Segmentation of Brain Internal Structures Automatically Using Non-Rigid Registration with Simultaneous Intensity and Geometric Match
|
|
|
- Bridget Ryan
- 6 years ago
- Views:
Transcription
1 Eighth International Conference on Intelligent Systems Design and Applications Segmentation of Brain Internal Structures Automatically Using Non-Rigid Registration with Simultaneous Intensity and Geometric Match Xiangbo LIN *,, Tianshuang QIU * 1, Su RUAN, Frederic Morain-Nicolier *.Department of Electronic Engineering, Dalian University of Technology, Dalian , China;. Dept. GE&II, IUT de Troyes, Université de Reims Champagne-Ardenne, Troyes Cedex, France; linxbo@dlut.edu.cn; qiutsh@dlut.edu.cn; su.ruan@univ-reims.fr ;frederic.nicolier@univ-reims.fr Abstract Segmentation of the brain internal structures is an important and a challenging task due to their small size, partial volume effects, and anatomical variability. In this paper we propose a method that segments automatically the deep brain internal structures from brain MRI images. It uses a combination of local affine transformation and optical flow based non-rigid registration, which has the advantages of modifying the larger geometric deformation and intensity differences simultaneously. Meanwhile the residual subtle differences decrease due to the high degree of freedom. Both simulated data and real data are used to validate the proposed method and the results are encouraging. It can be concluded that the image gray level of the corresponding structures plays an important role in registration based segmentation using intensity metric. Key words: Non-rigid registration; Brain structures segmentation; MRI 1. Introduction In recent years, the analysis of anatomical structures from medical images develops rapidly [1-3] due to the widespread research on brain functions and brain disorders. Brain internal structures play a central role in the intellectual capabilities of the human brain. Additionally, these structures are also relevant to a set of clinical conditions, such as Parkinson s and Creutzfeldt-Jakob diseases. However, segmenting these structures remains a challenging task due to their small size, partial volume effects, anatomical variability, and the lack of clearly defined edges [4]. Concerning about the reliability and the flexibility, manual delineation is still the main choice in clinical medicine at present despite the fact that many fully automatic and semiautomatic segmentation methods have been proposed in the literatures [2, 5-14]. However the large amount of data to be analyzed makes manual analysis of these images impractical. Moreover manual segmentation is prone to errors associated with inter-observer and intro-observer variability. Therefore it is necessary to find accurate automatic segmentation methods close to manual delineation. A variety of computer-assisted methods has been studied to automatically segment brain internal structures [5]. We can cite deformable models or active contour evolution based methods [6-9, 13], which can be good solutions to the problem because of their abilities to capture the information of the shapes or structures of interest. However the initialization of these methods prior to deformation remains difficult. Another crucial technology is image registration [1, 2, 10-12]. These methods rely on a reference image volume in which structures of interest have been carefully segmented by experts. To segment a new image volume, a transformation that registers the reference volume to the target volume is computed, which gives a spatial correspondence between the two image volumes. Then regions labeled in the reference volume can be projected onto the volume of interest. The key of this approach is to design a method being capable of computing the transformation in a reliable and accurate way. These methods take advantage of the prior knowledge, which is explicitly provided by the atlas (segmented reference volume), such as structure shape, relative positions between the structures. This allows helping the segmentation of the anatomical structures without clearly defined contours. The adopted method in this paper, Demons registration [15], belongs to intensity based non-rigid 1 Corresponding author: Tel: ; Fax: ; /08 $ IEEE DOI /ISDA
2 registration algorithm. It has high ability to model local deformation and has been used in medical image registration [15-18] successfully. However the requirements of intensity correspondence and small initial deformation between homologous structures put some limitations on it. In general global rigid or affine transformation and simple histogram match were often used to solve the problem [1, 19-22], but they were not ideal solutions. Therefore, we propose to implement the preprocess using a local affine transformation methods, which has the advantages of modifying the larger geometric deformation and intensity differences simultaneously [23]. Then the Demons registration with higher degree of freedom is used to cope with the residual subtle differences. 2. Methods The designed model of the local affine transformation to get the intensity match and the rough spatial match is m f ( x, y, t) + m 7 8 (1) = f ( m x + m y + m, m x + m y + m, t 1) Where the position parameters m, i = 1,,6 reflect i the spatial deformation, the brightness and contrast parameters m, m reflect the intensity variation 7 8 between the two images, and a temporal parameter t is used to distinguish the two images. The model parameters will be acquired after the error function T 2 E( m) = [ k c m ] x, y ω is minimized. ( k = f f + xf + yf, c =[ xf yf xf yf f f f 1] T, t x y x x y y x y ω denotes a small spatial neighborhood). After preprocess, a rough match is acquired. Then subtle differences are corrected using the Demons registration algorithm. The main idea of the Demons registration algorithm is to consider the non-rigid registration as a diffusion process. Although the behavior of the Demons can be designed in different ways, implementation method based on the optical flow theory is used for most medical image analysis. The assumption in optical flow field is intensity preservation in image moving process, that is I( x( t), y( t), t ) = const (2) Differentiating I with respect t results in a velocity vector ( m s) s v = (3) 2 2 s + ( m s) 2 Where ( m s) is an added term to keep the stability. When v is obtained, the atlas can be registered to the input image. More details about the local affine registration algorithm and the Demons algorithm can be found in [23] and [15]. The two assumptions of the Demons registration algorithm using optical equation are as follows: small deformation and intensity conservation between the reference image and the floating image. However it is hard to satisfy the assumptions in real image registration, especially in atlas-based registration, because the brain morphology and the intensities of any input image will be different comparing to the reference image. Fortunately the adopted local affine transformation is more effective to deal with larger deformation and can modify simultaneously the geometric shape and brightness/contrast differences despite its limited ability in registering subtle deformation due to the necessary size of the subregions. It shows that it is reasonable to combine the two registration algorithm to complete the segmentation task. After local affine transformation with adaptively intensity correction, the shape differences and the intensity variations between the reference image and the floating image will be small enough to satisfy the assumption of Demons algorithm. This enables the Demons registration algorithm more effectively to deal with subtle local differences. 3. Experiments and Results 3.1. Materials The reference image used in the experiment is the one from the Surgical Planning Laboratory of Harvard Medical School [24]. It consists of voxels with a spatial resolution of mm mm 1.5 mm. The test images were imaged with 1.5T GE scanner, and Axial 3D IR T1- weighted (TI/TR/TE: 600/10/2) acquired using a fast gradient echo with inversion recovery sequence. Each dataset (volume) consists of voxels, and the resolution of each voxel is mm mm 1.5 mm. Both larger intensity difference between the reference image and the test image due to different scan equipment and the individual morphology differences between subjects can be observed, that makes the registration difficult. Two experiments were carried out to get the quantitative assessment of the performance of the segmentation method. The first was synthetic deformation experiment, in which the SPL [24] standard reference image and the corresponding atlas 526
3 were deformed using an analytic harmonic deformation approach [25]. Then the reference image was registered to the deformed image to get the deformation field. Finally the acquired segmentation result using the deformation field was compared with the real segmentation, the deformed atlas. Ten volumes were used in this experiment. The second experiment consists of a performance comparison between the automatic segmentation and expert segmentation using ten real data Results Visual inspection and comparison. Figure 1 shows the registration performances before and after correcting subtle local differences using Demons registration. It can be seen that the difference is larger when only using local elastic registration (Figure 1(c)). After subtle local difference correction using Demons registration, the difference decreases. (a) (b) (c) (d) (e) Fig. 1. Comparison of the pure local elastic registration and the proposed method. (a) Internal region of reference image. (b) Result of local elastic registration. (c)differences between (a) and (b). (d) Result of the proposed method. (e) Differences between (a) and (d). Figure 2 shows a comparison between the Demons algorithm with global affine transform and histogram match and the Demons algorithm with the proposed preprocess method. Visual inspection of the segmentation results was performed by comparing gold standard (gray) and automatic (white) segmentation on caudate, putamen, pallidus and hippocampus. It can be clearly seen that better segmentation results are obtained with the proposed method. (a) (b) Fig. 2. Comparison of gold standard (gray) and automatic (white) segmentation on caudate, putamen, pallidus and hippocampus (a) using Demons algorithm with global affine transform and histogram match (b) using Demons algorithm with the proposed preprocess method Quantitative evaluation. validation criteria. To validate the results quantitatively, the criteria widely used in atlas-based segmentation [1-3, 13, 16], a kappa statistic based similarity index (KI) is adopted in this paper. The KI value measures the overlap ratio between the segmented structure and the ground truth, which is defined as KI=2TP/(2TP+FN+FP). The definitions of the parameters are as follows: TP = G E is the number of true positive; FP = G E is the number of false positive and FN = G E is the number of false negative. Where G is the ground truth segmentation of a given structure, E is the estimated segmentation of the same structure, and A denotes the complement of a set A. validation using synthetic deformation. It is hard to obtain real MRI images of the human brain with known segmentation results of the internal structures. Therefore, a quantitative assessment of the performance of the segmentation method requires the use of simulated data with known segmentation as the gold standard. The synthetic images came from the deformed SPL reference image, which was deformed using an analytic harmonic deformation approach [25]. 527
4 Table 1 shows the segmentation results on simulated data. It can be seen that there were great differences in cross correlation between the deformed images and the original reference image by the synthetic deformation. However the mean KI ± standard deviation for the four structures for ten cases was ± which was perfect results regardless how low the cross correlation was before registration between the reference image and the floating image. It showed that high registration quality could lead to high segmentation results. validation with expert delineation. Although validation with simulated data is a common method for its known segmentation, it is not sufficient to validate the approach. One drawback is that the deformation is not representative of the real inter-subject variability, and the other is that it doesn t consider the noise and artifact in real MRI images. So additional testing with real data must also be completed to demonstrate the ability of the algorithm to work under real-world conditions. For this reason, several real MRI datasets from different subjects were also used to test the performance of the approach. Table 2 shows the segmentation result using real brain MRI data. For caudate, the best overlap ratio was and the worst was , and the mean±standard deviation was ± for ten cases. For putamen, the best overlap ratio was and the worst was , and the mean±standard deviation was ± And for hippocampus, the best overlap ratio was and the worst was , and the mean± standard deviation was ± Comparison to the three structures, segmentation result for the pallidus was not very well. The reason may be due to its small size, which is easy to be influenced by the noise and artifact. cross correlation Table 1. Segmentation results on simulated data Caudate Putamen Pallidus Hippocampus before after L R L R L R L R KI cross correlation Table 2. Segmentation results on real data Caudate Putamen Pallidus Hippocampus before after L R L R L R L R KI 4. Discussion Two validation methods were used in this paper to evaluate the segmentation ability of the proposed nonrigid registration approach. The first was on the simulated data with known segmentation acquired 528
5 from the synthetic deformation, while the second was on real data with the expert delineation. Each validation method show their advantages and disadvantages. Validation on simulated data makes it possible generate known reference segmentation, thus the accuracy of the estimated solution can be evaluated. However there exist two main drawbacks. One is that the deformation is not representative of the real intersubject variability and the other is that it doesn t consider the noise and artifact in real MRI images. The second validation method is obviously in line with the actual situation. The main drawback of this validation is that the segmentation map, provided by an expert, cannot be considered as a perfect ground truth. So it can be seen that the first validation method has true reference segmentation but is not representative of the real situation, while the second validation method reflects the real situation but has no completely true reference segmentation. From the two experiment results, a phenomenon was also worthy of paying attention to. The segmentation result is very nice on simulated data in all cases and in all the four structures. While the segmentation result on real data is relatively not good enough on real data. It is possible that using expert segmentation as gold standard may have influence on the result, but it should not be the main reason. The actual reason should be due to the data differences. As it can be seen there are two main differences between the two data. One is that the deformation in simulated data couldn t reflect the real shape difference in different subjects while the real data could. However both the deformation are nonlinear, they should have the same characteristic. Thus it should not be the main reason. The other difference is that for simulated data, the homologous structures in reference image and floating image have the similar intensity. But for real data, these are not true because of the noise and interference introduced by the scanning procedure. This should be a serious problem for intensity based registration. The obtained better results using the proposed method have provided some evidences. Therefore, improving image quality and intensity correspondence of homologous structures should be a promising strategy for structure segmentation based on intensity based non-rigid registration. 5. Conclusion A combination of local affine transformation and Demons free-form transformation is used to segment automatically the brain internal structures in this paper. The segmentation results on both the simulated data and the real data are encouraging. It indicates that the proposed method could be a solution for segmenting brain internal structures which lack clearly defined intensity boundaries from human MRI images. Compared to global affine transform and simple histogram match, the selected local affine transform with intensity correction simultaneously provide a better initial conditions to the Demons registration algorithm. Furthermore, the results on the simulated data are superior to that on the real data due to the nice intensity correspondence between the homologous structures. It can be concluded that the image gray level of the corresponding structures plays an important role in registration based segmentation using intensity metric. 6. Acknowledgments This work is partly supported by National Natural Science Foundation of China (No , No , No ) and the Youth Foundation of Dalian University of Technology. 7. References [1] B. M. Dawant, S. L. Hartmann, J, -P. Thirion, et al. Automatic 3-D Segmentation of Internal Structures of the Head in MR Images Using a Combination of Similarity and Free-Form Transformations: Part I, Methodology and Validation on Normal Subjects. IEEE Transactions On Medical Imaging, 1999, 18(10): [2] Bruce Fischl, David H. Salat, Evelina Busa, et al.whole Brain Segmentation: Automated Labeling of Neuroanatomical Structures in the Human Brain. Neuron, 2002, 33(1): [3] Yan Xia, Keith Bettinger, Lin Shen, Allan L. Reiss. Automatic Segmentation of the Caudate Nucleus From Human Brain MR Images. IEEE Transactions on Medical Imaging, 2007, 26(4): [4] M. G. Linguraru, M. Á. G. Ballester, N. Ayache. Deformable Atlases for the Segmentation of Internal Brain Nuclei in Magnetic Resonance Imaging. International Journal of Computers, Communications & Control, 2007(II): [5] L. Clarke, R. Velthuizen, M. Camacho, et al. MRI segmentation: Methods and applications. Magn. Resonance Imag., 1995, 13: [6] T. McInerney D. Terzopoulos. Deformable models in medical image analysis: A survey. Med. Image Anal., 1996, (1): [7] P. Yushkevich, J. Piven, H. Hazlett, R. et al. Userguided 3-D active contour segmentation of anatomical structures: Significantly improved efficiency and reliability. NeuroImage, 2006, 31:
6 [8] Pitiot, H. Delingette, P. Thompson, N. Ayache. Expert knowledge-guided segmentation system for brain MRI. NeuroImage, 2004, 23: [9] T. F. Cootes, A. Hill, C. J. Taylor, J. Haslam. The use of active shape models for locating structures in medical images. Image and Vision Computing, 1994, 12(6): [10] Daniel Schwarz, Tomas Kasparek, Ivo Provaznik, Jiri Jarkovsky. A Deformable Registration Method for Automated Morphometry of MRI Brain Images in Neuropsychiatric Research. IEEE Transactions on Medical Imaging, 2007, 26(4): [11] D. V. Iosifescu, M. E. Shenton, S. K.Warfield, et al. An automated registration algorithm for measuring MRI subcortical brain structures. NeuroImage, 1997, 6: [12] A. Kelemen, G. Szekely, and G. Gerig. Elastic modelbased segmentation of 3-D neuroradiological data sets. IEEE Trans. Med. Imag., 1999, 18(10): [13] C. Baillard, P. Hellier, C. Barillot. Segmentation of brain 3D MR images using level sets and dense registration. Medical Image Analysis, 2001, 5: [14] V. Barra, J. Y. Boire. Automatic segmentation of subcortical brain structures in MR images using information fusion. IEEE Trans. Med. Imag., 2001, 20(7): [15] J. P. Thirion. Image matching as a diffusion process: an analogy with Maxwell s demons. Medical Image Analysis, 1998, 2(3): [16] V. Noblet, C. Heinrich, F. Heitz, et al. Retrospective evaluation of a topology preserving non-rigid registration method. Medical Image Analysis, 2006, 10: [17] Alexandre Guimond, Alexis Roche, Nicholas Ayache, et al. Three-Dimensional Multimodal Brain Warping Using the Demons Algorithm and Adaptive Intensity Corrections. IEEE Transactions On Medical Imaging, 2001, 20(1): [18] He Wang, Lei Dong, et al. Validation of an accelerated demons algorithm for deformable image registration in radiation therapy. Phys. Med. Biol., 2005, 50(12): [19] M. Bach Cuadra, M. De Craene, V. Duay, et al. Dense deformation field estimation for atlas-based segmentation of pathological MR brain images. Computer Methods And Programs In Biomedicine, 2006, 84: [20] Steven L. Hartmann, Mitchell H. Parks, Peter R. Martin, Benoit M. Dawant. Automatic 3-D Segmentation of Internal Structures of the Head in MR Images Using a Combination of Similarity and Free-Form Transformations: Part II, Validation on Severely Atrophied Brains. IEEE Transactions on Medical Imaging, 1999, 18(10): [21] P. Hellier*, C. Barillot, I. Corouge, et al. Retrospective Evaluation of Intersubject Brain Registration. IEEE Transactions on Medical Imaging, 2003, 22(9): [22] Parraga A, Susin A, Pettersson J, Macq B, De Craene M. Quality Assessment of Non-rigid Registration Methods for Atlas-based Segmentation in Head-neck Radiotherapy. Proceedings of IEEE International Conference on Acoustics, Speech, & Signal Processing, Honolulu, Hawaii, USA, Volume: 1, I-445 I-448. [23] Senthil Periaswamy, Hany Farid. Elastic Registration in the Presence of Intensity Variations. IEEE Transactions on Medical Imaging, 2003, 22(7): [24] R. Kikinis, et al. A digital brain atlas for surgical planning, model driven segmentation and teaching, IEEE Trans. Visual. Comput. Graph. 1996, 2 (3): [25] Weiguo Lu, et al Fast free-form deformable registration via calculus of variations. Phys. Med. Biol. 2004, 49:
Automatic Hippocampus Segmentation from Brain MRI Images
 International Journal of Computer Information Systems and Industrial Management Applications (IJCISIM) ISSN: 150-7988 Vol.1 (009), pp.39-48 http://www.mirlabs.org/ijcisim Automatic Hippocampus Segmentation
International Journal of Computer Information Systems and Industrial Management Applications (IJCISIM) ISSN: 150-7988 Vol.1 (009), pp.39-48 http://www.mirlabs.org/ijcisim Automatic Hippocampus Segmentation
CHAPTER 2. Morphometry on rodent brains. A.E.H. Scheenstra J. Dijkstra L. van der Weerd
 CHAPTER 2 Morphometry on rodent brains A.E.H. Scheenstra J. Dijkstra L. van der Weerd This chapter was adapted from: Volumetry and other quantitative measurements to assess the rodent brain, In vivo NMR
CHAPTER 2 Morphometry on rodent brains A.E.H. Scheenstra J. Dijkstra L. van der Weerd This chapter was adapted from: Volumetry and other quantitative measurements to assess the rodent brain, In vivo NMR
VALIDATION OF DIR. Raj Varadhan, PhD, DABMP Minneapolis Radiation Oncology
 VALIDATION OF DIR Raj Varadhan, PhD, DABMP Minneapolis Radiation Oncology Overview Basics: Registration Framework, Theory Discuss Validation techniques Using Synthetic CT data & Phantoms What metrics to
VALIDATION OF DIR Raj Varadhan, PhD, DABMP Minneapolis Radiation Oncology Overview Basics: Registration Framework, Theory Discuss Validation techniques Using Synthetic CT data & Phantoms What metrics to
Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge
 Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge Christian Wasserthal 1, Karin Engel 1, Karsten Rink 1 und André Brechmann
Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge Christian Wasserthal 1, Karin Engel 1, Karsten Rink 1 und André Brechmann
Semi-automated Basal Ganglia Segmentation Using Large Deformation Diffeomorphic Metric Mapping
 Semi-automated Basal Ganglia Segmentation Using Large Deformation Diffeomorphic Metric Mapping Ali Khan 1, Elizabeth Aylward 2, Patrick Barta 3, Michael Miller 4,andM.FaisalBeg 1 1 School of Engineering
Semi-automated Basal Ganglia Segmentation Using Large Deformation Diffeomorphic Metric Mapping Ali Khan 1, Elizabeth Aylward 2, Patrick Barta 3, Michael Miller 4,andM.FaisalBeg 1 1 School of Engineering
Multi-atlas labeling with population-specific template and non-local patch-based label fusion
 Multi-atlas labeling with population-specific template and non-local patch-based label fusion Vladimir Fonov, Pierrick Coupé, Simon Eskildsen, Jose Manjon, Louis Collins To cite this version: Vladimir
Multi-atlas labeling with population-specific template and non-local patch-based label fusion Vladimir Fonov, Pierrick Coupé, Simon Eskildsen, Jose Manjon, Louis Collins To cite this version: Vladimir
Curvature guided surface registration using level sets
 Curvature guided surface registration using level sets Marcel Lüthi, Thomas Albrecht, Thomas Vetter Department of Computer Science, University of Basel, Switzerland Abstract. We present a new approach
Curvature guided surface registration using level sets Marcel Lüthi, Thomas Albrecht, Thomas Vetter Department of Computer Science, University of Basel, Switzerland Abstract. We present a new approach
THE identification and delineation of brain structures from
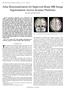 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 26, NO. 4, APRIL 2007 479 Atlas Renormalization for Improved Brain MR Image Segmentation Across Scanner Platforms Xiao Han and Bruce Fischl* Abstract Atlas-based
IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 26, NO. 4, APRIL 2007 479 Atlas Renormalization for Improved Brain MR Image Segmentation Across Scanner Platforms Xiao Han and Bruce Fischl* Abstract Atlas-based
The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy
 The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy Sokratis K. Makrogiannis, PhD From post-doctoral research at SBIA lab, Department of Radiology,
The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy Sokratis K. Makrogiannis, PhD From post-doctoral research at SBIA lab, Department of Radiology,
Automatic Subthalamic Nucleus Targeting for Deep Brain Stimulation. A Validation Study
 Automatic Subthalamic Nucleus Targeting for Deep Brain Stimulation. A Validation Study F. Javier Sánchez Castro a, Claudio Pollo a,b, Jean-Guy Villemure b, Jean-Philippe Thiran a a École Polytechnique
Automatic Subthalamic Nucleus Targeting for Deep Brain Stimulation. A Validation Study F. Javier Sánchez Castro a, Claudio Pollo a,b, Jean-Guy Villemure b, Jean-Philippe Thiran a a École Polytechnique
Elastically Deforming a Three-Dimensional Atlas to Match Anatomical Brain Images
 University of Pennsylvania ScholarlyCommons Technical Reports (CIS) Department of Computer & Information Science May 1993 Elastically Deforming a Three-Dimensional Atlas to Match Anatomical Brain Images
University of Pennsylvania ScholarlyCommons Technical Reports (CIS) Department of Computer & Information Science May 1993 Elastically Deforming a Three-Dimensional Atlas to Match Anatomical Brain Images
Subcortical Structure Segmentation using Probabilistic Atlas Priors
 Subcortical Structure Segmentation using Probabilistic Atlas Priors Sylvain Gouttard 1, Martin Styner 1,2, Sarang Joshi 3, Brad Davis 2, Rachel G. Smith 1, Heather Cody Hazlett 1, Guido Gerig 1,2 1 Department
Subcortical Structure Segmentation using Probabilistic Atlas Priors Sylvain Gouttard 1, Martin Styner 1,2, Sarang Joshi 3, Brad Davis 2, Rachel G. Smith 1, Heather Cody Hazlett 1, Guido Gerig 1,2 1 Department
PROSTATE CANCER DETECTION USING LABEL IMAGE CONSTRAINED MULTIATLAS SELECTION
 PROSTATE CANCER DETECTION USING LABEL IMAGE CONSTRAINED MULTIATLAS SELECTION Ms. Vaibhavi Nandkumar Jagtap 1, Mr. Santosh D. Kale 2 1 PG Scholar, 2 Assistant Professor, Department of Electronics and Telecommunication,
PROSTATE CANCER DETECTION USING LABEL IMAGE CONSTRAINED MULTIATLAS SELECTION Ms. Vaibhavi Nandkumar Jagtap 1, Mr. Santosh D. Kale 2 1 PG Scholar, 2 Assistant Professor, Department of Electronics and Telecommunication,
Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging
 1 CS 9 Final Project Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging Feiyu Chen Department of Electrical Engineering ABSTRACT Subject motion is a significant
1 CS 9 Final Project Classification of Subject Motion for Improved Reconstruction of Dynamic Magnetic Resonance Imaging Feiyu Chen Department of Electrical Engineering ABSTRACT Subject motion is a significant
2D Rigid Registration of MR Scans using the 1d Binary Projections
 2D Rigid Registration of MR Scans using the 1d Binary Projections Panos D. Kotsas Abstract This paper presents the application of a signal intensity independent registration criterion for 2D rigid body
2D Rigid Registration of MR Scans using the 1d Binary Projections Panos D. Kotsas Abstract This paper presents the application of a signal intensity independent registration criterion for 2D rigid body
NIH Public Access Author Manuscript Proc Soc Photo Opt Instrum Eng. Author manuscript; available in PMC 2014 October 07.
 NIH Public Access Author Manuscript Published in final edited form as: Proc Soc Photo Opt Instrum Eng. 2014 March 21; 9034: 903442. doi:10.1117/12.2042915. MRI Brain Tumor Segmentation and Necrosis Detection
NIH Public Access Author Manuscript Published in final edited form as: Proc Soc Photo Opt Instrum Eng. 2014 March 21; 9034: 903442. doi:10.1117/12.2042915. MRI Brain Tumor Segmentation and Necrosis Detection
Methodological progress in image registration for ventilation estimation, segmentation propagation and multi-modal fusion
 Methodological progress in image registration for ventilation estimation, segmentation propagation and multi-modal fusion Mattias P. Heinrich Julia A. Schnabel, Mark Jenkinson, Sir Michael Brady 2 Clinical
Methodological progress in image registration for ventilation estimation, segmentation propagation and multi-modal fusion Mattias P. Heinrich Julia A. Schnabel, Mark Jenkinson, Sir Michael Brady 2 Clinical
MARS: Multiple Atlases Robust Segmentation
 Software Release (1.0.1) Last updated: April 30, 2014. MARS: Multiple Atlases Robust Segmentation Guorong Wu, Minjeong Kim, Gerard Sanroma, and Dinggang Shen {grwu, mjkim, gerard_sanroma, dgshen}@med.unc.edu
Software Release (1.0.1) Last updated: April 30, 2014. MARS: Multiple Atlases Robust Segmentation Guorong Wu, Minjeong Kim, Gerard Sanroma, and Dinggang Shen {grwu, mjkim, gerard_sanroma, dgshen}@med.unc.edu
GLIRT: Groupwise and Longitudinal Image Registration Toolbox
 Software Release (1.0.1) Last updated: March. 30, 2011. GLIRT: Groupwise and Longitudinal Image Registration Toolbox Guorong Wu 1, Qian Wang 1,2, Hongjun Jia 1, and Dinggang Shen 1 1 Image Display, Enhancement,
Software Release (1.0.1) Last updated: March. 30, 2011. GLIRT: Groupwise and Longitudinal Image Registration Toolbox Guorong Wu 1, Qian Wang 1,2, Hongjun Jia 1, and Dinggang Shen 1 1 Image Display, Enhancement,
Neuroimaging and mathematical modelling Lesson 2: Voxel Based Morphometry
 Neuroimaging and mathematical modelling Lesson 2: Voxel Based Morphometry Nivedita Agarwal, MD Nivedita.agarwal@apss.tn.it Nivedita.agarwal@unitn.it Volume and surface morphometry Brain volume White matter
Neuroimaging and mathematical modelling Lesson 2: Voxel Based Morphometry Nivedita Agarwal, MD Nivedita.agarwal@apss.tn.it Nivedita.agarwal@unitn.it Volume and surface morphometry Brain volume White matter
Hybrid Spline-based Multimodal Registration using a Local Measure for Mutual Information
 Hybrid Spline-based Multimodal Registration using a Local Measure for Mutual Information Andreas Biesdorf 1, Stefan Wörz 1, Hans-Jürgen Kaiser 2, Karl Rohr 1 1 University of Heidelberg, BIOQUANT, IPMB,
Hybrid Spline-based Multimodal Registration using a Local Measure for Mutual Information Andreas Biesdorf 1, Stefan Wörz 1, Hans-Jürgen Kaiser 2, Karl Rohr 1 1 University of Heidelberg, BIOQUANT, IPMB,
Automatic Segmentation of Parotids from CT Scans Using Multiple Atlases
 Automatic Segmentation of Parotids from CT Scans Using Multiple Atlases Jinzhong Yang, Yongbin Zhang, Lifei Zhang, and Lei Dong Department of Radiation Physics, University of Texas MD Anderson Cancer Center
Automatic Segmentation of Parotids from CT Scans Using Multiple Atlases Jinzhong Yang, Yongbin Zhang, Lifei Zhang, and Lei Dong Department of Radiation Physics, University of Texas MD Anderson Cancer Center
Subcortical Structure Segmentation using Probabilistic Atlas Priors
 Subcortical Structure Segmentation using Probabilistic Atlas Priors Sylvain Gouttard a, Martin Styner a,b, Sarang Joshi c, Rachel G. Smith a, Heather Cody a, Guido Gerig a,b a Department of Psychiatry,
Subcortical Structure Segmentation using Probabilistic Atlas Priors Sylvain Gouttard a, Martin Styner a,b, Sarang Joshi c, Rachel G. Smith a, Heather Cody a, Guido Gerig a,b a Department of Psychiatry,
MR IMAGE SEGMENTATION
 MR IMAGE SEGMENTATION Prepared by : Monil Shah What is Segmentation? Partitioning a region or regions of interest in images such that each region corresponds to one or more anatomic structures Classification
MR IMAGE SEGMENTATION Prepared by : Monil Shah What is Segmentation? Partitioning a region or regions of interest in images such that each region corresponds to one or more anatomic structures Classification
Measuring longitudinal brain changes in humans and small animal models. Christos Davatzikos
 Measuring longitudinal brain changes in humans and small animal models Christos Davatzikos Section of Biomedical Image Analysis University of Pennsylvania (Radiology) http://www.rad.upenn.edu/sbia Computational
Measuring longitudinal brain changes in humans and small animal models Christos Davatzikos Section of Biomedical Image Analysis University of Pennsylvania (Radiology) http://www.rad.upenn.edu/sbia Computational
Multi-Atlas Segmentation of the Cardiac MR Right Ventricle
 Multi-Atlas Segmentation of the Cardiac MR Right Ventricle Yangming Ou, Jimit Doshi, Guray Erus, and Christos Davatzikos Section of Biomedical Image Analysis (SBIA) Department of Radiology, University
Multi-Atlas Segmentation of the Cardiac MR Right Ventricle Yangming Ou, Jimit Doshi, Guray Erus, and Christos Davatzikos Section of Biomedical Image Analysis (SBIA) Department of Radiology, University
3-D Compounding of B-Scan Ultrasound Images
 3-D Compounding of B-Scan Ultrasound Images Jochen F. Krücker, Charles R. Meyer, Theresa A. Tuthill, Gerald L. LeCarpentier, J. Brian Fowlkes, Paul L. Carson University of Michigan, Dept. of Radiology,
3-D Compounding of B-Scan Ultrasound Images Jochen F. Krücker, Charles R. Meyer, Theresa A. Tuthill, Gerald L. LeCarpentier, J. Brian Fowlkes, Paul L. Carson University of Michigan, Dept. of Radiology,
Fmri Spatial Processing
 Educational Course: Fmri Spatial Processing Ray Razlighi Jun. 8, 2014 Spatial Processing Spatial Re-alignment Geometric distortion correction Spatial Normalization Smoothing Why, When, How, Which Why is
Educational Course: Fmri Spatial Processing Ray Razlighi Jun. 8, 2014 Spatial Processing Spatial Re-alignment Geometric distortion correction Spatial Normalization Smoothing Why, When, How, Which Why is
Whole Body MRI Intensity Standardization
 Whole Body MRI Intensity Standardization Florian Jäger 1, László Nyúl 1, Bernd Frericks 2, Frank Wacker 2 and Joachim Hornegger 1 1 Institute of Pattern Recognition, University of Erlangen, {jaeger,nyul,hornegger}@informatik.uni-erlangen.de
Whole Body MRI Intensity Standardization Florian Jäger 1, László Nyúl 1, Bernd Frericks 2, Frank Wacker 2 and Joachim Hornegger 1 1 Institute of Pattern Recognition, University of Erlangen, {jaeger,nyul,hornegger}@informatik.uni-erlangen.de
City, University of London Institutional Repository
 City Research Online City, University of London Institutional Repository Citation: Doan, H., Slabaugh, G.G., Unal, G.B. & Fang, T. (2006). Semi-Automatic 3-D Segmentation of Anatomical Structures of Brain
City Research Online City, University of London Institutional Repository Citation: Doan, H., Slabaugh, G.G., Unal, G.B. & Fang, T. (2006). Semi-Automatic 3-D Segmentation of Anatomical Structures of Brain
Distance Transforms in Multi Channel MR Image Registration
 Distance Transforms in Multi Channel MR Image Registration Min Chen 1, Aaron Carass 1, John Bogovic 1, Pierre-Louis Bazin 2 and Jerry L. Prince 1 1 Image Analysis and Communications Laboratory, 2 The Laboratory
Distance Transforms in Multi Channel MR Image Registration Min Chen 1, Aaron Carass 1, John Bogovic 1, Pierre-Louis Bazin 2 and Jerry L. Prince 1 1 Image Analysis and Communications Laboratory, 2 The Laboratory
LOCAL-GLOBAL OPTICAL FLOW FOR IMAGE REGISTRATION
 LOCAL-GLOBAL OPTICAL FLOW FOR IMAGE REGISTRATION Ammar Zayouna Richard Comley Daming Shi Middlesex University School of Engineering and Information Sciences Middlesex University, London NW4 4BT, UK A.Zayouna@mdx.ac.uk
LOCAL-GLOBAL OPTICAL FLOW FOR IMAGE REGISTRATION Ammar Zayouna Richard Comley Daming Shi Middlesex University School of Engineering and Information Sciences Middlesex University, London NW4 4BT, UK A.Zayouna@mdx.ac.uk
Learning-based Neuroimage Registration
 Learning-based Neuroimage Registration Leonid Teverovskiy and Yanxi Liu 1 October 2004 CMU-CALD-04-108, CMU-RI-TR-04-59 School of Computer Science Carnegie Mellon University Pittsburgh, PA 15213 Abstract
Learning-based Neuroimage Registration Leonid Teverovskiy and Yanxi Liu 1 October 2004 CMU-CALD-04-108, CMU-RI-TR-04-59 School of Computer Science Carnegie Mellon University Pittsburgh, PA 15213 Abstract
Auto-contouring the Prostate for Online Adaptive Radiotherapy
 Auto-contouring the Prostate for Online Adaptive Radiotherapy Yan Zhou 1 and Xiao Han 1 Elekta Inc., Maryland Heights, MO, USA yan.zhou@elekta.com, xiao.han@elekta.com, Abstract. Among all the organs under
Auto-contouring the Prostate for Online Adaptive Radiotherapy Yan Zhou 1 and Xiao Han 1 Elekta Inc., Maryland Heights, MO, USA yan.zhou@elekta.com, xiao.han@elekta.com, Abstract. Among all the organs under
STIC AmSud Project. Graph cut based segmentation of cardiac ventricles in MRI: a shape-prior based approach
 STIC AmSud Project Graph cut based segmentation of cardiac ventricles in MRI: a shape-prior based approach Caroline Petitjean A joint work with Damien Grosgeorge, Pr Su Ruan, Pr JN Dacher, MD October 22,
STIC AmSud Project Graph cut based segmentation of cardiac ventricles in MRI: a shape-prior based approach Caroline Petitjean A joint work with Damien Grosgeorge, Pr Su Ruan, Pr JN Dacher, MD October 22,
1 Introduction Motivation and Aims Functional Imaging Computational Neuroanatomy... 12
 Contents 1 Introduction 10 1.1 Motivation and Aims....... 10 1.1.1 Functional Imaging.... 10 1.1.2 Computational Neuroanatomy... 12 1.2 Overview of Chapters... 14 2 Rigid Body Registration 18 2.1 Introduction.....
Contents 1 Introduction 10 1.1 Motivation and Aims....... 10 1.1.1 Functional Imaging.... 10 1.1.2 Computational Neuroanatomy... 12 1.2 Overview of Chapters... 14 2 Rigid Body Registration 18 2.1 Introduction.....
Where are we now? Structural MRI processing and analysis
 Where are we now? Structural MRI processing and analysis Pierre-Louis Bazin bazin@cbs.mpg.de Leipzig, Germany Structural MRI processing: why bother? Just use the standards? SPM FreeSurfer FSL However:
Where are we now? Structural MRI processing and analysis Pierre-Louis Bazin bazin@cbs.mpg.de Leipzig, Germany Structural MRI processing: why bother? Just use the standards? SPM FreeSurfer FSL However:
3D Brain Segmentation Using Active Appearance Models and Local Regressors
 3D Brain Segmentation Using Active Appearance Models and Local Regressors K.O. Babalola, T.F. Cootes, C.J. Twining, V. Petrovic, and C.J. Taylor Division of Imaging Science and Biomedical Engineering,
3D Brain Segmentation Using Active Appearance Models and Local Regressors K.O. Babalola, T.F. Cootes, C.J. Twining, V. Petrovic, and C.J. Taylor Division of Imaging Science and Biomedical Engineering,
Attribute Similarity and Mutual-Saliency Weighting for Registration and Label Fusion
 Attribute Similarity and Mutual-Saliency Weighting for Registration and Label Fusion Yangming Ou, Jimit Doshi, Guray Erus, and Christos Davatzikos Section of Biomedical Image Analysis (SBIA) Department
Attribute Similarity and Mutual-Saliency Weighting for Registration and Label Fusion Yangming Ou, Jimit Doshi, Guray Erus, and Christos Davatzikos Section of Biomedical Image Analysis (SBIA) Department
A Review on Label Image Constrained Multiatlas Selection
 A Review on Label Image Constrained Multiatlas Selection Ms. VAIBHAVI NANDKUMAR JAGTAP 1, Mr. SANTOSH D. KALE 2 1PG Scholar, Department of Electronics and Telecommunication, SVPM College of Engineering,
A Review on Label Image Constrained Multiatlas Selection Ms. VAIBHAVI NANDKUMAR JAGTAP 1, Mr. SANTOSH D. KALE 2 1PG Scholar, Department of Electronics and Telecommunication, SVPM College of Engineering,
Morphological Analysis of Brain Structures Using Spatial Normalization
 Morphological Analysis of Brain Structures Using Spatial Normalization C. Davatzikos 1, M. Vaillant 1, S. Resnick 2, J.L. Prince 3;1, S. Letovsky 1, and R.N. Bryan 1 1 Department of Radiology, Johns Hopkins
Morphological Analysis of Brain Structures Using Spatial Normalization C. Davatzikos 1, M. Vaillant 1, S. Resnick 2, J.L. Prince 3;1, S. Letovsky 1, and R.N. Bryan 1 1 Department of Radiology, Johns Hopkins
The Insight Toolkit. Image Registration Algorithms & Frameworks
 The Insight Toolkit Image Registration Algorithms & Frameworks Registration in ITK Image Registration Framework Multi Resolution Registration Framework Components PDE Based Registration FEM Based Registration
The Insight Toolkit Image Registration Algorithms & Frameworks Registration in ITK Image Registration Framework Multi Resolution Registration Framework Components PDE Based Registration FEM Based Registration
The organization of the human cerebral cortex estimated by intrinsic functional connectivity
 1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
Automatic Generation of Training Data for Brain Tissue Classification from MRI
 MICCAI-2002 1 Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. Cocosco, Alex P. Zijdenbos, and Alan C. Evans McConnell Brain Imaging Centre, Montreal Neurological
MICCAI-2002 1 Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. Cocosco, Alex P. Zijdenbos, and Alan C. Evans McConnell Brain Imaging Centre, Montreal Neurological
Integrated Approaches to Non-Rigid Registration in Medical Images
 Work. on Appl. of Comp. Vision, pg 102-108. 1 Integrated Approaches to Non-Rigid Registration in Medical Images Yongmei Wang and Lawrence H. Staib + Departments of Electrical Engineering and Diagnostic
Work. on Appl. of Comp. Vision, pg 102-108. 1 Integrated Approaches to Non-Rigid Registration in Medical Images Yongmei Wang and Lawrence H. Staib + Departments of Electrical Engineering and Diagnostic
MARS: Multiple Atlases Robust Segmentation
 Software Release (1.0.1) Last updated: April 30, 2014. MARS: Multiple Atlases Robust Segmentation Guorong Wu, Minjeong Kim, Gerard Sanroma, and Dinggang Shen {grwu, mjkim, gerard_sanroma, dgshen}@med.unc.edu
Software Release (1.0.1) Last updated: April 30, 2014. MARS: Multiple Atlases Robust Segmentation Guorong Wu, Minjeong Kim, Gerard Sanroma, and Dinggang Shen {grwu, mjkim, gerard_sanroma, dgshen}@med.unc.edu
Non-rigid Image Registration
 Overview Non-rigid Image Registration Introduction to image registration - he goal of image registration - Motivation for medical image registration - Classification of image registration - Nonrigid registration
Overview Non-rigid Image Registration Introduction to image registration - he goal of image registration - Motivation for medical image registration - Classification of image registration - Nonrigid registration
Probabilistic Registration of 3-D Medical Images
 Probabilistic Registration of 3-D Medical Images Mei Chen, Takeo Kanade, Dean Pomerleau, Jeff Schneider CMU-RI-TR-99-16 The Robotics Institute Carnegie Mellon University Pittsburgh, Pennsylvania 1513 July,
Probabilistic Registration of 3-D Medical Images Mei Chen, Takeo Kanade, Dean Pomerleau, Jeff Schneider CMU-RI-TR-99-16 The Robotics Institute Carnegie Mellon University Pittsburgh, Pennsylvania 1513 July,
Segmentation Using a Region Growing Thresholding
 Segmentation Using a Region Growing Thresholding Matei MANCAS 1, Bernard GOSSELIN 1, Benoît MACQ 2 1 Faculté Polytechnique de Mons, Circuit Theory and Signal Processing Laboratory Bâtiment MULTITEL/TCTS
Segmentation Using a Region Growing Thresholding Matei MANCAS 1, Bernard GOSSELIN 1, Benoît MACQ 2 1 Faculté Polytechnique de Mons, Circuit Theory and Signal Processing Laboratory Bâtiment MULTITEL/TCTS
Automated segmentation methods for liver analysis in oncology applications
 University of Szeged Department of Image Processing and Computer Graphics Automated segmentation methods for liver analysis in oncology applications Ph. D. Thesis László Ruskó Thesis Advisor Dr. Antal
University of Szeged Department of Image Processing and Computer Graphics Automated segmentation methods for liver analysis in oncology applications Ph. D. Thesis László Ruskó Thesis Advisor Dr. Antal
2 Michael E. Leventon and Sarah F. F. Gibson a b c d Fig. 1. (a, b) Two MR scans of a person's knee. Both images have high resolution in-plane, but ha
 Model Generation from Multiple Volumes using Constrained Elastic SurfaceNets Michael E. Leventon and Sarah F. F. Gibson 1 MIT Artificial Intelligence Laboratory, Cambridge, MA 02139, USA leventon@ai.mit.edu
Model Generation from Multiple Volumes using Constrained Elastic SurfaceNets Michael E. Leventon and Sarah F. F. Gibson 1 MIT Artificial Intelligence Laboratory, Cambridge, MA 02139, USA leventon@ai.mit.edu
Segmentation of 3D CT Volume Images Using a Single 2D Atlas
 Segmentation of 3D CT Volume Images Using a Single 2D Atlas Feng Ding 1, Wee Kheng Leow 1, and Shih-Chang Wang 2 1 Dept. of Computer Science, National University of Singapore, 3 Science Drive 2, Singapore
Segmentation of 3D CT Volume Images Using a Single 2D Atlas Feng Ding 1, Wee Kheng Leow 1, and Shih-Chang Wang 2 1 Dept. of Computer Science, National University of Singapore, 3 Science Drive 2, Singapore
Assessment of Reliability of Multi-site Neuroimaging via Traveling Phantom Study
 Assessment of Reliability of Multi-site Neuroimaging via Traveling Phantom Study Sylvain Gouttard 1, Martin Styner 2,3, Marcel Prastawa 1, Joseph Piven 3, and Guido Gerig 1 1 Scientific Computing and Imaging
Assessment of Reliability of Multi-site Neuroimaging via Traveling Phantom Study Sylvain Gouttard 1, Martin Styner 2,3, Marcel Prastawa 1, Joseph Piven 3, and Guido Gerig 1 1 Scientific Computing and Imaging
Introduction to Medical Image Processing
 Introduction to Medical Image Processing Δ Essential environments of a medical imaging system Subject Image Analysis Energy Imaging System Images Image Processing Feature Images Image processing may be
Introduction to Medical Image Processing Δ Essential environments of a medical imaging system Subject Image Analysis Energy Imaging System Images Image Processing Feature Images Image processing may be
Good Morning! Thank you for joining us
 Good Morning! Thank you for joining us Deformable Registration, Contour Propagation and Dose Mapping: 101 and 201 Marc Kessler, PhD, FAAPM The University of Michigan Conflict of Interest I receive direct
Good Morning! Thank you for joining us Deformable Registration, Contour Propagation and Dose Mapping: 101 and 201 Marc Kessler, PhD, FAAPM The University of Michigan Conflict of Interest I receive direct
Incorporating Non-rigid Registration into Expectation Maximization Algorithm to Segment MR Images
 Incorporating Non-rigid Registration into Expectation Maximization Algorithm to Segment MR Images Kilian M Pohl 1, William M. Wells 2, Alexandre Guimond 3, Kiyoto Kasai 4, Martha E. Shenton 2,4, Ron Kikinis
Incorporating Non-rigid Registration into Expectation Maximization Algorithm to Segment MR Images Kilian M Pohl 1, William M. Wells 2, Alexandre Guimond 3, Kiyoto Kasai 4, Martha E. Shenton 2,4, Ron Kikinis
Simultaneous Model-based Segmentation of Multiple Objects
 Simultaneous Model-based Segmentation of Multiple Objects Astrid Franz 1, Robin Wolz 1, Tobias Klinder 1,2, Cristian Lorenz 1, Hans Barschdorf 1, Thomas Blaffert 1, Sebastian P. M. Dries 1, Steffen Renisch
Simultaneous Model-based Segmentation of Multiple Objects Astrid Franz 1, Robin Wolz 1, Tobias Klinder 1,2, Cristian Lorenz 1, Hans Barschdorf 1, Thomas Blaffert 1, Sebastian P. M. Dries 1, Steffen Renisch
Non-rigid Image Registration using Electric Current Flow
 Non-rigid Image Registration using Electric Current Flow Shu Liao, Max W. K. Law and Albert C. S. Chung Lo Kwee-Seong Medical Image Analysis Laboratory, Department of Computer Science and Engineering,
Non-rigid Image Registration using Electric Current Flow Shu Liao, Max W. K. Law and Albert C. S. Chung Lo Kwee-Seong Medical Image Analysis Laboratory, Department of Computer Science and Engineering,
Medical Image Segmentation Using Descriptive Image Features
 YANG, YUAN, LI, YAN: MEDICAL IMAGE SEGMENTATION 1 Medical Image Segmentation Using Descriptive Image Features Meijuan Yang meijuan.yang@opt.ac.cn Yuan Yuan yuany@opt.ac.cn Xuelong Li xuelong_li@opt.ac.cn
YANG, YUAN, LI, YAN: MEDICAL IMAGE SEGMENTATION 1 Medical Image Segmentation Using Descriptive Image Features Meijuan Yang meijuan.yang@opt.ac.cn Yuan Yuan yuany@opt.ac.cn Xuelong Li xuelong_li@opt.ac.cn
Comparison Study of Clinical 3D MRI Brain Segmentation Evaluation
 Comparison Study of Clinical 3D MRI Brain Segmentation Evaluation Ting Song 1, Elsa D. Angelini 2, Brett D. Mensh 3, Andrew Laine 1 1 Heffner Biomedical Imaging Laboratory Department of Biomedical Engineering,
Comparison Study of Clinical 3D MRI Brain Segmentation Evaluation Ting Song 1, Elsa D. Angelini 2, Brett D. Mensh 3, Andrew Laine 1 1 Heffner Biomedical Imaging Laboratory Department of Biomedical Engineering,
Automatic MS Lesion Segmentation by Outlier Detection and Information Theoretic Region Partitioning Release 0.00
 Automatic MS Lesion Segmentation by Outlier Detection and Information Theoretic Region Partitioning Release 0.00 Marcel Prastawa 1 and Guido Gerig 1 Abstract July 17, 2008 1 Scientific Computing and Imaging
Automatic MS Lesion Segmentation by Outlier Detection and Information Theoretic Region Partitioning Release 0.00 Marcel Prastawa 1 and Guido Gerig 1 Abstract July 17, 2008 1 Scientific Computing and Imaging
Functional MRI in Clinical Research and Practice Preprocessing
 Functional MRI in Clinical Research and Practice Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization
Functional MRI in Clinical Research and Practice Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization
Elastic Registration with Partial Data
 Elastic Registration with Partial Data Senthil Periaswamy and Hany Farid Dartmouth College, Hanover, NH, 03755, USA Abstract. We have developed a general purpose registration algorithm for medical images
Elastic Registration with Partial Data Senthil Periaswamy and Hany Farid Dartmouth College, Hanover, NH, 03755, USA Abstract. We have developed a general purpose registration algorithm for medical images
Correspondence Detection Using Wavelet-Based Attribute Vectors
 Correspondence Detection Using Wavelet-Based Attribute Vectors Zhong Xue, Dinggang Shen, and Christos Davatzikos Section of Biomedical Image Analysis, Department of Radiology University of Pennsylvania,
Correspondence Detection Using Wavelet-Based Attribute Vectors Zhong Xue, Dinggang Shen, and Christos Davatzikos Section of Biomedical Image Analysis, Department of Radiology University of Pennsylvania,
SPM8 for Basic and Clinical Investigators. Preprocessing. fmri Preprocessing
 SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
Occluded Facial Expression Tracking
 Occluded Facial Expression Tracking Hugo Mercier 1, Julien Peyras 2, and Patrice Dalle 1 1 Institut de Recherche en Informatique de Toulouse 118, route de Narbonne, F-31062 Toulouse Cedex 9 2 Dipartimento
Occluded Facial Expression Tracking Hugo Mercier 1, Julien Peyras 2, and Patrice Dalle 1 1 Institut de Recherche en Informatique de Toulouse 118, route de Narbonne, F-31062 Toulouse Cedex 9 2 Dipartimento
Assessment of Reliability of Multi-site Neuroimaging Via Traveling Phantom Study
 Assessment of Reliability of Multi-site Neuroimaging Via Traveling Phantom Study Sylvain Gouttard 1, Martin Styner 2,3, Marcel Prastawa 1, Joseph Piven 3, and Guido Gerig 1 1 Scientific Computing and Imaging
Assessment of Reliability of Multi-site Neuroimaging Via Traveling Phantom Study Sylvain Gouttard 1, Martin Styner 2,3, Marcel Prastawa 1, Joseph Piven 3, and Guido Gerig 1 1 Scientific Computing and Imaging
Retrospective evaluation of inter-subject brain registration
 Retrospective evaluation of inter-subject brain registration P. Hellier 1, C. Barillot 1, I. Corouge 1,B.Gibaud 2, G. Le Goualher 2,3, L. Collins 3, A. Evans 3, G. Malandain 4, N. Ayache 4 1 Projet Vista,
Retrospective evaluation of inter-subject brain registration P. Hellier 1, C. Barillot 1, I. Corouge 1,B.Gibaud 2, G. Le Goualher 2,3, L. Collins 3, A. Evans 3, G. Malandain 4, N. Ayache 4 1 Projet Vista,
An Object-Based Volumetric Deformable Atlas for the Improved Localization of Neuroanatomy in MR Images
 An Object-Based Volumetric Deformable Atlas for the Improved Localization of Neuroanatomy in MR Images Tim McInerney 1,2 and Ron Kikinis 3 1 University of Toronto, Toronto, ON, Canada M5S 3H5 2 Massachusetts
An Object-Based Volumetric Deformable Atlas for the Improved Localization of Neuroanatomy in MR Images Tim McInerney 1,2 and Ron Kikinis 3 1 University of Toronto, Toronto, ON, Canada M5S 3H5 2 Massachusetts
Quantitative MRI of the Brain: Investigation of Cerebral Gray and White Matter Diseases
 Quantities Measured by MR - Quantitative MRI of the Brain: Investigation of Cerebral Gray and White Matter Diseases Static parameters (influenced by molecular environment): T, T* (transverse relaxation)
Quantities Measured by MR - Quantitative MRI of the Brain: Investigation of Cerebral Gray and White Matter Diseases Static parameters (influenced by molecular environment): T, T* (transverse relaxation)
Scene-Based Segmentation of Multiple Muscles from MRI in MITK
 Scene-Based Segmentation of Multiple Muscles from MRI in MITK Yan Geng 1, Sebastian Ullrich 2, Oliver Grottke 3, Rolf Rossaint 3, Torsten Kuhlen 2, Thomas M. Deserno 1 1 Department of Medical Informatics,
Scene-Based Segmentation of Multiple Muscles from MRI in MITK Yan Geng 1, Sebastian Ullrich 2, Oliver Grottke 3, Rolf Rossaint 3, Torsten Kuhlen 2, Thomas M. Deserno 1 1 Department of Medical Informatics,
Medical Image Registration by Maximization of Mutual Information
 Medical Image Registration by Maximization of Mutual Information EE 591 Introduction to Information Theory Instructor Dr. Donald Adjeroh Submitted by Senthil.P.Ramamurthy Damodaraswamy, Umamaheswari Introduction
Medical Image Registration by Maximization of Mutual Information EE 591 Introduction to Information Theory Instructor Dr. Donald Adjeroh Submitted by Senthil.P.Ramamurthy Damodaraswamy, Umamaheswari Introduction
SPM8 for Basic and Clinical Investigators. Preprocessing
 SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
SPM8 for Basic and Clinical Investigators Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial
Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm
 Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm Lin Chen and Gudrun Wagenknecht Central Institute for Electronics, Research Center Jülich, Jülich, Germany Email: l.chen@fz-juelich.de
Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm Lin Chen and Gudrun Wagenknecht Central Institute for Electronics, Research Center Jülich, Jülich, Germany Email: l.chen@fz-juelich.de
Elastic Registration in the Presence of Intensity Variations
 IEEE Transactions on Medical Imaging, 22(7):865-874, 2003 Elastic Registration in the Presence of Intensity Variations Senthil Periaswamy 1 and Hany Farid 1,2 1 Department of Computer Science and 2 Center
IEEE Transactions on Medical Imaging, 22(7):865-874, 2003 Elastic Registration in the Presence of Intensity Variations Senthil Periaswamy 1 and Hany Farid 1,2 1 Department of Computer Science and 2 Center
doi: /
 Yiting Xie ; Anthony P. Reeves; Single 3D cell segmentation from optical CT microscope images. Proc. SPIE 934, Medical Imaging 214: Image Processing, 9343B (March 21, 214); doi:1.1117/12.243852. (214)
Yiting Xie ; Anthony P. Reeves; Single 3D cell segmentation from optical CT microscope images. Proc. SPIE 934, Medical Imaging 214: Image Processing, 9343B (March 21, 214); doi:1.1117/12.243852. (214)
EPI Data Are Acquired Serially. EPI Data Are Acquired Serially 10/23/2011. Functional Connectivity Preprocessing. fmri Preprocessing
 Functional Connectivity Preprocessing Geometric distortion Head motion Geometric distortion Head motion EPI Data Are Acquired Serially EPI Data Are Acquired Serially descending 1 EPI Data Are Acquired
Functional Connectivity Preprocessing Geometric distortion Head motion Geometric distortion Head motion EPI Data Are Acquired Serially EPI Data Are Acquired Serially descending 1 EPI Data Are Acquired
Basic fmri Design and Analysis. Preprocessing
 Basic fmri Design and Analysis Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial filtering
Basic fmri Design and Analysis Preprocessing fmri Preprocessing Slice timing correction Geometric distortion correction Head motion correction Temporal filtering Intensity normalization Spatial filtering
3D Surface Reconstruction of the Brain based on Level Set Method
 3D Surface Reconstruction of the Brain based on Level Set Method Shijun Tang, Bill P. Buckles, and Kamesh Namuduri Department of Computer Science & Engineering Department of Electrical Engineering University
3D Surface Reconstruction of the Brain based on Level Set Method Shijun Tang, Bill P. Buckles, and Kamesh Namuduri Department of Computer Science & Engineering Department of Electrical Engineering University
Auto-Segmentation Using Deformable Image Registration. Disclosure. Objectives 8/4/2011
 Auto-Segmentation Using Deformable Image Registration Lei Dong, Ph.D. Dept. of Radiation Physics University of Texas MD Anderson Cancer Center, Houston, Texas AAPM Therapy Educational Course Aug. 4th 2011
Auto-Segmentation Using Deformable Image Registration Lei Dong, Ph.D. Dept. of Radiation Physics University of Texas MD Anderson Cancer Center, Houston, Texas AAPM Therapy Educational Course Aug. 4th 2011
A MORPHOLOGY-BASED FILTER STRUCTURE FOR EDGE-ENHANCING SMOOTHING
 Proceedings of the 1994 IEEE International Conference on Image Processing (ICIP-94), pp. 530-534. (Austin, Texas, 13-16 November 1994.) A MORPHOLOGY-BASED FILTER STRUCTURE FOR EDGE-ENHANCING SMOOTHING
Proceedings of the 1994 IEEE International Conference on Image Processing (ICIP-94), pp. 530-534. (Austin, Texas, 13-16 November 1994.) A MORPHOLOGY-BASED FILTER STRUCTURE FOR EDGE-ENHANCING SMOOTHING
Shape-based Diffeomorphic Registration on Hippocampal Surfaces Using Beltrami Holomorphic Flow
 Shape-based Diffeomorphic Registration on Hippocampal Surfaces Using Beltrami Holomorphic Flow Abstract. Finding meaningful 1-1 correspondences between hippocampal (HP) surfaces is an important but difficult
Shape-based Diffeomorphic Registration on Hippocampal Surfaces Using Beltrami Holomorphic Flow Abstract. Finding meaningful 1-1 correspondences between hippocampal (HP) surfaces is an important but difficult
Open Topology: A Toolkit for Brain Isosurface Correction
 Open Topology: A Toolkit for Brain Isosurface Correction Sylvain Jaume 1, Patrice Rondao 2, and Benoît Macq 2 1 National Institute of Research in Computer Science and Control, INRIA, France, sylvain@mit.edu,
Open Topology: A Toolkit for Brain Isosurface Correction Sylvain Jaume 1, Patrice Rondao 2, and Benoît Macq 2 1 National Institute of Research in Computer Science and Control, INRIA, France, sylvain@mit.edu,
Automatic Optimization of Segmentation Algorithms Through Simultaneous Truth and Performance Level Estimation (STAPLE)
 Automatic Optimization of Segmentation Algorithms Through Simultaneous Truth and Performance Level Estimation (STAPLE) Mahnaz Maddah, Kelly H. Zou, William M. Wells, Ron Kikinis, and Simon K. Warfield
Automatic Optimization of Segmentation Algorithms Through Simultaneous Truth and Performance Level Estimation (STAPLE) Mahnaz Maddah, Kelly H. Zou, William M. Wells, Ron Kikinis, and Simon K. Warfield
An Automated Image-based Method for Multi-Leaf Collimator Positioning Verification in Intensity Modulated Radiation Therapy
 An Automated Image-based Method for Multi-Leaf Collimator Positioning Verification in Intensity Modulated Radiation Therapy Chenyang Xu 1, Siemens Corporate Research, Inc., Princeton, NJ, USA Xiaolei Huang,
An Automated Image-based Method for Multi-Leaf Collimator Positioning Verification in Intensity Modulated Radiation Therapy Chenyang Xu 1, Siemens Corporate Research, Inc., Princeton, NJ, USA Xiaolei Huang,
Automatic Generation of Shape Models Using Nonrigid Registration with a Single Segmented Template Mesh
 Automatic Generation of Shape Models Using Nonrigid Registration with a Single Segmented Template Mesh Geremy Heitz, Torsten Rohlfing, and Calvin R. Maurer, Jr. Image Guidance Laboratories Department of
Automatic Generation of Shape Models Using Nonrigid Registration with a Single Segmented Template Mesh Geremy Heitz, Torsten Rohlfing, and Calvin R. Maurer, Jr. Image Guidance Laboratories Department of
Anatomy-Preserving Nonlinear Registration of Deep Brain ROIs Using Confidence-Based Block-Matching
 Anatomy-Preserving Nonlinear Registration of Deep Brain ROIs Using Confidence-Based Block-Matching Manik Bhattacharjee 1,2, Alain Pitiot 3,AlexisRoche 4, Didier Dormont 1,2, and Eric Bardinet 1,2,5 1 CNRS
Anatomy-Preserving Nonlinear Registration of Deep Brain ROIs Using Confidence-Based Block-Matching Manik Bhattacharjee 1,2, Alain Pitiot 3,AlexisRoche 4, Didier Dormont 1,2, and Eric Bardinet 1,2,5 1 CNRS
Image Registration I
 Image Registration I Comp 254 Spring 2002 Guido Gerig Image Registration: Motivation Motivation for Image Registration Combine images from different modalities (multi-modality registration), e.g. CT&MRI,
Image Registration I Comp 254 Spring 2002 Guido Gerig Image Registration: Motivation Motivation for Image Registration Combine images from different modalities (multi-modality registration), e.g. CT&MRI,
Applications of Elastic Functional and Shape Data Analysis
 Applications of Elastic Functional and Shape Data Analysis Quick Outline: 1. Functional Data 2. Shapes of Curves 3. Shapes of Surfaces BAYESIAN REGISTRATION MODEL Bayesian Model + Riemannian Geometry +
Applications of Elastic Functional and Shape Data Analysis Quick Outline: 1. Functional Data 2. Shapes of Curves 3. Shapes of Surfaces BAYESIAN REGISTRATION MODEL Bayesian Model + Riemannian Geometry +
Deformable Image Registration, Contour Propagation and Dose Mapping: 101 and 201. Please do not (re)redistribute
 Deformable Registration, Contour Deformable Registration, Contour Propagation and Dose Mapping: 101 and 201 Marc Kessler, PhD The University of Michigan Jean Pouliot, PhD University of California Learning
Deformable Registration, Contour Deformable Registration, Contour Propagation and Dose Mapping: 101 and 201 Marc Kessler, PhD The University of Michigan Jean Pouliot, PhD University of California Learning
Deformable Segmentation using Sparse Shape Representation. Shaoting Zhang
 Deformable Segmentation using Sparse Shape Representation Shaoting Zhang Introduction Outline Our methods Segmentation framework Sparse shape representation Applications 2D lung localization in X-ray 3D
Deformable Segmentation using Sparse Shape Representation Shaoting Zhang Introduction Outline Our methods Segmentation framework Sparse shape representation Applications 2D lung localization in X-ray 3D
Digital Volume Correlation for Materials Characterization
 19 th World Conference on Non-Destructive Testing 2016 Digital Volume Correlation for Materials Characterization Enrico QUINTANA, Phillip REU, Edward JIMENEZ, Kyle THOMPSON, Sharlotte KRAMER Sandia National
19 th World Conference on Non-Destructive Testing 2016 Digital Volume Correlation for Materials Characterization Enrico QUINTANA, Phillip REU, Edward JIMENEZ, Kyle THOMPSON, Sharlotte KRAMER Sandia National
Adaptive Fuzzy Connectedness-Based Medical Image Segmentation
 Adaptive Fuzzy Connectedness-Based Medical Image Segmentation Amol Pednekar Ioannis A. Kakadiaris Uday Kurkure Visual Computing Lab, Dept. of Computer Science, Univ. of Houston, Houston, TX, USA apedneka@bayou.uh.edu
Adaptive Fuzzy Connectedness-Based Medical Image Segmentation Amol Pednekar Ioannis A. Kakadiaris Uday Kurkure Visual Computing Lab, Dept. of Computer Science, Univ. of Houston, Houston, TX, USA apedneka@bayou.uh.edu
A Multiple-Layer Flexible Mesh Template Matching Method for Nonrigid Registration between a Pelvis Model and CT Images
 A Multiple-Layer Flexible Mesh Template Matching Method for Nonrigid Registration between a Pelvis Model and CT Images Jianhua Yao 1, Russell Taylor 2 1. Diagnostic Radiology Department, Clinical Center,
A Multiple-Layer Flexible Mesh Template Matching Method for Nonrigid Registration between a Pelvis Model and CT Images Jianhua Yao 1, Russell Taylor 2 1. Diagnostic Radiology Department, Clinical Center,
Segmenting Lesions in Multiple Sclerosis Patients James Chen, Jason Su
 Segmenting Lesions in Multiple Sclerosis Patients James Chen, Jason Su Radiologists and researchers spend countless hours tediously segmenting white matter lesions to diagnose and study brain diseases.
Segmenting Lesions in Multiple Sclerosis Patients James Chen, Jason Su Radiologists and researchers spend countless hours tediously segmenting white matter lesions to diagnose and study brain diseases.
Supplementary methods
 Supplementary methods This section provides additional technical details on the sample, the applied imaging and analysis steps and methods. Structural imaging Trained radiographers placed all participants
Supplementary methods This section provides additional technical details on the sample, the applied imaging and analysis steps and methods. Structural imaging Trained radiographers placed all participants
Multi-Atlas Brain MRI Segmentation with Multiway Cut
 Multi-Atlas Brain MRI Segmentation with Multiway Cut Duygu Sarikaya, Liang Zhao, and Jason J. Corso SUNY Buffalo, Computer Science and Engineering Department, 338 Davis Hall- Buffalo, New York, USA 14260-2500
Multi-Atlas Brain MRI Segmentation with Multiway Cut Duygu Sarikaya, Liang Zhao, and Jason J. Corso SUNY Buffalo, Computer Science and Engineering Department, 338 Davis Hall- Buffalo, New York, USA 14260-2500
Appearance-Based Segmentation of Medial Temporal Lobe Structures
 NeuroImage 17, 515 531 (2002) doi:10.1006/nimg.2002.1188 Appearance-Based Segmentation of Medial Temporal Lobe Structures S. Duchesne, 1 J. C. Pruessner, and D. L. Collins McConnell Brain Imaging Centre,
NeuroImage 17, 515 531 (2002) doi:10.1006/nimg.2002.1188 Appearance-Based Segmentation of Medial Temporal Lobe Structures S. Duchesne, 1 J. C. Pruessner, and D. L. Collins McConnell Brain Imaging Centre,
Deformetrica: a software for statistical analysis of anatomical shapes
 Deformetrica: a software for statistical analysis of anatomical shapes Alexandre Routier, Marcel Prastawa, Benjamin Charlier, Cédric Doucet, Joan Alexis Glaunès, Stanley Durrleman To cite this version:
Deformetrica: a software for statistical analysis of anatomical shapes Alexandre Routier, Marcel Prastawa, Benjamin Charlier, Cédric Doucet, Joan Alexis Glaunès, Stanley Durrleman To cite this version:
Fiber Selection from Diffusion Tensor Data based on Boolean Operators
 Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhof 1, G. Greiner 2, M. Buchfelder 3, C. Nimsky 4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
Fiber Selection from Diffusion Tensor Data based on Boolean Operators D. Merhof 1, G. Greiner 2, M. Buchfelder 3, C. Nimsky 4 1 Visual Computing, University of Konstanz, Konstanz, Germany 2 Computer Graphics
