Finding Hidden Patterns in DNA. What makes searching for frequent subsequences hard? Allowing for errors? All the places they could be hiding?
|
|
|
- Ashley Elliott
- 5 years ago
- Views:
Transcription
1 Finding Hidden Patterns in DNA What makes searching for frequent subsequences hard? Allowing for errors? All the places they could be hiding? 1
2 Initiating Transcription As a precursor to transcription (the reading of DNA to construct RNAs, that eventually leading to protein synthesis) special proteins bind to the DNA, and separate it to enable its reading. How do these proteins know where the coding genes are in order to bind? Genes are relatively rare O(1,000,000,000) bases/genome O(10000) genes/genome O(1000) bases/gene Approximately 1% of DNA codes for genes (10 10 /10 ) 2
3 Regulatory Regions RNA polymerases seek out regulatory or promoting regions located bp upstream from the coding region They work in conjunction with special proteins called transcription factors (TFs) whose presence enables gene expression Within these regions are the Transcription Factor Binding Sites (TFBS), special DNA sequence patterns known as motifs that are specific to a given transcription factor A Single TF can influence the expression of many genes. Through biological experiments one can infer, at least a subset of these affected genes. 3
4 Transcription Factor Binding Sites A TFBS can be located anywhere within the regulatory region. TFBS may vary slightly across different regulatory regions since non-essential bases could mutate Transcription factors are robust (they will still bind) in the presence of small sequence differences by a few bases 4
5 Identifying Motifs: Complications We don t know the motif sequence for every TF We don t know where it is located relative to a gene s start Moreover, motifs can differ slightly from one gene to the next We only know that it occurs somewhere near genes that share a TF How to discern a Motif s frequent similar pattern from random patterns? How is this problem different that finding frequent k-mers from Lecture 2? 5
6 Let's look for an Easy Motif 1 tagtggtcttttgagtgtagatccgaagggaaagtatttccaccagttcggggtcacccagcagggcagggtgacttaat 2 cgcgactcggcgctcacagttatcgcacgtttagaccaaaacggagttagatccgaaactggagtttaatcggagtcctt 3 gttacttgtgagcctggttagatccgaaatataattgttggctgcatagcggagctgacatacgagtaggggaaatgcgt 4 aacatcaggctttgattaaacaatttaagcacgtagatccgaattgacctgatgacaatacggaacatgccggctccggg 5 accaccggataggctgcttattagatccgaaaggtagtatcgtaataatggctcagccatgtcaatgtgcggcattccac 6 tagatccgaatcgatcgtgtttctccctctgtgggttaacgaggggtccgaccttgctcgcatgtgccgaacttgtaccc 7 gaaatggttcggtgcgatatcaggccgttctcttaacttggcggtgtagatccgaacgtctctggaggggtcgtgcgcta 8 atgtatactagacattctaacgctcgcttattggcggagaccatttgctccactacaagaggctactgtgtagatccgaa 9 ttcttacacccttctttagatccgaacctgttggcgccatcttcttttcgagtccttgtacctccatttgctctgatgac 10 ctacctatgtaaaacaacatctactaacgtagtccggtctttcctgatctgccctaacctacaggtagatccgaaattcg Problem: Given M sequences of length N find any k-mer that appears in each sequence. How would you go about finding a 10-mer that appears in every one of these strings? 6
7 Sneak Peek at the Answer 1 tagtggtcttttgagtgtagatccgaagggaaagtatttccaccagttcggggtcacccagcagggcagggtgacttaat 2 cgcgactcggcgctcacagttatcgcacgtttagaccaaaacggagttagatccgaaactggagtttaatcggagtcctt 3 gttacttgtgagcctggttagatccgaaatataattgttggctgcatagcggagctgacatacgagtaggggaaatgcgt 4 aacatcaggctttgattaaacaatttaagcacgtagatccgaattgacctgatgacaatacggaacatgccggctccggg 5 accaccggataggctgcttattagatccgaaaggtagtatcgtaataatggctcagccatgtcaatgtgcggcattccac 6 TAGATCCGAAtcgatcgtgtttctccctctgtgggttaacgaggggtccgaccttgctcgcatgtgccgaacttgtaccc 7 gaaatggttcggtgcgatatcaggccgttctcttaacttggcggtgtagatccgaacgtctctggaggggtcgtgcgcta 8 atgtatactagacattctaacgctcgcttattggcggagaccatttgctccactacaagaggctactgtgtagatccgaa 9 ttcttacacccttctttagatccgaacctgttggcgccatcttcttttcgagtccttgtacctccatttgctctgatgac 10 ctacctatgtaaaacaacatctactaacgtagtccggtctttcctgatctgccctaacctacaggtagatccgaaattcg Now that you've seen the answer, how would you find it? 7
8 Meet Mr Brute Force He's often the best starting point when approaching a problem He'll also serve as a straw-man when designing new approaches Though he's seldom elegant, he gets the job done Often, we can't afford to wait for him For our current problem a brute force solution would consider every k-mer position in all strings and see if they match. Given M sequences of length N, there are: position combinations to consider. (N k + 1) M How do you write M nested loops when M is a variable? 8
9 A Library of Helper Functions There's a tendancy to approach this problem with a series of nested for-loops, while the approach is valid, it doesn't generalize. It assumes a specific number of sequences. What we need is an iterator that generates all permutations of a sequence. This nested-for-loop iterator is called a Cartesian Product over sets. Python has a library to accomplish this 9
10 Using itertools import itertools for number in itertools.product("01", repeat=3): print ''.join(number)
11 All permutations of items from a list N = 0 for number in itertools.product(range(3), repeat=3): print number, N += 1 if (N % 5 == 0): print (0, 0, 0) (0, 0, 1) (0, 0, 2) (0, 1, 0) (0, 1, 1) (0, 1, 2) (0, 2, 0) (0, 2, 1) (0, 2, 2) (1, 0, 0) (1, 0, 1) (1, 0, 2) (1, 1, 0) (1, 1, 1) (1, 1, 2) (1, 2, 0) (1, 2, 1) (1, 2, 2) (2, 0, 0) (2, 0, 1) (2, 0, 2) (2, 1, 0) (2, 1, 1) (2, 1, 2) (2, 2, 0) (2, 2, 1) (2, 2, 2) 11
12 ('I', 'A', 1) ('I', 'A', 2) ('I', 'B', 1) ('I', 'B', 2) ('I', 'C', 1) ('I', 'C', 2) ('II', 'A', 1) ('II', 'A', 2) ('II', 'B', 1) ('II', 'B', 2) ('II', 'C', 1) ('II', 'C', 2) ('III', 'A', 1) ('III', 'A', 2) ('III', 'B', 1) ('III', 'B', 2) ('III', 'C', 1) ('III', 'C', 2) ('IV', 'A', 1) ('IV', 'A', 2) ('IV', 'B', 1) ('IV', 'B', 2) ('IV', 'C', 1) ('IV', 'C', 2) Permutations of mixed types for section in itertools.product(("i", "II", "III", "IV"),"ABC",range(1,3)): print section 12
13 Now let's try some Brute Force code sequences = [ 'tagtggtcttttgagtgtagatccgaagggaaagtatttccaccagttcggggtcacccagcagggcagggtgacttaat', 'cgcgactcggcgctcacagttatcgcacgtttagaccaaaacggagttagatccgaaactggagtttaatcggagtcctt', 'gttacttgtgagcctggttagatccgaaatataattgttggctgcatagcggagctgacatacgagtaggggaaatgcgt', 'aacatcaggctttgattaaacaatttaagcacgtagatccgaattgacctgatgacaatacggaacatgccggctccggg', 'accaccggataggctgcttattagatccgaaaggtagtatcgtaataatggctcagccatgtcaatgtgcggcattccac', 'tagatccgaatcgatcgtgtttctccctctgtgggttaacgaggggtccgaccttgctcgcatgtgccgaacttgtaccc', 'gaaatggttcggtgcgatatcaggccgttctcttaacttggcggtgtagatccgaacgtctctggaggggtcgtgcgcta', 'atgtatactagacattctaacgctcgcttattggcggagaccatttgctccactacaagaggctactgtgtagatccgaa', 'ttcttacacccttctttagatccgaacctgttggcgccatcttcttttcgagtccttgtacctccatttgctctgatgac', 'ctacctatgtaaaacaacatctactaacgtagtccggtctttcctgatctgccctaacctacaggtagatccgaaattcg'] def bruteforce(dna,k): """Finds a *k*-mer common to all sequences from a list of *dna* fragments with the same length""" M = len(dna) # how many sequences N = len(dna[0]) # length of sequences for offset in itertools.product(range(n-k+1), repeat=m): for i in xrange(1,len(offset)): if dna[0][offset[0]:offset[0]+k]!= dna[i][offset[i]:offset[i]+k]: break else: return offset, dna[0][offset[0]:offset[0]+10] 13
14 Test and then time it M = 4 position, motif = bruteforce(sequences[0:m], 10) print position, motif print for i in xrange(m): p = position[i] print sequences[i][:p]+sequences[i][p:p+10].upper()+sequences[i][p+10:] print %timeit bruteforce(sequences[0:m], 10) # you can try a larger value of M, but be prepared to wait (17, 47, 18, 33) tagatccgaa tagtggtcttttgagtgtagatccgaagggaaagtatttccaccagttcggggtcacccagcagggcagggtgacttaat cgcgactcggcgctcacagttatcgcacgtttagaccaaaacggagttagatccgaaactggagtttaatcggagtcctt gttacttgtgagcctggttagatccgaaatataattgttggctgcatagcggagctgacatacgagtaggggaaatgcgt aacatcaggctttgattaaacaatttaagcacgtagatccgaattgacctgatgacaatacggaacatgccggctccggg 1 loop, best of 3: 4.74 s per loop 14
15 Approximate Matching Now let's consider a more realistic motif finding problem, where the binding sites do not need to match exactly. 1 tagtggtcttttgagtgtagatctgaagggaaagtatttccaccagttcggggtcacccagcagggcagggtgacttaat 2 cgcgactcggcgctcacagttatcgcacgtttagaccaaaacggagttggatccgaaactggagtttaatcggagtcctt 3 gttacttgtgagcctggttagacccgaaatataattgttggctgcatagcggagctgacatacgagtaggggaaatgcgt 4 aacatcaggctttgattaaacaatttaagcacgtaaatccgaattgacctgatgacaatacggaacatgccggctccggg 5 accaccggataggctgcttattaggtccaaaaggtagtatcgtaataatggctcagccatgtcaatgtgcggcattccac 6 TAGATTCGAAtcgatcgtgtttctccctctgtgggttaacgaggggtccgaccttgctcgcatgtgccgaacttgtaccc 7 gaaatggttcggtgcgatatcaggccgttctcttaacttggcggtgcagatccgaacgtctctggaggggtcgtgcgcta 8 atgtatactagacattctaacgctcgcttattggcggagaccatttgctccactacaagaggctactgtgtagatccgta 9 ttcttacacccttctttagatccaaacctgttggcgccatcttcttttcgagtccttgtacctccatttgctctgatgac 10 ctacctatgtaaaacaacatctactaacgtagtccggtctttcctgatctgccctaacctacaggtcgatccgaaattcg Actually, none of the sequences have an unmodified copy of the original motif 15
16 Profile and Consensus How to find approximate string matches? Align candidate motifs by their start indexes s = ( s 1, s 2,, s t ) Construct a matrix profile with the frequencies of each nucleotide in columns Consensus nucleotide in each position has the highest score in each column 16
17 Consensus One can think of the consensus as an ancestor motif, from which mutated motifs emerged The distance between an actual motif and the consensus sequence is generally less than that for any two actual motifs Hamming distance is number of positions that differ between two strings 17
18 Consensus Properties A consensus string has a minimal hamming distance to all its source strings 18
19 Scoring Motifs Given s = ( s 1, s 2,, s t ) and DNA k j A,C,G,T i=1 Score(s, DNA) = max count(j, i) So our approach is back to brute force We consider every candidate motif in every string Return the set of indices with the highest score 19
20 Let's try again, and handle errors this time! def Score(s, DNA, k): """ compute the consensus SCORE of a given k-mer alignment given offsets into each DNA string. s = list of starting indices. DNA = list of nucleotide strings. k = Target Motif length """ score = 0 for i in xrange(k): # loop over string positions cnt = dict(zip("acgt",(0,0,0,0))) for j, sval in enumerate(s): base = DNA[j][sval+i] cnt[base] += 1 score += max(cnt.values()) return score def BruteForceMotifSearch(dna,k): M = len(dna) # how many sequences N = len(dna[0]) # length of sequences bestscore = 0 bestalignment = [] for offset in itertools.product(range(n-k+1), repeat=m): s = Score(offset,dna,k) if (s > bestscore): bestalignment = [p for p in offset] bestscore = s print bestalignment, bestscore 20
21 Test and time seqapprox = [ 'tagtggtcttttgagtgtagatctgaagggaaagtatttccaccagttcggggtcacccagcagggcagggtgacttaat', 'cgcgactcggcgctcacagttatcgcacgtttagaccaaaacggagttggatccgaaactggagtttaatcggagtcctt', 'gttacttgtgagcctggttagacccgaaatataattgttggctgcatagcggagctgacatacgagtaggggaaatgcgt', 'aacatcaggctttgattaaacaatttaagcacgtaaatccgaattgacctgatgacaatacggaacatgccggctccggg', 'accaccggataggctgcttattaggtccaaaaggtagtatcgtaataatggctcagccatgtcaatgtgcggcattccac', 'tagattcgaatcgatcgtgtttctccctctgtgggttaacgaggggtccgaccttgctcgcatgtgccgaacttgtaccc', 'gaaatggttcggtgcgatatcaggccgttctcttaacttggcggtgcagatccgaacgtctctggaggggtcgtgcgcta', 'atgtatactagacattctaacgctcgcttattggcggagaccatttgctccactacaagaggctactgtgtagatccgta', 'ttcttacacccttctttagatccaaacctgttggcgccatcttcttttcgagtccttgtacctccatttgctctgatgac', 'ctacctatgtaaaacaacatctactaacgtagtccggtctttcctgatctgccctaacctacaggtcgatccgaaattcg'] %timeit Score([17, 47, 18, 33, 21, 0, 46, 70, 16, 65], seqapprox, 10) %time BruteForceMotifSearch(seqApprox[0:4], 10) loops, best of 3: 40.6 µs per loop [17, 47, 18, 33] 36 CPU times: user 17min 22s, sys: 1.97 s, total: 17min 24s Wall time: 17min 24s 21
22 Running Time of BruteForceMotifSearch Search (N - k + 1) positions in each of M sequences, by examining (N - k + 1) sets of starting positions For each set of starting positions, the scoring function makes O(Mk) operations, so complexity is Mk(N k + 1 ) M = O(Mk N M ) That means that for M = 10, N = 80, k = 10 we must perform approximately computations Generously assuming 10 9 comps/sec it will require only secs > years ( ) Want to wait? 22 M
23 How conservative is this estimate? For the example we just did M = 4, N = 80, k = 10 So that gives operations Using our 10 9 operations per second estimate, it should have taken only 4 secs. Instead it took closer to 700 secs, which suggests we are getting around 5.85 million operations per second. So, in reality it will even take longer! 23
24 How can we find Motifs in our lifetime? Should we give up on Python and write in C? Assembly Language? Will biological insights save us this time? Are there other ways to find Motifs? Consider that if you knew what motif you were looking for, it would take only Is that significantly better? k(n k + 1)M = O(kNM) 24
Path Finding in Graphs. Problem Set #2 will be posted by tonight
 Path Finding in Graphs Problem Set #2 will be posted by tonight 1 From Last Time Two graphs representing 5-mers from the sequence "GACGGCGGCGCACGGCGCAA" Hamiltonian Path: Eulerian Path: Each k-mer is a
Path Finding in Graphs Problem Set #2 will be posted by tonight 1 From Last Time Two graphs representing 5-mers from the sequence "GACGGCGGCGCACGGCGCAA" Hamiltonian Path: Eulerian Path: Each k-mer is a
BCRANK: predicting binding site consensus from ranked DNA sequences
 BCRANK: predicting binding site consensus from ranked DNA sequences Adam Ameur October 30, 2017 1 Introduction This document describes the BCRANK R package. BCRANK[1] is a method that takes a ranked list
BCRANK: predicting binding site consensus from ranked DNA sequences Adam Ameur October 30, 2017 1 Introduction This document describes the BCRANK R package. BCRANK[1] is a method that takes a ranked list
ChIP-Seq Tutorial on Galaxy
 1 Introduction ChIP-Seq Tutorial on Galaxy 2 December 2010 (modified April 6, 2017) Rory Stark The aim of this practical is to give you some experience handling ChIP-Seq data. We will be working with data
1 Introduction ChIP-Seq Tutorial on Galaxy 2 December 2010 (modified April 6, 2017) Rory Stark The aim of this practical is to give you some experience handling ChIP-Seq data. We will be working with data
Randomized Algorithms
 Randomized Algorithms 1 Motif finding using a Profile Profile is generated by some consensus Use to find the best match of motif in each sequence These matches suggest a new consensus Reds are probabilities
Randomized Algorithms 1 Motif finding using a Profile Profile is generated by some consensus Use to find the best match of motif in each sequence These matches suggest a new consensus Reds are probabilities
Gene regulation. DNA is merely the blueprint Shared spatially (among all tissues) and temporally But cells manage to differentiate
 Gene regulation DNA is merely the blueprint Shared spatially (among all tissues) and temporally But cells manage to differentiate Especially but not only during developmental stage And cells respond to
Gene regulation DNA is merely the blueprint Shared spatially (among all tissues) and temporally But cells manage to differentiate Especially but not only during developmental stage And cells respond to
Twine User Guide. version 5/17/ Joseph Pearson, Ph.D. Stephen Crews Lab.
 Twine User Guide version 5/17/2013 http://labs.bio.unc.edu/crews/twine/ Joseph Pearson, Ph.D. Stephen Crews Lab http://www.unc.edu/~crews/ Copyright 2013 The University of North Carolina at Chapel Hill
Twine User Guide version 5/17/2013 http://labs.bio.unc.edu/crews/twine/ Joseph Pearson, Ph.D. Stephen Crews Lab http://www.unc.edu/~crews/ Copyright 2013 The University of North Carolina at Chapel Hill
Computational Genomics and Molecular Biology, Fall
 Computational Genomics and Molecular Biology, Fall 2015 1 Sequence Alignment Dannie Durand Pairwise Sequence Alignment The goal of pairwise sequence alignment is to establish a correspondence between the
Computational Genomics and Molecular Biology, Fall 2015 1 Sequence Alignment Dannie Durand Pairwise Sequence Alignment The goal of pairwise sequence alignment is to establish a correspondence between the
Special course in Computer Science: Advanced Text Algorithms
 Special course in Computer Science: Advanced Text Algorithms Lecture 6: Alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
Special course in Computer Science: Advanced Text Algorithms Lecture 6: Alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
Student: Yu Cheng (Jade) ICS 675 Assignment #1 October 25, # concatenates elem any symbol., ; ;
 Student: Yu Cheng (Jade) ICS 675 Assignment #1 October 25, 2009 Exercise 1 Algorithm time complexity analysis Question: What is the time complexity of the algorithm described below. Detail the answer using
Student: Yu Cheng (Jade) ICS 675 Assignment #1 October 25, 2009 Exercise 1 Algorithm time complexity analysis Question: What is the time complexity of the algorithm described below. Detail the answer using
CLC Server. End User USER MANUAL
 CLC Server End User USER MANUAL Manual for CLC Server 10.0.1 Windows, macos and Linux March 8, 2018 This software is for research purposes only. QIAGEN Aarhus Silkeborgvej 2 Prismet DK-8000 Aarhus C Denmark
CLC Server End User USER MANUAL Manual for CLC Server 10.0.1 Windows, macos and Linux March 8, 2018 This software is for research purposes only. QIAGEN Aarhus Silkeborgvej 2 Prismet DK-8000 Aarhus C Denmark
BLAST & Genome assembly
 BLAST & Genome assembly Solon P. Pissis Tomáš Flouri Heidelberg Institute for Theoretical Studies November 17, 2012 1 Introduction Introduction 2 BLAST What is BLAST? The algorithm 3 Genome assembly De
BLAST & Genome assembly Solon P. Pissis Tomáš Flouri Heidelberg Institute for Theoretical Studies November 17, 2012 1 Introduction Introduction 2 BLAST What is BLAST? The algorithm 3 Genome assembly De
Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014
 Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014 Dynamic programming is a group of mathematical methods used to sequentially split a complicated problem into
Dynamic Programming User Manual v1.0 Anton E. Weisstein, Truman State University Aug. 19, 2014 Dynamic programming is a group of mathematical methods used to sequentially split a complicated problem into
Biology 644: Bioinformatics
 A statistical Markov model in which the system being modeled is assumed to be a Markov process with unobserved (hidden) states in the training data. First used in speech and handwriting recognition In
A statistical Markov model in which the system being modeled is assumed to be a Markov process with unobserved (hidden) states in the training data. First used in speech and handwriting recognition In
7.36/7.91/20.390/20.490/6.802/6.874 PROBLEM SET 3. Gibbs Sampler, RNA secondary structure, Protein Structure with PyRosetta, Connections (25 Points)
 7.36/7.91/20.390/20.490/6.802/6.874 PROBLEM SET 3. Gibbs Sampler, RNA secondary structure, Protein Structure with PyRosetta, Connections (25 Points) Due: Thursday, April 3 th at noon. Python Scripts All
7.36/7.91/20.390/20.490/6.802/6.874 PROBLEM SET 3. Gibbs Sampler, RNA secondary structure, Protein Structure with PyRosetta, Connections (25 Points) Due: Thursday, April 3 th at noon. Python Scripts All
ViTraM: VIsualization of TRAnscriptional Modules
 ViTraM: VIsualization of TRAnscriptional Modules Version 2.0 October 1st, 2009 KULeuven, Belgium 1 Contents 1 INTRODUCTION AND INSTALLATION... 4 1.1 Introduction...4 1.2 Software structure...5 1.3 Requirements...5
ViTraM: VIsualization of TRAnscriptional Modules Version 2.0 October 1st, 2009 KULeuven, Belgium 1 Contents 1 INTRODUCTION AND INSTALLATION... 4 1.1 Introduction...4 1.2 Software structure...5 1.3 Requirements...5
4/26/16 Comp 555 Spring
 4/26/16 Comp 555 Spring 2016 1 RandomProfileMotifSearch is probably not the best way to find motifs. Depends on random guesses followed by a greedy optimization procedure. Major cost is Scoring, P(a P),
4/26/16 Comp 555 Spring 2016 1 RandomProfileMotifSearch is probably not the best way to find motifs. Depends on random guesses followed by a greedy optimization procedure. Major cost is Scoring, P(a P),
ViTraM: VIsualization of TRAnscriptional Modules
 ViTraM: VIsualization of TRAnscriptional Modules Version 1.0 June 1st, 2009 Hong Sun, Karen Lemmens, Tim Van den Bulcke, Kristof Engelen, Bart De Moor and Kathleen Marchal KULeuven, Belgium 1 Contents
ViTraM: VIsualization of TRAnscriptional Modules Version 1.0 June 1st, 2009 Hong Sun, Karen Lemmens, Tim Van den Bulcke, Kristof Engelen, Bart De Moor and Kathleen Marchal KULeuven, Belgium 1 Contents
Sequence Alignment. Ulf Leser
 Sequence Alignment Ulf Leser his Lecture Approximate String Matching Edit distance and alignment Computing global alignments Local alignment Ulf Leser: Bioinformatics, Summer Semester 2016 2 ene Function
Sequence Alignment Ulf Leser his Lecture Approximate String Matching Edit distance and alignment Computing global alignments Local alignment Ulf Leser: Bioinformatics, Summer Semester 2016 2 ene Function
Research Article An Improved Scoring Matrix for Multiple Sequence Alignment
 Hindawi Publishing Corporation Mathematical Problems in Engineering Volume 2012, Article ID 490649, 9 pages doi:10.1155/2012/490649 Research Article An Improved Scoring Matrix for Multiple Sequence Alignment
Hindawi Publishing Corporation Mathematical Problems in Engineering Volume 2012, Article ID 490649, 9 pages doi:10.1155/2012/490649 Research Article An Improved Scoring Matrix for Multiple Sequence Alignment
BLAST, Profile, and PSI-BLAST
 BLAST, Profile, and PSI-BLAST Jianlin Cheng, PhD School of Electrical Engineering and Computer Science University of Central Florida 26 Free for academic use Copyright @ Jianlin Cheng & original sources
BLAST, Profile, and PSI-BLAST Jianlin Cheng, PhD School of Electrical Engineering and Computer Science University of Central Florida 26 Free for academic use Copyright @ Jianlin Cheng & original sources
BLAST & Genome assembly
 BLAST & Genome assembly Solon P. Pissis Tomáš Flouri Heidelberg Institute for Theoretical Studies May 15, 2014 1 BLAST What is BLAST? The algorithm 2 Genome assembly De novo assembly Mapping assembly 3
BLAST & Genome assembly Solon P. Pissis Tomáš Flouri Heidelberg Institute for Theoretical Studies May 15, 2014 1 BLAST What is BLAST? The algorithm 2 Genome assembly De novo assembly Mapping assembly 3
MSCBIO 2070/02-710: Computational Genomics, Spring A4: spline, HMM, clustering, time-series data analysis, RNA-folding
 MSCBIO 2070/02-710:, Spring 2015 A4: spline, HMM, clustering, time-series data analysis, RNA-folding Due: April 13, 2015 by email to Silvia Liu (silvia.shuchang.liu@gmail.com) TA in charge: Silvia Liu
MSCBIO 2070/02-710:, Spring 2015 A4: spline, HMM, clustering, time-series data analysis, RNA-folding Due: April 13, 2015 by email to Silvia Liu (silvia.shuchang.liu@gmail.com) TA in charge: Silvia Liu
Locality-sensitive hashing and biological network alignment
 Locality-sensitive hashing and biological network alignment Laura LeGault - University of Wisconsin, Madison 12 May 2008 Abstract Large biological networks contain much information about the functionality
Locality-sensitive hashing and biological network alignment Laura LeGault - University of Wisconsin, Madison 12 May 2008 Abstract Large biological networks contain much information about the functionality
One report (in pdf format) addressing each of following questions.
 MSCBIO 2070/02-710: Computational Genomics, Spring 2016 HW1: Sequence alignment and Evolution Due: 24:00 EST, Feb 15, 2016 by autolab Your goals in this assignment are to 1. Complete a genome assembler
MSCBIO 2070/02-710: Computational Genomics, Spring 2016 HW1: Sequence alignment and Evolution Due: 24:00 EST, Feb 15, 2016 by autolab Your goals in this assignment are to 1. Complete a genome assembler
From Smith-Waterman to BLAST
 From Smith-Waterman to BLAST Jeremy Buhler July 23, 2015 Smith-Waterman is the fundamental tool that we use to decide how similar two sequences are. Isn t that all that BLAST does? In principle, it is
From Smith-Waterman to BLAST Jeremy Buhler July 23, 2015 Smith-Waterman is the fundamental tool that we use to decide how similar two sequences are. Isn t that all that BLAST does? In principle, it is
15.4 Longest common subsequence
 15.4 Longest common subsequence Biological applications often need to compare the DNA of two (or more) different organisms A strand of DNA consists of a string of molecules called bases, where the possible
15.4 Longest common subsequence Biological applications often need to compare the DNA of two (or more) different organisms A strand of DNA consists of a string of molecules called bases, where the possible
Sept. 9, An Introduction to Bioinformatics. Special Topics BSC5936:
 Special Topics BSC5936: An Introduction to Bioinformatics. Florida State University The Department of Biological Science www.bio.fsu.edu Sept. 9, 2003 The Dot Matrix Method Steven M. Thompson Florida State
Special Topics BSC5936: An Introduction to Bioinformatics. Florida State University The Department of Biological Science www.bio.fsu.edu Sept. 9, 2003 The Dot Matrix Method Steven M. Thompson Florida State
Min Wang. April, 2003
 Development of a co-regulated gene expression analysis tool (CREAT) By Min Wang April, 2003 Project Documentation Description of CREAT CREAT (coordinated regulatory element analysis tool) are developed
Development of a co-regulated gene expression analysis tool (CREAT) By Min Wang April, 2003 Project Documentation Description of CREAT CREAT (coordinated regulatory element analysis tool) are developed
Computational biology course IST 2015/2016
 Computational biology course IST 2015/2016 Introduc)on to Algorithms! Algorithms: problem- solving methods suitable for implementation as a computer program! Data structures: objects created to organize
Computational biology course IST 2015/2016 Introduc)on to Algorithms! Algorithms: problem- solving methods suitable for implementation as a computer program! Data structures: objects created to organize
Accelerating InDel Detection on Modern Multi-Core SIMD CPU Architecture
 Accelerating InDel Detection on Modern Multi-Core SIMD CPU Architecture Da Zhang Collaborators: Hao Wang, Kaixi Hou, Jing Zhang Advisor: Wu-chun Feng Evolution of Genome Sequencing1 In 20032: 1 human genome
Accelerating InDel Detection on Modern Multi-Core SIMD CPU Architecture Da Zhang Collaborators: Hao Wang, Kaixi Hou, Jing Zhang Advisor: Wu-chun Feng Evolution of Genome Sequencing1 In 20032: 1 human genome
Multiple Sequence Alignment Sum-of-Pairs and ClustalW. Ulf Leser
 Multiple Sequence Alignment Sum-of-Pairs and ClustalW Ulf Leser This Lecture Multiple Sequence Alignment The problem Theoretical approach: Sum-of-Pairs scores Practical approach: ClustalW Ulf Leser: Bioinformatics,
Multiple Sequence Alignment Sum-of-Pairs and ClustalW Ulf Leser This Lecture Multiple Sequence Alignment The problem Theoretical approach: Sum-of-Pairs scores Practical approach: ClustalW Ulf Leser: Bioinformatics,
The Dot Matrix Method
 Special Topics BS5936: An Introduction to Bioinformatics. Florida State niversity The Department of Biological Science www.bio.fsu.edu Sept. 9, 2003 The Dot Matrix Method Steven M. Thompson Florida State
Special Topics BS5936: An Introduction to Bioinformatics. Florida State niversity The Department of Biological Science www.bio.fsu.edu Sept. 9, 2003 The Dot Matrix Method Steven M. Thompson Florida State
Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA.
 Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA. Fasta is used to compare a protein or DNA sequence to all of the
Compares a sequence of protein to another sequence or database of a protein, or a sequence of DNA to another sequence or library of DNA. Fasta is used to compare a protein or DNA sequence to all of the
Lecture 2, Introduction to Python. Python Programming Language
 BINF 3360, Introduction to Computational Biology Lecture 2, Introduction to Python Young-Rae Cho Associate Professor Department of Computer Science Baylor University Python Programming Language Script
BINF 3360, Introduction to Computational Biology Lecture 2, Introduction to Python Young-Rae Cho Associate Professor Department of Computer Science Baylor University Python Programming Language Script
CSE 417 Dynamic Programming (pt 5) Multiple Inputs
 CSE 417 Dynamic Programming (pt 5) Multiple Inputs Reminders > HW5 due Wednesday Dynamic Programming Review > Apply the steps... optimal substructure: (small) set of solutions, constructed from solutions
CSE 417 Dynamic Programming (pt 5) Multiple Inputs Reminders > HW5 due Wednesday Dynamic Programming Review > Apply the steps... optimal substructure: (small) set of solutions, constructed from solutions
Long Read RNA-seq Mapper
 UNIVERSITY OF ZAGREB FACULTY OF ELECTRICAL ENGENEERING AND COMPUTING MASTER THESIS no. 1005 Long Read RNA-seq Mapper Josip Marić Zagreb, February 2015. Table of Contents 1. Introduction... 1 2. RNA Sequencing...
UNIVERSITY OF ZAGREB FACULTY OF ELECTRICAL ENGENEERING AND COMPUTING MASTER THESIS no. 1005 Long Read RNA-seq Mapper Josip Marić Zagreb, February 2015. Table of Contents 1. Introduction... 1 2. RNA Sequencing...
Dynamic Programming & Smith-Waterman algorithm
 m m Seminar: Classical Papers in Bioinformatics May 3rd, 2010 m m 1 2 3 m m Introduction m Definition is a method of solving problems by breaking them down into simpler steps problem need to contain overlapping
m m Seminar: Classical Papers in Bioinformatics May 3rd, 2010 m m 1 2 3 m m Introduction m Definition is a method of solving problems by breaking them down into simpler steps problem need to contain overlapping
Programming for Scientists (02-201) Prof: Phillip Compeau TA: Tim Wall
 Programming for Scientists (02-201) Prof: Phillip Compeau TA: Tim Wall Introduce Yourselves! Who Has Programmed Before? Class Goals 1. Learn the basics of programming (in Go). 2. Understand some algorithms
Programming for Scientists (02-201) Prof: Phillip Compeau TA: Tim Wall Introduce Yourselves! Who Has Programmed Before? Class Goals 1. Learn the basics of programming (in Go). 2. Understand some algorithms
ChIP-seq hands-on practical using Galaxy
 ChIP-seq hands-on practical using Galaxy In this exercise we will cover some of the basic NGS analysis steps for ChIP-seq using the Galaxy framework: Quality control Mapping of reads using Bowtie2 Peak-calling
ChIP-seq hands-on practical using Galaxy In this exercise we will cover some of the basic NGS analysis steps for ChIP-seq using the Galaxy framework: Quality control Mapping of reads using Bowtie2 Peak-calling
6.00 Introduction to Computer Science and Programming Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu 6.00 Introduction to Computer Science and Programming Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu 6.00 Introduction to Computer Science and Programming Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
CMSC 201 Fall 2016 Lab 13 More Recursion
 CMSC 201 Fall 2016 Lab 13 More Recursion Assignment: Lab 13 More Recursion Due Date: During discussion, December 5th through 8th Value: 10 points Part 1A: What is Recursion? So far this semester, we ve
CMSC 201 Fall 2016 Lab 13 More Recursion Assignment: Lab 13 More Recursion Due Date: During discussion, December 5th through 8th Value: 10 points Part 1A: What is Recursion? So far this semester, we ve
15.4 Longest common subsequence
 15.4 Longest common subsequence Biological applications often need to compare the DNA of two (or more) different organisms A strand of DNA consists of a string of molecules called bases, where the possible
15.4 Longest common subsequence Biological applications often need to compare the DNA of two (or more) different organisms A strand of DNA consists of a string of molecules called bases, where the possible
Finding Selection in All the Right Places TA Notes and Key Lab 9
 Objectives: Finding Selection in All the Right Places TA Notes and Key Lab 9 1. Use published genome data to look for evidence of selection in individual genes. 2. Understand the need for DNA sequence
Objectives: Finding Selection in All the Right Places TA Notes and Key Lab 9 1. Use published genome data to look for evidence of selection in individual genes. 2. Understand the need for DNA sequence
Using Real-valued Meta Classifiers to Integrate and Contextualize Binding Site Predictions
 Using Real-valued Meta Classifiers to Integrate and Contextualize Binding Site Predictions Offer Sharabi, Yi Sun, Mark Robinson, Rod Adams, Rene te Boekhorst, Alistair G. Rust, Neil Davey University of
Using Real-valued Meta Classifiers to Integrate and Contextualize Binding Site Predictions Offer Sharabi, Yi Sun, Mark Robinson, Rod Adams, Rene te Boekhorst, Alistair G. Rust, Neil Davey University of
Special course in Computer Science: Advanced Text Algorithms
 Special course in Computer Science: Advanced Text Algorithms Lecture 8: Multiple alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
Special course in Computer Science: Advanced Text Algorithms Lecture 8: Multiple alignments Elena Czeizler and Ion Petre Department of IT, Abo Akademi Computational Biomodelling Laboratory http://www.users.abo.fi/ipetre/textalg
ChIP-seq Analysis Practical
 ChIP-seq Analysis Practical Vladimir Teif (vteif@essex.ac.uk) An updated version of this document will be available at http://generegulation.info/index.php/teaching In this practical we will learn how
ChIP-seq Analysis Practical Vladimir Teif (vteif@essex.ac.uk) An updated version of this document will be available at http://generegulation.info/index.php/teaching In this practical we will learn how
ON HEURISTIC METHODS IN NEXT-GENERATION SEQUENCING DATA ANALYSIS
 ON HEURISTIC METHODS IN NEXT-GENERATION SEQUENCING DATA ANALYSIS Ivan Vogel Doctoral Degree Programme (1), FIT BUT E-mail: xvogel01@stud.fit.vutbr.cz Supervised by: Jaroslav Zendulka E-mail: zendulka@fit.vutbr.cz
ON HEURISTIC METHODS IN NEXT-GENERATION SEQUENCING DATA ANALYSIS Ivan Vogel Doctoral Degree Programme (1), FIT BUT E-mail: xvogel01@stud.fit.vutbr.cz Supervised by: Jaroslav Zendulka E-mail: zendulka@fit.vutbr.cz
CSI33 Data Structures
 Outline Department of Mathematics and Computer Science Bronx Community College September 6, 2017 Outline Outline 1 Chapter 2: Data Abstraction Outline Chapter 2: Data Abstraction 1 Chapter 2: Data Abstraction
Outline Department of Mathematics and Computer Science Bronx Community College September 6, 2017 Outline Outline 1 Chapter 2: Data Abstraction Outline Chapter 2: Data Abstraction 1 Chapter 2: Data Abstraction
Cache Policies. Philipp Koehn. 6 April 2018
 Cache Policies Philipp Koehn 6 April 2018 Memory Tradeoff 1 Fastest memory is on same chip as CPU... but it is not very big (say, 32 KB in L1 cache) Slowest memory is DRAM on different chips... but can
Cache Policies Philipp Koehn 6 April 2018 Memory Tradeoff 1 Fastest memory is on same chip as CPU... but it is not very big (say, 32 KB in L1 cache) Slowest memory is DRAM on different chips... but can
Lecture 9: Core String Edits and Alignments
 Biosequence Algorithms, Spring 2005 Lecture 9: Core String Edits and Alignments Pekka Kilpeläinen University of Kuopio Department of Computer Science BSA Lecture 9: String Edits and Alignments p.1/30 III:
Biosequence Algorithms, Spring 2005 Lecture 9: Core String Edits and Alignments Pekka Kilpeläinen University of Kuopio Department of Computer Science BSA Lecture 9: String Edits and Alignments p.1/30 III:
HIDDEN MARKOV MODELS AND SEQUENCE ALIGNMENT
 HIDDEN MARKOV MODELS AND SEQUENCE ALIGNMENT - Swarbhanu Chatterjee. Hidden Markov models are a sophisticated and flexible statistical tool for the study of protein models. Using HMMs to analyze proteins
HIDDEN MARKOV MODELS AND SEQUENCE ALIGNMENT - Swarbhanu Chatterjee. Hidden Markov models are a sophisticated and flexible statistical tool for the study of protein models. Using HMMs to analyze proteins
Properties of Biological Networks
 Properties of Biological Networks presented by: Ola Hamud June 12, 2013 Supervisor: Prof. Ron Pinter Based on: NETWORK BIOLOGY: UNDERSTANDING THE CELL S FUNCTIONAL ORGANIZATION By Albert-László Barabási
Properties of Biological Networks presented by: Ola Hamud June 12, 2013 Supervisor: Prof. Ron Pinter Based on: NETWORK BIOLOGY: UNDERSTANDING THE CELL S FUNCTIONAL ORGANIZATION By Albert-László Barabási
Building approximate overlap graphs for DNA assembly using random-permutations-based search.
 An algorithm is presented for fast construction of graphs of reads, where an edge between two reads indicates an approximate overlap between the reads. Since the algorithm finds approximate overlaps directly,
An algorithm is presented for fast construction of graphs of reads, where an edge between two reads indicates an approximate overlap between the reads. Since the algorithm finds approximate overlaps directly,
CS106A Handout 18 Winter February 3, 2014 Practice Midterm Exam
 CS106A Handout 18 Winter 2013-2014 February 3, 2014 Practice Midterm Exam This handout is intended to give you practice solving problems that are comparable in format and difficulty to those which will
CS106A Handout 18 Winter 2013-2014 February 3, 2014 Practice Midterm Exam This handout is intended to give you practice solving problems that are comparable in format and difficulty to those which will
Read Mapping. Slides by Carl Kingsford
 Read Mapping Slides by Carl Kingsford Bowtie Ultrafast and memory-efficient alignment of short DNA sequences to the human genome Ben Langmead, Cole Trapnell, Mihai Pop and Steven L Salzberg, Genome Biology
Read Mapping Slides by Carl Kingsford Bowtie Ultrafast and memory-efficient alignment of short DNA sequences to the human genome Ben Langmead, Cole Trapnell, Mihai Pop and Steven L Salzberg, Genome Biology
Running SNAP. The SNAP Team February 2012
 Running SNAP The SNAP Team February 2012 1 Introduction SNAP is a tool that is intended to serve as the read aligner in a gene sequencing pipeline. Its theory of operation is described in Faster and More
Running SNAP The SNAP Team February 2012 1 Introduction SNAP is a tool that is intended to serve as the read aligner in a gene sequencing pipeline. Its theory of operation is described in Faster and More
Initiate a PSI-BLAST search simply by choosing the option on the BLAST input form.
 1 2 Initiate a PSI-BLAST search simply by choosing the option on the BLAST input form. But note: invoking the algorithm is trivial. Using it correctly and interpreting the results, perhaps not so much.
1 2 Initiate a PSI-BLAST search simply by choosing the option on the BLAST input form. But note: invoking the algorithm is trivial. Using it correctly and interpreting the results, perhaps not so much.
Data Mining Technologies for Bioinformatics Sequences
 Data Mining Technologies for Bioinformatics Sequences Deepak Garg Computer Science and Engineering Department Thapar Institute of Engineering & Tecnology, Patiala Abstract Main tool used for sequence alignment
Data Mining Technologies for Bioinformatics Sequences Deepak Garg Computer Science and Engineering Department Thapar Institute of Engineering & Tecnology, Patiala Abstract Main tool used for sequence alignment
8/19/13. Computational problems. Introduction to Algorithm
 I519, Introduction to Introduction to Algorithm Yuzhen Ye (yye@indiana.edu) School of Informatics and Computing, IUB Computational problems A computational problem specifies an input-output relationship
I519, Introduction to Introduction to Algorithm Yuzhen Ye (yye@indiana.edu) School of Informatics and Computing, IUB Computational problems A computational problem specifies an input-output relationship
CS313 Exercise 4 Cover Page Fall 2017
 CS313 Exercise 4 Cover Page Fall 2017 Due by the start of class on Thursday, October 12, 2017. Name(s): In the TIME column, please estimate the time you spent on the parts of this exercise. Please try
CS313 Exercise 4 Cover Page Fall 2017 Due by the start of class on Thursday, October 12, 2017. Name(s): In the TIME column, please estimate the time you spent on the parts of this exercise. Please try
Dynamic Programming Course: A structure based flexible search method for motifs in RNA. By: Veksler, I., Ziv-Ukelson, M., Barash, D.
 Dynamic Programming Course: A structure based flexible search method for motifs in RNA By: Veksler, I., Ziv-Ukelson, M., Barash, D., Kedem, K Outline Background Motivation RNA s structure representations
Dynamic Programming Course: A structure based flexible search method for motifs in RNA By: Veksler, I., Ziv-Ukelson, M., Barash, D., Kedem, K Outline Background Motivation RNA s structure representations
1. R. Durbin, S. Eddy, A. Krogh und G. Mitchison: Biological sequence analysis, Cambridge, 1998
 7 Multiple Sequence Alignment The exposition was prepared by Clemens GrÃP pl, based on earlier versions by Daniel Huson, Knut Reinert, and Gunnar Klau. It is based on the following sources, which are all
7 Multiple Sequence Alignment The exposition was prepared by Clemens GrÃP pl, based on earlier versions by Daniel Huson, Knut Reinert, and Gunnar Klau. It is based on the following sources, which are all
Running SNAP. The SNAP Team October 2012
 Running SNAP The SNAP Team October 2012 1 Introduction SNAP is a tool that is intended to serve as the read aligner in a gene sequencing pipeline. Its theory of operation is described in Faster and More
Running SNAP The SNAP Team October 2012 1 Introduction SNAP is a tool that is intended to serve as the read aligner in a gene sequencing pipeline. Its theory of operation is described in Faster and More
Software and Hardware Acceleration of the Genomic Motif Finding Tool PhyloNet
 Department of Computer Science & Engineering 2008-18 Software and Hardware Acceleration of the Genomic Motif Finding Tool PhyloNet Authors: Justin Brown Corresponding Author: jtb1@cec.wustl.edu Type of
Department of Computer Science & Engineering 2008-18 Software and Hardware Acceleration of the Genomic Motif Finding Tool PhyloNet Authors: Justin Brown Corresponding Author: jtb1@cec.wustl.edu Type of
Boolean Network Modeling
 Boolean Network Modeling Bioinformatics: Sequence Analysis COMP 571 - Spring 2015 Luay Nakhleh, Rice University Gene Regulatory Networks Gene regulatory networks describe the molecules involved in gene
Boolean Network Modeling Bioinformatics: Sequence Analysis COMP 571 - Spring 2015 Luay Nakhleh, Rice University Gene Regulatory Networks Gene regulatory networks describe the molecules involved in gene
CSE200 Lecture 6: RECURSION
 Table of Contents Review of functions (using factorial example)... 1 Recursion... 1 Step by step run through of recursive factorial... 2 Recursion vs. iteration (for and while loops)... 3 Helper functions:...
Table of Contents Review of functions (using factorial example)... 1 Recursion... 1 Step by step run through of recursive factorial... 2 Recursion vs. iteration (for and while loops)... 3 Helper functions:...
1. R. Durbin, S. Eddy, A. Krogh und G. Mitchison: Biological sequence analysis, Cambridge, 1998
 7 Multiple Sequence Alignment The exposition was prepared by Clemens Gröpl, based on earlier versions by Daniel Huson, Knut Reinert, and Gunnar Klau. It is based on the following sources, which are all
7 Multiple Sequence Alignment The exposition was prepared by Clemens Gröpl, based on earlier versions by Daniel Huson, Knut Reinert, and Gunnar Klau. It is based on the following sources, which are all
Browser Exercises - I. Alignments and Comparative genomics
 Browser Exercises - I Alignments and Comparative genomics 1. Navigating to the Genome Browser (GBrowse) Note: For this exercise use http://www.tritrypdb.org a. Navigate to the Genome Browser (GBrowse)
Browser Exercises - I Alignments and Comparative genomics 1. Navigating to the Genome Browser (GBrowse) Note: For this exercise use http://www.tritrypdb.org a. Navigate to the Genome Browser (GBrowse)
Sequence clustering. Introduction. Clustering basics. Hierarchical clustering
 Sequence clustering Introduction Data clustering is one of the key tools used in various incarnations of data-mining - trying to make sense of large datasets. It is, thus, natural to ask whether clustering
Sequence clustering Introduction Data clustering is one of the key tools used in various incarnations of data-mining - trying to make sense of large datasets. It is, thus, natural to ask whether clustering
Transfer String Kernel for Cross-Context Sequence Specific DNA-Protein Binding Prediction. by Ritambhara Singh IIIT-Delhi June 10, 2016
 Transfer String Kernel for Cross-Context Sequence Specific DNA-Protein Binding Prediction by Ritambhara Singh IIIT-Delhi June 10, 2016 1 Biology in a Slide DNA RNA PROTEIN CELL ORGANISM 2 DNA and Diseases
Transfer String Kernel for Cross-Context Sequence Specific DNA-Protein Binding Prediction by Ritambhara Singh IIIT-Delhi June 10, 2016 1 Biology in a Slide DNA RNA PROTEIN CELL ORGANISM 2 DNA and Diseases
Easy visualization of the read coverage using the CoverageView package
 Easy visualization of the read coverage using the CoverageView package Ernesto Lowy European Bioinformatics Institute EMBL June 13, 2018 > options(width=40) > library(coverageview) 1 Introduction This
Easy visualization of the read coverage using the CoverageView package Ernesto Lowy European Bioinformatics Institute EMBL June 13, 2018 > options(width=40) > library(coverageview) 1 Introduction This
CSC 148 Lecture 3. Dynamic Typing, Scoping, and Namespaces. Recursion
 CSC 148 Lecture 3 Dynamic Typing, Scoping, and Namespaces Recursion Announcements Python Ramp Up Session Monday June 1st, 1 5pm. BA3195 This will be a more detailed introduction to the Python language
CSC 148 Lecture 3 Dynamic Typing, Scoping, and Namespaces Recursion Announcements Python Ramp Up Session Monday June 1st, 1 5pm. BA3195 This will be a more detailed introduction to the Python language
BMA Boolean Matrices as Model for Motif Kernels
 BMA Boolean Matrices as Model for Motif Kernels Jan Schröder, Manfred Schimmler, Heiko Schröder, Karsten Tischer Christian Albrechts Universität, Institut für Informatik, Kiel, Germany {jasc masch}@informatik.uni-kiel.de
BMA Boolean Matrices as Model for Motif Kernels Jan Schröder, Manfred Schimmler, Heiko Schröder, Karsten Tischer Christian Albrechts Universität, Institut für Informatik, Kiel, Germany {jasc masch}@informatik.uni-kiel.de
Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it.
 Sequence Alignments Overview Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it. Sequence alignment means arranging
Sequence Alignments Overview Sequence alignment is an essential concept for bioinformatics, as most of our data analysis and interpretation techniques make use of it. Sequence alignment means arranging
Bill Payment Optimization Algorithms
 Bill Payment Optimization Algorithms Ian Smeigh John Watts Students of Computer Science at the University of North Carolina Wilmington Abstract Bill payment is an area of life that all people have to deal
Bill Payment Optimization Algorithms Ian Smeigh John Watts Students of Computer Science at the University of North Carolina Wilmington Abstract Bill payment is an area of life that all people have to deal
Welfare Navigation Using Genetic Algorithm
 Welfare Navigation Using Genetic Algorithm David Erukhimovich and Yoel Zeldes Hebrew University of Jerusalem AI course final project Abstract Using standard navigation algorithms and applications (such
Welfare Navigation Using Genetic Algorithm David Erukhimovich and Yoel Zeldes Hebrew University of Jerusalem AI course final project Abstract Using standard navigation algorithms and applications (such
Sequence Alignment. GBIO0002 Archana Bhardwaj University of Liege
 Sequence Alignment GBIO0002 Archana Bhardwaj University of Liege 1 What is Sequence Alignment? A sequence alignment is a way of arranging the sequences of DNA, RNA, or protein to identify regions of similarity.
Sequence Alignment GBIO0002 Archana Bhardwaj University of Liege 1 What is Sequence Alignment? A sequence alignment is a way of arranging the sequences of DNA, RNA, or protein to identify regions of similarity.
CS216: Problem Set 1: Sequence Alignment
 1 of 5 1/18/2006 8:45 AM University of Virginia Computer Science CS216: Program and Data Representation, Spring 2006 Problem Set 1 Sequence Alignment 18 January 2006 Out: 18 January Due: Monday, 23 January
1 of 5 1/18/2006 8:45 AM University of Virginia Computer Science CS216: Program and Data Representation, Spring 2006 Problem Set 1 Sequence Alignment 18 January 2006 Out: 18 January Due: Monday, 23 January
On the Efficacy of Haskell for High Performance Computational Biology
 On the Efficacy of Haskell for High Performance Computational Biology Jacqueline Addesa Academic Advisors: Jeremy Archuleta, Wu chun Feng 1. Problem and Motivation Biologists can leverage the power of
On the Efficacy of Haskell for High Performance Computational Biology Jacqueline Addesa Academic Advisors: Jeremy Archuleta, Wu chun Feng 1. Problem and Motivation Biologists can leverage the power of
Database Searching Using BLAST
 Mahidol University Objectives SCMI512 Molecular Sequence Analysis Database Searching Using BLAST Lecture 2B After class, students should be able to: explain the FASTA algorithm for database searching explain
Mahidol University Objectives SCMI512 Molecular Sequence Analysis Database Searching Using BLAST Lecture 2B After class, students should be able to: explain the FASTA algorithm for database searching explain
BIOL591: Introduction to Bioinformatics Alignment of pairs of sequences
 BIOL591: Introduction to Bioinformatics Alignment of pairs of sequences Reading in text (Mount Bioinformatics): I must confess that the treatment in Mount of sequence alignment does not seem to me a model
BIOL591: Introduction to Bioinformatics Alignment of pairs of sequences Reading in text (Mount Bioinformatics): I must confess that the treatment in Mount of sequence alignment does not seem to me a model
Computer Science 121. Scientific Computing Winter 2016 Chapter 8 Loops
 Computer Science 121 Scientific Computing Winter 2016 Chapter 8 Loops Loops: Motivation We've already seen algorithms (findsmallest, Fibonacci, factorial) that repeat the same steps over and over Without
Computer Science 121 Scientific Computing Winter 2016 Chapter 8 Loops Loops: Motivation We've already seen algorithms (findsmallest, Fibonacci, factorial) that repeat the same steps over and over Without
CisGenome User s Manual
 CisGenome User s Manual 1. Overview 1.1 Basic Framework of CisGenome 1.2 Installation 1.3 Summary of All Functions 1.4 A Quick Start Analysis of a ChIP-chip Experiment 2. Genomics Toolbox I Establishing
CisGenome User s Manual 1. Overview 1.1 Basic Framework of CisGenome 1.2 Installation 1.3 Summary of All Functions 1.4 A Quick Start Analysis of a ChIP-chip Experiment 2. Genomics Toolbox I Establishing
Genome Browsers Guide
 Genome Browsers Guide Take a Class This guide supports the Galter Library class called Genome Browsers. See our Classes schedule for the next available offering. If this class is not on our upcoming schedule,
Genome Browsers Guide Take a Class This guide supports the Galter Library class called Genome Browsers. See our Classes schedule for the next available offering. If this class is not on our upcoming schedule,
Sequence alignment theory and applications Session 3: BLAST algorithm
 Sequence alignment theory and applications Session 3: BLAST algorithm Introduction to Bioinformatics online course : IBT Sonal Henson Learning Objectives Understand the principles of the BLAST algorithm
Sequence alignment theory and applications Session 3: BLAST algorithm Introduction to Bioinformatics online course : IBT Sonal Henson Learning Objectives Understand the principles of the BLAST algorithm
m6aviewer Version Documentation
 m6aviewer Version 1.6.0 Documentation Contents 1. About 2. Requirements 3. Launching m6aviewer 4. Running Time Estimates 5. Basic Peak Calling 6. Running Modes 7. Multiple Samples/Sample Replicates 8.
m6aviewer Version 1.6.0 Documentation Contents 1. About 2. Requirements 3. Launching m6aviewer 4. Running Time Estimates 5. Basic Peak Calling 6. Running Modes 7. Multiple Samples/Sample Replicates 8.
Using Biopython for Laboratory Analysis Pipelines
 Using Biopython for Laboratory Analysis Pipelines Brad Chapman 27 June 2003 What is Biopython? Official blurb The Biopython Project is an international association of developers of freely available Python
Using Biopython for Laboratory Analysis Pipelines Brad Chapman 27 June 2003 What is Biopython? Official blurb The Biopython Project is an international association of developers of freely available Python
Review of feature selection techniques in bioinformatics by Yvan Saeys, Iñaki Inza and Pedro Larrañaga.
 Americo Pereira, Jan Otto Review of feature selection techniques in bioinformatics by Yvan Saeys, Iñaki Inza and Pedro Larrañaga. ABSTRACT In this paper we want to explain what feature selection is and
Americo Pereira, Jan Otto Review of feature selection techniques in bioinformatics by Yvan Saeys, Iñaki Inza and Pedro Larrañaga. ABSTRACT In this paper we want to explain what feature selection is and
Introduction to Python (All the Basic Stuff)
 Introduction to Python (All the Basic Stuff) 1 Learning Objectives Python program development Command line, IDEs, file editing Language fundamentals Types & variables Expressions I/O Control flow Functions
Introduction to Python (All the Basic Stuff) 1 Learning Objectives Python program development Command line, IDEs, file editing Language fundamentals Types & variables Expressions I/O Control flow Functions
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 04: Variations of sequence alignments http://www.pitt.edu/~mcs2/teaching/biocomp/tutorials/global.html Slides adapted from Dr. Shaojie Zhang (University
EECS730: Introduction to Bioinformatics Lecture 04: Variations of sequence alignments http://www.pitt.edu/~mcs2/teaching/biocomp/tutorials/global.html Slides adapted from Dr. Shaojie Zhang (University
Program Synthesis. SWE 795, Spring 2017 Software Engineering Environments
 Program Synthesis SWE 795, Spring 2017 Software Engineering Environments Today HW3 is due next week in class! Part 1 (Lecture)(~50 mins) Break! Part 2 (Discussion)(~60 mins) Discussion of readings Part
Program Synthesis SWE 795, Spring 2017 Software Engineering Environments Today HW3 is due next week in class! Part 1 (Lecture)(~50 mins) Break! Part 2 (Discussion)(~60 mins) Discussion of readings Part
Shortest Path Algorithm
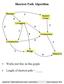 Shortest Path Algorithm C Works just fine on this graph. C Length of shortest path = Copyright 2005 DIMACS BioMath Connect Institute Robert Hochberg Dynamic Programming SP #1 Same Questions, Different
Shortest Path Algorithm C Works just fine on this graph. C Length of shortest path = Copyright 2005 DIMACS BioMath Connect Institute Robert Hochberg Dynamic Programming SP #1 Same Questions, Different
RAPID Programming of Pattern-Recognition Processors
 RAPID Programming of Pattern-Recognition Processors Kevin Angstadt Westley Weimer Kevin Skadron Department of Computer Science University of Virginia {angstadt, weimer, skadron}@cs.virginia.edu 6. April
RAPID Programming of Pattern-Recognition Processors Kevin Angstadt Westley Weimer Kevin Skadron Department of Computer Science University of Virginia {angstadt, weimer, skadron}@cs.virginia.edu 6. April
Finding local RNA motifs using covariance models
 Finding local RNA motifs using covariance models Sohrab P. Shah and Anne Condon Department of Computer Science, University of British Columbia, Vancouver, BC, Canada sshah, condon@cs.ubc.ca Technical Report
Finding local RNA motifs using covariance models Sohrab P. Shah and Anne Condon Department of Computer Science, University of British Columbia, Vancouver, BC, Canada sshah, condon@cs.ubc.ca Technical Report
Using Hidden Markov Models for Multiple Sequence Alignments Lab #3 Chem 389 Kelly M. Thayer
 Página 1 de 10 Using Hidden Markov Models for Multiple Sequence Alignments Lab #3 Chem 389 Kelly M. Thayer Resources: Bioinformatics, David Mount Ch. 4 Multiple Sequence Alignments http://www.netid.com/index.html
Página 1 de 10 Using Hidden Markov Models for Multiple Sequence Alignments Lab #3 Chem 389 Kelly M. Thayer Resources: Bioinformatics, David Mount Ch. 4 Multiple Sequence Alignments http://www.netid.com/index.html
B - Broken Track Page 1 of 8
 B - Broken Track There's a gap in the track! We need to make our robot even more intelligent so it won't get stuck, and can find the track again on its own. 2017 https://www.hamiltonbuhl.com/teacher-resources
B - Broken Track There's a gap in the track! We need to make our robot even more intelligent so it won't get stuck, and can find the track again on its own. 2017 https://www.hamiltonbuhl.com/teacher-resources
Evolutionary tree reconstruction (Chapter 10)
 Evolutionary tree reconstruction (Chapter 10) Early Evolutionary Studies Anatomical features were the dominant criteria used to derive evolutionary relationships between species since Darwin till early
Evolutionary tree reconstruction (Chapter 10) Early Evolutionary Studies Anatomical features were the dominant criteria used to derive evolutionary relationships between species since Darwin till early
Index Construction. Dictionary, postings, scalable indexing, dynamic indexing. Web Search
 Index Construction Dictionary, postings, scalable indexing, dynamic indexing Web Search 1 Overview Indexes Query Indexing Ranking Results Application Documents User Information analysis Query processing
Index Construction Dictionary, postings, scalable indexing, dynamic indexing Web Search 1 Overview Indexes Query Indexing Ranking Results Application Documents User Information analysis Query processing
Higher Order PWM for Modeling Transcription Factor Binding Sites
 San Jose State University SJSU ScholarWorks Master's Projects Master's Theses and Graduate Research Fall 2013 Higher Order PWM for Modeling Transcription Factor Binding Sites Dhivya Srinivasan Follow this
San Jose State University SJSU ScholarWorks Master's Projects Master's Theses and Graduate Research Fall 2013 Higher Order PWM for Modeling Transcription Factor Binding Sites Dhivya Srinivasan Follow this
Sequence comparison: Local alignment
 Sequence comparison: Local alignment Genome 559: Introuction to Statistical an Computational Genomics Prof. James H. Thomas http://faculty.washington.eu/jht/gs559_217/ Review global alignment en traceback
Sequence comparison: Local alignment Genome 559: Introuction to Statistical an Computational Genomics Prof. James H. Thomas http://faculty.washington.eu/jht/gs559_217/ Review global alignment en traceback
