. presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014
 BrainSuite presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014 David Shattuck Ahmanson-Lovelace Brain Mapping Center Department of Neurology David Geffen School of Medicine at
BrainSuite presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014 David Shattuck Ahmanson-Lovelace Brain Mapping Center Department of Neurology David Geffen School of Medicine at
CSC 2700: Scientific Computing
 CSC 2700: Scientific Computing Record and share your work: revision control systems Dr Frank Löffler Center for Computation and Technology Louisiana State University, Baton Rouge, LA Feb 13 2014 Overview
CSC 2700: Scientific Computing Record and share your work: revision control systems Dr Frank Löffler Center for Computation and Technology Louisiana State University, Baton Rouge, LA Feb 13 2014 Overview
Stereoscopic Atlas of Intrinsic Brain Networks (SAIBN)
 Stereoscopic Atlas of Intrinsic Brain Networks (SAIBN) Gonzalo M. Rojas, Marcelo Gálvez, Daniel Margulies, Cameron Craddock, Xavier Castellanos, Michael Milham 1.- Introduction Stereoscopic Atlas of Intrinsic
Stereoscopic Atlas of Intrinsic Brain Networks (SAIBN) Gonzalo M. Rojas, Marcelo Gálvez, Daniel Margulies, Cameron Craddock, Xavier Castellanos, Michael Milham 1.- Introduction Stereoscopic Atlas of Intrinsic
腦部結構影像 標準化 組織分割 體素型態 本週課程內容. Analysis Softwares. A Course of MRI
 本週課程內容 腦部結構影像 A Course of MRI 盧家鋒助理教授國立陽明大學物理治療暨輔助科技學系 alvin4016@ym.edu.tw 腦部結構影像 空間標準化 (Spatial normalization) 均勻度校正 (Bias correction) 組織分割 (Segmentation) 體素形態學分析 (Voxel-based morphometry, VBM) 影像平滑化
本週課程內容 腦部結構影像 A Course of MRI 盧家鋒助理教授國立陽明大學物理治療暨輔助科技學系 alvin4016@ym.edu.tw 腦部結構影像 空間標準化 (Spatial normalization) 均勻度校正 (Bias correction) 組織分割 (Segmentation) 體素形態學分析 (Voxel-based morphometry, VBM) 影像平滑化
google-search Documentation
 google-search Documentation Release 1.0.0 Anthony Hseb May 08, 2017 Contents 1 google-search 3 1.1 Features.................................................. 3 1.2 Credits..................................................
google-search Documentation Release 1.0.0 Anthony Hseb May 08, 2017 Contents 1 google-search 3 1.1 Features.................................................. 3 1.2 Credits..................................................
abstar Documentation Release Bryan Briney
 abstar Documentation Release 0.3.1 Bryan Briney Apr 26, 2018 Contents 1 Getting Started 3 2 Usage 7 3 About 13 4 Related Projects 15 5 Index 17 i ii AbStar is a core component of the Ab[x] Toolkit for
abstar Documentation Release 0.3.1 Bryan Briney Apr 26, 2018 Contents 1 Getting Started 3 2 Usage 7 3 About 13 4 Related Projects 15 5 Index 17 i ii AbStar is a core component of the Ab[x] Toolkit for
