Wavelet-Based Representation of Biological Shapes
|
|
|
- Jean Nicholson
- 6 years ago
- Views:
Transcription
1 Wavelet-Based Representation of Biological Shapes Bin Dong*, Yu Mao*, Ivo D. Dinov**, Zhuowen Tu**, Yonggang Shi**, Yalin Wang**, Arthur W. Toga** * Department of Mathematics, University of California, Los Angeles, CA ** Center for Computational Biology, Laboratory of Neuro Imaging 65 S. Charles Young Dr., #225, University of California, Los Angeles, Los Angeles, CA Abstract. Modeling, characterization and analysis of biological shape and form are important in many computational biology studies. Shape representation challenges span the spectrum from small scales (e.g., microarray imaging and protein structure) to the macro scale (e.g., neuroimaging of human brains). In this paper, we present a new approach for representation and analysis of biological shape using wavelets. We apply the new technique for multi-spectral shape decomposition and for studying shape variability between populations using brain cortical and subcortical surfaces. The wavelet-space induced shape representation allows us to study the multi-spectral nature of the shape geometry, topology and features. Our results are very promising and compared to the spherical-wavelets method generally provide a shape representation, which is more compact and allows utilization of diverse wavelet bases. Keywords. wavelets, shape, spherical harmonics, brain imaging, brain mapping. 1 Introduction 1.1 General Imaging, representation, geometric modeling and topologic characterization of shape and form are important components of Computational Biology. They apply across the vast length scales between genotypes to phenotypes, from the small scale of microarray imaging for genomics, to the larger scale of neuroimaging of human brains. Here we review the existent techniques and algorithms and present a new approach for representation and analysis of biological shape using wavelets. We apply the new method for multi-spectral shape decomposition and for studying shape variability between populations using brain cortical and subcortical surfaces. 1.2 Literature Review Recently, N. Hacker et al. used conformal mapping and spherical wavelets to analyze biological shapes (see [9, 10]). Their idea is first mapping the original shape onto a unit 2-sphere using a certain conformal mapping so that one obtains a - valued function f defined on the sphere; and then interpolate the function f onto the regular triangular mesh on the sphere (which is generated by recursively subdividing an icosahedron); and then finally, apply spherical wavelet transform to the interpolated function. The spherical wavelets they used were introduced by P. Schröder and W. Sweldens in [15], which were constructed using lifting scheme (see W. Sweldens [14] and F. Arandiga et. al. [1]). 2 In this paper, we also start from a - valued function f given by a certain mapping from to S. After that, we linearly interpolate the function f onto a triangular mesh, which is generated by recursively subdividing an octahedron in (not restricted on the sphere) and then transforming the mesh onto the sphere. This method was first introduced by E. Praun and H. Hoppe (see [1]) under the scope of computer graphics. The major advantage of it is that we can transform the subdivided octahedron to a unit square so that we obtain an image with - valued entries (which were called geometric image in [1]), and then we can apply traditional X-lets (e.g. wavelets, framelets, curvelets etc.) decomposition. In this way we have
2 plenty of good bases and frames to choose according the application we have. More details of this approach are given in the second part of this paper. 1. Brain Mapping Challenges Understanding the relationship between the structure and function of the human brain in vivo has been the driving motivation for many neurosciences research for centuries. The research efforts not only focus on studying normal development but also understanding alterations in various clinical populations including schizophrenia, Huntington's disease, Alzheimer's disease, Williams Syndrome, autism, stroke, chronic drug abuse, as well as pharmacological interventions. For instance, there are multiple studies underway to quantify the differences between the brain structure of schizophrenic patients and healthy individuals in different stages of this disease. Detection of these significant differences via neuroimaging studies is not only useful to elucidate the link between change in cognitive profile and change in brain structure, but also to improve diagnosis particularly in early stages of the disease. With the increasing interest in carrying out such studies with large numbers of subjects, there is a need for a unified framework for image segmentation to identify the structure of interest (e.g. caudate, ventricles, cerebral cortex, sulcal regions) (see Z. Tu et. al. [24]), and morphometric analysis which requires methods for shape representation, shape comparison, and change in shape measurement (see P. Thompson and A. Toga [20]). 1.4 Outline of the Paper The rest of the paper is organized as follows. Section 2 gives a detailed explanation of how we manage data, and how we build multiscaled representations for the shapes we have (i.e. hippocampus and cortical surfaces). We provide some experimental results in section : multiscaled sparse representation of shapes, curvature-like characterization of shapes using wavelet coefficients, and statistical analysis of healthy and unhealthy hippocampus/cortices. In section 4, we give discussions of the results in section and suggest possible future work. 2 Methods In this section we describe our test data and the specific approach we took to represent shape using wavelets, as well as the statistical analyses we carry on the wavelet-based shape decomposition to identify group, population, time or variation differences. 2.1 Data Cortical Models: Surface objects of normal subjects and Williams syndrome patients were used to explore the power of our method to synthesize the energy of the shape content in a few wavelet coefficients. The demographics of the population included age ( ), genders (approximately 50/50) and IQ scores, P. Thompson et. al. [21]. Nonbrain tissue (i.e., scalp, orbits) was removed from the images, and each image volume was resliced into a standard orientation who "tagged" 20 standardized anatomical landmarks in each subject's image data set that corresponded to the same 20 anatomical landmarks defined on the ICBM5 average brain (see Mazziotta et al. [12], Thompson et. al. [22]). Automated tissue segmentation was conducted for each volume data set to classify voxels as most representative of gray matter, white matter, CSF, or a background class (representing extracerebral voxels in the image) on the basis of signal intensity. The procedure fits a mixture of Gaussian distributions to the intensities in each image before assigning each voxel to the class with the highest probability, Shattuck et. al. [16]. Then each individual's cortical surface was extracted and three-dimensionally rendered using automated software, MacDonald [11]. Each resulting cortical surface was represented as a high-resolution mesh of 11,072 surface triangles spanning 65,56 surface points. Hippocampal surfaces: High-resolution MRI scans were acquired from 12 AD patients ages 68.4 ± 1.9 and 14 matched controls 71.4 ± 0.9, each scanned twice, 2.1 ± 0.4 years apart. D parametric mesh models of the left and right hippocampi and temporal horns were manually, Thompson et. al. [2]. For each scan, a radio frequency bias field correction algorithm eliminated intensity drifts due to scanner field inhomogeneity, using a histogram spline sharpening method, Sled et. al. [19]. Images were then normalized
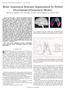
3 by transforming them to ICBM5 stereotaxic space, Evens et. al. [6], with automated image registration software, Collins et. al. []. To equalize image intensities across subjects, registered scans were histogramequalized. Each D surface is mapped to a unit sphere in R with 1-to-1 correspondence. For cortical surfaces, the conformal mapping method of Shi et. al. [17, 18] is used for hippocampal surfaces and that of Gu et. al. [8] (a) (b) (c) Fig. 1. Figure (a) is the shape we chose; (b) is the spherical function visualized as a spherical colored image; (c) is the geometric image. for cortical surfaces. For hippocampal surfaces, harmonic maps to the sphere are computed under the constraints of a set of automatically detected landmark curves (see [17, 18]). 2.2 Wavelet-Based Representation of Shapes From section 2.1, we got a function defined on the unit sphere S 2, which has vector values in R. However, the values of the function were only given on an irregular grid on the sphere. To apply the wavelet transform, we need to get the value of the function on a much more regular spherical grid. There are many approaches to get such kind of grids. The construction of the spherical mesh grid, sometimes called spherical triangular map, is an interesting subject itself (see e.g. Buss and Fillmore [2], and Praun (a) (b) (c) (d) (e) (f) (g) (h) Fig. 2. Two-levels wavelet decomposition: (a) and (e) are the low frequency coefficients of level one and two; (b)-(d) are wavelet coefficients of level 1 and band 1-; (f)-(h) are wavelet coefficients of level 2 and band 1-.
4 and Hoppe [1]). The basic idea is to start from a polyhedral base, which gives a simple but perfect grid on sphere, and then use some appropriate scheme to subdivide the mesh. A comparison of such techniques can be found in [1]. In our approach, we start from a recursive subdivision of the octahedral base. By mapping the subdivision grid onto the unit sphere, we get a regular grid structure on the sphere, and the function values on such a spherical grid can be obtained by linear interpolation. The reason we choose octahedron is that it can be unfolded to a plain image easily. Therefore, we can build a 1-1 map between a sphere and an image without too much distortion, and the data of a shape is transformed to a R - valued function defined on a plain image, which gives a geometric image (as shown in Figure 1). Since the mesh on the plain image is nothing but a Cartesian grid, a huge family of X-lets can be used to analyze properties of the geometric image. The wavelets that we shall use in the following experiments are Daubechies' Biorthogonal Wavelets [4]. We note that the boundary condition is a little complicated in this case. Topological saying, the two halves of each side of the image must be identified with each other. Thus, we need to setup corresponding boundary rules for the wavelet filters. Figure 2 shows how the decomposition is carried out to the geometric image we have. Since all low frequency and wavelet coefficients has x, y, and z three components, all coefficients are visualized as color images. 2. Multiscaled curvature-like characterization As shown in Figure 2 above, for each level and band of the wavelet coefficients, we have x, y, and z three components. The coefficient vectors reflect details of the shape at each position and scale. Indeed, the wavelet vectors can be treated as the displacement between the observed position and the predicted position calculated from the convolution of the wavelet filter and the scale coefficients of the neighboring vertices. Therefore, the direction of the wavelet vector gives us some information of the local geometric properties. For example, if we consider the wavelet vector at a local sunken area, then the approximated position interpolated from the neighboring vertices should be outer than the observed vertex, which means that the wavelet vector is pointing outwards. However, we cannot tell the geometric property of the shape from the wavelet coefficients directly, and a single wavelet coefficient itself is geometrically meaningless, so we combine the three components by calculating the inner product of the wavelet vector and the normal vector of the shape. As what we explained above, these inner products reflect geometric property at the corresponding position. In this way, a so called multiscaled curvature-like characterization of the shape is obtained: For given scale (or level), we compute normal of the shape at all positions under that scale and take the inner product of the normal with the wavelet coefficient vector. We then obtain a set of curvature-like coefficients within each level and band. The statistical analysis given in Section.2 is based on this representation. Figure shows how this representation can be used to find cortical sulci and gyri. (a) Fig.. Red regions in figure (a) and (b) are the sulcal and girial regions of a given cortex respectively. (b)
5 Results.1 Multiscaled Sparse Representation One of the most important properties of traditional wavelet transform is that it gives a multiscaled representation of the underlying function and the representation is sparse. We now show that our method as discussed in the previous section also gives a multiscaled sparse representation for the biological shapes we have. Figure 4 and 5 shows the multiscaled representation provided by the wavelet transform, and Figure 6 and 7 shows the sparcity of the representation, where one can see that even with only 2500 coefficients, the reconstructed shapes preserve most of the features of the original shapes. (a) (b) (c) (d) (e) (f) Fig. 4. Figures (a)-(e) present a multiscaled representation of the hippocampus from courser approximation to finer approximation. Figure (f) is the original hippocampus.
6 (a) (b) (c) (d) (e) (f) Fig. 5. Figures (a)-(e) present a multiscaled representation of the cortex from courser approximation to finer approximation. Figure (f) is the original cortex. (a) (b) (c) (d) Fig. 6. Figures (a)-(c) are the reconstructed hippocampus using 1000, 2500 and 5000 coefficients, and the relative errors of them from the original shape are e-004, e-005 and.91508e-005 respectively. Figure (d) is the original hippocampus. (a) (b) (c) (d) Fig. 7. Figures (a)-(c) are the reconstructed cortices using 1000, 2500 and 5000 coefficients, and the relative errors of them from the original shape are e-004, e-004 and e-004 respectively. Figure (d) is the original cortex. One advantage of our method over spherical wavelet transform in analyzing biological shapes is that we have a much more flexible choice of wavelets. In particular, we can choose one wavelet with very high vanishing moments so that the representation is very sparse. Figure 8 below shows a comparison of our method to spherical wavelets as used in [9, 10].
7 (a) (b) Fig. 8. Figures (a) and (b) are the decay of relative 2 l error verse number of coefficients used, where the underline shapes are the hippocampus and cortex as shown in Figure 6 and 7. The multiscaled sparse representation provided by the wavelet transform has many applications. For example, one can do shape compression, or in other words, feature dimension reduction for shapes. One can also do shape denoising via thresholding or shrinkage of wavelet coefficients. Since these kinds of applications are not of our main interest, we shall not explore them in further details..2 Non-Parametric Tests For given two groups of hippocampus, one from healthy population, the other from the one with Alzheimer s disease, we apply Wilcoxon s rank sum test (see e.g. [7]) to the multiscaled curvature-like representation of shapes to find regions on the shape where the two groups are different. The α -value we choose in the results below is We note that the tests are more reliable in higher levels than those in lower levels. This is because the shape corresponding to a higher level is a smoothed version of the original shape, which means we have more statistical inference of the object. As one can see from below that the higher the level is, the larger each area of significance will be. In Figure 9 below, we used the mean hippocampus as the reference shape, which is calculated by simply taking an average of all the vector values of hippocampus at every position. (a) (b) Fig. 9. Figures (a) and (b) show regions of significance of level 5 and 4 respectively. Here, each pair of hippocampus is viewed from the bottom.
8 4 Discussion The wavelet-based shape representation technique proposed here allows one to study the geometry, topology and features of general biological shapes using any of the standard wavelet-bases on real-valued Euclidean spaces. The results we obtained are robust and consistent across individuals and populations. In addition to direct representation and shape characterization, this technique allows us to compute mean shapes and improve the shape-analysis statistical power by concentrating the energy of the shapecharacteristics in few significant wavelet coefficients, Dinov et. al. [5]. We are in the process of validating the new methodology using larger number of subjects, different types of applications (e.g., studying the population-specific differences in the proportion of gyri to sulcal area) and quantitative comparison with spherical-harmonics, spherical wavelets and tensor-based morphometry techniques. The computational complexity of the algorithm is ON ( log N ) relative to the volume size N. We have a Matlab implementation that we are in the process of converting to stand-alone C++ code, which will be distributed by the end of 2007 on the CCB Software Download pages ( Software). We tested the actual computation time of the wavelet decomposition and reconstruction on a PC with Inter(R) Core(TM) 2, 2.1 GHz and 1G physical memory. For a given shape with 65,56 surface points, the computation time is 2-20 seconds, depending on the choice of basis and level of decomposition. 5 Acknowledgement This work is funded by the National Institutes of Health through the NIH Roadmap for Medical Research, Grant U54 RR Information on the National Centers for Biomedical Computing can be obtained from References 1. F. Arandiga, R. Donat and A. Harten: Multiresolution based on weighted averages of the hat function I: Linear Reconstruction Techniques. UCLA CAM Reports., No , Aug S. Buss and J. Fillmore. Spherical averages and applications to spherical splines and interpolation. ACM Transactions on Graphics, 20(2), pp , D. L. Collins, P. Neelin, T. Peters, A. C. Evans, Automatic D intersubject registration of MR volumetric data in standardized Talairach space. J. Comput. Assist. Tomogr. 18 (2), I. Daubechies. Ten Lectures on Wavelets. CBMS-NSF Lecture Notes nr. 61, SIAM, I. D. Dinov, J. W. Boscardin, M. S. Mega, E. L. Sowell, A. W. Toga. A Wavelet-Based Statistical Analysis of fmri data: I. Motivation and Data Distribution Modeling, NeuroInformatics, Humana Press, (4), 19-4, A. C. Evans, D. L. Collins, P. Neelin, D. MacDonald, M. Kamber, T. S. Marrett. Three-dimensional correlative imaging: applications in human brain mapping. In: Thatcher, R.W., Hallett, M., Zeffiro, T., John, E.R., Huerta, M. (Eds.), Functional Neuroimaging: Technical Foundations. Academic Press, San Diego, pp , J. D. Gibbons. Nonparametric Statistical Inference, 2nd Ed., M. Dekker, X. Gu, Y. Wang, T. F. Chan, P.M. Thompson and S.-T. Yau. Genus Zero Surface Conformal Mapping and Its Application to Brain Surface Mapping. IEEE Transaction on Medical Imaging, 2(8), Aug. 2004, pp a) < 9. Nain D, Hacker S, Bobick A, Tannenbaum. A multiscale D shape analysis using spherical wavelets. Proc MICCAI, Oct ; p Nain D, Hacker S, Bobick A, Tannenbaum. A shape-driven D segmentation using spherical wavelets. Proc MICCAI, Oct 2-5, To Appear. 11. D. MacDonald, A method for identifying geometrically simple surfaces from three dimensional images. PhD thesis, McGill University, Mazziotta JC, Toga AW, Evans AC, Fox PT, Lancaster J, Zilles K, Woods RP, Paus T, Simpson G, Pike B, Holmes CJ, Collins DL, Thompson PM, MacDonald D, Schormann T, Amunts K, Palomero-Gallagher N, Parsons L, Narr KL, Kabani N (2001) A probabilistic atlas and reference system for the human brain: International Consortium for Brain Mapping. Philos Trans R Soc Lond B Biol Sci 56: E. Praun and H. Hoppe, Spherical parametrization and remeshing. ACM Trans. Gr. Vol. 22 (), July W. Sweldens. The lifting scheme: A construction of second generation wavelets. SIAM J. Math. Anal., Vol 29 (2), , 1997.
9 15. P. Schröder and W. Sweldens. Sphereical wavelets: Efficiently representing functions on the sphere. Computer Graphics Proceedings, (SIGGRAPH 95), Pages , D. W. Shattuck, S. R. Sandor-Leahy, K. A. Schaper, D. A. Rottenberg, R. M. Leahy. Magnetic resonance image tissue classification using a partial volume model. NeuroImage 1: , Y. Shi, P. M. Thompson, G. I. de Zubicaray, S. Rose, Z. Tu, I. Dinov, A. W. Toga, Direct mapping of hippocampal surfaces with intrinsic shape context, NeuroImage, 7(), pp , Y. Shi, P. M. Thompson, I. Dinov, S. Osher, A. W. Toga, Direct cortical mapping via solving partial differential equations on implicit surfaces, Medical Image Analysis, 11(), pp , J. G. Sled, A. P. Zijdenbos, A. C. Evans, A non-parametric method for automatic correction of intensity non-uniformity in MRI data. IEEE Trans. Med. Imag. 17, Paul M. Thompson, Arthur W. Toga. A Framework for Computational Anatomy. Springer Berlin / Heidelberg, vol 5, no. 1, Paul M. Thompson, Agatha D. Lee, Rebecca A. Dutton, Jennifer A. Geaga, Kiralee M. Hayashi, Mark A. Eckert, Ursula Bellugi, Albert M. Galaburda, Julie R. Korenberg, Debra L. Mills, Arthur W. Toga, and Allan L. Reiss. Abnormal Cortical Complexity and Thickness Profiles Mapped in Williams Syndrome. The Journal of Neuroscience, vol. 25 no. 16, Thompson PM, Hayashi KM, de Zubicaray G, Janke AL, Rose SE, Semple J, Herman D, Hong MS, Dittmer S, Doddrell DM, Toga AW. Dynamics of gray matter loss in Alzheimer's disease. J Neurosci 2: , Paul M. Thompson, Kiralee M. Hayashi, Greig I. de Zubicaray, Andrew L. Janke, Stephen E. Rose, James Semple, Michael S. Hong, David H. Herman, David Gravano, David M. Doddrell and Arthur W. Toga, Mapping hippocampal and ventricular change in Alzheimer disease, Neuroimage, vol 22, no. 4, , Z. Tu, S. Zheng, A. Yuille, A. Reiss, R. Dutton, A.Lee, A.Galaburda, I. Dinov, P. Thompson, A. Toga, Automated Extraction of the Cortical Sulci Based on a Supervised Learning Approach, IEEE Tran. on Medical Imaging, vol. 26, no. 4, April, 2007
Wavelet-Based Representation of Biological Shapes
 Wavelet-Based Representation of Biological Shapes Bin Dong 1,YuMao 2, Ivo D. Dinov 3,ZhuowenTu 3, Yonggang Shi 3, Yalin Wang 3, and Arthur W. Toga 3 1 Department of Mathematics, University of California,
Wavelet-Based Representation of Biological Shapes Bin Dong 1,YuMao 2, Ivo D. Dinov 3,ZhuowenTu 3, Yonggang Shi 3, Yalin Wang 3, and Arthur W. Toga 3 1 Department of Mathematics, University of California,
Multivariate Tensor-based Brain Anatomical Surface Morphometry via Holomorphic One-Forms
 Multivariate Tensor-based Brain Anatomical Surface Morphometry via Holomorphic One-Forms Yalin Wang 1,2, Tony F. Chan 2, Arthur W. Toga 1, and Paul M. Thompson 1 1 Lab. of Neuro Imaging, UCLA School of
Multivariate Tensor-based Brain Anatomical Surface Morphometry via Holomorphic One-Forms Yalin Wang 1,2, Tony F. Chan 2, Arthur W. Toga 1, and Paul M. Thompson 1 1 Lab. of Neuro Imaging, UCLA School of
Brain Surface Conformal Parameterization with Algebraic Functions
 Brain Surface Conformal Parameterization with Algebraic Functions Yalin Wang 1,2, Xianfeng Gu 3, Tony F. Chan 1, Paul M. Thompson 2, and Shing-Tung Yau 4 1 Mathematics Department, UCLA, Los Angeles, CA
Brain Surface Conformal Parameterization with Algebraic Functions Yalin Wang 1,2, Xianfeng Gu 3, Tony F. Chan 1, Paul M. Thompson 2, and Shing-Tung Yau 4 1 Mathematics Department, UCLA, Los Angeles, CA
A Learning Based Algorithm for Automatic Extraction of the Cortical Sulci
 A Learning Based Algorithm for Automatic Extraction of the Cortical Sulci Songfeng Zheng 1, Zhuowen Tu 2, Alan L. Yuille 1, Allan L. Reiss 3, Rebecca A. Dutton 2, Agatha D. Lee 2, Albert M. Galaburda 4,
A Learning Based Algorithm for Automatic Extraction of the Cortical Sulci Songfeng Zheng 1, Zhuowen Tu 2, Alan L. Yuille 1, Allan L. Reiss 3, Rebecca A. Dutton 2, Agatha D. Lee 2, Albert M. Galaburda 4,
Detecting Changes In Non-Isotropic Images
 Detecting Changes In Non-Isotropic Images K.J. Worsley 1, M. Andermann 1, T. Koulis 1, D. MacDonald, 2 and A.C. Evans 2 August 4, 1999 1 Department of Mathematics and Statistics, 2 Montreal Neurological
Detecting Changes In Non-Isotropic Images K.J. Worsley 1, M. Andermann 1, T. Koulis 1, D. MacDonald, 2 and A.C. Evans 2 August 4, 1999 1 Department of Mathematics and Statistics, 2 Montreal Neurological
Automatic Generation of Training Data for Brain Tissue Classification from MRI
 Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. COCOSCO, Alex P. ZIJDENBOS, and Alan C. EVANS http://www.bic.mni.mcgill.ca/users/crisco/ McConnell Brain Imaging
Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. COCOSCO, Alex P. ZIJDENBOS, and Alan C. EVANS http://www.bic.mni.mcgill.ca/users/crisco/ McConnell Brain Imaging
Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm
 Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm Lin Chen and Gudrun Wagenknecht Central Institute for Electronics, Research Center Jülich, Jülich, Germany Email: l.chen@fz-juelich.de
Topology Correction for Brain Atlas Segmentation using a Multiscale Algorithm Lin Chen and Gudrun Wagenknecht Central Institute for Electronics, Research Center Jülich, Jülich, Germany Email: l.chen@fz-juelich.de
Multi-scale Voxel-based Morphometry via Weighted Spherical Harmonic Representation
 Multi-scale Voxel-based Morphometry via Weighted Spherical Harmonic Representation Moo K. Chung 1,2, Li Shen 4, Kim M. Dalton 2, and Richard J. Davidson 2,3 1 Department of Statistics, Biostatistics and
Multi-scale Voxel-based Morphometry via Weighted Spherical Harmonic Representation Moo K. Chung 1,2, Li Shen 4, Kim M. Dalton 2, and Richard J. Davidson 2,3 1 Department of Statistics, Biostatistics and
Bayesian Spherical Wavelet Shrinkage: Applications to Shape Analysis
 Bayesian Spherical Wavelet Shrinkage: Applications to Shape Analysis Xavier Le Faucheur a, Brani Vidakovic b and Allen Tannenbaum a a School of Electrical and Computer Engineering, b Department of Biomedical
Bayesian Spherical Wavelet Shrinkage: Applications to Shape Analysis Xavier Le Faucheur a, Brani Vidakovic b and Allen Tannenbaum a a School of Electrical and Computer Engineering, b Department of Biomedical
Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge
 Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge Christian Wasserthal 1, Karin Engel 1, Karsten Rink 1 und André Brechmann
Automatic segmentation of the cortical grey and white matter in MRI using a Region Growing approach based on anatomical knowledge Christian Wasserthal 1, Karin Engel 1, Karsten Rink 1 und André Brechmann
The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy
 The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy Sokratis K. Makrogiannis, PhD From post-doctoral research at SBIA lab, Department of Radiology,
The Anatomical Equivalence Class Formulation and its Application to Shape-based Computational Neuroanatomy Sokratis K. Makrogiannis, PhD From post-doctoral research at SBIA lab, Department of Radiology,
Brain Warping Via Landmark Points and Curves with a Level Set Representation
 Brain Warping Via Landmark Points and Curves with a Level Set Representation Andrew Y. Wang, Alex D. Leow, 2 Hillary D. Protas, Arthur W. Toga, Paul M. Thompson UCLA Laboratory of Neuro Imaging, Los Angeles,
Brain Warping Via Landmark Points and Curves with a Level Set Representation Andrew Y. Wang, Alex D. Leow, 2 Hillary D. Protas, Arthur W. Toga, Paul M. Thompson UCLA Laboratory of Neuro Imaging, Los Angeles,
Automatic Subcortical Segmentation Using a Contextual Model
 Automatic Subcortical Segmentation Using a Contextual Model Jonathan H. Morra 1, Zhuowen Tu 1, Liana G. Apostolova 1,2, Amity E. Green 1,2, Arthur W. Toga 1, and Paul M. Thompson 1 1 Laboratory of Neuro
Automatic Subcortical Segmentation Using a Contextual Model Jonathan H. Morra 1, Zhuowen Tu 1, Liana G. Apostolova 1,2, Amity E. Green 1,2, Arthur W. Toga 1, and Paul M. Thompson 1 1 Laboratory of Neuro
Shape-based Diffeomorphic Registration on Hippocampal Surfaces Using Beltrami Holomorphic Flow
 Shape-based Diffeomorphic Registration on Hippocampal Surfaces Using Beltrami Holomorphic Flow Abstract. Finding meaningful 1-1 correspondences between hippocampal (HP) surfaces is an important but difficult
Shape-based Diffeomorphic Registration on Hippocampal Surfaces Using Beltrami Holomorphic Flow Abstract. Finding meaningful 1-1 correspondences between hippocampal (HP) surfaces is an important but difficult
Research Proposal: Computational Geometry with Applications on Medical Images
 Research Proposal: Computational Geometry with Applications on Medical Images MEI-HENG YUEH yueh@nctu.edu.tw National Chiao Tung University 1 Introduction My research mainly focuses on the issues of computational
Research Proposal: Computational Geometry with Applications on Medical Images MEI-HENG YUEH yueh@nctu.edu.tw National Chiao Tung University 1 Introduction My research mainly focuses on the issues of computational
IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 29, NO. 12, DECEMBER
 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 29, NO. 12, DECEMBER 2010 2009 Robust Surface Reconstruction via Laplace-Beltrami Eigen-Projection and Boundary Deformation Yonggang Shi, Member, IEEE, Rongjie
IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 29, NO. 12, DECEMBER 2010 2009 Robust Surface Reconstruction via Laplace-Beltrami Eigen-Projection and Boundary Deformation Yonggang Shi, Member, IEEE, Rongjie
Adaptive Bayesian Shrinkage Model for Spherical Wavelet Based Denoising and Compression of Hippocampus Shapes
 Adaptive Bayesian Shrinkage Model for Spherical Wavelet Based Denoising and Compression of Hippocampus Shapes Xavier Le Faucheur 1, Brani Vidakovic 2, Delphine Nain 1, and Allen Tannenbaum 1 1 School of
Adaptive Bayesian Shrinkage Model for Spherical Wavelet Based Denoising and Compression of Hippocampus Shapes Xavier Le Faucheur 1, Brani Vidakovic 2, Delphine Nain 1, and Allen Tannenbaum 1 1 School of
Automatic Generation of Training Data for Brain Tissue Classification from MRI
 MICCAI-2002 1 Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. Cocosco, Alex P. Zijdenbos, and Alan C. Evans McConnell Brain Imaging Centre, Montreal Neurological
MICCAI-2002 1 Automatic Generation of Training Data for Brain Tissue Classification from MRI Chris A. Cocosco, Alex P. Zijdenbos, and Alan C. Evans McConnell Brain Imaging Centre, Montreal Neurological
Methods for data preprocessing
 Methods for data preprocessing John Ashburner Wellcome Trust Centre for Neuroimaging, 12 Queen Square, London, UK. Overview Voxel-Based Morphometry Morphometry in general Volumetrics VBM preprocessing
Methods for data preprocessing John Ashburner Wellcome Trust Centre for Neuroimaging, 12 Queen Square, London, UK. Overview Voxel-Based Morphometry Morphometry in general Volumetrics VBM preprocessing
Labeling the Brain Surface Using a Deformable Multiresolution Mesh
 Labeling the Brain Surface Using a Deformable Multiresolution Mesh Sylvain Jaume 1, Benoît Macq 2, and Simon K. Warfield 3 1 Multiresolution Modeling Group, California Institute of Technology, 256-80,
Labeling the Brain Surface Using a Deformable Multiresolution Mesh Sylvain Jaume 1, Benoît Macq 2, and Simon K. Warfield 3 1 Multiresolution Modeling Group, California Institute of Technology, 256-80,
Measuring longitudinal brain changes in humans and small animal models. Christos Davatzikos
 Measuring longitudinal brain changes in humans and small animal models Christos Davatzikos Section of Biomedical Image Analysis University of Pennsylvania (Radiology) http://www.rad.upenn.edu/sbia Computational
Measuring longitudinal brain changes in humans and small animal models Christos Davatzikos Section of Biomedical Image Analysis University of Pennsylvania (Radiology) http://www.rad.upenn.edu/sbia Computational
Multiple Cortical Surface Correspondence Using Pairwise Shape Similarity
 Multiple Cortical Surface Correspondence Using Pairwise Shape Similarity Pahal Dalal 1, Feng Shi 2, Dinggang Shen 2, and Song Wang 1 1 Department of Computer Science and Engineering, University of South
Multiple Cortical Surface Correspondence Using Pairwise Shape Similarity Pahal Dalal 1, Feng Shi 2, Dinggang Shen 2, and Song Wang 1 1 Department of Computer Science and Engineering, University of South
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit
 Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
Optimization of Brain Conformal Mapping with Landmarks
 Optimization of Brain Conformal Mapping with Landmarks Yalin Wang 1,LokMingLui 1,TonyF.Chan 1, and Paul M. Thompson 2 Mathematics Department, UCLA, Los Angeles, CA 90095, USA Lab. of Neuro Imaging, UCLA
Optimization of Brain Conformal Mapping with Landmarks Yalin Wang 1,LokMingLui 1,TonyF.Chan 1, and Paul M. Thompson 2 Mathematics Department, UCLA, Los Angeles, CA 90095, USA Lab. of Neuro Imaging, UCLA
Multi-atlas labeling with population-specific template and non-local patch-based label fusion
 Multi-atlas labeling with population-specific template and non-local patch-based label fusion Vladimir Fonov, Pierrick Coupé, Simon Eskildsen, Jose Manjon, Louis Collins To cite this version: Vladimir
Multi-atlas labeling with population-specific template and non-local patch-based label fusion Vladimir Fonov, Pierrick Coupé, Simon Eskildsen, Jose Manjon, Louis Collins To cite this version: Vladimir
Shape Analysis with Conformal Invariants for Multiply Connected Domains and its Application to Analyzing Brain Morphology
 Shape Analysis with Conformal Invariants for Multiply Connected Domains and its Application to Analyzing Brain Morphology Yalin Wang Dept of Neurology/Math UCLA ylwang@math.ucla.edu Xianfeng Gu Dept of
Shape Analysis with Conformal Invariants for Multiply Connected Domains and its Application to Analyzing Brain Morphology Yalin Wang Dept of Neurology/Math UCLA ylwang@math.ucla.edu Xianfeng Gu Dept of
COMBINATIONOF BRAIN CONFORMAL MAPPING AND LANDMARKS: A VARIATIONALAPPROACH
 COMBINATIONOF BRAIN CONFORMAL MAPPING AND LANDMARKS: A VARIATIONALAPPROACH Yalin Wang Mathematics Department, UCLA email: ylwang@math.ucla.edu Lok Ming Lui Mathematics Department, UCLA email: malmlui@math.ucla.edu
COMBINATIONOF BRAIN CONFORMAL MAPPING AND LANDMARKS: A VARIATIONALAPPROACH Yalin Wang Mathematics Department, UCLA email: ylwang@math.ucla.edu Lok Ming Lui Mathematics Department, UCLA email: malmlui@math.ucla.edu
UNC 4D Infant Cortical Surface Atlases, from Neonate to 6 Years of Age
 UNC 4D Infant Cortical Surface Atlases, from Neonate to 6 Years of Age Version 1.0 UNC 4D infant cortical surface atlases from neonate to 6 years of age contain 11 time points, including 1, 3, 6, 9, 12,
UNC 4D Infant Cortical Surface Atlases, from Neonate to 6 Years of Age Version 1.0 UNC 4D infant cortical surface atlases from neonate to 6 years of age contain 11 time points, including 1, 3, 6, 9, 12,
UCLA Advanced Neuroimaging Summer Program David W. Shattuck, PhD
 BrainSuite UCLA Advanced Neuroimaging Summer Program Presented 18 July 2013 David W. Shattuck, PhD Associate Professor Department of Neurology David Geffen School of Medicine at UCLA http://www.loni.ucla.edu/~shattuck/
BrainSuite UCLA Advanced Neuroimaging Summer Program Presented 18 July 2013 David W. Shattuck, PhD Associate Professor Department of Neurology David Geffen School of Medicine at UCLA http://www.loni.ucla.edu/~shattuck/
NIH Public Access Author Manuscript Conf Proc IEEE Eng Med Biol Soc. Author manuscript; available in PMC 2009 December 11.
 NIH Public Access Author Manuscript Published in final edited form as: Conf Proc IEEE Eng Med Biol Soc. 2004 ; 2: 1521 1524. doi:10.1109/iembs.2004.1403466. An Elliptic PDE Approach for Shape Characterization
NIH Public Access Author Manuscript Published in final edited form as: Conf Proc IEEE Eng Med Biol Soc. 2004 ; 2: 1521 1524. doi:10.1109/iembs.2004.1403466. An Elliptic PDE Approach for Shape Characterization
VOLUMETRIC HARMONIC MAP
 COMMUNICATIONS IN INFORMATION AND SYSTEMS c 2004 International Press Vol. 3, No. 3, pp. 191-202, March 2004 004 VOLUMETRIC HARMONIC MAP YALIN WANG, XIANFENG GU, AND SHING-TUNG YAU Abstract. We develop
COMMUNICATIONS IN INFORMATION AND SYSTEMS c 2004 International Press Vol. 3, No. 3, pp. 191-202, March 2004 004 VOLUMETRIC HARMONIC MAP YALIN WANG, XIANFENG GU, AND SHING-TUNG YAU Abstract. We develop
Comparison Study of Clinical 3D MRI Brain Segmentation Evaluation
 Comparison Study of Clinical 3D MRI Brain Segmentation Evaluation Ting Song 1, Elsa D. Angelini 2, Brett D. Mensh 3, Andrew Laine 1 1 Heffner Biomedical Imaging Laboratory Department of Biomedical Engineering,
Comparison Study of Clinical 3D MRI Brain Segmentation Evaluation Ting Song 1, Elsa D. Angelini 2, Brett D. Mensh 3, Andrew Laine 1 1 Heffner Biomedical Imaging Laboratory Department of Biomedical Engineering,
Neuroimaging and mathematical modelling Lesson 2: Voxel Based Morphometry
 Neuroimaging and mathematical modelling Lesson 2: Voxel Based Morphometry Nivedita Agarwal, MD Nivedita.agarwal@apss.tn.it Nivedita.agarwal@unitn.it Volume and surface morphometry Brain volume White matter
Neuroimaging and mathematical modelling Lesson 2: Voxel Based Morphometry Nivedita Agarwal, MD Nivedita.agarwal@apss.tn.it Nivedita.agarwal@unitn.it Volume and surface morphometry Brain volume White matter
Chapter 21 Structural MRI: Morphometry
 Chapter 21 Structural MRI: Morphometry Christian Gaser Abstract Human brains are characterised by considerable intersubject anatomical variability, which is of interest in both clinical practice and research.
Chapter 21 Structural MRI: Morphometry Christian Gaser Abstract Human brains are characterised by considerable intersubject anatomical variability, which is of interest in both clinical practice and research.
Morphological Analysis of Brain Structures Using Spatial Normalization
 Morphological Analysis of Brain Structures Using Spatial Normalization C. Davatzikos 1, M. Vaillant 1, S. Resnick 2, J.L. Prince 3;1, S. Letovsky 1, and R.N. Bryan 1 1 Department of Radiology, Johns Hopkins
Morphological Analysis of Brain Structures Using Spatial Normalization C. Davatzikos 1, M. Vaillant 1, S. Resnick 2, J.L. Prince 3;1, S. Letovsky 1, and R.N. Bryan 1 1 Department of Radiology, Johns Hopkins
CHAPTER 2. Morphometry on rodent brains. A.E.H. Scheenstra J. Dijkstra L. van der Weerd
 CHAPTER 2 Morphometry on rodent brains A.E.H. Scheenstra J. Dijkstra L. van der Weerd This chapter was adapted from: Volumetry and other quantitative measurements to assess the rodent brain, In vivo NMR
CHAPTER 2 Morphometry on rodent brains A.E.H. Scheenstra J. Dijkstra L. van der Weerd This chapter was adapted from: Volumetry and other quantitative measurements to assess the rodent brain, In vivo NMR
RESEARCH STATEMENT Ronald Lok Ming Lui
 Ronald Lok Ming Lui 1/11 RESEARCH STATEMENT Ronald Lok Ming Lui (malmlui@math.harvard.edu) The main focus of my research has been on problems in computational conformal geometry and human brain mapping.
Ronald Lok Ming Lui 1/11 RESEARCH STATEMENT Ronald Lok Ming Lui (malmlui@math.harvard.edu) The main focus of my research has been on problems in computational conformal geometry and human brain mapping.
Bayesian Spherical Wavelet Shrinkage: Applications to Shape Analysis
 Bayesian Spherical Wavelet Shrinkage: Applications to Shape Analysis Xavier Le Faucheur a, Brani Vidakovic b and Allen Tannenbaum a a School of Electrical and Computer Engineering, b Department of Biomedical
Bayesian Spherical Wavelet Shrinkage: Applications to Shape Analysis Xavier Le Faucheur a, Brani Vidakovic b and Allen Tannenbaum a a School of Electrical and Computer Engineering, b Department of Biomedical
Brain Surface Conformal Spherical Mapping
 Brain Surface Conformal Spherical Mapping Min Zhang Department of Industrial Engineering, Arizona State University mzhang33@asu.edu Abstract It is well known and proved that any genus zero surface can
Brain Surface Conformal Spherical Mapping Min Zhang Department of Industrial Engineering, Arizona State University mzhang33@asu.edu Abstract It is well known and proved that any genus zero surface can
SURFACE-BASED modeling is valuable in brain imaging to
 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 26, NO. 6, JUNE 2007 853 Brain Surface Conformal Parameterization Using Riemann Surface Structure Yalin Wang*, Member, IEEE, Lok Ming Lui, Xianfeng Gu, Kiralee
IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 26, NO. 6, JUNE 2007 853 Brain Surface Conformal Parameterization Using Riemann Surface Structure Yalin Wang*, Member, IEEE, Lok Ming Lui, Xianfeng Gu, Kiralee
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit
 Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
Knowledge-Based Segmentation of Brain MRI Scans Using the Insight Toolkit John Melonakos 1, Ramsey Al-Hakim 1, James Fallon 2 and Allen Tannenbaum 1 1 Georgia Institute of Technology, Atlanta GA 30332,
Joint Sulci Detection Using Graphical Models and Boosted Priors
 Joint Sulci Detection Using Graphical Models and Boosted Priors Yonggang Shi 1,ZhuowenTu 1, Allan L. Reiss 2, Rebecca A. Dutton 1, Agatha D. Lee 1, Albert M. Galaburda 3,IvoDinov 1, Paul M. Thompson 1,
Joint Sulci Detection Using Graphical Models and Boosted Priors Yonggang Shi 1,ZhuowenTu 1, Allan L. Reiss 2, Rebecca A. Dutton 1, Agatha D. Lee 1, Albert M. Galaburda 3,IvoDinov 1, Paul M. Thompson 1,
The organization of the human cerebral cortex estimated by intrinsic functional connectivity
 1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
1 The organization of the human cerebral cortex estimated by intrinsic functional connectivity Journal: Journal of Neurophysiology Author: B. T. Thomas Yeo, et al Link: https://www.ncbi.nlm.nih.gov/pubmed/21653723
A Fast and Accurate Method for Automated Cortical Surface Registration and Labeling
 A Fast and Accurate Method for Automated Cortical Surface Registration and Labeling Anand A Joshi 1, David W Shattuck 2 and Richard M Leahy 1 1 Signal and Image Processing Institute, University of Southern
A Fast and Accurate Method for Automated Cortical Surface Registration and Labeling Anand A Joshi 1, David W Shattuck 2 and Richard M Leahy 1 1 Signal and Image Processing Institute, University of Southern
Synthetic MRI Signal Standardization: Application to Multi-atlas Analysis
 Synthetic MRI Signal Standardization: Application to Multi-atlas Analysis Juan Eugenio Iglesias 1,IvoDinov 2, Jaskaran Singh 2, Gregory Tong 2, and Zhuowen Tu 2 1 Medical Imaging Informatics, University
Synthetic MRI Signal Standardization: Application to Multi-atlas Analysis Juan Eugenio Iglesias 1,IvoDinov 2, Jaskaran Singh 2, Gregory Tong 2, and Zhuowen Tu 2 1 Medical Imaging Informatics, University
Open Topology: A Toolkit for Brain Isosurface Correction
 Open Topology: A Toolkit for Brain Isosurface Correction Sylvain Jaume 1, Patrice Rondao 2, and Benoît Macq 2 1 National Institute of Research in Computer Science and Control, INRIA, France, sylvain@mit.edu,
Open Topology: A Toolkit for Brain Isosurface Correction Sylvain Jaume 1, Patrice Rondao 2, and Benoît Macq 2 1 National Institute of Research in Computer Science and Control, INRIA, France, sylvain@mit.edu,
STATISTICAL ATLAS-BASED SUB-VOXEL SEGMENTATION OF 3D BRAIN MRI
 STATISTICA ATAS-BASED SUB-VOXE SEGMENTATION OF 3D BRAIN MRI Marcel Bosc 1,2, Fabrice Heitz 1, Jean-Paul Armspach 2 (1) SIIT UMR-7005 CNRS / Strasbourg I University, 67400 Illkirch, France (2) IPB UMR-7004
STATISTICA ATAS-BASED SUB-VOXE SEGMENTATION OF 3D BRAIN MRI Marcel Bosc 1,2, Fabrice Heitz 1, Jean-Paul Armspach 2 (1) SIIT UMR-7005 CNRS / Strasbourg I University, 67400 Illkirch, France (2) IPB UMR-7004
Multiresolution Meshes. COS 526 Tom Funkhouser, Fall 2016 Slides by Guskov, Praun, Sweldens, etc.
 Multiresolution Meshes COS 526 Tom Funkhouser, Fall 2016 Slides by Guskov, Praun, Sweldens, etc. Motivation Huge meshes are difficult to render store transmit edit Multiresolution Meshes! [Guskov et al.]
Multiresolution Meshes COS 526 Tom Funkhouser, Fall 2016 Slides by Guskov, Praun, Sweldens, etc. Motivation Huge meshes are difficult to render store transmit edit Multiresolution Meshes! [Guskov et al.]
Conformal Slit Mapping and Its Applications to Brain Surface Parameterization
 Conformal Slit Mapping and Its Applications to Brain Surface Parameterization Yalin Wang 1,2,XianfengGu 3,TonyF.Chan 2,PaulM.Thompson 1, and Shing-Tung Yau 4 1 Lab. of Neuro Imaging, UCLA School of Medicine,
Conformal Slit Mapping and Its Applications to Brain Surface Parameterization Yalin Wang 1,2,XianfengGu 3,TonyF.Chan 2,PaulM.Thompson 1, and Shing-Tung Yau 4 1 Lab. of Neuro Imaging, UCLA School of Medicine,
An Intensity Consistent Approach to the Cross Sectional Analysis of Deformation Tensor Derived Maps of Brain Shape
 An Intensity Consistent Approach to the Cross Sectional Analysis of Deformation Tensor Derived Maps of Brain Shape C. Studholme, V. Cardenas, A. Maudsley, and M. Weiner U.C.S.F., Dept of Radiology, VAMC
An Intensity Consistent Approach to the Cross Sectional Analysis of Deformation Tensor Derived Maps of Brain Shape C. Studholme, V. Cardenas, A. Maudsley, and M. Weiner U.C.S.F., Dept of Radiology, VAMC
Cortical surface thickness as a classifier: Boosting for autism classification
 Cortical surface thickness as a classifier: Boosting for autism classification Vikas Singh 1, Lopamudra Mukherjee 2, and Moo K. Chung 1 1 Biostatistics and Medical Informatics, University of Wisconsin-Madison,
Cortical surface thickness as a classifier: Boosting for autism classification Vikas Singh 1, Lopamudra Mukherjee 2, and Moo K. Chung 1 1 Biostatistics and Medical Informatics, University of Wisconsin-Madison,
Simultaneous Cortical Surface Labeling and Sulcal Curve Extraction
 Simultaneous Cortical Surface Labeling and Sulcal Curve Extraction Zhen Yang a, Aaron Carass a, Chen Chen a, Jerry L Prince a,b a Electrical and Computer Engineering, b Biomedical Engineering, Johns Hopkins
Simultaneous Cortical Surface Labeling and Sulcal Curve Extraction Zhen Yang a, Aaron Carass a, Chen Chen a, Jerry L Prince a,b a Electrical and Computer Engineering, b Biomedical Engineering, Johns Hopkins
Surface-based Analysis: Inter-subject Registration and Smoothing
 Surface-based Analysis: Inter-subject Registration and Smoothing Outline Exploratory Spatial Analysis Coordinate Systems 3D (Volumetric) 2D (Surface-based) Inter-subject registration Volume-based Surface-based
Surface-based Analysis: Inter-subject Registration and Smoothing Outline Exploratory Spatial Analysis Coordinate Systems 3D (Volumetric) 2D (Surface-based) Inter-subject registration Volume-based Surface-based
Human Brain Mapping with Conformal Geometry and Multivariate Tensor-based Morphometry
 Human Brain Mapping with Conformal Geometry and Multivariate Tensor-based Morphometry Jie Shi 1, Paul M. Thompson 2, Yalin Wang 1 1 School of Computing, Informatics and Decision Systems Engineering, ASU,
Human Brain Mapping with Conformal Geometry and Multivariate Tensor-based Morphometry Jie Shi 1, Paul M. Thompson 2, Yalin Wang 1 1 School of Computing, Informatics and Decision Systems Engineering, ASU,
DEVELOPMENT OF REALISTIC HEAD MODELS FOR ELECTRO- MAGNETIC SOURCE IMAGING OF THE HUMAN BRAIN
 DEVELOPMENT OF REALISTIC HEAD MODELS FOR ELECTRO- MAGNETIC SOURCE IMAGING OF THE HUMAN BRAIN Z. "! #$! Acar, N.G. Gençer Department of Electrical and Electronics Engineering, Middle East Technical University,
DEVELOPMENT OF REALISTIC HEAD MODELS FOR ELECTRO- MAGNETIC SOURCE IMAGING OF THE HUMAN BRAIN Z. "! #$! Acar, N.G. Gençer Department of Electrical and Electronics Engineering, Middle East Technical University,
Automated Surface Matching using Mutual Information Applied to Riemann Surface Structures
 Automated Surface Matching using Mutual Information Applied to Riemann Surface Structures Yalin Wang 1, Ming-Chang Chiang 2, and Paul M. Thompson 2 1 Mathematics Department, UCLA, Los Angeles, CA 90095,
Automated Surface Matching using Mutual Information Applied to Riemann Surface Structures Yalin Wang 1, Ming-Chang Chiang 2, and Paul M. Thompson 2 1 Mathematics Department, UCLA, Los Angeles, CA 90095,
Applications of Elastic Functional and Shape Data Analysis
 Applications of Elastic Functional and Shape Data Analysis Quick Outline: 1. Functional Data 2. Shapes of Curves 3. Shapes of Surfaces BAYESIAN REGISTRATION MODEL Bayesian Model + Riemannian Geometry +
Applications of Elastic Functional and Shape Data Analysis Quick Outline: 1. Functional Data 2. Shapes of Curves 3. Shapes of Surfaces BAYESIAN REGISTRATION MODEL Bayesian Model + Riemannian Geometry +
Performance Evaluation of the TINA Medical Image Segmentation Algorithm on Brainweb Simulated Images
 Tina Memo No. 2008-003 Internal Memo Performance Evaluation of the TINA Medical Image Segmentation Algorithm on Brainweb Simulated Images P. A. Bromiley Last updated 20 / 12 / 2007 Imaging Science and
Tina Memo No. 2008-003 Internal Memo Performance Evaluation of the TINA Medical Image Segmentation Algorithm on Brainweb Simulated Images P. A. Bromiley Last updated 20 / 12 / 2007 Imaging Science and
CORTICAL sulci are important structures of the brain
 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 26, NO. 4, APRIL 2007 541 Automated Extraction of the Cortical Sulci Based on a Supervised Learning Approach Zhuowen Tu, Songfeng Zheng, Alan L. Yuille, Allan
IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 26, NO. 4, APRIL 2007 541 Automated Extraction of the Cortical Sulci Based on a Supervised Learning Approach Zhuowen Tu, Songfeng Zheng, Alan L. Yuille, Allan
NIH Public Access Author Manuscript Med Image Comput Comput Assist Interv. Author manuscript; available in PMC 2013 May 07.
 NIH Public Access Author Manuscript Published in final edited form as: Med Image Comput Comput Assist Interv. 2005 ; 8(0 2): 459 467. Multiscale 3D Shape Analysis using Spherical Wavelets Delphine Nain
NIH Public Access Author Manuscript Published in final edited form as: Med Image Comput Comput Assist Interv. 2005 ; 8(0 2): 459 467. Multiscale 3D Shape Analysis using Spherical Wavelets Delphine Nain
Fast Interactive Region of Interest Selection for Volume Visualization
 Fast Interactive Region of Interest Selection for Volume Visualization Dominik Sibbing and Leif Kobbelt Lehrstuhl für Informatik 8, RWTH Aachen, 20 Aachen Email: {sibbing,kobbelt}@informatik.rwth-aachen.de
Fast Interactive Region of Interest Selection for Volume Visualization Dominik Sibbing and Leif Kobbelt Lehrstuhl für Informatik 8, RWTH Aachen, 20 Aachen Email: {sibbing,kobbelt}@informatik.rwth-aachen.de
Computational QC Geometry: A tool for Medical Morphometry, Computer Graphics & Vision
 Computational QC Geometry: A tool for Medical Morphometry, Computer Graphics & Vision Part II of the sequel of 2 talks. Computation C/QC geometry was presented by Tony F. Chan Ronald Lok Ming Lui Department
Computational QC Geometry: A tool for Medical Morphometry, Computer Graphics & Vision Part II of the sequel of 2 talks. Computation C/QC geometry was presented by Tony F. Chan Ronald Lok Ming Lui Department
Matching 3D Shapes Using 2D Conformal Representations
 Matching 3D Shapes Using 2D Conformal Representations Xianfeng Gu 1 and Baba C. Vemuri 2 Computer and Information Science and Engineering, Gainesville, FL 32611-6120, USA {gu,vemuri}@cise.ufl.edu Abstract.
Matching 3D Shapes Using 2D Conformal Representations Xianfeng Gu 1 and Baba C. Vemuri 2 Computer and Information Science and Engineering, Gainesville, FL 32611-6120, USA {gu,vemuri}@cise.ufl.edu Abstract.
Parameterization of 3D Brain Structures for Statistical Shape Analysis
 Parameterization of 3D Brain Structures for Statistical Shape Analysis Litao Zhu and Tianzi Jiang * National Laboratory of Pattern Recognition, Institute of Automation, Chinese Academy of Sciences, Beijing
Parameterization of 3D Brain Structures for Statistical Shape Analysis Litao Zhu and Tianzi Jiang * National Laboratory of Pattern Recognition, Institute of Automation, Chinese Academy of Sciences, Beijing
Automated Graph-Based Analysis and Correction of Cortical Volume Topology
 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 20, NO. 11, NOVEMBER 2001 1167 Automated Graph-Based Analysis and Correction of Cortical Volume Topology David W. Shattuck* and Richard M. Leahy Abstract The
IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 20, NO. 11, NOVEMBER 2001 1167 Automated Graph-Based Analysis and Correction of Cortical Volume Topology David W. Shattuck* and Richard M. Leahy Abstract The
Towards Learning-Based Holistic Brain Image Segmentation
 Towards Learning-Based Holistic Brain Image Segmentation Zhuowen Tu Lab of Neuro Imaging University of California, Los Angeles in collaboration with (A. Toga, P. Thompson et al.) Supported by (NIH CCB
Towards Learning-Based Holistic Brain Image Segmentation Zhuowen Tu Lab of Neuro Imaging University of California, Los Angeles in collaboration with (A. Toga, P. Thompson et al.) Supported by (NIH CCB
Comets v. 1.0 Matlab toolbox. User Manual. Written by Young-Jin Jung
 Comets v. 1.0 Matlab toolbox User Manual Written by Young-Jin Jung (cometstool@gmail.com) Comets A Matlab toolbox for simulating electric field due to tdcs Version 1.0, Feb. 1, 2012 Downloadable at http://www.cometstool.com
Comets v. 1.0 Matlab toolbox User Manual Written by Young-Jin Jung (cometstool@gmail.com) Comets A Matlab toolbox for simulating electric field due to tdcs Version 1.0, Feb. 1, 2012 Downloadable at http://www.cometstool.com
Norbert Schuff VA Medical Center and UCSF
 Norbert Schuff Medical Center and UCSF Norbert.schuff@ucsf.edu Medical Imaging Informatics N.Schuff Course # 170.03 Slide 1/67 Objective Learn the principle segmentation techniques Understand the role
Norbert Schuff Medical Center and UCSF Norbert.schuff@ucsf.edu Medical Imaging Informatics N.Schuff Course # 170.03 Slide 1/67 Objective Learn the principle segmentation techniques Understand the role
Neuroimage Processing
 Neuroimage Processing Instructor: Moo K. Chung mkchung@wisc.edu Lecture 2-3. General Linear Models (GLM) Voxel-based Morphometry (VBM) September 11, 2009 What is GLM The general linear model (GLM) is a
Neuroimage Processing Instructor: Moo K. Chung mkchung@wisc.edu Lecture 2-3. General Linear Models (GLM) Voxel-based Morphometry (VBM) September 11, 2009 What is GLM The general linear model (GLM) is a
Quantitative MRI of the Brain: Investigation of Cerebral Gray and White Matter Diseases
 Quantities Measured by MR - Quantitative MRI of the Brain: Investigation of Cerebral Gray and White Matter Diseases Static parameters (influenced by molecular environment): T, T* (transverse relaxation)
Quantities Measured by MR - Quantitative MRI of the Brain: Investigation of Cerebral Gray and White Matter Diseases Static parameters (influenced by molecular environment): T, T* (transverse relaxation)
RECENT developments in brain imaging have accelerated
 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 23, NO. 8, AUGUST 2004 949 Genus Zero Surface Conformal Mapping and Its Application to Brain Surface Mapping Xianfeng Gu, Yalin Wang*, Tony F. Chan, Paul M. Thompson,
IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 23, NO. 8, AUGUST 2004 949 Genus Zero Surface Conformal Mapping and Its Application to Brain Surface Mapping Xianfeng Gu, Yalin Wang*, Tony F. Chan, Paul M. Thompson,
Transformation Model and Constraints Cause Bias in Statistics on Deformation Fields
 Transformation Model and Constraints Cause Bias in Statistics on Deformation Fields Torsten Rohlfing Neuroscience Program, SRI International, Menlo Park, CA, USA torsten@synapse.sri.com http://www.stanford.edu/
Transformation Model and Constraints Cause Bias in Statistics on Deformation Fields Torsten Rohlfing Neuroscience Program, SRI International, Menlo Park, CA, USA torsten@synapse.sri.com http://www.stanford.edu/
Automated MR Image Analysis Pipelines
 Automated MR Image Analysis Pipelines Andy Simmons Centre for Neuroimaging Sciences, Kings College London Institute of Psychiatry. NIHR Biomedical Research Centre for Mental Health at IoP & SLAM. Neuroimaging
Automated MR Image Analysis Pipelines Andy Simmons Centre for Neuroimaging Sciences, Kings College London Institute of Psychiatry. NIHR Biomedical Research Centre for Mental Health at IoP & SLAM. Neuroimaging
arxiv: v1 [cs.cv] 6 Jun 2017
![arxiv: v1 [cs.cv] 6 Jun 2017 arxiv: v1 [cs.cv] 6 Jun 2017](/thumbs/96/128307097.jpg) Volume Calculation of CT lung Lesions based on Halton Low-discrepancy Sequences Liansheng Wang a, Shusheng Li a, and Shuo Li b a Department of Computer Science, Xiamen University, Xiamen, China b Dept.
Volume Calculation of CT lung Lesions based on Halton Low-discrepancy Sequences Liansheng Wang a, Shusheng Li a, and Shuo Li b a Department of Computer Science, Xiamen University, Xiamen, China b Dept.
Automatic MS Lesion Segmentation by Outlier Detection and Information Theoretic Region Partitioning Release 0.00
 Automatic MS Lesion Segmentation by Outlier Detection and Information Theoretic Region Partitioning Release 0.00 Marcel Prastawa 1 and Guido Gerig 1 Abstract July 17, 2008 1 Scientific Computing and Imaging
Automatic MS Lesion Segmentation by Outlier Detection and Information Theoretic Region Partitioning Release 0.00 Marcel Prastawa 1 and Guido Gerig 1 Abstract July 17, 2008 1 Scientific Computing and Imaging
Parameterization of Triangular Meshes with Virtual Boundaries
 Parameterization of Triangular Meshes with Virtual Boundaries Yunjin Lee 1;Λ Hyoung Seok Kim 2;y Seungyong Lee 1;z 1 Department of Computer Science and Engineering Pohang University of Science and Technology
Parameterization of Triangular Meshes with Virtual Boundaries Yunjin Lee 1;Λ Hyoung Seok Kim 2;y Seungyong Lee 1;z 1 Department of Computer Science and Engineering Pohang University of Science and Technology
CS 229 Final Project Report Learning to Decode Cognitive States of Rat using Functional Magnetic Resonance Imaging Time Series
 CS 229 Final Project Report Learning to Decode Cognitive States of Rat using Functional Magnetic Resonance Imaging Time Series Jingyuan Chen //Department of Electrical Engineering, cjy2010@stanford.edu//
CS 229 Final Project Report Learning to Decode Cognitive States of Rat using Functional Magnetic Resonance Imaging Time Series Jingyuan Chen //Department of Electrical Engineering, cjy2010@stanford.edu//
A Parameterization-based Numerical Method for Isotropic and Anisotropic Diffusion Smoothing on Non-Flat Surfaces
 1 A Parameterization-based Numerical Method for Isotropic and Anisotropic Diffusion Smoothing on Non-Flat Surfaces Anand A. Joshi, David W. Shattuck, Paul M. Thompson and Richard M. Leahy Abstract Neuroimaging
1 A Parameterization-based Numerical Method for Isotropic and Anisotropic Diffusion Smoothing on Non-Flat Surfaces Anand A. Joshi, David W. Shattuck, Paul M. Thompson and Richard M. Leahy Abstract Neuroimaging
3D Surface Matching and Registration through Shape Images
 3D Surface Matching and Registration through Shape Images Zhaoqiang Lai and Jing Hua Department of Computer Science, Wayne State University, Detroit, MI 48202, USA Abstract. In this paper, we present a
3D Surface Matching and Registration through Shape Images Zhaoqiang Lai and Jing Hua Department of Computer Science, Wayne State University, Detroit, MI 48202, USA Abstract. In this paper, we present a
IMA Preprint Series # 2078
 A GEOMETRIC METHOD FOR AUTOMATIC EXTRACTION OF SULCAL FUNDI By C.-Y. Kao M. Hofer G. Sapiro J. Stern and D. Rottenberg IMA Preprint Series # 2078 ( November 2005 ) INSTITUTE FOR MATHEMATICS AND ITS APPLICATIONS
A GEOMETRIC METHOD FOR AUTOMATIC EXTRACTION OF SULCAL FUNDI By C.-Y. Kao M. Hofer G. Sapiro J. Stern and D. Rottenberg IMA Preprint Series # 2078 ( November 2005 ) INSTITUTE FOR MATHEMATICS AND ITS APPLICATIONS
Preprocessing II: Between Subjects John Ashburner
 Preprocessing II: Between Subjects John Ashburner Pre-processing Overview Statistics or whatever fmri time-series Anatomical MRI Template Smoothed Estimate Spatial Norm Motion Correct Smooth Coregister
Preprocessing II: Between Subjects John Ashburner Pre-processing Overview Statistics or whatever fmri time-series Anatomical MRI Template Smoothed Estimate Spatial Norm Motion Correct Smooth Coregister
Discriminative, Semantic Segmentation of Brain Tissue in MR Images
 Discriminative, Semantic Segmentation of Brain Tissue in MR Images Zhao Yi 1, Antonio Criminisi 2, Jamie Shotton 2, and Andrew Blake 2 1 University of California, Los Angeles, USA. zyi@ucla.edu. 2 Microsoft
Discriminative, Semantic Segmentation of Brain Tissue in MR Images Zhao Yi 1, Antonio Criminisi 2, Jamie Shotton 2, and Andrew Blake 2 1 University of California, Los Angeles, USA. zyi@ucla.edu. 2 Microsoft
Using a Statistical Shape Model to Extract Sulcal Curves on the Outer Cortex of the Human Brain
 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 21, NO. 5, MAY 2002 513 Using a Statistical Shape Model to Extract Sulcal Curves on the Outer Cortex of the Human Brain Xiaodong Tao, Jerry L. Prince, and Christos
IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 21, NO. 5, MAY 2002 513 Using a Statistical Shape Model to Extract Sulcal Curves on the Outer Cortex of the Human Brain Xiaodong Tao, Jerry L. Prince, and Christos
Topology Preserving Brain Tissue Segmentation Using Graph Cuts
 Topology Preserving Brain Tissue Segmentation Using Graph Cuts Xinyang Liu 1, Pierre-Louis Bazin 2, Aaron Carass 3, and Jerry Prince 3 1 Brigham and Women s Hospital, Boston, MA 1 xinyang@bwh.harvard.edu
Topology Preserving Brain Tissue Segmentation Using Graph Cuts Xinyang Liu 1, Pierre-Louis Bazin 2, Aaron Carass 3, and Jerry Prince 3 1 Brigham and Women s Hospital, Boston, MA 1 xinyang@bwh.harvard.edu
BrainSuite. presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014
 BrainSuite presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014 David Shattuck Ahmanson-Lovelace Brain Mapping Center Department of Neurology David Geffen School of Medicine at
BrainSuite presented at the UCLA/NITP Advanced Neuroimaging Summer Program 29 July 2014 David Shattuck Ahmanson-Lovelace Brain Mapping Center Department of Neurology David Geffen School of Medicine at
Adaptive Fuzzy Watermarking for 3D Models
 International Conference on Computational Intelligence and Multimedia Applications 2007 Adaptive Fuzzy Watermarking for 3D Models Mukesh Motwani.*, Nikhil Beke +, Abhijit Bhoite +, Pushkar Apte +, Frederick
International Conference on Computational Intelligence and Multimedia Applications 2007 Adaptive Fuzzy Watermarking for 3D Models Mukesh Motwani.*, Nikhil Beke +, Abhijit Bhoite +, Pushkar Apte +, Frederick
Learning-based Neuroimage Registration
 Learning-based Neuroimage Registration Leonid Teverovskiy and Yanxi Liu 1 October 2004 CMU-CALD-04-108, CMU-RI-TR-04-59 School of Computer Science Carnegie Mellon University Pittsburgh, PA 15213 Abstract
Learning-based Neuroimage Registration Leonid Teverovskiy and Yanxi Liu 1 October 2004 CMU-CALD-04-108, CMU-RI-TR-04-59 School of Computer Science Carnegie Mellon University Pittsburgh, PA 15213 Abstract
A Binary Entropy Measure to Assess Nonrigid Registration Algorithms
 A Binary Entropy Measure to Assess Nonrigid Registration Algorithms Simon K. Warfield 1, Jan Rexilius 1, Petra S. Huppi 2, Terrie E. Inder 3, Erik G. Miller 1, William M. Wells III 1, Gary P. Zientara
A Binary Entropy Measure to Assess Nonrigid Registration Algorithms Simon K. Warfield 1, Jan Rexilius 1, Petra S. Huppi 2, Terrie E. Inder 3, Erik G. Miller 1, William M. Wells III 1, Gary P. Zientara
Simultaneous surface and volumetric registration using harmonic maps
 Simultaneous surface and volumetric registration using harmonic maps Anand A. Joshi a, David W. Shattuck b, Paul M. Thompson b and Richard M. Leahy a a University of Southern California, Los Angeles, CA
Simultaneous surface and volumetric registration using harmonic maps Anand A. Joshi a, David W. Shattuck b, Paul M. Thompson b and Richard M. Leahy a a University of Southern California, Los Angeles, CA
3D Brain Segmentation Using Active Appearance Models and Local Regressors
 3D Brain Segmentation Using Active Appearance Models and Local Regressors K.O. Babalola, T.F. Cootes, C.J. Twining, V. Petrovic, and C.J. Taylor Division of Imaging Science and Biomedical Engineering,
3D Brain Segmentation Using Active Appearance Models and Local Regressors K.O. Babalola, T.F. Cootes, C.J. Twining, V. Petrovic, and C.J. Taylor Division of Imaging Science and Biomedical Engineering,
SEGMENTING subcortical structures from 3-D brain images
 IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 27, NO. 4, APRIL 2008 495 Brain Anatomical Structure Segmentation by Hybrid Discriminative/Generative Models Zhuowen Tu, Katherine L. Narr, Piotr Dollár, Ivo
IEEE TRANSACTIONS ON MEDICAL IMAGING, VOL. 27, NO. 4, APRIL 2008 495 Brain Anatomical Structure Segmentation by Hybrid Discriminative/Generative Models Zhuowen Tu, Katherine L. Narr, Piotr Dollár, Ivo
Matching 3D Shapes Using 2D Conformal Representations
 Matching 3D Shapes Using 2D Conformal Representations Xianfeng Gu 1 and Baba C. Vemuri 2 Computer and Information Science and Engineering, Gainesville, FL 32611-6120, USA {gu,vemuri}@cise.ufl.edu Matching
Matching 3D Shapes Using 2D Conformal Representations Xianfeng Gu 1 and Baba C. Vemuri 2 Computer and Information Science and Engineering, Gainesville, FL 32611-6120, USA {gu,vemuri}@cise.ufl.edu Matching
Progressive Geometry Compression. Andrei Khodakovsky Peter Schröder Wim Sweldens
 Progressive Geometry Compression Andrei Khodakovsky Peter Schröder Wim Sweldens Motivation Large (up to billions of vertices), finely detailed, arbitrary topology surfaces Difficult manageability of such
Progressive Geometry Compression Andrei Khodakovsky Peter Schröder Wim Sweldens Motivation Large (up to billions of vertices), finely detailed, arbitrary topology surfaces Difficult manageability of such
Form follows func-on. Which one of them can fly? And why? Why would anyone bother about brain structure, when we can do func5onal imaging?
 Why would anyone bother about brain structure, when we can do func5onal imaging? Form follows func-on Which one of them can fly? And why? h;p://animals.na5onalgeographic.com Why would anyone bother about
Why would anyone bother about brain structure, when we can do func5onal imaging? Form follows func-on Which one of them can fly? And why? h;p://animals.na5onalgeographic.com Why would anyone bother about
Structural MRI analysis
 Structural MRI analysis volumetry and voxel-based morphometry cortical thickness measurements structural covariance network mapping Boris Bernhardt, PhD Department of Social Neuroscience, MPI-CBS bernhardt@cbs.mpg.de
Structural MRI analysis volumetry and voxel-based morphometry cortical thickness measurements structural covariance network mapping Boris Bernhardt, PhD Department of Social Neuroscience, MPI-CBS bernhardt@cbs.mpg.de
Fmri Spatial Processing
 Educational Course: Fmri Spatial Processing Ray Razlighi Jun. 8, 2014 Spatial Processing Spatial Re-alignment Geometric distortion correction Spatial Normalization Smoothing Why, When, How, Which Why is
Educational Course: Fmri Spatial Processing Ray Razlighi Jun. 8, 2014 Spatial Processing Spatial Re-alignment Geometric distortion correction Spatial Normalization Smoothing Why, When, How, Which Why is
G 2 Interpolation for Polar Surfaces
 1 G 2 Interpolation for Polar Surfaces Jianzhong Wang 1, Fuhua Cheng 2,3 1 University of Kentucky, jwangf@uky.edu 2 University of Kentucky, cheng@cs.uky.edu 3 National Tsinhua University ABSTRACT In this
1 G 2 Interpolation for Polar Surfaces Jianzhong Wang 1, Fuhua Cheng 2,3 1 University of Kentucky, jwangf@uky.edu 2 University of Kentucky, cheng@cs.uky.edu 3 National Tsinhua University ABSTRACT In this
Multiscale 3D Shape Analysis Using Spherical Wavelets
 Multiscale 3D Shape Analysis Using Spherical Wavelets Delphine Nain 1,StevenHaker 2, Aaron Bobick 1, and Allen R. Tannenbaum 3 1 College of Computing, Georgia Institute of Technology, Atlanta, GA 30332-0280
Multiscale 3D Shape Analysis Using Spherical Wavelets Delphine Nain 1,StevenHaker 2, Aaron Bobick 1, and Allen R. Tannenbaum 3 1 College of Computing, Georgia Institute of Technology, Atlanta, GA 30332-0280
Automatic Segmentation of Neonatal Brain MRI
 Automatic Segmentation of Neonatal Brain MRI 1 Marcel Prastawa, 2 John Gilmore, 3 Weili Lin, and 1,2 Guido Gerig Department of 1 Computer Science, 2 Psychiatry, and 3 Radiology University of North Carolina,
Automatic Segmentation of Neonatal Brain MRI 1 Marcel Prastawa, 2 John Gilmore, 3 Weili Lin, and 1,2 Guido Gerig Department of 1 Computer Science, 2 Psychiatry, and 3 Radiology University of North Carolina,
Manifold Parameterization
 Manifold Parameterization Lei Zhang 1,2, Ligang Liu 1,2, Zhongping Ji 1,2, and Guojin Wang 1,2 1 Department of Mathematics, Zhejiang University, Hangzhou 310027, China 2 State Key Lab of CAD&CG, Zhejiang
Manifold Parameterization Lei Zhang 1,2, Ligang Liu 1,2, Zhongping Ji 1,2, and Guojin Wang 1,2 1 Department of Mathematics, Zhejiang University, Hangzhou 310027, China 2 State Key Lab of CAD&CG, Zhejiang
