Exercises: Motif Searching
|
|
|
- Jeffrey Newton
- 5 years ago
- Views:
Transcription
1 Exercises: Motif Searching Version
2 Exercises: Motif Searching 2 Licence This manual is , Simon Andrews. This manual is distributed under the creative commons Attribution-Non-Commercial-Share Alike 2.0 licence. This means that you are free: to copy, distribute, display, and perform the work to make derivative works Under the following conditions: Attribution. You must give the original author credit. Non-Commercial. You may not use this work for commercial purposes. Share Alike. If you alter, transform, or build upon this work, you may distribute the resulting work only under a licence identical to this one. Please note that: For any reuse or distribution, you must make clear to others the licence terms of this work. Any of these conditions can be waived if you get permission from the copyright holder. Nothing in this license impairs or restricts the author's moral rights. Full details of this licence can be found at
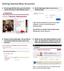
3 Exercises: Motif Searching 3 Promoter Analysis In this exercise you are given a list of genes coming out of a previous experiment and you are going to analyse their promoter sequences to see if they contain any enrichment motifs. You will also analyse them for occurrences of known motifs. Exercise 1a: Retrieving sequences You should find a file called promoter_gene_list.txt in your data folder, this contains a list of genes of interest. To retrieve the promoter sequences for these genes we re going to use the Ensembl biomart system. Open a web browser and navigate to ensembl.org/biomart/ Choose the Ensembl Genes database to query and the Mouse genes dataset. Click on the Filters on the left of the screen, and then expand the GENE section on the right Copy and paste your gene list into the Input external references ID list box and tick the checkbox next to it. Change the ID type in the dropdown to Gene Name(s). You can press the Count button at the top to check your genes are being found. You should see around 100 genes in your dataset Click on the Attributes link on the left and change the type of output from Features to Sequences Expand the SEQUENCES section and select Flank (Gene) as the sequence type. In the flank options tick the Upstream flank box and set the value to 500 Expand the Header Information section Under Gene Information deselect Ensembl Gene ID and select Gene Name Under Transcript Information deselect Transcript stable ID Click on the Results button at the top of the page. You should see something similar to the text below. If yours looks different check your attributes preferences carefully. >Vrk3 GGCATGGCTCCGTGTCTCCGTACCTCAGCATGGCTGGCAACCCAAAGAACATATGCTTTC CAGGATCCCAAGAAAACCCAACTCCTCATGAGGAGGGCTCCCTCAGAGTACAGGGGGAAA TTCCTCCGAGGAAGTTCCGGTTCCAGAAACTTCTCCTCAAATTTATCCCACCCAGGTAAC TCCGCGCCAAATTCCAGGACCCACCACCTGTGGTCTCCACCCACTGAGCCCTCTAGGAAG TAGAGAGGAAACAGATTCATGGCTAAGTGACACCAAGGTTGGATGGCCCACTACCCTGTT ATAGGACTACTTTCACCTTCTACACCCCGAAAATTCCATTTGGAGGTGACAGCGGGTTGC CATAGATACAGTTTGAAGGCAAAACTGAGATTACAGAGAGAGCACAGAAGCAAGGAGAGT TTAAAAATGACAACAGGACCTTTGATTGGTGGCTGTCACGTATTTCATCAAGATTGACGT CAGACATGCGCAGTAGAAGG >Tat TTTTTCCTTGGAGTGCGGTTGAATTTTTGTGGAGATTCCCATTGTCCATAGCAAATCCCA CATGCAGTAGATGAATAGTCAACCAAGCTCTTATCATTGCCCATGTCCAGGAGTGAGACT GGAGTGCACAGAGAAAGCACTACCAGAAGGACCCTCATTTTAATGCATTTCTGTACTGTT AACACAATAGTTTTCATAACAAAATAAGGACAATAAAGTTAATCGCTGTCACCCCAACTG CCCTCCACAGTTTCCTGAAACAGCTCACCAAATGACCAGAAAGAGTTAACACAGAGAATT CCAGATAACGTGATGTGTGGACAAATCCCAGATTGGATGGTGGCCCAGGGTTGGGGTGAG AAACAGCAGAGTGTGGGGGGGGGGAGTGGGGGGAGGTCCAGGGGAGGGGCTTAGTCCTCA CTCTCAACCAATAGCATAAGGGCTTCAGGCCCAACGCCCATTGGCTGAAACTATTTCAAG GGTCAGGACCGCACCTGAAC
4 Exercises: Motif Searching 4 Export your file by using the option at the top of the Results page to Export all results to File FASTA (press the Go button to start the export). Save the results to a file called promoter_sequences.txt. Exercise 1b: Repeatmasking Since repetitive sequences could cause false positive discoveries within our data we are going to use the repeatmasker service to remove them from our promoter data. Navigate to Click on the Repeat Masking link in the services section on the left hand side. Select the file you downloaded in Exercise 1a Change the sensitivity to quick Change the DNA source to Mouse Press Submit sequence to start the search Once the search has completed scroll to the bottom of the results page and find the link to the masked file. View the masked file in a browser, then save it to a file called promoter_sequences_masked.txt Exercise 2: Novel motif discovery We can now use the repeat-masked sequences from exercise 1 to search for novel motifs. Navigate to meme-suite.org and select MEME from the Motif Discovery section on the left Upload your masked sequence file Set the number of motifs to be found to 5 Run the search View the HTML output Note that due to the time it takes for MEME to process this type of search you should not wait for it to complete but continue with exercise 3. If it still hasn t finished when you ve completed that then you can look at the example result we ve provided you with instead of waiting for it to complete. Exercise 3: Enrichment of known motifs Rather than searching for a novel motif we can use the same repeat-masked sequences and search these against the known motif database to look for enrichment of known transcription factor binding sites. Navigate to meme-suite.org and select AME from the Motif Enrichment section on the left Upload your masked sequence file Set the motif database to be JASPAR (NON-REDUNDANT) DNA Run the search Compare the results with what you saw in the MEME search results. Could you have got the same answer from AME without having to run MEME?
5 Exercises: Motif Searching 5 Exercise 4: Differential Motif Searching For this exercise you will do a comparison of two sets of CpG island sequences. We have already prepared these for you and there are two files called positive_cgi_set.txt and negative_cgi_set.txt. To start with we want to check that there is no gross compositional difference between the two sets of sequences since this would override any differences in specific motifs. To do this we will use the Compter kmer analysis tool to visualise their composition Navigate to Select the two CGI sequence files as the first and second search sequence sets. Set the background to be pre-calculated mouse Press Run compter to start the analysis. Look at the results and see if you can see any obvious differences in composition between the two sequence sets. We will then use DREME to compare these and compare the top hit to known motifs to see if we can find a match. Navigate to meme-suite.org and select DREME from the Motif Discovery section on the left Change the control sequence option to User provided sequences Select your positive and negative sequence sets and run the search Do you see a convincing hit returned? How good is the detected motif at distinguishing the two groups? From the top hit press the right pointing arrow under the Submit column and submit the motif to TomTom. Select the JASPAR (NON-REDUNDANT) DNA database for the target motifs. Look at the TomTom results and see if the motif identified looks like any existing known motifs.
Finding and Exporting Data. BioMart
 September 2017 Finding and Exporting Data Not sure what tool to use to find and export data? BioMart is used to retrieve data for complex queries, involving a few or many genes or even complete genomes.
September 2017 Finding and Exporting Data Not sure what tool to use to find and export data? BioMart is used to retrieve data for complex queries, involving a few or many genes or even complete genomes.
Genome Environment Browser (GEB) user guide
 Genome Environment Browser (GEB) user guide GEB is a Java application developed to provide a dynamic graphical interface to visualise the distribution of genome features and chromosome-wide experimental
Genome Environment Browser (GEB) user guide GEB is a Java application developed to provide a dynamic graphical interface to visualise the distribution of genome features and chromosome-wide experimental
Introduction to Genome Browsers
 Introduction to Genome Browsers Rolando Garcia-Milian, MLS, AHIP (Rolando.milian@ufl.edu) Department of Biomedical and Health Information Services Health Sciences Center Libraries, University of Florida
Introduction to Genome Browsers Rolando Garcia-Milian, MLS, AHIP (Rolando.milian@ufl.edu) Department of Biomedical and Health Information Services Health Sciences Center Libraries, University of Florida
Once file and folders are added to your Module Content area you will need to link to them using the Item tool.
 VITAL how to guides elearning Unit Last updated: 01.10.2010 Course Files tool Overview Course Files tool enables you to: Quickly copy large numbers of files into a VITAL module. Files can be dragged and
VITAL how to guides elearning Unit Last updated: 01.10.2010 Course Files tool Overview Course Files tool enables you to: Quickly copy large numbers of files into a VITAL module. Files can be dragged and
MetaStorm: User Manual
 MetaStorm: User Manual User Account: First, either log in as a guest or login to your user account. If you login as a guest, you can visualize public MetaStorm projects, but can not run any analysis. To
MetaStorm: User Manual User Account: First, either log in as a guest or login to your user account. If you login as a guest, you can visualize public MetaStorm projects, but can not run any analysis. To
How To: Run the ENCODE histone ChIP- seq analysis pipeline on DNAnexus
 How To: Run the ENCODE histone ChIP- seq analysis pipeline on DNAnexus Overview: In this exercise, we will run the ENCODE Uniform Processing ChIP- seq Pipeline on a small test dataset containing reads
How To: Run the ENCODE histone ChIP- seq analysis pipeline on DNAnexus Overview: In this exercise, we will run the ENCODE Uniform Processing ChIP- seq Pipeline on a small test dataset containing reads
BioMart: a research data management tool for the biomedical sciences
 Yale University From the SelectedWorks of Rolando Garcia-Milian 2014 BioMart: a research data management tool for the biomedical sciences Rolando Garcia-Milian, Yale University Available at: https://works.bepress.com/rolando_garciamilian/2/
Yale University From the SelectedWorks of Rolando Garcia-Milian 2014 BioMart: a research data management tool for the biomedical sciences Rolando Garcia-Milian, Yale University Available at: https://works.bepress.com/rolando_garciamilian/2/
BovineMine Documentation
 BovineMine Documentation Release 1.0 Deepak Unni, Aditi Tayal, Colin Diesh, Christine Elsik, Darren Hag Oct 06, 2017 Contents 1 Tutorial 3 1.1 Overview.................................................
BovineMine Documentation Release 1.0 Deepak Unni, Aditi Tayal, Colin Diesh, Christine Elsik, Darren Hag Oct 06, 2017 Contents 1 Tutorial 3 1.1 Overview.................................................
Talend Data Preparation Free Desktop. Getting Started Guide V2.1
 Talend Data Free Desktop Getting Guide V2.1 1 Talend Data Training Getting Guide To navigate to a specific location within this guide, click one of the boxes below. Overview of Data Access Data And Getting
Talend Data Free Desktop Getting Guide V2.1 1 Talend Data Training Getting Guide To navigate to a specific location within this guide, click one of the boxes below. Overview of Data Access Data And Getting
m6aviewer Version Documentation
 m6aviewer Version 1.6.0 Documentation Contents 1. About 2. Requirements 3. Launching m6aviewer 4. Running Time Estimates 5. Basic Peak Calling 6. Running Modes 7. Multiple Samples/Sample Replicates 8.
m6aviewer Version 1.6.0 Documentation Contents 1. About 2. Requirements 3. Launching m6aviewer 4. Running Time Estimates 5. Basic Peak Calling 6. Running Modes 7. Multiple Samples/Sample Replicates 8.
Tutorial 1: Using Excel to find unique values in a list
 Tutorial 1: Using Excel to find unique values in a list It is not uncommon to have a list of data that contains redundant values. Genes with multiple transcript isoforms is one example. If you are only
Tutorial 1: Using Excel to find unique values in a list It is not uncommon to have a list of data that contains redundant values. Genes with multiple transcript isoforms is one example. If you are only
Tutorial 1: Exploring the UCSC Genome Browser
 Last updated: May 12, 2011 Tutorial 1: Exploring the UCSC Genome Browser Open the homepage of the UCSC Genome Browser at: http://genome.ucsc.edu/ In the blue bar at the top, click on the Genomes link.
Last updated: May 12, 2011 Tutorial 1: Exploring the UCSC Genome Browser Open the homepage of the UCSC Genome Browser at: http://genome.ucsc.edu/ In the blue bar at the top, click on the Genomes link.
Exercises: Analysing RNA-Seq data
 Exercises: Analysing RNA-Seq data Version 2018-03 Exercises: Analysing RNA-Seq data 2 Licence This manual is 2011-18, Simon Andrews, Laura Biggins. This manual is distributed under the creative commons
Exercises: Analysing RNA-Seq data Version 2018-03 Exercises: Analysing RNA-Seq data 2 Licence This manual is 2011-18, Simon Andrews, Laura Biggins. This manual is distributed under the creative commons
Content Collection. How to Access Content Collection. From the homepage: From a course:
 Content Collection What is Content Management? Blackboard s Content Collection is a file repository which allows faculty to store, manage, and share content within personal user folders, course folders,
Content Collection What is Content Management? Blackboard s Content Collection is a file repository which allows faculty to store, manage, and share content within personal user folders, course folders,
ChIP-seq Analysis Practical
 ChIP-seq Analysis Practical Vladimir Teif (vteif@essex.ac.uk) An updated version of this document will be available at http://generegulation.info/index.php/teaching In this practical we will learn how
ChIP-seq Analysis Practical Vladimir Teif (vteif@essex.ac.uk) An updated version of this document will be available at http://generegulation.info/index.php/teaching In this practical we will learn how
ChIP-seq practical: peak detection and peak annotation. Mali Salmon-Divon Remco Loos Myrto Kostadima
 ChIP-seq practical: peak detection and peak annotation Mali Salmon-Divon Remco Loos Myrto Kostadima March 2012 Introduction The goal of this hands-on session is to perform some basic tasks in the analysis
ChIP-seq practical: peak detection and peak annotation Mali Salmon-Divon Remco Loos Myrto Kostadima March 2012 Introduction The goal of this hands-on session is to perform some basic tasks in the analysis
Documentation Tool Tutorial Tutorial for Creating a Documentation Tool Interactive in the Texas Gateway
 Tutorial for Creating a Documentation Tool Interactive in the Texas Gateway Introduction The Documentation Tool interactive serves as a wizard that can help learners easily document and evaluate goal driven
Tutorial for Creating a Documentation Tool Interactive in the Texas Gateway Introduction The Documentation Tool interactive serves as a wizard that can help learners easily document and evaluate goal driven
Importing from Blackboard Learn Grade Center Data to Banner 9 User Learning Scenarios
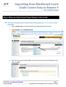 Importing from Blackboard Learn Grade Center Data to Banner 9 User Learning Scenarios Step 1: Make sure Final Grade Column Displays Letter Grade Ensure your final grade column in Grade Center has letter
Importing from Blackboard Learn Grade Center Data to Banner 9 User Learning Scenarios Step 1: Make sure Final Grade Column Displays Letter Grade Ensure your final grade column in Grade Center has letter
National Monuments Service: Wreck Database Viewer
 National Monuments Service: Wreck Database Viewer Introduction HELP DOCUMENT Welcome to the Wreck Viewer map / search facility that provides access to the records of wrecks held by the National Monuments
National Monuments Service: Wreck Database Viewer Introduction HELP DOCUMENT Welcome to the Wreck Viewer map / search facility that provides access to the records of wrecks held by the National Monuments
Data Visualisation with Google Fusion Tables
 Data Visualisation with Google Fusion Tables Workshop Exercises Dr Luc Small 12 April 2017 1.5 Data Visualisation with Google Fusion Tables Page 1 of 33 1 Introduction Google Fusion Tables is shaping up
Data Visualisation with Google Fusion Tables Workshop Exercises Dr Luc Small 12 April 2017 1.5 Data Visualisation with Google Fusion Tables Page 1 of 33 1 Introduction Google Fusion Tables is shaping up
Mouse Atlas DiscoverySpace Tutorial
 Mouse Atlas DiscoverySpace Tutorial Example 1: Foregut vs. Hindgut 1. Find all the tags that are exclusive to the Foregut (SM107) and not in Hindgut (SM112).. pg 2 2. Identify Tags that are up regulated
Mouse Atlas DiscoverySpace Tutorial Example 1: Foregut vs. Hindgut 1. Find all the tags that are exclusive to the Foregut (SM107) and not in Hindgut (SM112).. pg 2 2. Identify Tags that are up regulated
Browser Exercises - I. Alignments and Comparative genomics
 Browser Exercises - I Alignments and Comparative genomics 1. Navigating to the Genome Browser (GBrowse) Note: For this exercise use http://www.tritrypdb.org a. Navigate to the Genome Browser (GBrowse)
Browser Exercises - I Alignments and Comparative genomics 1. Navigating to the Genome Browser (GBrowse) Note: For this exercise use http://www.tritrypdb.org a. Navigate to the Genome Browser (GBrowse)
Exercises. Biological Data Analysis Using InterMine workshop exercises with answers
 Exercises Biological Data Analysis Using InterMine workshop exercises with answers Exercise1: Faceted Search Use HumanMine for this exercise 1. Search for one or more of the following using the keyword
Exercises Biological Data Analysis Using InterMine workshop exercises with answers Exercise1: Faceted Search Use HumanMine for this exercise 1. Search for one or more of the following using the keyword
General Help & Instructions to use with Examples
 General Help & Instructions to use with Examples Contents Types of Searches and their Purposes... 2 Basic Search:... 2 Advance search option... 6 List Search:... 7 Details Page... 8 Results Grid functionalities:...
General Help & Instructions to use with Examples Contents Types of Searches and their Purposes... 2 Basic Search:... 2 Advance search option... 6 List Search:... 7 Details Page... 8 Results Grid functionalities:...
ChIP-Seq Tutorial on Galaxy
 1 Introduction ChIP-Seq Tutorial on Galaxy 2 December 2010 (modified April 6, 2017) Rory Stark The aim of this practical is to give you some experience handling ChIP-Seq data. We will be working with data
1 Introduction ChIP-Seq Tutorial on Galaxy 2 December 2010 (modified April 6, 2017) Rory Stark The aim of this practical is to give you some experience handling ChIP-Seq data. We will be working with data
Membership Portal Manual
 Membership Portal Manual Table of Contents Login... 4 Contact Tab... 6 Contact Information Dropdown...6 Features on the Contact Information Dropdown... 6 Account Information Dropdown...6 Features on the
Membership Portal Manual Table of Contents Login... 4 Contact Tab... 6 Contact Information Dropdown...6 Features on the Contact Information Dropdown... 6 Account Information Dropdown...6 Features on the
Creating and Using Genome Assemblies Tutorial
 Creating and Using Genome Assemblies Tutorial Release 8.1 Golden Helix, Inc. March 18, 2014 Contents 1. Create a Genome Assembly for Danio rerio 2 2. Building Annotation Sources 5 A. Creating a Reference
Creating and Using Genome Assemblies Tutorial Release 8.1 Golden Helix, Inc. March 18, 2014 Contents 1. Create a Genome Assembly for Danio rerio 2 2. Building Annotation Sources 5 A. Creating a Reference
CU Careers: Step-by-Step Guide
 CU Careers: Step-by-Step Guide Search Committee Experience This guide describes actions that Search Committee Members can perform in the evaluation process of candidates. It also describes how department
CU Careers: Step-by-Step Guide Search Committee Experience This guide describes actions that Search Committee Members can perform in the evaluation process of candidates. It also describes how department
Document Search. Chapter 4. 09/17 Chapter 4 - Page 1. Copyright University of Pittsburgh. All rights reserved.
 Chapter 4 09/17 Chapter 4 - Page 1 Section Objectives At the end of this section, you should be able to: Perform a simple and an advanced search Search for and view requisition, purchase order and invoice
Chapter 4 09/17 Chapter 4 - Page 1 Section Objectives At the end of this section, you should be able to: Perform a simple and an advanced search Search for and view requisition, purchase order and invoice
Mascot Insight is a new application designed to help you to organise and manage your Mascot search and quantitation results. Mascot Insight provides
 1 Mascot Insight is a new application designed to help you to organise and manage your Mascot search and quantitation results. Mascot Insight provides ways to flexibly merge your Mascot search and quantitation
1 Mascot Insight is a new application designed to help you to organise and manage your Mascot search and quantitation results. Mascot Insight provides ways to flexibly merge your Mascot search and quantitation
Student Records Training Level IIA
 ` Student Records Training Level IIA Assigning Overloads... 2 Assigning Student Specific Permissions... 4 Assigning Class Permission Numbers... 6 Changing Classes to Pass/Fail... 8 Adding a Class... 12
` Student Records Training Level IIA Assigning Overloads... 2 Assigning Student Specific Permissions... 4 Assigning Class Permission Numbers... 6 Changing Classes to Pass/Fail... 8 Adding a Class... 12
Job Aid. Remote Access BAIRS Printing and Saving a Report. Table of Contents
 Remote Access BAIRS Printing and Saving a Report Table of Contents Remote Access BAIRS Printing a Report PDF HTML... 2 Remote Access BAIRS Printing a Report Export to PDF Interactive Reporting... 3 Remote
Remote Access BAIRS Printing and Saving a Report Table of Contents Remote Access BAIRS Printing a Report PDF HTML... 2 Remote Access BAIRS Printing a Report Export to PDF Interactive Reporting... 3 Remote
Lab 5: Dreamweaver CS5, Uploading your Web site
 Lab 5: Dreamweaver CS5, Uploading your Web site Setting up Local and Remote Information: 1. Launch Dreamweaver 2. Choose site->new site 3. By Site Name give your site a name. Make sure the name has no
Lab 5: Dreamweaver CS5, Uploading your Web site Setting up Local and Remote Information: 1. Launch Dreamweaver 2. Choose site->new site 3. By Site Name give your site a name. Make sure the name has no
Click on OneDrive on the menu bar at the top to display your Documents home page.
 Getting started with OneDrive Information Services Getting started with OneDrive What is OneDrive @ University of Edinburgh? OneDrive @ University of Edinburgh is a cloud storage area you can use to share
Getting started with OneDrive Information Services Getting started with OneDrive What is OneDrive @ University of Edinburgh? OneDrive @ University of Edinburgh is a cloud storage area you can use to share
Analysing High Throughput Sequencing Data with SeqMonk
 Analysing High Throughput Sequencing Data with SeqMonk Version 2017-01 Analysing High Throughput Sequencing Data with SeqMonk 2 Licence This manual is 2008-17, Simon Andrews. This manual is distributed
Analysing High Throughput Sequencing Data with SeqMonk Version 2017-01 Analysing High Throughput Sequencing Data with SeqMonk 2 Licence This manual is 2008-17, Simon Andrews. This manual is distributed
ChIP-seq hands-on practical using Galaxy
 ChIP-seq hands-on practical using Galaxy In this exercise we will cover some of the basic NGS analysis steps for ChIP-seq using the Galaxy framework: Quality control Mapping of reads using Bowtie2 Peak-calling
ChIP-seq hands-on practical using Galaxy In this exercise we will cover some of the basic NGS analysis steps for ChIP-seq using the Galaxy framework: Quality control Mapping of reads using Bowtie2 Peak-calling
Creating An Account With Outlook
 Creating An Email Account With Outlook In this lesson we will show you how to create an email account and go through some of the basic functionality of the account. Lesson 1: Accessing The Website and
Creating An Email Account With Outlook In this lesson we will show you how to create an email account and go through some of the basic functionality of the account. Lesson 1: Accessing The Website and
DIRECTV RESIDENTIAL EXPERIENCE
 DIRECTV RESIDENTIAL EXPERIENCE Guest Welcome Screen PRO Quick Start Guide Table of Contents Overview... 3 Content Types... 4 DRE Theme... 5 Create a Theme... 5 Assign a Theme... 6 DRE Home Screen... 7
DIRECTV RESIDENTIAL EXPERIENCE Guest Welcome Screen PRO Quick Start Guide Table of Contents Overview... 3 Content Types... 4 DRE Theme... 5 Create a Theme... 5 Assign a Theme... 6 DRE Home Screen... 7
Administrative Training Mura CMS Version 5.6
 Administrative Training Mura CMS Version 5.6 Published: March 9, 2012 Table of Contents Mura CMS Overview! 6 Dashboard!... 6 Site Manager!... 6 Drafts!... 6 Components!... 6 Categories!... 6 Content Collections:
Administrative Training Mura CMS Version 5.6 Published: March 9, 2012 Table of Contents Mura CMS Overview! 6 Dashboard!... 6 Site Manager!... 6 Drafts!... 6 Components!... 6 Categories!... 6 Content Collections:
Protocol: peak-calling for ChIP-seq data / segmentation analysis for histone modification data
 Protocol: peak-calling for ChIP-seq data / segmentation analysis for histone modification data Table of Contents Protocol: peak-calling for ChIP-seq data / segmentation analysis for histone modification
Protocol: peak-calling for ChIP-seq data / segmentation analysis for histone modification data Table of Contents Protocol: peak-calling for ChIP-seq data / segmentation analysis for histone modification
Cyverse tutorial 1 Logging in to Cyverse and data management. Open an Internet browser window and navigate to the Cyverse discovery environment:
 Cyverse tutorial 1 Logging in to Cyverse and data management Open an Internet browser window and navigate to the Cyverse discovery environment: https://de.cyverse.org/de/ Click Log in with your CyVerse
Cyverse tutorial 1 Logging in to Cyverse and data management Open an Internet browser window and navigate to the Cyverse discovery environment: https://de.cyverse.org/de/ Click Log in with your CyVerse
Visual Dialogue User Guide. Version 6.0
 Visual Dialogue User Guide Version 6.0 2013 Pitney Bowes Software Inc. All rights reserved. This document may contain confidential and proprietary information belonging to Pitney Bowes Inc. and/or its
Visual Dialogue User Guide Version 6.0 2013 Pitney Bowes Software Inc. All rights reserved. This document may contain confidential and proprietary information belonging to Pitney Bowes Inc. and/or its
Student eportfolio Step-by-step Guide Student eportfolio Step-by-step Guide
 Student eportfolio Step-by-step Guide Section 1 Adding an Experience Section 2 File Manager and Artefacts Section 3 Adding an Artefact Section 4 eportfolio Views Section 5 Releasing your eportfolio to
Student eportfolio Step-by-step Guide Section 1 Adding an Experience Section 2 File Manager and Artefacts Section 3 Adding an Artefact Section 4 eportfolio Views Section 5 Releasing your eportfolio to
7.36/7.91/20.390/20.490/6.802/6.874 PROBLEM SET 3. Gibbs Sampler, RNA secondary structure, Protein Structure with PyRosetta, Connections (25 Points)
 7.36/7.91/20.390/20.490/6.802/6.874 PROBLEM SET 3. Gibbs Sampler, RNA secondary structure, Protein Structure with PyRosetta, Connections (25 Points) Due: Thursday, April 3 th at noon. Python Scripts All
7.36/7.91/20.390/20.490/6.802/6.874 PROBLEM SET 3. Gibbs Sampler, RNA secondary structure, Protein Structure with PyRosetta, Connections (25 Points) Due: Thursday, April 3 th at noon. Python Scripts All
Working with the Document Library
 Working with the Document Library The HQ Document Library The Document Library is a vital part of the complete document management system in HQ. The fields created using the Document Library may be accessible
Working with the Document Library The HQ Document Library The Document Library is a vital part of the complete document management system in HQ. The fields created using the Document Library may be accessible
QUICK REFERENCE GUIDE
 Folders new projects. Organise your folders to find files quickly and easily 1 Look in your yellow storage Folders it can be organised into simple folder structures to help with browsing 2 Click on your
Folders new projects. Organise your folders to find files quickly and easily 1 Look in your yellow storage Folders it can be organised into simple folder structures to help with browsing 2 Click on your
How to Export a Report in Cognos Analytics
 IBM Cognos Analytics How to Export a Report in Cognos Analytics Reports viewed in IBM Cognos Analytics can be exported in many formats including Excel. Some of the steps for exporting are different depending
IBM Cognos Analytics How to Export a Report in Cognos Analytics Reports viewed in IBM Cognos Analytics can be exported in many formats including Excel. Some of the steps for exporting are different depending
1 Installing the ZENworks Content Reporting Package
 ZENworks Content Reporting Package November 21, 2013 Novell The ZENworks Content Reporting Package is a ZENworks Reporting 5 domain, a set of predefined ad hoc views, and a set of default reports that
ZENworks Content Reporting Package November 21, 2013 Novell The ZENworks Content Reporting Package is a ZENworks Reporting 5 domain, a set of predefined ad hoc views, and a set of default reports that
How to use earray to create custom content for the SureSelect Target Enrichment platform. Page 1
 How to use earray to create custom content for the SureSelect Target Enrichment platform Page 1 Getting Started Access earray Access earray at: https://earray.chem.agilent.com/earray/ Log in to earray,
How to use earray to create custom content for the SureSelect Target Enrichment platform Page 1 Getting Started Access earray Access earray at: https://earray.chem.agilent.com/earray/ Log in to earray,
Submitting a Form 11 using the ROS Offline Application
 Submitting a Form 11 using the ROS Offline Application To work on your Form 11 offline you must use the ROS Offline Application. Instructions on how to install the ROS Offline Application are available
Submitting a Form 11 using the ROS Offline Application To work on your Form 11 offline you must use the ROS Offline Application. Instructions on how to install the ROS Offline Application are available
Analyzing ChIP- Seq Data in Galaxy
 Analyzing ChIP- Seq Data in Galaxy Lauren Mills RISS ABSTRACT Step- by- step guide to basic ChIP- Seq analysis using the Galaxy platform. Table of Contents Introduction... 3 Links to helpful information...
Analyzing ChIP- Seq Data in Galaxy Lauren Mills RISS ABSTRACT Step- by- step guide to basic ChIP- Seq analysis using the Galaxy platform. Table of Contents Introduction... 3 Links to helpful information...
Drupal 7 guide CONTENTS. p. 2 Logging In
 Drupal 7 guide Drupal is a widely used, open-source, free platform that has an easy-to-use content management system for updating websites. This guide was created by the Health Communication Core (www.healthcommcore.org)
Drupal 7 guide Drupal is a widely used, open-source, free platform that has an easy-to-use content management system for updating websites. This guide was created by the Health Communication Core (www.healthcommcore.org)
Creating an Account and Using Universal Job Match
 Creating an Account and Using Universal Job Match In this hand-out we will take you through setting up an account with UJM and show you some of the functionality If you haven t already, create an email
Creating an Account and Using Universal Job Match In this hand-out we will take you through setting up an account with UJM and show you some of the functionality If you haven t already, create an email
epigenomegateway.wustl.edu
 Everything can be found at epigenomegateway.wustl.edu REFERENCES 1. Zhou X, et al., Nature Methods 8, 989-990 (2011) 2. Zhou X & Wang T, Current Protocols in Bioinformatics Unit 10.10 (2012) 3. Zhou X,
Everything can be found at epigenomegateway.wustl.edu REFERENCES 1. Zhou X, et al., Nature Methods 8, 989-990 (2011) 2. Zhou X & Wang T, Current Protocols in Bioinformatics Unit 10.10 (2012) 3. Zhou X,
ChIP-seq hands-on practical using Galaxy
 ChIP-seq hands-on practical using Galaxy In this exercise we will cover some of the basic NGS analysis steps for ChIP-seq using the Galaxy framework: Quality control Mapping of reads using Bowtie2 Peak-calling
ChIP-seq hands-on practical using Galaxy In this exercise we will cover some of the basic NGS analysis steps for ChIP-seq using the Galaxy framework: Quality control Mapping of reads using Bowtie2 Peak-calling
Blackbaud Direct Marketing New Features Guide
 Blackbaud Direct Marketing New Features Guide 05/09/2018 Blackbaud Direct Marketing 5.0 Blackbaud Direct Marketing New Features US 2018 Blackbaud, Inc. This publication, or any part thereof, may not be
Blackbaud Direct Marketing New Features Guide 05/09/2018 Blackbaud Direct Marketing 5.0 Blackbaud Direct Marketing New Features US 2018 Blackbaud, Inc. This publication, or any part thereof, may not be
On the go? Take Premier with you.
 On the go? Take Premier with you. Premier Online Care mobile enhancements Release Notes May 2017 Manage your accounts on the fly Easily manage your Premier accounts from your smartphone. When Premier detects
On the go? Take Premier with you. Premier Online Care mobile enhancements Release Notes May 2017 Manage your accounts on the fly Easily manage your Premier accounts from your smartphone. When Premier detects
TurningPoint AnyWhere
 TurningPoint AnyWhere TurningPoint Blackboard Registration Tool Making the Tool Available 1. From the Control Panel, select click Customization >>Tool Availability. 2. From the Tools list, check Registration
TurningPoint AnyWhere TurningPoint Blackboard Registration Tool Making the Tool Available 1. From the Control Panel, select click Customization >>Tool Availability. 2. From the Tools list, check Registration
Lab 2 Building on Linux
 Lab 2 Building on Linux Assignment Details Assigned: January 28 th, 2013. Due: January 30 th, 2013 at midnight. Background This assignment should introduce the basic development tools on Linux. This assumes
Lab 2 Building on Linux Assignment Details Assigned: January 28 th, 2013. Due: January 30 th, 2013 at midnight. Background This assignment should introduce the basic development tools on Linux. This assumes
Creating & Using Forms in Contensis 15 th March 2016
 Creating & Using Forms in Contensis 15 th March 2016 Contents Introduction... 2 Creating the actual form... 2 Fields that can be added to the form... 3 To add a field to a form... 4 Adding further fields
Creating & Using Forms in Contensis 15 th March 2016 Contents Introduction... 2 Creating the actual form... 2 Fields that can be added to the form... 3 To add a field to a form... 4 Adding further fields
Mapping RNA sequence data (Part 1: using pathogen portal s RNAseq pipeline) Exercise 6
 Mapping RNA sequence data (Part 1: using pathogen portal s RNAseq pipeline) Exercise 6 The goal of this exercise is to retrieve an RNA-seq dataset in FASTQ format and run it through an RNA-sequence analysis
Mapping RNA sequence data (Part 1: using pathogen portal s RNAseq pipeline) Exercise 6 The goal of this exercise is to retrieve an RNA-seq dataset in FASTQ format and run it through an RNA-sequence analysis
Focus Group Analysis
 Focus Group Analysis Contents FOCUS GROUP ANALYSIS... 1 HOW CAN MAXQDA SUPPORT FOCUS GROUP ANALYSIS?... 1 IMPORTING FOCUS GROUP TRANSCRIPTS... 1 TRANFORMING AN ALREADY IMPORTED TEXT INTO A FOCUS GROUP
Focus Group Analysis Contents FOCUS GROUP ANALYSIS... 1 HOW CAN MAXQDA SUPPORT FOCUS GROUP ANALYSIS?... 1 IMPORTING FOCUS GROUP TRANSCRIPTS... 1 TRANFORMING AN ALREADY IMPORTED TEXT INTO A FOCUS GROUP
CLC Server. End User USER MANUAL
 CLC Server End User USER MANUAL Manual for CLC Server 10.0.1 Windows, macos and Linux March 8, 2018 This software is for research purposes only. QIAGEN Aarhus Silkeborgvej 2 Prismet DK-8000 Aarhus C Denmark
CLC Server End User USER MANUAL Manual for CLC Server 10.0.1 Windows, macos and Linux March 8, 2018 This software is for research purposes only. QIAGEN Aarhus Silkeborgvej 2 Prismet DK-8000 Aarhus C Denmark
HymenopteraMine Documentation
 HymenopteraMine Documentation Release 1.0 Aditi Tayal, Deepak Unni, Colin Diesh, Chris Elsik, Darren Hagen Apr 06, 2017 Contents 1 Welcome to HymenopteraMine 3 1.1 Overview of HymenopteraMine.....................................
HymenopteraMine Documentation Release 1.0 Aditi Tayal, Deepak Unni, Colin Diesh, Chris Elsik, Darren Hagen Apr 06, 2017 Contents 1 Welcome to HymenopteraMine 3 1.1 Overview of HymenopteraMine.....................................
Working with Attributes
 Working with Attributes QGIS Tutorials and Tips Author Ujaval Gandhi http://www.spatialthoughts.com This work is licensed under a Creative Commons Attribution 4.0 International License. Working with Attributes
Working with Attributes QGIS Tutorials and Tips Author Ujaval Gandhi http://www.spatialthoughts.com This work is licensed under a Creative Commons Attribution 4.0 International License. Working with Attributes
Survey Creation Workflow These are the high level steps that are followed to successfully create and deploy a new survey:
 Overview of Survey Administration The first thing you see when you open up your browser to the Ultimate Survey Software is the Login Page. You will find that you see three icons at the top of the page,
Overview of Survey Administration The first thing you see when you open up your browser to the Ultimate Survey Software is the Login Page. You will find that you see three icons at the top of the page,
Module 1 Artemis. Introduction. Aims IF YOU DON T UNDERSTAND, PLEASE ASK! -1-
 Module 1 Artemis Introduction Artemis is a DNA viewer and annotation tool, free to download and use, written by Kim Rutherford from the Sanger Institute (Rutherford et al., 2000). The program allows the
Module 1 Artemis Introduction Artemis is a DNA viewer and annotation tool, free to download and use, written by Kim Rutherford from the Sanger Institute (Rutherford et al., 2000). The program allows the
SharePoint Bridges Agency User Training
 SharePoint 2010 What is SharePoint? SharePoint was designed to assist organizations in sharing various types of content and information. SharePoint 2010 allows you to manage content and business processes
SharePoint 2010 What is SharePoint? SharePoint was designed to assist organizations in sharing various types of content and information. SharePoint 2010 allows you to manage content and business processes
Structural Bioinformatics
 Structural Bioinformatics Elucidation of the 3D structures of biomolecules. Analysis and comparison of biomolecular structures. Prediction of biomolecular recognition. Handles three-dimensional (3-D) structures.
Structural Bioinformatics Elucidation of the 3D structures of biomolecules. Analysis and comparison of biomolecular structures. Prediction of biomolecular recognition. Handles three-dimensional (3-D) structures.
Creating Dashboard Widgets. Version: 16.0
 Creating Dashboard Widgets Version: 16.0 Copyright 2017 Intellicus Technologies This document and its content is copyrighted material of Intellicus Technologies. The content may not be copied or derived
Creating Dashboard Widgets Version: 16.0 Copyright 2017 Intellicus Technologies This document and its content is copyrighted material of Intellicus Technologies. The content may not be copied or derived
User s Guide: BIOS Online Mapping Tool to access the California Freshwater Species Database
 User s Guide: BIOS Online Mapping Tool to access the California Freshwater Species Database Last updated: December 2017 Tutorial created by: The Nature Conservancy Topics covered Part 1: Exporting a tabular
User s Guide: BIOS Online Mapping Tool to access the California Freshwater Species Database Last updated: December 2017 Tutorial created by: The Nature Conservancy Topics covered Part 1: Exporting a tabular
In this lesson you will learn how to:
 LESSON 5: CREATING BUTTON STATES OBJECTIVES In this lesson you will learn how to: use FreeHand layers to create navigation buttons export layers from FreeHand to Flash create and edit symbols and instances
LESSON 5: CREATING BUTTON STATES OBJECTIVES In this lesson you will learn how to: use FreeHand layers to create navigation buttons export layers from FreeHand to Flash create and edit symbols and instances
The address is:
 M a n ti s U s e r G u i d e L o g i n p a g e The address is: http://discoverysupport.reply.it/mantis/login_page.php Just enter your username and password and hit the login button. There is also a Save
M a n ti s U s e r G u i d e L o g i n p a g e The address is: http://discoverysupport.reply.it/mantis/login_page.php Just enter your username and password and hit the login button. There is also a Save
GEP Project Management System: TSS Project Submission
 GEP Project Management System: TSS Project Submission Author Wilson Leung wleung@wustl.edu Document History Initial Draft 08/21/2015 Version GEP Project Management System (Version alpha) Introduction In
GEP Project Management System: TSS Project Submission Author Wilson Leung wleung@wustl.edu Document History Initial Draft 08/21/2015 Version GEP Project Management System (Version alpha) Introduction In
Basic Query for Human Resources
 Basic Query for Human Resources Open browser Log into PeopleSoft Human Resources: Go to: https://cubshr9.clemson.edu/psp/hpprd/?cmd=login Enter your Novell ID and Password Click Sign In Navigation into
Basic Query for Human Resources Open browser Log into PeopleSoft Human Resources: Go to: https://cubshr9.clemson.edu/psp/hpprd/?cmd=login Enter your Novell ID and Password Click Sign In Navigation into
Introduction to SAGA GIS
 GIS Tutorial ID: IGET_RS_001 This tutorial has been developed by BVIEER as part of the IGET web portal intended to provide easy access to geospatial education. This tutorial is released under the Creative
GIS Tutorial ID: IGET_RS_001 This tutorial has been developed by BVIEER as part of the IGET web portal intended to provide easy access to geospatial education. This tutorial is released under the Creative
Cytidine-to-Uridine Recognizing Editor for Chloroplasts
 For Chloroplasts Cytidine-to-Uridine Recognizing Editor for Chloroplasts A Chloroplasts C-to-U RNA editing site prediction tool A User Manual Pufeng Du, Liyan Jia and Yanda Li MOE Key Laboratory of Bioinformatics
For Chloroplasts Cytidine-to-Uridine Recognizing Editor for Chloroplasts A Chloroplasts C-to-U RNA editing site prediction tool A User Manual Pufeng Du, Liyan Jia and Yanda Li MOE Key Laboratory of Bioinformatics
HMIS Guide to Running & Retrieving the CSV APR 2017 Export in ClientTrack An HMIS End User Training Resource
 2018 HMIS Guide to Running & Retrieving the CSV APR 2017 Export in ClientTrack An HMIS End User Training Resource This guide will demonstrate how to run the CSV APR 2017 Export and prepare files for submission
2018 HMIS Guide to Running & Retrieving the CSV APR 2017 Export in ClientTrack An HMIS End User Training Resource This guide will demonstrate how to run the CSV APR 2017 Export and prepare files for submission
More Skills 11 Export Queries to Other File Formats
 = CHAPTER 2 Access More Skills 11 Export Queries to Other File Formats Data from a table or query can be exported into file formats that are opened with other applications such as Excel and Internet Explorer.
= CHAPTER 2 Access More Skills 11 Export Queries to Other File Formats Data from a table or query can be exported into file formats that are opened with other applications such as Excel and Internet Explorer.
DAVID hands-on. by Ester Feldmesser, June 2017
 DAVID hands-on by Ester Feldmesser, June 2017 1. Go to the DAVID website (http://david.abcc.ncifcrf.gov/) 2. Press on Start Analysis: 3. Choose the Upload tab in the left panel: 4. Download the k-means5_arabidopsis.txt
DAVID hands-on by Ester Feldmesser, June 2017 1. Go to the DAVID website (http://david.abcc.ncifcrf.gov/) 2. Press on Start Analysis: 3. Choose the Upload tab in the left panel: 4. Download the k-means5_arabidopsis.txt
Wilson Leung 01/03/2018 An Introduction to NCBI BLAST. Prerequisites: Detecting and Interpreting Genetic Homology: Lecture Notes on Alignment
 An Introduction to NCBI BLAST Prerequisites: Detecting and Interpreting Genetic Homology: Lecture Notes on Alignment Resources: The BLAST web server is available at https://blast.ncbi.nlm.nih.gov/blast.cgi
An Introduction to NCBI BLAST Prerequisites: Detecting and Interpreting Genetic Homology: Lecture Notes on Alignment Resources: The BLAST web server is available at https://blast.ncbi.nlm.nih.gov/blast.cgi
Getting Started (New Accounts)
 Getting Started (New Accounts) 1. On any page with the menu, go to the faculty section and choose Faculty Website Access. 2. On the login page, make sure you are on Windows Login. Login with the username
Getting Started (New Accounts) 1. On any page with the menu, go to the faculty section and choose Faculty Website Access. 2. On the login page, make sure you are on Windows Login. Login with the username
Adding Pages. Adding pages to your website is simple and powerful! In just a few minutes you can create a page that: Highlights a special event
 Adding Pages Adding pages to your website is simple and powerful! In just a few minutes you can create a page that: Highlights a special event Collects entries on a registration form for a promotional
Adding Pages Adding pages to your website is simple and powerful! In just a few minutes you can create a page that: Highlights a special event Collects entries on a registration form for a promotional
Generate a Report on Submitted Invoices
 1 / 6 How To Generate a Report on Submitted Invoices 1. Go to the OB10 Portal - Go to www.ob10.com. - Click on the Login button. - Enter your login credentials. - If you need help, refer to the How to
1 / 6 How To Generate a Report on Submitted Invoices 1. Go to the OB10 Portal - Go to www.ob10.com. - Click on the Login button. - Enter your login credentials. - If you need help, refer to the How to
Ion AmpliSeq Designer: Getting Started
 Ion AmpliSeq Designer: Getting Started USER GUIDE Publication Number MAN0010907 Revision F.0 For Research Use Only. Not for use in diagnostic procedures. Manufacturer: Life Technologies Corporation Carlsbad,
Ion AmpliSeq Designer: Getting Started USER GUIDE Publication Number MAN0010907 Revision F.0 For Research Use Only. Not for use in diagnostic procedures. Manufacturer: Life Technologies Corporation Carlsbad,
Using Galaxy for NGS Analyses Luce Skrabanek
 Using Galaxy for NGS Analyses Luce Skrabanek Registering for a Galaxy account Before we begin, first create an account on the main public Galaxy portal. Go to: https://main.g2.bx.psu.edu/ Under the User
Using Galaxy for NGS Analyses Luce Skrabanek Registering for a Galaxy account Before we begin, first create an account on the main public Galaxy portal. Go to: https://main.g2.bx.psu.edu/ Under the User
You can make certain sections of the text clickable by creating hyperlinks. Once clicked, these links navigate users to different
 You can make certain sections of the text clickable by creating hyperlinks. Once clicked, these links navigate users to different pages or, as described in working with anchors, to different sections of
You can make certain sections of the text clickable by creating hyperlinks. Once clicked, these links navigate users to different pages or, as described in working with anchors, to different sections of
Proteome Comparison: A fine-grained tool for comparative genomics
 Proteome Comparison: A fine-grained tool for comparative genomics In addition to the Protein Family Sorter that allows researchers to examine up to the protein families from up to 500 genomes at a time,
Proteome Comparison: A fine-grained tool for comparative genomics In addition to the Protein Family Sorter that allows researchers to examine up to the protein families from up to 500 genomes at a time,
Lession #5: Adding a New Salary Using Batch Uploads
 Lession #5: Adding a New Salary Using Batch Uploads In Lesson #4, we discussed the function of adding or updating a salary online through IWAS. In this lesson, we ll discuss performing the same function,
Lession #5: Adding a New Salary Using Batch Uploads In Lesson #4, we discussed the function of adding or updating a salary online through IWAS. In this lesson, we ll discuss performing the same function,
Figure 1 Forms category in the Insert panel. You set up a form by inserting it and configuring options through the Properties panel.
 Adobe Dreamweaver CS6 Project 3 guide How to create forms You can use forms to interact with or gather information from site visitors. With forms, visitors can provide feedback, sign a guest book, take
Adobe Dreamweaver CS6 Project 3 guide How to create forms You can use forms to interact with or gather information from site visitors. With forms, visitors can provide feedback, sign a guest book, take
1. CONTENTS 2. OVERVIEW CANDIDATES Add Candidate Update Candidate Details CVs & CV to Send...
 User Guide 1. CONTENTS 2. OVERVIEW... 4 3. CANDIDATES... 4 3.1. Add Candidate... 4 3.2. Update Candidate Details... 4 3.3. CVs & CV to Send... 4 3.4. Apply Candidate to Vacancy... 5 3.5. Quick Links...
User Guide 1. CONTENTS 2. OVERVIEW... 4 3. CANDIDATES... 4 3.1. Add Candidate... 4 3.2. Update Candidate Details... 4 3.3. CVs & CV to Send... 4 3.4. Apply Candidate to Vacancy... 5 3.5. Quick Links...
CAL 9-2: Café Soylent Green Chapter 12
 CAL 9-2: Café Soylent Green Chapter 12 This version is for those students who are using Dreamweaver CC. You will be completing the Forms Tutorial from your textbook, Chapter 12 however, you will be skipping
CAL 9-2: Café Soylent Green Chapter 12 This version is for those students who are using Dreamweaver CC. You will be completing the Forms Tutorial from your textbook, Chapter 12 however, you will be skipping
Web CMS Sub Administrator Training
 Web CMS Sub Administrator Training - Introduction... 2 User Administration... 2 User Roles... 2 Administrator... 2 Sub Administrator... 2 Content Contributor... 2 Site User... 3 Overview of User Management...
Web CMS Sub Administrator Training - Introduction... 2 User Administration... 2 User Roles... 2 Administrator... 2 Sub Administrator... 2 Content Contributor... 2 Site User... 3 Overview of User Management...
Submitting Assignments
 Submitting Assignments Blackboard s assignments feature allows the instructor to assign coursework for you to submit electronically. First, you need to locate the assignment. Your instructor will place
Submitting Assignments Blackboard s assignments feature allows the instructor to assign coursework for you to submit electronically. First, you need to locate the assignment. Your instructor will place
Precalculus An Investigation of Functions
 Precalculus An Investigation of Functions David Lippman Melonie Rasmussen Edition 1.3 This book is also available to read free online at http://www.opentextbookstore.com/precalc/ If you want a printed
Precalculus An Investigation of Functions David Lippman Melonie Rasmussen Edition 1.3 This book is also available to read free online at http://www.opentextbookstore.com/precalc/ If you want a printed
Quick Reference Guide for Students: Applying for Course or Campus Transfer
 Quick Reference Guide for Students: Applying for Course or Campus Transfer This guide contains information for current Monash students applying for a Course or Campus Transfer using a new online course
Quick Reference Guide for Students: Applying for Course or Campus Transfer This guide contains information for current Monash students applying for a Course or Campus Transfer using a new online course
Accommodations Upload Quick Guide Oklahoma School Testing Program & College and Career Readiness Assessments Spring 2018
 Accommodations Upload Quick Guide Oklahoma School Testing Program & College and Career Readiness Assessments Spring 2018 1 Table of Contents Extracting the emetric Report in OK EdPlan... 3 Uploading to
Accommodations Upload Quick Guide Oklahoma School Testing Program & College and Career Readiness Assessments Spring 2018 1 Table of Contents Extracting the emetric Report in OK EdPlan... 3 Uploading to
USER GUIDE. PowerPhoto CRM 2011
 USER GUIDE PowerPhoto CRM 2011 Contents Placing the PowerPhoto Contol on an Entity Moving PowerPhoto on the Form Configuring PowerPhoto to Support Multiple Photos Using PowerPhoto Add Drag and Drop Paste
USER GUIDE PowerPhoto CRM 2011 Contents Placing the PowerPhoto Contol on an Entity Moving PowerPhoto on the Form Configuring PowerPhoto to Support Multiple Photos Using PowerPhoto Add Drag and Drop Paste
USING THE BU BRAIN CLONE SCHEDULE OF CLASSES QUERY
 BU Brain Clone Schedule of Classes Query Use the BU Brain Clone Query (titled BU Brain Clone Schedule of Classes Query and stored on the On Demand Server in the * BU Canned Queries/Catalog & Course Schedules
BU Brain Clone Schedule of Classes Query Use the BU Brain Clone Query (titled BU Brain Clone Schedule of Classes Query and stored on the On Demand Server in the * BU Canned Queries/Catalog & Course Schedules
Course Exercises for the Content Management System. Grazyna Whalley, Laurence Cornford June 2014 AP-CMS2.0. University of Sheffield
 Course Exercises for the Content Management System. Grazyna Whalley, Laurence Cornford June 2014 AP-CMS2.0 University of Sheffield PART 1 1.1 Getting Started 1. Log on to the computer with your usual username
Course Exercises for the Content Management System. Grazyna Whalley, Laurence Cornford June 2014 AP-CMS2.0 University of Sheffield PART 1 1.1 Getting Started 1. Log on to the computer with your usual username
