BrainPrint : Identifying Subjects by their Brain
|
|
|
- Gillian Edwards
- 5 years ago
- Views:
Transcription
1 BrainPrint : Identifying Subjects by their Brain Christian Wachinger 1,2, Polina Golland 1, Martin Reuter 1,2 1 Computer Science and Artificial Intelligence Lab, MIT 2 Massachusetts General Hospital, Harvard Medical School Abstract. Introducing BrainPrint, a compact and discriminative representation of anatomical structures in the brain. BrainPrint captures shape information of an ensemble of cortical and subcortical structures by solving the 2D and 3D Laplace-Beltrami operator on triangular (boundary) and tetrahedral (volumetric) meshes. We derive a robust classifier for this representation that identifies the subject in a new scan, based on a database of brain scans. In an example dataset containing over 3000 MRI scans, we show that BrainPrint captures unique information about the subject s anatomy and permits to correctly classify a scan with an accuracy of over 99.8%. All processing steps for obtaining the compact representation are fully automated making this processing framework particularly attractive for handling large datasets. 1 Introduction Is it possible to identify an individual based on their brain? Are cortical folding patterns unique to a person, similar to a fingerprint? While the unique complexity of the brain may indicate that an unambiguous identification should be possible, there is currently little empirical research that can speak to these questions. One difficulty for identifying the subject of a given brain is that longitudinal changes caused by aging or disease may significantly alter the brain morphometry. Additionally, scanning artifacts, inhomogeneities, and different imaging protocols can cause changes in intensity values in magnetic resonance scans, further complicating the identification. Therefore, a subject-specific brain signature must be both stable across time and insensitive to imaging artifacts. Moreover, it needs to provide a holistic representation of the brain to ensure subject identification even if certain parts change. Finally, small changes in the brain should map to small changes in the representation to permit a robust identification. Here, we introduce BrainPrint, a holistic representation of the brain anatomy, containing the shape information of an ensemble of cortical and subcortical structures. The inclusion of only shape information has the advantage to remain independent from the local intensity values in the scan. Moreover, the variety of the different structures included in the BrainPrint yields an extensive characterization of the brain anatomy. We quantify the shape information by calculating the spectrum of the Laplace-Beltrami operator (LBO) on both triangular meshes that represent boundary surfaces, e.g., the white matter surface, and tetrahedral
2 2 C. Wachinger, P. Golland, M. Reuter meshes for volumetric representations of individual structures. We then derive a classifier that identifies a subject from an MRI scan based on its BrainPrint. We achieve robustness in the identification by letting each brain structure vote independently for the subject s identity. Not only does our classifier identify previously encountered subjects with high accuracy, but it also determines whether a query brain belongs to an unknown subject, not yet represented in the existing database. An alternative approach to calculate the similarity between scans could be based on image registration [3, 5]. However, real applications of such identification methods require large datasets and the cost for aligning a new scan to all scans in the database becomes prohibitive for a large number of scans. Brain- Print introduces a new framework that is especially beneficial when working with large datasets widely available today. The first step extracts information from the image, based on the segmentation of anatomical structures. The second step transfers this information into a compact and discriminative representation, the BrainPrint. Any further processing is conducted on this representation, which takes less memory and permits easier calculations and comparisons than the original scan. 1.1 Related Work A 3D object can be represented by the space that it occupies (3D volume representation, e.g., voxels, tetrahedra meshes) or by representing its boundary (2D surface representation, e.g., triangle meshes). Reuter et al. [10] introduced the shapedna and demonstrated that the spectra of 3D solid objects and their 2D boundary surfaces contain complementary information: the spectra of the 2D boundary surface was capable of distinguishing two isospectral 3D solids (GWW-prisms). Therefore, we propose to combine the information from both the 3D solid and 2D boundary shape representations. While there has been previous work analyzing the shapedna for single brain structures [1, 9, 11], to the best of our knowledge this is the first study that evaluates its application to cortical structures and a wide range of subcortical structures. Importantly, we investigate the joint modeling of the ensemble. Additionally, most prior work computes the shapedna for triangular surface meshes [1, 8], while we also work with tetrahedral volume tessellations. Given that the Laplace spectra are isometry invariant, the 2D boundary representation alone may yield a weaker descriptor, due to the large set of potential (near-) isometric deformations. For example, a closed 2D surface with a protrusion pointing inwards yields the same descriptor as one with the protrusion pointing outwards, while the spectra of the enclosed 3D solids differ. 2 Shape Descriptor We segment anatomical structures from brain scans with FreeSurfer [2]. Next, we compute a compact shape representation that captures important shape information and facilitates the further processing. Since image intensity varies across
3 BrainPrint: Identifying Subjects by their Brain 3 Volume 3.2 x Subject Mean LGI Subject Fig. 1: Mean and standard deviation of the volume (left) and of the mean local gyrification index (right) of the cortex for 40 subjects. Statistics are calculated over several longitudinal scans per subject. scans, we focus on geometrical properties. Example representations are the volume and the local gyrification index (LGI) of a structure. While volume will be affected by brain atrophy, quantifying the gyrification may be more robust to longitudinal changes, assuming that the folding patterns of the brain remain stable. The LGI was used previously to identify gyral abnormalities [12]. We transform this local measure into a global shape descriptor by computing the mean LGI over the surface. Fig. 1 shows the mean and standard deviation of these measures calculated from several longitudinal scans per subject. The large variance and overlap across subjects indicates that such representations are not well suited for identifying subjects. In this work we use the shapedna [10] as a shape descriptor, which performed among the best in a recent comparison of methods for non-rigid 3D shape retrieval [6]. The ShapeDNA is computed from the intrinsic geometry of an object by calculating the Laplace-Beltrami spectrum. Considering the Laplace- Beltrami operator, we obtain the spectrum by solving the Laplacian eigenvalue problem (Helmholtz equation) f = λf using the finite element method. The solution consists of eigenvalue λ i R and eigenfunction f i pairs (sorted by eigenvalues, 0 λ 1 λ 2...). To be independent of the objects scale, we normalize the eigenvalues λ = vol 2 D λ, where vol is the Riemannian volume of the D-dimensional manifold (i.e., the area for 2D surfaces) [10]. The first l non-zero eigenvalues form the shapedna: λ = (λ 1,..., λ l ). The eigenvalues are isometry invariant with respect to the Riemannian manifold, meaning that length-preserving deformations will not change the spectrum. This important property permits the comparison of subjects by directly comparing the shapedna, without the need for alignment. While isometric noncongruent surfaces exist (e.g., bending a sheet of paper), two solid bodies embedded in R 3 are isometric if and only if they are congruent (translated, rotated and mirrored). A second property is that the spectrum continuously changes with topology-preserving deformations of the object boundary. Fig. 2 illustrates the eigenfunctions of the cerebral cortex boundary. The eigenfunctions show natural
4 4 C. Wachinger, P. Golland, M. Reuter Fig. 2: Left cerebral cortex and first eigenfunctions of the LBO calculated on the surface (yellow positive, red negative, and green zero). vibrations of the shape when oscillating at a frequency specified by the square root of the eigenvalue. We compute the spectra for all cortical and subcortical structures on the 2D boundary surfaces (triangle meshes) and additionally on the full 3D solid (tetrahedra meshes) for the cortical structures (white and pial surfaces in both hemispheres), forming the BrainPrint Λ = (λ1,..., λη ). Triangle meshes of the cortical surfaces are obtained automatically for each hemisphere using FreeSurfer. Surface meshes of subcortical structures are constructed via marching cubes from the FreeSurfer subcortical segmentation. To construct tetrahedral meshes, we remove handles from the surface meshes, uniformly resample the output to 60K vertices, and create the volumetric mesh with the gmsh package [4]. We use the linear finite element method [10] with Neumann boundary condition (zero normal derivative) to compute the spectra of the tetrahedral meshes. 3 Classifier We derive a classifier to assign a new scan to one of the subjects in the database. Since the segmentation or tessellation of specific structures may fail in certain cases, we propose a robust classifier that handles missing information. We build a classifier by combining the results from weak classifiers operating on specific brain structures. Assuming n subjects C1,..., Cn and N scans in a database (N n, for repeated scans of subjects). Each scan has its associated BrainPrint Λ1,..., ΛN. Let Sk {1,..., N } denote scans for subject Ck. The probability that a the new scan with BrainPrint Λ shows subject Ck is Y p(λ Ck ) p(ck ) p(ck Λ) = P p(λs Ck ), ν p(λ Cν ) p(cν ) s=1,...,η (1) where we assume a uniform class probability p(ck ) 1 and the conditional independence of structures given the subject. The likelihood is multivariate Pnormal distributed p(λs Ck ) N (λs ; µks, Σs ) with the subject mean µks = S1k i Sk λis for structure s. Since we only have a few samples per class, we estimate a global diagonal covariance matrix Σs across all scans for each structure. Weighting distances by the variance helps to prevent the domination by higher eigenvalues
5 BrainPrint: Identifying Subjects by their Brain 5 Classification Rate Cortical Triangular Cortical Tetrahedral Cortical Both Selection All All+Diff Number Eigenvalues Classification Rate Cortical Triangular Cortical Tetrahedral Cortical Both Selection All All+Diff Number Eigenvalues Fig. 3: Classification results for product classifier (left) and voting classifier (right) under variable number of eigenvalues and feature sets. that exhibit higher variation. The subject identity with the highest probability is assigned to the scan k = arg max p(c k Λ). (2) k The posterior probability of this classifier is the product of the posterior probabilities across all structures, cf. Eq. (1), which may be problematic for structures with low discriminative power. Many subcortical structures do not carry much distinctive shape information and can therefore negatively influence the overall probability. We therefore propose a second classifier that is specifically adapted to working with structures that are not very discriminative. Increased robustness is achieved by voting for each structure independently ks = arg max p(λ s C k ), s {1,..., η}, (3) k with the final vote set to the mode of the vote distribution. 4 Results We perform experiments on data from the Alzheimer s Disease Neuroimaging Initiative (ADNI) [7]. We work with over 3000 scans from almost 700 subjects, where each subject has between three and six longitudinal scans. Each T1- weighted image from the dataset is processed independently with FreeSurfer. We calculate 36 shape descriptors for subcortical structures and 8 descriptors for cortical structures (left/right, white/gray matter, 2D/3D). Additionally, we calculate the lateral differences of shapedna between left and right cortical structures to quantify asymmetries, resulting in 4 additional descriptors. We perform leave-one-out experiments by removing one scan from the dataset and by aiming to recover the correct identity. Fig. 3 reports the classification results for the product classifier in Eq.(2) and the structure-specific voting in Eq.(3). We report classification results as a function of the number of
6 6 C. Wachinger, P. Golland, M. Reuter Structures Votes Subject included Subject excluded Boundary Fig. 4: Left: Subject (color) voted for by each structure (row) for each scan (column). Cortical structures in first 8 rows, subcortical features below. Optimal feature response would show a color gradient from blue to red, since scans are sorted by subject. Right: Number of votes for the winning subject identity when the correct subject is included (blue) in the database and when it is excluded (green). Decision boundary at 4 votes (red) yields a 0.49% false negative rate. eigenvalues used to represent the shape. Additionally, we vary the set of brain structures in BrainPrint: cortical structures with triangular meshes (4), cortical structures with tetrahedral meshes (4), cortical structures for both mesh types (8), a selection of structures with the highest individual performances (15), all structures (44), and all structures with the lateral differences of cortical structures (48). The number of structures is shown in parentheses. The results demonstrate a clear difference between the two classifiers. The product classifier achieves the best performance when working with cortical triangular meshes. Adding more features, especially when working with all features, dramatically reduces the classification results. We observe an opposite behavior when working with the structure-specific voting. Subcortical structures alone yield the worst performance in this case. The combination of 3D solid and 2D boundary descriptors leads to a clear improvement. A further improvement is gained by adding subcortical structures. To further study this behavior, we examine the candidate subject that each structure votes for in Fig. 4. Each column corresponds to one scan and each row to one structure. The color indicates the subject number. Scans were sorted by subject; a perfect feature should show a color gradient from blue to red. The first 8 rows correspond to cortical structures, which exhibit the best performance. The remaining 36 rows show subcortical structures that perform worse than cortical structures and vary in their discriminative power. This explains the poor performance of the product classifier for the whole feature set, as weak features can obscure good features. In contrast, weak features do not degrade the performance of the voting classifier as long as weak features show no bias for a specific subject. The best performance of over 99.8% is achieved for 50 eigenvalues on all features with the additional difference features. For comparison, the classification rate for the mean LGI on both hemispheres is 1.0% for the product and 3.9% for the voting classifier. The classification rate for the volume, calculated from all cortical and subcortical structures, is 0.03% for the product and 0.6% for the voting classifier, confirming results from Fig. 1.
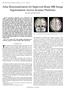
7 BrainPrint: Identifying Subjects by their Brain 7 Fig. 5: Coronal and axial slices from two misclassified scans. White matter segmentation is shown in yellow. Fig. 5 shows the two scans for which BrainPrint does not correctly identify the subject identity. These subjects show strong atrophy and imaging artifacts, resulting in pronounced segmentation errors. Manual correction in FreeSurfer or reacquisition to avoid motion artifacts may therefore improve the above results. As an additional experiment, we evaluate the possibility to determine whether a subject is not contained in the database. We study the number of votes the winning subject receives in Fig. 4, once when the subject of the scan is included in the database and once when the subject is excluded. If the subject in the current scan exists in the database, the scan receives about 15 votes for the winning subject class. If the subject is not contained in the database, the number of votes for the winner does not surpass 4. Setting 4 votes as our decision boundary results in only a 0.49% error (false negative) of concluding incorrectly that a subject is not in the database. The false positive rate is zero. 5 Discussion and Conclusions The high classification accuracy of BrainPrint suggests that brain structures are unique to individuals and can be used for identification. Since our study only includes data on subjects followed over a period of up to 36 months, we cannot currently assess how the accuracy of BrainPrint changes across the entire lifespan of a subject. Unfortunately, such data sets are not yet available. However, since subjects with Alzheimer s disease in our dataset demonstrate pronounced neurodegeneration in a relatively short time, we are optimistic that BrainPrint will remain robust for comparison across longer time periods. The identification accuracy may raise concerns about privacy issues when publicly distributing de-faced or skull-stripped brain scans together with diagnosis and other sensitive information. Yet, we currently do not think that BrainPrint interferes with anonymization because at least a second scan with knowledge of the identity needs to be available to connect to the private information. In terms of its practical applications, we see BrainPrint as an aid when handling large datasets. Identifying similar images in an efficient way can provide the launchpad for a more detailed follow-up analysis, e.g., calculation or prediction of localized growth and shrinkage patterns. Since most of our re-
8 8 C. Wachinger, P. Golland, M. Reuter trieval errors are related to incorrect segmentations, our approach could also be used as an automatic quality control. Furthermore, BrainPrint can help identify anonymization errors (mismatch of subject identity), which are difficult to detect and can impede longitudinal studies. Finally, the presented framework of image understanding and compact characterization is relevant for handling large datasets in other fields and not limited to neuroscience. Acknowledgements: This work was supported in part by the Humboldt foundation, the Martinos Center for Biomedical Imaging (P41-RR014075, P41- EB015896), the National Alliance for Medical Image Computing (U54-EB005149) and the NeuroImaging Analysis Center (P41-EB015902). We thank Anna Rieckmann for revising the manuscript and the Alzheimer s Disease Neuroimaging Initiative (ADNI) for image data. References 1. Bates, J., Pafundi, D., Kanel, P., Liu, X., Mio, W.: Spectral signatures of point clouds and applications to detection of alzheimer s disease through neuroimaging. In: IEEE International Symposium on Biomedical Imaging. pp (2011) 2. Fischl, B., Salat, D.H., Busa, E., Albert, M., Dieterich, M., Haselgrove, C., van der Kouwe, A., Killiany, R., Kennedy, D., Klaveness, S., Montillo, A., Makris, N., Rosen, B., Dale, A.M.: Whole brain segmentation: automated labeling of neuroanatomical structures in the human brain. Neuron 33(3), (2002) 3. Gerber, S., Tasdizen, T., Fletcher, P.T., Joshi, S., Whitaker, R.: Manifold modeling for brain population analysis. Medical Image Analysis 14(5), (2010) 4. Geuzaine, C., Remacle, J.F.: Gmsh: A 3-d finite element mesh generator with builtin pre-and post-processing facilities. International Journal for Numerical Methods in Engineering 79(11), (2009) 5. Hamm, J., Ye, D.H., Verma, R., Davatzikos, C.: Gram: A framework for geodesic registration on anatomical manifolds. Med. Image Analysis 14(5), (2010) 6. Lian, Z., Godil, A., Bustos, B., Daoudi, M., Hermans, J., Kawamura, S., Kurita, Y., Lavoué, G., Van Nguyen, H., Ohbuchi, R., et al.: A comparison of methods for non-rigid 3D shape retrieval. Pattern Recognition 46, (2012) 7. Mueller, S.G., Weiner, M.W., Thal, L.J., et al: The alzheimer s disease neuroimaging initiative. Neuroimaging Clinics of North America 15(4), (2005) 8. Niethammer, M., Reuter, M., Wolter, F.E., Bouix, S., Peinecke, N., Koo, M.S., Shenton, M.: Global medical shape analysis using the Laplace-Beltrami spectrum. In: Ayache, N., Ourselin, S., Maeder, A.J. (eds.) MICCAI LNCS, vol. 4791, pp Springer, Heidelberg (2007) 9. Reuter, M., Niethammer, M., Wolter, F.E., Bouix, S., Shenton, M.: Global medical shape analysis using the volumetric Laplace spectrum. In: International Conference on Cyberworlds, NASA-GEM Workshop. pp (2007) 10. Reuter, M., Wolter, F.E., Peinecke, N.: Laplace-Beltrami spectra as Shape-DNA of surfaces and solids. Computer-Aided Design 38(4), (2006) 11. Reuter, M., Wolter, F.E., Shenton, M., Niethammer, M.: Laplace-Beltrami eigenvalues and topological features of eigenfunctions for statistical shape analysis. Computer-Aided Design 41(10), (2009) 12. Schaer, M., Cuadra, M.B., Tamarit, L., Lazeyras, F., Eliez, S., Thiran, J.: A surface-based approach to quantify local cortical gyrification. Medical Imaging, IEEE Transactions on 27(2), (2008)
BrainPrint: Identifying Subjects by Their Brain
 BrainPrint: Identifying Subjects by Their Brain The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation As Published Publisher Wachinger,
BrainPrint: Identifying Subjects by Their Brain The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation As Published Publisher Wachinger,
BrainPrint in the Computer-Aided Diagnosis of Alzheimer s Disease
 BrainPrint in the Computer-Aided Diagnosis of Alzheimer s Disease C. Wachinger 1,2, K. Batmanghelich 1, P. Golland 1, M. Reuter 1,2 1 Computer Science and Artificial Intelligence Lab, MIT 2 Massachusetts
BrainPrint in the Computer-Aided Diagnosis of Alzheimer s Disease C. Wachinger 1,2, K. Batmanghelich 1, P. Golland 1, M. Reuter 1,2 1 Computer Science and Artificial Intelligence Lab, MIT 2 Massachusetts
The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy
 The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy Sokratis K. Makrogiannis, PhD From post-doctoral research at SBIA lab, Department of Radiology,
The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy Sokratis K. Makrogiannis, PhD From post-doctoral research at SBIA lab, Department of Radiology,
The organization of the human cerebral cortex estimated by intrinsic functional connectivity
 1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
SURFACE RECONSTRUCTION OF EX-VIVO HUMAN V1 THROUGH IDENTIFICATION OF THE STRIA OF GENNARI USING MRI AT 7T
 SURFACE RECONSTRUCTION OF EX-VIVO HUMAN V1 THROUGH IDENTIFICATION OF THE STRIA OF GENNARI USING MRI AT 7T Oliver P. Hinds 1, Jonathan R. Polimeni 2, Megan L. Blackwell 3, Christopher J. Wiggins 3, Graham
SURFACE RECONSTRUCTION OF EX-VIVO HUMAN V1 THROUGH IDENTIFICATION OF THE STRIA OF GENNARI USING MRI AT 7T Oliver P. Hinds 1, Jonathan R. Polimeni 2, Megan L. Blackwell 3, Christopher J. Wiggins 3, Graham
Multiple Cortical Surface Correspondence Using Pairwise Shape Similarity
 Multiple Cortical Surface Correspondence Using Pairwise Shape Similarity Pahal Dalal 1, Feng Shi 2, Dinggang Shen 2, and Song Wang 1 1 Department of Computer Science and Engineering, University of South
Multiple Cortical Surface Correspondence Using Pairwise Shape Similarity Pahal Dalal 1, Feng Shi 2, Dinggang Shen 2, and Song Wang 1 1 Department of Computer Science and Engineering, University of South
Quantitative MRI of the Brain: Investigation of Cerebral Gray and White Matter Diseases
 Quantities Measured by MR - Quantitative MRI of the Brain: Investigation of Cerebral Gray and White Matter Diseases Static parameters (influenced by molecular environment): T, T* (transverse relaxation)
Quantities Measured by MR - Quantitative MRI of the Brain: Investigation of Cerebral Gray and White Matter Diseases Static parameters (influenced by molecular environment): T, T* (transverse relaxation)
Regional Manifold Learning for Deformable Registration of Brain MR Images
 Regional Manifold Learning for Deformable Registration of Brain MR Images Dong Hye Ye, Jihun Hamm, Dongjin Kwon, Christos Davatzikos, and Kilian M. Pohl Department of Radiology, University of Pennsylvania,
Regional Manifold Learning for Deformable Registration of Brain MR Images Dong Hye Ye, Jihun Hamm, Dongjin Kwon, Christos Davatzikos, and Kilian M. Pohl Department of Radiology, University of Pennsylvania,
Population Modeling in Neuroscience Using Computer Vision and Machine Learning to learn from Brain Images. Overview. Image Registration
 Population Modeling in Neuroscience Using Computer Vision and Machine Learning to learn from Brain Images Overview 1. Part 1: Theory 1. 2. Learning 2. Part 2: Applications ernst.schwartz@meduniwien.ac.at
Population Modeling in Neuroscience Using Computer Vision and Machine Learning to learn from Brain Images Overview 1. Part 1: Theory 1. 2. Learning 2. Part 2: Applications ernst.schwartz@meduniwien.ac.at
Automatic Caudate Segmentation by Hybrid Generative/Discriminative Models
 Automatic Caudate Segmentation by Hybrid Generative/Discriminative Models Zhuowen Tu, Mubeena Mirza, Ivo Dinov, and Arthur W. Toga Lab of Neuro Imaging, School of Medicine University of California, Los
Automatic Caudate Segmentation by Hybrid Generative/Discriminative Models Zhuowen Tu, Mubeena Mirza, Ivo Dinov, and Arthur W. Toga Lab of Neuro Imaging, School of Medicine University of California, Los
Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge
 Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge Christian Wasserthal 1, Karin Engel 1, Karsten Rink 1 und André Brechmann
Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge Christian Wasserthal 1, Karin Engel 1, Karsten Rink 1 und André Brechmann
Open Topology: A Toolkit for Brain Isosurface Correction
 Open Topology: A Toolkit for Brain Isosurface Correction Sylvain Jaume 1, Patrice Rondao 2, and Benoît Macq 2 1 National Institute of Research in Computer Science and Control, INRIA, France, sylvain@mit.edu,
Open Topology: A Toolkit for Brain Isosurface Correction Sylvain Jaume 1, Patrice Rondao 2, and Benoît Macq 2 1 National Institute of Research in Computer Science and Control, INRIA, France, sylvain@mit.edu,
Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging
 1 CS 9 Final Project Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging Feiyu Chen Department of Electrical Engineering ABSTRACT Subject motion is a significant
1 CS 9 Final Project Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging Feiyu Chen Department of Electrical Engineering ABSTRACT Subject motion is a significant
Measuring longitudinal brain changes in humans and small animal models. Christos Davatzikos
 Measuring longitudinal brain changes in humans and small animal models Christos Davatzikos Section of Biomedical Image Analysis University of Pennsylvania (Radiology) http://www.rad.upenn.edu/sbia Computational
Measuring longitudinal brain changes in humans and small animal models Christos Davatzikos Section of Biomedical Image Analysis University of Pennsylvania (Radiology) http://www.rad.upenn.edu/sbia Computational
Towards Whole Brain Segmentation by a Hybrid Model
 Towards Whole Brain Segmentation by a Hybrid Model Zhuowen Tu and Arthur W. Toga Lab of Neuro Imaging, School of Medicine University of California, Los Angeles, USA Abstract. Segmenting cortical and sub-cortical
Towards Whole Brain Segmentation by a Hybrid Model Zhuowen Tu and Arthur W. Toga Lab of Neuro Imaging, School of Medicine University of California, Los Angeles, USA Abstract. Segmenting cortical and sub-cortical
Ensemble registration: Combining groupwise registration and segmentation
 PURWANI, COOTES, TWINING: ENSEMBLE REGISTRATION 1 Ensemble registration: Combining groupwise registration and segmentation Sri Purwani 1,2 sri.purwani@postgrad.manchester.ac.uk Tim Cootes 1 t.cootes@manchester.ac.uk
PURWANI, COOTES, TWINING: ENSEMBLE REGISTRATION 1 Ensemble registration: Combining groupwise registration and segmentation Sri Purwani 1,2 sri.purwani@postgrad.manchester.ac.uk Tim Cootes 1 t.cootes@manchester.ac.uk
A combined Surface And VOlumetric Registration (SAVOR) framework to study cortical biomarkers and volumetric imaging data
 A combined Surface And VOlumetric Registration (SAVOR) framework to study cortical biomarkers and volumetric imaging data Eli Gibson 1, Ali R. Khan 1, and Mirza Faisal Beg 1 School of Engineering Science,
A combined Surface And VOlumetric Registration (SAVOR) framework to study cortical biomarkers and volumetric imaging data Eli Gibson 1, Ali R. Khan 1, and Mirza Faisal Beg 1 School of Engineering Science,
1 Introduction Motivation and Aims Functional Imaging Computational Neuroanatomy... 12
 Contents 1 Introduction 10 1.1 Motivation and Aims....... 10 1.1.1 Functional Imaging.... 10 1.1.2 Computational Neuroanatomy... 12 1.2 Overview of Chapters... 14 2 Rigid Body Registration 18 2.1 Introduction.....
Contents 1 Introduction 10 1.1 Motivation and Aims....... 10 1.1.1 Functional Imaging.... 10 1.1.2 Computational Neuroanatomy... 12 1.2 Overview of Chapters... 14 2 Rigid Body Registration 18 2.1 Introduction.....
MULTIVARIATE TEXTURE DISCRIMINATION USING A PRINCIPAL GEODESIC CLASSIFIER
 MULTIVARIATE TEXTURE DISCRIMINATION USING A PRINCIPAL GEODESIC CLASSIFIER A.Shabbir 1, 2 and G.Verdoolaege 1, 3 1 Department of Applied Physics, Ghent University, B-9000 Ghent, Belgium 2 Max Planck Institute
MULTIVARIATE TEXTURE DISCRIMINATION USING A PRINCIPAL GEODESIC CLASSIFIER A.Shabbir 1, 2 and G.Verdoolaege 1, 3 1 Department of Applied Physics, Ghent University, B-9000 Ghent, Belgium 2 Max Planck Institute
Predicting Activation Across Individuals with Resting- State Functional Connectivity Based Multi-Atlas Label Fusion
 Predicting Activation Across Individuals with Resting- State Functional Connectivity Based Multi-Atlas Label Fusion The MIT Faculty has made this article openly available. Please share how this access
Predicting Activation Across Individuals with Resting- State Functional Connectivity Based Multi-Atlas Label Fusion The MIT Faculty has made this article openly available. Please share how this access
UNC 4D Infant Cortical Surface Atlases, from Neonate to 6 Years of Age
 UNC 4D Infant Cortical Surface Atlases, from Neonate to 6 Years of Age Version 1.0 UNC 4D infant cortical surface atlases from neonate to 6 years of age contain 11 time points, including 1, 3, 6, 9, 12,
UNC 4D Infant Cortical Surface Atlases, from Neonate to 6 Years of Age Version 1.0 UNC 4D infant cortical surface atlases from neonate to 6 years of age contain 11 time points, including 1, 3, 6, 9, 12,
Shape-based Diffeomorphic Registration on Hippocampal Surfaces Using Beltrami Holomorphic Flow
 Shape-based Diffeomorphic Registration on Hippocampal Surfaces Using Beltrami Holomorphic Flow Abstract. Finding meaningful 1-1 correspondences between hippocampal (HP) surfaces is an important but difficult
Shape-based Diffeomorphic Registration on Hippocampal Surfaces Using Beltrami Holomorphic Flow Abstract. Finding meaningful 1-1 correspondences between hippocampal (HP) surfaces is an important but difficult
CHAPTER 2. Morphometry on rodent brains. A.E.H. Scheenstra J. Dijkstra L. van der Weerd
 CHAPTER 2 Morphometry on rodent brains A.E.H. Scheenstra J. Dijkstra L. van der Weerd This chapter was adapted from: Volumetry and other quantitative measurements to assess the rodent brain, In vivo NMR
CHAPTER 2 Morphometry on rodent brains A.E.H. Scheenstra J. Dijkstra L. van der Weerd This chapter was adapted from: Volumetry and other quantitative measurements to assess the rodent brain, In vivo NMR
Applications of Elastic Functional and Shape Data Analysis
 Applications of Elastic Functional and Shape Data Analysis Quick Outline: 1. Functional Data 2. Shapes of Curves 3. Shapes of Surfaces BAYESIAN REGISTRATION MODEL Bayesian Model + Riemannian Geometry +
Applications of Elastic Functional and Shape Data Analysis Quick Outline: 1. Functional Data 2. Shapes of Curves 3. Shapes of Surfaces BAYESIAN REGISTRATION MODEL Bayesian Model + Riemannian Geometry +
Correspondence Detection Using Wavelet-Based Attribute Vectors
 Correspondence Detection Using Wavelet-Based Attribute Vectors Zhong Xue, Dinggang Shen, and Christos Davatzikos Section of Biomedical Image Analysis, Department of Radiology University of Pennsylvania,
Correspondence Detection Using Wavelet-Based Attribute Vectors Zhong Xue, Dinggang Shen, and Christos Davatzikos Section of Biomedical Image Analysis, Department of Radiology University of Pennsylvania,
THE identification and delineation of brain structures from
 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 26, NO. 4, APRIL 2007 479 Atlas Renormalization for Improved Brain MR Image Segmentation Across Scanner Platforms Xiao Han and Bruce Fischl* Abstract Atlas-based
IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 26, NO. 4, APRIL 2007 479 Atlas Renormalization for Improved Brain MR Image Segmentation Across Scanner Platforms Xiao Han and Bruce Fischl* Abstract Atlas-based
PBSI: A symmetric probabilistic extension of the Boundary Shift Integral
 PBSI: A symmetric probabilistic extension of the Boundary Shift Integral Christian Ledig 1, Robin Wolz 1, Paul Aljabar 1,2, Jyrki Lötjönen 3, Daniel Rueckert 1 1 Department of Computing, Imperial College
PBSI: A symmetric probabilistic extension of the Boundary Shift Integral Christian Ledig 1, Robin Wolz 1, Paul Aljabar 1,2, Jyrki Lötjönen 3, Daniel Rueckert 1 1 Department of Computing, Imperial College
FreeSurfer: Troubleshooting surfer.nmr.mgh.harvard.edu
 FreeSurfer: Troubleshooting surfer.nmr.mgh.harvard.edu 1 Hard and Soft Failures Categories of errors: Hard & Soft Failures Hard = recon-all quits before it finishes Soft = recon-all finishes but results
FreeSurfer: Troubleshooting surfer.nmr.mgh.harvard.edu 1 Hard and Soft Failures Categories of errors: Hard & Soft Failures Hard = recon-all quits before it finishes Soft = recon-all finishes but results
Mid-Space-Independent Symmetric Data Term for Pairwise Deformable Image Registration
 Mid-Space-Independent Symmetric Data Term for Pairwise Deformable Image Registration Iman Aganj 1, Juan Eugenio Iglesias, Martin Reuter 1,3, Mert Rory Sabuncu 1,3, and Bruce Fischl 1,3 1 Athinoula A. Martinos
Mid-Space-Independent Symmetric Data Term for Pairwise Deformable Image Registration Iman Aganj 1, Juan Eugenio Iglesias, Martin Reuter 1,3, Mert Rory Sabuncu 1,3, and Bruce Fischl 1,3 1 Athinoula A. Martinos
3D Brain Segmentation Using Active Appearance Models and Local Regressors
 3D Brain Segmentation Using Active Appearance Models and Local Regressors K.O. Babalola, T.F. Cootes, C.J. Twining, V. Petrovic, and C.J. Taylor Division of Imaging Science and Biomedical Engineering,
3D Brain Segmentation Using Active Appearance Models and Local Regressors K.O. Babalola, T.F. Cootes, C.J. Twining, V. Petrovic, and C.J. Taylor Division of Imaging Science and Biomedical Engineering,
MARS: Multiple Atlases Robust Segmentation
 Software Release (1.0.1) Last updated: April 30, 2014. MARS: Multiple Atlases Robust Segmentation Guorong Wu, Minjeong Kim, Gerard Sanroma, and Dinggang Shen {grwu, mjkim, gerard_sanroma, dgshen}@med.unc.edu
Software Release (1.0.1) Last updated: April 30, 2014. MARS: Multiple Atlases Robust Segmentation Guorong Wu, Minjeong Kim, Gerard Sanroma, and Dinggang Shen {grwu, mjkim, gerard_sanroma, dgshen}@med.unc.edu
NIH Public Access Author Manuscript Med Image Comput Comput Assist Interv. Author manuscript; available in PMC 2009 December 4.
 NIH Public Access Author Manuscript Med Image Comput Comput Assist Interv. Author manuscript; available in PMC 2009 December 4. Published in final edited form as: Med Image Comput Comput Assist Interv.
NIH Public Access Author Manuscript Med Image Comput Comput Assist Interv. Author manuscript; available in PMC 2009 December 4. Published in final edited form as: Med Image Comput Comput Assist Interv.
Methods for data preprocessing
 Methods for data preprocessing John Ashburner Wellcome Trust Centre for Neuroimaging, 12 Queen Square, London, UK. Overview Voxel-Based Morphometry Morphometry in general Volumetrics VBM preprocessing
Methods for data preprocessing John Ashburner Wellcome Trust Centre for Neuroimaging, 12 Queen Square, London, UK. Overview Voxel-Based Morphometry Morphometry in general Volumetrics VBM preprocessing
Structural MRI analysis
 Structural MRI analysis volumetry and voxel-based morphometry cortical thickness measurements structural covariance network mapping Boris Bernhardt, PhD Department of Social Neuroscience, MPI-CBS bernhardt@cbs.mpg.de
Structural MRI analysis volumetry and voxel-based morphometry cortical thickness measurements structural covariance network mapping Boris Bernhardt, PhD Department of Social Neuroscience, MPI-CBS bernhardt@cbs.mpg.de
Incorporating Non-rigid Registration into Expectation Maximization Algorithm to Segment MR Images
 Incorporating Non-rigid Registration into Expectation Maximization Algorithm to Segment MR Images Kilian M Pohl 1, William M. Wells 2, Alexandre Guimond 3, Kiyoto Kasai 4, Martha E. Shenton 2,4, Ron Kikinis
Incorporating Non-rigid Registration into Expectation Maximization Algorithm to Segment MR Images Kilian M Pohl 1, William M. Wells 2, Alexandre Guimond 3, Kiyoto Kasai 4, Martha E. Shenton 2,4, Ron Kikinis
NIH Public Access Author Manuscript Proc IEEE Int Symp Biomed Imaging. Author manuscript; available in PMC 2014 November 15.
 NIH Public Access Author Manuscript Published in final edited form as: Proc IEEE Int Symp Biomed Imaging. 2013 April ; 2013: 748 751. doi:10.1109/isbi.2013.6556583. BRAIN TUMOR SEGMENTATION WITH SYMMETRIC
NIH Public Access Author Manuscript Published in final edited form as: Proc IEEE Int Symp Biomed Imaging. 2013 April ; 2013: 748 751. doi:10.1109/isbi.2013.6556583. BRAIN TUMOR SEGMENTATION WITH SYMMETRIC
MR IMAGE SEGMENTATION
 MR IMAGE SEGMENTATION Prepared by : Monil Shah What is Segmentation? Partitioning a region or regions of interest in images such that each region corresponds to one or more anatomic structures Classification
MR IMAGE SEGMENTATION Prepared by : Monil Shah What is Segmentation? Partitioning a region or regions of interest in images such that each region corresponds to one or more anatomic structures Classification
Context-sensitive Classification Forests for Segmentation of Brain Tumor Tissues
 Context-sensitive Classification Forests for Segmentation of Brain Tumor Tissues D. Zikic, B. Glocker, E. Konukoglu, J. Shotton, A. Criminisi, D. H. Ye, C. Demiralp 3, O. M. Thomas 4,5, T. Das 4, R. Jena
Context-sensitive Classification Forests for Segmentation of Brain Tumor Tissues D. Zikic, B. Glocker, E. Konukoglu, J. Shotton, A. Criminisi, D. H. Ye, C. Demiralp 3, O. M. Thomas 4,5, T. Das 4, R. Jena
Manifold Learning-based Data Sampling for Model Training
 Manifold Learning-based Data Sampling for Model Training Shuqing Chen 1, Sabrina Dorn 2, Michael Lell 3, Marc Kachelrieß 2,Andreas Maier 1 1 Pattern Recognition Lab, FAU Erlangen-Nürnberg 2 German Cancer
Manifold Learning-based Data Sampling for Model Training Shuqing Chen 1, Sabrina Dorn 2, Michael Lell 3, Marc Kachelrieß 2,Andreas Maier 1 1 Pattern Recognition Lab, FAU Erlangen-Nürnberg 2 German Cancer
Sampling-Based Ensemble Segmentation against Inter-operator Variability
 Sampling-Based Ensemble Segmentation against Inter-operator Variability Jing Huo 1, Kazunori Okada, Whitney Pope 1, Matthew Brown 1 1 Center for Computer vision and Imaging Biomarkers, Department of Radiological
Sampling-Based Ensemble Segmentation against Inter-operator Variability Jing Huo 1, Kazunori Okada, Whitney Pope 1, Matthew Brown 1 1 Center for Computer vision and Imaging Biomarkers, Department of Radiological
Subvoxel Segmentation and Representation of Brain Cortex Using Fuzzy Clustering and Gradient Vector Diffusion
 Subvoxel Segmentation and Representation of Brain Cortex Using Fuzzy Clustering and Gradient Vector Diffusion Ming-Ching Chang Xiaodong Tao GE Global Research Center {changm, taox} @ research.ge.com SPIE
Subvoxel Segmentation and Representation of Brain Cortex Using Fuzzy Clustering and Gradient Vector Diffusion Ming-Ching Chang Xiaodong Tao GE Global Research Center {changm, taox} @ research.ge.com SPIE
Manifold Learning of Brain MRIs by Deep Learning
 Manifold Learning of Brain MRIs by Deep Learning Tom Brosch 1,3 and Roger Tam 2,3 for the Alzheimer s Disease Neuroimaging Initiative 1 Department of Electrical and Computer Engineering 2 Department of
Manifold Learning of Brain MRIs by Deep Learning Tom Brosch 1,3 and Roger Tam 2,3 for the Alzheimer s Disease Neuroimaging Initiative 1 Department of Electrical and Computer Engineering 2 Department of
Automated MR Image Analysis Pipelines
 Automated MR Image Analysis Pipelines Andy Simmons Centre for Neuroimaging Sciences, Kings College London Institute of Psychiatry. NIHR Biomedical Research Centre for Mental Health at IoP & SLAM. Neuroimaging
Automated MR Image Analysis Pipelines Andy Simmons Centre for Neuroimaging Sciences, Kings College London Institute of Psychiatry. NIHR Biomedical Research Centre for Mental Health at IoP & SLAM. Neuroimaging
CS/NEUR125 Brains, Minds, and Machines. Due: Wednesday, April 5
 CS/NEUR125 Brains, Minds, and Machines Lab 8: Using fmri to Discover Language Areas in the Brain Due: Wednesday, April 5 In this lab, you will analyze fmri data from an experiment that was designed to
CS/NEUR125 Brains, Minds, and Machines Lab 8: Using fmri to Discover Language Areas in the Brain Due: Wednesday, April 5 In this lab, you will analyze fmri data from an experiment that was designed to
Introduc)on to FreeSurfer h0p://surfer.nmr.mgh.harvard.edu. Jenni Pacheco.
 Introduc)on to FreeSurfer h0p://surfer.nmr.mgh.harvard.edu Jenni Pacheco jpacheco@mail.utexas.edu Outline FreeSurfer ROI Terminology ROI Sta)s)cs Files ROI Studies Volumetric/Area Intensity FreeSurfer
Introduc)on to FreeSurfer h0p://surfer.nmr.mgh.harvard.edu Jenni Pacheco jpacheco@mail.utexas.edu Outline FreeSurfer ROI Terminology ROI Sta)s)cs Files ROI Studies Volumetric/Area Intensity FreeSurfer
Pathology Hinting as the Combination of Automatic Segmentation with a Statistical Shape Model
 Pathology Hinting as the Combination of Automatic Segmentation with a Statistical Shape Model Pascal A. Dufour 12,HannanAbdillahi 3, Lala Ceklic 3,Ute Wolf-Schnurrbusch 23,JensKowal 12 1 ARTORG Center
Pathology Hinting as the Combination of Automatic Segmentation with a Statistical Shape Model Pascal A. Dufour 12,HannanAbdillahi 3, Lala Ceklic 3,Ute Wolf-Schnurrbusch 23,JensKowal 12 1 ARTORG Center
Norbert Schuff VA Medical Center and UCSF
 Norbert Schuff Medical Center and UCSF Norbert.schuff@ucsf.edu Medical Imaging Informatics N.Schuff Course # 170.03 Slide 1/67 Objective Learn the principle segmentation techniques Understand the role
Norbert Schuff Medical Center and UCSF Norbert.schuff@ucsf.edu Medical Imaging Informatics N.Schuff Course # 170.03 Slide 1/67 Objective Learn the principle segmentation techniques Understand the role
A Model-Independent, Multi-Image Approach to MR Inhomogeneity Correction
 Tina Memo No. 2007-003 Published in Proc. MIUA 2007 A Model-Independent, Multi-Image Approach to MR Inhomogeneity Correction P. A. Bromiley and N.A. Thacker Last updated 13 / 4 / 2007 Imaging Science and
Tina Memo No. 2007-003 Published in Proc. MIUA 2007 A Model-Independent, Multi-Image Approach to MR Inhomogeneity Correction P. A. Bromiley and N.A. Thacker Last updated 13 / 4 / 2007 Imaging Science and
Fully Automatic Multi-organ Segmentation based on Multi-boost Learning and Statistical Shape Model Search
 Fully Automatic Multi-organ Segmentation based on Multi-boost Learning and Statistical Shape Model Search Baochun He, Cheng Huang, Fucang Jia Shenzhen Institutes of Advanced Technology, Chinese Academy
Fully Automatic Multi-organ Segmentation based on Multi-boost Learning and Statistical Shape Model Search Baochun He, Cheng Huang, Fucang Jia Shenzhen Institutes of Advanced Technology, Chinese Academy
Attribute Similarity and Mutual-Saliency Weighting for Registration and Label Fusion
 Attribute Similarity and Mutual-Saliency Weighting for Registration and Label Fusion Yangming Ou, Jimit Doshi, Guray Erus, and Christos Davatzikos Section of Biomedical Image Analysis (SBIA) Department
Attribute Similarity and Mutual-Saliency Weighting for Registration and Label Fusion Yangming Ou, Jimit Doshi, Guray Erus, and Christos Davatzikos Section of Biomedical Image Analysis (SBIA) Department
Attention modulates spatial priority maps in human occipital, parietal, and frontal cortex
 Attention modulates spatial priority maps in human occipital, parietal, and frontal cortex Thomas C. Sprague 1 and John T. Serences 1,2 1 Neuroscience Graduate Program, University of California San Diego
Attention modulates spatial priority maps in human occipital, parietal, and frontal cortex Thomas C. Sprague 1 and John T. Serences 1,2 1 Neuroscience Graduate Program, University of California San Diego
Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques
 Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques Sea Chen Department of Biomedical Engineering Advisors: Dr. Charles A. Bouman and Dr. Mark J. Lowe S. Chen Final Exam October
Analysis of Functional MRI Timeseries Data Using Signal Processing Techniques Sea Chen Department of Biomedical Engineering Advisors: Dr. Charles A. Bouman and Dr. Mark J. Lowe S. Chen Final Exam October
Anatomical landmark and region mapping based on a template surface deformation for foot bone morphology
 Anatomical landmark and region mapping based on a template surface deformation for foot bone morphology Jaeil Kim 1, Sang Gyo Seo 2, Dong Yeon Lee 2, Jinah Park 1 1 Department of Computer Science, KAIST,
Anatomical landmark and region mapping based on a template surface deformation for foot bone morphology Jaeil Kim 1, Sang Gyo Seo 2, Dong Yeon Lee 2, Jinah Park 1 1 Department of Computer Science, KAIST,
K-Means Clustering Using Localized Histogram Analysis
 K-Means Clustering Using Localized Histogram Analysis Michael Bryson University of South Carolina, Department of Computer Science Columbia, SC brysonm@cse.sc.edu Abstract. The first step required for many
K-Means Clustering Using Localized Histogram Analysis Michael Bryson University of South Carolina, Department of Computer Science Columbia, SC brysonm@cse.sc.edu Abstract. The first step required for many
Cortical Surface Shape Analysis Based on Spherical Wavelet Transformation
 Cortical Surface Shape Analysis Based on Spherical Wavelet ransformation Peng Yu 1, Xiao Han 2, Florent Ségonne 3, 4, Rudolph Pienaar 3, Randy L. Buckner 3, 5, 6 Polina Golland 4, P. Ellen Grant 3, 7 1,
Cortical Surface Shape Analysis Based on Spherical Wavelet ransformation Peng Yu 1, Xiao Han 2, Florent Ségonne 3, 4, Rudolph Pienaar 3, Randy L. Buckner 3, 5, 6 Polina Golland 4, P. Ellen Grant 3, 7 1,
Automatic Generation of Training Data for Brain Tissue Classification from MRI
 Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. COCOSCO, Alex P. ZIJDENBOS, and Alan C. EVANS http://www.bic.mni.mcgill.ca/users/crisco/ McConnell Brain Imaging
Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. COCOSCO, Alex P. ZIJDENBOS, and Alan C. EVANS http://www.bic.mni.mcgill.ca/users/crisco/ McConnell Brain Imaging
Automatic Generation of Training Data for Brain Tissue Classification from MRI
 MICCAI-2002 1 Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. Cocosco, Alex P. Zijdenbos, and Alan C. Evans McConnell Brain Imaging Centre, Montreal Neurological
MICCAI-2002 1 Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. Cocosco, Alex P. Zijdenbos, and Alan C. Evans McConnell Brain Imaging Centre, Montreal Neurological
Automatic Subcortical Segmentation Using a Contextual Model
 Automatic Subcortical Segmentation Using a Contextual Model Jonathan H. Morra 1, Zhuowen Tu 1, Liana G. Apostolova 1,2, Amity E. Green 1,2, Arthur W. Toga 1, and Paul M. Thompson 1 1 Laboratory of Neuro
Automatic Subcortical Segmentation Using a Contextual Model Jonathan H. Morra 1, Zhuowen Tu 1, Liana G. Apostolova 1,2, Amity E. Green 1,2, Arthur W. Toga 1, and Paul M. Thompson 1 1 Laboratory of Neuro
Statistical Shape Analysis of Anatomical Structures. Polina Golland
 Statistical Shape Analysis of Anatomical Structures by Polina Golland B.A., Technion, Israel (1993) M.Sc., Technion, Israel (1995) Submitted to the Department of Electrical Engineering and Computer Science
Statistical Shape Analysis of Anatomical Structures by Polina Golland B.A., Technion, Israel (1993) M.Sc., Technion, Israel (1995) Submitted to the Department of Electrical Engineering and Computer Science
Pathology Hinting as the Combination of Automatic Segmentation with a Statistical Shape Model
 Pathology Hinting as the Combination of Automatic Segmentation with a Statistical Shape Model Pascal A. Dufour 1,2, Hannan Abdillahi 3, Lala Ceklic 3, Ute Wolf-Schnurrbusch 2,3, and Jens Kowal 1,2 1 ARTORG
Pathology Hinting as the Combination of Automatic Segmentation with a Statistical Shape Model Pascal A. Dufour 1,2, Hannan Abdillahi 3, Lala Ceklic 3, Ute Wolf-Schnurrbusch 2,3, and Jens Kowal 1,2 1 ARTORG
The Allen Human Brain Atlas offers three types of searches to allow a user to: (1) obtain gene expression data for specific genes (or probes) of
 Microarray Data MICROARRAY DATA Gene Search Boolean Syntax Differential Search Mouse Differential Search Search Results Gene Classification Correlative Search Download Search Results Data Visualization
Microarray Data MICROARRAY DATA Gene Search Boolean Syntax Differential Search Mouse Differential Search Search Results Gene Classification Correlative Search Download Search Results Data Visualization
GENERAL AUTOMATED FLAW DETECTION SCHEME FOR NDE X-RAY IMAGES
 GENERAL AUTOMATED FLAW DETECTION SCHEME FOR NDE X-RAY IMAGES Karl W. Ulmer and John P. Basart Center for Nondestructive Evaluation Department of Electrical and Computer Engineering Iowa State University
GENERAL AUTOMATED FLAW DETECTION SCHEME FOR NDE X-RAY IMAGES Karl W. Ulmer and John P. Basart Center for Nondestructive Evaluation Department of Electrical and Computer Engineering Iowa State University
Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm
 Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm Lin Chen and Gudrun Wagenknecht Central Institute for Electronics, Research Center Jülich, Jülich, Germany Email: l.chen@fz-juelich.de
Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm Lin Chen and Gudrun Wagenknecht Central Institute for Electronics, Research Center Jülich, Jülich, Germany Email: l.chen@fz-juelich.de
Learning-based Neuroimage Registration
 Learning-based Neuroimage Registration Leonid Teverovskiy and Yanxi Liu 1 October 2004 CMU-CALD-04-108, CMU-RI-TR-04-59 School of Computer Science Carnegie Mellon University Pittsburgh, PA 15213 Abstract
Learning-based Neuroimage Registration Leonid Teverovskiy and Yanxi Liu 1 October 2004 CMU-CALD-04-108, CMU-RI-TR-04-59 School of Computer Science Carnegie Mellon University Pittsburgh, PA 15213 Abstract
2 Michael E. Leventon and Sarah F. F. Gibson a b c d Fig. 1. (a, b) Two MR scans of a person's knee. Both images have high resolution in-plane, but ha
 Model Generation from Multiple Volumes using Constrained Elastic SurfaceNets Michael E. Leventon and Sarah F. F. Gibson 1 MIT Artificial Intelligence Laboratory, Cambridge, MA 02139, USA leventon@ai.mit.edu
Model Generation from Multiple Volumes using Constrained Elastic SurfaceNets Michael E. Leventon and Sarah F. F. Gibson 1 MIT Artificial Intelligence Laboratory, Cambridge, MA 02139, USA leventon@ai.mit.edu
Spatio-Temporal Registration of Biomedical Images by Computational Methods
 Spatio-Temporal Registration of Biomedical Images by Computational Methods Francisco P. M. Oliveira, João Manuel R. S. Tavares tavares@fe.up.pt, www.fe.up.pt/~tavares Outline 1. Introduction 2. Spatial
Spatio-Temporal Registration of Biomedical Images by Computational Methods Francisco P. M. Oliveira, João Manuel R. S. Tavares tavares@fe.up.pt, www.fe.up.pt/~tavares Outline 1. Introduction 2. Spatial
ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION
 ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION Abstract: MIP Project Report Spring 2013 Gaurav Mittal 201232644 This is a detailed report about the course project, which was to implement
ADAPTIVE GRAPH CUTS WITH TISSUE PRIORS FOR BRAIN MRI SEGMENTATION Abstract: MIP Project Report Spring 2013 Gaurav Mittal 201232644 This is a detailed report about the course project, which was to implement
HHS Public Access Author manuscript IEEE Trans Med Imaging. Author manuscript; available in PMC 2015 June 14.
 Cortical Surface Shape Analysis Based on Spherical Wavelet Transformation Peng Yu 1, Xiao Han 2, Florent Ségonne 3,4, Rudolph Pienaar 3, Randy L. Buckner 3,5,6, Polina Golland 4, P. Ellen Grant 3,7, and
Cortical Surface Shape Analysis Based on Spherical Wavelet Transformation Peng Yu 1, Xiao Han 2, Florent Ségonne 3,4, Rudolph Pienaar 3, Randy L. Buckner 3,5,6, Polina Golland 4, P. Ellen Grant 3,7, and
Lilla Zöllei A.A. Martinos Center, MGH; Boston, MA
 Lilla Zöllei lzollei@nmr.mgh.harvard.edu A.A. Martinos Center, MGH; Boston, MA Bruce Fischl Gheorghe Postelnicu Jean Augustinack Anastasia Yendiki Allison Stevens Kristen Huber Sita Kakonoori + the FreeSurfer
Lilla Zöllei lzollei@nmr.mgh.harvard.edu A.A. Martinos Center, MGH; Boston, MA Bruce Fischl Gheorghe Postelnicu Jean Augustinack Anastasia Yendiki Allison Stevens Kristen Huber Sita Kakonoori + the FreeSurfer
Performance Issues in Shape Classification
 Performance Issues in Shape Classification Samson J. Timoner 1, Pollina Golland 1, Ron Kikinis 2, Martha E. Shenton 3, W. Eric L. Grimson 1, and William M. Wells III 1,2 1 MIT AI Laboratory, Cambridge
Performance Issues in Shape Classification Samson J. Timoner 1, Pollina Golland 1, Ron Kikinis 2, Martha E. Shenton 3, W. Eric L. Grimson 1, and William M. Wells III 1,2 1 MIT AI Laboratory, Cambridge
A Multiple-Layer Flexible Mesh Template Matching Method for Nonrigid Registration between a Pelvis Model and CT Images
 A Multiple-Layer Flexible Mesh Template Matching Method for Nonrigid Registration between a Pelvis Model and CT Images Jianhua Yao 1, Russell Taylor 2 1. Diagnostic Radiology Department, Clinical Center,
A Multiple-Layer Flexible Mesh Template Matching Method for Nonrigid Registration between a Pelvis Model and CT Images Jianhua Yao 1, Russell Taylor 2 1. Diagnostic Radiology Department, Clinical Center,
A COMPARISON OF WAVELET-BASED AND RIDGELET- BASED TEXTURE CLASSIFICATION OF TISSUES IN COMPUTED TOMOGRAPHY
 A COMPARISON OF WAVELET-BASED AND RIDGELET- BASED TEXTURE CLASSIFICATION OF TISSUES IN COMPUTED TOMOGRAPHY Lindsay Semler Lucia Dettori Intelligent Multimedia Processing Laboratory School of Computer Scienve,
A COMPARISON OF WAVELET-BASED AND RIDGELET- BASED TEXTURE CLASSIFICATION OF TISSUES IN COMPUTED TOMOGRAPHY Lindsay Semler Lucia Dettori Intelligent Multimedia Processing Laboratory School of Computer Scienve,
EVIDENCE suggests that morphological changes of neuroanatomical
 582 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 26, NO. 4, APRIL 2007 Cortical Surface Shape Analysis Based on Spherical Wavelets Peng Yu, Student Member, IEEE, P. Ellen Grant, Yuan Qi, Xiao Han, Florent
582 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 26, NO. 4, APRIL 2007 Cortical Surface Shape Analysis Based on Spherical Wavelets Peng Yu, Student Member, IEEE, P. Ellen Grant, Yuan Qi, Xiao Han, Florent
Surface-based Analysis: Inter-subject Registration and Smoothing
 Surface-based Analysis: Inter-subject Registration and Smoothing Outline Exploratory Spatial Analysis Coordinate Systems 3D (Volumetric) 2D (Surface-based) Inter-subject registration Volume-based Surface-based
Surface-based Analysis: Inter-subject Registration and Smoothing Outline Exploratory Spatial Analysis Coordinate Systems 3D (Volumetric) 2D (Surface-based) Inter-subject registration Volume-based Surface-based
GLIRT: Groupwise and Longitudinal Image Registration Toolbox
 Software Release (1.0.1) Last updated: March. 30, 2011. GLIRT: Groupwise and Longitudinal Image Registration Toolbox Guorong Wu 1, Qian Wang 1,2, Hongjun Jia 1, and Dinggang Shen 1 1 Image Display, Enhancement,
Software Release (1.0.1) Last updated: March. 30, 2011. GLIRT: Groupwise and Longitudinal Image Registration Toolbox Guorong Wu 1, Qian Wang 1,2, Hongjun Jia 1, and Dinggang Shen 1 1 Image Display, Enhancement,
Recognizing Deformable Shapes. Salvador Ruiz Correa UW Ph.D. Graduate Researcher at Children s Hospital
 Recognizing Deformable Shapes Salvador Ruiz Correa UW Ph.D. Graduate Researcher at Children s Hospital Input 3-D Object Goal We are interested in developing algorithms for recognizing and classifying deformable
Recognizing Deformable Shapes Salvador Ruiz Correa UW Ph.D. Graduate Researcher at Children s Hospital Input 3-D Object Goal We are interested in developing algorithms for recognizing and classifying deformable
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit
 Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
GPU-Based Implementation of a Computational Model of Cerebral Cortex Folding
 GPU-Based Implementation of a Computational Model of Cerebral Cortex Folding Jingxin Nie 1, Kaiming 1,2, Gang Li 1, Lei Guo 1, Tianming Liu 2 1 School of Automation, Northwestern Polytechnical University,
GPU-Based Implementation of a Computational Model of Cerebral Cortex Folding Jingxin Nie 1, Kaiming 1,2, Gang Li 1, Lei Guo 1, Tianming Liu 2 1 School of Automation, Northwestern Polytechnical University,
Computational Methods in NeuroImage Analysis!
 Computational Methods in NeuroImage Analysis! Instructor: Moo K. Chung" mkchung@wisc.edu" Lecture 8" Geometric computation" October 29, 2010" NOTICE! Final Exam: December 3 9:00-12:00am (35%)" Topics:
Computational Methods in NeuroImage Analysis! Instructor: Moo K. Chung" mkchung@wisc.edu" Lecture 8" Geometric computation" October 29, 2010" NOTICE! Final Exam: December 3 9:00-12:00am (35%)" Topics:
Analyzing Anatomical Structures: Leveraging Multiple Sources of Knowledge
 Analyzing Anatomical Structures: Leveraging Multiple Sources of Knowledge Eric Grimson and Polina Golland Computer Science and Artificial Intelligence Laboratory, Massachusetts Institute of Technology,
Analyzing Anatomical Structures: Leveraging Multiple Sources of Knowledge Eric Grimson and Polina Golland Computer Science and Artificial Intelligence Laboratory, Massachusetts Institute of Technology,
Topology-Invariant Similarity and Diffusion Geometry
 1 Topology-Invariant Similarity and Diffusion Geometry Lecture 7 Alexander & Michael Bronstein tosca.cs.technion.ac.il/book Numerical geometry of non-rigid shapes Stanford University, Winter 2009 Intrinsic
1 Topology-Invariant Similarity and Diffusion Geometry Lecture 7 Alexander & Michael Bronstein tosca.cs.technion.ac.il/book Numerical geometry of non-rigid shapes Stanford University, Winter 2009 Intrinsic
Robustness of Selective Desensitization Perceptron Against Irrelevant and Partially Relevant Features in Pattern Classification
 Robustness of Selective Desensitization Perceptron Against Irrelevant and Partially Relevant Features in Pattern Classification Tomohiro Tanno, Kazumasa Horie, Jun Izawa, and Masahiko Morita University
Robustness of Selective Desensitization Perceptron Against Irrelevant and Partially Relevant Features in Pattern Classification Tomohiro Tanno, Kazumasa Horie, Jun Izawa, and Masahiko Morita University
Simultaneous Model-based Segmentation of Multiple Objects
 Simultaneous Model-based Segmentation of Multiple Objects Astrid Franz 1, Robin Wolz 1, Tobias Klinder 1,2, Cristian Lorenz 1, Hans Barschdorf 1, Thomas Blaffert 1, Sebastian P. M. Dries 1, Steffen Renisch
Simultaneous Model-based Segmentation of Multiple Objects Astrid Franz 1, Robin Wolz 1, Tobias Klinder 1,2, Cristian Lorenz 1, Hans Barschdorf 1, Thomas Blaffert 1, Sebastian P. M. Dries 1, Steffen Renisch
Multi-Atlas Segmentation of the Cardiac MR Right Ventricle
 Multi-Atlas Segmentation of the Cardiac MR Right Ventricle Yangming Ou, Jimit Doshi, Guray Erus, and Christos Davatzikos Section of Biomedical Image Analysis (SBIA) Department of Radiology, University
Multi-Atlas Segmentation of the Cardiac MR Right Ventricle Yangming Ou, Jimit Doshi, Guray Erus, and Christos Davatzikos Section of Biomedical Image Analysis (SBIA) Department of Radiology, University
Reliability in multi-site structural MRI studies: Effects of gradient non-linearity correction on phantom and human data
 www.elsevier.com/locate/ynimg NeuroImage 30 (2006) 436 443 Reliability in multi-site structural MRI studies: Effects of gradient non-linearity correction on phantom and human data Jorge Jovicich, a, *
www.elsevier.com/locate/ynimg NeuroImage 30 (2006) 436 443 Reliability in multi-site structural MRI studies: Effects of gradient non-linearity correction on phantom and human data Jorge Jovicich, a, *
VOLUMETRIC HARMONIC MAP
 COMMUNICATIONS IN INFORMATION AND SYSTEMS c 2004 International Press Vol. 3, No. 3, pp. 191-202, March 2004 004 VOLUMETRIC HARMONIC MAP YALIN WANG, XIANFENG GU, AND SHING-TUNG YAU Abstract. We develop
COMMUNICATIONS IN INFORMATION AND SYSTEMS c 2004 International Press Vol. 3, No. 3, pp. 191-202, March 2004 004 VOLUMETRIC HARMONIC MAP YALIN WANG, XIANFENG GU, AND SHING-TUNG YAU Abstract. We develop
An Introduction To Automatic Tissue Classification Of Brain MRI. Colm Elliott Mar 2014
 An Introduction To Automatic Tissue Classification Of Brain MRI Colm Elliott Mar 2014 Tissue Classification Tissue classification is part of many processing pipelines. We often want to classify each voxel
An Introduction To Automatic Tissue Classification Of Brain MRI Colm Elliott Mar 2014 Tissue Classification Tissue classification is part of many processing pipelines. We often want to classify each voxel
Brain Surface Conformal Spherical Mapping
 Brain Surface Conformal Spherical Mapping Min Zhang Department of Industrial Engineering, Arizona State University mzhang33@asu.edu Abstract It is well known and proved that any genus zero surface can
Brain Surface Conformal Spherical Mapping Min Zhang Department of Industrial Engineering, Arizona State University mzhang33@asu.edu Abstract It is well known and proved that any genus zero surface can
Mapping Multi-Modal Routine Imaging Data to a Single Reference via Multiple Templates
 Mapping Multi-Modal Routine Imaging Data to a Single Reference via Multiple Templates Johannes Hofmanninger 1, Bjoern Menze 2, Marc-André Weber 3,4 and Georg Langs 1 1 Department of Biomedical imaging
Mapping Multi-Modal Routine Imaging Data to a Single Reference via Multiple Templates Johannes Hofmanninger 1, Bjoern Menze 2, Marc-André Weber 3,4 and Georg Langs 1 1 Department of Biomedical imaging
H. Mirzaalian, L. Ning, P. Savadjiev, O. Pasternak, S. Bouix, O. Michailovich, M. Kubicki, C-F Westin, M.E.Shenton, Y. Rathi
 Harmonizing diffusion MRI data from multiple scanners H. Mirzaalian, L. Ning, P. Savadjiev, O. Pasternak, S. Bouix, O. Michailovich, M. Kubicki, C-F Westin, M.E.Shenton, Y. Rathi Synopsis Diffusion MRI
Harmonizing diffusion MRI data from multiple scanners H. Mirzaalian, L. Ning, P. Savadjiev, O. Pasternak, S. Bouix, O. Michailovich, M. Kubicki, C-F Westin, M.E.Shenton, Y. Rathi Synopsis Diffusion MRI
CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS
 CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS CHAPTER 4 CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS 4.1 Introduction Optical character recognition is one of
CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS CHAPTER 4 CLASSIFICATION WITH RADIAL BASIS AND PROBABILISTIC NEURAL NETWORKS 4.1 Introduction Optical character recognition is one of
Manifold Learning: Applications in Neuroimaging
 Your own logo here Manifold Learning: Applications in Neuroimaging Robin Wolz 23/09/2011 Overview Manifold learning for Atlas Propagation Multi-atlas segmentation Challenges LEAP Manifold learning for
Your own logo here Manifold Learning: Applications in Neuroimaging Robin Wolz 23/09/2011 Overview Manifold learning for Atlas Propagation Multi-atlas segmentation Challenges LEAP Manifold learning for
3D Volume Mesh Generation of Human Organs Using Surface Geometries Created from the Visible Human Data Set
 3D Volume Mesh Generation of Human Organs Using Surface Geometries Created from the Visible Human Data Set John M. Sullivan, Jr., Ziji Wu, and Anand Kulkarni Worcester Polytechnic Institute Worcester,
3D Volume Mesh Generation of Human Organs Using Surface Geometries Created from the Visible Human Data Set John M. Sullivan, Jr., Ziji Wu, and Anand Kulkarni Worcester Polytechnic Institute Worcester,
Deformable Registration of Cortical Structures via Hybrid Volumetric and Surface Warping
 Deformable Registration of Cortical Structures via Hybrid Volumetric and Surface Warping Tianming Liu, Dinggang Shen, and Christos Davatzikos Section of Biomedical Image Analysis, Department of Radiology,
Deformable Registration of Cortical Structures via Hybrid Volumetric and Surface Warping Tianming Liu, Dinggang Shen, and Christos Davatzikos Section of Biomedical Image Analysis, Department of Radiology,
Structural Segmentation
 Structural Segmentation FAST tissue-type segmentation FIRST sub-cortical structure segmentation FSL-VBM voxelwise grey-matter density analysis SIENA atrophy analysis FAST FMRIB s Automated Segmentation
Structural Segmentation FAST tissue-type segmentation FIRST sub-cortical structure segmentation FSL-VBM voxelwise grey-matter density analysis SIENA atrophy analysis FAST FMRIB s Automated Segmentation
White Matter Lesion Segmentation (WMLS) Manual
 White Matter Lesion Segmentation (WMLS) Manual 1. Introduction White matter lesions (WMLs) are brain abnormalities that appear in different brain diseases, such as multiple sclerosis (MS), head injury,
White Matter Lesion Segmentation (WMLS) Manual 1. Introduction White matter lesions (WMLs) are brain abnormalities that appear in different brain diseases, such as multiple sclerosis (MS), head injury,
Deformetrica: a software for statistical analysis of anatomical shapes
 Deformetrica: a software for statistical analysis of anatomical shapes Alexandre Routier, Marcel Prastawa, Benjamin Charlier, Cédric Doucet, Joan Alexis Glaunès, Stanley Durrleman To cite this version:
Deformetrica: a software for statistical analysis of anatomical shapes Alexandre Routier, Marcel Prastawa, Benjamin Charlier, Cédric Doucet, Joan Alexis Glaunès, Stanley Durrleman To cite this version:
A Generative Model for Image Segmentation Based on Label Fusion Mert R. Sabuncu*, B. T. Thomas Yeo, Koen Van Leemput, Bruce Fischl, and Polina Golland
 1714 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 29, NO. 10, OCTOBER 2010 A Generative Model for Image Segmentation Based on Label Fusion Mert R. Sabuncu*, B. T. Thomas Yeo, Koen Van Leemput, Bruce Fischl,
1714 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 29, NO. 10, OCTOBER 2010 A Generative Model for Image Segmentation Based on Label Fusion Mert R. Sabuncu*, B. T. Thomas Yeo, Koen Van Leemput, Bruce Fischl,
Conference Biomedical Engineering
 Automatic Medical Image Analysis for Measuring Bone Thickness and Density M. Kovalovs *, A. Glazs Image Processing and Computer Graphics Department, Riga Technical University, Latvia * E-mail: mihails.kovalovs@rtu.lv
Automatic Medical Image Analysis for Measuring Bone Thickness and Density M. Kovalovs *, A. Glazs Image Processing and Computer Graphics Department, Riga Technical University, Latvia * E-mail: mihails.kovalovs@rtu.lv
Chapter 4. Clustering Core Atoms by Location
 Chapter 4. Clustering Core Atoms by Location In this chapter, a process for sampling core atoms in space is developed, so that the analytic techniques in section 3C can be applied to local collections
Chapter 4. Clustering Core Atoms by Location In this chapter, a process for sampling core atoms in space is developed, so that the analytic techniques in section 3C can be applied to local collections
